Abstract
The cocirculation of different HIV types and groups can lead to dual infections and recombinants, which hinder diagnosis and therapeutic management. We designed two multiplex PCRs (mPCRs) coupled with capillary electrophoresis to facilitate the detection of such infections. The first, MMO2, targets three variants (HIV-1/M, HIV-1/O, and HIV-2), and the second, MMO, targets HIV-1/M and HIV-1/O. These mPCRs were validated on DNA and RNA extracts from 19 HIV-1/M, 12 HIV-1/O, and 13 HIV-2 cultures and from mixtures simulating dual infections. They were then assessed with DNA and RNA extracts from samples of 47 clinical monoinfections and HIV-1/M+O dual infections or infections with HIV-1/MO recombinants. Both mPCRs had excellent specificity. Sensitivities ranged from 80 to 100% for in vitro samples and from 58 to 100% for clinical samples, with the results obtained depending on the material used and the region of the genome concerned. Sensitivity was generally lower for DNA than for RNA and for amplifications of the integrase and matrix regions. In terms of global detection (at least one target gene for each strain), both mPCRs yielded a detection rate of 100% for in vitro samples. MMO2 detected 100% of the clinical strains from DNA and 97% from RNA, whereas MMO detected 100% of the strains from both materials. Thus, for in vitro and clinical samples, MMO2 was a useful tool for detecting dual infections with HIV-1 and HIV-2 (referred to as HIV-1+HIV-2) and HIV-1/M+O, and MMO was useful for detecting both MO dual infections and MO mosaic patterns.
INTRODUCTION
HIV displays considerable genetic diversity due to its simian origin, specific structural traits, replication mechanisms involving the low fidelity of the reverse transcriptase (RT) (1), host pressure, and the possibility of recombination events (2).
The current HIV classification includes two types, HIV-1 and HIV-2, which are further subdivided into groups (groups M to P and A to I, respectively) (3, 4). HIV-1/M (“major”) is pandemic, whereas HIV-1/O, HIV-1/N, and HIV-1/P are much rarer (10,000 to 30,000, 16, and 2 cases, respectively) and are endemic in West-Central Africa and countries historically linked to this area (5, 6). HIV-2 is endemic to West Africa, where only groups A and B are epidemic (4, 7). HIV-2 groups C to I are each found in one (or two, for group F) individual living in Ivory Coast, Liberia, or Sierra Leone (7–11). In these regions, the cocirculation of HIV-1/M and HIV-2 or HIV-1/O leads to dual infections (12–21). In West Africa, dual infections with HIV-1 and HIV-2 (referred to as HIV-1+HIV-2) are thought to account for 1 to 8% of HIV infections (17, 22), although no HIV-1/2 recombinant has yet been described. In West-Central Africa, including Cameroon in particular, the frequency of HIV-1/M+O dual infections has been estimated to be 10% in the HIV-1/O-seropositive population (20, 23, 24). However, in contrast to what has been reported for HIV-2, HIV-1/M+O dual infections have been shown to lead to the circulation of recombinants with or without their parental strains (25–29).
Dual infections and recombinants have major consequences for the course of the worldwide HIV epidemic and are essential indicators for molecular epidemiology. They also have an impact on clinical practice, because they may interfere with the results obtained with serological and molecular tools for diagnosis and patient follow-up. Some serological tools are less sensitive to HIV-1/O (30) or HIV-2 (22, 31) and may behave similarly with recombinants carrying a group O-derived envelope or with cryptic HIV-1+HIV-2 infections (17). Similarly, variants may affect the determination of viral load (VL) by quantitative RT-PCR (32–35). This justifies the use of nonspecific primers to prevent the misquantification or lack of detection of one of the variants in cases of dual infection or recombinants involving minor variants. Conversely, specific techniques targeting variants may also be required for diagnosing and following up the mother-to-child transmission of dual infections or for monitoring the overgrowth of one variant with respect to the other to facilitate the adaptation of treatment and to prevent virological failure.
Variants can also have an impact on antiretroviral therapy (ART) through their natural polymorphism. HIV-1/O strains are naturally resistant to nonnucleoside RT inhibitors (NNRTI), (36), and HIV-2 is naturally resistant to NNRTI, enfuvirtide (T20), and some protease inhibitors (4). Consequently, dual infections with these viruses and recombinants, if undetected, will affect the response to ART. Correct detection is therefore essential to prevent the development and selection of resistant mutants by ensuring that the patient receives the most appropriate treatment.
Current tools for detecting dual infections and recombinant forms remain inadequate. Serological methods based on Western blotting or rapid diagnostic tests can differentiate between HIV-1 and -2 infections but are not specific enough (15, 31, 37). Homemade V3 serotyping assays are useful for discriminating between HIV-1 and -2, HIV-1/M subtype B and non-subtype B, and HIV-1/O, -N, and -P infections (37–39) but are performed only in specialized laboratories and do not allow the genomic characterization of viral strains. Most of the molecular tools developed to date were designed to detect intra-HIV-1/M dual infections and recombinants. These tools include the heteroduplex tracking assay (HTA) (21, 40, 41), the multiregion hybridization assay (MHA) (42), and type- or group-specific PCR followed by sequencing and phylogenetic analyses, for which the number of experiments to be performed depends on the number of target strains and genomic regions considered. The advent of next-generation sequencing (NGS) has made it possible to carry out both nonspecific and specific population sequencing and has facilitated the detection of dual infections and recombination (43, 44), but it remains expensive and requires specific facilities. No molecular tool has yet been developed for detecting HIV-1+HIV-2 and HIV-1/M+O dual infections or HIV-1/MO recombinants in a single experiment.
We aimed to develop a multiplex PCR (mPCR) strategy followed by capillary electrophoresis (CE) for the detection of dual infections involving HIV-1/M, HIV-1/O, and HIV-2 and for the specific detection of HIV-1/MO recombinants. We used in vitro samples to develop the method, which we then tested on clinical samples.
MATERIALS AND METHODS
mPCRs.
