Abstract
Arylfluorosulfates have appeared only rarely in the literature and have not been explored as probes for covalent conjugation to proteins, possibly because they were assumed to possess high reactivity, as with other sulfur(VI) halides. However, we find that arylfluorosulfates become reactive only under certain circumstances, e.g., when fluoride displacement by a nucleophile is facilitated. Herein, we explore the reactivity of structurally simple arylfluorosulfates towards the proteome of human cells. We demonstrate that the protein reactivity of arylfluorosulfates is lower than that of the corresponding aryl sulfonyl fluorides, which are better characterized with regard to proteome reactivity. We discovered that simple hydrophobic arylfluorosulfates selectively react with a few members of the intracellular lipid binding protein (iLBP) family. A central function of iLBPs is to deliver small-molecule ligands to nuclear hormone receptors. Arylfluorosulfate probe 1 reacts with a conserved tyrosine residue in the ligand-binding site of a subset of iLBPs. Arylfluorosulfate probes 3 and 4, featuring a biphenyl core, very selectively and efficiently modify cellular retinoic acid binding protein 2 (CRABP2), both in vitro and in living cells. The x-ray crystal structure of the CRABP2–4 conjugate, when considered together with binding site mutagenesis experiments, provides insight into how CRABP2 might activate arylfluorosulfates toward site-specific reaction. Treatment of breast cancer cells with probe 4 attenuates nuclear hormone receptor activity mediated by retinoic acid, an endogenous client lipid of CRABP2. Our findings demonstrate that arylfluorosulfates can selectively target single iLBPs, making them useful for understanding iLBP function.
INTRODUCTION
Among sulfur(VI) halides, arylfluorosulfates (Ar-O-SO2-F) and aryl sulfonyl fluorides (Ar-SO2-F) are especially interesting because they are relatively unreactive toward hydrolysis, reduction, nucleophilic substitution, thermolysis, etc.1–13 When reactions do occur, they appear to occur exclusively at the sulfur center.1–3,14–17 A sulfur(VI) fluoride exchange (SuFEx) reaction1 occurs when a fluoride anion is displaced from a sulfur center by a nucleophile, and these are understudied with regard to other protein bioconjugation reactions. In contrast to other well-studied electrophiles that modify proteins, such as fluorophosphonates,18,19 vinyl sulfones,20–22 and acrylamides,23,24 the sulfur(VI) fluoride functional groups, and especially arylfluorosulfates, are exceedingly weak electrophiles that require protein binding-associated activation to become reactive (see below).
The contrast between the high kinetic stability of arylfluorosulfates and their activatable reactivity when bound to a protein likely stems from stabilization of the departing fluoride ion by local environmental factors (e.g., by interacting with a hydrogen bond donor within the protein binding site and/or by electric field interfacial effects within the protein binding site to extract the fluoride anion and/or an aqueous-hydrophobic interface) during sulfur(VI) attack by a proximal protein-bound nucleophile.1,25–38 These properties suggest that sulfur exchange reactions have potential to become context-dependent click reactions.39–41
Unlike arylfluorosulfates, the sulfur fluoride exchange reactivity of aryl sulfonyl fluorides with proteins has been extensively explored in the 1960s by pioneering studies of Baker4–13 and is exemplified by their covalent inhibition of numerous proteins, including enzymes (dihydrofolate reductase,2,3 fatty acid amide hydrolase,42 and serine proteases)43,44 and non-enzymes (transthyretin).17 Sulfonyl fluorides react with the side chains of various amino acids, such as serine, lysine and tyrosine, depending on the protein context.1,2,14–16,45 We recently reported fluorogenic arylfluorosulfates A1 and A2, which bind to human transthyretin non-covalently, and enable fluorescence imaging of this protein.46 The low reactivity of these arylfluorosulfates towards the HeLa cell human proteome was hinted at by the small number of fluorescent A1 or A2 conjugate bands in the SDS PAGE whole HeLa cell proteome analysis (Figure S1). The results also suggested that, when arylfluorosulfates exhibit moderate to high affinity reversible binding to a given protein, they can selectively react, as long as the protein binding site provides the means to activate the arylfluorosulfates to undergo the SuFEx reaction (see below) with a proximal nucleophile. Herein, we compare the basal levels of the proteome reactivity of aryl sulfonyl fluorides with that of arylfluorosulfates, and then focus on studying the latter more extensively.
We find that arylfluorosulfates are substantially less reactive in acid-base catalyzed SuFEx reactions with the human cell proteome in comparison to the better-known aryl sulfonyl fluorides. The simple arylfluorosulfates employed in this manuscript chemoselectively react with a conserved tyrosine phenolic group within the binding site of intracellular lipid binding proteins (iLBPs). A central function of the non-enzyme iLBPs is to deliver hydrophobic small molecule organic ligands to nuclear hormone receptors—transcription factors that are activated upon ligand binding.47–51 Arylfluorosulfate probes 3 and 4, both featuring a biphenyl core, covalently modify cellular retinoic acid binding protein 2 (CRABP2) with high efficiency and selectivity relative to the other iLBPs, both in vitro and in living cells. The crystal structure of the CRABP2–4 conjugate at 1.75 Å resolution, along with CRABP2 binding site mutagenesis / SuFEx data, provide insight as to how binding might activate arylfluorosulfate probes 3 and 4 and hint at how CRABP2 might bind these small molecules reversibly and selectively. Importantly, probe 4 inhibits CRABP2 and RARα-mediated retinoic acid (RA) signaling in a breast cancer cell line.
