Abstract
Although Anaplastic Large Cell Lymphomas (ALCL) carrying Anaplastic Lymphoma Kinase (ALK) have a relatively good prognosis, aggressive forms exist. We have identified a novel translocation, causing the fusion of the TRAF1 and ALK genes, in one patient who presented with a leukemic ALK+ ALCL (ALCL-11). To uncover the mechanisms leading to high-grade ALCL, we developed a human Patient Derived Tumorgraft (hPDT) line. Molecular characterization of primary and PDT cells demonstrated the activation of ALK and of NFkB pathways. Genomic studies of ALCL-11 showed the TP53 loss and the in vivo subclonal expansion of lymphoma cells lacking PRDM1/Blimp-1 and with c-MYC gene amplification. The treatment with proteasome inhibitors of TRAF1-ALK cells led to down-regulation of p50/p52 and lymphoma growth inhibition. Moreover a NFkB gene set classifier stratified ALCL in distinct subsets with different clinical outcome. Moreover, a selective ALK inhibitor (CEP28122) resulted in a significant clinical response of hPDT mice, but the disease could not be eradicated.
These data indicate that the activation of NFkB signaling contributes to the neoplastic phenotype of TRAF1-ALK ALCL. ALCL hPDTs are invaluable to validate the role of druggable molecules, predict therapeutic responses and are helpful tools for the implementation of patient specific therapies.
Keywords: anaplastic large cell lymphoma (ALCL), Patient Derived Tumorgraft (PDT), Tyrosine kinase fusion proteins, personalized medicine, targeted therapy
INTRODUCTION
Anaplastic Large Cell Lymphomas (ALCL) are a distinct subset of mature T cell lymphoma and they can be subclassified based on the presence chromosomal translocations affecting the Anaplastic Lymphoma Kinase (ALK) gene {Falini, 2009 #737; Swerdlow SH, 2008 #1092}. In the 2008 revised WHO classification, ALK+ ALCL and cutaneous ALCL received distinct designations, while ALK- ALCL remained as a provisional entity in anticipation of additional data {Swerdlow SH, 2008 #1092}.
De novo ALK+ ALCL are clinically aggressive lymphomas, most frequently occurring in the first 3 decades of life, with a stage III–IV disease, systemic symptoms, and extranodal involvement (60%) and a typical male predominance {Savage, 2008 #1044},{Schmitz, 2010 #1194}. Although the long-term clinical outcome for the majority of ALK+ ALCL patients is relatively favorable {Savage, 2008 #1044;Schmitz, 2010 #1194}, some patients can have a highly aggressive disease. Clinical development can include rapid nodal dissemination, extranodal involvement or even frank leukemic presentation {Monaco, 2007 #939;Liang, 2013 #1195}. Moreover, 20–30% of ALK+ ALCL patients relapse and eventually require high dose chemotherapies and bone marrow transplantation {Savage, 2008 #1044;Schmitz, 2010 #1194;Ferreri, 2012 #1193}.
The majority of ALK+ ALCL cases carry the t(2;5)(p23;q35) translocation, which fuses the 3’ portion of the ALK into the nucleophosmin (NPM) gene. As a result of this translocation, affected cells display ectopic expression of the NPM-ALK fusion protein within the cytoplasm and nucleus. Approximately 20% of cases of ALK+ ALCL, however, exhibit different ALK fusions with a cytoplasmic localization {Barreca, 2011 #618}. The N-terminus regions of all ALK chimera encode unique dimerization domains. These are critical for the constitutive activation of the kinases and required for ALK-mediated transformation. It is believed that ALK partners do not contribute otherwise to ALK lymphomagenesis. One notable exception has been identified in the TFG-ALK fusion, in which the TFG region can interact with NEMO and TANK leading to the NFkB activation {Miranda, 2006 #1197}.
ALK+ ALCL have more often a stable karyotypes {Boi, 2013 #1207}, supporting the hypothesis that they are highly addicted to ALK signaling and may not require multiple and synergizing alterations. Conversely, aggressive ALK+ cases have a large spectrum of chromosomal defects, frequently involving chromosome 8q, suggesting that the deregulated expression of MYC might leads to unfavorable clinical outcome {Grewal, 2007 #790;Monaco, 2007 #939;Liang, 2013 #1195;Moritake, 2011 #941}.
Patient Derived Tumorgraft (PDT) in heavily immuno-compromised animals [i.e. NOD.Cg-PrkdcscidIl2rgtm1Wjl/SzJ (NSG) mice] are a valuable instrument to study human cancers {Shultz, 2012 #1058}. These mice display a high rate of engraftment {Quintana, 2008 #1010} and provide a host environment capable to sustain the survival of neoplastic as well as normal human elements {Shultz, 2012 #1058}. The expansion of primary tumor cells provides abundant pathological tissues and the opportunity to test ad hoc protocols in a reasonable time-frame {Tentler, 2012 #1110}. Lastly, PDT enable the discovery of genetic lesions, which can be targeted by specific compounds {Garber, 2009 #1180} in ad hoc preclinical therapeutic protocols {Vilas-Zornoza, 2012 #1259;Cheng, 2011 #666;Rubio-Viqueira, 2006 #1034}.
Here, we have studied the tumorigenic properties of a novel TRAF1-ALK chimera {Feldman, 2013 #1226;Tabbo’, 2013 #1328}, which can elicit the inappropriate activation of ALK and NFkB pathways. Multiple genomic defects (loss of TP53, and Blimp1 and MYC amplification) were associated to a leukemic and chemo-resistant TRAF1-ALK ALCL patient. Notably, the usage of a specific ALK inhibitor could prolong the survival of ALCL PDT bearing mice, but was unable to eradicate the disease. Overall, PDT models represent a novel tool to validate therapeutic protocols and to predict clinical responses, particularly in the setting of refractory patients.
MATERIALS AND METHODS
Patients Selection and immunohistochemistry
Fresh and/or viable cryopreserved samples from primary ALCL were obtained at the time of diagnosis, before treatment, or at relapse after chemotherapy from the Universities of Perugia, Turin and Leuven. Diagnoses were assigned according to the WHO classification by at least two experienced pathologists. Informed consents were obtained following the recommendations of local ethical committees. Representative formalin-fixed tumor sections and/or tissue microarrays (TMAs) were processed for immunohistochemical (IHC) analyses on a semi-automated stainer {Piva, 2010 #8623}. List of the antibodies and staining conditions are provided in Supplemental Table S1.
Fluorescence in situ hybridization analysis
Cytogenetic and fluorescence in situ hybridization (FISH) followed routine methods. Interphase FISH was performed on FFPE sections. FFPE sections were pretreated with SPOT-Light Tissue Pretreatment Kit (Life Techologies), following manufacturer’s protocol. Probes applied for FISH included LSI ALK, LSI MYC, LSI TP53/CEP17 (Abbott Molecular, Ottigne, Belgium or Rome, Italy) and home-brewed bacterial artificial chromosome (BAC) clones flanking TRAF1 or BRCA1 genes (Supplemental S2), selected from www.ensembl.org, or PRDM1 gene, kindly provided by Dr. Laura Pasqualucci (Columbia University, New York, NY, USA). Non-commercial probes were labeled with SpectrumOrange- and SpectrumGreen-d-UTP (Abbott Molecular) using random priming. FISH images were acquired with a fluorescence microscope equipped with an Axiophot 2 camera (Carl Zeiss Microscopy, Jena, Germany) and a MetaSystems ISIS imaging system (MetaSystems, Altlussheim, Germany). Approximately 100 interphase cells were evaluated in each analysis.
RNA-Seq library preparation and Bioinformatics Analysis
RNA-seq was performed as previously described {Palomero, 2014 #1294} Supplementary Information). Bionformatics analyses were accomplished using dedicated fusion detection tools (Bellerophontes {Abate, 2012 #596}, Defuse {McPherson, 2011 #10377}, ChimeraScan {Iyer, 2011 #828} and TxFuse {Singh, 2012 #1061}).
5′ Race and RT-qPCR
Race {Ma, 2000 #900} and RT-qPCR {Agnelli, 2012 #600} were performed as described. The PCR cycling conditions were: 95°C for 5 minutes, followed by 40 cycles at 94°C for 10 seconds and 60 or 62°C for 30 seconds. Primer sequences are available if requested.
Mice and mice treatment
NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ (NSG) mice were bred within the Molecular Biotechnology Center (MBC) Animal Resource, under strict specific and opportunistic pathogen free conditions. Animal protocols were reviewed and approved by the Animal Committee of the University of Turin. CEP28122 (Cephalon, 100 mg/kg body weight BID) were administered by oral gavage. Magnetic resonance imaging and data analysis and postprocessing are described in the Supplementary Information.
Tumor grafting and harvesting
Leukemic ALCL cells were obtained from a peripheral blood sample of a refractory ALK+ ALCL patient (Supplementary Table S1). For the initial tumorgraft implants, lymphoma cells (10–12.5 × 106, 200 μl of Dulbecco’s phosphate buffered saline) were injected intravenously (i.v., tail vein), or intraperitoneally (i.p.) or subcutaneously (s.c.) in a total of 5- to 8-week-old anaesthetized NSG mice (Rompun 0.05μl/g e Zoletil 1.6μg/g i.m.). Secondary transplants were performed by injecting 1 × 106 ALCL cells i.v. or alternatively implanting s.c. tumor lymphoma fragments (2 to 4, 3×3×1 mm fragments) {Shultz, 2007 #1059}. Implants’ growth was assessed either by palpation (s.c) and by magnetic resonance imaging (MRI). Presence and immunophenotype of human blood circulating cells were determined by multicolor flow cytometry {Brusa, 2013 #1258}. Recipient animals were checked regularly and sacrificed at early signs of distress. At harvesting, mice were sacrificed in a CO2 chamber and tissue grafts were collected for histologic evaluation, re-grafting, cryopreservation, or snap-freezing in liquid nitrogen.