HIV-1+HIV-2 and HIV-1/M+O dual infections raise different issues, as HIV-1/M+O dual infections can lead to the production of recombinants, which has never been reported for HIV-1+HIV-2 dual infections. We therefore designed the following two multiplex PCRs: the first, MMO2, targeted the integrase (IN) and envelope (gp41) regions of the HIV-1/M, HIV-1/O, and HIV-2 genomes, whereas the second, MMO, targeted five genomic regions of HIV-1/M and HIV-1/O, corresponding to the matrix (MA), protease (PR), integrase (IN), SUgp120 (gp120), and TMgp41 (gp41) regions (see the supplemental methods and Fig. S1 in the supplemental material).
These mPCRs were validated and assessed with DNAs extracted from infected cells in vitro (in vitro samples) or from peripheral blood mononuclear cells (PBMCs) isolated from clinical samples (clinical samples) and with RNAs extracted from viral supernatants (in vitro samples) or plasma samples (clinical samples) (see the supplemental material). PCR and RT-PCR were performed on all samples as described in the supplemental material.
CE.
We used a model CEQ8000 automated Sanger sequencer (Beckman Coulter) to analyze the mPCR products. For each pair of primers, according to amplicon size, we labeled one primer with one of the following three dyes compatible with the CEQ8000 laser detection channels: Alexa Fluor 750 (AF750; channel D2), Cy5.5 (channel D3), and ATTO647N (AT647N; channel D4). Channel D1 was used to detect the MM1000 molecular ladder (MapMarker1000, D1 labeled; BioVentures, Inc.) for the generation of a calibration curve and the assignment of peak sizes.
We also used an electrophoresis internal control (IC) to validate the migration process and to make it possible to compare samples by calculating the normalized ratio (R) of the intensity of each peak to the IC intensity. The IC consisted of three PCR products, each labeled with a different dye, resulting from the amplification of gp41 and gp120 (channel D2 [720 nucleotides] and channel D4 [424 nucleotides], respectively) from the HIV-2 group B (HIV-2/B) strain DIL and of PR (channel D3 [653 nucleotides]) from the HIV-2 group A (HIV-2/A) strain ROD. These fragments were chosen because they should not interfere with the multiplex amplicons during electrophoresis. The mPCR products were loaded onto a 96-well plate as follows: 23.7 μl sample loading solution (SLS; deionized formamide) plus 0.3 μl MM1000 plus 1 μl IC plus 1 μl mPCR amplicon. The running conditions were as follows: denaturation at 90°C for 120 s, injection for 60 s at 2 kV, and separation at 50°C and 5 kV for 100 min. The electropherograms obtained were analyzed with the GenomLab GeXP genetic analysis system, with baseline slope and relative peak height thresholds set at 10%. Peak coordinates corresponding to amplicon size (S) and peak intensity (I) were noted, and normalized ratios (R) between samples and IC peak intensities were calculated according to the dyes used for labeling.
Samples.
Each step in mPCR development was assessed with archived DNAs (extracted from endpoint cells or PBMCs) and circulating RNAs (extracted from virus supernatants or plasma samples) from viruses cultured in the laboratory or from clinical samples.
(i) In vitro samples.
We used the following four reference strains to validate the choice of primers and the first multiplexing steps: NL4.3 (HIV-1/M), RBF189 (HIV-1/O), ROD (HIV-2/A), and DIL (HIV-2/B). We further validated the mPCR criteria and interpretation rules on a larger panel of strains including 19 HIV-1/M, 12 HIV-1/O, and 13 HIV-2 strains and reflecting intragroup and intratype genetic diversities. For 21 strains, we were able to test both DNA and RNA, and for the others we tested only one type of material. We thus analyzed 32 DNA and 33 RNA extracts (see Table S2 in the supplemental material).
We assessed the ability of the mPCRs to detect several variants simultaneously in mixed genetic materials that simulated dual infections. For archived forms, we prepared five HIV-1/M+O and five HIV-1/M+HIV-2 mixtures from DNA samples for the evaluation of MMO2 and 10 HIV-1/M+O mixtures for the evaluation of MMO (see Table S3 in the supplemental material). The mixtures were based on the same input as that for single DNA samples (2 μl/PCR mixture). For circulating forms, we chose five HIV-1/M and five HIV-1/O strains and prepared five crossed ranges by mixing supernatants (100% plus 0%, 75% plus 25%, 50% plus 50%, 25% plus 75%, and 0% plus 100%) to simulate dual replicative infections (see Table S3). The 100% inputs were obtained by diluting supernatants in HIV-negative human plasma before extraction, to reach a VL of 105 copies/ml. The same method was applied to the five HIV-1/M–HIV-2 mixtures.
(ii) Clinical samples.
We assessed the clinical sensitivity and specificity of different variants of the mPCRs by challenging them with a panel of 31 HIV-1/M, 12 HIV-1/O, and 4 HIV-2 strains from clinical monoinfections. This involved the analysis of 15 DNA and 35 RNA extracts (see Table S4 in the supplemental material). Three DNA extracts from HIV-negative patients were used as controls and for the evaluation of background. Plasma VL was determined with an m2000SP/m2000RT system (Abbott, Rungis, France) for HIV-1/M and HIV-1/O and by an in-house technique for HIV-2 (45).
We then tested the mPCRs on eight samples obtained from patients identified in our laboratory by conventional molecular biology techniques [group M- and group O-specific (RT-)PCR followed by sequencing, as previously described (29)] as having a dual HIV-1/M+O infection or being infected with an HIV-1/MO recombinant (see Table 4).
TABLE 4.
mPCR-CE results for clinical monoinfections
| Sample material | MMO2 assay |
MMO assay |
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sensitivitya |
Specificity (%) |
Global detectiona,b | Sensitivitya |
Specificity (%) |
Global detectiona,b | |||||||
| HIV-1/M | HIV-1/O | HIV-2 | HIV-1/M | HIV-1/O | HIV-2 | HIV-1/M | HIV-1/O | HIV-1/M | HIV-1/O | |||
| DNA | 10/12 (83) | 3/3 (100) | 100 | 100 | 15/15 (100) | 7/12 (58) | 2/3 (67) | 100 | 100 | 15/15 (100) | ||
| RNA | 18/19 (95) | 11/12 (92) | 3/4 (75) | 100 | 100 | 100 | 34/35 (97) | 16/19 (84) | 9/12 (75) | 100 | 100 | 31/31 (100) |
Values shown are number of positive results/number of samples tested (%).