EXPERIMENTAL SECTION
Lysate preparation and fluorescence electrophoresis gel analysis
HeLa cells prepared under different transfection and probe-labeling conditions were collected by scraping the culture dishes. Cells were washed three times with phosphate buffered saline (PBS; pH 7.4) to remove residual free arylfluorosulfate probe, and sonicated in PBS to generate whole-cell lysates.52 The total protein concentration of lysates was measured using the micro bicinchoninic acid (BCA) assay (Thermo Scientific). A copper(I)-catalyzed azide-alkyne cycloaddition or CuAAC click reaction was used to covalently attach a fluorophore to the alkyne moiety of the protein-conjugates derived from arylfluorosulfates (Reaction not conducted for native polyacrylamide gel electrophoresis (PAGE) experiments).41,53–57 SDS loading buffer [or Native-PAGE Sample Buffer (Thermo Scientific) for native PAGE experiments] was then added to each sample, and the samples were boiled for 5 min (No boiling step was performed for native PAGE experiments). Proteins were resolved by SDS-PAGE gel (4–12%, Thermo Scientific) or using a Native-PAGE gel (Thermo Scientific). For fluorescence visualization, wet slab gels were scanned using a ChemiDoc MP Imaging system (Bio-Rad).
Affinity capture of protein conjugates via CuAAC click reaction
Addition of biotin-azide (B10184, Thermo Scientific) or rhodamine-azide58 to the alkyne moiety on the arylfluorosulfate probe-labeled proteins in whole-cell lysates was carried out as previously published56,57,59 with minor modifications. For appending rhodamine-azide to the alkyne moiety of the conjugate, 50 µL of 1–2 mg/mL lysate was used for each reaction. 20 mM CuSO4 solution was pre-incubated with 20 mM 2-[4-({bis[(1-tert-butyl-1H-1,2,3-triazol-4-yl)methyl]amino}methyl)-1H-1,2,3- triazol-1-yl]acetic acid57 (BTTAA) solution (1:2) before this solution was added to the lysates. The reagents for CuAAC in lysates were added in the following order: 1 µL of 5 mM rhodamine-azide in DMSO, 7.5 µL of Cu-BTTAA (1:2) mixture, and 2.5 µL of 100 mM sodium ascorbate solution (freshly prepared). The reaction mixtures were then briefly stirred and incubated at 30 °C for 1 h. For appending biotin-azide to the alkyne moiety, 1000 µL of 2 mg/mL lysate was used for each reaction. The reagent amounts were proportionally increased, except that 5 mM biotin-azide stock solution was used instead of a 5 mM rhodamine-azide solution.
Western blot analysis
The wet slab gels were transferred to polyvinylidene difluoride (PVDF) membranes (Bio-Rad), which were subsequently blocked with Odyssey Blocking Buffer (LI-COR Corp.) for 1 h at 25 °C or overnight at 4°C. Western blot analysis was performed with a rat monoclonal α-fatty acid binding protein 5 (FABP5) antibody (MAB3077, R&D systems), a rabbit polyclonal α-CRABP2 antibody (PA5-27451, Thermo Scientific), a rabbit polyclonal α-FABP4 antibody (PA5-13452, Thermo Scientific) or a rabbit polyclonal α-FABP3 antibody (ARP-47482-P050, Avivasysbio), followed by secondary α-rat or α-rabbit antibodies (IRDye 800 or IRDye 680, LI-COR Corp.). The PVDF membranes were scanned using the LI-COR Odyssey Imager (LI-COR Corp.).
Stable isotope labeling by amino acids in cell culture (SILAC)-mass spectrometry (MS) analysis
Dulbecco's Modified Eagle Medium (DMEM) for SILAC and amino acids were purchased from ThermoFisher and heavy isotope-labeled media was supplemented with 13C6,15N4 L-arginine, and 15C6 L-lysine. Heavy isotope-labeled HeLa cells were treated with a given arylfluorosulfate probe (40 µM) and light isotope-labeled HeLa cells were treated with vehicle (DMSO, final concentration 0.2%) for 16 h. Heavy and light cells were collected, washed in PBS three times and lysed by sonication (see above for details). After determining the total protein concentration of lysates by microBCA assay (Thermo Scientific), 2 mg/mL heavy and light proteomes were mixed 1:1, and probe-labeled proteins were conjugated to a biotin-azide tag (B10184, Thermo Scientific) by a CuAAC click reaction. The treated proteomes were then pelleted by MeOH/CHCl3 (3:1) precipitation. Excess reagents and biotin-azide were removed by MeOH/CHCl3 (1:1) washing/sonication steps. Washed pellets were briefly dried, and resuspended in 6 M urea / 25 mM ammonium bicarbonate solution, 2.5% SDS solution. The solubilized proteomes were treated with dithiothreitol (7.5 mM) at 65 °C for 15 min, briefly cooled on ice, and treated with iodoacetamide (30 mM) at 25 °C for 30 min in the dark. Probe-labeled proteins were enriched using streptavidin beads (20347, Thermo Scientific) and washed once with 1% SDS in PBS and 3 times with PBS. Enriched proteins were digested on-bead with trypsin (V5111, Promega) and the resulting tryptic peptides were analyzed by liquid chromatography-mass spectrometry (LC-MS) on a Q-Exactive mass spectrometer equipped with an EASY-nLC 1000 system (Thermo Scientific). Multi-dimensional protein identification technology (MuDPIT) experiments were performed by 5 min sequential injections of 0, 20, 50, 80, 100 % buffer C (500 mM ammonium acetate in buffer A) and a final step of 90% buffer C / 10% buffer B (20 % water, 80 % acetonitrile, 0.1 % formic acid, v/v/v) and each step followed by a gradient from buffer A (95 % water, 5 % acetonitrile, 0.1 % formic acid) to buffer B. Electrospray was performed directly from the analytical column by applying a voltage of 2.5 kV with an inlet capillary temperature of 275°C. Data-dependent acquisition of MS/MS spectra were performed with the following settings: eluted peptides were scanned from 400 to 1800 m/z with a resolution of 30000, with the mass spectrometer in a data dependent acquisition mode. The top ten peaks for each full scan were fragmented by higher-energy collisional dissociation (HCD) using a normalized collision energy of 30%, a 100 ms activation time, and a resolution of 7500. Dynamic exclusion parameters were 1 repeat count, 30 ms repeat duration, 500 exclusion list size, 120 s exclusion duration, and exclusion width between 0.51 and 1.51. Peptide identification was performed using the Integrated Proteomics Pipeline Suite (IP2, Integrated Proteomics Applications, Inc.) and SILAC quantification was performed using CIMAGE as described previously.60 To confidently identify targets of our arylfluorosulfate probes in the SILAC mass spectrometry experiments, we manually applied a filter that a given protein had to be identified with at least two peptides in at least 2 out of 3 replicates.
