Abstract
The contribution of CXCL12/CXCR4/CXCR7 axis to cancer progression has been increasingly recognized. However, its role in thyroid cancer development remains unclear. The present study aimed to examine the expression and function of CXCL12 and its receptors in thyroid cancer. The expression of CXCL12/CXCR4/CXCR7 in human tissue specimens of papillary, follicular, medullary, and anaplastic thyroid carcinoma, follicular adenoma, Hashimoto's thyroiditis and nodular goiter were examined by immunohistochemistry using a tissue microarray. CXCR4 and CXCR7 were over-expressed in human thyroid cancer cells K1 by transduction of recombinant lentivirus. The effect of overexpression of CXCR4 and CXCR7 on K1 cell proliferation and invasion and the molecular mechanism underlying the effect were investigated. CXCL12 was exclusively expressed in papillary thyroid carcinoma tissue but absent in other types of thyroid malignancies and benign lesions. CXCR7 was widely expressed in the endothelial cells of all types of malignancy but only occasionally detected in benign lesions. CXCR4 was expressed in 62.5% of papillary thyroid carcinoma tissue specimens and in 30–40% of other types of malignancy, and it was either absent or weakly expressed in benign lesions. CXCL12 stimulated the invasion and migration of K1 cells overexpressing CXCR4, but did not affect K1 cells overexpressing CXCR7. K1 cell proliferation was not affected by overexpression of CXCR4 or CXCR7. Overexpression of CXCR4 in K1 cells significantly increased AKT and ERK phosphorylation and markedly induced the expression and activity of matrix metalloproteinase-2 (MMP-2). Thus, CXCL12 may be an effective diagnostic marker for papillary thyroid carcinoma, and CXCL12/CXCR4/CXCR7 axis may contribute to thyroid cancer development by regulating cancer cell migration and invasion via AKT and ERK signaling and MMP-2 activation.
Keywords: AKT, CXCL12/CXCR4/CXCR7, ERK, invasion, matrix metalloproteinase-2, thyroid cancer
Introduction
The incidence of thyroid cancer has increased continuously worldwide for the past 3 decades (1,2); particularly, the incidence increases dramatically in Chinese women (3). Thyroid cancer is ranked the fourth most common cancer among women in urban Beijing in 2012, whereas it was not even in the top 10 cancers in 1999 (3). Papillary thyroid carcinoma (PTC) exclusively accounts for the increase in the incidence of total thyroid cancer, whereas the incidences of follicular thyroid carcinoma (FTC), medullary thyroid carcinoma (MTC), and anaplastic thyroid carcinoma (ATC) have not changed substantially (2). The mechanism underlying thyroid cancer progression remains unclear. CXCL12/CXCR4/CXCR7 axis has been found to play a critical role in cancer metastasis and progression in many types of cancer (4), thus, may also contribute to thyroid cancer progression.
CXCL12, which is also named as stromal cell-derived factor-1 (SDF-1), is a CXC-chemokine and was originally identified from bone marrow stromal cells (5). In addition to the well-recognized receptor CXCR4 (6,7), CXCR7 has recently been identified as another receptor for CXCL12 (8,9). The molecular mechanism underlying the biological effect of CXCL12 is associated with CXCR4-mediated activation of G protein-coupled signaling molecules, including ERK1/2, MAPK, JNK and AKT (10,11). The function of CXCR7 remains unclear; some reports suggested that it may be a co-receptor for CXCR4 to enhance CXCL12-mediated G-protein signaling (12,13); others indicated that CXCR7 may behave as a decoy receptor to sequester CXCL12, resulting in a CXCL12 gradient so to facilitate the stimulation of CXCR4-associated downstream signaling (14,15). The role of CXCL12/ CXCR4/CXCR7 axis in cancer metastasis and progression has been increasingly recognized in many types of cancer (4). Both in vitro and in vivo studies have shown that the role of CXCL12/CXCR4 axis in cancer progression is mainly to facilitate metastasis and mobilization of cancer cells (4).
The protein expression of CXCL12/CXCR4/CXCR7 has been confirmed in multiple human cancers. In prostate cancer, CXCR7 expression increases as cancer aggressiveness increases, and CXCL12 level at the preferential metastatic sites, such as the bone, liver, and kidney, were higher than that at non-metastatic tissue (16,17). In breast cancer, CXCR4 is highly expressed in cancer tissue but absent in normal breast tissue (18). Strong CXCR7 expression has been found in tumor-associated blood vessels in breast and lung cancer tissue but not on normal vasculature (19). In thyroid cancer, the expressions of CXCL12, CXCR4 and CXCR7 have also been detected in cancer tissue (20–23). However, the contribution of CXCL12/CXCR4/CXCR7 axis to thyroid cancer progression and the possible CXCR4-mediated signaling pathways in thyroid cancer cells remain unclear. The present study was performed to fill this knowledge gap. The aim of this study was to investigate the expression of CXCL12, CXCR4 and CXCR7 in malignant and benign thyroid lesions and examine the role of CXCR4 and CXCR7 in human thyroid cancer cell proliferation, migration and invasion.
Materials and methods
Tissue specimen
The present study was approved by the Institutional Review Board of Shanghai Cancer Center of Fudan University. Signed informed consent was obtained from each patient. Surgical tissue specimens of PTC (40 cases), FTC (10 cases), MTC (10 cases), ATC (10 cases), follicular adenoma (FA, 10 cases), Hashimoto's thyroiditis (HT, 10 cases), and nodular goiter (NG, 10 cases) were collected. The malignancy of thyroid cancer tissue was confirmed by histopathological examination. The benign thyroid tissue specimens were also confirmed to be free of malignancy. Paraffin-embedded tissue specimens were used to prepare a tissue microarray (TMA) for immunohistochemical staining. Each TMA contained 120 (12×10) cores and 2 cores of each sample were printed on a TMA. The core size was 1 mm (diameter) × 4 μm (thickness).
