Abstract
The health care-associated pneumonia (HCAP) criteria have a limited ability to predict pneumonia caused by drug-resistant bacteria and favor the overutilization of broad-spectrum antibiotics. We aimed to derive and validate a clinical prediction score with an improved ability to predict the risk of pneumonia due to drug-resistant pathogens compared to that of HCAP criteria. A derivation cohort of 200 microbiologically confirmed pneumonia cases in 2011 and 2012 was identified retrospectively. Risk factors for pneumonia due to drug-resistant pathogens were evaluated by logistic regression, and a novel prediction score (the drug resistance in pneumonia [DRIP] score) was derived. The score was then validated in a prospective, observational cohort of 200 microbiologically confirmed cases of pneumonia at four U.S. centers in 2013 and 2014. The DRIP score (area under the receiver operator curve [AUROC], 0.88 [95% confidence interval {CI}, 0.82 to 0.93]) performed significantly better (P = 0.02) than the HCAP criteria (AUROC, 0.72 [95% CI, 0.64 to 0.79]). At a threshold of ≥4 points, the DRIP score demonstrated a sensitivity of 0.82 (95% CI, 0.67 to 0.88), a specificity of 0.81 (95% CI, 0.73 to 0.87), a positive predictive value (PPV) of 0.68 (95% CI, 0.56 to 0.78), and a negative predictive value (NPV) of 0.90 (95% CI, 0.81 to 0.93). By comparison, the performance of HCAP criteria was less favorable: sensitivity was 0.79 (95% CI, 0.67 to 0.88), specificity was 0.65 (95% CI, 0.56 to 0.73), PPV was 0.53 (95% CI, 0.42 to 0.63), and NPV was 0.86 (95% CI, 0.77 to 0.92). The overall accuracy of the HCAP criteria was 69.5% (95% CI, 62.5 to 75.7%), whereas that of the DRIP score was 81.5% (95% CI, 74.2 to 85.6%) (P = 0.005). Unnecessary extended-spectrum antibiotics were recommended 46% less frequently by applying the DRIP score (25/200, 12.5%) than by use of HCAP criteria (47/200, 23.5%) (P = 0.004), without increasing the rate at which inadequate treatment recommendations were made. The DRIP score was more predictive of the risk of pneumonia due to drug-resistant pathogens than HCAP criteria and may have the potential to decrease antibiotic overutilization in patients with pneumonia. Validation in larger cohorts of patients with pneumonia due to all causes is necessary.
INTRODUCTION
Pneumonia is the leading infectious cause of death in the United States and a major contributor to health care resource utilization (1, 2). Effective initial empirical antibiotic selection has been associated with improved outcomes (3, 4). In patients with community-acquired pneumonia (CAP), treatment with beta-lactam–macrolide combinations or a fluoroquinolone with activity against respiratory pathogens is recommended on the basis of the pathogens that are commonly isolated and their benefit for the prevention of mortality (5–7). However, there has been increasing concern over pneumonia due to CAP drug-resistant pathogens (DRPs) (8). Prediction of the risk of pneumonia due to DRPs and the appropriate selection of extended-spectrum antibiotics targeting methicillin-resistant Staphylococcus aureus (MRSA), Pseudomonas aeruginosa, and other drug-resistant members of the family Enterobacteriaceae are challenging for clinicians.
To identify patients at risk of infection caused by DRPs, the 2005 American Thoracic Society/Infectious Diseases Society of America (ATS/IDSA) guidelines introduced a new subset of CAP, health care-associated pneumonia (HCAP) (9). Treatment with extended-spectrum antibiotics is recommended for patients meeting at least one of the HCAP criteria (9). Following publication of these guidelines, the rate of use of vancomycin and extended-spectrum antibiotics with activity against Gram-negative bacteria for the treatment of pneumonia doubled (10, 11). However, the outcomes among patients that meet the HCAP criteria and that are treated with guideline-concordant therapy have not improved (12–14). This may be in part due to the poor ability of the HCAP criteria to predict pneumonia due to DRPs (15–17). The unnecessary use of extended-spectrum antibiotics is associated with negative collateral effects, including increased costs (12, 18) and lengths of stay (12, 19, 20), drug toxicity (21–24), Clostridium difficile infection (25), and resistance (26).
Better methods of predicting the risk of pneumonia due to DRPs while limiting the unnecessary use of extended-spectrum antibiotics are needed. Although several alternative predictive scores have been proposed (27–33), none has demonstrated significantly improved specificity compared to that of the HCAP criteria (34) and few validation studies with U.S. populations have been performed. Here we describe the derivation of the drug resistance in pneumonia (DRIP) score, a novel model to predict the risk of pneumonia due to DRPs, and the results of a prospective validation in a multicenter U.S. cohort.
MATERIALS AND METHODS
Approval for the study was independently granted by the institutional review boards (IRBs) of Intermountain Healthcare, the Mayo Clinic in Arizona, and the University of Connecticut Health Center. Informed consent was obtained from patients when required by the IRBs.
Derivation cohort.
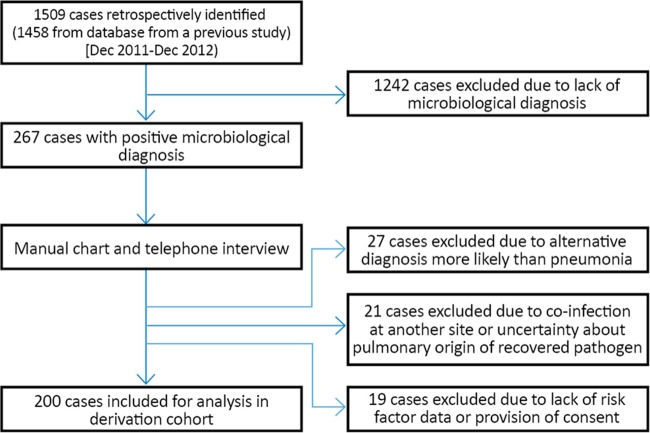
A derivation cohort comprised of 200 patients with microbiologically confirmed cases of pneumonia was identified retrospectively by applying the study criteria to a cohort of 1,509 patients identified in a previous observational study (35) (Fig. 1). The derivation cohort consisted of patients with pneumonia admitted to seven hospitals in Utah from December 2011 to December 2012. After electronic screening of a database containing the records of pneumonia patients assembled for a previous study (35) by microbiological diagnosis, the medical records of 267 cases were ultimately manually reviewed by investigators in order to assemble the final derivation cohort. Patients were contacted by telephone if risk factor data were not available in the medical record. Uncertainty related to enrollment eligibility, pneumonia diagnosis, or microbiological validity was arbitrated by an independent study investigator.
FIG 1.
Derivation cohort enrollment.
Pneumonia was defined clinically as two or more clinical signs or symptoms (temperature, <36.0°C or >38.0°C; respiratory rate, >20 breaths per minute; room air oxygen saturation, <90%; partial pressure of O2 in arterial blood, <60 mm Hg; cough; sputum production; white blood cell count, <4,000/μl or >10,000/μl; bandemia, >10%) plus radiographic evidence of new parenchymal opacity or cavitation. Cases were excluded if a diagnosis other than pneumonia was considered more likely by investigators.
Microbiological sources included cultures of an adequate sputum sample (36) or tracheal aspirate, bronchoalveolar lavage (BAL) fluid, pleural fluid, or blood sample; urinary antigen tests for Legionella pneumophila and Streptococcus pneumoniae; and a PCR film array for Bordetella pertussis, Mycoplasma pneumoniae, and Chlamydophila pneumoniae. Organisms recovered from cultures of sputum were excluded on the basis of suspected colonization if a second, alternative pathogen was concomitantly recovered from a microbiological specimen (blood or BAL fluid) collected by an invasive means. Organisms recovered from cultures of blood were excluded if bacteremia from an alternate source, such as the urinary tract, was deemed more likely.
Cases with a positive microbiological diagnosis were included only if an organism consistent with a bacterial respiratory pathogen was identified. The following organisms were excluded: coagulase-negative staphylococci, Micrococcus spp., Neisseria spp., Enterococcus spp., diphtheroids, and non-Streptococcus pneumoniae alpha-hemolytic streptococci, except in patients with a lung abscess or parapneumonic empyema. Organisms from infections with viral and fungal etiologies were excluded. Organisms were categorized as DRPs if they were resistant to drug regimens recommended for the treatment of CAP, e.g., if they were resistant to either ceftriaxone plus azithromycin or levofloxacin. Because the rates of levofloxacin resistance or ceftriaxone resistance among the S. pneumoniae isolates at the study institutions were low (0% to 5%), isolates from patients with positive pneumococcal urinary antigen results were categorized as CAP drug sensitive.
Risk factors for pneumonia due to DRPs.
