ABSTRACT
VP8 is a major tegument protein of bovine herpesvirus 1 (BoHV-1) and is essential for viral replication in cattle. The protein undergoes phosphorylation after transcription through cellular casein kinase 2 (CK2) and a viral kinase, US3. In this study, a virus containing a mutated VP8 protein that is not phosphorylated by CK2 and US3 (BoHV-1-YmVP8) was constructed by homologous recombination in mammalian cells. When BoHV-1-YmVP8-infected cells were observed by transmission electron microscopy, blocking phosphorylation of VP8 was found to impair viral DNA encapsidation, resulting in release of incomplete viral particles to the extracellular environment. Consequently, less infectious virus was produced by the mutant virus than by wild-type (WT) virus. A comparison of mutant and WT VP8 by confocal microscopy revealed that mutant VP8 is nuclear throughout infection while WT VP8 is nuclear early during infection and is associated with the Golgi apparatus at later stages. This, together with the observation that mutant VP8 is present in virions, albeit in smaller amounts, suggests that the incorporation of VP8 may occur at two stages. The first takes place without the need for phosphorylation and before or during nuclear egress of capsids, whereas the second occurs in the Golgi apparatus and requires phosphorylation of VP8. The results indicate that phosphorylated VP8 plays a role in viral DNA encapsidation and in the secondary virion incorporation of VP8. To perform these functions, the cellular localization of VP8 is adjusted based on the phosphorylation status.
IMPORTANCE In this study, phosphorylation of VP8 was shown to have a function in BoHV-1 replication. A virus containing a mutated VP8 protein that is not phosphorylated by CK2 and US3 (BoHV-1-YmVP8) produced smaller numbers of infectious virions than wild-type (WT) virus. The maturation and egress of WT and mutant BoHV-1 were studied, showing a process similar to that reported for other alphaherpesviruses. Interestingly, lack of phosphorylation of VP8 by CK2 and US3 resulted in reduced incorporation of viral DNA into capsids during mutant BoHV-1 infection, as well as lower numbers of extracellular virions. Furthermore, mutant VP8 remained nuclear throughout infection, in contrast to WT VP8, which is nuclear at early stages and Golgi apparatus associated late during infection. This correlates with smaller amounts of mutant VP8 in virions and suggests for the first time that VP8 may be assembled into the virions at two stages, with the latter dependent on phosphorylation.
INTRODUCTION
Bovine herpesvirus 1 (BoHV-1), a member of the Alphaherpesviridae, causes infectious bovine rhinotracheitis (IBR) in cattle, as well as conjunctivitis, vulvovaginitis, and balanoposthitis. The herpesvirus particle is composed of a capsid containing the double-stranded DNA genome, which is surrounded by a tegument layer and an envelope containing viral glycoproteins (1). During the herpesvirus life cycle, genome synthesis, capsid assembly, DNA incorporation (2), and primary tegumentation (3) occur in the nuclei of infected cells. Subsequently, nucleocapsids leave the nucleus by budding through the nuclear membrane via an envelopment–de-envelopment pathway (4, 5). Capsids acquire tegument proteins, envelope proteins, and glycoproteins in the cytoplasm and finally egress via the cellular secretory pathway (6). During the maturation process, viral components are sequentially incorporated into viral particles in a strictly controlled manner.
Tegument proteins have various functions, including regulation of transcription (4), kinase functions (5), and virus assembly (7). The BoHV-1 unique long 47 (UL47) gene product (6), VP8, is a 97-kDa tegument protein that is indispensable for viral replication in host animals (8). While VP8 is the most abundant viral protein (9), the mechanism of incorporation of VP8 into virions is unknown. The cellular localization of VP8 changes at different stages of infection, indicating varying roles of the protein during the virus life cycle. Navigated by an N-terminal nuclear localization signal (NLS) (10, 11), virion-derived VP8 targets to the nucleus immediately upon BoHV-1 infection (9, 12). At late stages of infection, VP8 accumulates in the cytoplasm (9), but no cytoplasmic function has been identified for VP8. In transfected cells, VP8 is mainly restricted to the nucleus, and the nuclear VP8 appears to remodel the promyelocytic leukemia nuclear body (PML-NB) (13), which is a suppressor of human herpesvirus 1 (HHV-1) replication (14). VP8 also has the capacity to interact with intronless mRNAs of BoHV-1 (15).
In BoHV-1-infected cells, VP8 is highly expressed and extensively phosphorylated. Phosphorylation is an important posttranslational modification for VP8 to maintain PML-remodeling capacity; however, it does not directly affect the nuclear localization of VP8 (13). This is in contrast to other proteins, where phosphorylation changes their cellular localization. For example, VP22 of HHV-1 has the capacity to perform nuclear-cytoplasmic shuttling during infection, and the nonphosphorylated form localizes to the cytoplasm while the phosphorylated form localizes to the nucleus (16, 17). VP8 is mainly phosphorylated by a viral protein, US3, and a cellular protein, casein kinase 2 (CK2), through distinct target residues (13, 18). By site-specific mutagenesis of BoHV-1 VP8, serine 16 was identified as essential for phosphorylation by US3, and 7 residues (threonine 65, serine 66, serine 79, serine 80, serine 82, serine 88, and threonine 107) were critical for phosphorylation by CK2 (13).
Phosphorylation on BoHV-1 proteins has been reported to correlate with viral replication and protein incorporation. Blocking tyrosine phosphorylation of glycoprotein E (gE), an envelope protein, reduces viral replication (19). Removing tyrosine phosphorylation of VP22 decreases the amount of VP22 incorporated into virions (20). However, how phosphorylation benefits viral replication and determines protein incorporation is not clear. The presence of VP8 in the cytoplasm and the nucleus indicates that the protein has different functions, corresponding to the cellular localization. These functions might be related to the extensive phosphorylation of VP8, mediated by serine/threonine kinases CK2 and US3. This is also supported by a previous result showing that US3 associates with nuclear VP8 but not with cytoplasmic VP8 (18).
To study the role of VP8 phosphorylation in BoHV-1 replication, we constructed a recombinant BoHV-1 containing a mutant VP8 that is not phosphorylated by CK2 or US3 (BoHV-1-YmVP8). This mutant virus showed reduced DNA encapsidation and virus production. Furthermore, VP8 was found to be nuclear in early stages and Golgi apparatus associated late during infection, while mutant VP8 remained nuclear throughout infection, which indicates that the localization of VP8 to the Golgi apparatus requires phosphorylation. Furthermore, as mutant VP8 was incorporated into virions, but in smaller amounts, this suggests that VP8 may be incorporated into virons at two stages during BoHV-1 infection.
MATERIALS AND METHODS
Cells and virus.
Madin-Darby bovine kidney (MDBK) cells and primary fetal bovine testis (FBT) cells were cultured in Eagle's minimum essential medium (MEM) (Gibco, Life Technologies, Burlington, ON, Canada) supplemented with 10% fetal bovine serum (FBS) (Gibco). Production of all viral stocks was carried out in MDBK cells as previously described (9). Briefly, virus infections were accomplished by rocking 150-cm2 85 to 90% confluent cell monolayers with BoHV-1 in 10 ml MEM at 37°C, which was replaced after 1 h with 10 ml MEM supplemented with 2% FBS, followed by further incubation at 37°C. The virus titer was determined by plaque titration in 24-well plates overlaid with 8% low-melting-point agarose in MEM (8, 9).
Antibodies.
Monoclonal antibodies specific for gB, gC, and gD (21) and polyclonal antibodies specific for VP8 (9), VP22 (22), VP5 (8), and US3 (22) were raised as described previously. A monoclonal anti-Golgi apparatus 58,000-molecular-weight (58K) protein antibody was purchased from Sigma-Aldrich (St. Louis, MO, USA). IRDye 680RD goat anti-rabbit IgG and IRDye 800CW goat anti-mouse IgG were purchased from Li-Cor Biosciences (Lincoln, NE, USA). Alexa 488-conjugated goat anti-mouse IgG and Alexa 633-conjugated goat anti-rabbit IgG were purchased from Life Technologies.
Construction of recombinant viruses.
Recombinant viruses were constructed by homologous recombination between WT strain 108 (9) viral DNA and recombinant PCR products within FBT cells. UL47 was mutated (mUL47) to remove all essential phosphorylation sites within VP8 by replacing serine 16, threonine 65, serine 66, serine 79, serine 80, serine 82, serine 88, and threonine 107 with alanine (13). Briefly, plasmids with yellow fluorescent protein (YFP) fused in frame with both UL47 and mUL47, together with flanking regions of homology, were constructed and used to generate DNA fragments (Fig. 1A). A DNA fragment encoding YFP was PCR amplified from pEYFP-N1 using primers 5′-CGAGCTCAAGCTTCGAATTCTGCAGTC-3′ and 5′-GAAAGATCTCGCTTGTACAGCTCGTCCATGCC-3′, cut with HindIII and BglII, and cloned into pFLAG-VP8 and pFLAG-Mut-VP8 (13), similarly cut, giving pYFP-UL47 and pYFP-mUL47. The recognition sequences for restriction enzymes in the primers are underlined. The 5′ homology region was amplified from BoHV-1 DNA by PCR using primers 5′-GAAGGTACCGCTGGCCTTTGCGCATATGTACG-3′ and 5′-GAAACCGGTGCGTCTAAGGGCGCCTAGAA-3′ and inserted into pYFP-UL47 and pYFP-mUL47 after digestion with KpnI and AgeI, resulting in pUL48-YFP-UL47 and pUL48-YFP-mUL47. For transfection, DNA fragments were generated by PCR using the primers 5′-CGTGTTCGTTTCGCTGTACTATGC-3′ and 5′-CAGTAAATCAGGGAGCCCATTGAG-3′; these fragments contained about 950 bp of homologous DNA for recombination on either side of the YFP and mutagenized UL47.
