Abstract
Hydrophobic signal sequences target secretory polypeptides to a protein-conducting channel formed by a heterotrimeric membrane protein complex, the prokaryotic SecY or eukaryotic Sec61 complex. How signal sequences are recognized is poorly understood, particularly because they are diverse in sequence and length. Structures of the inactive channel show that the largest subunit, SecY or Sec61α, consists of two halves that form an hourglass-shaped pore with a constriction in the middle of the membrane and a lateral gate that faces lipid1-10. The cytoplasmic funnel is empty, while the extracellular funnel is filled with a plug domain. In bacteria, the SecY channel associates with the translating ribosome in co-translational translocation, and with the SecA ATPase in post-translational translocation 11. How a translocating polypeptide inserts into the channel is uncertain, as cryo-EM structures of the active channel have a relatively low resolution (~10Å) or are of insufficient quality 6-8. Here we report a crystal structure of the active channel, assembled from SecY complex, the SecA ATPase, and a segment of a secretory protein fused into SecA. The translocating protein segment inserts into the channel as a loop, displacing the plug domain. The hydrophobic core of the signal sequence forms a helix that sits in a groove outside the lateral gate, while the following polypeptide segment intercalates into the gate. The C-terminal section of the polypeptide loop is located in the channel, surrounded by residues of the pore ring. Thus, during translocation, the hydrophobic segments of signal sequences, and probably bilayer-spanning domains of nascent membrane proteins, exit the lateral gate and dock at a specific site that faces the lipid phase.
To determine the structure of an active SecY channel, we initially generated in Escherichia coli a translocation intermediate, consisting of SecA, SecY complex, and a short segment of a secretory protein fused to a fast-folding green fluorescent protein (GFP) (Extended Data Fig.1a). Although this complex could be purified 12, it failed to crystallize. We therefore reduced the complexity of the system by fusing a short segment of a secretory protein directly into SecA. The segment contains the signal sequence of OmpA and a short polypeptide following it, and was inserted into the tip of the two-helix finger of SecA (SecA-OAIns; Fig. 1a; Extended Data Fig. 1b), because the finger was seen to protrude into the cytoplasmic cavity of SecY in a structure of SecA/SecY complex lacking a translocation substrate 9. Using E. coli SecA-OAIns and E. coli SecY complex, the inserted secretory protein segment was indeed translocated to the periplasm in E. coli, as demonstrated by the formation of a disulfide bridge between a cysteine introduced C-terminally of the signal sequence and a cysteine placed into the plug domain of SecY (Extended Data Fig.2a). This disulfide bridge formed spontaneously, i.e. without addition of an exogenous oxidant. The introduction of Gln residues into the hydrophobic core of the signal sequence abolished disulfide bridge formation (Extended Data Fig.2b), demonstrating that an intact signal sequence is required for translocation of the polypeptide segment. Similar results were obtained with Bacillus subtilis SecA-OAIns and Geobacillus thermodenitrificans SecYE (Extended Data Fig.2c), a complex of increased thermostability that is functional in E. coli. After optimization (Extended Data Fig. 2d-f), the construct chosen for crystallization contained 49 residues inserted into the two-helix finger of B. subtilis SecA, with a cysteine at position +7 in the region following the signal sequence of 20 residues. Channel insertion of the secretory protein segment was similar to that observed with the physiological system, containing wild type SecA and a GFP-fusion to a secretory protein fragment (Extended Data Fig.1), except that the latter requires an additional polypeptide segment to span the SecA molecule. Thus, our simplified system is a faithful mimic of normal initiation of protein translocation. Binding of SecA to the SecY complex seems to be sufficient to cause polypeptide chain insertion into the channel, similar to how ribosome binding allows nascent chain insertion in cotranslational translocation 13. In our system, disulfide crosslinking at the periplasmic side made channel insertion irreversible.
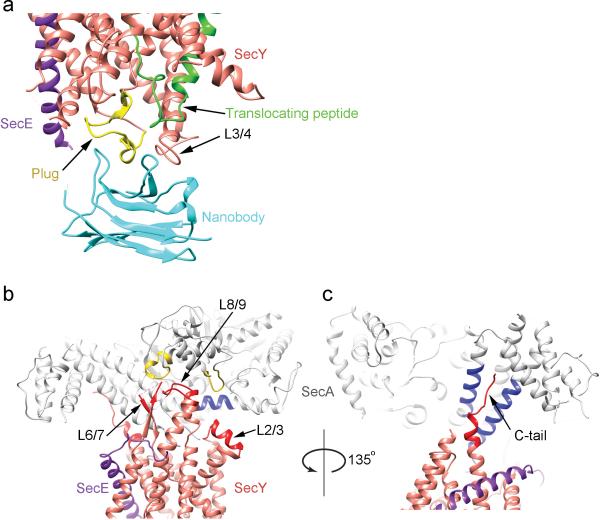
Figure 1. Architecture of the active SecY channel.
a, A secretory protein segment was inserted into the two-helix finger of the SecA ATPase (SecA-OAIns). The segment contains a linker (grey), the signal sequence of OmpA, consisting of the N-, H-, and C-regions (in red, black, and purple letters, respectively), and a region (in light green) that includes a unique cysteine (yellow star). Residues in the signal sequence are numbered backwards from the cleavage site. The fused segment inserts into the SecY channel in vivo and spontaneously forms a disulfide bridge with a cysteine in the plug. This complex was used for structure determination. b, Ribbon diagram of the complex, viewed from the side. The numbers refer to TMs of SecY. The lines indicate the membrane boundaries. A nanobody was used for crystallization.
The disulfide-bridged complex of B. subtilis SecA-OAIns and G. thermodenitrificans SecYE was purified and crystallized in the presence of ADP and BeFx (Extended Data Fig. 3), conditions that lock SecA into a conformation close to its ATP-bound state and maximize SecA's affinity for the channel 9,14. The diffraction of the crystals was improved by the use of single-domain antibody fragments (nanobodies), raised against G. thermodenitrificans SecYE and selected for binding to periplasmic loops of the SecA-OAIns/SecYE complex, and by soaking crystals with a Ta6Br12 metal ion cluster. The structure was determined from multi-wavelength anomalous diffraction (MAD) data obtained with a crystal that diffracted to a resolution of 3.70 Å along one axis and 4.48 Å along the other two (Fig. 1b; Extended Data Fig.4; Extended Data Table 1). An initial experimental electron density map had a resolution of ~5.5Å. This map was improved by density modification and molecular replacement using higher resolution structures of SecA, SecYE, and the nanobody, followed by cycles of model building and refinement. Inclusion of the model-refined Ta6Br12 clusters as the resolved heavy atom substructure for recalculation of MAD phases did not further improve the map. Nevertheless, the final map allowed the unambiguous placement of all TMs of SecY and SecE and of many other regions. The translocating polypeptide segment could be built into the map without model bias (Extended Data Fig. 4c). As expected, the nanobody bound to the periplasmic side of SecY, interacting with both the plug and the loop between TMs 3 and 4 (Fig. 1b; Extended Data Fig.5a).
The structure of the active channel shows that SecA undergoes relatively small conformational changes compared to a Thermotoga maritima complex lacking a translocating chain 9 (Extended Data Fig.6). SecA binds to the cytoplasmic loop between TMs 8 and 9 and the C-terminal tail of SecY (Extended Data Fig. 5b, c). It probably binds only weakly to the loop between TMs 6 and 7, as its tip is disordered. In contrast to the ribosome 6-8, SecA also binds to the N-terminal half of SecY, i.e. the loop between TMs 2 and 3 (Extended Data Fig.5b). Thus, at least in its ATP-bound state, SecA prevents large relative movements of the two halves of SecY.
SecY also undergoes relatively small changes, except at the lateral gate (Fig. 2). Compared with the idle Methanocaldococcus jannaschii or Thermus thermophilus channels 1,3, only TM7 (M.j.) or TM7 and TM8 (T.t.) significantly shift their positions (Fig. 2a; Extended Data Figs. 7a). Compared to the SecA-bound T. maritima channel 9, the periplasmic ends of TMs 3 and 7 move towards each other and TM 7 tilts by 10° relative to the plane of the membrane (Fig. 2b), changes that generate a pocket for the signal sequence (see below). In both structures, the lateral gate is partially open (compare Figs. 2a and 2b).