After reviewing the available literature, the following risk factors for pneumonia due to DRPs were identified and included in the data collection: prior antibiotic use (3, 16, 28, 29, 31, 32, 37–42), prior hospitalization (an inpatient stay of more than 48 h) (16, 31–33, 37, 39–41, 43–46), residence in a long-term-care facility (including facilities providing long-term acute care and facilities providing skilled nursing care and inpatient rehabilitation but not assisted living or group home facilities) (16, 29, 31, 33, 37, 38, 40, 44, 45, 47), tube feeding (nasogastric/jejunal feeding or feeding via a percutaneous gastrostomy tube) (3, 31, 32, 41), chronic lung disease (defined as chronic obstructive, interstitial, or other structural disease, including bronchiectasis but excluding cystic fibrosis) (17, 29, 31, 38, 39, 42, 44, 45), immunosuppression (by daily treatment with systemic corticosteroids consisting of ≥15 mg prednisone; with immunomodulating agents, such as calcineurin inhibitors and mycophenolic acid, or with biologic agents, such as rituximab and tumor necrosis factor inhibitors, or because of heritable immunodeficiency, such as common variable immune deficiency; active hematologic or myeloproliferative malignancy; neutropenia [absolute neutrophil count, <1,000 cells/μl]; or a CD4 count of <200 cells/μl in patients with HIV infection or AIDS) (31, 32, 37–39), chronic kidney disease (creatinine clearance, ≤60 μg/min) (33, 37, 42, 44, 45), active wound care at the time of admission (31, 48), infusion therapy (chemotherapy or treatment with antibiotics or other intravenous medications) (29), poor functional status (defined as a Karnofsky score of <70 or nonambulatory status) (3, 28, 32, 47), aspiration risk (17), diabetes mellitus (29, 40), current smoking (39), severity of pneumonia (defined in three different ways: intensive care unit [ICU] admission, mechanical ventilator support, and predicted percent 30 day all-cause mortality using the electronic adaptation of the CURB-65 [eCURB] score [49]) (29, 30, 33, 40), cerebrovascular disease (40, 44), cognitive impairment (40, 42), prior colonization with a DRP or MRSA strain (16, 29, 32, 50, 51), gastric acid suppression (with histamine H2 antagonists or proton pump inhibitors) within 14 days (32), and the presence of an indwelling catheter (47). In order to evaluate the temporal association with certain exposures, we collected quantitative data on the time from the previous hospitalization, nursing home residence, and antibiotic use, all within the last 180 days.
Statistical analysis.
Univariate analysis of categorical variables was performed using the χ2 test or Fisher's exact test, and continuous variables were analyzed using the unpaired t test or Mann-Whitney U test for nonparametrically distributed data. A P value of <0.05 was considered significant, and all analyses were two-tailed. The correlation between the timing of antibiotic use, hospitalization, and residence in a long-term-care facility and pneumonia due to a DRP was evaluated using a grouped data plot analysis and logistic regression to find the temporal cutoff point in the continuous data with the highest odds ratio for pneumonia due to a DRP. The 95% confidence intervals (CIs) for proportions and the significance of differences between proportions were calculated by the Z test and the modified Wilson method, respectively (52) (Vassarstats.net [R. Lowry, Poughkeepsie, NY, 2015]).
Logistic regression models were constructed with DRPs. An initial set of candidate risk factor variables was selected a priori on the basis of the factors reproducibly associated with pneumonia due to DRPs in the literature. Ten variables significant at a P value of <0.20 in univariate analysis were entered into the initial model and then sequentially examined for their contribution to the model. Goodness of fit was confirmed using the Hosmer-Lemeshow calculation. Test performance characteristics, including the area under the receiver operator curve (AUROC), were computed using standard methods. The significance of the difference between AUROCs was determined by the Z test (53). The SPSS Statistics for Windows program, version 22.0 (IBM Corp., Armonk, NY), was used for all other statistical analysis, except where noted above.
A clinical prediction score was derived by the sequential inclusion of risk factors and evaluation of test performance characteristics (sensitivity, specificity, and AUROC) after each addition. Initial derivation models included only factors that were statistically significantly associated with pneumonia due to DRPs in the final regression model. Several probabilistic models weighting risk factors according to the regression β coefficient were evaluated. Cumulative models that simply summed equally weighted variables were also evaluated. Ultimately, the optimum test performance was achieved by use of a hybrid cumulative and probabilistic model, built by including four major risk factors identified by regression analysis, weighted at two points each, plus a broad set of minor risk factors weighted equally at one point each. The HCAP criteria and six other previously described prediction models (the Park [31], Shindo [32], Shorr [33], Schreiber [38], Aliberti [44], and Niederman [54] models) were applied to the derivation cohort, and test performance characteristics were compared to the novel prediction score.
Validation cohort.
The novel prediction score was then validated in an independent multicenter, observational cohort. Patients with pneumonia admitted to four geographically separate U.S. tertiary care facilities from May 2013 until May 2014 were prospectively identified. The inclusion criteria and clinical and radiographic definitions of pneumonia used for the derivation cohort were applied to patients within 48 h of admission to determine enrollment eligibility. Cases were screened for enrollment by real-time review of hospital admissions and microbiology data. Only cases with a positive microbiological diagnosis were included in the analysis. Demographic, microbiology, and risk factor data were collected via manual medical record review. Risk factor data that were required for calculation of the DRIP score but that were not available in the electronic medical record were collected during a bedside interview with the patient or designee after informed consent was obtained. The validation study was observational, and investigators did not influence clinical management, nor was microbiological testing standardized across the enrollment centers. Enrollment in the validation cohort was terminated during the 12th month of a planned 1-year enrollment period after accrual of a number of cases equivalent to the number in the derivation cohort after screening approximately 1,450 admissions. Twenty-six cases were enrolled at the Mayo Clinic Arizona site, 47 cases were enrolled at the University of Connecticut, 35 cases were enrolled at LDS Hospital (Salt Lake City, UT), and 92 were enrolled at the Intermountain Medical Center (Murray, UT). The novel prediction score, HCAP criteria, and seven previously published prediction models (42) were applied to the validation cohort, and the sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV), accuracy, and AUROC were calculated.
RESULTS
Derivation cohort.
The mean age for the derivation cohort was 63.1 years; most patients had multiple comorbidities, of which chronic pulmonary disease (41.5%) was the most common (Table 1). A total of 17.5% of the patients had poor functional status, and 12.5% were immunosuppressed. The median predicted 30-day mortality according to the eCURB score was 11.2%, corresponding to pneumonia severity index class IV (55). The observed in-hospital mortality rate was 8.5%. Twenty-one percent of patients met the HCAP criteria. Roughly 60% of the patients received initial empirical beta-lactam or macrolide therapy, and approximately 40% of patients received antipseudomonal or anti-MRSA therapy. DRPs were recovered from 25% of cases; MRSA comprised 11.7% of all organisms, and Pseudomonas aeruginosa accounted for 6.1%. Overall, Streptococcus pneumoniae was the most common pathogen identified (44.1%) (Table 2).
TABLE 1.