Detection of at least one gene for each target strain.
RESULTS
Assessment of mPCR-CE for detection of single variants in vitro.
After validating the mPCR-CE combination (see Fig. S3 in the supplemental material), we determined the sensitivities and specificities of the mPCRs with single DNA and RNA extracts corresponding to the 19 HIV-1/M, 12 HIV-1/O, and 13 HIV-2 strains from in vitro samples (see Table S2) used for the definition of validation criteria and interpretation rules (see the supplemental material). We carried out the following two analyses: a stringent analysis, in which we considered the detection of all target genes on the electropherogram for each strain, and a less stringent analysis based on the detection of at least one target gene for each strain.
(i) All-target detection.
For the MMO2 assay, we obtained a sensitivity of 91% for HIV-1/M, 91% for HIV-1/O, and 80% for HIV-2 by using DNA (Table 1). With RNA, this mPCR had a sensitivity of 100%, regardless of the group or type. Specificity, based on the absence of amplification of a genomic region from a type or group other than those targeted, was 100% for both DNA and RNA.
TABLE 1.
mPCR-CE results for single in vitro samples
| Sample material | MMO2 assay |
MMO assay |
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sensitivitya |
Specificity (%) |
Global detectiona,b | Sensitivitya |
Specificity (%) |
Global detectiona,b | |||||||
| HIV-1/M | HIV-1/O | HIV-2 | HIV-1/M | HIV-1/O | HIV-2 | HIV-1/M | HIV-1/O | HIV-1/M | HIV-1/O | |||
| DNA | 10/11 (91) | 10/11 (91) | 8/10 (80) | 100 | 100 | 100 | 32/32 (100) | 11/11 (100) | 11/11 (100) | 100 | 100 | 22/22 (100) |
| RNA | 11/11 (100) | 10/10 (100) | 12/12 (100) | 100 | 100 | 100 | 33/33 (100) | 11/11 (100) | 10/10 (100) | 100 | 80c | 21/21 (100) |
Values shown are number of positive results/number of samples tested (%).
Detection of at least one gene for each target strain.
Two HIV-1/M samples presented a peak corresponding to PR-O.
The sensitivity of the MMO assay on both DNA and RNA for both groups M and O was 100%. This mPCR had a specificity of 100% on DNA for both groups and specificities of 100% and 80% on RNA for groups M and O, respectively.
We used mixtures of DNA or RNA samples (crossed dilution panels) to mimic dual infections and to evaluate the ability of the mPCRs to detect such infections. For MMO2, the sensitivity with DNA was 80% for both HIV-1/M+O and HIV-1/M+HIV-2 mixtures (Table 2). This mPCR had a sensitivity of 100% with RNA, regardless of the mixture and the dilution. For MMO, the sensitivity was 90% with DNA and 100% with RNA, regardless of the mixture and the dilution.
TABLE 2.
mPCR-CE results for dual in vitro samples
| Sample material | MMO2 assay |
MMO assay |
|||
|---|---|---|---|---|---|
| Sensitivitya |
Global detectionb | Sensitivitya | Global detectionb | ||
| HIV-1/M+O | HIV-1+HIV-2 | HIV-1/M+O | |||
| DNA | 4/5 (80)c | 4/5 (80)c | 10/10 (100) | 9/10 (90)d | 10/10 (100) |
| RNA | 5/5 (100) | 5/5 (100) | 10/10 (100) | 5/5 (100) | 5/5 (100) |
Values shown are number of positive results/number of samples tested (%).
Detection of at least one gene for each target strain.
The detection of IN-M and IN-2A failed in one mixture each.
gp120-M detection failed in one mixture.
Focusing on genetic diversity regardless of the nature of the template, MMO2 detected 95% of HIV-1/M strains (1 strain from subtype CRF02 was not detected), 92% of HIV-1/O strains (1 strain from clade B was not detected), and 85% of HIV-2 strains (2 strains from group A were not detected) (Table 3). MMO detected 100% of both group M and O strains (Table 1). MMO2 had a specificity of 100% for each type and group, and MMO had a specificity of 100% for HIV-1/M and 89% for HIV-1/O.
TABLE 3.
Summary of mPCR-CE detection failures by strain
| Strain | Subtype, clade, or groupa | VLb (log10) | Failed MMO2 target |
Failed MMO target |
||||||
|---|---|---|---|---|---|---|---|---|---|---|
|
In vitro samples |
Clinical samples |
In vitro samples |
Clinical samples |
|||||||
| DNA | RNA | DNA | RNA | DNA | RNA | DNA | RNA | |||
| 5_M | CRF02 | INc | ||||||||
| 7_M | CRF01 | |||||||||
| 34_M | B | 3.2 | INc | MA, PR, IN,c gp41 | ||||||
| 35_M | B | 3.4 | MA, gp120, gp41 | |||||||
| 38_M | A1 | 3.9 | INc | MA, PR, gp120 | ||||||
| 40_M | B | 4.3 | MA | |||||||
| 24_M | CRF11 | 4.3 | MA,c gp120 | |||||||
| 42_M | B | 4.4 | MA | |||||||
| 25_M | B | 4.4 | IN | |||||||
| 26_M | B | 4.6 | gp41c | gp41c | ||||||
| 13_O | B | 3.1 | IN, gp41 | |||||||
| 7_O | B | 3.3 | INc | IN | ||||||
| 1_O | A | 3.7 | gp41 | |||||||
| 19_O | A | 4.1 | PR | |||||||
| 18_O | A | 4.7 | PR | |||||||
| 4_2 | A | INc | ||||||||
| 7_2 | A | 3.2 | IN | IN | ||||||
The viral subtype (HIV-1/M), clade (HIV-1/O), or group (HIV-2) is given.