FIG 1.
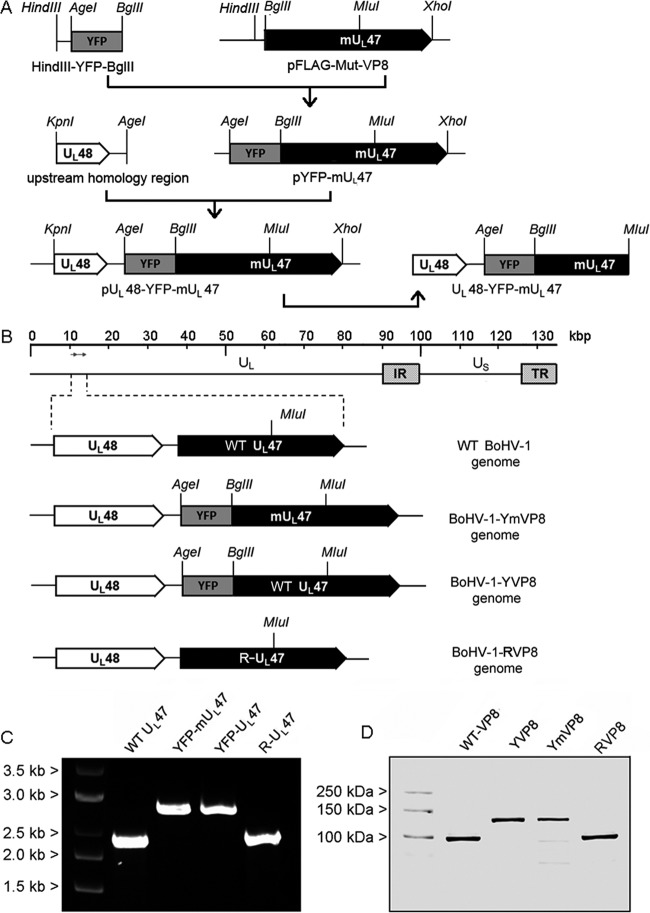
Schematic representations of DNA constructs and the genomes of recombinant viruses. (A) Development of DNA fragments for homologous recombination. The locations of restriction sites are indicated on each construct. (B) Genomes of recombinant viruses. The BoHV-1 genome consists of a unique long (UL) region and a unique short (US) region bracketed by inverted-repeat sequences (IR and TR). The UL48 and UL47 ORFs are highlighted. (C) PCR analysis of the viral genomes. Purified genomic DNA of WT BoHV-1, BoHV-1-YVP8, BoHV-1-YmVP8, and BoHV-1-RVP8 were used as PCR templates. PCR products covering from about 1 kbp of upstream flanking sequence of UL47 to the MluI restriction site of the UL47 gene were amplified using primers 5′-CGTGTTCGTTTCGCTGTACTATGC-3′ and 5′-CAGTAAATCAGGGAGCCCATTGAG-3′. The PCR products were separated in a 1% agarose gel with a DNA marker. (D) Western blot of VP8 proteins. MDBK cells were infected with WT BoHV-1, BoHV-1-YVP8, BoHV-1-YmVP8, or BoHV-1-RVP8 at an MOI of 1 for 20 h. Whole-protein extracts from cell lysates were analyzed by Western blotting using polyclonal anti-VP8 antibody and IRDye 600RD-conjugated secondary antibody. Molecular mass markers are shown on the left.
To introduce the YFP-UL47 mutant into the viral genome (Fig. 1B), 2 μg of PCR fragment was cotransfected with 1 μg of wild-type WT BoHV-1 (WT BoHV-1) genomic DNA into FBT cells with Lipofectamine and Plus reagent (Life Technologies). YFP-positive plaques were plaque purified several times to eliminate contamination from wild-type virus. The resulting virus (BoHV-1-YmVP8) encompasses an YFP-tagged mutant VP8. The same procedure was performed to introduce YFP-UL47 into the viral genome, resulting in a virus (BoHV-1-YVP8) expressing YFP-tagged WT-VP8.
To produce a revertant virus with WT UL47, homologous DNA fragments containing 921 bp of DNA upstream of the UL47 gene and 1,308 bp from the starting codon of the UL47 open reading frame (ORF) were PCR amplified from the WT BoHV-1 genome by using the primers 5′-CGTGTTCGTTTCGCTGTACTATGC-3′ and 5′-CAGTAAATCAGGGAGCCCATTGAG-3′. Two micrograms of this DNA was cotransfected with 1 μg of genomic DNA of BoHV-1-YmVP8 into FBT cells as described above. YFP-negative plaques were plaque purified several times to eliminate contamination from mutant virus. In the revertant virus (BoHV-1-RVP8) genome, the WT UL47 ORF was repaired.
The PCR, restriction digestions, and ligations were carried out with Q5 Hot Start High-Fidelity DNA Polymerase (NEB, Ipswich, MA, USA), restriction enzymes (NEB), and T4 DNA ligase (NEB). The DNA sequencing was performed by the NRC-Plant Biotechnology Institute (Saskatoon, SK, Canada).
Immunoprecipitation.
MDBK cells were pretreated with phosphate-free Dulbecco's modified Eagle's medium (DMEM) (Life Technologies) for 3 h prior to virus infection and subsequent incubation with [32P]orthophosphate (PerkinElmer, Woodbridge, ON, Canada). Cell lysates were precleared with protein G-Sepharose (GE Healthcare, Burnaby, BC, Canada) and then incubated with anti-VP8 polyclonal antibody and protein G-Sepharose overnight at 4°C. The protein G-Sepharose was washed three times with wash buffer (50 mM Tris-HCl, 150 mM NaCl, pH 7.4) and boiled for 5 min with SDS-PAGE sample buffer. The samples were separated in SDS-PAGE gels. The gels were subsequently dried and exposed to Imaging Screen K for visualization on a Molecular Imager FX (Bio-Rad, Mississauga, ON, Canada). ProtoBlue Safe (National Diagnostics, Atlanta, GA, USA) was used to stain the gels.
Western blotting.
Whole-cell extracts were prepared by suspension of cell pellets in radioimmunoprecipitation assay (RIPA) buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1% NP-40, 1% deoxycholate, 0.1% SDS) supplemented with protease inhibitor cocktail (Sigma-Aldrich) and phosphatase inhibitor cocktail (EMD Millipore). The cell lysates, clarified by centrifugation at 13,000 × g for 10 min at 4°C, were boiled in SDS-PAGE sample buffer for 5 min. After separation by SDS-PAGE, the proteins were transferred to nitrocellulose membranes and incubated with the appropriate antibodies. The membranes were washed and incubated with IRDye 600RD/800CW-conjugated secondary antibodies at a 1:20,000 dilution and scanned with an Odyssey CLx infrared imaging system (Li-Cor Biosciences).
Densitometry was used to quantify the proteins in the SDS-PAGE gels and Western blots. Capsid protein VP5 was used to normalize the amount of input viruses. The relative quantity (RQ) of each protein was calculated by dividing the density of the viral protein of interest (Dint) by the density of VP5 (DVP5) in the same lane. The relative difference of the protein of interest (RD) was used to illustrate the changes in quantity of that protein between the recombinant viruses (sample) and wild-type virus (WT). These calculations are presented in the following equations: RD = (RDsample/RQWT) × 100% and RQ = Dint/DVPS.
Confocal microscopy.
MDBK cells, cultured on Permanox 2-well chamber slides (Thermo Fisher Scientific, Waltham, MA, USA), were infected with viruses or mock infected. At the indicated time points, the cells were washed three times with phosphate-buffered saline (PBS) (136.9 mM NaCl, 2.7 mM KCl, 7.0 mM Na3PO4, and 0.9 mM Na3PO4, pH 7.4) and fixed with 4% paraformaldehyde for 20 min, followed by three washes with PBS. Subsequently, the cells were permeabilized with 0.1% Triton X-100 in PBS for 20 min, washed with PBS, and then incubated with 1% normal goat serum (Gibco) in PBS for 30 min at room temperature. The cells were incubated with primary antibodies at the appropriate dilutions for 2 h at room temperature, followed by washing with PBS and incubation with Alexa Fluor-conjugated antibodies (Life Technologies) at a 1:500 dilution for 1 h at room temperature. Finally, the slides were mounted with ProLong Gold Antifade Mountant with DAPI (4′,6-diamidino-2-phenylindole) (Life Technologies) prior to examination with a Leica SP5 confocal microscope (Leica Microsystems CMS GmbH, Mannheim, Germany) equipped with an external argon ion 488-/633-/461-nm laser.