Demographic and clinical data
| Characteristic | Value for group |
|
|---|---|---|
| Derivation group (n = 200) | Validation group (n = 200) | |
| Mean (SD) age (yr) | 63.1 (16.7) | 65.2 (17.5) |
| No. (%) of male patients | 118 (59.0) | 103 (51.50) |
| No. (%) of patients with the following comorbidity: | ||
| Chronic pulmonary disease | 83 (41.5) | 86 (43.0) |
| Renal insufficiency | 64 (32.0) | 67 (33.5) |
| Congestive heart failure | 46 (23.0) | 42 (21.0) |
| Hepatic disease | 21 (10.5) | 18 (9.0) |
| Cognitive impairment | 22 (11.0) | 25 (12.5) |
| Diabetes mellitus | 55 (27.5) | 47 (23.5) |
| Neurological disease | 37 (18.5) | 25 (12.5) |
| Poor functional status | 35 (17.5) | 27 (13.5) |
| Influenza-like illness | 6 (3.0) | 12 (6.0) |
| Active malignancya | 14 (7.0) | 26 (13.0) |
| Mean (SD) no. of comorbidities | 1.7 (1.2) | 1.7 (1.2) |
| No. (%) of patients in whom HCAP criteria were met | 42 (21.0) | 99 (49.5) |
| No. (%) of patients with: | ||
| Sepsis | 167 (83.5) | 154 (77.0) |
| Positive-pressure ventilationb | 70 (35.0) | 57 (28.5) |
| Mean (SD) eCURB score (%) for predicted 30-day mortality | 11.2c (12.9) | 12.9 (16.6) |
| No. (%) of patients with in-hospital mortality | 17 (8.5) | 18 (9.0) |
| No. (%) of patients hospitalized in the last 180 days | 57 (28.5) | 93 (46.5) |
| Median (25%–75% IQRd) no. of days prior | 65 (20.8–119.5) | 37 (14.0–72.5) |
| No. (%) of patients in long-term care facility in the last 180 days | 31 (15.5) | 42 (21.0) |
| Median (25%–75% IQR) no. of days prior | 1 (1.0–1.0) | 1 (1.0–4.0) |
| No. (%) of patients receiving antibiotics in the last 180 days | 71 (35.5) | 109 (54.5 |
| Median (25%–75% IQR) no. of days prior | 60 (21.5–96.3) | 20 (1.25–56.8) |
| No. (%) of patients receiving: | ||
| Hemodialysis | 7 (3.5) | 10 (5.0) |
| Infusion therapy | 14 (7.0) | 7 (3.5) |
| Wound care | 20 (10.0) | 19 (9.5) |
| Immunosuppression | 25 (12.5) | 40 (20.0) |
| Gastric acid suppression | 70 (35.0) | 99 (49.5) |
| Tube feedinge | 12 (6.0) | 22 (11.0) |
| No. (%) of patients with: | ||
| History of infection with a DRP | 22 (11.0) | 28 (14.0) |
| History of MRSA colonization | 15 (7.5) | 19 (9.5) |
| No. (%) of patients for whom the initial antibiotic choice was a (an): | ||
| Macrolide | 128 (64.0) | 74 (37.0) |
| Beta-lactam | 119 (59.5) | 78 (39.0) |
| Antipseudomonal | 87 (43.5) | 110 (55.0) |
| Fluoroquinolone | 55 (27.5) | 89 (44.5) |
| Doxycycline | 0 (0) | 1 (0.5) |
| Aminoglycoside | 0 (0) | 1 (0.5) |
| Anti-MRSA agent | 77 (38.5) | 104 (52.0) |
| No. (%) of patients in whom the initial antibiotic therapy was appropriate | 182 (91.0) | 178 (89.0) |
| No. (%) of patients infected with a DRP | 50 (25.0) | 66 (33.0) |
Solid organ tumor or hematologic malignancy excluding in situ and skin cancers; not in remission.
Includes mechanical ventilation and noninvasive modalities.
Equivalent to pneumonia severity index (PSI) class IV (55).
IQR, interquartile range.
Includes all routes of enteral feeding, including nasogastric/jejunal and percutaneous gastrostomy.
TABLE 2.
Microbiology
| Source or organism | Derivation group (n = 213) |
Validation group (n = 218) |
||
|---|---|---|---|---|
| No. of patientsa | % of patients | No. of patientsa | % of patients | |
| Sources | ||||
| Sputum | 72 | 33.8 | 110 | 50.5 |
| Tracheal aspirate | 15 | 7.0 | 4 | 1.8 |
| BAL fluid | 10 | 4.7 | 7 | 3.2 |
| Pleural fluid | 3 | 1.4 | 9 | 4.1 |
| Blood | 67 | 31.4 | 45 | 20.6 |
| Urine antigen test | 44 | 20.7 | 33 | 15.1 |
| Respiratory film array PCR | 1 | 0.5 | 9 | 4.1 |
| Organisms | ||||
| Gram-positive bacteria | 154 | 72.3 | 128 | 60.1 |
| Streptococcus pneumoniae | 94 (0) | 44.1 | 54 (0) | 24.8 |
| MRSA | 25 (25) | 11.7 | 32 (32) | 14.7 |
| MSSA | 20 (0) | 9.4 | 30 (0) | 13.8 |
| Group A beta-hemolytic streptococcus | 5 (0) | 2.3 | 3 (0) | 1.4 |
| Streptococcus anginosus group | 3 (0) | 1.4 | 6 (0) | 2.8 |
| Other Gram-positive bacteria | 7 (0) | 3.3 | 3 (1) | 1.4 |
| Gram-negative bacteria | 49 | 23.0 | 73 | 34.3 |
| Pseudomonas aeruginosa | 13 (13) | 6.1 | 21 (21) | 9.6 |
| Escherichia coli | 10 (3) | 4.7 | 14 (8) | 6.4 |
| Klebsiella pneumoniae | 5 (1) | 2.3 | 7 (0) | 3.2 |
| Klebsiella oxytoca | 0 (0) | 0.0 | 4 (0) | 1.8 |
| Haemophilus influenzae | 9 (0) | 4.2 | 6 (0) | 2.8 |
| Enterobacter aerogenes | 2 (1) | 0.9 | 4 (2) | 1.8 |
| Enterobacter cloacae | 1 (0) | 0.5 | 2 (1) | 0.9 |
| Moraxella catarrhalis | 0 (0) | 0.0 | 3 (0) | 1.4 |
| Stenotrophomonas maltophilia | 3 (3) | 1.4 | 2 (2) | 0.9 |
| Serratia marcescens | 0 (0) | 0.0 | 3 (0) | 1.4 |
| Other Gram-negative bacteria | 6 (4) | 2.8 | 7 (4) | 3.2 |
| Atypical bacteria | 7 | 3.3 | 15 | 6.9 |
| Mycoplasma pneumoniae | 1 (0) | 0.5 | 8 (0) | 3.7 |
| Legionella pneumophila | 6 (0) | 2.8 | 6 (0) | 2.8 |
| Chlamydophila pneumoniae | 0 (0) | 0.0 | 1 (0) | 0.5 |
| Other bacteria | ||||
| Peptostreptococcus sp. | 1 (0) | 0.5 | 1 (0) | 0.5 |
| Prevotella sp. | 1 (1) | 0.5 | 0 (0) | 0.0 |
| Veillonella sp. | 1 (1) | 0.5 | 0 (0) | 0.0 |
| Mycobacterium avium-M. intracellulare complex | 0 (0) | 0.0 | 1 (1) | 0.5 |
| Ceftriaxone- and macrolide-sensitive bacteria | 165 | 77.5 | 150 | 68.8 |
| Fluoroquinolone-sensitive bacteria | 173 | 81.2 | 164 | 75.2 |
| DRPs (all isolates) | 52 | 24.4 | 72 | 33 |
| DRPs (cases) | 50 | 25 | 66 | 33 |
Values in parentheses in this column indicate the number of patients infected with DRPs.
The following factors were associated with pneumonia due to DRPs in the univariate analysis: chronic pulmonary disease, poor functional status, number of comorbidities, hospitalization within the last 180 days, residence in a long-term-care facility in the last 180 days, antibiotic use in the last 180 days, wound care, gastric acid suppression, tube feeding, any infection with a DRP in the last year, and MRSA colonization in the last year. Temporal analysis revealed that both hospitalization and antibiotic use within 60 days of hospital admission for pneumonia were associated with the strongest risk of pneumonia due to DRPs, whereas the risk of pneumonia due to DRPs was the strongest for patients admitted directly from a long-term-care facility.
Specific predictors of pneumonia due to MRSA and Gram-negative DRPs were also evaluated. In the univariate analysis, poor functional status, recent hospitalization, residence in a long-term-care facility, antibiotic use, wound care, gastric acid suppression, tube feeding, prior infection with a DRP, and MRSA colonization were associated with pneumonia due to MRSA. In a multiple logistic regression model, only residence in a long-term-care facility (adjusted odds ratio [AOR], 7.45 [95% CI, 1.85 to 30.0]) and wound care in the last 30 days (AOR, 4.79 [95% CI, 1.23 to 18.8]) remained significant. Risk factors that remained significantly associated with pneumonia due to Gram-negative DRPs in a multivariable regression model included chronic pulmonary disease (AOR, 5.6 [95% CI, 1.48 to 21.17]; P < 0.011), antibiotic use in the last 60 days (AOR, 5.32 [95% CI, 1.86 to 15.26]; P < 0.002), tube feeding (AOR, 12.93 [95% CI, 2.28 to 73.37]; P < 0.004), and prior infection with a DRP (AOR, 5.24 [95% CI, 1.56 to 17.66]; P < 0.008).
In a logistic regression analysis for pneumonia due to DRPs, prior antibiotic use within 60 days (AOR, 10.5 [95% CI, 2.25 to 22.9], P < 0.001), tube feeding (AOR, 34.6 [95% CI, 2.8 to 424.8], P = 0.006), a history of DRP in the year prior to admission (AOR, 20.2 [95% CI, 4.7 to 87.3]; P < 0.001), residence in a long-term-care facility (AOR, 3.9 [95% CI, 1.1 to 13.6]; P = 0.03), and chronic pulmonary disease (AOR, 3.1 [95% CI, 1.2 to 8.4] correlated most closely with pneumonia due to DRPs. A novel clinical prediction rule, the drug resistance in pneumonia (DRIP) score, was ultimately derived by including four major risk factors for pneumonia due to DRPs (antibiotic use within 60 days, tube feeding, residence in a long-term-care facility, and prior infection with a DRP), each of which was assigned 2 points, and six minor risk factors (chronic pulmonary disease, prior hospitalization within 60 days, poor functional status, MRSA colonization, wound care, and gastric acid suppression), each of which was assigned 1 point, for a total of 14 possible points (Table 3).