Tandem mass spectrometry analysis
Arylfluorosulfate probe-labeled proteins were precipitated using MeOH/CHCl3 as described above. The protein pellets were resuspended in 8 M urea / 50 mM Tris pH 8.0, reduced with 10 mM DTT for 30 min, alkylated with 12 mM iodoacetamide for 30 min in the dark and finally diluted to 2 M urea. Proteins were digested with trypsin (1:50 ratio) for 18 h at 37 °C. LC-MS analysis was carried out using the instrument and settings described above and a single gradient from buffer A to 80% buffer B. Peptide identification was carried out using IP2, followed by manual assignments of the MS2 spectra of the probe-labeled peptides.
Plasmid construction
The DNA for human CRABP2, FABP5, FABP3 and FABP4 were synthesized by integrated DNA technologies (gBlocks) and subcloned into the pET22b vector using PIPE cloning methods61 for the expression of C-terminal His6-tagged recombinant proteins. Wild-type FABP4 became disulfide-crosslinked during purification and deletion of the Cys2 residue eliminated this problem. The DNA for CRABP2 was subcloned into a pET28b vector using PIPE cloning methods61 for the expression of N-terminal His6-tagged recombinant protein with a tobacco etch virus (TEV) protease cleavage site between the His6-tag and CRABP2. All mutations were introduced by QuikChange site-directed mutagenesis per the manufacturer’s instructions (Agilent Technologies). Plasmids of GFP-CRABP2 and FABP5-GFP were kindly provided by Professor Noa Noy from Case Western Reserve University.62,63
Protein expression and purification
The pET22b vectors encoding FABP5, FABP4, FABP3 and CRABP2 variants were transformed into BL21 Star (DE3) competent cells (Invitrogen). When the bacteria reached an OD600 of 0.6–0.8, protein expression was induced with 1 mM isopropyl β-D-1-thiogalactopyranoside (IPTG, BioPineer Inc.) for 4 h at 37 °C. His6-tagged proteins were affinity purified using Ni-NTA resin (Qiagen) in PBS containing 10 mM imidazole and were eluted with PBS containing 500 mM imidazole. Eluted proteins were further purified by size-exclusion chromatography (Superdex 200, GE Healthcare) in PBS. Purified proteins were analyzed for purity by SDS-PAGE and Coomassie blue staining. FABP5 lacking any tags was from Sino Biological Inc. (12581-HNAE).
Labeling kinetics
Labeling rates were measured by incubating recombinant CRABP2-His6 (2.5 µM) with varying concentrations (7, 10, 20, 40, 60 or 100 µM) of arylfluorosulfate probe 3 (25 °C) in 50 mM sodium phosphate buffer (pH 8.0), 100 mM NaCl. The reaction was quenched at different time points (t) by the addition of SDS loading buffer followed by boiling for 5 min. The boiled samples were resolved by SDS-PAGE and the wet slab gels were scanned to visualize the fluorescence signal from the probe 3-CRABP2 conjugate using a ChemiDoc MP Imaging system (Bio-Rad). The fluorescence intensity (Ft) of the probe 3-CRABP2 conjugate was quantified with ImageJ64 and plotted as a function of time to derive the observed rate constant (kobs) using the equation,
where F∞ is the fluorescence intensity of the fully labeled probe 3-CRABP2 conjugate. Steady-state kinetic parameters were derived by fitting kobs to the equation,
where kinact is the rate constant of the chemical step, and Ki is the probe 3 concentration that provides half of the maximal rate.65 The diarylsulfate ester product appears to be very stable to hydrolysis.1,25
Crystallization and structure determination of CRABP2–4 conjugate
The N-terminal His6-tagged CRABP2 with a TEV protease cleavage site was expressed and affinity purified using Ni-NTA resin following the above-mentioned protocol. The eluted His6-TEV-CRABP2 protein was dialyzed against PBS at 4°C overnight, and then cleaved by Ac-TEV protease (Thermo Scientific) following the manufacturer’s instructions. Ac-TEV protease and uncleaved His6-TEV-CRABP2 were removed by Ni-NTA resin and the cleaved, tag-less CRABP2 was obtained in the flow through. The purified tag-less CRABP2 (25 µM) was modified by probe 4 (100 µM) for 2 h and the probe 4-CRABP2 conjugate was purified by size-exclusion chromatography (Superdex 200, GE Healthcare) in PBS. The purified CRABP2–4 conjugate was concentrated to 13 mg/mL before being screened for crystallization on the high-throughput robotic Rigaku CrystalMation platform at the Joint Center for Structural Genomics (JCSG) facility at The Scripps Research Institute. Crystals of the CRABP2–4 conjugate were obtained via the sitting-drop vapor-diffusion method at 20 °C in a crystallization reagent consisting of 20% (w/v) PEG3350 and 200 mM KH2PO4. Crystals were cryoprotected by brief immersion in a mixture of mother liquor:glycerol (70:30) prior to flash-cooling in liquid nitrogen.