The log-rank (Mentel-Cox) test was used to compare survival rates among different populations. The cumulative probability of overall survival was plotted as a curve according to the Kaplan-Meier method.
Antibodies Western Blotting and immune-precipitation assays
The following primary antibodies were used for Western blotting: anti-ALK (Life Technology, Monza, IT; mouse 1:2000), TRAF1 (Santa Cruz, rabbit 1:1000) and TRAF2 (Santa Cruz, Heidelberg DE; mouse 1:1000) anti-Actin Millipore (Merk-Millipore, Milano IT; mouse 1:2000), and anti MYC (rabbit 1:1000), antiphospho-STAT3 (rabbit 1:1000), anti STAT3-Y705 (rabbit 1:1000), anti SHP2 (rabbit 1:1000), anti SHP2-Y542 (rabbit 1:1000), anti ERK1/2-Thr202/Tyr204 (rabbit 1:1000), p50 NFkB (rabbit 1:1000), and p52NFkB (rabbit 1:1000) from Cell Signaling Technology (Danvers, MA). Western blotting and immunoiprecipaitation studies were performed as previously described {Zamo, 2002 #5422;Piva, 2006 #1201;Boccalatte, 2009 #639}.
Cell cultures and viability
Human ALCL cells (TS-SUP-M2, JB-6, Karpass 299, L82 and SU-DHL-1) were cultured under standard conditions (37°C in humidified atmosphere, with 5% CO2) in RPMI 1640 (Sigma-Aldrich, St Luis, MO, USA) and supplemented with 10% fetal calf serum (Lonza, Rockland, ME, USA), 2 mM glutamine, 100 U/ml penicillin and 100μg/ml streptomycin (Eurobio Biotechnology, Les Ulis, France). Human embryonal kidney cells HEK-293T (ATCC, Manassas, VA, USA) were cultivated in supplemented Dulbecco modified Eagle medium (DMEM). Cell viability was measured by ATP-lite using Perkin Elmer luminometer. Bortezomib, Tripolide, and Carfilzomib (Selleck, Houston, TX, USA), CEP28122 (Teva, West Chester, PA, USA) and Crizotinib (Chemitek, Indianapolis, IN, USA) were used for in vitro or in vivo studies.
ΔTRAF1-ALK (ΔT/A) and ΔNPM-ALK cassettes were constructed by PCR including the first 299 aa of TRAF1 and 100aa of NPM link to a short peptide corresponding to the intracytoplasmic region of the ALK gene (first 21 aa of the juxta-membrane region). Transfection of HEK-293T cells was performed with Effectene reagent (Qiagen, Valencia, CA, USA), according to the manufacturer’s instructions.
Luciferase analysis
Cells were transfected with multiple plasmids in association with pGL3 SOCS3 or IgK–HIV-kB-driven luciferase reporter and Renilla (1 to 40 ratio) constructs to test the TRAF1-ALK NFkB mediated transcriptional activity. After transfection (36h), cell lysates were prepared from 4×104 cells and dispensed in 96 well/plate, following the manufactorer’s instructions. Luciferase expression levels were determined following the recommended protocol (Dual-Luciferase Reporter Assay, Promega, Milano, Italy).
Gene expression profiling, processing and data analysis of microarray data
A gene expression profiling (GEP) dataset {Agnelli, 2012 #600} was analyzed as previously described {Fu, 2007 #769;Agnelli, 2012 #600}. The neoplastic profiles were derived from ALCL (ALK-, no.=24; ALK+, no.=30), PTCL-NOS (no.=74) and angioimmunoblastic T cell lymphoma (AITL, no.=41). For the extension/validation analysis we used the total of de novo 59 ALCL samples (46 clinically annotated) of the Lymphoma Leukemia Molecular Profiling Project and the International Peripheral T-cell Lymphoma Project {Iqbal, 2014 #1329}. Hierarchical clustering and dendrogram were generated with the GenePattern2.0 {Reich, 2006 #1214} suite (Pearson correlation distance measure and pairwise average-linkage clustering method). NFkB related genes were assessed by gene set enrichment analysis (GSEA) {Subramanian, 2005 #3408}. We compared the 69 normal T cells and the 169 neoplastic samples for enrichment of up-regulated NFkB target gene sets. The enriched gene sets were established combining the NFkB signatures from the Molecular Signatures Database (MsigDB) {Subramanian, 2005 #3408}, the Transcriptional Regulatory Element Database {Jiang, 2007 #1220} and NFkB target genes reported in large B-cell lymphoma {Feuerhake, 2005 #1213}, breast cancer {Lee, 2011 #1215} and T cells {Nagar, 2010 #1216}. We randomized the dataset by permuting the gene sets (10000 times) to assess the statistical significance. Overall survival significance was assessed by log-rank test. Significance associations between categorical data were calculated by Fisher exact test.
Genome-wide DNA profiling
Fresh and/or cryopreserved primary (ALCL11 [nodal and leukemic], NE433870L [diagnostic cervical LN]) and PDT lymphoma samples (ALCL11-PDT T1-1017) underwent DNA profiling using the Genome-Wide Human SNP Array 6.0 (Affymetrix, Santa Clara, CA, USA). Copy Number Variation (CNV) was determined {Rinaldi, 2013 #1221}.
RESULTS
Discovery of a TRAF1-ALK variant chimera
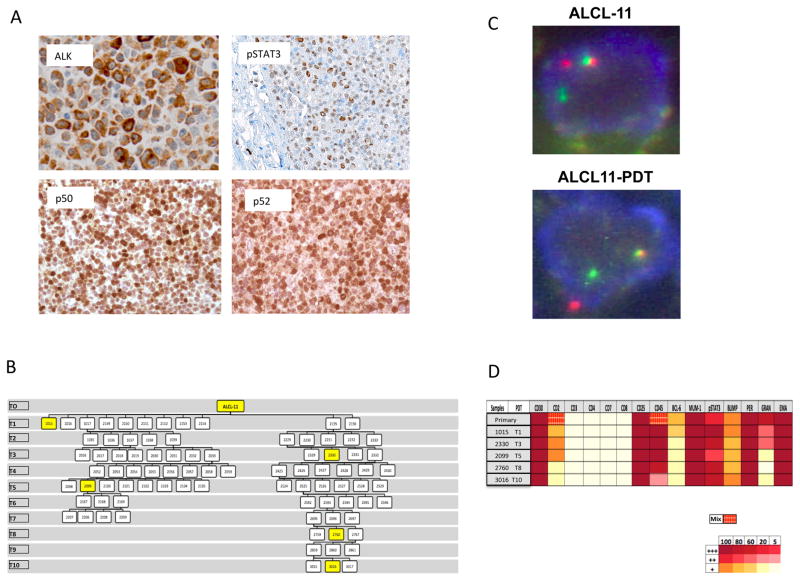
Aggressive forms and rare leukemic presentation of ALK+ ALCL have been reported {Savage, 2008 #1044;Schmitz, 2010 #1194;Ferreri, 2012 #1193}. To discover additional pathogenetic lesions associated to aggressive ALK+ ALCL cases, we selected a chemorefractory patient (ALCL-11) who rapidly evolved to a leukemic phase (Fig. S1A). An initial workout on the nodal ALCL lesion showed the presence of back to back lymphoma cells, displaying an intense cytoplasmic ALK positivity and a strong nuclear immunoreactivity for pSTAT3, p50 and p52 NFkB proteins (Figure 1a). These findings were consistent with a variant ALK+ translocation leading to the constitutive activation of JAK-STAT3 as well as NFkB pathways.
Fig 1. The ALCL-11 Patient Derived Tumorgraft line mimics its corresponding primary tissues sample.
(A) Immunohistochemistry stains performed on primary diagnostic sample (ALCL-11) show a strong ALK cytoplasmic staining with an antibody (D5F3, Cell Signaling Technology) specific for an intracytoplasmic peptide of the human ALK gene (x 400). Tissue sections stained with specific antibody against human phospoSTAT3 (pSTAT3, x100), p50 and p52 demonstrate a variable number lymphoma cells with nuclear positivity (x400). These analyses were performed on the diagnostic/primary tissue sample of the ALCL-11 patient.
(B) PDT engraftment and serial tumor propagation in NSG mice. Two distinct engraftments are depicted with their serial passages. Intersecting lines define the relationship among different tumors. Yellow boxes show mice the lymphoma tissue samples used for IHC stains.
(C) FISH analysis was performed on the primary tissue and in a representative PDT (ALCL-11 PDT1-2152) sample using a commercial kit and break apart set recognizing the 2p23 regions (Abbott Molecular/Vysis LSI ALK Break Apart FISH Probe Kit).
(D) Immunohistochemical stains were performed with specific antibodies recognizing T-cell associated and/or restricted antigens on the primary and serial PDT tissue samples (see Table S1). Percentage of positive cells and intensity of staining are indicated in the above diagram. Broken patterns indicate the presence of heterogeneous staining patterns.