Immunohistochemistry
TMAs were incubated with the following primary antibodies: monoclonal mouse anti-human CXCR4 (1:400; Abcam, Cambridge, MA, USA), anti-human CXCR7 (1:50; R&D Systems, Inc., Minneapolis, MN, USA), or anti-human CXCL12 (1:50; R&D Systems). Briefly, 4-μm-thick sections of TMAs were transferred to adhesive slides and maintained in a drying oven at 60°C for 2 h. The standard heat epitope was deparaffinized using EZ Prep (Ventana Medical Systems, Inc., Tucson, AZ, USA) at 75°C for 4 min. During the heat pretreatment step, cell-conditioning solutions (Ventana Medical Systems) containing Tris/Borate/EDTA were used at 100°C for 92 min. The antibodies were pre-diluted, and the tissue sections were incubated with the diluted antibodies at 37°C for 16 min (for CXCL12) and 30 min (for CXCR7). Subsequently, the tissue sections were incubated with an appropriate reagent from the ultraView Universal DAB kit (Ventana Medical Systems) at 37°C for 8 min and counterstained with Harris's haematoxylin for 2 min. The stained TMAs were observed under a microscope. The staining intensity (CXCL12 in tumor cells and CXCR7 in endothelial cells) was scored based on the following criteria: 0 represents no staining or faint staining intensity in 10% cells; 1+ represents faint staining in >10% of cells; 2+ represents moderate staining; 3+ represents strong staining. The tissue specimen was considered positive for CXCL12 or CXCR7 when the staining intensity score was 1+, 2+, or 3+ and negative when the score was 0.
Cell culture
Human thyroid cancer K1 cells were purchased from Guangzhou Jenniobio Biotechnology Co., Ltd. (Guangzhou, China). The cells were cultured in Dulbecco's modified Eagle's media (DMEM; HyClone Laboratories, Inc., Logan, UT, USA) supplemented with 10% fetal bovine serum (FBS; Invitrogen, Carlsbad, CA, USA), 100 units/ml penicillin and 100 μg/ml streptomycin (Thermo Fisher Scientific, Waltham, MA, USA) at 37°C in 5% CO2.
Recombinant lentivirus construction and transduction
Recombinant lentivirus expressing CXCR4 or CXCR7 was constructed using the ViruaPower™ Lentiviral Expression systems (Invitrogen). K1 cells were seeded in 24-well plates at the density of 1×105 cells/well and incubated overnight. The culture media were replaced with 2 ml fresh media containing 6 μg/ml polybrene (Sigma, St. Louis, MO, USA) and the recombinant lentivirus, and then the cells were incubated with polybrene and the recombinant lentivirus at 37°C for 4 h. The media were replaced with 2 ml fresh media and the cells were incubated overnight. Lentivirus transduction efficiency was determined by lentivirus expressing green fluorescent protein (GFP). Apparent GFP expression appeared 48 h after transduction and GFP level reached the peak 72 h after transduction. Thus, K1 cells were cultured for 72 h after transduction with recombinant lentivirus and then used for RT-PCR, proliferation and Transwell invasion assay.
Transwell invasion assay
Cell invasion was determined by using 24-well Transwell invasion chamber (8 μm pore size; Corning, Tewksbury, MA, USA). A total of 200 μl cell suspensions in serum-free media (5×104/well) were added in the upper chambers. The lower chambers were filled with 800 μl DMEM media containing either 10% FBS or 100 ng/ml CXCL12. After incubation for 24 h, the cells on the upper surface of the membrane in the upper chamber were removed by cotton swaps and the cells on the other side of the membrane were fixed with methanol for 30 min, air dried and stained with 0.1% crystal violet solution for 10 min. The penetrated cells were then counted under a microscope. Three to 5 fields were randomly selected on each membrane and the average number of cells was used. The experiment was repeated at least 3 times.
Cell proliferation
Cell proliferation was determined using the Cell Counting kit-8 (CCK-8 kit; Dojindo Laboratories, Kumamoto, Japan). Cells were plated in 96-well plates at a density of 8×103 cells/well in 100 μl serum-free media containing 100 ng/ml recombinant CXCL12 (PeproTech, Rocky Hill, NJ, USA) or media containing 10% FBS. The cells were incubated for 24 or 48 h. After incubation, 10 μl of CCK-8 solution was added to each well and incubated at 37°C for 1–4 h. The media were then removed and the dark blue formazan crystals were dissolved in 150 μl of DMSO. The absorbance at 450 nm was measured in a microplate reader (Corning Costar, Corning, NY, USA, USA). The experiment was repeated independently at least 3 times.
Western blot analysis
Cells (2×105 cells/well) were cultured in 6-well plates and incubated at 37°C until confluency. After being washed twice with ice-cold PBS, the cells were lysed in RIPA buffer (Beyotime Institute of Biotechnology, Haimen, China) containing protease inhibitor cocktail (1:1,000 dilution; Sigma) and phosphatase inhibitor cocktail (1:1,000 dilution; Sigma). The cell lysate was centrifuged at 12,000 × g for 10 min at 4°C. Protein concentration in the supernatants was determined using a BCA kit (Beyotime Institute of Biotechnology). A total of 30 μg protein extract were separated by SDS-PAGE and transferred to PVDF membranes. The membrane was blocked for 1 h at room temperature in 5% non-fat dry milk in TBST (10 mmol/l Tris, pH 8.0, 165 mmol/l NaCl, 0.05% Tween-20). The membrane was then incubated with the following primary antibodies: rabbit anti-human CXCR4, CXCR7, p-Akt1 (B-1), p-ERK (B-5), β-actin, GAPDH and ERK1/2 (Santa Cruz Biotechnology, Santa Cruz, CA, USA) at room temperature for 1 h or at 4°C overnight. After extensive washing, the membrane was incubated with the secondary antibody, goat anti-rabbit IgG conjugated with horseradish peroxidase (Cell Signaling Technology, Danvers, MA, USA), at a dilution of 1:5,000 at room temperature for 1.5 h. Target proteins were visualized using an enhanced chemiluminescence kit (Beyotime Institute of Biotechnology). For quantification, the intensity of protein signal was evaluated using Quantity One system (Bio-Rad Laboratories, Hercules, CA, USA). The experiment was repeated at least 3 times.