Transmission electron microscopy (TEM).
MDBK cells in T75 flasks were infected with viruses at a multiplicity of infection (MOI) of 1. At 15 h postinfection (hpi), the cells were harvested with trypsin, washed in cold PBS, and pelleted at 500 × g for 10 min at 4°C. The cell pellets were fixed with 2.5% glutaraldehyde in PBS for 4 h and postfixed with 1% osmium tetraoxide for 4 h. After washing with PBS for 30 min, the fixed samples were dehydrated in graded concentrated ethanol (50, 70, 90, and 100%) and polymerized with propylene oxide for 1 h. Subsequently, the pellets were embedded in Epon 812, followed by polymerization for 3 days at 60°C. Ultrathin sections with a thickness of 50 to 70 nm prepared by a Reichert-Jung Ultracut E Ultramicrotome (Reichert-Jung, Vienna, Austria) were mounted on 200-mesh carbon-coated grids and poststained with 2% uranyl acetate for 10 min and 1% lead citrate for 40 min. After washing with water and air drying, the specimens were observed with a Philips CM10 transmission electron microscope (Philips Electron Optics, Eindhoven, Netherlands).
Virus purification.
MDBK cells cultured in T150 flasks were infected with BoHV-1, BoHV-1-YVP8, or BoHV-1-YmVP8 at an MOI of 1. The medium was harvested when over 90% of the cells showed cytopathic effect and centrifuged at 3,000 × g for 30 min at 4°C to remove cell debris. The viruses were pelleted by centrifugation at 25,000 rpm for 2 h at 4°C in a Beckman Coulter SW 32 Ti Rotor (Beckman Coulter Inc., Atlanta, GA, USA). The virus pellets were resuspended in a small volume of TNE buffer (50 mM Tris-HCl, pH 7.4, 100 mM NaCl, and 0.1 mM EDTA) overnight. The virus suspensions were loaded on top of a 10 to 60% potassium sodium tartrate gradient in TNE buffer and centrifuged at 25,000 rpm for 2 h at 4°C in a Beckman Coulter SW 41 Ti rotor.
Statistical analysis.
Data were analyzed using Microsoft Excel 2010. Standard deviations were calculated based on the entire population of each group and are shown as error bars. A two-tailed t test was used to determine the statistical differences between two groups. Differences were considered statistically significant at P values of >0.01 and ≤0.05 and statistically highly significant at P values of ≤0.01.
RESULTS
Construction of recombinant BoHV-1.
To study the impact of phosphorylation of VP8 during BoHV-1 infection, we constructed a virus expressing mutant VP8 (Mut-VP8) in which all essential phosphorylation sites for CK2 and US3 were removed by point mutations. YFP N-terminally fused to VP8 and Mut-VP8 served as a selection marker. Integration of DNA was confirmed by PCR (Fig. 1C) and DNA sequencing. The PCR fragments were sequenced and found to be identical to the WT sequence of UL47 in the case of YFP-UL-47 and revertant UL47 (R-UL47) and to the mutated sequence in the case of YFP-mUL47. VP8 proteins from infected cell lysates were analyzed by Western blotting. YVP8 (YFP-VP8) and YmVP8 (YFP-mutant VP8) are about 27.5 kDa larger than WT VP8 and RVP8 (revertant VP8), which correlates with the apparent molecular mass of YFP (Fig. 1D).
Blocking phosphorylation of VP8 impairs the production of BoHV-1.
The phosphorylation status of WT VP8, YVP8, YmVP8, and RVP8 in infected cells was investigated by immunoprecipitation. [32P]orthophosphate-labeled VP8 proteins were purified from infected MDBK cells, and equal amounts were loaded onto SDS-PAGE gels (Fig. 2A). WT VP8 was phosphorylated in the infected cells, and YVP8 had a phosphorylation level similar to that of WT VP8. However, YmVP8 showed a very substantial reduction in phosphorylation. Phosphorylation was restored in RVP8 to a level similar to those of WT VP8 and YVP8.
FIG 2.
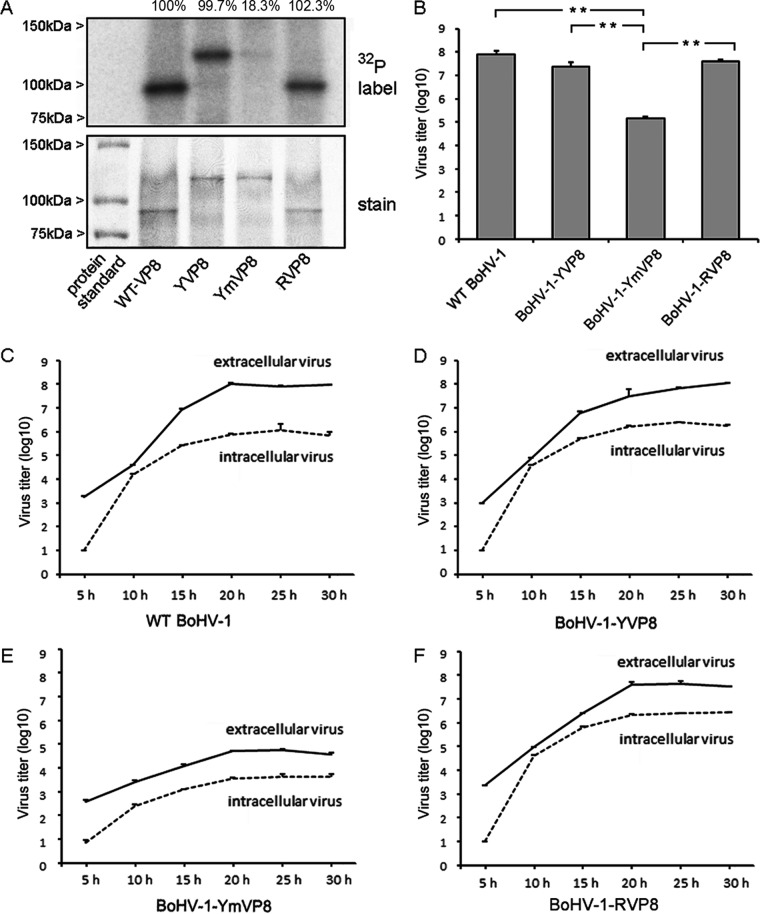
Phosphorylation status of VP8 and growth characteristics of viruses in MDBK cells. (A) Analysis of VP8 proteins by immunoprecipitation. MDBK cells were infected with WT BoHV-1, BoHV-1-YVP8, BoHV-1-YmVP8, and BoHV-1-RVP8 at an MOI of 1 and labeled with [32P]orthophosphate. Cell lysates were collected at 20 hpi and used for VP8 purification by incubation with anti-VP8 polyclonal antibody and protein G Sepharose. (Top) The samples were separated by SDS-PAGE and exposed to Imaging Screen K. (Bottom) The gels were stained with ProtoBlue Safe to indicate the amount of protein loading. The relative difference of each sample from WT VP8 is shown as a percentage at the top. (B) Titration of viruses. MDBK cells were infected with viruses at an MOI of 1. The supernatant and cells were collected at 24 hpi. Viruses from infected cells and supernatants were quantified by plaque titration on MDBK cell monolayers. The data were analyzed by two-tailed t test. The statistical significance of the difference between the values is shown; **, P ≤ 0.01. (C to F) Single-step growth curve of viruses. MDBK cells were infected with viruses at an MOI of 1, and the cell culture medium and cells were harvested separately at the indicated time points. The titer of infectious virus progeny in each sample was determined by plaque assay on MDBK cells. The error bars indicate standard deviations.
Production of WT BoHV-1, BoHV-1-YVP8, BoHV-1-YmVP8, and BoHV-1-RVP8 was analyzed by titrating the viruses from the combined supernatants and cells (Fig. 2B). The titer of BoHV-1-YmVP8 was significantly lower than that of WT BoHV-1. The titers were restored to WT levels by repairing VP8 in BoHV-1-RVP8, and the BoHV-1-YVP8 titer was equal to that of WT BoHV-1. To verify the intracellular and extracellular production of the mutant virus, viruses from the cells and supernatants were collected separately every 5 h (up to 30 hpi) and subjected to one-step growth analysis (Fig. 2C to F). The intracellular WT BoHV-1 titer rapidly increased from 5 hpi (10 PFU/ml) to 15 hpi (105.4 PFU/ml), indicating a major replication stage during this time period. Subsequently, a moderate increase in titer was observed from 15 hpi (105.4 PFU/ml) to 25 hpi (106.0 PFU/ml), followed by a slight decrease during the next 5 h (from 106.0 to 105.8 PFU/ml). Concomitantly, the extracellular virus titer increased substantially between 10 hpi (104.6 PFU/ml) and 20 hpi (108.0 PFU/ml) (Fig. 2C). This suggests that major egress takes place from 10 to 20 hpi. BoHV-1-YVP8 had a growth pattern similar to that of WT BoHV-1 (Fig. 2D). In contrast, the intracellular BoHV-1-YmVP8 titer increased constantly and slowly from 5 hpi (100.8 PFU/ml) to 30 hpi (103.6 PFU/ml), and the extracellular virus titer increased concurrently from 5 hpi (102.6 PFU/ml) to 30 hpi (104.6 PFU/ml) (Fig. 2E). Repair of the phosphorylation residues on VP8 brought the virus growth curve to a pattern similar to that observed for WT BoHV-1 (Fig. 2F).