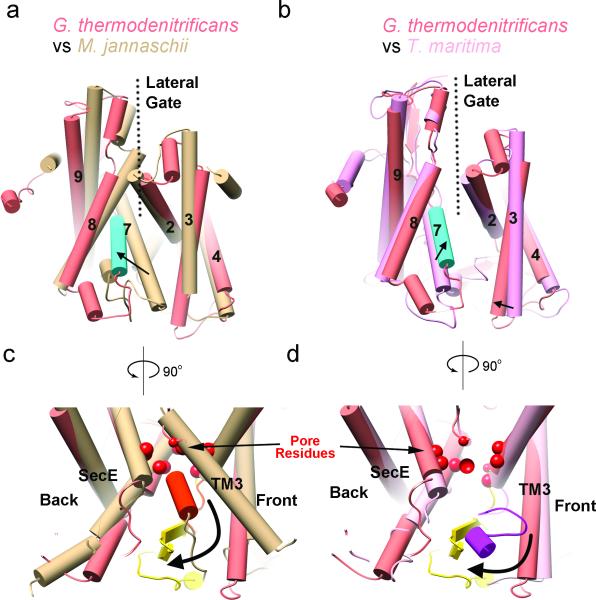
Figure 2. Conformational changes of the SecY channel.
a, Comparison of the lateral gate of the active G. thermodenitrificans channel (salmon, except for TM7 in cyan) with the closed SecY channel of M. jannaschii (in tan). The largest changes are indicated by arrows. The translocating peptide segment is omitted for clarity. b, As in a, but a comparison with that of the SecA-bound T. maritima channel lacking a translocating polypeptide (pink). c, Comparison of the plugs (orange for idle M. jannaschii channel and yellow for the active G. thermodenitrificans channel). Pore residues are shown as red spheres. d, As in c, but comparison with the inactive T. maritima channel (plug in magenta).
In the active G. thermodenitrificans SecY channel, the plug consists of two β-strands and therefore differs from the α-helical structures observed in other species 1,4,5,9 (Extended Data Fig. 8a). Such variability is consistent with the fact that the amino acid sequence of the plug region is least conserved 1, and that plug deletions cause neighboring polypeptide regions to form new plug domains 15. Different plug structures can probably be tolerated, as long as they fill the extracellular cavity of the channel, so that the closed state of the channel is stabilized and small molecules cannot pass through it. However, it is possible that the plug has different conformations in the closed and active channels.
Whereas the plug is close to the central constriction in the closed M. jannaschii channel 1, in the active G. thermodenitrificans channel it moves to the periplasmic side and towards the back of the channel, away from the lateral gate (Fig. 2c). The plug comes close to the TM of SecE, consistent with disulfide crosslinking experiments 16,17. In a SecA-SecY structure lacking a translocation substrate 9, the plug moves a smaller distance and towards the front (Fig. 2d), partially sealing the opened lateral gate (Extended Data Fig. 8b). In an intact membrane, this would prevent surrounding lipid molecules from moving through the lateral gate into the extracellular cavity. The plug is likely flexible in the active channel, but in our crystal structure, it is confined both by the disulfide bond to the translocating chain and by the interaction with the nanobody (Extended Data Fig.5a). Indeed, in a 6.5Å resolution structure determined without nanobody, the plug is shifted further towards the back, although there are otherwise only small differences (Extended Data Fig. 9a).
The signal sequence of the secretory protein segment forms a helix that is tilted ~45° relative to the plane of the membrane (Fig. 3a). The positively charged N-terminus of the signal sequence (N-region) is in the same plane as hydrophilic residues of SecY. In an intact membrane, the N-region could interact with the negatively charged head groups of the phospholipid bilayer. This interaction may retain the N-terminus on the cytoplasmic side of the membrane while the C-terminal end of the signal sequence moves through the channel, resulting in loop insertion of the translocating polypeptide.
Figure 3. The polypeptide in the channel.
a, Cut-away side view of the translocating polypeptide. The N-, H-, and C-regions of the signal sequence are indicated. b, View perpendicular to the axis of the signal sequence helix. c, Side view showing the translocating polypeptide together with a surface representation of the channel, with hydrophilic and hydrophobic residues in blue and orange, respectively. d, Side view of a cut through a surface representation model, with the main chain of the translocating chain shown as blue sticks. The residues located in the pore ring (red) are indicated. The plug is in orange. e, Top view of a slab, showing pore residues and Gly19 of the translocating chain as stick and balls with a mesh surface representation. Pore residues displaced in the active channel are in yellow. f, As in e, but with the closed M. jannaschii channel. I75 and I170 correspond to V74 and L179 in G. thermodenitrificans, respectively.
The hydrophobic core of the signal sequence (H-region; residues −17 to −6 of the original OmpA sequence; Fig. 1a) sits in a groove outside the lateral gate of SecY and forms a helix that runs almost parallel to TM2 (Fig. 3a, b). Some residues make van der Waals contacts with hydrophobic amino acids in TM2 (Extended Data Fig. 7c), but most face detergent (Fig. 3b), and would be in contact with hydrocarbon chains of phospholipids in an intact membrane.
The C-terminal region of the signal sequence (C-region; residues −5 to −1; Fig. 1a) replaces the periplasmic end of TM7 in the closed M. jannaschii channel (Extended Data Fig. 7b). This segment forms a distorted, amphipathic α-helix that is intercalated between TM7 and TM3 into the periplasmic side of the lateral gate (Fig. 3a,c). In our model, the side chains of Thr(−5) and Ser(−2) point into the periplasmic cavity previously occupied by the plug in the closed channel (Extended Data Fig. 7d). The hydrophobic residues Val(−4) and Ala(−3) face detergent/lipid. The lateral gate is thus sealed by the C-region from surrounding lipid molecules, which could otherwise pass through a large opening generated by the displacement of the plug from the front (Figs. 3c). After signal sequence cleavage, the periplasmic parts of TMs 7 and 3 probably move towards each other and seal the lateral gate.
The hydrophilic polypeptide segment following the signal sequence adopts a partially extended conformation with a loop in the periplasmic cavity, centered on the cysteine used for crosslinking to the plug. The signal sequence cleavage site is located inside the channel (Fig. 3c), likely inaccessible to the periplasmically disposed active site of signal peptidase 18. Thus, at some point during translocation, the C-region of the signal sequence probably must adopt a more extended conformation, a change also suggested by experiments with synthetic signal peptides 19,20.
The polypeptide chain inside the channel is perpendicular to the plane of the membrane (Fig. 3a). The two strands of the hairpin formed by the translocating polypeptide do not interact with one another, so that during translocation the C-terminal part of the hairpin could move unimpeded through the center of the channel. Our model places residue Gly(+19), or one of the neighboring residues (Ala18 or Gly20), of the translocating polypeptide inside the pore ring (Fig. 3d). The density around Gly19 is particularly strong, indicating that this segment is confined by the surrounding four pore ring residues (Ile78, Ile183, Ile275, Ile404). The ring is wider than in the idle M. jannaschii or SecA-bound T. maritima channels (diameters 8.8Å versus 5.6Å or 6.6Å, respectively). Crystallization may have favored the presence of small amino acids in the pore, minimizing its expansion by the presence of a translocating chain. However, even a small increase in pore diameter would allow the passage of amino acids with larger side chains. The pore ring residues fit snugly around the translocating polypeptide (Fig. 3e), confirming that they form a “gasket” that maintains the permeability barrier for ions and other small molecules during translocation 21. Consistent with disulfide crosslinking experiments 22, only pore ring residues contact the translocating chain (Fig. 3d). Thus, the hourglass-shape of the channel minimizes interactions with the translocation substrate, facilitating its movement through the channel.
The pore ring of the idle M. jannaschii channel contains two additional residues (Ile75 and Ile170; Fig. 3f). In the active channel, the corresponding residues (Val74, Leu179) are displaced (Fig. 3e). The pore “ring” is thus open at the lateral gate between Ile78 in TM2 and Ile275 in TM7 (Fig. 3e). These features suggest that a translocating polypeptide segment continuously encounters the hydrocarbon chains of surrounding lipids; when sufficiently hydrophobic, the segment will partition into the lipid phase and become a TM domain of a membrane protein 23,24.
Our crystal structure likely reflects the physiological situation of a translocating polypeptide. Five of the seven residues of the N-terminal linker are invisible and thus likely flexible, allowing unrestricted interaction of the signal sequence with the channel. In addition, most polypeptide segments following the signal sequence are in a relaxed conformation, unconstrained by fusion to SecA, or the disulfide bridge to the plug. The disulfide bridge helps to stabilize the signal sequence in the channel, but it likely does not lead to gross distortions, because the plug is mobile and the disulfide bridge is formed spontaneously in vivo.