Quantitative reverse-transcription polymerase chain reaction
Total RNA was extracted from K1 cells using the RNAiso Plus kit (Takara Bio, Shiga, Japan) according to the manufacturer's instruction. A total of 500 ng total RNA was reverse transcribed to cDNA using the reverse transcriptase M-MLV RT (Takara) and random primers in a 20 μl reaction system. The RT product (1 μl) was used as the template for quantitative PCR (qPCR). The SYBR Premix Ex Taq™ II system (Takara Bio) was used for qPCR reaction. The primer sequences are: MMP-2 forward, 5′-CGACCACAGCCAACTACGAT-3′ and reverse, 5′-GTCAGGAGAGGCCCCATAGA-3′; MMP-9 forward, 5′-TCTATGGTCCTCGCCCTGAA-3′ and reverse, 5′-CATCGTCCACCGGACTCAAA-3′. The qPCR condition was 95°C for 2 min and then 40 cycles of 95°C for 10 sec, 60°C for 30 sec, and 70°C for 45 sec. Human β-actin was used as the internal control. The qPCR was performed on the Applied Biosystems 7500 detection system. Cycle threshold (Ct) values for all samples were determined. The ΔΔCt method was used to calculate the expression levels of target genes in experimental groups relative to control group.
Gel zymography
Culture media were mixed with non-reducing loading buffer (4% SDS, 0.25 mmol/l Tris-HCl, 40% glycerol, 0.1% bromophenol blue) at 1:3 ratio, and 15 μl of the mixture was loaded in 10% SDS-PAGE gel containing 1 mg/ml gelatin. After electrophoresis, the gel was washed twice, 45 min each time, in a washing buffer (2.5% Triton X-100, 50 mmol/l Tris-HCl, 5 mmol/l CaCl2, 1 μmol/l ZnCl2, pH 7.6), and then washed again twice, 20 min each time, in a washing buffer (50 mmol/l Tris-HCl, 5 mmol/l CaCl2, 1 μmol/l ZnCl2, pH 7.6). The gel was incubated in the reaction buffer (50 mmol/l Tris-HCl, 5 mmol/l CaCl2, 1 μmol/l ZnCl2, 0.02% Brij-35, pH 7.6) at 37°C for 42 h, stained with Coomassie blue (0.05% Coomassie blue, 30% methanol, 10% acetate) for 3 h, and de-stained with buffer A (30% methanol and 10% acetate), buffer B (20% methanol and 10% acetate) and buffer C (10% methanol and 5% acetate) for 0.5, 1 h and 2 h, respectively.
Statistical analysis
Continuous variables are presented as mean ± standard deviation. The statistical analysis was performed using the software SPSS 19.0. The Chi-square test and the Fisher's exact test were used for categorical variables. P-value was 2-sided and P<0.05 was considered statistically significant.
Results
The expression of CXCR4, CXCL12 and CXCR7 in thyroid tissue specimens
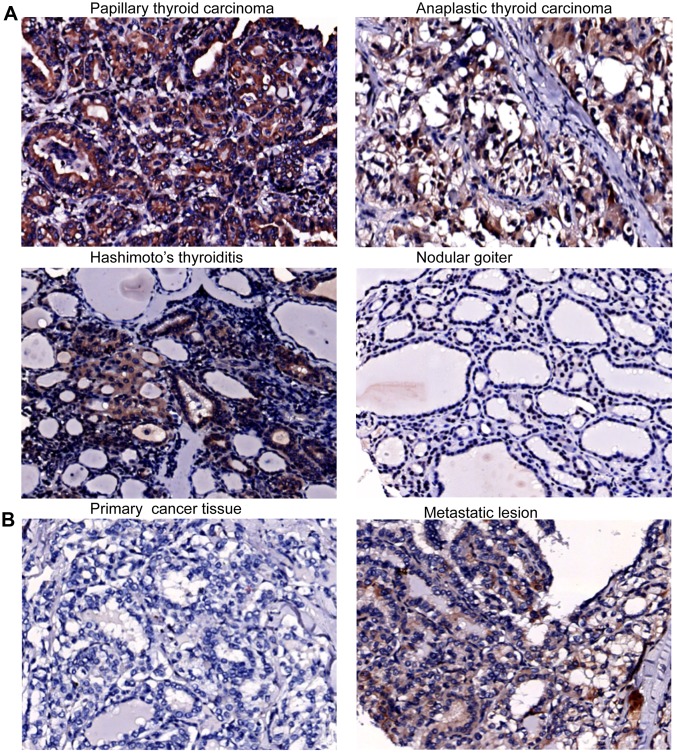
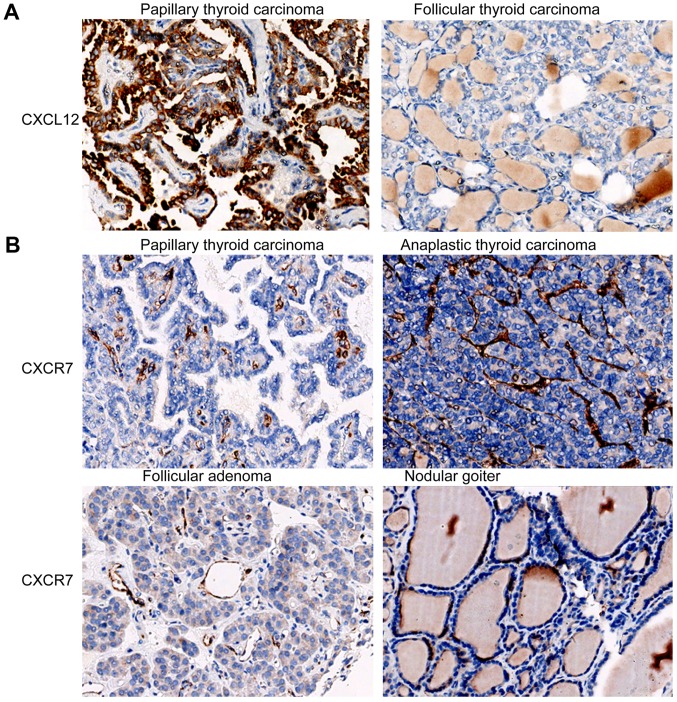
The expression of CXCR4, CXCL12 and CXCR7 varied in malignant and benign thyroid tissue specimens. CXCR4 was expressed in 62.5% of PTC specimens and in 30–40% of other types of thyroid malignancy, and benign lesions also expressed CXCR4 (Table I). The CXCR4 staining signals were diffuse and appeared in the cytoplasm of cancer cells in PTC and ATC specimens, and in HT and NG specimens, CXCR4 was predominantly in follicular cells (Fig. 1A). The proportion of specimens with positive CXCR4 in PTC with lymph node metastasis (55%) was not significantly different from that (85%) in PTC without lymph node metastasis (Table I). Notably, 3 cases of CXCR4 positive PTC with lymph node metastasis showed stronger CXCR4 staining in the metastatic lesions than in the primary thyroid cancer tissue (Fig. 1B). CXCL12 expression was found in 75% of PTC specimens and 10% of FA specimens, but was absent in FTC, MTC, ATC, HT and NG specimens (Table I). Additionally, the proportion of cases with positive CXCL12 (80%) in PTC with lymph node metastasis was similar to that (70%) in PTC without lymph node metastasis (Table I). CXCL12 staining signals were in the cytoplasm of cancer cells (Fig. 2A). These results suggest that papillary thyroid cancer cells may secret CXCL12, but other types of thyroid cancer cells may not. CXCR7 was commonly expressed in malignant thyroid tissue specimens. The proportions of cases with positive CXCR7 in PTC (97.5%), FTC (100%), MTC (100%) and ATC (100%) specimens were significantly higher than those in the benign thyroid specimens, HT (20%) and NG (10%) (All P<0.05; Table I). The proportion of positive CXCR7 in FA was 80%. Notably, CXCR7 staining signals were mainly located in the blood vessels in thyroid tissue (Fig. 2B).