Phosphorylation affects the incorporation and cellular localization of VP8 during infection.
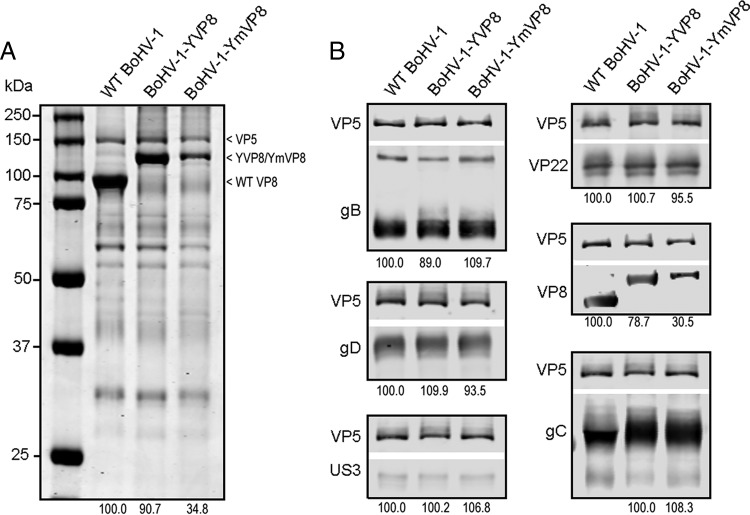
Deletion of VP8 impairs the incorporation of gB, gD, VP22, and especially gC into BoHV-1 (8). To determine whether phosphorylation of VP8 plays a role in incorporation of major glycoproteins or tegument proteins, whole-protein extracts from purified virions were analyzed by SDS-PAGE (Fig. 3A) and Western blotting (Fig. 3B). After densitometry, the normalized values for the BoHV-1-YVP8 and BoHV-1-YmVP8 proteins were compared to the corresponding values for the WT virus. The overall profiles of the three viruses were similar, with the exception that the incorporation of YmVP8 was lower than that of WT VP8 and YVP8 and that the molecular masses of YVP8 and YmVP8 were about 27.5 kDa higher (Fig. 3A) than that of VP8. The packaging of VP8 into BoHV-1-YVP8 and BoHV-1-YmVP8 virions was reduced by 10% and 65%, respectively. There were no obvious changes in glycoprotein (gB, gC, and gD) and tegument protein (VP22 and US3) amounts when analyzed by Western blotting. The value of gC from BoHV-1-YmVP8 was compared to that of BoHV-1-YVP8, because gC (91 kDa) was partially masked by VP8 (97 kDa) in the WT BoHV-1 lane. The densities of YVP8 and YmVP8 were 21% and 70% lower than that of WT VP8 (Fig. 3B), respectively, indicating that the reduced packaging of VP8 may be due to a block in phosphorylation.
FIG 3.
Influence of blocking VP8 phosphorylation on virion composition. (A) Solubilized proteins of purified extracellular virions of BoHV-1, BoHV-1-YVP8, and BoHV-1-YmVP8 were analyzed by SDS-PAGE, and the proteins were stained with ProtoBlue Safe. The densities of VP5 and VP8 were analyzed by densitometry. The percentages of YVP8 and YmVP8 in comparison to WT VP8 are indicated below the samples. (B) Western blotting of viral proteins. Solubilized proteins of purified extracellular WT BoHV-1, BoHV-1-YVP8, and BoHV-1-YmVP8 virions were analyzed by Western blotting. The proteins were detected with polyclonal anti-VP5, -VP22, -VP8, and -US3 antibodies and monoclonal anti-gB, -gC, and -gD antibodies, followed by IRDye 680RD goat anti-rabbit IgG and IRDye 800CW goat anti-mouse IgG, respectively. Each protein was probed simultaneously with VP5. The blots were analyzed by densitometry; the top and bottom of each gel show the images taken of VP5 and the tested proteins, respectively.
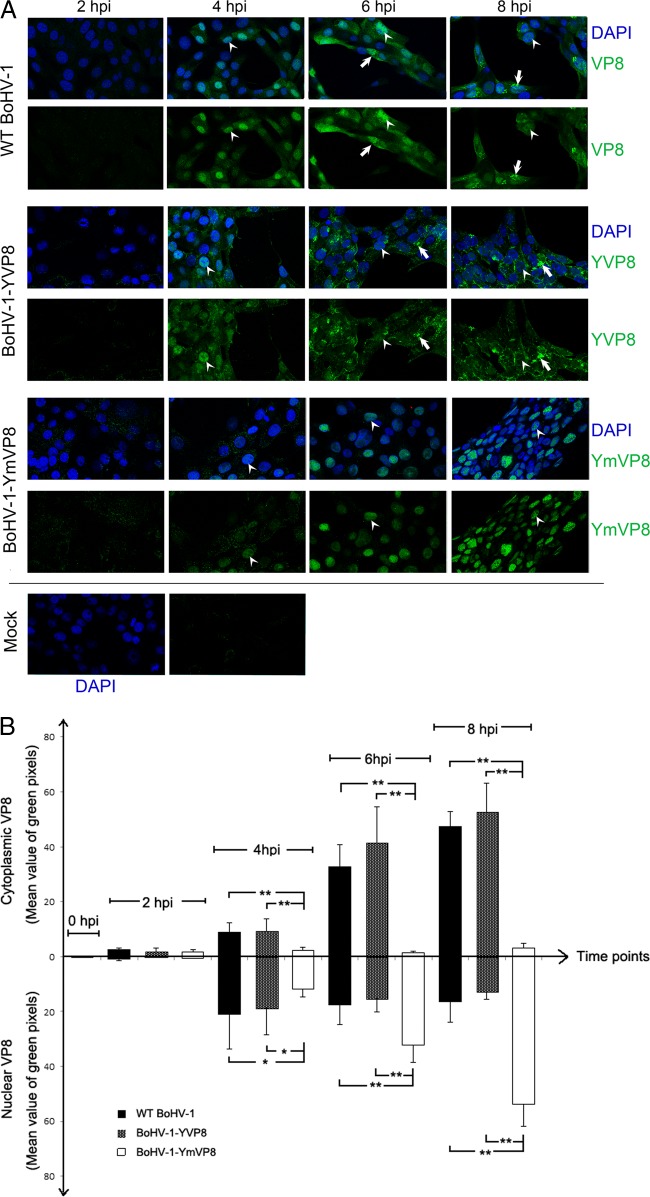
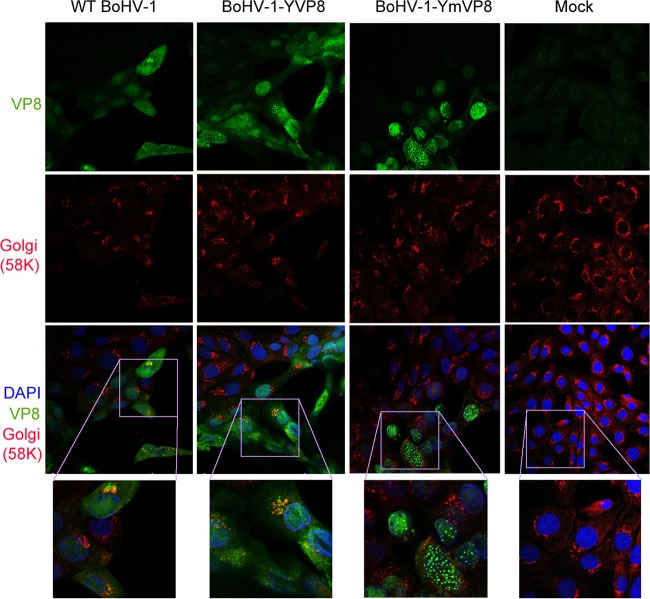
The localization of VP8 in infected MDBK cells was observed by confocal microscopy (Fig. 4A). WT VP8 first appeared in the cytoplasm at 2 hpi. At 4 hpi, the majority of WT VP8 localized in the nucleus, and the level of nuclear VP8 slightly decreased during the following 4 h. In contrast, the cytoplasmic VP8 population increased slowly before 4 hpi and increased relatively quickly from 6 to 8 hpi, ultimately exceeding the nuclear population. A similar trend in YFP-VP8 distribution was found in BoHV-1-YVP8-infected cells (Fig. 4B). In BoHV-1-YmVP8-infected cells, the nonphosphorylated Mut-VP8 first appeared in the cytoplasm at 2 hpi. Nuclear Mut-VP8 started to increase from 2 to 8 hpi, while the cytoplasmic level remained the same (Fig. 4B). Similar results with respect to the distribution of VP8 proteins were found in infected embryonic bovine tracheal (EBTr) cells (data not shown). To track the location of VP8 in the cytoplasm, infected cells were labeled with monoclonal anti-58K protein antibody, which is used as a marker of the Golgi apparatus (23). In WT BoHV-1- and BoHV-1-YVP8-infected cells, VP8 accumulated in the cytoplasm and localized to the Golgi network, while YmVP8 did not localize to the cytoplasm or Golgi network of BoHV-1-YmVP8-infected cells (Fig. 5).