TABLE 3.
DRIP score calculation
| Characteristic | No. of points |
|---|---|
| Major risk factors | |
| Antibiotic use within previous 60 days | 2 |
| Residence in a long-term-care facility | 2 |
| Tube feeding | 2 |
| Prior infection with a DRP (1 yr) | 2 |
| Minor risk factors | |
| Hospitalization within previous 60 days | 1 |
| Chronic pulmonary disease | 1 |
| Poor functional status | 1 |
| Gastric acid suppression | 1 |
| Wound care | 1 |
| MRSA colonization (1 yr) | 1 |
| Total no. of points possible | 14 |
In the derivation cohort, the DRIP score was predictive of pneumonia due to DRPs, demonstrated by an AUROC value of 0.90 (95% CI, 0.84 to 0.95) (Fig. 2). This was higher than that obtained by the use of the HCAP criteria (AUROC, 0.60 [95% CI, 0.50 to 0.70]) and all other predictive models evaluated (Table 4). The DRIP score best differentiated a low risk from a high risk of pneumonia due to a DRP at a threshold of ≥4 points with a sensitivity of 0.76, a specificity of 0.91, a positive predictive value (PPV) of 0.73, and a negative predictive value (NPV) of 0.92. Using this threshold, the overall accuracy (the percentage of cases in which the antibiotic spectrum that would have been recommended on the basis of the DRIP score classification was appropriate for the recovered organism) was 87.0% (95% CI, 81.4 to 91.2%), while an inadequate spectrum (undertreatment) would have been recommended in 6.0% (95% CI, 3.3 to 10.5%) of cases, and an unnecessary spectrum (overtreatment) would have been recommended in 7.0% (95% CI, 4.0 to 11.7%) of cases.
FIG 2.
Receiver operator curves for derivation cohort.
TABLE 4.
Test performance with derivation cohort
| Prediction model | Cutoff score | Sensitivity (95% CI) | Specificity (95% CI) | PPV (95% CI) | NPV (95% CI) | AUROC (95% CI) | Inadequate spectrum treatment % (95% CI) | Overall accuracy % (95% CI) | Unnecessary spectrum treatment % (95% CI) |
|---|---|---|---|---|---|---|---|---|---|
| DRIP score | 4 | 0.76 (0.62–0.86) | 0.91 (0.85–0.95) | 0.73 (0.59–0.84) | 0.92 (0.86–0.96) | 0.9 (0.84–0.95) | 6.0 (3.3–10.5) | 87.0 (81.4–91.2) | 7.0 (4.0–11.7) |
| HCAP criteria | 1 | 0.36 (0.23–0.51) | 0.84 (0.77–0.89) | 0.43 (0.28–0.59) | 0.80 (0.72–0.85) | 0.6 (0.5–0.7) | 16.0 (11.4–22.0) | 72.0 (65.2–78.0) | 12.0 (8.0–17.5) |
| Schreiber | 2 | 0.52 (0.38–0.66) | 0.79 (0.71–0.85) | 0.45 (0.32–0.58) | 0.83 (0.76–0.89) | 0.68 (0.6–0.77) | 12.0 (8.0–17.5) | 72.0 (65.2–78.0) | 16.0 (11.4–22.0) |
| Shorr | 1 | 0.9 (0.77–0.96) | 0.29 (0.22–0.37) | 0.3 (0.23–0.38) | 0.90 (0.77–0.96) | 0.71 (0.63–0.8) | 2.5 (0.9–6.1) | 44.0 (37.1–51.2) | 53.5 (46.3–60.5) |
| Niederman | 2 | 0.82 (0.68–0.91) | 0.66 (0.58–0.73) | 0.45 (0.34–0.55) | 0.92 (0.84–0.96) | 0.8 (0.72–0.87) | 4.5 (2.2–8.6) | 70.0 (63.1–76.2) | 25.5 (19.7–32.2) |
| Shindo | 3 | 0.64 (0.49–0.77) | 0.79 (0.72–0.85) | 0.51 (0.38–0.63) | 0.87 (0.8–0.92) | 0.77 (0.68–0.86) | 9.0 (5.6–14.1) | 75.5 (68.8–81.2) | 15.5 (10.9–21.4) |
| Aliberti | 3 | 0.7 (0.55–0.82) | 0.57 (0.48–0.65) | 0.35 (0.26–0.45) | 0.85 (0.76–0.91) | 0.66 (0.57–0.74) | 7.5 (4.4–12.3) | 60.0 (52.8–66.8) | 32.5 (26.2–39.5) |
| Park | 3 | 0.5 (0.36–0.64) | 0.86 (0.79–0.91) | 0.54 (0.39–0.69) | 0.84 (0.77–0.89) | 0.75 (0.67–0.84) | 12.5 (8.4–18.1) | 77.0 (70.4–82.5) | 10.5 (6.8–15.8) |
When the performance characteristics of the DRIP score and the HCAP criteria were modeled for the full derivation cohort, including both microbiology-negative and -positive cases (n = 1,509), the estimated measures of diagnostic accuracy again favored the DRIP score over the HCAP criteria: sensitivities, 0.76 and 0.36 for the DRIP score and HCAP criteria, respectively; specificities, 0.89 and 0.80 for the DRIP score and HCAP criteria, respectively; PPVs, 0.19 and 0.06 for the DRIP score and HCAP criteria, respectively; and NPVs, 0.99 and 0.97 for the DRIP score and HCAP criteria, respectively.
Validation cohort.
In the validation cohort, the mean patient age was 65.2 years (Table 1). Forty-three percent of the patients had chronic pulmonary disease (43%), 13.5% had poor functional status, and 20.0% were immunosuppressed. Nearly half met the HCAP criteria (49.5%), and the number of patients that received extended-spectrum antibiotics was greater than the number that received a regimen recommended for the treatment of CAP. Positive-pressure ventilation was delivered to 28.5% of patients, and the median predicted risk of 30-day mortality according to the eCURB score was 16.6%. The observed in-hospital mortality rate was 9.0%. DRPs were recovered from 33.0% of cases; MRSA comprised 14.7% of all isolates, and P. aeruginosa comprised 9.6% of all isolates (Table 2). S. pneumoniae comprised 24.8% of the isolates.
When applied to the validation cohort, the DRIP score showed a strong correlation with pneumonia due to DRPs at all scoring levels: the AUROC value was 0.88 (95% CI, 0.82 to 0.93) (Fig. 3). Again, the DRIP score performed significantly better than the HCAP criteria (AUROC, 0.72 [95% CI, 0.64 to 0.79]; P = 0.02) and the other models when applied to the same cohort (Table 5). At a threshold of ≥4 points, the DRIP score differentiated a high risk versus a low risk of pneumonia due to drug-resistant pathogens with a sensitivity of 0.82 (95% CI, 0.67 to 0.88), a specificity of 0.81 (95% CI, 0.73 to 0.87), a PPV of 0.68 (95% CI, 0.56 to 0.78), and an NPV of 0.90 (95% CI, 0.81 to 0.93). By comparison, the performance of the HCAP criteria was less favorable: sensitivity was 0.79 (95% CI, 0.67 to 0.88), specificity was 0.65 (95% CI, 0.56 to 0.73), PPV was 0.53 (95% CI, 0.42 to 0.63), and NPV was 0.86 (95% CI, 0.77 to 0.92). The overall accuracy differed between the DRIP score and the HCAP criteria: 81.5% (95% CI, 74.2 to 85.6%) and 69.5% (95% CI, 62.5 to 75.7%), respectively (P = 0.005). Use of the DRIP score rather than the HCAP criteria would have resulted in a 46% relative reduction in the rate of unnecessary antibiotic spectrum use without an increase in the rate of inadequate treatment (rates of use of unnecessary spectrum, 23.5% and 12.5% for the HCAP criteria and the DRIP score, respectively; absolute difference, 11.0 percentage points [95% CI, 3.5 to 18.4 percentage points]) (P = 0.004).
FIG 3.
Receiver operator curves for validation cohort.
TABLE 5.