Diffraction data were collected at beamline 12-2 at the Stanford Synchrotron Radiation Lightsource (SSRL) using an X-ray beam of wavelength 0.9784 Å. Frames were indexed and integrated using the XDS package, the space group was assigned as P212121 using Pointless, and the data were scaled using Scala.66 Reflection intensities were converted to amplitudes using Truncate.67 Five percent of reflections were flagged for model cross-validation using Rfree. The structure of the CRABP2–4 conjugate was solved via molecular replacement in Phaser68 using an initial model of apo-CRABP2 derived from PDB entry 2FS6.69 Two copies of CRABP2 were found in the asymmetric unit. The model was refined to 1.75 Å resolution in Refmac570–72 using TLS parameters and hydrogen atoms included at riding positions. Ligand restraints were generated using the GRADE server.73
Quantitative RT-PCR (qPCR)
MCF-7 cells were cultured in DMEM supplemented with charcoal-treated fetal bovine serum (FBS), 1% glutamine, 1% sodium pyruvate, 1% non-essential amino acids, 1% HEPES, 10 µg/ml Insulin and 1% penicillin/streptomycin. Cells were treated with retinoic acid (100 nM) and compound 4 as described at 37 °C, washed with Dulbecco’s phosphate-buffered saline, and RNA was extracted using the Quick-RNA MiniPrep Plus Kit (Zymo Research). qPCR reactions were performed on cDNA prepared from 500 ng of total cellular RNA using the QuantiTect Reverse Transcription Kit (Qiagen). The 2× Fast SYBR Green qPCR Master Mix (Biotool), cDNA, and appropriate human primers (CRBP1; ctccagtcactccccgaaat and cgtcctgcacgatctctttg; FABP5: tggccaagccagattgtatc and ctgccatcagctgtggtttc, CRABP2: caaggttggggaggagtttg and gttccccatcgttggtcagt) purchased from Integrated DNA Technologies were used for amplifications (5 min at 50 °C, 6 min at 95 °C, followed by 45 cycles of 15 s at 95 °C, 1 min at 60 °C) in an ABI 7900HT Fast Real Time PCR machine. Transcripts were normalized to the housekeeping gene RPLP2 and all measurements were performed in triplicate. Data were analyzed using the RQ Manager and DataAssist 2.0 software packages (ABI).
RESULTS
Arylfluorosulfates were synthesized directly from the phenols and SO2F2 following the procedure of Dong et al.1 Briefly, phenols dissolved in basic (Et3N) dichloromethane were treated with SO2F2 gas via an inflated balloon. Pure products were generally obtained in quantitative yield without column chromatography. For the probes that contain a polyethylene glycol (PEG) moiety, a high performance liquid chromatography (HPLC) purification step was used to remove residual impurities. To facilitate the detection of the protein-arylfluorosulfate conjugates of interest, we synthesized arylfluorosulfates containing a terminal alkyne moiety. This alkyne allows us to employ the CuAAC bioconjugation method (click reaction)40,54 to attach a tag to the protein-arylfluorosulfate conjugates. A non-environmentally sensitive fluorescent visualization tag was attached for SDS-PAGE analysis or a biotin-azide tag was conjugated for affinity purification-mass spectrometry evaluation.58
We first investigated whether simple arylfluorosulfate probes functionalized with an alkyne could covalently modify proteins in living human cells. Probe 1 has a solubilizing PEG substructure, whereas probe 2 lacks this feature. Probes 1 and 2 emerged from efforts to optimize arylfluorosulfate preparation, and were not “designed” to probe proteome reactivity. HeLa cells were treated with vehicle or arylfluorosulfate probe 1 or 2 (40 µM, Figure 1a) for 16 h at 37 °C. Cell lysates were then subjected to the CuAAC click reaction to attach a fluorescent rhodamine-azide to the arylfluorosulfate probe-protein conjugate(s).55,58 After separation by SDS-PAGE, the protein(s) that reacted with the arylfluorosulfate probes could be distinguished by their in-gel fluorescence (Figure 1b, left panel).
Figure 1.