Table I.
The expression of CXCL12, CXCR7 and CXCR4 in malignant and benign thyroid tissue specimens.
| Total number | CXCL12+ n (%) | CXCR7+ n (%) | CXCR4+ n (%) | |
|---|---|---|---|---|
| PTC | 40 | 30 (75) | 39 (97.5) | 25 (62.5) |
| LN− | 20 | 14 (70) | 19 (95) | 17 (85) |
| LN+ | 20 | 16 (80) | 20 (100) | 11 (55) |
| FTC | 10 | 0 (0) | 10 (100) | 3 (30) |
| MTC | 10 | 0 (0) | 10 (100) | 4 (40) |
| ATC | 10 | 0 (0) | 10 (100) | 4 (40) |
| FA | 10 | 1 (10)a | 8 (80) | 2 (20)a |
| HT | 10 | 0 (0)a | 2 (20)a | 4 (40) |
| NG | 10 | 0 (0)a | 1 (10)a | 0 (0)a |
PTC, papillary thyroid carcinoma; LN, lymph node metastasis; FTC, follicular thyroid carcinoma; MTC, medullary thyroid carcinoma; ATC, anaplastic thyroid carcinoma; FA, follicular adenoma; HT, Hashimoto's thyroiditis; NG, nodular goiter.
Fisher's exact test revealed significant difference between the indicated groups vs. PTC. P<0.05.
Figure 1.
Immunohistochemical staining of CXCR4 of thyroid tissue specimens. (A) Images of immunohistochemical staining with anti-CXCR4 for tissue specimens of papillary thyroid carcinoma, anaplastic thyroid carcinoma, Hashimoto's thyroiditis and nodular goiter. (B) Stronger CXCR4 staining in metastatic lesions than in the primary lesions.
Figure 2.
Immunohistochemical staining of XCL12 and CXCR7 of thyroid tissue specimens. (A) Images of immunohistochemical staining with anti-CXCL12 for tissue specimens of papillary thyroid carcinoma and follicular thyroid carcinoma. (B) Images of immunohistochemical staining with anti-CXCR7 for tissue specimens of papillary thyroid carcinoma, anaplastic thyroid carcinoma, follicular adenoma and nodular goiter.
Further examination of the staining intensity of the positive specimens revealed that the intensity of CXCR7 staining signals in the benign thyroid tissue specimens (FA, HT and NG) was weak, and different types of thyroid malignancies presented various CXCR7 staining intensity (Fig. 2 and Table II). The majority of ATC specimens (90%) showed moderate or strong CXCR7 expression, and for other types of thyroid malignancy, including PTC, FTC and MTC, there was 50–70% of cases with moderate or strong CXCR7 staining (Table II). CXCR4 expression level was low in benign lesions including FA, HT and NG. There was only one case of moderate or strong CXCR4 staining in the benign lesions (Table II). All the positive FTC specimens showed weak CXCR4 staining. In PTC, MTC and ATC specimens, the proportions of weak staining were similar to those of moderate or strong staining (Table II).
Table II.
The intensity of CXCL12, CXCR7 and CXCR4 staining in thyroid tissue specimens.
| CXCL12 | CXCR7 | CXCR4 | |||||
|---|---|---|---|---|---|---|---|
|
|
|
|
|||||
| Total number | Weak n (%) | Moderate or strong n (%) | Weak n (%) | Moderate or strong n (%) | Weak n (%) | Moderate or strong n (%) | |
| PTC | 40 | 15 (37.5) | 15 (37.5) | 17 (42.5) | 22 (55) | 11 (27.5) | 14 (35) |
| LN− | 20 | 7 (35) | 7 (35) | 9 (45) | 10 (50) | 6 (30) | 11 (55) |
| LN+ | 20 | 8 (40) | 8 (40) | 8 (40) | 12 (60) | 6 (30) | 5 (25) |
| FTC | 10 | 0 (0) | 0 (0) | 5 (50) | 5 (50) | 3 (30) | 0 (0) |
| MTC | 10 | 0 (0) | 0 (0) | 3 (30) | 7 (70) | 2 (20) | 2 (20) |
| ATC | 10 | 0 (0) | 0 (0) | 1 (10) | 9 (90) | 2 (20) | 2 (20) |
| FA | 10 | 1 (10) | 0 (0) | 5 (50) | 3 (30) | 2 (20) | 0 (0) |
| HT | 10 | 0 (0) | 0 (0) | 1 (10) | 1 (10) | 3 (30) | 1 (10) |
| NG | 10 | 0 (0) | 0 (0) | 1 (10) | 0 (0) | 0 (0) | 0 (0) |
PTC, papillary thyroid carcinoma; LN, lymph node metastasis; FTC, follicular thyroid carcinoma; MTC, medullary thyroid carcinoma; ATC, anaplastic thyroid carcinoma; FA, follicular adenoma; HT, Hashimoto's thyroiditis; NG, nodular goiter.