Test performance with validation cohort
| Prediction model | Cutoff score | Sensitivity (95% CI) | Specificity (95% CI) | PPV (95% CI) | NPV (95% CI) | AUROC (95% CI) | Inadequate spectrum treatment % (95% CI) | Overall accuracy % (95% CI) | Unnecessary spectrum treatment % (95% CI) |
|---|---|---|---|---|---|---|---|---|---|
| DRIP | 4 | 0.82 (0.67–0.88) | 0.81 (0.73–0.87) | 0.68 (0.56–0.78) | 0.90 (0.81–0.93) | 0.88 (0.82–0.93) | 6.0 (3.5–10.2) | 81.5 (74.2–85.6) | 12.5 (8.4–18.1) |
| HCAP | 1 | 0.79 (0.67–0.88) | 0.65 (0.56–0.73) | 0.53 (0.42–0.63) | 0.86 (0.77–0.92) | 0.72 (0.64–0.79) | 7.0 (4.2–11.4) | 69.5 (62.5–75.7) | 23.5 (17.9–30.1) |
| Schreiber | 2 | 0.71 (0.55–0.79) | 0.66 (0.58–0.74) | 0.51 (0.39–0.61) | 0.82 (0.72–0.88) | 0.74 (0.67–0.81) | 9.5 (6.8–15.8) | 68.0 (60.0–73.4) | 22.5 (17.0–29.0) |
| Shorr | 1 | 0.88 (0.77–0.94) | 0.4 (0.31–0.48) | 0.42 (0.34–0.50) | 0.87 (0.75–0.94) | 0.77 (0.70–0.85) | 4.0 (1.9–8.0) | 55.5 (48.3–62.5) | 40.5 (33.7–47.7) |
| Niederman | 2 | 0.91 (0.81–0.96) | 0.53 (0.44–0.62) | 0.49 (0.40–0.58) | 0.92 (0.83–0.97) | 0.80 (0.74–0.87) | 3.0 (1.2–6.7) | 65.5 (58.4–72.0) | 31.5 (25.2–38.5) |
| Shindo | 3 | 0.83 (0.72–0.91) | 0.6 (0.51–0.68) | 0.5 (0.41–0.60) | 0.88 (0.79–0.94) | 0.79 (0.73–0.86) | 5.5 (2.9–9.9) | 67.5 (60.5–73.8) | 27.0 (21.1–33.8) |
| Shindo + 1 MRSA-specific factor | ≥3 and 2 + 1 MRSA-specific factor | 0.77 (0.65—0.86) | 0.72 (0.63–0.79) | 0.57 (0.46–0.68) | 0.86 (0.78–0.92) | NA | 7.5 (4.4–12.3) | 73.5 (67.0–79.4) | 19.0 (14.0–25.3) |
| Aliberti | 0.5 | 0.88 (0.77–0.94) | 0.55 (0.46–0.64) | 0.49 (0.40–0.58) | 0.90 (0.81–0.95) | 0.73 (0.66–0.8) | 4.0 (1.9–8.0) | 66.0 (58.9–72.4) | 30.0 (23.8–36.9) |
| Park | 3 | 0.77 (0.65–0.86) | 0.72 (0.63–0.79) | 0.57 (0.46–0.68) | 0.86 (0.78–0.92) | 0.82 (0.75–0.88) | 7.5 (4.4–12.3) | 73.5 (66.7–79.4) | 19.0 (14.0–25.3) |
| PES | 5 | 0.71 (0.59–0.81) | 0.65 (0.56–0.73) | 0.50 (0.40–0.60) | 0.82 (0.73–0.89) | 0.74 (0.67–0.81) | 9.5 (6.0–14.7) | 67.0 (0.60–0.73) | 23.5 (17.9–30.1) |
All cases with false-negative results (DRIP score of <4 and pneumonia due to DRPs) and false-positive results (DRIP score of ≥4 but no pneumonia due to DRPs) were reviewed in detail to identify potential risk factors for future study. MRSA was the most common DRP recovered in cases with DRIP scores of <4 (41.6%), while non-lactose-fermenting Gram-negative rods, including P. aeruginosa, were present in 33.3% of cases. Many of the cases with false-negative results had severe chronic pulmonary disease requiring supplemental oxygen, systemic or inhaled steroids, or structural abnormalities. Intravenous drug use, homelessness, and/or a psychiatric comorbidity was observed in several of the cases of pneumonia due to MRSA with DRIP scores of <4. The majority of the cases with false-positive results and DRIP scores of 4 were due to S. pneumoniae or methicillin-susceptible S. aureus (MSSA). The ability of the DRIP score to predict pneumonia due to MRSA was also compared to that of two prediction tools (32, 40) designed to specifically identify the risk of pneumonia due to MRSA in the validation cohort (Table 6).
TABLE 6.
MRSA-specific prediction
| Prediction model | Cutoff score | Sensitivity (95% CI) | Specificity (95% CI) | PPV (95% CI) | NPV (95% CI) | Lack of anti-MRSA % (95% CI) | Overall accuracy % (95% CI) | Unnecessary anti-MRSA treatment % (95% CI) |
|---|---|---|---|---|---|---|---|---|
| DRIP score for MRSA | 4 | 0.84 (0.66–0.94) | 0.69 (0.61–0.76) | 0.34 (0.24–0.46) | 0.96 (0.90–0.98) | 2.5 (0.9–6.1) | 81.5 (74.2–85.6) | 26.0 (20.2–32.8) |
| Shorr | 2 | 0.97 (0.82–1) | 0.30 (0.23–0.37) | 0.21 (0.15–0.28) | 0.98 (0.88–1) | 0.5 (0.03–3.2) | 40.5 (33.7–47.7) | 59.0 (51.8–65.8) |
| Shindo + 1 MRSA-specific factor | ≥3 and 2 + 1 MRSA-specific factor | 0.84 (0.66–0.94) | 0.63 (0.55–0.70) | 0.30 (0.21–0.41) | 0.95 (0.89–0.98) | 2.5 (0.9–6.1) | 66.5 (59.5–72.9) | 31.0 (24.8–38.0) |
DISCUSSION
The DRIP score is a novel cumulative, probabilistic model for predicting the risk of pneumonia due to drug-resistant bacteria. In both a retrospective, multicenter derivation cohort and a prospective observational multicenter validation cohort, the DRIP score demonstrated performance characteristics superior to those of the HCAP criteria. At a threshold score of ≥4, the DRIP score effectively differentiated a high and a low probability of the risk of pneumonia due to DRPs, suggesting its utility as a clinical decision tool to guide empirical antibiotic selection. This threshold is fulfilled by the presence of either 2 major risk factors, 1 major risk factor plus 2 minor risk factors, or 4 minor risk factors.
The DRIP score addresses several important limitations of the HCAP criteria and further advances the foundational work of other models for prediction of the risk of pneumonia due to DRP. First, the lack of specificity of the model with the HCAP criteria has resulted in dramatic increases in the unnecessary use of broad-spectrum antibiotics. The results of this study suggest that the DRIP score may have the potential to reduce the rate of recommendation of the use of broad-spectrum antibiotics but retain a sensitivity sufficient to avoid inadequate initial antibiotic treatment. There are several possible explanations for the improved performance characteristics of the DRIP score compared to those of the HCAP criteria and, to a lesser extent, those of the other models. The primary contributor may be the inclusion of a broader set of risk factors that have all been reproducibly associated with pneumonia due to DRPs in previous studies. While no single factor is strongly predictive, inclusion of a larger number of factors improved the specificity of the model. Other contributors include differences in the definition of a DRP, variations in cohort characteristics, and selection of a treatment threshold that optimized both sensitivity and sensitivity.
This study also clarifies previously identified risk factors. For example, prior antibiotic use is not included in the model with the HCAP criteria but appears to be a major contributor to the development of resistance. Although this association was observed for antibiotic use up to 6 months prior to the onset of pneumonia, we found that the most predictive interval was 60 days prior to admission. This is consistent with the findings of other studies, in which the development of antibiotic resistance was most common for patients who had been given antibiotics within the previous 60 days (26, 56, 57). In contrast to the HCAP criteria, we also found that hospitalization within the previous 60 days was also the optimum interval for the risk of pneumonia due to DRPs. This may represent the interval in which the burden of oropharyngeal colonization with potential DRPs is the highest before repopulation with normal flora (46). We were unable to further elucidate some risk factors. For example, ICU admission may portend an additional risk of pneumonia due to DRPs beyond that of hospitalization alone. Similarly, we observed that the risk of pneumonia due to DRPs appears to increase with the severity of chronic pulmonary disease, but this effect was most prominent for Gram-negative DRPs. Immunosuppression has been associated with the risk of pneumonia due to DRPs, but we did not observe this correlation. This may be definitional and colinear with severe pulmonary disease in patients requiring systemic corticosteroids. It is possible that if it is defined as such, severe chronic pulmonary disease would be better categorized as a major risk factor for pneumonia due to DRPs. Further research is needed to quantify the risk of pneumonia due to DRPs by specific immune-modulating therapies and dose.