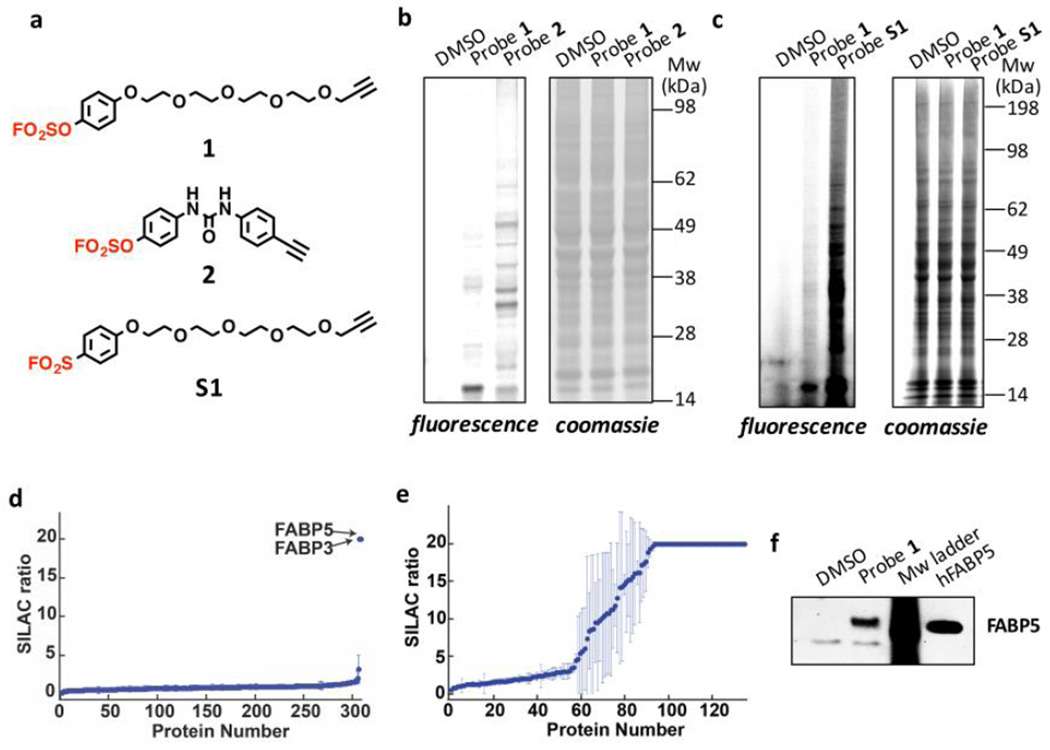
Evaluating the proteome reactivity of arylfluorosulfate probes. (a) Alkyne-functionalized arylfluorosulfate probes 1 and 2 and aryl sulfonyl fluoride probe S1. The arylfluorosulfate and aryl sulfonyl fluoride group are shown in red. (b-c) In-gel fluorescence evaluation of the reactivity of probes 1, 2 and S1 in HeLa cells after cell lysis and incorporation of rhodamine-azide using CuAAC. Left panel: in-gel fluorescence, right panel: Coomassie blue staining. (d-e) SILAC ratio plots for proteins identified in experiments comparing cells treated with 1 (d) or 2 (e) versus DMSO (no probe). Proteins with median SILAC ratio ≥ 5 (probe/DMSO) are designated as probe-labeled targets. Ratio ≥ 20 are listed as 20; results display the average of triplicate SILAC experiments performed in HeLa cells. (f) Western blot analysis of probe 1 modified proteins after incorporation of biotin-azide using CuAAC Click and affinity purification using streptavidin agarose beads. Recombinant tag-less human FABP5 serves as the positive control.
Structurally distinct arylfluorosulfate probes 1 and 2 exhibited discrete conjugation selectivities in the HeLa cell proteome (Figure 1b). The simple arylfluorosulfate probe 1, which is expected to bind reversibly to HeLa cell proteins with a generally lower equilibrium binding constant (because it is less hydrophobic) relative to 2, was more selective towards a protein(s) in the 15 kDa band. Arylfluorosulfate probe 2 comprising two aromatic rings separated by a urea linker modified a larger set of proteins. The time- and concentration-dependence of the labeling of the 15 kDa band by probe 1 showed that the covalent capture rate is slow (Figure S2). We next compared the proteome reactivity of 1 to the otherwise identical aryl sulfonyl fluoride probe S1 (Figure 1a; both at 40 µM) after a 16 h incubation period. This comparison revealed that 1 labeled very few proteins, whereas S1 formed conjugates with numerous proteome members, reflecting the overall muted reactivity of arylfluorosulfates in comparison to aryl sulfonyl fluorides (Figure 1c).2,3,14–16 Subjecting HEK293T cells to 1 for up to 48 h did not result in the substantial modification of the proteome relative to treatment by S1, which is significantly more reactive over all time intervals examined (Figure S3).