FIG 4.
Localization of VP8 proteins in infected MDBK cells. MDBK cell monolayers were infected with WT BoHV-1, BoHV-1-YVP8, or BoHV-1-YmVP8 at an MOI of 5 or mock infected and processed for immunofluorescence staining every 2 h until 8 hpi. (A) VP8 was labeled with polyclonal anti-VP8 antibody and Alexa 488-conjugated antibody. DNA was labeled with DAPI. The arrowheads indicate nuclear VP8, and the arrows indicate cytoplasmic VP8. (B) The cell images for each time point were analyzed with a biological image-processing program, Fiji (34). The average green value within a selected area in an image was calculated by the program. At each time point, the cytoplasm and the nuclei of 10 cells were selected and analyzed. The mean values are shown. The data were analyzed by two-tailed t test. The statistical significance of the difference between the values is shown; *, 0.01 < P ≤ 0.05; **, P ≤ 0.01. The error bars indicate standard deviations.
FIG 5.
Colocalization of cytoplasmic VP8 with the Golgi apparatus. MDBK cell monolayers were infected with WT BoHV-1, BoHV-1-YVP8, or BoHV-1-YmVP8 at an MOI of 0.001 until plaques were well developed or mock infected. WT VP8 was labeled with monoclonal anti-VP8 antibody and Alexa 488-conjugated antibody. The Golgi apparatus was identified with polyclonal anti-58K protein antibody and Alexa 633-conjugated antibody. The DNA was labeled with DAPI. The cells were observed under a Leica SP5 confocal microscope. The boxed areas are enlarged in the bottom row.
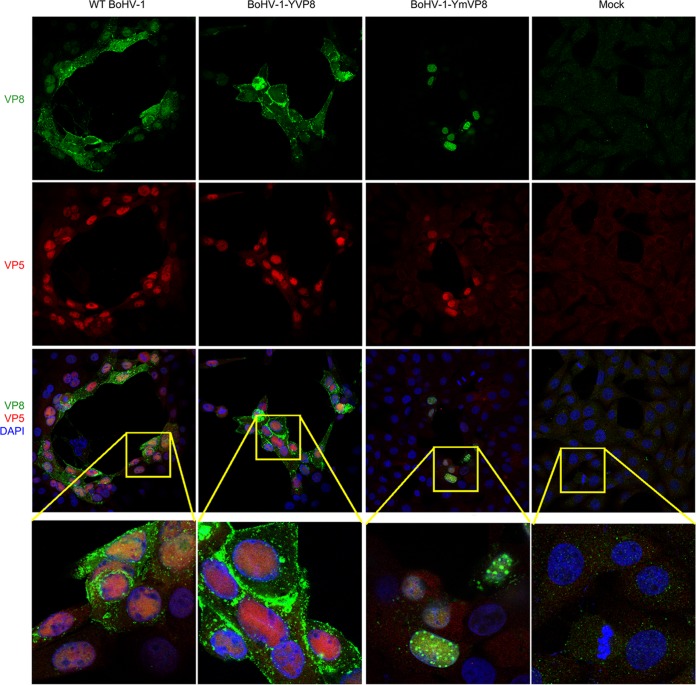
To examine whether Golgi apparatus association of VP8 resulted from translocation of virus particles, capsid protein VP5 was labeled with polyclonal anti-VP5 antibody in infected MDBK cells. In WT BoHV-1-infected cells, the VP5 signal was mostly in the nucleus and did not associate with cytoplasmic VP8, indicating that the majority of cytoplasmic VP8 was not virus associated (Fig. 6). The distributions of VP5 in WT BoHV-1- and BoHV-1-YmVP8-infected cells were not different. YmVP8 showed a punctate pattern, but VP5 did not, indicating that the speckled YmVP8 was not virus associated.
FIG 6.
Coimmunostaining of VP8 and VP5. MDBK cell monolayers were infected with WT BoHV-1, BoHV-1-YVP8, or BoHV-1-YmVP8 at an MOI of 0.001 until plaques were well developed or mock infected. WT VP8 was labeled with monoclonal anti-VP8 antibody and Alexa 488-conjugated antibody. VP5 was identified with polyclonal anti-VP5 antibody and Alexa 633-conjugated antibody. DNA was labeled with DAPI. The cells were observed under a Leica SP5 confocal microscope. The boxed areas are enlarged in the bottom row.
Phosphorylation of VP8 benefits the viral DNA content.
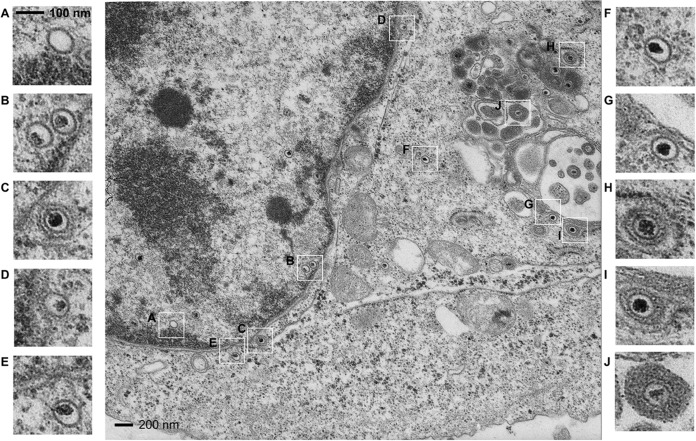
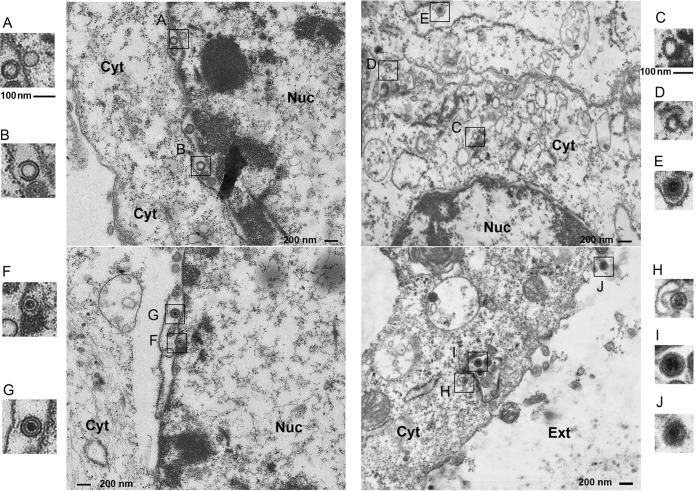
To further study the effects of VP8 phosphorylation, the BoHV-1 life cycle was studied by transmission electron microcopy. Figure 7 demonstrates the life cycle of WT BoHV-1 from empty nucleocapsids to cytoplasmic enveloped particles. Capsid formation and DNA encapsidation took place in the nucleus (Fig. 7A and B). The intranuclear capsids exited from the nucleus to the cytoplasm by budding through the two leaflets of the nuclear membrane. First, these nucleocapsids were in intimate contact with the inner nuclear membrane, while the membrane bent inward to enclose the capsid (Fig. 7B), resulting in a primary enveloped virus in the perinuclear space (Fig. 7C). These capsids were surrounded by a smooth envelope. The capsids were released from the perinuclear space by fusion of the primary envelope with the outer nuclear membrane (Fig. 7D), so that the viral particles lost the primary envelope (Fig. 7E) and became free naked capsids in the cytoplasm (Fig. 7F). The cytoplasmic capsids migrated toward to the Golgi apparatus area (Fig. 7G) and obtained the secondary envelope by being wrapped with the Golgi membrane (Fig. 7H and I). Viral capsids were surrounded by a thick layer of electron-dense material in the Golgi apparatus, and the Golgi apparatus-derived envelope contained visible surface projections compared with the perinuclear envelope (Fig. 7H to J). BoHV-1-YVP8-infected cells were also observed by transmission electron microscopy, and the virus exhibited the same maturation as WT virus (data not shown).
FIG 7.
Transmission electron microscopy of cells infected with WT BoHV-1. MDBK cells infected with WT BoHV-1 at an MOI of 1 were collected and processed at 15 hpi and observed with a Philips CM10 transmission electron microscope. The boxed areas in the central image are enlarged on the right and left.