In contrast to other studies (29, 33, 40), we did not find a significant association between the severity of illness, defined by either eCURB, ICU admission, or the presence of sepsis plus mechanical ventilation, and DRPs. This may be because patients evaluated in the emergency department not requiring admission were not included or because DRPs may not, in fact, be more virulent than other organisms. However, we recognize that in clinical practice, a tolerance for prediction of false-negative results resulting in inadequate initial therapy is conditioned upon a clinical margin for error. A strength of the Niederman model (30) is the tiered strategy, in which the drug recommendation is based in part on severity. We investigated this effect in our cohort, applying a lower threshold of 3 for broad-spectrum antibiotic treatment for patients with a high severity of illness, and found that the sensitivity (0.86) and the rate of false-negative results (4.5% [95% CI, 2.2 to 8.6%]) were modestly improved but were done so at the expense of decreased specificity (0.74) and an increase in the rate of potential overtreatment (17.5% [95% CI, 12.6 to 23.6%]) approaching that observed by the use of the HCAP criteria.
Like other studies, we found a significant overlap between risk factors for pneumonia due to MRSA and Gram-negative DRPs (32, 40, 58). Unlike in the study of Shindo et al. (32), neither hemodialysis nor congestive heart failure was associated with pneumonia due to MRSA in our cohort. Attempts to incorporate the MRSA-specific criteria identified in other studies (32, 40) into the model with the DRIP score resulted in significant reductions in both overall sensitivity and specificity. Consequently, the DRIP score is not able to discriminate the risk of pneumonia due to MRSA from that due to Gram-negative DRPs. However, because of its increased overall specificity, the DRIP score would still conserve anti-MRSA therapy compared to other models. When the predictive value of the DRIP score for the risk of pneumonia due to MRSA was compared to that of the Shorr model at a threshold of ≥2 points and the Shindo algorithm recommending anti-MRSA coverage for scores of ≥3 and a score of 2 plus at least one MRSA-specific factor, the DRIP score exhibited a good balance of sensitivity and specificity. The Shindo model demonstrated performance characteristics very similar to those demonstrated by the DRIP score. The score developed by Shorr et al. for predicting the risk of pneumonia due to MRSA was highly sensitive at the threshold recommended by the authors, but it was nonspecific, recommending unnecessary anti-MRSA therapy in more than half of the cases. At thresholds consisting of a score of 4 or greater, the Shorr model performed similarly to the model with the DRIP score and Shindo's algorithm. In clinical practice, a reasonable approach may be to combine empirical anti-MRSA coverage for patients at risk for pneumonia due to DRPs with rapid de-escalation, leveraging the strong negative predictive value of PCR for MRSA in a nasal swab specimen (50, 59).
A strength of this study is the inclusion of multiple U.S. centers with diverse demographic and microbiological populations. By design, the characteristics of the patients in the validation cohort differed from those of the patients in the derivation cohort in order to confirm that the DRIP score would perform adequately with a population different from the one with which it was derived. The prospective design of the validation cohort facilitated rigorous clinical and radiographic confirmation of pneumonia and allowed the accurate capture of risk factors, such as prior antibiotic use, not reliably available in medical records. Because true validation of any diagnostic tool requires a “gold standard” reference method (60), we included only patients with a microbiological diagnosis because microbiology-negative cases are technically not evaluable with respect to the reference, e.g., the presence of a DRP or not. However, this methodology has several limitations.
The derivation and validation cohorts were limited by their small sample sizes, which may affect the generalizability of the results of this study. Despite careful efforts to identify and include bacterial culture-positive cases, not all patients underwent standardized microbiological testing. Inclusion of organisms identified by sputum cultures may also be inaccurate due to the detection of colonizing species rather than true lower respiratory tract pathogens. Finally, the test performance characteristics obtained when the DRIP score was applied to the validation cohort do not reflect those that would be obtained in a real-world application. Evaluation with a larger cohort of patients with pneumonia due to all causes is needed to confirm these results.
Conclusion.
As the microbiology of community-acquired pneumonia evolves, prediction of the risk of disease due to drug-resistant pathogens remains challenging for clinicians. Studies have failed to show improvements in outcomes when empirical extended-spectrum antibiotic therapy is prescribed on the basis of the use of the HCAP criteria; improved methods of prediction are needed. In the prospective multicenter U.S. cohort of microbiology-positive pneumonia cases described here, use of a model with a novel scoring approach, the DRIP scoring approach, was more predictive of the risk of pneumonia due to DRP than use of the HCAP criteria, and the DRIP score may have the potential to decrease antibiotic overutilization. Additional observational validation with larger cohorts of patients with pneumonia due to all causes will be necessary in order to confirm this. Furthermore, prospective implementation studies evaluating mortality and other outcome measures are warranted in order to determine whether the DRIP score or other scores are clinically useful.
ACKNOWLEDGMENTS
B.J.W., N.C.D., M.M., H.R.V., E.S., and K.D. conceived of and designed the study. All authors acquired, analyzed, or interpreted the data. B.J.W., N.C.D., and E.S. drafted the manuscript. All authors critically reviewed the final manuscript.
All authors have completed and submitted the ICMJE Form for Disclosure of Potential Conflicts of Interest. N.C.D. reports the receipt of grants or consultation fees from Cerexa and Pfizer in the past but has no current relationship with these companies. E.S. and K.D. report the receipt of grants from Pfizer unrelated to the current study. B.J.W. reports the receipt of consultation fees from Allergan unrelated to this study. No other financial disclosures are reported.
Funding Statement
A portion of this study was funded by a grant from the Intermountain Research and Medical Foundation, which had no role in study design and conduct of the study, data collection or interpretation, manuscript preparation, review, or decision to submit for publication.
REFERENCES
- 1.Buie VC, Owings MF, DeFrances CJ, Golosinskiy A. 2010. National Hospital Discharge Survey: 2006 summary. Vital Health Stat 13 2010:1–79. [PubMed] [Google Scholar]
- 2.Heron M, Hoyert DL, Murphy SL, Xu JQ, Kochanek KD, Tejada-Vera B. 2009. Deaths: final data for 2006. Natl Vital Stat Rep 57:1–134. [PubMed] [Google Scholar]
- 3.Shindo Y, Sato S, Maruyama E, Ohashi T, Ogawa M, Hashimoto N, Imaizumi K, Sato T, Hasegawa Y. 2009. Health-care-associated pneumonia among hospitalized patients in a Japanese community hospital. Chest 135:633–640. doi: 10.1378/chest.08-1357. [DOI] [PubMed] [Google Scholar]
- 4.Zilberberg MD, Shorr AF, Micek ST, Mody SH, Kollef MH. 2008. Antimicrobial therapy escalation and hospital mortality among patients with health-care-associated pneumonia: a single-center experience. Chest 134:963–968. doi: 10.1378/chest.08-0842. [DOI] [PubMed] [Google Scholar]
- 5.Dean NC, Bateman KA, Donnelly SM, Silver MP, Snow GL, Hale D. 2006. Improved clinical outcomes with utilization of a community-acquired pneumonia guideline. Chest 130:794–799. doi: 10.1378/chest.130.3.794. [DOI] [PubMed] [Google Scholar]