Overexpression of CXCR4 or CXCR7 in thyroid cancer cells stimulates K1 cell invasion but does not affect K1 cell proliferation
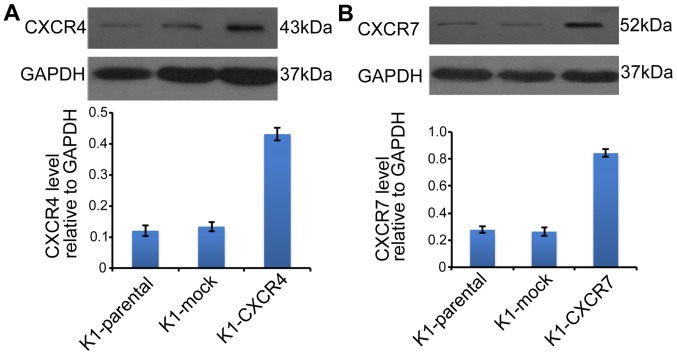
The protein levels of CXCR4 (Fig. 3A) and CXCR7 (Fig. 3B) were markedly increased in K1 cells transduced with CXCR4 or CXCR7 recombinant lentivirus. CXCL12 (100 ng/ml) markedly stimulated the invasion of K1 cells overexpressing CXCR4 (K1-CXCR4); the CXCL12-induced K1-CXCR4 invasion was completely blocked by functional neutralizing antibody against CXCR4 (Fig. 4A). However, CXCL12 had no effects on the invasion of K1 cells overexpressing CXCR7 (K1-CXCR7) (data not shown). In addition, 10% FBS did not induce the invasion of K1-CXCR4 and K1-CXCR7 cells. These results suggest that CXCR4 and CXCR7 may interact with CXCL12 differently in K1 cells. CXCL12 did not stimulate the proliferation of K1-CXCR4 and K1-CXCR7 cells (data not shown). The proliferation rate of K1 cells in media containing 10% FBS was not affected by overexpression of CXCR4 or CXCR7 (Fig. 4B).
Figure 3.
Overexpression of CXCR4 and CXCR7 in K1 cells. (A) Representative image of western blot analysis of CXCR4 expression in K1 parental, mock-transfected, CXCR4-overexpression cells. (B) Representative image of western blot analysis of CXCR7 K1 parental, mock-transfected, CXCR7-overexpressing cells. Cells were harvested 72 h after transduction with CXCR4 or CXCR7 recombinant lentivirus, and protein extract was separated on SDS-PAGE gel and transferred to PVDF membrane. CXCR4 and CXCR7 protein were detected by anti-CXCR4 and anti-CXCR7. Densitometry was analyzed by Quantity One system.
Figure 4.
Overexpression of CXCR4 and CXCR7 induces K1 cell invasion but has no effect on cell proliferation. (A) Overexpression of CXCR4 increased CXCL12-induced K1 cell invasion. Cells were treated with 100 ng/ml CXCL12 for 24 h. The concentration of control IgG and anti-CXCR4 was 10 μg/ml. The number of cells invaded through the membrane was counted 24 h after incubation with CXCL12. Three to 5 fields were randomly selected on each membrane and the average number of cells was used. The experiment was repeated at least 3 times. (B) Overexpression of CXCR4 or CXCR7 did not affect K1 cell proliferation. Cells were incubated in media containing 10% FBS, and cell growth was determined by using CCK-8 kit.
Overexpression of CXCR4 or CXCR7 upregulates AKT and ERK phosphorylation and MMP-2 expression and activity
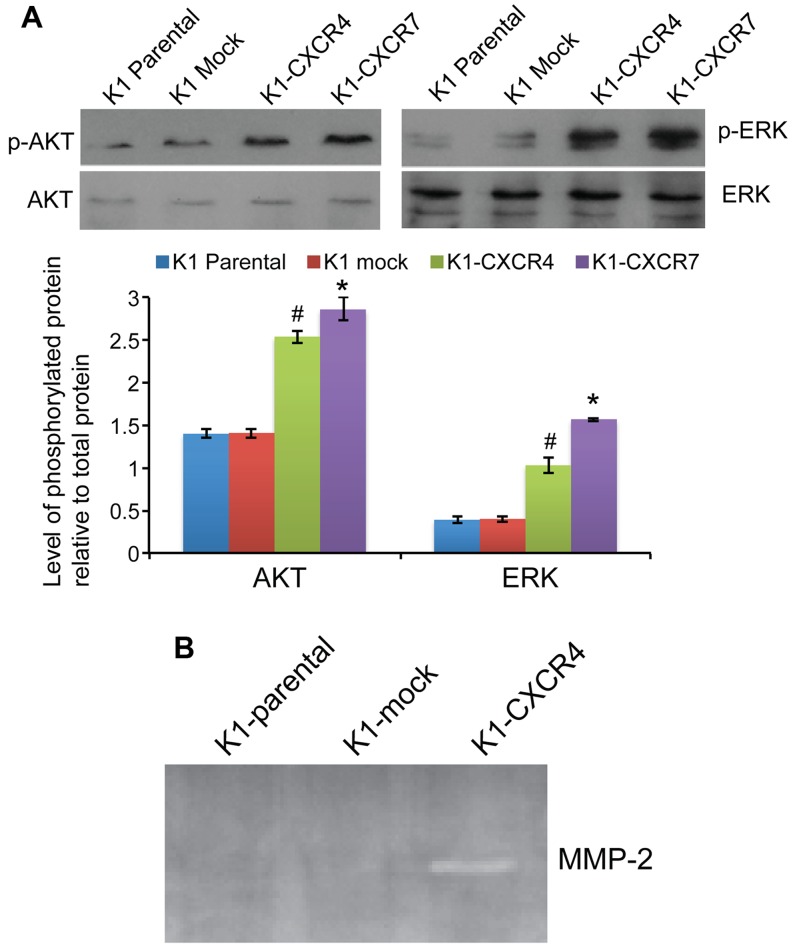
The phosphorylation levels of both AKT and ERK were significantly increased in K1 cells overexpressing CXCR4 or CXCR7 compared with mock transfected cells (all P<0.05; Fig. 5A). RT-PCR revealed that overexpression of CXCR4 markedly increased the mRNA expression of matrix metalloproteinase (MMP)-2 in K1 cells by 908.2±10.9-fold compared with K1-mock cells, whereas MMP-9 expression in K1 cells was not affected by CXCR4 or CXCR7 overexpression. Consistently, zymography assay showed that MMP-2 activity was markedly increased in K1-CXCR4 cells compared with parental or mock-transfected K1 cells (Fig. 5B).