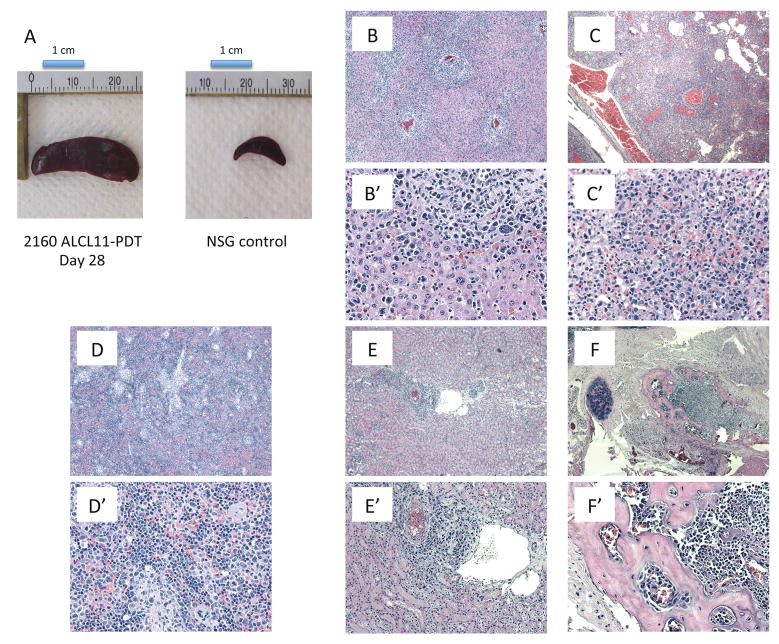
To establish a model representative of ALK variant ALCL, we first cultured the lymphoma cells with stromal mesenchymal elements and/or PHA-stimulated PBMC cell conditioned media. This strategy was however unsuccessful (data not shown). As alternative approach, we implanted fresh leukemic cells into NSG mice. These resulted in the engraftment and growth ALCL cells in all mice (100%, Figure 1b) (ALCL-11-PTD) and lymphoma graft cells (s.c. tissue fragments or i.v. of single cells from pathological spleens) could be successfully propagated in vivo (up to 13 consecutive passages). At autopsy, mice showed a disseminated infiltration of the lymphoid organs (i.e. spleen, bone marrow) and different parenchymal tissues with minor (kidneys) or extensive architectural effacements (liver) (Fig. S2). When we attempted to culture in vitro the ALCL-11 PDT cells (from passage T1 to T8, with or without stromal elements and with PHA-conditional media) no sustainable growth could be obtained (90% apoptotic cells after 96 h), suggesting that the growth of ALCL-11 PDT requires additional signals provided by the mouse-host cells.
Next, we performed a FISH to assess and correlate the ALK genomic configuration of diagnostic and PDT derived tissues. This analysis demonstrated the presence of the genomic rearrangement of the ALK locus, in both primary and PDT cells (Figure 1c). We also applied a panel of antibodies (Table S1) on the diagnostic biopsy and in representative tissue samples (s.c. tumor masses or spleen), derived from two independent primary engraftment lines (Figure 1b). This approach showed that all lymphoma cells had a similar immunophenotype (Figure 1d and Fig. S3). No difference were observed from different tissue samples (liver, spleen etc.) and/or from animals implanted at different passages and/or via alternative routes (subcutaneously or i.v.). Interestingly, variable expression of CD2 and CD45 antigens was observed in both primary ALCL and PDT tissues, suggesting a certain degree of intraclonal diversity (Figure 1d and S3).
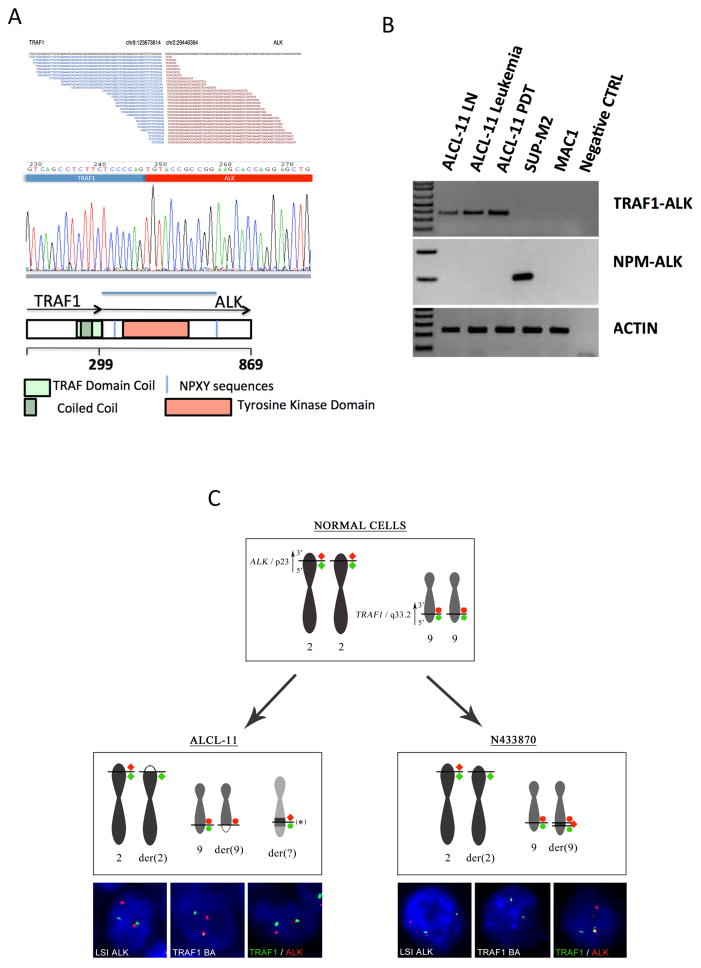
Since ALCL-11 cells did not display any common variant fusion transcript, we used a next-generation sequencing RNAseq approach and discovered 29 reads spanning across a junction breakpoint corresponding to the TRAF1 and ALK genes. These predicted a novel chimeric TRAF1-ALK transcript encoding the first five exons of the TRAF1 gene fused in frame to the exons 20–29 of ALK (Figure 2a) {Inghirami, 2012 #1263; Tabbo, 2013 #1093}. Validation by Sanger DNA sequencing and RT-PCR confirmed the presence of TRAF1-ALK transcripts of primary, relapsed and ALCL11-PDT samples (Figure 2b). These data were then validated by additional RNAseq analyses of samples derived from additional PDT (data not shown).
Figure 2. ALCL11 harbors the novel TRAF1-ALK fusion protein and leads to the constitutive activation of ALK signaling pathway.
(A) TRAF1–ALK gene fusion was defined by RNAseq (with 29 split reads overlapping junction breakpoint across TRAF1 and ALK). The genomic coordinates are shown. TRAF1 exon 5 and ALK exon 20 reading frames were conserved (central panel). The TRAF1-ALK chimera includes the TRAF1 and Coiled-coil domains, fused to the ALK intra-cytoplasmic region (lower panel).
(B) TRAF1-ALK fusion transcripts are detected in primary, leukemic and ALCL11-PDT samples by RT-PCR. ALK + (SUP-M2) and ALK- (MAC1) lines were used as controls. Negative controls correspond to samples lacking cDNA templates.
(C) TRAF1 BA assay (see supplementary T2S) was designed to validate molecular results by FISH. Case NE 433870L showed a normal hybridization pattern of TRAF1 BA in all analyzed interphase cells. To test if the TRAF1-ALK fusion might occur by an insertion of the 3′ALK within the TRAF1 locus in case 2, we applied probes covering the 5′end of TRAF1 (SO-labeled) and the 3′end of ALK (SG-labeled) (Upper right panel). Examination with MYC, TP53 and BLIMP1 of NE 433870L diagnostic tissue showed a normal hybridization pattern for all three probes (data not shown). ALCL-11 revealed split TRAF1 BA signals in approximately 50% of interphase cells (Lower panel);
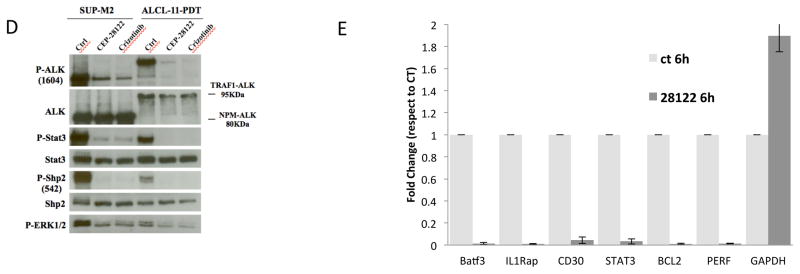
(D) TRAF1-ALK signaling is inhibited by anti-ALK inhibitors. PDT cells (PDT-3-2330) and SUP-M2 as control were treated in vitro (6hr, Crizotinib or CEP28122, 200nM). Total cell lysates were immune-blotted with the indicated antibodies.
(E) Panel shows the normalized levels of mRNA expression of STAT3 and representative STAT3-responsive genes in untreated or (anti-ALK) CEP28122 treated ALCL-11 leukemic cells (200nM, 6hr treatment). Expression levels were determined using a qRT-PCR approach as previously described {Piva, 2010 #8623;Agnelli, 2012 #599}. Data are depicted as 2^ddCt.5. Expression levels GAPDH of untreated (reference value) and CEP28122 treated cells are reported.