To identify the proteins modified by probes 1 and 2, we employed stable isotope labeling by amino acids in cell culture (SILAC) coupled with affinity enrichment, tryptic digestion, and liquid chromatography-tandem mass spectrometry (see Figure S4 for experimental scheme).74 SILAC allows for quantification of protein conjugates enriched by affinity chromatography in comparison to a background of non-conjugated proteins. Light isotope-labeled HeLa cells were treated with DMSO, whereas heavy isotope-labeled HeLa cells were treated with probes 1 or 2 (40 µM) for 16 h at 37 °C. Quantification of the affinity purification-mass spectrometry data revealed that probe 1 conjugated with fatty acid binding protein (FABP) 3 and FABP5 from the iLBP family, both of which exhibited heavy/light ratios exceeding 5 (Figures 1d and S5 and Table S1). For probe 2, many more HeLa cell proteins exhibited a heavy/light conjugate ratio above 5 (Figures 1e and S6 and Table S2), consistent with the SDS-PAGE-gel visualized by fluorescence imaging (Figure 1b). FABP5 was again observed, suggesting that it can also form a conjugate with probe 2. CRABP2, another iLBP, exhibited a heavy/light ratio of 20, but peptides from CRABP2 were identified in only one of three replicate experiments (Figure S6), and therefore it did not pass our initial filter cutoff. Collectively, these data are consistent with probes 1 and 2 labeling several of the 15 kDa iLBPs.47,50,62
Since probe 1 exhibited greater selectivity towards the 15 kDa band, we chose probe 1 for further characterization of the targeted protein(s) by western blot analysis. To provide additional support that probe 1 targets FABP5 and FABP3, conjugate formation between probe 1 and these iLBPs was studied using antibodies that are selective for FABP5 or FABP3. We treated HeLa cells with probe 1 or vehicle and then affinity purified the probe-labeled protein(s) conjugates from cell lysates using biotin tags (see Figure S7 for experimental scheme). The samples were then subjected to western blot analysis using either an anti-FABP5 or anti-FABP3 antibody. Probe 1 treated samples showed a band for FABP5 (Figure 1f), but no signal when the blot was probed using the FABP3 antibody (Figure S8), suggesting that the 15 kDa band contains at least FABP5. It is possible that the FABP3 antibody was insufficiently sensitive to detect FABP3 in comparison to SILAC mass spectrometry experiments. In this regard, quantification of protein abundances in HeLa cells suggests that the expression level of FABP5 is higher than that of CRABP2 and FABP3.75
To further scrutinize the SILAC-MS and Western blot data, we incubated recombinant iLBPs CRABP2, FABP5, or FABP3 individually with probe 1. The conjugate resulting from probe 1 (100 µM) reacting with CRABP2 (2 µM) formed quantitatively within 48 h (Figure S9), as discerned by liquid chromatography-electrospray ionization mass spectrometry (LC-ESI-MS; see Figure S10 for the appropriate quantification control experiments). However, the conjugate derived from FABP5 formed in only about 80% yield after 48 h under identical conditions (Figure S11). Interestingly, recombinant FABP3 showed no observable conjugate formation (Figure S12), suggesting that the labeling of FABP3 by probe 1 is very inefficient, at least with the recombinant protein in buffer.
Despite differences in function and ligand specificity, there is significant structural homology amongst iLBP family members.49,76 A phylogenetic analysis of human FABPs and CRABPs demonstrates that CRABPs and FABPs are closely related proteins.49 The similarities between CRABP2 and FABP5 are also evident in the way they bind ligands containing a carboxylic acid. Three amino acids—Arg111/109, Arg132/129, and Tyr134/131 (numbering for CRABP2 and FABP5, respectively)—are known to hydrogen bond with the endogenous ligands, e.g., RA and long chain fatty acids. An alignment of the available crystal structures of apo CRABP2, FABP5 and FABP4 indicates that the orientation of the Arg~Arg~Tyr carboxylic acid binding modules are almost identical (Figure 2a).69,77,78 Given the structural similarity of FABP4, CRABP2 and FABP5, we hypothesized that arylfluorosulfate probes 1 and 2 might also be able to covalently target FABP4 in living systems. Since HeLa cells do not express FABP4, we tested this hypothesis using 3T3-L1 mouse adipocytes, which express FABP4, but not FABP3, at high levels.79 Living cells were treated with probe 1 (40 µM for 16 h at 37 °C) and cell lysates were reacted with rhodamine-azide using a CuAAC click reaction. SDS-PAGE in-gel fluorescence analysis revealed that probe 1 covalently labeled a 15 kDa band (Figure S13a). A biotin click reaction with the cell lysate, followed by affinity purification and western blot analysis confirmed that this band contained FABP4 (Figure S13b), showing that 1 had conjugated to mouse FABP4 in 3T3-L1 cells. Recombinant human FABP4 also reacted with probe 1, affording a 90% modification yield after 48 h at 25 °C, as discerned by LC-ESI-MS (Figure S14).
Figure 2.