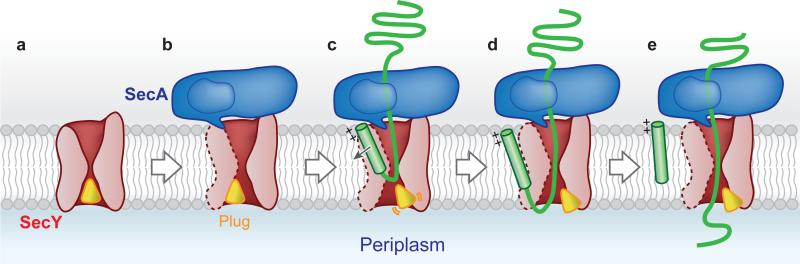
The crystal structure of the active channel leads to a refined model for post-translational protein translocation in bacteria (Fig. 4). The SecY channel is initially in the idle state, with the plug in the center and the lateral gate closed (Fig. 4a). Binding of SecA primes the channel for the arrival of a secretory protein precursor: the lateral gate is partially opened, the pore ring widened, and the plug domain moved towards the front (Fig. 4b). Next, the secretory protein inserts into the channel as a loop, with the C-terminal section of the polypeptide hairpin in the pore proper, surrounded by pore ring residues (Figs. 4c,d). During subsequent cycles of ATP hydrolysis, SecA uses a “push-and-slide” mechanism to move the C-terminal part of the polypeptide loop through the pore 14. Eventually, the signal sequence is cleaved by signal peptidase (Fig. 4e).
Figure 4. Scheme of SecA-mediated protein translocation.
Stage ‘a’ corresponds to the closed channel 1-3, stage ‘b” to the structure of the inactive SecA-SecY complex 9, stage ‘c’ represents an intermediate to stage ‘d’, which corresponds to the structure of the active channel. Stage ‘e’ is attained after signal sequence cleavage. The translocating polypeptide is shown in green and the signal sequence as a green cylinder. The lateral gate of the channel is shown as a broken line on the left. The clamp of SecA is indicated.
During loop insertion, the H-region of the signal sequence moves through the cytoplasmic part of the lateral gate and ends up in a hydrophobic groove on the outside, while the following hydrophilic segment crosses the lateral opening of the pore ring. A signal sequence might move through a partially open gate in an extended conformation, or it could move through a widened gate as a preformed helix. The latter possibility is suggested by a ~10Å-resolution EM structure of a ribosome/nascent chain/channel complex, in which a signal sequence helix was seen inside the lateral gate 6, probably prevented from exit by a disulfide bridge between a cysteine immediately following the signal sequence and a cysteine in the plug. The groove on the outside of the lateral gate site appears to be a general binding site for hydrophobic sequences, as indicated by a 8.5Å resolution structure of a complex in which the OmpA signal sequence was replaced by that of DsbA (Extended Data Fig. 9b,c), and EM structures of ribosome-channel complexes, in which TMs of nascent membrane proteins are located at about the same position 7,25 (Extended Data Fig. 7e). Like TMs, signal sequences appear to be recognized mostly by lipid partitioning, consistent with their ability to be crosslinked to lipids 26,27, and the correlation between the partitioning of synthetic peptides into hydrophobic solvents and their function as signal sequences in vivo 28. Nevertheless, amino acid interactions with TM2 of the channel may contribute to the recognition of signal sequences. This may explain why some amino acids occur more frequently than others in signal sequences, even when they have about the same hydrophobicity: leucine is preferred over isoleucine and valine 29, perhaps because its extended side chain can make tighter van der Waals contacts with residues in TM2. Whereas signal sequences of translocating secretory proteins would tend to stay in the binding pocket until they are cleaved off, the more hydrophobic TMs of membrane proteins could move away once the connecting loop to the polypeptide segment inside the channel pore attains adequate length.
While this paper was under review, a cryo-EM structure was published describing an active ribosome-bound mammalian Sec61 channel containing a short secretory polypeptide segment 30. The authors conclude that the signal sequence helix replaces TM2 of Sec61α in the idle channel, implying that the signal sequence is intercalated into the lateral gate and raising the possibility that the conformational changes differ greatly from those in our system. However, a superposition of the two active channels based on secondary structure matching shows that they are actually very similar, with only moderate differences at the periplasmic/luminal ends of some TMs (Extended Data Fig. 7f). Importantly, in both cases the signal sequence helix docks to the same site outside the lateral gate and runs parallel to TM2 (Extended Data Fig. 7f). Thus, regardless of the organism and mode of translocation, lipid partitioning appears to be the major mechanism by which signal sequences are recognized.
METHODS
Protein translocation and disulfide crosslinking assays
To test in vivo OmpA-GFP translocation by E. coli SecA and E. coli SecY complex (three-component system), E. coli strain EP52 or EP62 (Δrmf ΔompT secY-CBP) 12 was transformed with pACYC-SecYEG expressing E. coli SecYEG complex with SecY containing a unique Cys at position at 68 (ref. 21). The cells also expressed OmpA-GFP from pBAD-OmpA-GFP under an arabinose-inducible promoter 12. In all constructs, including those used in the two-component system, position −1 of the signal sequence was mutated to Tyr to prevent signal peptide cleavage 31. The cells were grown at 37°C to log phase in LB medium supplemented with 100 μg/mL ampicillin and 40 μg/mL chloramphenicol. OmpA-GFP expression was induced by addition of 0.15% L-arabinose for 1 hr. Where indicated, 0.3 mM copper(II) 1,10-phenanthroline (CuPh3) was added to the bacterial culture for 15 min at room temperature. The culture was treated with 10 mM N-ethyl maleimide (NEM) for 30 min on ice to block free cysteines. The cells were lysed in SDS sample buffer. Samples (equivalent amounts based on OD600nm of E. coli cultures) were subjected to non-reducing SDS-PAGE and analyzed by western blotting with anti-SecY- or GFP-antibodies. Where indicated, the samples were treated with 2% β-mercaptoethanol (β-ME) at 50°C for 20 min prior to SDS-PAGE. To test the translocation of E. coli SecA-OAIns fusion with E. coli SecY complex (two-component system), EP52 cells were transformed with pACYC-SecYEG and pBAD-EcSecA-OAIns. The cells were grown to log phage in LB medium supplemented with 100 μg/mL ampicillin and 40 μg/mL chloramphenicol, and SecA-OAIns expression was induced by addition of 0.1% L-arabinose for 4 hrs at room temperature or 30 min at 37°C. Disulfide crosslinking and non-reducing SDS-PAGE analysis were performed as described above.
Translocation by B. subtilis SecA and G. thermodenitrificans SecY complex was tested similarly. In the case of the three-component system, E. coli strain EP51 (Δrmf ΔompT) 21 was transformed with pTet-G. thermodenitrificans SecEHis8/YHis6, which expresses bicistronic G. thermodenitrificans SecE and SecY genes (SecY contains a Cys at position 60) under a tetracycline-inducible promoter 21. The cells were also transformed with pBAD-OmpA-GFP/B. subtilis SecA, a modified version of pBAD-OmpA, which contains an additional ribosome binding site and the B. subtilis SecA gene (for bicitronic expression) following OmpA-GFP. In the case of the two-component system, EP51 cells were transformed with pTet-G. thermodenitrificans SecEHis8/YHis6 and pBAD-B. subtilis SecA-OAIns. The cells were grown to log phase at 37°C, and the expression of G. thermodenitrificans SecYE was induced by addition of 200 ng/mL anhydrotetracycline for 1.5–2 hrs at 37°C. Then, the expression of OmpA-GFP/BsSecA or BsSecA-OAIns was induced by addition of 0.2% L-arabinose for 1–3 hrs. The cells were collected and subjected to SDS-PAGE followed by western blotting as described above. G. thermodenitrificans SecY was detected using anti-His antibodies.
Nanobody library generation
Purified G. thermodenitrificans SecYE in DDM-containing buffer was injected into an alpaca to elicit an immune response. A male alpaca (V. pacos) was purchased locally, maintained in pasture, and immunized following a protocol authorized by the Tufts University Cummings Veterinary School Institutional Animal Care and Use Committee. Following five rounds of immunization, total RNA was isolated from ~106 fresh peripheral blood lymphocytes, using the RNeasy Plus Mini Kit (Qiagen), following the manufacturer's instructions. Total RNA was used to synthesize a cDNA library using SuperScript III reverse transcriptase (ThermoFisher Scientific, Waltham MA) with a combination of random hexamers, oligo(dT), and gene-specific primers. The variable fragments of heavy chain antibodies (VHHs) segments were further amplified from this cDNA library using primers specific to the VHH region 32. PCR products were pooled, digested with NotI-HF and AscI (NEB, Ipswich, MA), gel purified, ligated into a M13 phagemid vector (pJSC), and transformed via electroporation into TG1 E. coli (Agilent, Santa Clara, CA). Library complexity was assessed by serial dilution and plating on 2YT agar plates supplemented with 2% glucose and 10 μg/ml ampicillin.