To determine the frequency of TRAF1-ALK among variant ALK+ ALCL, a total of 12 archival cases with cytoplasmic ALK staining were studied by RT-qPCR or 5′-RACE PCR on mRNA extracted from either FFPE or frozen tissue samples. One case, corresponding to an 11-year old Belgian patient (NE 433870L) displayed TRAF1-ALK mRNA transcripts identical to the one detected in ALCL-11 (Figure. S4), suggesting similar breakpoints {Feldman, 2013 #16682}. ALK fusions were documented in the remaining cases using a break part probe or RT-PCR (ATIC-ALK, TFG-ALK) {Cools, 2002 #695}. To further characterize the TRAF1-ALK rearrangement, we designed TRAF1 Break Apart (BA) probes. This FISH assay revealed split signals in the interphase lymphoma cells of the diagnostic ALCL-11 sample (~50% of the total cells). Conversely, the NE 433870L sample showed a normal hybridization pattern. Next we applied two alternative probes covering the 5′end of TRAF1 (SO-labeled) and the 3′end of ALK (SG-labeled) demonstrating the co-localization of red/green signals (Figure 2c). Globally, these data document the presence of genomic translocations involving the TRAF1 and the ALK loci, occurring with alternative modalities.
The TRAF1-ALK fusion leads to the constitutive activation of ALK and NFkB pathways
To define the oncogenic properties of TRAF1-ALK, we initially studied the phosphorylation status of the fusion protein and known ALK adaptors. As shown in Figure 2d, untreated ALCL-11 cells displayed a phosphorylated ALK band (~ 95kDa), which was substantially reduced in samples treated with ALK inhibitors (CEP28122 {Cheng, 2011 #666} and Crizotinib). As expected, the loss of ALK signaling led also to the down-regulation of pSTAT3 and pShp2 {Tabbo, 2012 #13783}, without substantial changes in cell viability and total protein expression (STAT3 and Shp2) within the first hour of treatment (6hr). Next, we evaluated the mRNA expression of genes whose expression is known to correlate with ALK-STAT3 signaling {Piva, 2010 #8623}, demonstrating that anti-ALK treatment led to their robust down-regulation (Fig. 2e). Similar data were observed with NPM-ALK positive cells lines (Figure S5a,b).
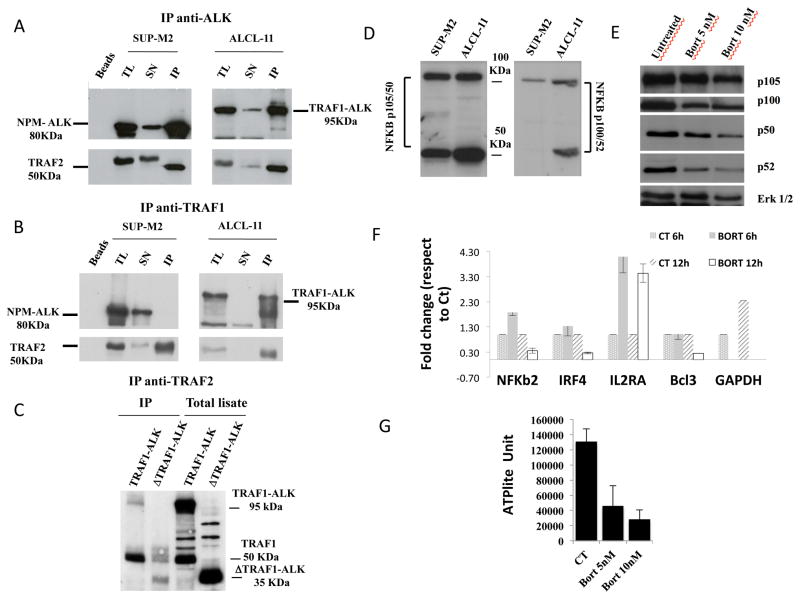
Since TRAF1 interacts with TRAF2, we first demonstrated that TRAF1-ALK could bind TRAF2 (Figure 3a,b). Because the intra-cytoplasmic region of ALK contains a TRAF2 interacting domain {Horie, 2004 #805}, we constructed a truncated form of TRAF1-ALK, lacking the TRAF2 ALK docking site (ΔTRAF1-ALK or ΔT/A). Using an antibody recognizing the N-terminus of TRAF1, we demonstrated that both WT and TRAF1_ALK could co-precipitate TRAF2 (Figure 3c). Knowing that TRAF1-ALK cells express NFkB, p50 and p52 (Figure 3d), we then tested whether ΔTRAF1-ALK could elicit a NFkB mediated transcription via a “bona fide” luciferase reporter cassette in cells co-expressing CD30 {Horie, 2004 #805}. HEK293T cells were transfected with different ALK cassettes in presence of CD30. As previously reported {Horie, 2004 #805}, NPM-ALK but not ELM4-ALK down-regulated the CD30 mediated NFkB activation (Figure 3e). Moreover, the forced expression of ΔTRAF1-ALK but not of ΔNPM-ALK enhanced the CD30-mediated NFkB luciferase expression (Figure 3f). To prove that the ΔTRAF1-ALK NFkB signaling requires p50 and/or p52, we co-transfected specific p50 and p52 shRNA cassettes, demonstrating decreased levels of transcription when they were coexpressed with ΔTRAF1-ALK (Figure 3f). Lastly, we treated ALCL-11 and/or ALCL-11 PDT cells with a proteasome inhibitor (Bortezomib) known to inhibit the NFkB signaling {Hideshima, 2009 #1199}, demonstrating a dose- and time-dependent down-regulation of both p50 and p52 protein levels (Figure 3g) and known NFkB regulated genes (starting from 12 hrs of incubation, Figure 3h). These later changes were associated to decreased cell viability (Figure 3J) and a concomitant reduction of intracellular ATP content measured by relative firefly luciferase activity (PerkinElmer ATPLite, Figure S6c). Since Bortezomib can inhibit multiple targets, we tested alternative proteasome inhibitors demonstrating similar findings in ALCL-11 or NPM-ALK cell lines (Figure S6).
Figure 3. TRAF1-ALK leads to the constitutive activation of NFkB transcription members.
(A–B) Lysates from a representative NPM-ALK cell line (SUP-M2 and Karpas 299) and leukemic ALCL-11 cells were immune-precipitated with an anti-ALK (Panel A) and anti-TRAF1 (B) antibodies. Immuno-complexes were resolved and then blotted with the indicated antibodies. Immuno-precipitates in presence of mouse serum are depicted as well (Beads). TL: Total Lysates, SN: Supernatant and IP Immune Precipitation. Protein molecular weights are shown. (C) Lysates from HEK293T cells, transfected with the full length TRAF1-ALK and ΔTRAF1-ALK, were immuno-precipitated with an anti-TRAF2 antibodies. Immuno-complexes were resolved and blotted with anti-TRAF1 antibodies. The expression of the corresponding proteins prior the immune-precipitation is shown (Total lysate) Predicted protein molecular weights are indicated.
(D) ALCL-11 cells show constitutive activation of the NFkB pathway. Primary ALCL-11 cells were immune-blotted with antibodies recognizing p105/p50 and p100/p52 NFkB. NPM-ALK+ ALCL cell lines were used as control.
(E) Luciferase Expression of transfected HEK-293T cells. Cells were transfected with the indicated cassettes expressing CD30, NFkB-ROS, CD30+NPM-ALK (active form), CD30+K210R NPM_ALK (inactive form) and CD30+ EML4-ALK.
(F) Knock-down of p50 or p52 impairs the NFkB mediated luciferase expression via CD30 signaling. HEK-293T cells were transfected with the indicated cassettes in presence of a CD30 expression vector.
(G) Protein expression of untreated and treated (Bortezomib 5nM/10nM) ALCL-11-PDTs was determined at 36 hr by Western Blotting with specific antibodies.
(H) Panel shows the normalized levels of mRNA expression of NFkB-regulated genes in untreated (control) or Bortezomib treated ALCL-11 leukemic cells (10nM, at 6 and 12 hr of culture). Expression levels were determined using a qRT-PCR approach as previously described {Piva, 2010 #8623;Agnelli, 2012 #599}. Data are depicted as 2^ddCt. Expression levels GAPDH of 6 hr treated (reference value) and 12 hr treated cells are reported.
(J) ALCL-11 cells are sensitive to Bortezomib. Primary cells (1×105/ml) were treated with increasing dose of the drug, overtime. Data have been normalized to control DMSO treated cells. DMSO viability decreased over time with a 30–40% spontaneous cell death at 36hrs.
Collectively these data demonstrate that TRAF1-ALK can elicit the NFkB pathways, which contributes the TRAF1-ALK phenotype.