- 6.McCabe C, Kirchner C, Zhang H, Daley J, Fisman DN. 2009. Guideline-concordant therapy and reduced mortality and length of stay in adults with community-acquired pneumonia: playing by the rules. Arch Intern Med 169:1525–1531. doi: 10.1001/archinternmed.2009.259. [DOI] [PubMed] [Google Scholar]
- 7.Mandell LA, Wunderink RG, Anzueto A, Bartlett JG, Campbell GD, Dean NC, Dowell SF, File TM Jr, Musher DM, Niederman MS, Torres A, Whitney CG. 2007. Infectious Diseases Society of America/American Thoracic Society consensus guidelines on the management of community-acquired pneumonia in adults. Clin Infect Dis 44(Suppl 2):S27–S72. doi: 10.1086/511159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kollef MH, Shorr A, Tabak YP, Gupta V, Liu LZ, Johannes RS. 2005. Epidemiology and outcomes of health-care-associated pneumonia: results from a large US database of culture-positive pneumonia. Chest 128:3854–3862. doi: 10.1378/chest.128.6.3854. [DOI] [PubMed] [Google Scholar]
- 9.American Thoracic Society, Infectious Diseases Society of America. 2005. Guidelines for the management of adults with hospital-acquired, ventilator-associated, and healthcare-associated pneumonia. Am J Respir Crit Care Med 171:388–416. doi: 10.1164/rccm.200405-644ST. [DOI] [PubMed] [Google Scholar]
- 10.Berger A, Edelsberg J, Oster G, Huang X, Weber DJ. 2014. Patterns of initial antibiotic therapy for community-acquired pneumonia in U.S. hospitals, 2000 to 2009. Am J Med Sci 347:347–356. doi: 10.1097/MAJ.0b013e318294833f. [DOI] [PubMed] [Google Scholar]
- 11.Jones BE, Jones M, Huttner B, Stoddard G, Brown KA, Stevens V, Greene T, Sauer B, Madaras-Kelly K, Rubin M, Goetz MB, Samore M. 2015. Trends in antibiotic use and nosocomial pathogens in hospitalized veterans with pneumonia at 128 medical centers, 2006-2010. Clin Infect Dis 61:1403–1410. doi: 10.1093/cid/civ629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Attridge RT, Frei CR, Restrepo MI, Lawson KA, Ryan L, Pugh MJ, Anzueto A, Mortensen EM. 2011. Guideline-concordant therapy and outcomes in healthcare-associated pneumonia. Eur Respir J 38:878–887. doi: 10.1183/09031936.00141110. [DOI] [PubMed] [Google Scholar]
- 13.Kett DH, Cano E, Quartin AA, Mangino JE, Zervos MJ, Peyrani P, Cely CM, Ford KD, Scerpella EG, Ramirez JA, Improving Medicine through Pathway Assessment of Critical Therapy of Hospital-Acquired Pneumonia (IMPACT-HAP) Investigators. 2011. Implementation of guidelines for management of possible multidrug-resistant pneumonia in intensive care: an observational, multicentre cohort study. Lancet Infect Dis 11:181–189. doi: 10.1016/S1473-3099(10)70314-5. [DOI] [PubMed] [Google Scholar]
- 14.Troitino AX, Porhomayon J, El-Solh AA. 2013. Guideline-concordant antimicrobial therapy for healthcare-associated pneumonia: a systematic review and meta-analysis. Lung 191:229–237. doi: 10.1007/s00408-013-9462-8. [DOI] [PubMed] [Google Scholar]
- 15.Chalmers JD, Rother C, Salih W, Ewig S. 2014. Healthcare-associated pneumonia does not accurately identify potentially resistant pathogens: a systematic review and meta-analysis. Clin Infect Dis 58:330–339. doi: 10.1093/cid/cit734. [DOI] [PubMed] [Google Scholar]
- 16.Gross AE, Van Schooneveld TC, Olsen KM, Rupp ME, Bui TH, Forsung E, Kalil AC. 2014. Epidemiology and predictors of multidrug-resistant community-acquired and health care-associated pneumonia. Antimicrob Agents Chemother 58:5262–5268. doi: 10.1128/AAC.02582-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Chalmers JD, Taylor JK, Singanayagam A, Fleming GB, Akram AR, Mandal P, Choudhury G, Hill AT. 2011. Epidemiology, antibiotic therapy, and clinical outcomes in health care-associated pneumonia: a UK cohort study. Clin Infect Dis 53:107–113. doi: 10.1093/cid/cir274. [DOI] [PubMed] [Google Scholar]
- 18.Chen JI, Slater LN, Kurdgelashvili G, Husain KO, Gentry CA. 2013. Outcomes of health care-associated pneumonia empirically treated with guideline-concordant regimens versus community-acquired pneumonia guideline-concordant regimens for patients admitted to acute care wards from home. Ann Pharmacother 47:9–19. doi: 10.1345/aph.1R322. [DOI] [PubMed] [Google Scholar]
- 19.El Solh AA, Akinnusi ME, Alfarah Z, Patel A. 2009. Effect of antibiotic guidelines on outcomes of hospitalized patients with nursing home-acquired pneumonia. J Am Geriatr Soc 57:1030–1035. doi: 10.1111/j.1532-5415.2009.02279.x. [DOI] [PubMed] [Google Scholar]
- 20.Taylor SP, Taylor BT. 2013. Health care-associated pneumonia in haemodialysis patients: clinical outcomes in patients treated with narrow versus broad spectrum antibiotic therapy. Respirology 18:364–368. doi: 10.1111/j.1440-1843.2012.02306.x. [DOI] [PubMed] [Google Scholar]
- 21.Burgess LD, Drew RH. 2014. Comparison of the incidence of vancomycin-induced nephrotoxicity in hospitalized patients with and without concomitant piperacillin-tazobactam. Pharmacotherapy 34:670–676. doi: 10.1002/phar.1442. [DOI] [PubMed] [Google Scholar]
- 22.Gomes DM, Smotherman C, Birch A, Dupree L, Della Vecchia BJ, Kraemer DF, Jankowski CA. 2014. Comparison of acute kidney injury during treatment with vancomycin in combination with piperacillin-tazobactam or cefepime. Pharmacotherapy 34:662–669. doi: 10.1002/phar.1428. [DOI] [PubMed] [Google Scholar]
- 23.Jeffres MN, Isakow W, Doherty JA, Micek ST, Kollef MH. 2007. A retrospective analysis of possible renal toxicity associated with vancomycin in patients with health care-associated methicillin-resistant Staphylococcus aureus pneumonia. Clin Ther 29:1107–1115. doi: 10.1016/j.clinthera.2007.06.014. [DOI] [PubMed] [Google Scholar]
- 24.Meaney CJ, Hynicka LM, Tsoukleris MG. 2014. Vancomycin-associated nephrotoxicity in adult medicine patients: incidence, outcomes, and risk factors. Pharmacotherapy 34:653–661. doi: 10.1002/phar.1423. [DOI] [PubMed] [Google Scholar]
- 25.Chalmers JD, Al-Khairalla M, Short PM, Fardon TC, Winter JH. 2010. Proposed changes to management of lower respiratory tract infections in response to the Clostridium difficile epidemic. J Antimicrob Chemother 65:608–618. doi: 10.1093/jac/dkq038. [DOI] [PubMed] [Google Scholar]
- 26.Costelloe C, Metcalfe C, Lovering A, Mant D, Hay AD. 2010. Effect of antibiotic prescribing in primary care on antimicrobial resistance in individual patients: systematic review and meta-analysis. BMJ 340:c2096. doi: 10.1136/bmj.c2096. [DOI] [PubMed] [Google Scholar]
- 27.Aliberti S, Zanaboni AM, Blasi F. 2012. Pneumonia in the community caused by multidrug-resistant organisms: keep working on probabilistic scores. Clin Infect Dis 54:1519–1520. doi: 10.1093/cid/cis260. [DOI] [PubMed] [Google Scholar]
- 28.El Solh AA, Pietrantoni C, Bhat A, Bhora M, Berbary E. 2004. Indicators of potentially drug-resistant bacteria in severe nursing home-acquired pneumonia. Clin Infect Dis 39:474–480. doi: 10.1086/422317. [DOI] [PubMed] [Google Scholar]
- 29.Madaras-Kelly KJ, Remington RE, Fan VS, Sloan KL. 2012. Predicting antibiotic resistance to community-acquired pneumonia antibiotics in culture-positive patients with healthcare-associated pneumonia. J Hosp Med 7:195–202. doi: 10.1002/jhm.942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Maruyama T, Fujisawa T, Okuno M, Toyoshima H, Tsutsui K, Maeda H, Yuda H, Yoshida M, Kobayashi H, Taguchi O, Gabazza EC, Takei Y, Miyashita N, Ihara T, Brito V, Niederman MS. 2013. A new strategy for healthcare-associated pneumonia: a 2-year prospective multicenter cohort study using risk factors for multidrug-resistant pathogens to select initial empiric therapy. Clin Infect Dis 57:1373–1383. doi: 10.1093/cid/cit571. [DOI] [PubMed] [Google Scholar]
- 31.Park SC, Kim EY, Kang YA, Park MS, Kim YS, Kim SK, Chang J, Jung JY. 2013. Validation of a scoring tool to predict drug-resistant pathogens in hospitalised pneumonia patients. Int J Tuberc Lung Dis 17:704–709. doi: 10.5588/ijtld.12.0723. [DOI] [PubMed] [Google Scholar]
- 32.Shindo Y, Ito R, Kobayashi D, Ando M, Ichikawa M, Shiraki A, Goto Y, Fukui Y, Iwaki M, Okumura J, Yamaguchi I, Yagi T, Tanikawa Y, Sugino Y, Shindoh J, Ogasawara T, Nomura F, Saka H, Yamamoto M, Taniguchi H, Suzuki R, Saito H, Kawamura T, Hasegawa Y. 2013. Risk factors for drug-resistant pathogens in community-acquired and healthcare-associated pneumonia. Am J Respir Crit Care Med 188:985–995. doi: 10.1164/rccm.201301-0079OC. [DOI] [PubMed] [Google Scholar]