Selection of nanobodies by phage display
Purified SecYE and SecA-OAIns/SecYE proteins were biotinylated via coupling to primary amines with a fivefold molar excess of Chromalink NHS biotin reagent (Solulink, San Diego, CA) for 90 min. Excess biotin reagent was removed using a ZeBa desalting column (ThermoFisher Scientific, Waltham MA). Twenty μg of each protein were mixed with 100 μl MyOne Streptavidin T1 Dynabeads (ThermoFisher Scientific) blocked with 2% BSA. The beads were incubated with 200 ul of 1013 pfu/ml phage for 1 hr at room temperature. Non-binding phage were washed away and bound phage was eluted first by incubating with 1ml of saturated ER2738 culture, followed by 200mM pH2.2 glycine. The elutions were neutralized, pooled, and plated onto 2YT agar plates supplemented with 2% glucose, 5 μg/ml tetracycline, and 10 μg/ml ampicillin. A second round panning was performed with 2 μg of each protein and 40 μl MyOne Streptavidin T1 Dynabeads. All procedures were conducted in 20 mM Tris-HCl pH7.5, 150 mM NaCl, 10% glycerol, 0.02% DDM for SecYE, and with 20mM HEPES-KOH pH7.0, 150 mM NaCl, 10% glycerol, 0.02% DDM, 5mM MgCl2, 0.1mM ADP/BeFx for SecA-OAIns/SecYE. For each protein, 95 clones were sequenced, and sequences that appeared more than 5 times were selected for subsequent validation.
Nanobody screening
Thirteen distinct families of nanobodies directed against G. thermodenitrificans SecYE were identified by DNA sequencing. Twenty-two nanobody clones were sub-cloned into the pHEN6 vector 33, which adds an N-terminal pelB sequence and C-terminal sortase and His6 tags (LPETGG-His6). The proteins were expressed in 5ml E. coli cultures. After Ni-resin purification, the nanobodies were labeled with Alexa 555 (Invitrogen) by sortase reaction 34. The labeled nanobodies (1 μM concentration) were mixed with G. thermodenitrificans SecA-OAIns/SecYE complex or with SecYE alone at a molar ratio of 2:1. Nanobody binding was monitored by a shift of the peak in size-exclusion chromatography coupled with a fluorescence detector (Shimadzu). All 22 nanobodies bound to SecYE, but only nanobody AYC08 had a high affinity for SecA-OAIns/SecYE. The binding of AYC08 to free SecYE was weaker than to SecA-OAIns/SecYE, indicating that AYC08 interacts with the periplasmic side of SecY and that the binding epitope is only fully exposed in the active channel.
Protein Expression and Purification
E. coli strain EP51 was transformed with pTet- G. thermodenitrificans SecEHis8/Y and pBAD-B. subtilis SecA-OAIns49(L7). Residues 202-213 in the loop between TMs 5 and 6 of SecY were replaced by the sequence TFGGLN. Cells were grown in LB medium supplemented with 100 μg/mL ampicillin, 40 μg/mL chloramphenicol, and 0.5% glycerol at 37°C until OD 600nm reached 0.6–0.7. The expression of the G. thermodenitrificans SecYE was induced by addition of 200 ng/mL anhydrotetracycline, and cells were incubated for 1.5 hrs at 37°C and additional 1 h at 22°C. Then 0.15% L-arabinose was added to the culture to express B. subtilis SecA-OAIns overnight at 16°C. Cells were harvested by centrifugation and stored at −80°C until use.
The cells were suspended in buffer A (20 mM Tris-HCl pH7.5, 150 mM NaCl, 1 mM phenylmethanesulfonyl fluoride; IPTG) and lysed in a microfluidizer (Microfluidics). The membranes were pelleted by ultracentrifugation, washed once with buffer B (20 mM Tris-HCl pH7.5, 300 mM NaCl), and solubilized with 1% n-dodecyl β-D-maltoside (DDM; Anatrace) in buffer C (20 mM Tris-HCl pH7.5, 150 mM NaCl, 10% glycerol). After 1 hr incubation at 4 °C, the solution was clarified by ultracentrifugation. The supernatant was loaded onto a 5 ml POROS-MC20 column (Applied Biosystems) pre-charged with CoCl2. After washing with 15 ml of buffer D (as buffer C, but with 0.02% DDM) containing 10 mM imidazole and 5 ml of buffer D containing 15 mM imidazole, the protein was eluted with 5.5 ml of buffer D containing 250mM imidazole. Immediately after elution, 1 mM ADP/BeFx, 5 mM MgCl2, and 0.1 mg/ml E.coli polar lipids (25 mg/ml stock dissolved in 1% DDM) were added. To cleave the GFP-strep tag, 3C protease was added at a ratio of 1:30 (w:w) and mixture was incubated overnight at 4 °C. The sample was diluted 1:1 (v:v) with buffer E (20 mM Tris-HCl pH7.5, 10% glycerol, 0.02% DDM, 5 mM MgCl2) and loaded onto a Mono Q 10/100 column (GE Healthcare). The protein was eluted with a gradient of 15%-35% buffer F (20 mM Tris-HCl pH7.5, 1 M NaCl, 10% glycerol, 0.02% DDM, 5 mM MgCl2). The peak fractions were collected, and 0.1 mg/ml E .coli polar lipids and 1 mM ADP/BeFx were added. The protein was concentrated with an Amicon filter (100 kD MWCO, EMD Millipore) and loaded onto a Superdex 200 10/300 column (GE Healthcare) in buffer G (20 mM HEPES-KOH pH7.0, 100 mM NaCl, 10% glycerol, 0.02% DDM, 5 mM MgCl2, 1 mM ADP/BeFx). The peak fractions were concentrated to ~12 mg/ml, aliquoted, and flash-frozen in liquid nitrogen. The protein was stored at −80 °C and thawed right before crystallization.
The plasmid coding for nanobody AYC08 was transformed into WK6 cells. The cells were grown in 2×YT medium at 37 °C and protein expression was induced with 0.5 mM IPTG at OD600 0.6. The incubation was continued overnight at 30 °C. About 5 grams of cells were obtained from 1 L of culture. After suspension in 15 ml of TES buffer (200 mM Tris-HCl pH8.0, 500 μM EDTA, 500 mM sucrose), the sample was diluted 1:3 (v:v) in ice-cold water for 3 hrs to induce cell lysis. After centrifugation, the supernatant was mixed with 5 ml of Ni-NTA resin (Qiagen) and incubated at 4 °C for 1 hr. The protein was eluted with 250mM imidazole. The protein was further purified by gel filtration on a Superdex 200 10/300 column in 20 mM HEPES-KOH pH7.0, 100 mM NaCl, 10% glycerol, 5 mM MgCl2). The purified nanobody was concentrated to 30 mg/ml.
Crystallization
Initial crystals were obtained with a complex of G. thermodenitrificans SecYE and either B. subtilis SecA-OAIns and or B. subtilis SecA-DsbAIns (the latter contains the signal sequence of DsbA instead of that of OmpA). Only small crystals were obtained and diffracted to a maximum resolution of 6.5 Å at the Synchrotron X-ray source. A variety of approaches were tested to improve the crystals, such as inserting different hydrophilic proteins into each of the periplasmic loops of SecY (T4 lysozyme 35, cytochrome b562 36, P1/P4 domain of SecD/F 37, ROP helical bundle 38), truncating SecY loops, using various detergents, co-expressing SecG, and employing Fab-fragments of monoclonal antibodies generated against SecY. However, crystals with improved diffraction were only obtained when the complex was co-crystallized with nanobody AYC08.