Figure 5.
The effect of overexpression of CXCR4 or CXCR7 on AKT and ERK phosphorylation and MMP-2 activity. (A) Overexpression of CXCR4 or CXCR7 increased AKT and ERK phosphorylation. Cells were harvested 72 h after transduction with CXCR4 or CXCR7 recombinant lentivirus, and protein extract was separated on SDS-PAGE gel and transferred to PVDF membrane. Densitometry was analyzed by Quantity One system. *P<0.05, significant difference between K1-CXCR7 and K1 mock. #P<0.05, significant difference between K1-CXCR4 and K1 mock. (B) Zymography assay of MMP activity. Culture media were collected and loaded in SDS-PAGE gel containing 1 mg/ml gelatin. After electrophoresis, the gel was incubated with MMP reaction buffer. The gel was then stained and de-stained. The experiment was repeated 3 times. Representative gel image is presented.
Discussion
In the present study, we found that CXCL12 was exclusively expressed in PTC specimens but was not expressed in benign thyroid specimens and other types of thyroid malignancy, including FTC, MTC and ATC. These findings indicate that CXCL12 may be an effective biomarker for PTC. Consistently, Chung et al (20) recently demonstrated that CXCL12 was exclusively overexpressed in human PTC regardless of histological subtypes compared with non-cancerous thyroid lesions, and the sensitivity and specificity of using CXCL12 as a diagnostic marker for PTC was 90.8 and 96.8%, respectively (20). Jung et al (21) also reported that 90% of follicular PTC specimens were positive for CXCL12, whereas only 10.5% of follicular thyroid neoplasm specimens showed positive CXCL12. The results of the present study showed that neither non-cancerous thyroid lesions nor malignancy of FTC, MTC and ATC expressed CXCL12, further supporting the exclusive expression of CXCL12 in PTC and the diagnostic value of CXCL12 for PTC.
We found positive CXCL12 and CXCR7 staining in 75 and 97.5% of PTC specimens, respectively. Similarly, Liu et al (22) showed that positive immunohistochemical staining of CXCL12 and CXCR7 in 69.6 and 65.8% of PTC specimens, respectively, and the expression of the ligand and the receptor was positively correlated with lymph node metastasis of PTC. Wagner et al (23) semi-quantitatively measured the immunohistochemical staining intensity of CXCR7 in human PTC specimens and found that PTCs with lymph node metastasis showed high intensity of staining for CXCR7. In contrast, we did not find any correlation of CXCR7 expression or the intensity of CXCR7 expression to PTC lymph node metastasis. The discrepancy may be associated with the facts that antibodies against CXCR7 used in the studies were from different manufacturers and/or minor difference in immunohistochemical staining procedures could result in subtle variation of staining. Thus, the association between CXCR7 expression and PTC lymph node metastasis needs to be further investigated.
To the best of our knowledge, this study is the first demonstrating that CXCR7 was also widely expressed in thyroid malignancy of FTC, MTC and ATC in addition to PTC, and CXCR7 was only weakly expressed in benign thyroid lesions. Notably, the intensity of CXCR7 expression was particularly high in ATC. These results indicate that CXCR7 function may vary in different types of thyroid malignancy. We found that the CXCR7 staining signals were mainly at endothelium of malignant thyroid tissue specimens. Immunohistochemistry of human breast and lung cancer tissue also revealed extensive CXCR7 expression on tumor-associated blood vessels and cancer cells (19). Thus, CXCR7 may contribute to thyroid cancer development by regulating angiogenesis.
The present study found that CXCR4 was either absent (NG) or expressed weakly in benign lesions (FA and HT) but showed strong expression in PTC, MTC and ATC specimens. De Falco et al (25) showed that CXCR4 was overexpressed in ATC specimens compared with normal thyroid tissue by real-time PCR and immunohistochemistry. Wagner et al (23) found that high CXCR4 expression was significantly associated with large tumor size of PTC specimens. Interestingly, we found that some PTC specimens showed stronger CXCR4 staining in lymph node metastasis than in the primary thyroid cancer tissue. Similarly, Lu et al (24) also showed that higher CXCR4 expression in the lymph node metastatic lesions than in the primary tumor tissues of esophageal squamous cell cancer. These findings indicate that CXCR4 positive thyroid cancer cells may have strong migratory potential. Our results from the in vitro experiments appear to support this hypothesis.
The in vitro experiments in this study showed that CXCL12 induced K1-CXCR4 cell invasion, whereas did not affect the invasion of K1 cells overexpressing CXCR7. These results indicate that CXCR4 and CXCR7 may have distinct function in thyroid cancer cells. CXCL12/CXCR4 axis-mediated migration has been observed in human anaplastic thyroid cancer cells, and CXCL12-induced anaplastic thyroid cancer cell migration is associated with AKT and ERK activation (25,26). Similarly, the present study also showed that AKT and ERK phosphorylation was significantly increased in K1 cells overexpressing CXCR4. Thus, CXCL12/CXCR4 axis appears to promote thyroid cancer cell migration by activating AKT and ERK signaling pathways. However, the function of CXCR7 in cancer development remains unclear. CXCR7 alone may not directly trigger the downstream signaling, instead, it may function as a co-receptor for CXCR4 or as a decoy receptor to stimulate or enhance CXCR4-associated downstream signaling (12–15). The results of this study appear to support the co-receptor or decoy receptor hypothesis regarding CXCR7 function, because CXCL12 failed to stimulate the invasion of K1 cells overexpressing CXCR7, but significantly induced K1-CXCR4 cell invasion. Levoye et al (12) investigated the cooperation of CXCR4 and CXCR7 in T cells in the presence of CXCL12 found that CXCR7 alone did not trigger the CXCL12-induced downstream signaling events, whereas co-expression of CXCR4 and CXCR7 resulted in CXCR4/CXCR7 heterodimer formation, which regulated CXCL12-promoted chemotaxis.