NFkB is activated in a subset of primary ALK positive and ALK negative ALCL
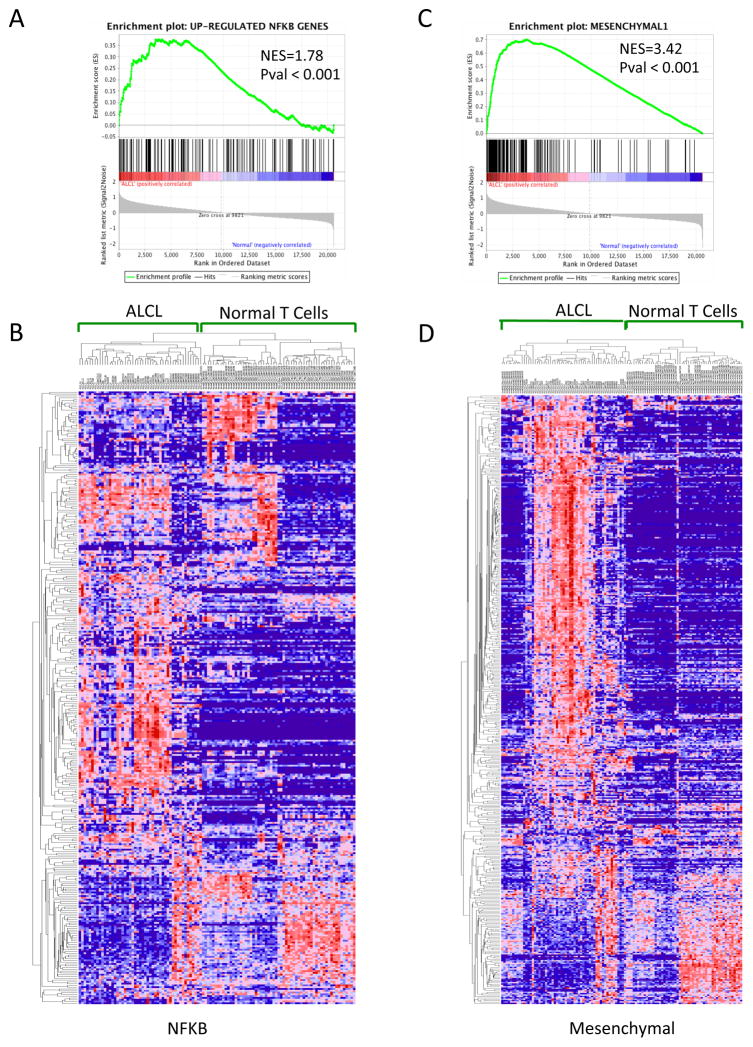
Since NFkB signaling plays a critical role in regulating stromal-tumor cells responses, we undertook a bioinformatics analysis to investigate whether ALCL primary samples displayed a unique NFkB signature. We first created a NFkB classifier combing established NFkB gene set lists (152 genes, Table S3, see Material and Methods) and tested it in a large cohort of ALCL samples and normal purified T cells {Agnelli, 2012 #600}. As expected NFkB transcripts from primary ALCL samples were significantly enriched (ES=0.38, NES=1.78, p-value < 0.001, FDR < 0.001) compared to purified normal, resting and activated T-cells (Figure. S7a,b). The robustness of this gene set classifier was similarly validated in a cohort (GSE12195) of diffuse large B-cell lymphoma (DLBCL) samples (Figure S7c) {Compagno, 2009 #1217}. We then demonstrated that the NFkB classifier could divided the ALCL in two different subsets (Figure 4a), which were similarly stratified uisng a mesenchymal signature (Figure 4b) {Lenz, 2008 #1225}. Notably, these substs displayed a strong correlation (Figure 4b,c). Findings were cross-validated with a second and independent ALCL cohort (Figure S7d–e) {Iqbal, 2014 #1329}. More importantly, we proved that the ALCL subgroups, which were defined by either the stromal or NFkB signatures, displayed unique clinical outcomes and that ALK+ ALCL preferentially enriched among patients with a better outcome.
Figure 4.
Lastly, when we applied specific antibodies against p50 and p52 in independent set of 98 ALCL, we found that lymphoma cells (anti-p52: 20/95, 21%; anti-p50: 19/98, 19%) and non neoplastic elements (endothelial and stromal cells) were NFkB positive (anti-p52: 87/98, 87%; p50: 13/89, 14%, Figure S8), suggesting that both ALCL and host cells can contribute to the NFkB signature.
The TRAF1-ALK PDT model bears high-risk genomic lesions
Although many ALK+ ALCL patients reach clinical remission, few refractory {Savage, 2008 #1044} and rare aggressive forms, carrying translocations or copy number gains of MYC {Monaco, 2007 #939}, have been described {Grewal, 2007 #790;Monaco, 2007 #939;Liang, 2013 #1195;Moritake, 2011 #941}.
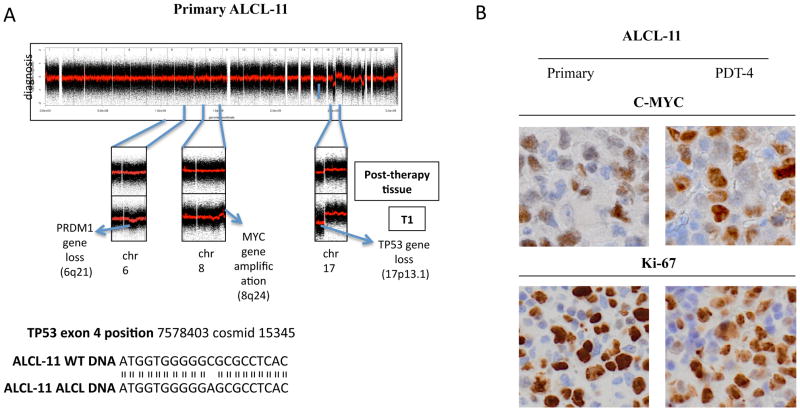
To identify genomic aberrations associated to TRAF1-ALK cases, we performed a Copy Number Variation (CNV) analysis. ALCL-11 leukemic and T1 and T3 PDT samples showed the heterozygous loss of 17p13 region and a concomitant missense mutation within TP53 gene exon 4 (Figure 5a). Comparison of the DNA profiles of primary, relapsed (post-therapy) and PDT tissues documented the loss of 20q (100% all samples) and a region spanning including the PRDM1/Blimp1 gene (6q, 30% PDTs). Interestingly, PDT samples displayed an amplification at 8q (42% of all cells), a region containing the c-MYC gene, a finding further confirmed by FISH (Table S4) and by a strong nuclear expression of MYC (>90%) (Figure 5b). On the contrary, the NE 433870L TRAF1-ALK ALCL sample showed minor CNV changes and a normal genomic configuration of MYC, PRDM1/Blimp1 and TP53 genes. This later patient had a remarcable response to chemotherapy and remain in clinical remission (Figure 1S), suggesting that the acqusition of multiple genetic defects may predict for clinical outcomes.
Figure 5. GSEA identifies distinct subsets of ALCL among PTCL and normal samples.
(A and B) Unsupervised hierarchical clustering (NFKB and Mesenchymal1 gene set) classifies ALCL in two distinct groups, respectively NKFBclus1/NKFBclus1 and MesClus1/MesClus2, (Pearson correlation distance measure, pairwise average-linkage clustering) with a significant association between the two groups (pvalue < 0.0001, Fisher exact test)
(C and D) Overall survival (OS) analysis of the two molecularly defined clusters for both NFKB and Mesenchymal gene set (M1=Mch1, M2=Mch2, NK1=NFKB1, NK2=NFKB2, see figure S7) in an independent cohort of 46 ALCL. Tables show a significant correlation between the NFKB/Mesenchymal groups and the ALK status in ALCL.
ALK inhibition can improve the outcome of ALCL-11 PDT mice
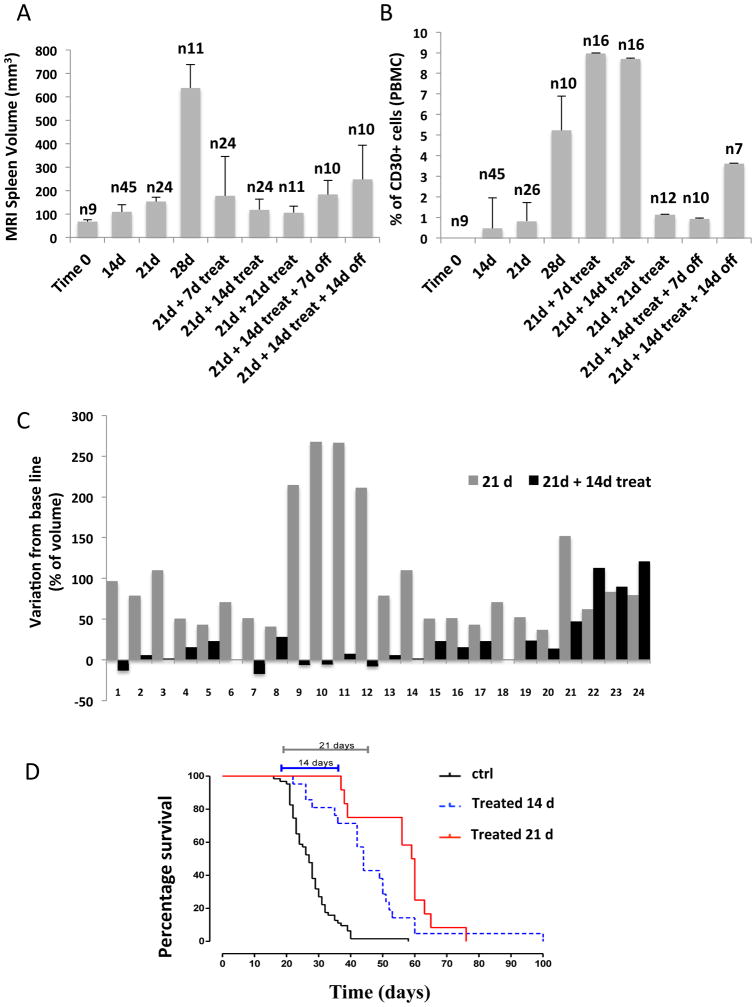
Since the management of chemo-refractory ALCL patients remains problematic, we tested the therapeutic efficacy of a selected ALK inhibitor in our ALCL-11 PDT model. Mice were i.v. tail injected (tumor cells 1×106). Engraftment and disease evolution were assessed by enumerating circulating CD30+ cells and by total body magnetic resonance imaging (MRI). As a therapeutic scheme, we selected a 14- or 21-day protocol (CEP28122 100mg/kg/bid), proven to eradicate disease in NPM-ALK PDT model obtained from a CHOP chemo-refractory ALCL patient {Cheng, 2011 #666}. As shown in Figure 6 (a and b), untreated mice had an increasing number of circulating CD30 cells and enlarging spleens (Figure S9) and 62% them succumbed at day 28 (90% after 37 from injection). While 88% of treated mice (21 from injection +7 treatment) were alive at day 28 and disease progression was controlled in mice treated for consecutive 14 days (Figure 6c). With a longer treatment (21 days) both spleen size and the percentage of circulating CD30 cells decreased, with an improvement of the over survival (p<0.0001). Nevertheless, when the inhibitor was withdrawn, circulating lymphoma cells and spleen sizes rose again (Figure 6a,b and Figure S10) and mice eventually succumbed with disease (90% at day 60). A similar scenario was obtained in a xenograft NPM-ALK cell line model adopting an identical therapeutic schedule (14 day treatment, Figure S11).