The complex of G. thermodenitrificans SecYE and B. subtilis SecA-OAIns was mixed with nanobody AYC08 at a molar ratio of 1:1.2 with addition of 1 mg/ml lipids (42 mg/ml 1,2-dipalmitoyl-sn-glycero-3-phosphoglycerol (DPPG) plus 1,2-dipalmitoylsn-glycero-3-phosphoethanolamine (DPPE) (3:1) suspension in 0.5% DDM) 39. The mixture was incubated at 4 °C overnight and clarified by ultracentrifugation before setting up crystallization trays. The initial crystal screening yielded several crystal forms. Three of them were readily reproducible, but all diffracted to a maximum of ~6 Å resolution. Heavy atom compounds were screened for crystal soaking and the Ta6Br12 cluster improved the resolution limit of one crystal form. The best crystals were obtained with the hanging drop method, mixing 0.5 μl of the protein solution and 0.5 μl of well solution (21-24% polyethylene glycol (PEG) 1500, 100mM Tris-HCl pH8.5, 50-100 mM MgAc2, 2% 2-methyl-2,4-pentandiol; MPD) and using 24 well VDX plates with 500 μl well solution. The crystals were grown at 22°C over a week. The Ta6Br12 powder (Jena Bioscience) was suspended in buffer at a concentration of 20 mM and added to the crystallization drops at a final concentration of ~2 mM. After overnight incubation, the crystals were flash-frozen in liquid nitrogen
X-ray data collection and structure determination
Hundreds of crystals were screened at NE-CAT and GM/CA-CAT of the Advanced Photon Source (APS, Argonne National Laboratory). The diffraction of the crystals decayed rapidly and was weak, caused by strong heavy atom absorption. However, a complete set of three-wavelength MAD data from a single crystal could be collected at GM/CA-CAT. The data were processed with the XDS package40 and analyzed by the program Aimless41. The crystal belongs to the P6122 space group. The diffraction was anisotropic. Along axis c*, the diffraction went beyond 3.70 Å (I/sigma=2.0), whereas along the axes a* and b*, the diffraction was limited to 4.48 Å (I/sigma=1.5). The data were initially processed to 3.9 Å. Anisotropy correction was applied in the different programs used in the following calculations. Molecular replacement was used to locate SecA, SecYE, and the nanobody, employing as search models B. subtilis SecA (PDB ID: 1TF5), T. maritima SecYE (PDB ID:3DIN), and an anti-DHFR nanobody (PDB ID: 4EIG), respectively. The crystal contained one complex per asymmetric unit with a solvent content of 69%. Heavy atom sites were identified with the molecular replacement results by MR-SAD in Phaser42 and refined without molecular replacement models in Sharp (Global Phasing Limited). The experimental map based on the positions of the heavy metal ion clusters contained useful phase information to 5.5 Å (judged by phasing power of 1). The overall figure of merit was 0.44 (acentric) and 0.42 (centric). The phases were extended to 3.9 Å through density modification, using the programs Resolve43 and CNS44. After density modification, most of the α-helices were well resolved and some of the large side chains were visible in B-factor sharpened maps. To improve the density map, model phases were combined with experimental phases. Models for SecA and the nanobody molecules were placed first and modified according to the density map. Model phases were then combined with experimental phases to improve the density map. The SecY and SecE molecules were then built with the guidance of B-factor sharpened maps and the crystal structures of T. thermophilus 4 and T. maritima SecYE9. A density map for tracing the translocating peptide was generated by combining phase information from MAD phasing, and models of SecA, nanobody, and SecYE (Extended Data Fig. 4c). The signal sequence was initially modeled as an ideal poly-Ala helix and placed into the density map. The registry of the signal sequence helix was then determined from the density for two aromatic residues (Phe(−7) and Tyr(−1)). The following 23 amino acids of the translocating peptide were traced in a B-factor sharpened density map. The registry of this segment was determined on the basis of density for an aromatic residue (Phe3), several negatively charged residues (Glu4, Asp5, Glu10, and Glu12) surrounding the positively charged Ta6Br12 cluster, and the cysteine engaged in the disulfide bond (Cys7). Our model places Gly19 inside the pore ring, but it is possible that the registry is off by one residue (Ala18 or Gly20 would be in the pore), an ambiguity that does not alter the interpretation of the model. Annealing to a temperature of 2,500 K was applied to the model at an early stage of refinement with the program Phenix45. The individual XYZ and group B factors were refined by using both the Phenix and Refmac546 programs. Secondary structure was tightly constrained during refinement. The SAD likelihood function implemented in Refmac5 was used to refine the heavy atom clusters together with the protein model. The model was improved by molecular replacement and multiple cycles of model building and refinement. Manual adjustments were made in COOT47 and refinement was performed in reciprocal space. At a late stage of refinement, the diffraction data were extended to 3.7 Å, which helped to resolve some regions in the density map, e.g. of the translocating peptide. We also tested the data with ellipsoidal truncation processed by the Anisotropy server (http://services.mbi.ucla.edu/anisoscale/). The truncated data produced better density maps, which were used for refinement as well. The final model was refined to an Rwork of 30.6% and an Rfree of 32.5% and showed good geometry (Extended Data Table 1). An attempt was made to include the model-refined Ta6Br12 clusters as the resolved heavy atom substructure for recalculation of SAD and MAD experimental phases. However, this did not result in improvement of the map, in part because individual metal atoms of the clusters could not be accurately positioned, as observed in other cases of similar resolution 48. While the centers of the clusters are well defined, the individual metal atom positions in our model should be considered to be very approximate. To ensure that the R-factor is not dominated by the heavy metal ion clusters, we tested its sensitivity to changes in the protein model. Rfree increased by 0.013-0.017 upon deletion of any of the TMs of SecY or of the signal sequence helix. A similar increase was observed when the same analysis was performed for three different membrane proteins of similar size (2ZD9, 4CZB, 4CDI; 1,078-1,655 amino acids), the structure of which was determined in the absence of heavy metal ion clusters at similar resolution (3.5-4Å), solvent content (0.63-0.85), and R-factor (0.3-0.34). Model validation was performed by using PHENIX. The following regions could not be traced: residues 1-15, 244-247, 262-264, 271-272, 620-626, and 635-712 of SecA, residues 1-12, 145-146, 200-213, 244-260, and 291-308, 390-391, and 396-398 of SecY, residues 1-3, 22-23, 58-60 of SecE, residues 30-31, 42, and 100 of the nanobody, and the first 5 residues of the linker preceding the signal sequence. In addition, some amino acid side chains were not well resolved, so they were modeled as Ala. Some density close to metal ion cluster sites 15 to 18 remains unexplained. Figures showing the structures were generated with Chimera49. All the X-ray crystallographic software was maintained by SBGrid50.
Extended Data
Extended Data Figure 1. Generation of translocation intermediates with a three-component system.
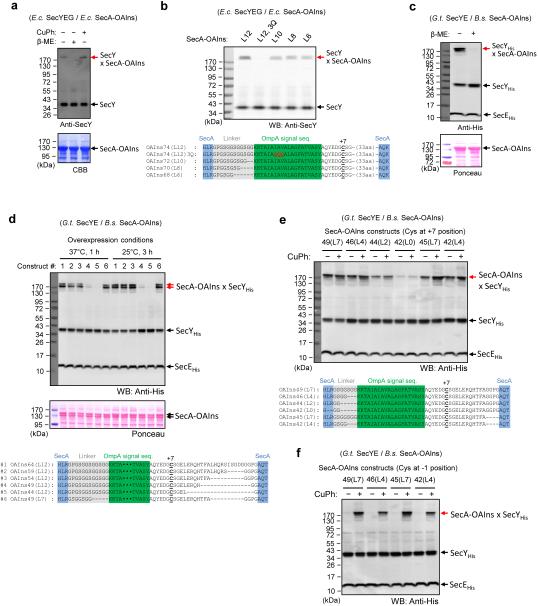
a, Strategy to generate SecA-dependent translocation intermediates in E. coli cells. The intermediates are assembled from E. coli SecA, E. coli SecY complex, and substrate that contains an N-terminal OmpA signal sequence and C-terminal superfolder GFP51 (sfGFP). After loop insertion into the SecY channel, translocation of the C-terminus is stalled by the folded sfGFP. Insertion is monitored by disulfide bond formation between a pair of cysteines introduced into the substrate and the plug of SecY (yellow stars). b, Scheme showing the simplified system, in which a secretory protein segment is fused into the two-helix finger of SecA. c, Sequence of the substrate used in a. The −1 position of the original signal sequence was changed to Tyr to prevent signal sequence cleavage. The position of the cysteine and the length of the translocated segment were varied (here shown for Cys at position +3 and 58 amino acids length). d, Variation of the Cys position with a translocationg segment of 58 residues. Where indicated, disulfide crosslinks to SecY with a Cys at position 68 (OmpA-GFPxSecY) were induced by the oxidant copper phenanthroline (CuPh) prior to harvesting the cells. The samples were analyzed by non-reducing SDS-PAGE, followed by western blotting (WB) with anti-SecY and anti-GFP antibodies. e, As in d, but in the absence of oxidant, with Cys at different positions and variation of the length of the translocated segment. Asterisks indicate nonspecific bands. f, As in e, but with E. coli cells expressing B. subtilis (B.s.) SecA and G. thermodenitrificans (G.t.) SecYE. The substrate contained a Cys at position +7, and SecY a Cys at position 60. The red arrows indicate spontaneously generated disulfide crosslinks (GFP sometimes does not unfold in SDS, resulting in two bands). The OmpA-GFP constructs contained a C-terminal Strep-tag that was detected by StrepTactin conjugated with horse radish peroxidase (HRP).