In addition, this study found that overexpression of CXCR4 markedly induced MMP-2 expression and its activity in K1 cells. The association of CXCL12/CXCR4 axis and MMP-2 activation has been indicated in CXCL12-induced lung alveolar epithelial cell migration. Ghosh et al (27) found that blockade of CXCR4 by the specific antagonist AMD-3100 inhibited epithelial cell migration and decreased MMP-2 activity. Ying et al (28) also suggested that CXCL12/CXCR4 axis might promote pancreatic cancer invasion by activating p38 mitogen-activated protein kinase and upregulating MMP-2 activity. Thus, during thyroid cancer progression, CXCL12 may bind to CXCR4 to activate AKT and ERK signaling pathways, thus, upregulating MMP-2 and ultimately promoting cancer cell migration and invasion. CXCR7 may facilitate or enhance the binding of CXCL12 to CXCR4.
In conclusion, immunohistochemistry revealed that CXCL12 was exclusively expressed in human PTC tissue and CXCR7 was widely expressed in the endothelial cells of PTC, FTC, MTC, ATC and FA tissue specimens, but only occasionally found in HT and NG tissue specimens. CXCR4 was commonly expressed in thyroid cancer, but only weakly expressed in benign thyroid lesions. CXCL12/CXCR4/CXCR7 axis may contribute to thyroid cancer development by regulating cancer cell migration/invasion via AKT and ERK signaling and MMP-2 activation.
Acknowledgements
The present study was supported by grants from the National Natural Science Foundation of China (grant no. 81102044).
References
- 1.Kilfoy BA, Zheng T, Holford TR, Han X, Ward MH, Sjodin A, Zhang Y, Bai Y, Zhu C, Guo GL, et al. International patterns and trends in thyroid cancer incidence, 1973–2002. Cancer Causes Control. 2009;20:525–531. doi: 10.1007/s10552-008-9260-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Pellegriti G, Frasca F, Regalbuto C, Squatrito S, Vigneri R. Worldwide increasing incidence of thyroid cancer: Update on epidemiology and risk factors. J Cancer Epidemiol. 2013;2013:965212. doi: 10.1155/2013/965212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Yang L, Yuan Y, Sun T, Li H, Wang N. Population-based cancer incidence analysis in Beijing, 2008–2012. Chin J Cancer Res. 2015;27:13–21. doi: 10.3978/j.issn.1000-9604.2015.01.07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Sun X, Cheng G, Hao M, Zheng J, Zhou X, Zhang J, Taichman RS, Pienta KJ, Wang J. CXCL12/CXCR4/CXCR7 chemokine axis and cancer progression. Cancer Metastasis Rev. 2010;29:709–722. doi: 10.1007/s10555-010-9256-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bleul CC, Fuhlbrigge RC, Casasnovas JM, Aiuti A, Springer TA. A highly efficacious lymphocyte chemoattractant, stromal cell-derived factor 1 (SDF-1) J Exp Med. 1996;184:1101–1109. doi: 10.1084/jem.184.3.1101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Caruz A, Samsom M, Alonso JM, Alcami J, Baleux F, Virelizier JL, Parmentier M, Arenzana-Seisdedos F. Genomic organization and promoter characterization of human CXCR4 gene. FEBS Lett. 1998;426:271–278. doi: 10.1016/S0014-5793(98)00359-7. [DOI] [PubMed] [Google Scholar]
- 7.Gupta SK, Pillarisetti K. Cutting edge: CXCR4-Lo: molecular cloning and functional expression of a novel human CXCR4 splice variant. J Immunol. 1999;163:2368–2372. [PubMed] [Google Scholar]
- 8.Burns JM, Summers BC, Wang Y, Melikian A, Berahovich R, Miao Z, Penfold ME, Sunshine MJ, Littman DR, Kuo CJ, et al. A novel chemokine receptor for SDF-1 and I-TAC involved in cell survival, cell adhesion, and tumor development. J Exp Med. 2006;203:2201–2213. doi: 10.1084/jem.20052144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Balabanian K, Lagane B, Infantino S, Chow KY, Harriague J, Moepps B, Arenzana-Seisdedos F, Thelen M, Bachelerie F. The chemokine SDF-1/CXCL12 binds to and signals through the orphan receptor RDC1 in T lymphocytes. J Biol Chem. 2005;280:35760–35766. doi: 10.1074/jbc.M508234200. [DOI] [PubMed] [Google Scholar]
- 10.Lu DY, Tang CH, Yeh WL, Wong KL, Lin CP, Chen YH, Lai CH, Chen YF, Leung YM, Fu WM. SDF-1alpha up-regulates interleukin-6 through CXCR4, PI3K/Akt, ERK, and NF-kappaB-dependent pathway in microglia. Eur J Pharmacol. 2009;613:146–154. doi: 10.1016/j.ejphar.2009.03.001. [DOI] [PubMed] [Google Scholar]
- 11.Roland J, Murphy BJ, Ahr B, Robert-Hebmann V, Delauzun V, Nye KE, Devaux C, Biard-Piechaczyk M. Role of the intra-cellular domains of CXCR4 in SDF-1-mediated signaling. Blood. 2003;101:399–406. doi: 10.1182/blood-2002-03-0978. [DOI] [PubMed] [Google Scholar]
- 12.Levoye A, Balabanian K, Baleux F, Bachelerie F, Lagane B. CXCR7 heterodimerizes with CXCR4 and regulates CXCL12-mediated G protein signaling. Blood. 2009;113:6085–6093. doi: 10.1182/blood-2008-12-196618. [DOI] [PubMed] [Google Scholar]