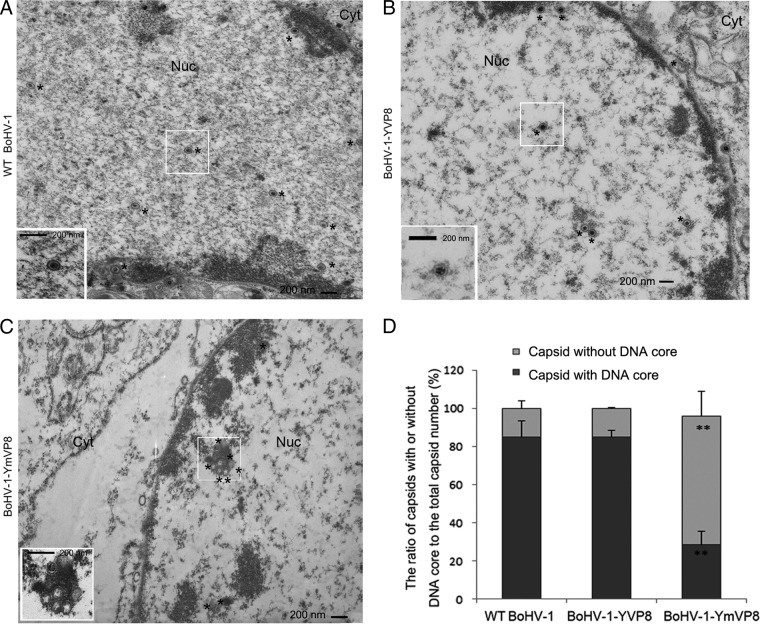
Compared with WT BoHV-1 (Fig. 8A) and BoHV-1-YVP8 (Fig. 8B) nuclear capsids, which mostly contained DNA cores at 15 hpi, BoHV-1-YmVP8-infected cells had more empty nucleocapsids present in the nucleus (Fig. 8C). Five views were captured in the nuclei of cells infected with WT BoHV-1, BoHV-1-YFP, or BoHV-1-YmVP8. The capsids with and without DNA cores were counted in each view. Over 80% of the WT BoHV-1 and BoHV-1-YVP8 nucleocapsids had DNA incorporated, while less than 30% of the BoHV-1-YmVP8 nucleocapsids had DNA cores (Fig. 8D). These results suggest that phosphorylated VP8 might play a role in promoting production or encapsidation of viral DNA.
FIG 8.
Analysis of the impact of blocking phosphorylation of VP8 on viral DNA content. (A to C) MDBK cells infected with WT BoHV-1, BoHV-1-YVP8, or BoHV-1-YmVP8 at an MOI of 1 were collected and processed at 15 hpi and observed with a Philips CM10 transmission electron microscope. The locations of nuclear capsids are marked with asterisks. Cyt, cytoplasmic area; Nuc, nuclear area. The boxed areas are enlarged in the bottom left corners. (D) The capsids from five views of the nuclear areas infected with WT BoHV-1, BoHV-1-YVP8, or BoHV-1-YmVP8 were counted. The data were analyzed by two-tailed t test. The statistical significance of the difference between the values is shown; **, P ≤ 0.01. The error bars indicate standard deviations.
Phosphorylation on VP8 does not affect virus particle release from the nucleus.
In BoHV-1-YmVP8-infected cells, both empty capsids (Fig. 9A to D) and DNA-containing capsids (Fig. 9F to I) egressed from the nucleus through the envelopment–de-envelopment pathway. Empty nucleocapsids migrated toward the inner nuclear membrane (Fig. 9A), which subsequently enclosed the capsids, forming a primary envelope surrounding the empty capsids. The particles were presented as two-layer halos in the perinuclear space (Fig. 9A and B). DNA-free capsids (Fig. 9C and D) and DNA-containing capsids (Fig. 9E, H, and I) were found in the cytoplasm. Figure 9H shows that DNA-containing capsids were captured by the Golgi apparatus structure in the cytoplasm for further maturation and eventual release out of the cells (Fig. 9J). There was no apparent evidence for blockage of empty-capsid egress from the nucleus in BoHV-1-YmVP8-infected cells.
FIG 9.
Transmission electron microscopy of cells infected with BoHV-1-YmVP8. MDBK cells infected with BoHV-1-YmVP8 at an MOI of 1 were collected and processed at 15 hpi and observed with a Philips CM10 transmission electron microscope. The boxed areas from the large images are enlarged on the right and left. Cyt, cytoplasmic area; Nuc, nucleus area; Ext, extracellular area.
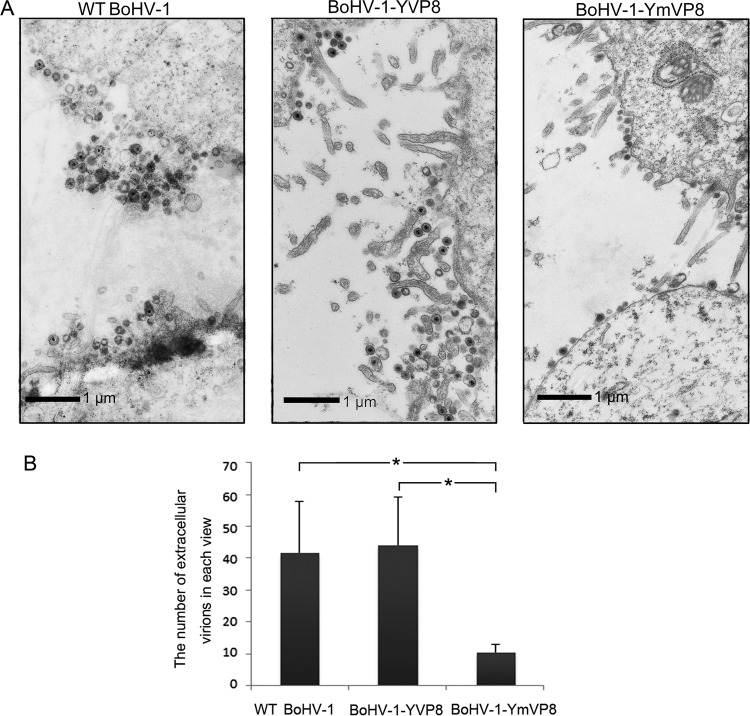
The amounts of extracellular WT BoHV-1, BoHV-1-YVP8, and BoHV-1-YmVP8 are presented in Fig. 10. In agreement with the viral growth characteristics (Fig. 2B), BoHV-1-YmVP8-infected cells were surrounded by apparently smaller amounts of extracellular virions than cells infected with WT BoHV-1 or BoHV-1-YVP8 (Fig. 10A). Five views of extracellular WT BoHV-1, BoHV-1-YVP8, and BoHV-1-YmVP8 were captured, and the mature virus particles were counted. The numbers of released WT BoHV-1 virions and BoHV-1-YVP8 virions were significantly higher than those of BoHV-1-YmVP8 (Fig. 10B). The ultrastructures of the three viruses did not have any obvious differences (data not shown).
FIG 10.
Analysis of extracellular viruses. (A) MDBK cells infected with WT BoHV-1, BoHV-1-YVP8, or BoHV-1-YmVP8 at an MOI of 1 were collected and processed at 15 hpi and observed with a Philips CM10 transmission electron microscope. The extracellular viral particles are displayed. (B) The viral particles from five views of extracellular areas of each treatment were counted. The data were analyzed by two-tailed t test. The statistical significance of the difference between the values is shown; *, 0.01 < P ≤ 0.05. The error bars indicate standard deviations.
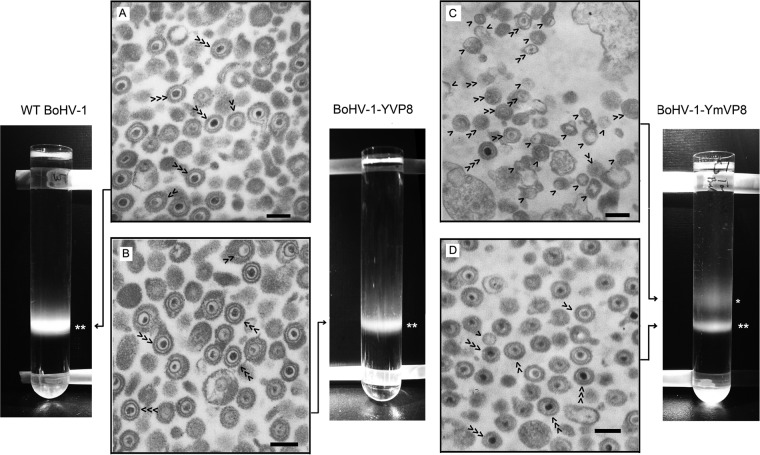
To further confirm the presence of empty virions in BoHV-1-YmVP8-infected cells, viral particles released to the cell culture medium were collected and separated in a sodium potassium tartrate density gradient (Fig. 11). WT BoHV-1 and BoHV-1-YVP8 consisted of a single virus band located at 40% to 50% of the density gradient, while BoHV-1-YmVP8 had two visible bands. The lower band had the same density as wild-type virus. These bands contained mature virions with DNA cores (Fig. 11A, B, and D). The upper band of BoHV-1-YmVP8 contained incomplete virions without DNA cores or with partial genomes (Fig. 11C), which thus had a lower density than the complete virions. The band with complete WT BoHV-1 virions was thicker and brighter than that of BoHV-1-YmVP8. These data are consistent with the results shown in Fig. 2 and 10, which demonstrated lower virus production by BoHV-1-YmVP8.