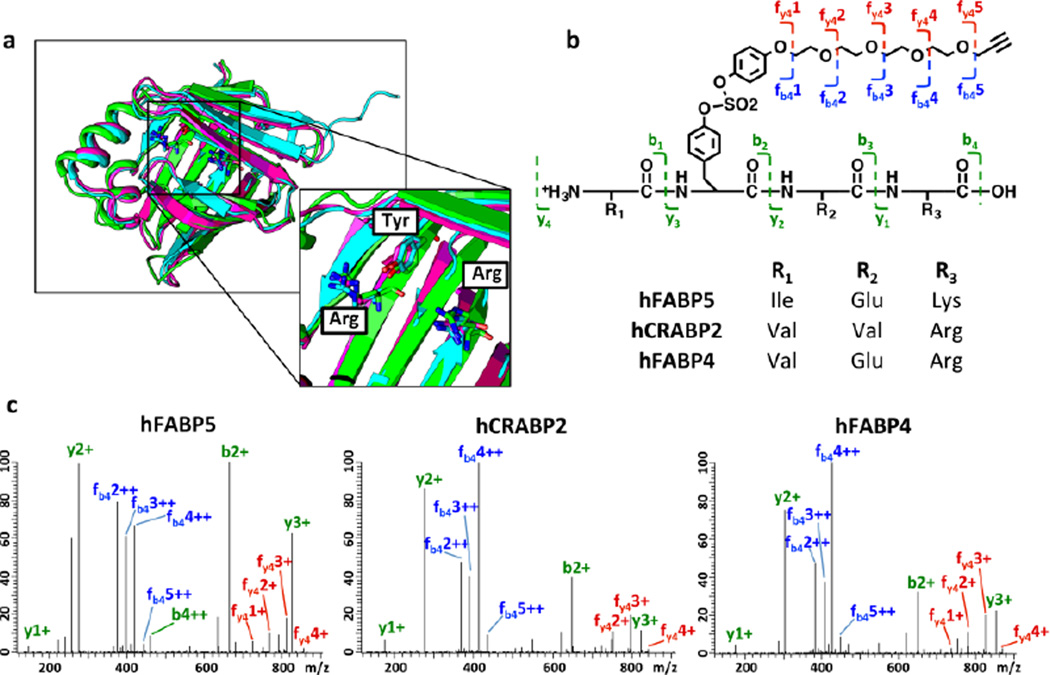
(a) Structural alignment of apo CRABP2 (PDBID: 2FS6, green), FABP5 (4LKP, magenta) and FABP4 (3RZY, cyan) in ribbon format by superposition of their protein backbones. Protein side chains of the Arg~Arg~Tyr module are depicted in stick format. (b-c) Tyrosine modification in CRABP2, FABP5 and FABP4 by probe 1, demonstrated by LC-MS/MS analysis. Recombinant proteins (20 µM) were incubated with probe 1 (100 µM) at 25 °C until the modification reached ≥ 80% completion (observed by LC-ESI-MS). Representative fragmentation (MS2) patterns for a peptide containing tyrosine modified with probe 1 is shown for each iLBP analyzed. Identified b and y ions are indicated. A complete list of tryptic peptides identified for each protein, as well as the assignment of the MS2 fragment ion can be found in Supplemental Table S3–S8.
To determine which iLBP amino acid residue reacted with probe 1, we subjected recombinant CRABP2, FABP5 and FABP4 modified by probe 1 to tandem mass spectrometry analysis.46,65,80 Modification by probe 1 was observed at the tyrosine that is part of the conserved proximal Arg~Arg~Tyr carboxylic acid binding motif in all three proteins (Figures 2b–c and Tables S3–S8). These results suggest that other iLBP members that contain this structural motif (including FABP3, 7, 8, 9, 12, and CRABP1; Figure S15) may also react with an appropriately designed arylfluorosulfate via this Tyr preference, although this hypothesis remains to be substantiated given that the structurally simple probe 1 does not seem to modify FABP3 in buffer, and the substrate binding pocket that contains the Arg~Arg~Tyr motif in FABP3 seems readily accessible for probe 1.
The modification of iLBPs by probe 1 was very slow (Figure S2), but this is not surprising considering that probe 1 is structurally simple. The use of structurally more complex probes should increase the equilibrium binding affinity, reduce the binding off-rate, and increase the iLBP residence time, possibly increasing the iLBP modification efficiency. Structural complementarity between the probe and the iLBP should also influence activation efficiency, putatively accomplished by stabilization of the departing F− anion during the SuFEx reaction, thereby increasing the rate of conjugation.1,25,27,29–31,81
Replacing the phenyl substructure of probe 1 by a biphenyl substructure, affording probes 3 and 4 (Figures 3a and 4a, respectively), dramatically increased the conjugation rate with recombinant CRABP2 – this was expected due to the anticipated increase in the equilibrium binding affinity of the probe. Covalent modification of CRABP2 (2 µM) by probe 3 (100 µM) reached completion in 1 h in pH 8.0 buffer at 25 °C (Figure S16), whereas prolonged incubation of probe 3 (100 µM) with recombinant FABP3 (2 µM), FABP5 (2 µM) or FABP4 (2 µM) for up to 24 h did not lead to significant conjugate formation, as discerned by LC-ESI-MS analysis (Figure S17). Probe 3 (100 µM) also efficiently modified recombinant CRABP2 (2 µM) spiked into HeLa cell lysate (total protein = 2 mg/mL), and as expected, only very low non-specific labeling of the HeLa cell proteome occurred (Figure S18).
Figure 3.