Plasma viral loads for clinical RNA samples and corresponding plasma viral loads for clinical DNA samples.
Detection failure due to the cutoff ratio used.
(ii) Single-target detection.
Based on the detection of at least one target gene for each different strain, which should be enough to suspect a dual infection or infection with a recombinant due to the excellent specificity, both mPCRs detected single viruses and two-virus mixtures in 100% of cases (Table 1).
Assessment of mPCR-CE for detection of variants in clinical samples.
We assessed the mPCRs under clinical conditions by using DNAs and RNAs from samples of well-defined clinical monoinfections with 31 HIV-1/M, 12 HIV-1/O, and 4 HIV-2 strains (see Table S4 in the supplemental material).
(i) All-target detection.
No HIV-2 extract was available for testing of the MMO2 assay. This mPCR had a sensitivity with DNA of 83% for group M and 100% for group O. With RNA, it had a sensitivity of 95% for HIV-1/M, 92% for HIV-1/O, and 75% for HIV-2 and a specificity of 100% (Table 4).
With DNA, the MMO assay had sensitivities of 58% and 67% for group M and group O, respectively. With RNA, it had a sensitivity of 84% for group M and 75% for group O. It had a specificity of 100%.
Focusing on DNA samples, MMO2 failed to detect 2 of 12 HIV-1/M strains; MMO missed 5 of 12 HIV-1/M and 1 of 3 HIV-1/O strains (Table 3). In the absence of an intracellular DNA VL, we looked for a possible link between low levels of viral RNA in plasma and the lack of DNA detection. The three samples (34_M, 35_M, and 38_M) presenting multiple-gene detection failure with MMO were among those with the lowest VL (3.2, 3.4, and 3.9 log10 copies/ml, respectively) (Table 3; see Table S4 in the supplemental material), with no IN-M detection by either mPCR for sample 34_M due to the cutoff ratio used. The other two samples, 40_M and 42_M, with only one missed target (MA-M), had higher corresponding VL (4.3 and 4.4 log10 copies/ml) (Table 3).
With RNA samples, MMO2 missed 1 of 19 HIV-1/M strains, 1 of 12 HIV-1/O strains, and 1 of 4 HIV-2 strains, whereas MMO missed 3 of 19 HIV-1/M strains and 3 of 12 HIV-1/O strains (Table 3). Detection of HIV-1/M failed for three samples (24_M, 25_M, and 26_M), with VL between 4.3 and 4.6 log10 copies/ml, whereas six samples with correct detection of all the MMO target regions had lower VL. For sample 26_M (4.6 log10 copies/ml), gp41-M was not detected by either mPCR due to the cutoff ratio used. For HIV-1/O, detection failures were noted for one sample (13_O), with the lowest VL, for MMO2 and for two samples (7_O and 1_O) with low VL values (3.3 and 3.7 log10 copies/ml, respectively) and one sample with a higher VL (18_O; 4.7 log10 copies/ml) for MMO (Table 3).
(ii) Single-target detection.
Based on detection of at least one gene, MMO2 was able to detect variant monoinfections in 100% (15/15 samples) and 97% (34/35 samples) of DNA and RNA samples, respectively, and MMO detected 100% of cases for both materials (Table 4).
Testing of mPCR-CE on genuine dual infections and recombinants.
We tested the mPCR-CE assay with previously characterized samples from two patients with dual HIV-1/M+O infections and six patients infected with an HIV-1/MO recombinant, with or without the parental strains (Fig. 1). No HIV-1+HIV-2 samples were available.
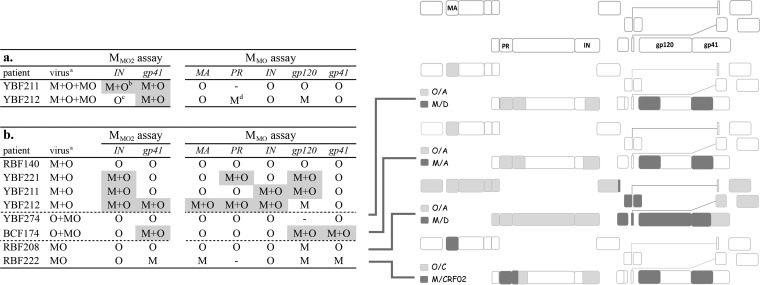
FIG 1.
(Left) mPCR-CE results for clinical HIV-M+O dual infections and/or infections with an HIV-MO recombinant. (a) DNA forms; (b) RNA forms. (Right) Partial or whole-genome recombination patterns associated with each recombinant. The HIV-1 genome is given above the recombinants, with the genomic targets of the mPCRs (MA, PR, IN, gp120, and gp41) indicated. a, abbreviations are as follows: M+O, HIV-1/M+O dual infection; MO, HIV-1/MO recombinant; M+O+MO, HIV-1/M+O dual infection plus HIV-1/MO recombinant; O+MO, HIV-1/O plus HIV-1/MO recombinant. b, detection of HIV-1/M and HIV-1/O; c, detection of HIV-1/O only; d, detection of HIV-1/M only.
With DNA (available only for YBF211 and YBF212), only MMO2 detected the dual infections based on at least one gene (Fig. 1a). For the YBF212 sample, MMO showed that M and O viral populations were present, but not for the same genomic region.
With RNA, the RBF140 sample corresponded to a dual HIV-1/M+O infection, but only HIV-1/O was detected by both multiplex PCRs. For YBF221, YBF211, and YBF212, both multiplex PCRs detected the presence of RNAs from both groups M and O for at least one gene (Fig. 1b).
For the detection of RNAs of the recombinant forms with or without the parental strains, only HIV-1/O was detected in sample YBF274, whereas both the HIV-1/O and HIV-1/M moieties of the recombinants were detected in samples BCF174, RBF208, and RBF222 by use of both mPCR-CE assays (Fig. 1b).
Taking a positive result for two strains for at least one gene to be suggestive of a replicative dual infection with or without a replicative recombinant, both mPCRs identified the same four possible dual infections among the six tested. If the results for all target genes were taken into account, then MMO2 identified two suspected replicative recombinants among the four tested, and MMO identified three of four.