FIG 11.
Gradient sedimentation analysis and TEM observation of extracellular viruses. Extracellular viral particles were loaded onto 10 to 60% potassium sodium tartrate gradients in TNE buffer and sedimented by centrifugation at 25,000 rpm for 2 h at 4°C in a SW41 rotor. The virus bands were visualized by passing a light beam from the top through the tartrate gradient in a dark environment. The double asterisks indicate bands containing complete viruses with higher density, and the single asterisk indicates a band containing incomplete viruses with lower density. (A to D) The virus bands were aspired and observed by TEM. The single arrowheads indicate virions without a DNA core, the double arrowheads indicate virions with an incomplete DNA core, and the triple arrowheads indicate virions with a complete DNA core (bars, 200 nm).
DISCUSSION
To characterize the function of phosphorylation on VP8 and eventually the impact on the life cycle of BoHV-1, a recombinant BoHV-1 expressing mutant VP8 was constructed. The mutant VP8, which has been previously studied in vitro (13) outside the context of infection, is not phosphorylated by CK2 or US3, as amino acid residues at the essential sites for the kinases have been replaced with alanines. The results showed that blocking phosphorylation on VP8 leads to reduced virus production (Fig. 2B). Accordingly, a lower number of complete BoHV-1-YmVP8 particles than WT BoHV-1 particles were found adjacent to infected cells (Fig. 10). According to the results of the single-step growth curve, there may be at least two reasons for this reduction. First, the replication of the VP8 mutant virus was impaired compared to that of wild-type virus. From 5 to 15 hpi, intracellular BoHV-1-YmVP8 virions increased 163-fold (from 100.8 PFU/ml to 103.1 PFU/ml), whereas intracellular WT BoHV-1 experienced a more rapid amplification, ∼27,000-fold (from 10 PFU/ml to 105.4 PFU/ml). As a result, during this period, the amount of extracellular BoHV-1-YmVP8 was lower than that of extracellular WT BoHV-1. Second, the mature mutant virus was less efficiently released from the host cells than the wild-type virus. From 10 to 25 hpi, extracellular WT virus showed a rapid increase of ∼2,000-fold (from 104.6 PFU/ml to 107.9 PFU/ml), which was associated with a slower increase in intracellular virus. In contrast, the extracellular mutant virus increased ∼20-fold (from 103.4 PFU/ml to 104.8 PFU/ml) (Fig. 2C and E). Repairing phosphorylation on VP8 (Fig. 2A) restored the virus titer (Fig. 2B) and growth property (Fig. 2F), suggesting the attenuated virus growth in BoHV-1-YmVP8 was due to diminished phosphorylation on VP8.
The ultrastructural study of virus-infected cells revealed a potential function of phosphorylated VP8 in BoHV-1 maturation. The maturation process of WT BoHV-1 from the nucleus to the cytoplasm is displayed in Fig. 7. Capsid formation and DNA incorporation occurred in the nucleus. Nuclear capsids traveled through the nuclear membrane via an envelopment–de-envelopment process and obtained most of the tegument and envelope in the cytoplasm. This maturation and egress model is in agreement with the one described previously for other herpesviruses, such as HHV-1 (24), human cytomegalovirus (HCMV) (25), and pseudorabies virus (PRV) (26). In the nuclei of BoHV-1-YmVP8-infected cells, most capsids did not contain a DNA core, indicating that phosphorylated VP8 may promote viral DNA production or packaging into capsids; however, this is not an absolute requirement, as a certain number of DNA-containing particles were found in the nucleus, cytoplasm, and extracellular environment. Encapsidation of herpesvirus DNA involves cooperation of many viral proteins that cleave viral DNA and introduce the cleaved DNA into capsids (27). The UL25 gene product of HHV-1 encompasses a capsid-binding domain through which the protein targets capsids and propels DNA into the capsid (28). Viral structural proteins L1 and L2 of PRV are required for assembly of viral DNA into capsids (29). For BoHV-1, phosphorylated VP8 appeared to be a contributing factor for DNA replication or encapsidation. DNA incorporation was not critical for nuclear egress of capsids, and neither was phosphorylation of VP8, because empty capsids were present in the perinuclear space and cytoplasm. In BoHV-1-YmVP8-infected cells, empty capsids merged with the inner nuclear membrane and obtained a primary envelope similar to that of DNA-containing capsids. Ultimately, the DNA-free particles were released to the extracellular environment, resulting in an additional, lower-density virus band in a potassium tartrate gradient (Fig. 11).
The inefficient incorporation of mutant VP8 into the virion indicates that optimal VP8 packaging requires phosphorylation and that this may have an impact on virion maturation and release. It also suggests that VP8 may be incorporated at two stages, the first occurring in the nucleus and the second taking place in the cytoplasm. The YmVP8 from the mutant virus was mainly localized in the nucleus during infection (Fig. 4), indicating that the virion-associated YmVP8 was incorporated prior to or during nuclear egress and that phosphorylation by CK2 and US3 is not required for the primary incorporation of VP8. This conclusion is also supported by data from a previous study using VP8-specific immunogold staining that showed accumulation of VP8 protein around the nuclear capsids, as well as along the outer nuclear membrane (9). The addition of certain tegument components of herpesviruses to capsids prior to or during nuclear egress is referred to as primary tegumentation (30). In HHV-1, UL36, UL37, ICP0, and ICP4 have been identified as primary tegument proteins that are present on the capsids before they reach the cytoplasm (30). The UL31 and UL34 gene products of PrV are incorporated in the primary enveloped capsids and eventually dissociate from the capsids during the later process (31). US3 of PrV is another primary tegument protein that, however, does not dissociate from the viral particle during de-envelopment (32).
Both viral particles (Fig. 7) and WT VP8 protein (Fig. 5) accumulated in the Golgi apparatus area during the late stages of infection, indicating that the Golgi apparatus is a major site for VP8 incorporation. Previous evidence that immunogold particle-labeled VP8 localized in the Golgi apparatus area (9) also supports these results. The secondary incorporation of VP8 requires the protein to be phosphorylated by CK2 and/or US3, as without CK2- and/or US3-mediated phosphorylation, it did not associate with the Golgi apparatus (Fig. 5). It is less likely that the Golgi apparatus-associated VP8 is the result of viral particle transport, as the cytoplasmic localization of VP8 was independent of the VP5 protein, a major component of viral capsids (Fig. 6).
The fact that VP8 localizes in the cytoplasm and associates with the Golgi apparatus only during infection but remains mainly in the nucleus in VP8-transfected cells (13) suggests that VP8 needs the presence of other viral components for direction to the Golgi apparatus. Blocking phosphorylation on VP8 inhibited the protein from targeting the Golgi apparatus, which likely resulted in disruption of the secondary incorporation of VP8 and ultimately reduced infectious-virus production. In contrast, tegument protein VP13/14, a gene product of HHV-1 UL47 and a homologue of VP8, assembles mainly in the nucleus. The majority of VP13/14 localizes in the nucleus during all infection stages, and it forms punctate patterns in the late stages. The relative cytoplasmic intensity of VP13/14 is very low in comparison to the nuclear intensity, which may result from viral particle migration (33).
In summary, this study demonstrates that blocking phosphorylation of VP8 may have a negative impact on viral DNA encapsidation and reduce viral replication efficiency. Phosphorylation was found to be essential for the cytoplasmic incorporation of VP8 into virions, but not for primary packaging of VP8 in the nucleus. Ultimately, the impairment in cytoplasmic maturation of the virus reduced the efficiency of virus release.
ACKNOWLEDGMENTS
We thank Guosheng Liu for kind assistance with TEM experiments.
This research was supported by grant 90887-2010 RGPIN from the Natural Sciences and Engineering Research Council of Canada. Kuan Zhang was supported by a scholarship from the China Scholarship Council.
Footnotes
This is VIDO manuscript number 753.