Extended Data Figure 2. Generation of translocation complexes with SecA-substrate fusion constructs.
a, Translocation complexes were generated as indicated in the scheme in Extended Data Fig. 1b. An E. coli SecA-substrate fusion (SecA-OAIns74 (L12)) was overexpressed together with E. coli SecY complex in E. coli cells. SecA-OAIns74 (L12) contains 74 amino acids inserted into the two-helix finger of SecA, including a linker of 12 residues, and a GFP tag following SecA. Translocation of the substrate segment was monitored by spontaneous disulfide crosslinking between a cysteine at position +7 (with respect to the original signal peptidase cleavage site) and a cysteine at position 68 in the plug of SecY. Where indicated, β-mercaptoethanol (β-ME) was added to reduce the disulfide bond. The samples were analyzed by non-reducing SDS-PAGE and western blotting with anti-SecY antibodies. The overexpression of SecA-OAIns was monitored by the fluorescence of GFP (data not shown) and staining with Coomassie blue (CBB, lower panel). b, As in a, but with E. coli SecA-OAIns constructs containing from 6 to 12 residues in the linker (L6-L12) or mutations (3Q) in the H-region of the signal sequence. Expression of SecA-OAIns was verified by the strong green fluorescence of cell lysates, caused by GFP fused to the C-terminus of SecA (not shown). The lower panel shows the sequences of the SecA-inserted segments. c, As in a, but with B. subtilis SecA-OAIns74 (L12) and G. thermodenitrificans SecYE. SecY and SecA were detected by western blotting with anti-His antibodies and Ponceau staining, respectively. d, As in c, but with B. subtilis SecA-OAIns containing different inserted segments. SecA-OAIns was expressed for under different conditions, as indicated. Expression of SecA-OAIns was verified by green fluorescence of GFP fused to the C-terminus of SecA (not shown) and Ponceau staining (second panel). The sequences of the constructs are shown in the lowest panel. e, As in d, but with different constructs, the sequences of which are shown in the lower panel. Where indicated, copper phenanthroline (CuPh) was added to the cells to induce disulfide bridge formation. SecA-OAIns49(L7) was used for crystallization. f, As in e, but with a Cys at position −1 (the last residue of the OmpA signal sequence) instead of position +7. Note that in this case disulfide formation does not occur spontaneously.
Extended Data Figure 3. Purification of an active translocation complex.
a, Scheme of the purification protocol. b, Elution of the G. thermodenitrificans SecYE – B. subtilis SecA-OAIns complex from a Superdex200 column during the last chromatography step. c, Non-reducing SDS-PAGE analysis of samples taken during the purification procedure and of fractions indicated with red numbers in b. Lane 1, molecular weight markers. Lane 2, sample analyzed after IMAC. Lane 3, sample after cleavage of the GFP tag. Lane 4, sample after anion exchange chromatography (MonoQ).
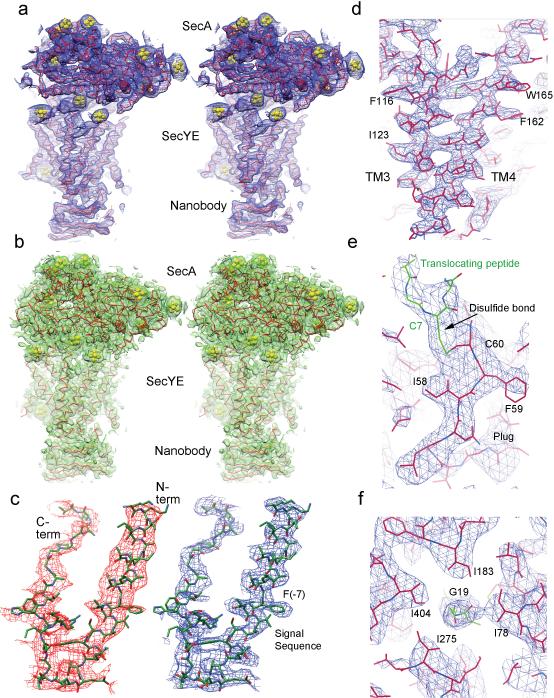
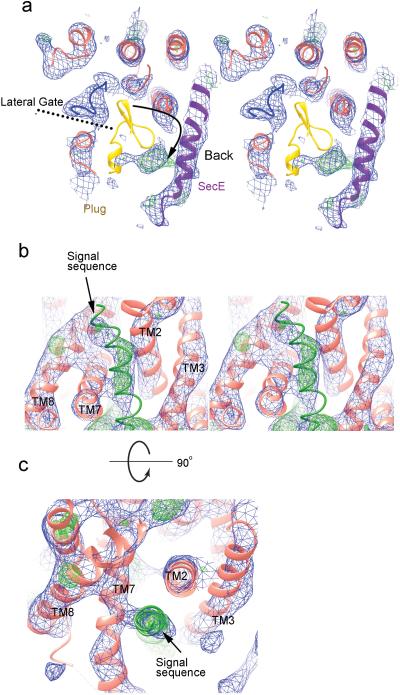
Extended Data Figure 4. Electron density map and refined model.
a, Stereo view of the unsharpened density map (2FO-FC; 1σ) of the entire complex. Heavy metal ion clusters are shown in yellow. b, As in a, but with the density map derived from MAD phasing after density modification. c, SigmaA-weighted phase-combined 2FO-FC density maps of the translocating peptide region. The left panel shows an omit map calculated without a model for the translocating peptide. The right panel shows a map calculated with the model. Phe (−7) is one of the residues used to determine the registry of the helix. d, A side view of the density for TMs 3 and 4. e, Density showing the disulfide crosslink between the plug and translocating chain. f, Top view of Gly19 of the translocating chain surrounded by pore residues.
Extended Data Figure 5. Interactions of the nanobody and SecA with SecY.
a, The nanobody binds to the plug and to the loop between TMs 3 and 4 (L3/4). b, The polypeptide crosslinking domain (PPXD; in yellow) of SecA interacts with the loop between TMs 8 and 9 of SecY (L8/9; in red), and the long helix of the helical scaffold domain (HSD; in blue) with the loop between TMs 2 and 3 (L2/3; in red). The loop between TMs 6 and 7 of SecY (L6/7; in red) does not seem to make strong contact with SecA. c, Two helices of the HSD interact with the C-terminal tail of SecY (C-tail; in red).
Extended Data Figure 6. Comparison of the conformations of SecA in the active G. thermodenitrificans and inactive T. maritima complexes.
The domains of SecA in the G. thermodenitrificans complex are labeled with different colors (nucleotide binding domain 1 (NBD1), blue; nucleotide binding domain 2 (NBD2), cyan; helical scaffold domain (HSD), brown; helical wing domain (HWD), grey; polypeptide crosslinking domain (PPXD), yellow). SecA in the T. maritima complex is shown in pink. The left panel shows a top view (the channel would be underneath), and the right panel a side view with the two-helix finger (part of HSD) indicated.
Extended Data Figure 7. Localization of signal sequences in the G. thermodenitrificans SecY and mammalian Sec61 channels and of a TM domain in the mammalian Sec61 channel.
a, In the active channel (salmon), the signal sequence displaces TM7 and TM8 in the idle T. thermophilus channel (cyan). b, As in a, but comparison with the idle M. jannaschii channel (tan). The C-region of the signal sequence takes the position of TM7. c, Side view of the interactions of the H-region of the signal sequence with TM2 of G. thermodenitrificans SecY. Interacting amino acids are indicated. d, Stereo view showing the intercalation of the C-region into the periplasmic side of the lateral gate. Residues of the amphipathic helix are indicated. e, The active G. thermodenitrificans channel (in salmon) was aligned with a mammalian channel (in grey) containing a nascent membrane protein (PDB ID: 4CG6) using secondary structure matching 47. The signal sequence in the bacterial channel is shown in green, and the TM segment of the nascent membrane protein in yellow. f, As in e, but comparison of the active G. thermodenitrificans channel with a mammalian Sec61 channel (light blue) containing a secretory protein fragment (PDB ID 3JC2). The signal sequences are shown in green and yellow, respectively.
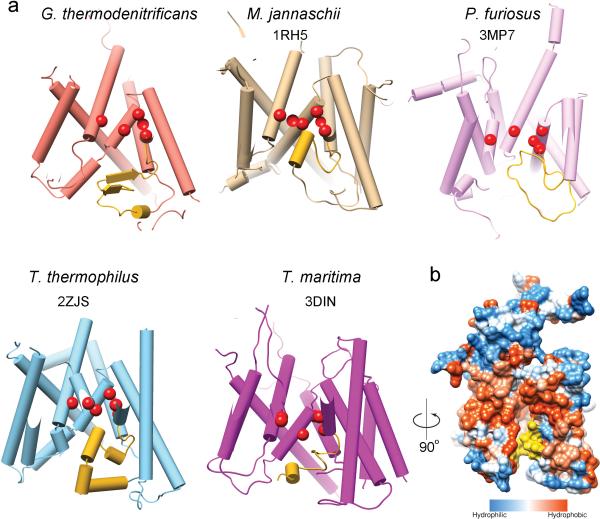
Extended Data Figure 8. The plug domains in SecY channels.
a, The plugs in channels of different organisms have different structures. Shown are side views with the plugs in yellow and pore residues as red spheres. PDB IDs are given below the names of the organisms. For the G. thermodenitrificans channel, the translocating peptide segment was omitted. b, In the inactive T. maritima SecY channel, the plug (in yellow) is at the front of the channel, partially sealing the periplasmic side of the lateral gate. Shown is a side view in a surface representation, with hydrophilic and hydrophobic residues in blue and orange, respectively.