Figure 6. CGH analysis of primary and PDT ALCL-11 show the emergency of subclonal population lacking TP53 and MYC amplification.
(A) SNP-array plots of ALCL11-PDT identified a los at 17p13.1 (TP53) and 6q21 (PRDM1) and a gene amplification at 8q24 (MYC). X-axis: chromosome localization and physical mapping; Y-axis: signal indicating copy number status at each locus. DNA sequencing of normal and pathological DNA derived form Patient ALCL-11.
(B) IHC stains of primary and PDTs (ALCL-11-PDT-T4) tissue samples by an anti-MYC (Y69, Epitomics, Burlingame, CA) and Ki-67 (MIB-1, Dako Italia S.p.A, Italy) Mab. Original magnification x400.
These data demonstrate that ALK+ chemorefractory leukemic forms can be susceptible to anti-ALK kinase inhibitors, remaining addicted to ALK signaling.
DISCUSSION
We have provided evidence of the pathogenetic role of a novel TRAF1-ALK translocation and potential utility of anti-ALK inhibitors in aggressive leukemic forms of ALK+ ALCL. The TRAF1-ALK fusion chimera leads to the concomitant activation of ALK and NFkB pathways, which contribute to the maintenance of the neoplastic phenotype. Moreover, NFkB as well as mesenchymal gene set classifiers can stratify ALCL patients and predict clinical outcomes.
It is well established that ALK fusions are powerful oncogenes {Barreca, 2011 #618;Zhang, 2013 #1260} leading to the constitutive activation of JAK/STAT, PI3K/AKT, Ras/ERK1-2 and PLC-γ pathways {Barreca, 2011 #618;Tabbo, 2012 #13783}. Moreover, ALK chimera can interact with TRAF2 and modulate the CD30-mediate activation of NFkB {Horie, 2004 #805}. However, the contribution of NFkB signaling in ALCL tumorigenesis remains controversial. ALK+ ALCL cell lines have been reported to be devoid of p50 nuclear-binding activity {Martinez-Delgado, 2005 #1208;Bargou, 1996 #1209}. On the contrary, Bcl-3 and p52 are seen in primary ALCL {Mathas, 2005 #1210} and micro-dissected ALCL lymphoma cells preferentially express NFkB target genes {Eckerle, 2009 #10607}. Our data have confirmed these findings and demonstrated that a gene data set classifier of NFkB can reproducibly stratify ALCL, irrespectively of their ALK status. Interestingly, when we applied a mesenchymal gene set, known to stratify DLBCL and to predict clinical outcome, ALCL were also clustered in two major subgroups, which strongly correlated with those defined by the NFkB signature. These data suggest that NFkB signaling may dictate specific responses in lymphoma as well as in host cells. The fact that ALCL subgroups have unique clinical outcomes is relevant and may provide a novel tool to define poor responders with ALK+ ALCL.
Indeed, the pharmacological inhibition of the proteasome degradation known to modulate the NFkB pathway could represent a novel therapeutic modality. NFkB inhibitors could target either the lymphoma cells (TRAF1-ALK) or the host elements with a modality previously reported in other settings {Hiruma, 2009 #1261;Lwin, 2007 #1262}. Targeting the microeviroment has been recently proposed as analogous strategy in ALCL patients {Laimer, 2012 #872}.
Since many ALK- and a sizable subset of ALK+ALCL (20–30%) fail to reach clinical remission and prolong disease free survival {Savage, 2008 #1044;Grewal, 2007 #1181}, predicting biomarkers are highly desirable. The loss of TP53, PRDM1/Blimp1 {Boi, 2013 #1207} and TP63 {Vasmatzis, 2012 #1132} and deregulated expression of MYC {Monaco, 2007 #939} might pinpoint at diagnosis refractory/relapse or even aggressive forms of ALCL. In our PDT model, the emergency of subclones carrying defects of PRDM1/Blimp1 and MYC support the hypothesis that these lesions may have a relevant pathogenetic role and might occur as additional event along tumorigenesis contributing to more aggressive phenotype. A scenario previously described in acute lymphoblastic/leukemic PDT models {Clappier, 2011 #685}.
We have described the properties of a novel TRAF1-ALK fusion showing the unique contribution of its N-terminal TRAF1 region and demonstrated by ALK and NFkB can contribute to the neoplastic phenotype. The generation of PDT should represent a powerful approach to design more efficacious therapies in individual patients, particularly in the setting of relapses or refractoriness.
Supplementary Material
Figure 7. Single therapy with anti-ALK inhibitor leads to a partial response of ALCL1-11-PDT mice.
(A) ALCL-11 PDT2-6 (injected with 1×106 lymphoma cells, i.v) were analyzed using a dedicated MRI unit to determine the volume of the spleen as a surrogate of lymphoma/leukemia burden. Mice were treated with CEP28122, 100mg/kg/BID starting day 21th from the i.v. inoculum and scanned a different intervals (7, 14 or 21 days). A cohort of treated mice (14 days) were then released and followed overtime (7 or 14 days off). Numbers of mice are indicated.
(B) ALCL-11 PDT2-6 (injected with 1×106 lymphoma cells, i.v) were bled and blood circulating human CD45+CD30+ cells were determined by flow cytometry. Mice were treated with CEP28122, 100mg/kg/BID as indicate above. Numbers of mice are indicated.
(C) Effect of CEP28122 treatment in ALCL-11 PDT. Waterfall plot of mice injected with 1×106 ALCL-11 cells (i.v. day 21, grey bars) and CEP28122 response (day 21+14, back bars) after two weeks of treatment, compared with tumor volume at baseline (day 21), in 24 cases.
(D) Overall survival of ALCL-11-PDT treated with anti-ALK (CEP28122). Mice were randomized and treated as indicated with 100mg/kg/BID CEP28122 as indicated.
Acknowledgments
GI is supported by the Italian Association for Cancer Research (AIRC) Special Program in Clinical Molecular Oncology, Milan (5x1000 No. 10007); Regione Piemonte (ONCOPROT, CIPE 25/2005); ImmOnc (Innovative approaches to boost the immune responses, Programma Operativo Regionale, Piattaforme Innovative BIO F.E.S.R. 2007/13, Asse 1 ‘Ricerca e innovazione’ della LR 34/2004) and the Oncology Program of Compagnia di San Paolo, Torino. SP and BF by the Italian Association for Cancer Research (AIRC) Special Program in Clinical Molecular Oncology, Milan (5x1000 No. 10007); RR by Partnership for Cure, NIH 1 P50 MH094267-01, NIH 1 U54 CA121852-05, NIH 1R01CA164152-01. FB is sponsored by the Oncosuisse KLS-02403-02-2009 (Bern, Switzerland); Anna Lisa Stiftung (Ascona, Switzerland); Nelia and Amadeo Barletta Foundation (Lausanne, Switzerland); RP by Rete Oncologica del Piemonte e della Valle d’Aosta. LDS by National Institutes of Health (USA) grant CA034196. M.B. and M.T., I.L., F.T. F.dG., R.C and L.B. are enrolled in the PhD program (Pharmaceutical Sciences, University of Geneva, Switzerland and Molecular Medicine, University of Torino, respectively). We thank Drs. Vigliani C, Fioravanti A., and Mossino M. for their technical support and to Dr Casano J. for the constructive revision of the manuscript.
Footnotes
CONFLICT OF INTEREST
Authors must declare to not have any competing financial interests in relation to the work described.
Supplementary information is available at Leukemia’s website.