Detection failures according to the nature of the variant.
The mPCR-CE method was globally evaluated on 115 extracts—47 DNA extracts and 68 RNA extracts—corresponding to 45 HIV-1/M, 21 HIV-1/O, and 16 HIV-2 isolates (in vitro viral strains and clinical samples from monoinfections).
Considering all the HIV-1/M data and the results of both mPCRs for all targets, regardless of the nature of the template (DNA or RNA), detection failure was observed for 20 (8%) of the 265 target genes tested (detection of five targets for the 53 HIV-1/M samples). These failures (Table 3) occurred mostly with MMO (16/20 failures) and with clinical DNA samples (14/20 failures) and involved the MA and IN regions in 11 of 20 cases. Detection failure was due to the ratio cutoff used in 7 of the 20 cases.
Seven of the 180 HIV-1/O targets (4%) and 3 of the 52 HIV-2 targets (6%) were missed. The nature of the material used (DNA or RNA) had no clear impact on mPCR. However, IN was again the target region accounting for the largest number of detection failures (3/7 failures for HIV-1/O and 3/3 failures for HIV-2). In two cases, detection failure resulted from the cutoff ratios used. The lack of detection was also strain dependent, occurring twice with 7_O and twice with 7_2.
DISCUSSION
Little is currently known about the true prevalence of HIV-1+HIV-2 and HIV-1/M+O dual infections or of HIV-1/MO recombinants. The detection of dual infections and recombinants involving major HIV variants is essential for accurate diagnosis and virological follow-up and for appropriate treatment. In this study, we aimed to develop a simple, cost-effective tool for the detection of such infections. Dual HIV-1+HIV-2 and HIV-1/M+O infections have different implications, as HIV-1/M+O infections have been linked to the generation of MO recombinants. We therefore designed two mPCR-CE assays: one (MMO2) for the rapid detection or confirmation of dual infections involving HIV-1/M, HIV-1/O, and HIV-2 and one (MMO) for the specific detection of HIV-1/MO recombinants. The principal challenge was achieving a sufficient sensitivity in terms of analytical (i.e., according to viral load) and clinical (i.e., to encompass the broad genetic diversity of HIV) performances for both circulating and integrated forms.
Development and evaluation of mPCR-CE on in vitro samples.
We first validated the tool on viral culture samples so as to cover a broad range of genetic diversity without being limited by the amount of material available. The global specificity of mPCR-CE was high, as was the sensitivity (80 to 100%), on both DNA and RNA and with both simple material and samples simulating dual infections. The sensitivity increased to 100% when we considered the detection of at least one or two species for a single genomic region as sufficient to suspect a dual infection or the presence of a recombinant form.
Assessment of mPCR-CE on clinical samples.
We further assessed this tool in clinical situations. We obtained a specificity of 100% for both mPCRs with both DNAs and RNAs from monoinfections and a high sensitivity with RNAs (75% to 95%) in the stringent analysis. Based on the lowest VL of the samples giving positive results (see the supplemental material), we estimated the limit of detection for the two multiplex PCRs to be 3.7 log10 copies/ml for HIV-1/M and close to 3 log10 copies/ml for HIV-1/O and HIV-2. A broader range of sensitivity was recorded for DNA (58% to 100%) due to the smaller amounts of viral DNA in cells (data not shown).
If we considered the detection of at least one target gene for each different strain, regardless of VL, for the RNA samples and DNA samples, then the sensitivity of MMO2 was 100% for HIV-1/M and HIV-2 and 97% for HIV-1/O, and that of MMO was 100% for both group M and group O.
Testing of mPCRs on dual infections and recombinants.
We then tested the mPCR strategy on real HIV-1/M+O dual infections and HIV-1/MO recombinants. Both mPCRs identified four of the six dual infections based on findings for at least one gene with an RNA template, and MMO2 identified both dual infections with a DNA template. MMO detected the two MO recombinants circulating alone and one of the two recombinants associated with the parental HIV-1/O strain for all the target genes compared to sequencing results. In its current state, MMO2 is not the test of choice for detecting MO recombinants, as demonstrated for the RBF208 recombinant, which was identified as an HIV-1/O strain (but consistent with its recombination pattern) rather than as a recombinant. MMO is the preferred test for the detection of HIV-1/MO.
Relevance and limitations of mPCR-CE.
Given the large number of PCRs required to target several genomic regions in various types and groups, multiplexing the primers was the best choice for simplifying sample handling and decreasing the experimental time and costs in routine use. However, the detection of mPCR amplicons made it necessary to consider several important issues, such as amplicon number and size. For example, it would have been difficult to carry out real-time PCR or DNA microarray analysis due to the genetic diversity of HIV, which would have made probe design very difficult, and due to high costs. Unlike the nonspecific detection techniques used in microbiology (46), CE has a high resolution and sensitivity and can be carried out with several dyes. This approach has already been validated and used by microbiologists for the diagnosis of respiratory diseases (47) and is simple to use with automated sequencers. However, standardization and rules for the analysis were required, and an internal control was used to validate the migration process and to normalize peak size. This approach, which is cheaper and less time-consuming than the 12 uniplex reactions required to obtain the same results, could be implemented in routine practice for the rapid detection of dual infections and recombinants after serological screening (in case of dual seroreactivities or based on discordance in seromolecular diagnosis algorithms).
The mPCRs performed well for the detection of at least one target, but these reactions were subject to four types of limitation. First, their sensitivity with DNA templates depended on the intracellular viral load. Second, some regions performed less well than others, with the IN and MA regions, in particular, accounting for 55% of failures for HIV-1/M samples. Third, clinical sensitivity is challenged by the considerable genetic diversity of HIV, as observed for the four strains giving false-negative results for the same region, regardless of the type of material and the mPCR used (Table 3). Finally, some failures (9/30 failures) were due to the analysis rules used to normalize the interpretation of electropherograms and to eliminate background noise (which was greater for DNA samples), resulting in a lower sensitivity in some cases.