Extended Data Figure 9. Structures of the active G. thermodenitrificans complex determined without nanobody or with a different signal sequence.
a, Stereo view of density maps at 6.5Å resolution for the active complex in the absence of nanobody. Shown is a 2FO-FC density map at 1σ (blue mesh) and a difference map (FO-FC) at 3σ (green mesh), both calculated by molecular replacement with a model lacking the plug. Strong positive density is seen close to SecE, likely corresponding to parts of the plug. The arrow indicates the movement of the plug from the position in the structure with nanobody to the density seen in the structure without nanobody. b, Stereo views of density maps at ~8.5 Å resolution for the active G. thermodenitrificans complex in which the OmpA signal sequence was replaced by that of DsbA. Shown is a side view of the 2FO-FC density map at 1σ (blue mesh) and a difference map (Fo-Fc) at 3σ (green mesh), both calculated by molecular replacement with a model lacking the signal sequence. Note that the model for the OmpA signal sequence fits well into the density corresponding to the DsbA signal sequence. c, As in b, but top view and not in stereo.
Extended Data Table 1.
Data collection and refinement statistics for MAD structures
| SecA-OAIns/SecYE + Ta6Br12 | |
|---|---|
| Data collection | GM/CAT-CAT @APS |
| Space group | P6122 |
| Cell dimensions | |
| a, b, c (Å) | 127.798,127.798,554.772 |
| α, β, γ (°) | 90, 90, 120 |
| Remote | Inflection | Peak | |
|---|---|---|---|
| Wavelength | 1.2782 | 1.2552 | 1.2548 |
| Resolution (Å) | 3.70 (3.80-3.70) | 4.00 (4.10-4.00) | 4.50 (4.62-4.50) |
| Rsym or Rmerge | 0.084 (>1) | 0.095 (>1) | 0.128 (>1) |
| I/σI | 9.30 (0.4) | 9.69 (0.9) | 11.90 (1.0) |
| Completeness (%) | 99 (100) | 99 (100) | 99 (100) |
| CC1/2 | 0.999 (0.57) | 0.998 (0.80) | 0.999 (0.92) |
| Redundancy | 10.6 (10.2) | 10.6 (10.4) | 21.3 (21.6) |
| Refinement | |||
| Resolution (Å) | 3.7 | ||
| No. reflections | 53812 | ||
| Rwork/ Rfree | 0.306/0.325 | ||
| No. atoms | 9888 | ||
| Protein | 9532 | ||
| Ligand/ion | 356 | ||
| Water | N/A | ||
| B-factors | 194.00 | ||
| Protein | 190.24 | ||
| Ligand/ion | 300.34 | ||
| Water | N/A | ||
| R.m.s deviations | |||
| Bond lengths (Å) | 0.007 | ||
| Bond angles (°) | 1.15 | ||
| Ramachandran | |||
| Favored / allowed / outliers (%) | 91.3 / 6.8 / 1.9 | ||
*Highest resolution shell is shown in parenthesis.
Supplementary Material
Acknowledgments
We thank Christina Martone and for help with nanobody generation and cloning, Charles Shoemaker and Jean Mukherjee at Tufts Cummings Veterinary School for their assistance in alpaca immunization, Andrew Whynot for cloning G. thermodenitrificans SecY, Hiroshi Suzuki and Thomas Walz for help with fluorescence size exclusion chromatography. We thank the staff at NE-CAT of the Advanced Photon Source, the SBGrid consortium at Harvard Medical School, the organizers of the CCP4/APS summer school 2015, and the beam host at GMCA-CAT. We thank Adrian Salic and Thomas Guettler for critical reading of the manuscript. The work was supported by National Institute of Health grants to T.A.R. (GM052586) and by a Pioneer Award to H.P. T.A.R. is a Howard Hughes Medical Institute investigator.
Footnotes
Author Contributions
E.P. designed the SecA-substrate fusion constructs, performed biochemical tests, and obtained the initial crystals; L.L. optimized constructs and crystals, performed biochemical tests, screened nanobodies and determined the crystal structures; J.L., J.I., and H.P. generated and cloned nanobodies; T.A.R., L.L., and E.P interpreted the structure and wrote the manuscript. T.A.R. supervised the project.
The crystal structure determined in this study has been depositied in the RCSB Protein Data Bank under the accession number 5EUL. The authors declare no competing financial interests. Readers are welcome to comment on the online version of the paper.
Online Content Methods, along with any additional Extended Data display items and Source data, are available in the online version of the paper; references unique to these sections appear only in the online paper.
Accession codes of structures used in this paper
1RH5, 3MP7, 2ZJS, 3DIN, 5AWW, 3JC2, 4CG6. The new structure described in this paper has the code 5EUL.
References
- 1.van den Berg B, et al. X-ray structure of a protein-conducting channel. Nature. 2004;427:36–44. doi: 10.1038/nature02218. [DOI] [PubMed] [Google Scholar]
- 2.Breyton C, Haase W, Rapoport TA, Kuhlbrandt W, Collinson I. Threedimensional structure of the bacterial protein-translocation complex SecYEG. Nature. 2002;418:662–665. doi: 10.1038/nature00827. [DOI] [PubMed] [Google Scholar]
- 3.Tanaka Y, et al. Crystal Structures of SecYEG in Lipidic Cubic Phase Elucidate a Precise Resting and a Peptide-Bound State. Cell reports. 2015;13:1561–1568. doi: 10.1016/j.celrep.2015.10.025. [DOI] [PubMed] [Google Scholar]
- 4.Tsukazaki T, et al. Conformational transition of Sec machinery inferred from bacterial SecYE structures. Nature. 2008;455:988–991. doi: 10.1038/nature07421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Egea PF, Stroud RM. Lateral opening of a translocon upon entry of protein suggests the mechanism of insertion into membranes. Proceedings of the National Academy of Sciences of the United States of America. 2010;107:17182–17187. doi: 10.1073/pnas.1012556107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Park E, et al. Structure of the SecY channel during initiation of protein translocation. Nature. 2014;506:102–106. doi: 10.1038/nature12720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Gogala M, et al. Structures of the Sec61 complex engaged in nascent peptide translocation or membrane insertion. Nature. 2014;506:107–110. doi: 10.1038/nature12950. [DOI] [PubMed] [Google Scholar]
- 8.Voorhees RM, Fernandez IS, Scheres SH, Hegde RS. Structure of the mammalian ribosome-Sec61 complex to 3.4 A resolution. Cell. 2014;157:1632–1643. doi: 10.1016/j.cell.2014.05.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zimmer J, Nam Y, Rapoport TA. Structure of a complex of the ATPase SecA and the protein-translocation channel. Nature. 2008;455:936–943. doi: 10.1038/nature07335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Pfeffer S, et al. Structure of the native Sec61 protein-conducting channel. Nature communications. 2015;6:8403. doi: 10.1038/ncomms9403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Park E, Rapoport TA. Mechanisms of Sec61/SecY-mediated protein translocation across membranes. Annual review of biophysics. 2012;41:21–40. doi: 10.1146/annurev-biophys-050511-102312. [DOI] [PubMed] [Google Scholar]
- 12.Park E, Rapoport TA. Bacterial protein translocation requires only one copy of the SecY complex in vivo. The Journal of cell biology. 2012;198:881–893. doi: 10.1083/jcb.201205140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Jungnickel B, Rapoport TA. A posttargeting signal sequence recognition event in the endoplasmic reticulum membrane. Cell. 1995;82:261–270. doi: 10.1016/0092-8674(95)90313-5. [DOI] [PubMed] [Google Scholar]
- 14.Bauer BW, Shemesh T, Chen Y, Rapoport TA. A “Push and Slide” Mechanism Allows Sequence-Insensitive Translocation of Secretory Proteins by the SecA ATPase. Cell. 2014;157:1416–1429. doi: 10.1016/j.cell.2014.03.063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Li W, et al. The plug domain of the SecY protein stabilizes the closed state of the translocation channel and maintains a membrane seal. Molecular cell. 2007;26:511–521. doi: 10.1016/j.molcel.2007.05.002. [DOI] [PubMed] [Google Scholar]
- 16.Harris CR, Silhavy TJ. Mapping an interface of SecY (PrlA) and SecE (PrlG) by using synthetic phenotypes and in vivo cross-linking. J. Bacteriol. 1999;181:3438–3444. doi: 10.1128/jb.181.11.3438-3444.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Tam PC, Maillard AP, Chan KK, Duong F. Investigating the SecY plug movement at the SecYEG translocation channel. The EMBO journal. 2005;24:3380–3388. doi: 10.1038/sj.emboj.7600804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Paetzel M. Structure and mechanism of Escherichia coli type I signal peptidase. Biochimica et biophysica acta. 2014;1843:1497–1508. doi: 10.1016/j.bbamcr.2013.12.003. [DOI] [PubMed] [Google Scholar]