References
- 1.Falini B, Martelli MP. Anaplastic large cell lymphoma: changes in the World Health Organization classification and perspectives for targeted therapy. Haematologica. 2009;94(7):897–900. doi: 10.3324/haematol.2009.008250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Swerdlow SHCE, Harris NL, Jaffe ES, Pileri SA, Stein H, Thiele J, et al. WHO classification of tumors of haemotolopoietic and lymphoid tissues. 4. Stylus Publishing, LLC; Sterling, VA: 2008. [Google Scholar]
- 3.Savage KJ, Harris NL, Vose JM, Ullrich F, Jaffe ES, Connors JM, et al. ALK- anaplastic large-cell lymphoma is clinically and immunophenotypically different from both ALK+ ALCL and peripheral T-cell lymphoma, not otherwise specified: report from the International Peripheral T-Cell Lymphoma Project. Blood. 2008;111(12):5496–5504. doi: 10.1182/blood-2008-01-134270. [DOI] [PubMed] [Google Scholar]
- 4.Schmitz N, Trumper L, Ziepert M, Nickelsen M, Ho AD, Metzner B, et al. Treatment and prognosis of mature T-cell and NK-cell lymphoma: an analysis of patients with T-cell lymphoma treated in studies of the German High-Grade Non-Hodgkin Lymphoma Study Group. Blood. 2010;116(18):3418–3425. doi: 10.1182/blood-2010-02-270785. [DOI] [PubMed] [Google Scholar]
- 5.Monaco S, Tsao L, Murty VV, Nandula SV, Donovan V, Oesterheld J, et al. Pediatric ALK+ anaplastic large cell lymphoma with t(3;8)(q26.2;q24) translocation and c-myc rearrangement terminating in a leukemic phase. Am J Hematol. 2007;82(1):59–64. doi: 10.1002/ajh.20758. [DOI] [PubMed] [Google Scholar]
- 6.Liang X, Branchford B, Greffe B, McGavran L, Carstens B, Meltesen L, et al. Dual ALK and MYC rearrangements leading to an aggressive variant of anaplastic large cell lymphoma. J Pediatr Oncol. 2013;35(5):e209–213. doi: 10.1097/MPH.0b013e3182815046. [DOI] [PubMed] [Google Scholar]
- 7.Ferreri AJ, Govi S, Pileri SA, Savage KJ. Anaplastic large cell lymphoma, ALK-positive. Crit Rev Oncology Hematol. 2012;83(2):293–302. doi: 10.1016/j.critrevonc.2012.02.005. [DOI] [PubMed] [Google Scholar]
- 8.Barreca A, Lasorsa E, Riera L, Machiorlatti R, Piva R, Ponzoni M, et al. Anaplastic lymphoma kinase in human cancer. J Mol Endocrinol. 2011;47(1):R11–23. doi: 10.1530/JME-11-0004. [DOI] [PubMed] [Google Scholar]
- 9.Miranda C, Roccato E, Raho G, Pagliardini S, Pierotti MA, Greco A. The TFG protein, involved in oncogenic rearrangements, interacts with TANK and NEMO, two proteins involved in the NF-kappaB pathway. J Cell Physiol. 2006;208(1):154–160. doi: 10.1002/jcp.20644. [DOI] [PubMed] [Google Scholar]
- 10.Boi M, Rinaldi A, Kwee I, Bonetti P, Todaro M, Tabbo F, et al. PRDM1/BLIMP1 is commonly inactivated in anaplastic large T-cell lymphoma. Blood. 2013;122(15):2683–2693. doi: 10.1182/blood-2013-04-497933. [DOI] [PubMed] [Google Scholar]
- 11.Grewal JS, Smith LB, Winegarden JD, 3rd, Krauss JC, Tworek JA, Schnitzer B. Highly aggressive ALK-positive anaplastic large cell lymphoma with a leukemic phase and multi-organ involvement: a report of three cases and a review of the literature. Ann Hematol. 2007;86(7):499–508. doi: 10.1007/s00277-007-0289-3. [DOI] [PubMed] [Google Scholar]
- 12.Moritake H, Shimonodan H, Marutsuka K, Kamimura S, Kojima H, Nunoi H. C-MYC rearrangement may induce an aggressive phenotype in anaplastic lymphoma kinase positive anaplastic large cell lymphoma: Identification of a novel fusion gene ALO17/C-MYC. Am J Hematol. 2011;86(1):75–78. doi: 10.1002/ajh.21887. [DOI] [PubMed] [Google Scholar]
- 13.Shultz LD, Brehm MA, Garcia-Martinez JV, Greiner DL. Humanized mice for immune system investigation: progress, promise and challenges. Nat Rev Immunol. 2012;12(11):786–798. doi: 10.1038/nri3311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Quintana E, Shackleton M, Sabel MS, Fullen DR, Johnson TM, Morrison SJ. Efficient tumour formation by single human melanoma cells. Nature. 2008;456(7222):593–598. doi: 10.1038/nature07567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Tentler JJ, Tan AC, Weekes CD, Jimeno A, Leong S, Pitts TM, et al. Patient-derived tumour xenografts as models for oncology drug development. Nat Rev Clin Oncol. 2012;9(6):338–350. doi: 10.1038/nrclinonc.2012.61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Garber K. From human to mouse and back: ‘tumorgraft’ models surge in popularity. J Natl Cancer Inst. 2009;101(1):6–8. doi: 10.1093/jnci/djn481. [DOI] [PubMed] [Google Scholar]
- 17.Vilas-Zornoza A, Agirre X, Abizanda G, Moreno C, Segura V, De Martino Rodriguez A, et al. Preclinical activity of LBH589 alone or in combination with chemotherapy in a xenogeneic mouse model of human acute lymphoblastic leukemia. Leukemia. 2012;26(7):1517–1526. doi: 10.1038/leu.2012.31. [DOI] [PubMed] [Google Scholar]
- 18.Cheng M, Quail MR, Gingrich DE, Ott GR, Lu L, Wan W, et al. CEP-28122, a highly potent and selective orally active inhibitor of anaplastic lymphoma kinase with antitumor activity in experimental models of human cancers. Mol Cancer Ther. 2012;11(3):670–679. doi: 10.1158/1535-7163.MCT-11-0776. [DOI] [PubMed] [Google Scholar]
- 19.Rubio-Viqueira B, Jimeno A, Cusatis G, Zhang X, Iacobuzio-Donahue C, Karikari C, et al. An in vivo platform for translational drug development in pancreatic cancer. Clinical Cancer Res. 2006;12(15):4652–4661. doi: 10.1158/1078-0432.CCR-06-0113. [DOI] [PubMed] [Google Scholar]
- 20.Feldman AL, Vasmatzis G, Asmann YW, Davila J, Middha S, Eckloff BW, et al. Novel TRAF1-ALK fusion identified by deep RNA sequencing of anaplastic large cell lymphoma. Genes Chromosomes Cancer. 2013;52(11):1097–1102. doi: 10.1002/gcc.22104. [DOI] [PubMed] [Google Scholar]
- 21.Tabbo F, Barreca A, Machioriatti R, Messana K, Landra I, Abate F, et al. Humanized NOD/Scid/IL2g-/- tumor grafts recapitulate primary anaplastic large cell lymphoma. AACR Annual Meeting. 2013 abstract 3853. [Google Scholar]
- 22.Piva R, Agnelli L, Pellegrino E, Todoerti K, Grosso V, Tamagno I, et al. Gene expression profiling uncovers molecular classifiers for the recognition of anaplastic large-cell lymphoma within peripheral T-cell neoplasms. J Clin Oncol. 2010;28(9):1583–1590. doi: 10.1200/JCO.2008.20.9759. [DOI] [PubMed] [Google Scholar]
- 23.Palomero T, Couronne L, Khiabanian H, Kim MY, Ambesi-Impiombato A, Perez-Garcia A, et al. Recurrent mutations in epigenetic regulators, RHOA and FYN kinase in peripheral T cell lymphomas. Nat Genet. 2014;46(2):166–170. doi: 10.1038/ng.2873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Abate F, Acquaviva A, Paciello G, Foti C, Ficarra E, Ferrarini A, et al. Bellerophontes: an RNA-Seq data analysis framework for chimeric transcripts discovery based on accurate fusion model. Bioinformatics. 2012;28(16):2114–2121. doi: 10.1093/bioinformatics/bts334. [DOI] [PubMed] [Google Scholar]
- 25.McPherson A, Hormozdiari F, Zayed A, Giuliany R, Ha G, Sun MG, et al. deFuse: an algorithm for gene fusion discovery in tumor RNA-Seq data. PLoS Comput Biology. 2011;7(5):e1001138. doi: 10.1371/journal.pcbi.1001138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Iyer MK, Chinnaiyan AM, Maher CA. ChimeraScan: a tool for identifying chimeric transcription in sequencing data. Bioinformatics. 2011;27(20):2903–2904. doi: 10.1093/bioinformatics/btr467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Singh D, Chan JM, Zoppoli P, Niola F, Sullivan R, Castano A, et al. Transforming fusions of FGFR and TACC genes in human glioblastoma. Science. 2012;337(6099):1231–1235. doi: 10.1126/science.1220834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ma Z, Cools J, Marynen P, Cui X, Siebert R, Gesk S, et al. Inv(2)(p23q35) in anaplastic large-cell lymphoma induces constitutive anaplastic lymphoma kinase (ALK) tyrosine kinase activation by fusion to ATIC, an enzyme involved in purine nucleotide biosynthesis. Blood. 2000;95(6):2144–2149. [PubMed] [Google Scholar]
- 29.Agnelli L, Mereu E, Pellegrino E, Limongi T, Kwee I, Bergaggio E, et al. Identification of a 3-gene model as a powerful diagnostic tool for the recognition of ALK-negative anaplastic large-cell lymphoma. Blood. 2012;120(6):1274–1281. doi: 10.1182/blood-2012-01-405555. [DOI] [PubMed] [Google Scholar]