In conclusion, our findings highlight the difficulties involved in the development of a perfect tool, with 100% specificity and sensitivity for all variants, for use with both DNA and RNA and for multiple targets. However, the original technique developed here can identify HIV-1+HIV-2 dual infections, HIV-1/M+O dual infections, and HIV-1/MO-related recombinants based on the detection of at least one gene of each variant sought. It is therefore potentially useful for the detection of these atypical infections (and of single variants in monoinfections), for evaluating their epidemiological impact and improving diagnosis and virological and therapeutic management, and for physiopathological studies.
Supplementary Material
ACKNOWLEDGMENTS
Funding was obtained from the Centre Hospitalier Universitaire de Rouen and the Institut National de Veille Sanitaire (InVS).
Footnotes
Supplemental material for this article may be found at http://dx.doi.org/10.1128/JCM.03222-15.
REFERENCES
- 1.Mansky LM, Temin HM. 1995. Lower in vivo mutation rate of human immunodeficiency virus type 1 than that predicted from the fidelity of purified reverse transcriptase. J Virol 69:5087–5094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Simon-Loriere E, Rossolillo P, Negroni M. 2011. RNA structures, genomic organization and selection of recombinant HIV. RNA Biol 8:280–286. doi: 10.4161/rna.8.2.15193. [DOI] [PubMed] [Google Scholar]
- 3.Plantier J-C, Leoz M, Dickerson JE, De Oliveira F, Cordonnier F, Lemée V, Damond F, Robertson DL, Simon F. 2009. A new human immunodeficiency virus derived from gorillas. Nat Med 15:871–872. doi: 10.1038/nm.2016. [DOI] [PubMed] [Google Scholar]
- 4.de Silva TI, Cotten M, Rowland-Jones SL. 2008. HIV-2: the forgotten AIDS virus. Trends Microbiol 16:588–595. doi: 10.1016/j.tim.2008.09.003. [DOI] [PubMed] [Google Scholar]
- 5.Mourez T, Simon F, Plantier J-C. 2013. Non-M variants of human immunodeficiency virus type 1. Clin Microbiol Rev 26:448–461. doi: 10.1128/CMR.00012-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Péré H, Roques P, Talla F, Meillo H, Charpentier C, Bélec L. 2015. Sustained virological failure in Cameroonese patient infected by HIV-1 group N evidenced by sequence-based genotyping assay. AIDS 29:1267–1269. doi: 10.1097/QAD.0000000000000704. [DOI] [PubMed] [Google Scholar]
- 7.Ayouba A, Akoua-Koffi C, Calvignac-Spencer S, Esteban A, Locatelli S, Li H, Li Y, Hahn BH, Delaporte E, Leendertz FH, Peeters M. 2013. Evidence for continuing cross-species transmission of SIVsmm to humans: characterization of a new HIV-2 lineage in rural Côte d'Ivoire. AIDS 27:2488–2491. doi: 10.1097/01.aids.0000432443.22684.50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Smith SM, Christian D, de Lame V, Shah U, Austin L, Gautam R, Gautam A, Apetrei C, Marx PA. 2008. Isolation of a new HIV-2 group in the US. Retrovirology 5:103. doi: 10.1186/1742-4690-5-103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gao F, Yue L, Robertson DL, Hill SC, Hui H, Biggar RJ, Neequaye AE, Whelan TM, Ho DD, Shaw GM. 1994. Genetic diversity of human immunodeficiency virus type 2: evidence for distinct sequence subtypes with differences in virus biology. J Virol 68:7433–7447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chen Z, Luckay A, Sodora DL, Telfer P, Reed P, Gettie A, Kanu JM, Sadek RF, Yee J, Ho DD, Zhang L, Marx PA. 1997. Human immunodeficiency virus type 2 (HIV-2) seroprevalence and characterization of a distinct HIV-2 genetic subtype from the natural range of simian immunodeficiency virus-infected sooty mangabeys. J Virol 71:3953–3960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Damond F, Worobey M, Campa P, Farfara I, Colin G, Matheron S, Brun-Vézinet F, Robertson DL, Simon F. 2004. Identification of a highly divergent HIV type 2 and proposal for a change in HIV type 2 classification. AIDS Res Hum Retroviruses 20:666–672. doi: 10.1089/0889222041217392. [DOI] [PubMed] [Google Scholar]
- 12.Cot MC, Poulain M, Delagneau JF, Peeters M, Brun-Vézinet F. 1988. Dual HIV-1 and HIV-2 infection in West Africa supported by synthetic peptide analysis. AIDS Res Hum Retroviruses 4:239–241. doi: 10.1089/aid.1988.4.239. [DOI] [PubMed] [Google Scholar]
- 13.Evans LA, Moreau J, Odehouri K, Seto D, Thomson-Honnebier G, Legg H, Barboza A, Cheng-Mayer C, Levy JA. 1988. Simultaneous isolation of HIV-1 and HIV-2 from an AIDS patient. Lancet ii:1389–1391. [DOI] [PubMed] [Google Scholar]
- 14.George JR, Ou CY, Parekh B, Brattegaard K, Brown V, Boateng E, De Cock KM. 1992. Prevalence of HIV-1 and HIV-2 mixed infections in Côte d'Ivoire. Lancet 340:337–339. doi: 10.1016/0140-6736(92)91406-X. [DOI] [PubMed] [Google Scholar]
- 15.Rouet F, Ekouevi DK, Inwoley A, Chaix M-L, Burgard M, Bequet L, Viho I, Leroy V, Simon F, Dabis F, Rouzioux C. 2004. Field evaluation of a rapid human immunodeficiency virus (HIV) serial serologic testing algorithm for diagnosis and differentiation of HIV type 1 (HIV-1), HIV-2, and dual HIV-1–HIV-2 infections in West African pregnant women. J Clin Microbiol 42:4147–4153. doi: 10.1128/JCM.42.9.4147-4153.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Plantier J-C, Lemée V, Dorval I, Gueudin M, Braun J, Hutin P, Ruffault A, Simon F. 2004. HIV-1 group M superinfection in an HIV-1 group O-infected patient. AIDS 18:2444–2446. [PubMed] [Google Scholar]