- 19.Yamamoto Y, et al. Conformational requirement of signal sequences functioning in yeast: circular dichroism and 1H nuclear magnetic resonance studies of synthetic peptides. Biochemistry. 1990;29:8998–9006. doi: 10.1021/bi00490a017. [DOI] [PubMed] [Google Scholar]
- 20.Rizo J, Blanco FJ, Kobe B, Bruch MD, Gierasch LM. Conformational Behavior of Escherichia coli OmpA Signal Peptides in Membrane Mimetic Environments. Biochemistry. 1993;32:4881–4894. doi: 10.1021/bi00069a025. [DOI] [PubMed] [Google Scholar]
- 21.Park E, Rapoport TA. Preserving the membrane barrier for small molecules during bacterial protein translocation. Nature. 2011;473:239–242. doi: 10.1038/nature10014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Cannon KS, Or E, Clemons WM, Jr., Shibata Y, Rapoport TA. Disulfide bridge formation between SecY and a translocating polypeptide localizes the translocation pore to the center of SecY. The Journal of cell biology. 2005;169:219–225. doi: 10.1083/jcb.200412019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Heinrich SU, Mothes W, Brunner J, Rapoport TA. The Sec61p complex mediates the integration of a membrane protein by allowing lipid partitioning of the transmembrane domain. Cell. 2000;102:233–244. doi: 10.1016/s0092-8674(00)00028-3. [DOI] [PubMed] [Google Scholar]
- 24.Hessa T, et al. Recognition of transmembrane helices by the endoplasmic reticulum translocon. Nature. 2005;433:377–381. doi: 10.1038/nature03216. [DOI] [PubMed] [Google Scholar]
- 25.Bischoff L, Wickles S, Berninghausen O, van der Sluis EO, Beckmann R. Visualization of a polytopic membrane protein during SecY-mediated membrane insertion. Nature communications. 2014;5:4103. doi: 10.1038/ncomms5103. [DOI] [PubMed] [Google Scholar]
- 26.Martoglio B, Hofmann MW, Brunner J, Dobberstein B. The proteinconducting channel in the membrane of the endoplasmic reticulum is open laterally toward the lipid bilayer. Cell. 1995;81:207–214. doi: 10.1016/0092-8674(95)90330-5. [DOI] [PubMed] [Google Scholar]
- 27.Plath K, Mothes W, Wilkinson BM, Stirling CJ, Rapoport TA. Signal sequence recognition in posttranslational protein transport across the yeast ER membrane. Cell. 1998;94:795–807. doi: 10.1016/s0092-8674(00)81738-9. [DOI] [PubMed] [Google Scholar]
- 28.McKnight CJ, Briggs MS, Gierasch LM. Functional and Nonfunctional Lamb Signal Sequences Can Be Distinguished by Their Biophysical Properties. The Journal of biological chemistry. 1989;264:17293–17297. [PubMed] [Google Scholar]
- 29.Nielsen H, Engelbrecht J, Brunak S, von Heijne G. Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein engineering. 1997;10:1–6. doi: 10.1093/protein/10.1.1. [DOI] [PubMed] [Google Scholar]
- 30.Voorhees RM, Hegde RS. Structure of the Sec61 channel opened by a signal sequence. Science. 2016;351:88–91. doi: 10.1126/science.aad4992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Fikes JD, Barkocy-Gallagher GA, Klapper DG, Bassford PJ. Maturation of Escherichia coli Maltose-Binding Protein by Signal Peptidase-I in vivo - Sequence Requirements for Efficient Processing and Demonstration of an Alternate Cleavage Site. The Journal of biological chemistry. 1990;265:3417–3423. [PubMed] [Google Scholar]
- 32.Maass DR, Sepulveda J, Pernthaner A, Shoemaker CB. Alpaca (Lama pacos) as a convenient source of recombinant camelid heavy chain antibodies (VHHs). Journal of immunological methods. 2007;324:13–25. doi: 10.1016/j.jim.2007.04.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Arbabi Ghahroudi M, Desmyter A, Wyns L, Hamers R, Muyldermans S. Selection and identification of single domain antibody fragments from camel heavy-chain antibodies. FEBS letters. 1997;414:521–526. doi: 10.1016/s0014-5793(97)01062-4. [DOI] [PubMed] [Google Scholar]
- 34.Guimaraes CP, et al. Site-specific C-terminal and internal loop labeling of proteins using sortase-mediated reactions. Nature protocols. 2013;8:1787–1799. doi: 10.1038/nprot.2013.101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Rosenbaum DM, et al. GPCR engineering yields high-resolution structural insights into beta2-adrenergic receptor function. Science. 2007;318:1266–1273. doi: 10.1126/science.1150609. [DOI] [PubMed] [Google Scholar]
- 36.Chun E, et al. Fusion partner toolchest for the stabilization and crystallization of G protein-coupled receptors. Structure. 2012;20:967–976. doi: 10.1016/j.str.2012.04.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Tsukazaki T, et al. Structure and function of a membrane component SecDF that enhances protein export. Nature. 2011;474:235–238. doi: 10.1038/nature09980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Hari SB, Byeon C, Lavinder JJ, Magliery TJ. Cysteine-free Rop: a fourhelix bundle core mutant has wild-type stability and structure but dramatically different unfolding kinetics. Protein science : a publication of the Protein Society. 2010;19:670–679. doi: 10.1002/pro.342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Gourdon P, et al. HiLiDe—Systematic Approach to Membrane Protein Crystallization in Lipid and Detergent. Cryst. Growth Des. 2011;11:2098–2106. [Google Scholar]
- 40.Kabsch W. Xds. Acta crystallographica. Section D, Biological crystallography. 2010;66:125–132. doi: 10.1107/S0907444909047337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Evans PR, Murshudov GN. How good are my data and what is the resolution? Acta crystallographica. Section D, Biological crystallography. 2013;69:1204–1214. doi: 10.1107/S0907444913000061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.McCoy AJ. Solving structures of protein complexes by molecular replacement with Phaser. Acta crystallographica. Section D, Biological crystallography. 2007;63:32–41. doi: 10.1107/S0907444906045975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Terwilliger TC. Maximum-likelihood density modification. Acta crystallographica. Section D, Biological crystallography. 2000;56:965–972. doi: 10.1107/S0907444900005072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Brunger AT. Version 1.2 of the Crystallography and NMR system. Nat. Protoc. 2007;2:2728–2733. doi: 10.1038/nprot.2007.406. [DOI] [PubMed] [Google Scholar]
- 45.Adams PD, et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta crystallographica. Section D, Biological crystallography. 2010;66:213–221. doi: 10.1107/S0907444909052925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Murshudov GN, et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta crystallographica. Section D, Biological crystallography. 2011;67:355–367. doi: 10.1107/S0907444911001314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Emsley P, Lohkamp B, Scott WG, Cowtan K. Features and development of Coot. Acta crystallographica. Section D, Biological crystallography. 2010;66:486–501. doi: 10.1107/S0907444910007493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Banumathi S, Dauter M, Dauter Z. Phasing at high resolution using Ta6Br12 cluster. Acta crystallographica. Section D, Biological crystallography. 2003;59:492–498. doi: 10.1107/s0907444903000064. [DOI] [PubMed] [Google Scholar]
- 49.Pettersen EF, et al. UCSF Chimera--a visualization system for exploratory research and analysis. Journal of computational chemistry. 2004;25:1605–1612. doi: 10.1002/jcc.20084. [DOI] [PubMed] [Google Scholar]
- 50.Morin A, et al. Collaboration gets the most out of software. eLife. 2013;2:e01456. doi: 10.7554/eLife.01456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Pedelacq JD, Cabantous S, Tran T, Terwilliger TC, Waldo GS. Engineering and characterization of a superfolder green fluorescent protein. Nature biotechnology. 2006;24:79–88. doi: 10.1038/nbt1172. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.