REFERENCES
- 1.Levings RL, Roth JA. 2013. Immunity to bovine herpesvirus 1. I. Viral lifecycle and innate immunity. Anim Health Res Rev 14:88–102. doi: 10.1017/S1466252313000042. [DOI] [PubMed] [Google Scholar]
- 2.Bosse JB, Virding S, Thiberge SY, Scherer J, Wodrich H, Ruzsics Z, Koszinowski UH, Enquist LW. 2014. Nuclear herpesvirus capsid motility is not dependent on F-actin. MBio 5:e01909-01914. doi: 10.1128/mBio.01909-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Shen S, Jia X, Guo H, Deng H. 2015. Tegument protein ORF33 of a gammaherpesvirus is associated with intranuclear capsids at an early stage of the tegumentation process. J Virol 89:5288–5297. doi: 10.1128/JVI.00079-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Yu X, Li W, Liu L, Che Y, Cun W, Wu W, He C, Shao C, Li Q. 2008. Functional analysis of transcriptional regulation of herpes simplex virus type 1 tegument protein VP22. Sci China C Life Sci 51:966–972. doi: 10.1007/s11427-008-0127-4. [DOI] [PubMed] [Google Scholar]
- 5.Kato A, Liu Z, Minowa A, Imai T, Tanaka M, Sugimoto K, Nishiyama Y, Arii J, Kawaguchi Y. 2011. Herpes simplex virus 1 protein kinase Us3 and major tegument protein UL47 reciprocally regulate their subcellular localization in infected cells. J Virol 85:9599–9613. doi: 10.1128/JVI.00845-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Carpenter DE, Misra V. 1991. The most abundant protein in bovine herpes 1 virions is a homologue of herpes simplex virus type 1 UL47. J Gen Virol 72:3077–3084. doi: 10.1099/0022-1317-72-12-3077. [DOI] [PubMed] [Google Scholar]
- 7.Potel C, Elliott G. 2005. Phosphorylation of the herpes simplex virus tegument protein VP22 has no effect on incorporation of VP22 into the virus but is involved in optimal expression and virion packaging of ICP0. J Virol 79:14057–14068. doi: 10.1128/JVI.79.22.14057-14068.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lobanov VA, Maher-Sturgess SL, Snider MG, Lawman Z, Babiuk LA. 2010. A UL47 gene deletion mutant of bovine herpesvirus type 1 exhibits impaired growth in cell culture and lack of virulence in cattle. J Virol 84:445–458. doi: 10.1128/JVI.01544-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.van Drunen Littel-van den Hurk S, Garzon S, van den Hurk JV, Babiuk LA, Tijssen P. 1995. The role of the major tegument protein VP8 of bovine herpesvirus-1 in infection and immunity. Virology 206:413–425. doi: 10.1016/S0042-6822(95)80057-3. [DOI] [PubMed] [Google Scholar]
- 10.Zheng C, Brownlie R, Babiuk LA, van Drunen Littel-van den Hurk S. 2004. Characterization of nuclear localization and export signals of the major tegument protein VP8 of bovine herpesvirus-1. Virology 324:327–339. doi: 10.1016/j.virol.2004.03.042. [DOI] [PubMed] [Google Scholar]
- 11.Verhagen J, Donnelly M, Elliott G. 2006. Characterization of a novel transferable CRM-1-independent nuclear export signal in a herpesvirus tegument protein that shuttles between the nucleus and cytoplasm. J Virol 80:10021–10035. doi: 10.1128/JVI.01322-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Verhagen J, Hutchinson I, Elliott G. 2006. Nucleocytoplasmic shuttling of bovine herpesvirus 1 UL47 protein in infected cells. J Virol 80:1059–1063. doi: 10.1128/JVI.80.2.1059-1063.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Zhang K, Afroz S, Brownlie R, Snider M, van Drunen Littel-van den Hurk S. 2015. Regulation and function of phosphorylation on VP8, the major tegument protein of bovine herpesvirus-1. J Virol 89:4598–4611. doi: 10.1128/JVI.03180-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Everett RD, Rechter S, Papior P, Tavalai N, Stamminger T, Orr A. 2006. PML contributes to a cellular mechanism of repression of herpes simplex virus type 1 infection that is inactivated by ICP0. J Virol 80:7995–8005. doi: 10.1128/JVI.00734-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Islam A, Schulz S, Afroz S, Babiuk LA, van Drunen Littel-van den Hurk S. 2015. Interaction of VP8 with mRNAs of bovine herpesvirus-1. Virus Res 197:116–126. doi: 10.1016/j.virusres.2014.12.017. [DOI] [PubMed] [Google Scholar]
- 16.Pomeranz LE, Blaho JA. 1999. Modified VP22 localizes to the cell nucleus during synchronized herpes simplex virus type 1 infection. J Virol 73:6769–6781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Geiss BJ, Tavis JE, Metzger LM, Leib DA, Morrison LA. 2001. Temporal regulation of herpes simplex virus type 2 VP22 expression and phosphorylation. J Virol 75:10721–10729. doi: 10.1128/JVI.75.22.10721-10729.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Labiuk SL, Babiuk LA, van Drunen Littel-van den Hurk S. 2009. Major tegument protein VP8 of bovine herpesvirus 1 is phosphorylated by viral US3 and cellular CK2 protein kinases. J Gen Virol 90:2829–2839. doi: 10.1099/vir.0.013532-0. [DOI] [PubMed] [Google Scholar]
- 19.Shaw AM, Braun L, Frew T, Hurley DJ, Rowland RR, Chase CC. 2000. A role for bovine herpesvirus 1 (BHV-1) glycoprotein E (gE) tyrosine phosphorylation in replication of BHV-1 wild-type virus but not BHV-1 gE deletion mutant virus. Virology 268:159–166. doi: 10.1006/viro.1999.0164. [DOI] [PubMed] [Google Scholar]
- 20.Ren X, Harms JS, Splitter GA. 2001. Tyrosine phosphorylation of bovine herpesvirus 1 tegument protein VP22 correlates with the incorporation of VP22 into virions. J Virol 75:9010–9017. doi: 10.1128/JVI.75.19.9010-9017.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Li Y, van Drunen Littel-van den Hurk S, Babiuk LA, Liang X. 1995. Characterization of cell-binding properties of bovine herpesvirus 1 glycoproteins B, C, and D: identification of a dual cell-binding function of gB. J Virol 69:4758–4768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Labiuk SL, Lobanov V, Lawman Z, Snider M, Babiuk LA. 2010. Bovine herpesvirus-1 US3 protein kinase: critical residues and involvement in the phosphorylation of VP22. J Gen Virol 91:1117–1126. doi: 10.1099/vir.0.016600-0. [DOI] [PubMed] [Google Scholar]
- 23.Bashour A-M, Bloom GS. 1998. 58K, a microtubule-binding Golgi protein, is a formiminotransferase cyclodeaminase. J Biol Chem 273:19612–19617. doi: 10.1074/jbc.273.31.19612. [DOI] [PubMed] [Google Scholar]
- 24.Guan Y, Guo L, Yang E, Liao Y, Liu L, Che Y, Zhang Y, Wang L, Wang J, Li Q. 2014. HSV-1 nucleocapsid egress mediated by UL31 in association with UL34 is impeded by cellular transmembrane protein 140. Virology 464-465:1–10. doi: 10.1016/j.virol.2014.06.034. [DOI] [PubMed] [Google Scholar]
- 25.Mettenleiter TC. 2002. Herpesvirus assembly and egress. J Virol 76:1537–1547. doi: 10.1128/JVI.76.4.1537-1547.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Schulz KS, Klupp BG, Granzow H, Passvogel L, Mettenleiter TC. 2015. Herpesvirus nuclear egress: Pseudorabies virus can simultaneously induce nuclear envelope breakdown and exit the nucleus via the envelopment-deenvelopment-pathway. Virus Res 209:76–86. doi: 10.1016/j.virusres.2015.02.001. [DOI] [PubMed] [Google Scholar]
- 27.Visalli RJ, van Zeijl M. 2003. DNA encapsidation as a target for anti-herpesvirus drug therapy. Antivir Res 59:73–87. doi: 10.1016/S0166-3542(03)00108-6. [DOI] [PubMed] [Google Scholar]
- 28.Cockrell SK, Sanchez ME, Erazo A, Homa FL. 2009. Role of the UL25 protein in herpes simplex virus DNA encapsidation. J Virol 83:47–57. doi: 10.1128/JVI.01889-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kawana K, Yoshikawa H, Taketani Y, Yoshiike K, Kanda T. 1998. In vitro construction of pseudovirions of human papillomavirus type 16: incorporation of plasmid DNA into reassembled L1/L2 capsids. J Virol 72:10298–10300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Henaff D, Remillard-Labrosse G, Loret S, Lippe R. 2013. Analysis of the early steps of herpes simplex virus 1 capsid tegumentation. J Virol 87:4895–4906. doi: 10.1128/JVI.03292-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Fuchs W, Klupp BG, Granzow H, Osterrieder N, Mettenleiter TC. 2002. The interacting UL31 and UL34 gene products of pseudorabies virus are involved in egress from the host-cell nucleus and represent components of primary enveloped but not mature virions. J Virol 76:364–378. doi: 10.1128/JVI.76.1.364-378.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Granzow H, Klupp BG, Mettenleiter TC. 2004. The pseudorabies virus US3 protein is a component of primary and of mature virions. J Virol 78:1314–1323. doi: 10.1128/JVI.78.3.1314-1323.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Donnelly M, Elliott G. 2001. Fluorescent tagging of herpes simplex virus tegument protein VP13/14 in virus infection. J Virol 75:2575–2583. doi: 10.1128/JVI.75.6.2575-2583.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B. 2012. Fiji: an open-source platform for biological-image analysis. Nat Methods 9:676–682. doi: 10.1038/nmeth.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]