- 17.Landman R, Damond F, Gerbe J, Brun-Vézinet F, Yeni P, Matheron S. 2009. Immunovirological and therapeutic follow-up of HIV-1/HIV-2-dually seropositive patients. AIDS 23:426–428. doi: 10.1097/QAD.0b013e328321305a. [DOI] [PubMed] [Google Scholar]
- 18.Günthard HF, Huber M, Kuster H, Shah C, Schüpbach J, Trkola A, Böni J. 2009. HIV-1 superinfection in an HIV-2-infected woman with subsequent control of HIV-1 plasma viremia. Clin Infect Dis 48:e117–e120. doi: 10.1086/598987. [DOI] [PubMed] [Google Scholar]
- 19.Janssens W, Fransen K, Loussert-Ajaka I, Heyndrickx L, Ivens T, Eberle J, Nkengasong J. 1995. Diagnosis of HIV-1 group O infection by polymerase chain reaction. Lancet 346:451–452. [DOI] [PubMed] [Google Scholar]
- 20.Takehisa J, Zekeng L, Ido E, Mboudjeka I, Moriyama H, Miura T, Yamashita M, Gürtler LG, Hayami M, Kaptué L. 1998. Various types of HIV mixed infections in Cameroon. Virology 245:1–10. doi: 10.1006/viro.1998.9141. [DOI] [PubMed] [Google Scholar]
- 21.van der Kuyl AC, Cornelissen M. 2007. Identifying HIV-1 dual infections. Retrovirology 4:67. doi: 10.1186/1742-4690-4-67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ekouevi DK, Balestre E, Coffie PA, Minta D, Messou E, Sawadogo A, Minga A, Sow PS, Bissagnene E, Eholie SP, Gottlieb GS, Dabis F, IeDEA West Africa Collaboration. 2013. Characteristics of HIV-2 and HIV-1/HIV-2 dually seropositive adults in West Africa presenting for care and antiretroviral therapy: the IeDEA-West Africa HIV-2 Cohort Study. PLoS One 8:e66135. doi: 10.1371/journal.pone.0066135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Vessière A, Rousset D, Kfutwah A, Leoz M, Depatureaux A, Simon F, Plantier J-C. 2010. Diagnosis and monitoring of HIV-1 group O-infected patients in Cameroun. J Acquir Immune Defic Syndr 53:107–110. doi: 10.1097/QAI.0b013e3181b97ec1. [DOI] [PubMed] [Google Scholar]
- 24.Vergne L, Bourgeois A, Mpoudi-Ngole E, Mougnutou R, Mbuagbaw J, Liegeois F, Laurent C, Butel C, Zekeng L, Delaporte E, Peeters M. 2003. Biological and genetic characteristics of HIV infections in Cameroon reveals dual group M and O infections and a correlation between SI-inducing phenotype of the predominant CRF02_AG variant and disease stage. Virology 310:254–266. doi: 10.1016/S0042-6822(03)00167-3. [DOI] [PubMed] [Google Scholar]
- 25.Peeters M, Liegeois F, Torimiro N, Bourgeois A, Mpoudi E, Vergne L, Saman E, Delaporte E, Saragosti S. 1999. Characterization of a highly replicative intergroup M/O human immunodeficiency virus type 1 recombinant isolated from a Cameroonian patient. J Virol 73:7368–7375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Takehisa J, Zekeng L, Ido E, Yamaguchi-Kabata Y, Mboudjeka I, Harada Y, Miura T, Kaptu L, Hayami M. 1999. Human immunodeficiency virus type 1 intergroup (M/O) recombination in Cameroon. J Virol 73:6810–6820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Yamaguchi J, Bodelle P, Vallari AS, Coffey R, McArthur CP, Schochetman G, Devare SG, Brennan CA. 2004. HIV infections in northwestern Cameroon: identification of HIV type 1 group O and dual HIV type 1 group M and group O infections. AIDS Res Hum Retroviruses 20:944–957. doi: 10.1089/aid.2004.20.944. [DOI] [PubMed] [Google Scholar]
- 28.Vessière A, Leoz M, Brodard V, Strady C, Lemée V, Depatureaux A, Simon F, Plantier J-C. 2010. First evidence of a HIV-1 M/O recombinant form circulating outside Cameroon. AIDS 24:1079–1082. doi: 10.1097/QAD.0b013e3283355659. [DOI] [PubMed] [Google Scholar]
- 29.Ngoupo PA, Sadeuh-Mba SA, De Oliveira F, Ngono V, Ngono L, Tchendjou P, Penlap V, Mourez T, Njouom R, Kfutwah A, Plantier J-C. 2016. First evidence of transmission of an HIV-1 M/O intergroup recombinant virus. AIDS 30:1–8. doi: 10.1097/QAD.0000000000000880. [DOI] [PubMed] [Google Scholar]
- 30.Plantier JC, Djemai M, Lemee V, Reggiani A, Leoz M, Burc L, Vessiere A, Rousset D, Poveda JD, Henquell C, Gautheret-Dejean A, Barin F. 2009. Census and analysis of persistent false-negative results in serological diagnosis of human immunodeficiency virus type 1 group O infections. J Clin Microbiol 47:2906–2911. doi: 10.1128/JCM.00602-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Gautheret-Dejean A, Bocobza J, Brunet S, Damond F, Plantier J-C, Barin F. 2015. Performance of rapid tests for discrimination between HIV-1 and/or HIV-2 infections. J Med Virol 87:2061–2066. doi: 10.1002/jmv.24282. [DOI] [PubMed] [Google Scholar]
- 32.Gueudin M, Plantier J-C, Lemée V, Schmitt MP, Chartier L, Bourlet T, Ruffault A, Damond F, Vray M, Simon F. 2007. Evaluation of the Roche Cobas TaqMan and Abbott RealTime extraction-quantification systems for HIV-1 subtypes. J Acquir Immune Defic Syndr 44:500–505. doi: 10.1097/QAI.0b013e31803260df. [DOI] [PubMed] [Google Scholar]