- 33.Shorr AF, Zilberberg MD, Micek ST, Kollef MH. 2008. Prediction of infection due to antibiotic-resistant bacteria by select risk factors for health care-associated pneumonia. Arch Intern Med 168:2205–2210. doi: 10.1001/archinte.168.20.2205. [DOI] [PubMed] [Google Scholar]
- 34.Self WH, Wunderink RG, Williams DJ, Barrett TW, Baughman AH, Grijalva CG. 2015. Comparison of clinical prediction models for resistant bacteria in community-onset pneumonia. Acad Emerg Med 22:730–740. doi: 10.1111/acem.12672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Dean NC, Jones BE, Ferraro JP, Vines CG, Haug PJ. 2013. Performance and utilization of an emergency department electronic screening tool for pneumonia. JAMA Intern Med 173:699–701. doi: 10.1001/jamainternmed.2013.3299. [DOI] [PubMed] [Google Scholar]
- 36.Bartlett R. 1974. Medical microbiology: quality cost and clinical relevance. John Wiley & Sons, Inc., New York, NY. [Google Scholar]
- 37.Shorr AF, Zilberberg MD, Reichley R, Kan J, Hoban A, Hoffman J, Micek ST, Kollef MH. 2012. Validation of a clinical score for assessing the risk of resistant pathogens in patients with pneumonia presenting to the emergency department. Clin Infect Dis 54:193–198. doi: 10.1093/cid/cir813. [DOI] [PubMed] [Google Scholar]
- 38.Schreiber MP, Chan CM, Shorr AF. 2010. Resistant pathogens in nonnosocomial pneumonia and respiratory failure: is it time to refine the definition of health-care-associated pneumonia? Chest 137:1283–1288. doi: 10.1378/chest.09-2434. [DOI] [PubMed] [Google Scholar]
- 39.Wooten DA, Winston LG. 2013. Risk factors for methicillin-resistant Staphylococcus aureus in patients with community-onset and hospital-onset pneumonia. Respir Med 107:1266–1270. doi: 10.1016/j.rmed.2013.05.006. [DOI] [PubMed] [Google Scholar]
- 40.Shorr AF, Myers DE, Huang DB, Nathanson BH, Emons MF, Kollef MH. 2013. A risk score for identifying methicillin-resistant Staphylococcus aureus in patients presenting to the hospital with pneumonia. BMC Infect Dis 13:268. doi: 10.1186/1471-2334-13-268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Jung JY, Park MS, Kim YS, Park BH, Kim SK, Chang J, Kang YA. 2011. Healthcare-associated pneumonia among hospitalized patients in a Korean tertiary hospital. BMC Infect Dis 11:61. doi: 10.1186/1471-2334-11-61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Prina E, Ranzani OT, Polverino E, Cilloniz C, Ferrer M, Fernandez L, Puig de la Bellacasa J, Menendez R, Mensa J, Torres A. 2015. Risk factors associated with potentially antibiotic-resistant pathogens in community-acquired pneumonia. Ann Am Thorac Soc 12:153–160. doi: 10.1513/AnnalsATS.201407-305OC. [DOI] [PubMed] [Google Scholar]
- 43.Venditti M, Falcone M, Corrao S, Licata G, Serra P. 2009. Outcomes of patients hospitalized with community-acquired, health care-associated, and hospital-acquired pneumonia. Ann Intern Med 150:19–26. doi: 10.7326/0003-4819-150-1-200901060-00005. [DOI] [PubMed] [Google Scholar]
- 44.Aliberti S, Di Pasquale M, Zanaboni AM, Cosentini R, Brambilla AM, Seghezzi S, Tarsia P, Mantero M, Blasi F. 2012. Stratifying risk factors for multidrug-resistant pathogens in hospitalized patients coming from the community with pneumonia. Clin Infect Dis 54:470–478. doi: 10.1093/cid/cir840. [DOI] [PubMed] [Google Scholar]
- 45.Aliberti S, Cilloniz C, Chalmers JD, Zanaboni AM, Cosentini R, Tarsia P, Pesci A, Blasi F, Torres A. 2013. Multidrug-resistant pathogens in hospitalised patients coming from the community with pneumonia: a European perspective. Thorax 68:997–999. doi: 10.1136/thoraxjnl-2013-203384. [DOI] [PubMed] [Google Scholar]
- 46.Filius PM, Gyssens IC, Kershof IM, Roovers PJ, Ott A, Vulto AG, Verbrugh HA, Endtz HP. 2005. Colonization and resistance dynamics of Gram-negative bacteria in patients during and after hospitalization. Antimicrob Agents Chemother 49:2879–2886. doi: 10.1128/AAC.49.7.2879-2886.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Fisch J, Lansing B, Wang L, Symons K, Cherian K, McNamara S, Mody L. 2012. New acquisition of antibiotic-resistant organisms in skilled nursing facilities. J Clin Microbiol 50:1698–1703. doi: 10.1128/JCM.06469-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Webb BJ, Dangerfield BS, Pasha JS, Agrwal N, Vikram HR. 2012. Guideline-concordant antibiotic therapy and clinical outcomes in healthcare-associated pneumonia. Respir Med 106:1606–1612. doi: 10.1016/j.rmed.2012.08.003. [DOI] [PubMed] [Google Scholar]
- 49.Jones BE, Jones J, Bewick T, Lim WS, Aronsky D, Brown SM, Boersma WG, van der Eerden MM, Dean NC. 2011. CURB-65 pneumonia severity assessment adapted for electronic decision support. Chest 140:156–163. doi: 10.1378/chest.10-1296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Dangerfield B, Chung A, Webb B, Seville MT. 2014. Predictive value of methicillin-resistant Staphylococcus aureus (MRSA) nasal swab PCR assay for MRSA pneumonia. Antimicrob Agents Chemother 58:859–864. doi: 10.1128/AAC.01805-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Scanvic A, Denic L, Gaillon S, Giry P, Andremont A, Lucet JC. 2001. Duration of colonization by methicillin-resistant Staphylococcus aureus after hospital discharge and risk factors for prolonged carriage. Clin Infect Dis 32:1393–1398. doi: 10.1086/320151. [DOI] [PubMed] [Google Scholar]
- 52.Newcombe RG. 1998. Two-sided confidence intervals for the single proportion: comparison of seven methods. Stat Med 17:857–872. doi: 10.1002/(SICI)1097-0258(19980430)17:8<857::AID-SIM777>3.0.CO;2-E. [DOI] [PubMed] [Google Scholar]
- 53.Hanley JA, McNeil BJ. 1982. The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 143:29–36. doi: 10.1148/radiology.143.1.7063747. [DOI] [PubMed] [Google Scholar]
- 54.Brito V, Niederman MS. 2009. Healthcare-associated pneumonia is a heterogeneous disease, and all patients do not need the same broad-spectrum antibiotic therapy as complex nosocomial pneumonia. Curr Opin Infect Dis 22:316–325. doi: 10.1097/QCO.0b013e328329fa4e. [DOI] [PubMed] [Google Scholar]
- 55.Fine MJ, Auble TE, Yealy DM, Hanusa BH, Weissfeld LA, Singer DE, Coley CM, Marrie TJ, Kapoor WN. 1997. A prediction rule to identify low-risk patients with community-acquired pneumonia. N Engl J Med 336:243–250. doi: 10.1056/NEJM199701233360402. [DOI] [PubMed] [Google Scholar]
- 56.Goldstein RC, Husk G, Jodlowski T, Mildvan D, Perlman DC, Ruhe JJ. 2014. Fluoroquinolone- and ceftriaxone-based therapy of community-acquired pneumonia in hospitalized patients: the risk of subsequent isolation of multidrug-resistant organisms. Am J Infect Control 42:539–541. doi: 10.1016/j.ajic.2014.01.005. [DOI] [PubMed] [Google Scholar]
- 57.Kuster SP, Rudnick W, Shigayeva A, Green K, Baqi M, Gold WL, Lovinsky R, Muller MP, Powis JE, Rau N, Simor AE, Walmsley SL, Low DE, McGeer A, Toronto Invasive Bacterial Diseases Network. 2014. Previous antibiotic exposure and antimicrobial resistance in invasive pneumococcal disease: results from prospective surveillance. Clin Infect Dis 59:944–952. doi: 10.1093/cid/ciu497. [DOI] [PubMed] [Google Scholar]
- 58.Metersky M, Frei CR, Mortensen EM. 2015. Predictors of Pseudomonas and methicillin-resistant Staphylococcus aureus in hospitalized patients with healthcare-associated pneumonia. Respirology 21:157–163. doi: 10.1111/resp.12651. [DOI] [PubMed] [Google Scholar]
- 59.Boyce JM, Pop OF, Abreu-Lanfranco O, Hung WY, Fisher A, Karjoo A, Thompson B, Protopapas Z. 2013. A trial of discontinuation of empiric vancomycin therapy in patients with suspected methicillin-resistant Staphylococcus aureus health care-associated pneumonia. Antimicrob Agents Chemother 57:1163–1168. doi: 10.1128/AAC.01965-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.TDR Diagnostics Evaluation Expert Panel, Banoo S, Bell D, Bossuyt P, Herring A, Mabey D, Poole F, Smith PG, Sriram N, Wongsrichanalai C, Linke R, O'Brien R, Perkins M, Cunningham J, Matsoso P, Nathanson CM, Olliaro P, Peeling RW, Ramsay A. 2010. Evaluation of diagnostic tests for infectious diseases: general principles. Nat Rev Microbiol 8:S17–S29. [PubMed] [Google Scholar]