- 30.Shultz LD, Ishikawa F, Greiner DL. Humanized mice in translational biomedical research. Nat Rev Immunology. 2007;7(2):118–130. doi: 10.1038/nri2017. [DOI] [PubMed] [Google Scholar]
- 31.Brusa D, Serra S, Coscia M, Rossi D, D’Arena G, Laurenti L, et al. The PD-1/PD-L1 axis contributes to T-cell dysfunction in chronic lymphocytic leukemia. Haematologica. 2013;98(6):953–963. doi: 10.3324/haematol.2012.077537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zamo A, Chiarle R, Piva R, Howes J, Fan Y, Chilosi M, et al. Anaplastic lymphoma kinase (ALK) activates Stat3 and protects hematopoietic cells from cell death. Oncogene. 2002;21(7):1038–1047. doi: 10.1038/sj.onc.1205152. [DOI] [PubMed] [Google Scholar]
- 33.Piva R, Chiarle R, Manazza AD, Taulli R, Simmons W, Ambrogio C, et al. Ablation of oncogenic ALK is a viable therapeutic approach for anaplastic large-cell lymphomas. Blood. 2006;107(2):689–697. doi: 10.1182/blood-2005-05-2125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Boccalatte FE, Voena C, Riganti C, Bosia A, D’Amico L, Riera L, et al. The enzymatic activity of 5-aminoimidazole-4-carboxamide ribonucleotide formyltransferase/IMP cyclohydrolase is enhanced by NPM-ALK: new insights in ALK-mediated pathogenesis and the treatment of ALCL. Blood. 2009;113(12):2776–2790. doi: 10.1182/blood-2008-06-161018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Fu L, Medico E. FLAME, a novel fuzzy clustering method for the analysis of DNA microarray data. BMC Bioinformatics. 2007;8:3. doi: 10.1186/1471-2105-8-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Iqbal J, Wright G, Wang C, Rosenwald A, Gascoyne RD, Weisenburger DD, et al. Gene expression signatures delineate biological and prognostic subgroups in peripheral T-cell lymphoma. Blood. 2014;123(19):2915–2923. doi: 10.1182/blood-2013-11-536359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Reich M, Liefeld T, Gould J, Lerner J, Tamayo P, Mesirov JP. GenePattern 2.0. Nat Genet. 2006;38(5):500–501. doi: 10.1038/ng0506-500. [DOI] [PubMed] [Google Scholar]
- 38.Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Nat Acad Sci USA. 2005;102(43):15545–15550. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Jiang C, Xuan Z, Zhao F, Zhang MQ. TRED: a transcriptional regulatory element database, new entries and other development. Nucleic Acids Res. 2007;35(Database issue):D137–140. doi: 10.1093/nar/gkl1041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Feuerhake F, Kutok JL, Monti S, Chen W, LaCasce AS, Cattoretti G, et al. NFkappaB activity, function, and target-gene signatures in primary mediastinal large B-cell lymphoma and diffuse large B-cell lymphoma subtypes. Blood. 2005;106(4):1392–1399. doi: 10.1182/blood-2004-12-4901. [DOI] [PubMed] [Google Scholar]
- 41.Lee ST, Li Z, Wu Z, Aau M, Guan P, Karuturi RK, et al. Context-specific regulation of NF-kappaB target gene expression by EZH2 in breast cancers. Mol Cell. 2011;43(5):798–810. doi: 10.1016/j.molcel.2011.08.011. [DOI] [PubMed] [Google Scholar]
- 42.Nagar M, Jacob-Hirsch J, Vernitsky H, Berkun Y, Ben-Horin S, Amariglio N, et al. TNF activates a NF-kappaB-regulated cellular program in human CD45RA- regulatory T cells that modulates their suppressive function. J Immunol. 2010;184(7):3570–3581. doi: 10.4049/jimmunol.0902070. [DOI] [PubMed] [Google Scholar]
- 43.Rinaldi A, Kwee I, Young KH, Zucca E, Gaidano G, Forconi F, et al. Genome-wide high resolution DNA profiling of hairy cell leukaemia. Br J Haematol. 2013;162(4):566–569. doi: 10.1111/bjh.12393. [DOI] [PubMed] [Google Scholar]
- 44.Cools J, Wlodarska I, Somers R, Mentens N, Pedeutour F, Maes B, et al. Identification of novel fusion partners of ALK, the anaplastic lymphoma kinase, in anaplastic large-cell lymphoma and inflammatory myofibroblastic tumor. Genes Chromosomes Cancer. 2002 Aug;34(4):354–362. doi: 10.1002/gcc.10033. [DOI] [PubMed] [Google Scholar]
- 45.Tabbo F, Barreca A, Piva R, Inghirami G European TCLSG. ALK Signaling and Target Therapy in Anaplastic Large Cell Lymphoma. Front Oncol. 2012;2:41. doi: 10.3389/fonc.2012.00041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Horie R, Watanabe M, Ishida T, Koiwa T, Aizawa S, Itoh K, et al. The NPM-ALK oncoprotein abrogates CD30 signaling and constitutive NF-kappaB activation in anaplastic large cell lymphoma. Cancer cell. 2004;5(4):353–364. doi: 10.1016/s1535-6108(04)00084-4. [DOI] [PubMed] [Google Scholar]
- 47.Hideshima T, Ikeda H, Chauhan D, Okawa Y, Raje N, Podar K, et al. Bortezomib induces canonical nuclear factor-kappaB activation in multiple myeloma cells. Blood. 2009;114(5):1046–1052. doi: 10.1182/blood-2009-01-199604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Compagno M, Lim WK, Grunn A, Nandula SV, Brahmachary M, Shen Q, et al. Mutations of multiple genes cause deregulation of NF-kappaB in diffuse large B-cell lymphoma. Nature. 2009;459(7247):717–721. doi: 10.1038/nature07968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Lenz G, Wright G, Dave SS, Xiao W, Powell J, Zhao H, et al. Stromal gene signatures in large-B-cell lymphomas. N Eng J Med. 2008;359(22):2313–2323. doi: 10.1056/NEJMoa0802885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Zhang Q, Wei F, Wang HY, Liu X, Roy D, Xiong QB, et al. The potent oncogene NPM-ALK mediates malignant transformation of normal human CD4(+) T lymphocytes. Am J Pathol. 2013;183(6):1971–1980. doi: 10.1016/j.ajpath.2013.08.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Martinez-Delgado B, Cuadros M, Honrado E, Ruiz de la Parte A, Roncador G, Alves J, et al. Differential expression of NF-kappaB pathway genes among peripheral T-cell lymphomas. Leukemia. 2005;19(12):2254–2263. doi: 10.1038/sj.leu.2403960. [DOI] [PubMed] [Google Scholar]
- 52.Bargou RC, Leng C, Krappmann D, Emmerich F, Mapara MY, Bommert K, et al. High-level nuclear NF-kappa B and Oct-2 is a common feature of cultured Hodgkin/Reed-Sternberg cells. Blood. 1996;87(10):4340–4347. [PubMed] [Google Scholar]
- 53.Mathas S, Johrens K, Joos S, Lietz A, Hummel F, Janz M, et al. Elevated NF-kappaB p50 complex formation and Bcl-3 expression in classical Hodgkin, anaplastic large-cell, and other peripheral T-cell lymphomas. Blood. 2005;106(13):4287–4293. doi: 10.1182/blood-2004-09-3620. [DOI] [PubMed] [Google Scholar]
- 54.Eckerle S, Brune V, Doring C, Tiacci E, Bohle V, Sundstrom C, et al. Gene expression profiling of isolated tumour cells from anaplastic large cell lymphomas: insights into its cellular origin, pathogenesis and relation to Hodgkin lymphoma. Leukemia. 2009;23(11):2129–2138. doi: 10.1038/leu.2009.161. [DOI] [PubMed] [Google Scholar]
- 55.Hiruma Y, Honjo T, Jelinek DF, Windle JJ, Shin J, Roodman GD, et al. Increased signaling through p62 in the marrow microenvironment increases myeloma cell growth and osteoclast formation. Blood. 2009;113(20):4894–4902. doi: 10.1182/blood-2008-08-173948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Lwin T, Hazlehurst LA, Li Z, Dessureault S, Sotomayor E, Moscinski LC, et al. Bone marrow stromal cells prevent apoptosis of lymphoma cells by upregulation of anti-apoptotic proteins associated with activation of NF-kappaB (RelB/p52) in non-Hodgkin’s lymphoma cells. Leukemia. 2007;21(7):1521–1531. doi: 10.1038/sj.leu.2404723. [DOI] [PubMed] [Google Scholar]
- 57.Laimer D, Dolznig H, Kollmann K, Vesely PW, Schlederer M, Merkel O, et al. PDGFR blockade is a rational and effective therapy for NPM-ALK-driven lymphomas. Nat Med. 2012;18(11):1699–1704. doi: 10.1038/nm.2966. [DOI] [PubMed] [Google Scholar]
- 58.Vasmatzis G, Johnson SH, Knudson RA, Ketterling RP, Braggio E, Fonseca R, et al. Genome-wide analysis reveals recurrent structural abnormalities of TP63 and other p53-related genes in peripheral T-cell lymphomas. Blood. 2012;120(11):2280–2289. doi: 10.1182/blood-2012-03-419937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Clappier E, Gerby B, Sigaux F, Delord M, Touzri F, Hernandez L, et al. Clonal selection in xenografted human T cell acute lymphoblastic leukemia recapitulates gain of malignancy at relapse. J Exp Med. 2011;208(4):653–661. doi: 10.1084/jem.20110105. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.