- 33.Gueudin M, Leoz M, Lemée V, De Oliveira F, Vessière A, Kfutwah A, Plantier J-C. 2012. A new real-time quantitative PCR for diagnosis and monitoring of HIV-1 group O infection. J Clin Microbiol 50:831–836. doi: 10.1128/JCM.05669-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Delarue S, Didier E, Damond F, Ponscarme D, Brengle-Pesce K, Resche-Rigon M, Vray M, Gueudin M, Simon F. 2013. Highly sensitive plasma RNA quantification by real-time PCR in HIV-2 group A and group B infection. J Clin Virol 58:461–467. doi: 10.1016/j.jcv.2013.08.003. [DOI] [PubMed] [Google Scholar]
- 35.Avettand-Fenoel V, Damond F, Gueudin M, Matheron S, Melard A, Collin G, Descamps D, Chaix ML, Rouzioux C, Plantier JC, ANRS-CO5 HIV-2 and ANRS-AC11 Quantification Working Group . 2014. New sensitive one-step real-time duplex PCR method for group A and B HIV-2 RNA load. J Clin Microbiol 52:3017–3022. doi: 10.1128/JCM.00724-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Descamps D, Collin G, Letourneur F, Apetrei C, Damond F, Loussert-Ajaka I, Simon F, Saragosti S, Brun-Vézinet F. 1997. Susceptibility of human immunodeficiency virus type 1 group O isolates to antiretroviral agents: in vitro phenotypic and genotypic analyses. J Virol 71:8893–8898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Tchounga BK, Inwoley A, Coffie PA, Minta D, Messou E, Bado G, Minga A, Hawerlander D, Kane C, Eholie SP, Dabis F, Ekouevi DK, WADA Collaboration. 2014. Re-testing and misclassification of HIV-2 and HIV-1&2 dually reactive patients among the HIV-2 cohort of the West African Database to evaluate AIDS collaboration. J Int AIDS Soc 17:19064. doi: 10.7448/IAS.17.1.19064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Simon F, Souquière S, Damond F, Kfutwah A, Makuwa M, Leroy E, Rouquet P, Berthier JL, Rigoulet J, Lecu A, Telfer PT, Pandrea I, Plantier JC, Barré-Sinoussi F, Roques P, Müller-Trutwin MC, Apetrei C. 2001. Synthetic peptide strategy for the detection of and discrimination among highly divergent primate lentiviruses. AIDS Res Hum Retroviruses 17:937–952. doi: 10.1089/088922201750290050. [DOI] [PubMed] [Google Scholar]
- 39.Barin F, Plantier J-C, Brand D, Brunet S, Moreau A, Liandier B, Thierry D, Cazein F, Lot F, Semaille C, Desenclos J-C. 2006. Human immunodeficiency virus serotyping on dried serum spots as a screening tool for the surveillance of the AIDS epidemic. J Med Virol 78(Suppl 1):S13–S18. doi: 10.1002/jmv.20600. [DOI] [PubMed] [Google Scholar]
- 40.Ariën KK, Abraha A, Quiñones-Mateu ME, Kestens L, Vanham G, Arts EJ. 2005. The replicative fitness of primary human immunodeficiency virus type 1 (HIV-1) group M, HIV-1 group O, and HIV-2 isolates. J Virol 79:8979–8990. doi: 10.1128/JVI.79.14.8979-8990.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Manigart O, Courgnaud VR, Sanou O, Val A, Nagot DN, Meda N, Delaporte E, Peeters M, Van de Perre P. 2004. HIV-1 superinfections in a cohort of commercial sex workers in Burkina Faso as assessed by an autologous heteroduplex mobility procedure. AIDS 18:1645–1651. doi: 10.1097/01.aids.0000131333.30548.db. [DOI] [PubMed] [Google Scholar]
- 42.Hoelscher M, Dowling WE, Sanders-Buell E, Carr JK, Harris ME, Thomschke A, Robb ML, Birx DL, McCutchan FE. 2002. Detection of HIV-1 subtypes, recombinants, and dual infections in east Africa by a multi-region hybridization assay. AIDS 16:2055–2064. doi: 10.1097/00002030-200210180-00011. [DOI] [PubMed] [Google Scholar]
- 43.Pacold M, Smith D, Little S, Cheng PM, Jordan P, Ignacio C, Richman D, Pond SK. 2010. Comparison of methods to detect HIV dual infection. AIDS Res Hum Retroviruses 26:1291–1298. doi: 10.1089/aid.2010.0042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Routh A, Johnson JE. 2014. Discovery of functional genomic motifs in viruses with ViReMa—a virus recombination mapper—for analysis of next-generation sequencing data. Nucleic Acids Res 42:e11. doi: 10.1093/nar/gkt916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Damond F, Benard A, Balotta C, Böni J, Cotten M, Duque V, Ferns B, Garson J, Gomes P, Gonçalves F, Gottlieb G, Kupfer B, Ruelle J, Rodés B, Soriano V, Wainberg M, Taieb A, Matheron S, Chene G, Brun-Vézinet F, ACHI(E)V(2E) Study Group. 2011. An international collaboration to standardize HIV-2 viral load assays: results from the 2009 ACHI(E)V(2E) quality control study. J Clin Microbiol 49:3491–3497. doi: 10.1128/JCM.02389-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Rademaker JL, Hoste B, Louws FJ, Kersters K, Swings J, Vauterin L, Vauterin P, de Bruijn FJ. 2000. Comparison of AFLP and rep-PCR genomic fingerprinting with DNA-DNA homology studies: Xanthomonas as a model system. Int J Syst Evol Microbiol 50:665–677. doi: 10.1099/00207713-50-2-665. [DOI] [PubMed] [Google Scholar]
- 47.Dabisch-Ruthe M, Vollmer T, Adams O, Knabbe C, Dreier J. 2012. Comparison of three multiplex PCR assays for the detection of respiratory viral infections: evaluation of xTAG respiratory virus panel fast assay, RespiFinder 19 assay and RespiFinder SMART 22 assay. BMC Infect Dis 12:163. doi: 10.1186/1471-2334-12-163. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.