- 13.Sierro F, Biben C, Martínez-Muñoz L, Mellado M, Ransohoff RM, Li M, Woehl B, Leung H, Groom J, Batten M, et al. Disrupted cardiac development but normal hematopoiesis in mice deficient in the second CXCL12/SDF-1 receptor, CXCR7. Proc Natl Acad Sci USA. 2007;104:14759–14764. doi: 10.1073/pnas.0702229104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Boldajipour B, Mahabaleshwar H, Kardash E, Reichman-Fried M, Blaser H, Minina S, Wilson D, Xu Q, Raz E. Control of chemokine-guided cell migration by ligand sequestration. Cell. 2008;132:463–473. doi: 10.1016/j.cell.2007.12.034. [DOI] [PubMed] [Google Scholar]
- 15.Dambly-Chaudière C, Cubedo N, Ghysen A. Control of cell migration in the development of the posterior lateral line: Antagonistic interactions between the chemokine receptors CXCR4 and CXCR7/RDC1. BMC Dev Biol. 2007;7:23. doi: 10.1186/1471-213X-7-23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Wang J, Shiozawa Y, Wang J, Wang Y, Jung Y, Pienta KJ, Mehra R, Loberg R, Taichman RS. The role of CXCR7/ RDC1 as a chemokine receptor for CXCL12/SDF-1 in prostate cancer. J Biol Chem. 2008;283:4283–4294. doi: 10.1074/jbc.M707465200. [DOI] [PubMed] [Google Scholar]
- 17.Sun YX, Schneider A, Jung Y, Wang J, Dai J, Wang J, Cook K, Osman NI, Koh-Paige AJ, Shim H, et al. Skeletal localization and neutralization of the SDF-1(CXCL12)/CXCR4 axis blocks prostate cancer metastasis and growth in osseous sites in vivo. J Bone Miner Res. 2005;20:318–329. doi: 10.1359/JBMR.041109. [DOI] [PubMed] [Google Scholar]
- 18.Müller A, Homey B, Soto H, Ge N, Catron D, Buchanan ME, McClanahan T, Murphy E, Yuan W, Wagner SN, et al. Involvement of chemokine receptors in breast cancer metastasis. Nature. 2001;410:50–56. doi: 10.1038/35065016. [DOI] [PubMed] [Google Scholar]
- 19.Miao Z, Luker KE, Summers BC, Berahovich R, Bhojani MS, Rehemtulla A, Kleer CG, Essner JJ, Nasevicius A, Luker GD, et al. CXCR7 (RDC1) promotes breast and lung tumor growth in vivo and is expressed on tumor-associated vasculature. Proc Natl Acad Sci USA. 2007;104:15735–15740. doi: 10.1073/pnas.0610444104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Chung SY, Park ES, Park SY, Song JY, Ryu HS. CXC motif ligand 12 as a novel diagnostic marker for papillary thyroid carcinoma. Head Neck. 2014;36:1005–1012. doi: 10.1002/hed.23404. [DOI] [PubMed] [Google Scholar]
- 21.Jung YY, Park IA, Kim MA, Min HS, Won JK, Ryu HS. Application of chemokine CXC motif ligand 12 as a novel diagnostic marker in preoperative fine-needle aspiration biopsy for papillary thyroid carcinoma. Acta Cytol. 2013;57:447–454. doi: 10.1159/000351305. [DOI] [PubMed] [Google Scholar]
- 22.Liu Z, Sun DX, Teng XY, Xu WX, Meng XP, Wang BS. Expression of stromal cell-derived factor 1 and CXCR7 in papillary thyroid carcinoma. Endocr Pathol. 2012;23:247–253. doi: 10.1007/s12022-012-9223-x. [DOI] [PubMed] [Google Scholar]
- 23.Wagner PL, Moo TA, Arora N, Liu YF, Zarnegar R, Scognamiglio T, Fahey TJ., III The chemokine receptors CXCR4 and CCR7 are associated with tumor size and pathologic indicators of tumor aggressiveness in papillary thyroid carcinoma. Ann Surg Oncol. 2008;15:2833–2841. doi: 10.1245/s10434-008-0064-2. [DOI] [PubMed] [Google Scholar]
- 24.Lu CL, Guo J, Gu J, Ge D, Hou YY, Lin ZW, Ding JY. CXCR4 heterogeneous expression in esophageal squamous cell cancer and stronger metastatic potential with CXCR4-positive cancer cells. Dis Esophagus. 2014;27:294–302. doi: 10.1111/dote.12100. [DOI] [PubMed] [Google Scholar]
- 25.De Falco V, Guarino V, Avilla E, Castellone MD, Salerno P, Salvatore G, Faviana P, Basolo F, Santoro M, Melillo RM. Biological role and potential therapeutic targeting of the chemokine receptor CXCR4 in undifferentiated thyroid cancer. Cancer Res. 2007;67:11821–11829. doi: 10.1158/0008-5472.CAN-07-0899. [DOI] [PubMed] [Google Scholar]
- 26.Hwang JH, Hwang JH, Chung HK, Kim DW, Hwang ES, Suh JM, Kim H, You KH, Kwon OY, Ro HK, et al. CXC chemokine receptor 4 expression and function in human anaplastic thyroid cancer cells. J Clin Endocrinol Metab. 2003;88:408–416. doi: 10.1210/jc.2002-021381. [DOI] [PubMed] [Google Scholar]
- 27.Ghosh MC, Makena PS, Gorantla V, Sinclair SE, Waters CM. CXCR4 regulates migration of lung alveolar epithelial cells through activation of Rac1 and matrix metalloproteinase-2. Am J Physiol Lung Cell Mol Physiol. 2012;302:L846–L856. doi: 10.1152/ajplung.00321.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ying X, Jing L, Ma S, Li Q, Luo X, Pan Z, Feng Y, Feng P. GSK3β mediates pancreatic cancer cell invasion in vitro via the CXCR4/MMP-2 pathway. Cancer Cell Int. 2015;15:70. doi: 10.1186/s12935-015-0216-y. [DOI] [PMC free article] [PubMed] [Google Scholar]