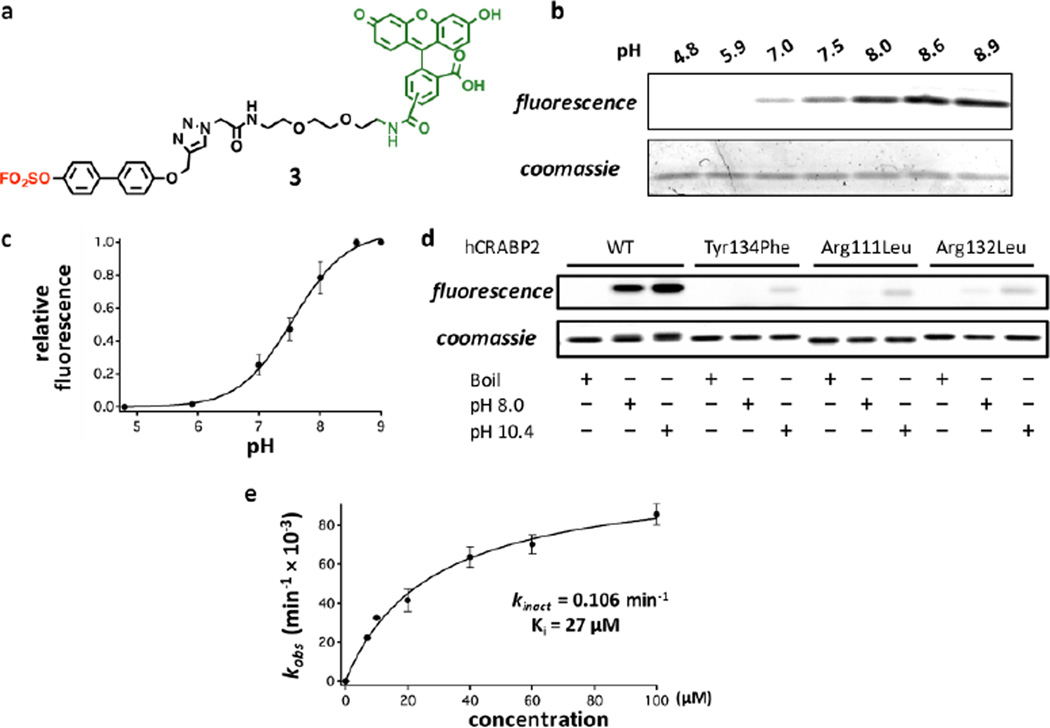
Characterization of the efficient chemoselective modification of CRABP2 by probe 3. (a) Arylfluorosulfate probe 3 bearing a biphenyl moiety. The arylfluorosulfate group is shown in red and the fluorescein moiety is shown in green. (b) pH dependence of modification of CRABP2 (2 µM) by probe 3 (10 µM) in a 10 min reaction period. Top panel: in-gel fluorescence, bottom panel: Coomassie blue staining. (c) Quantification of the relative labeling efficiency from (b) results in an apparent pKa of 7.6 for the Tyr134 phenol group. (d) Comparison of the labeling efficiency for WT, Tyr134Phe, Arg111Leu and Arg132Leu CRABP2 in pH 8.0 and pH 10.4 buffers. Recombinant CRABP2 proteins (2 µM) were incubated with probe 3 (100 µM) at 25 °C for 10 min before quenching the reaction. Top panel: in-gel fluorescence, bottom panel: Coomassie blue staining. (e) Kinetics of probe 3 labeling of CRABP2. Kinetic parameters are indicated. See Experimental Section for definition and calculation of kinetic parameters.
Figure 4.
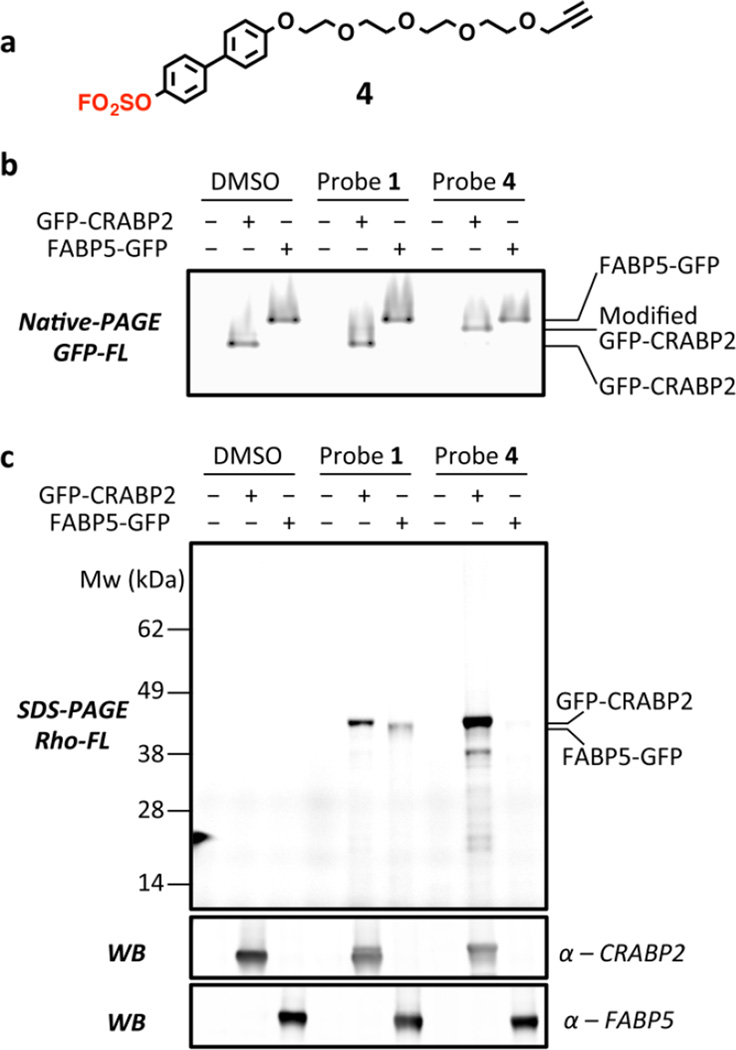
Characterization of the selective modification of overexpressed GFP-CRABP2 by probe 4 in HEK293T cells. (a) Alkyne-functionalized arylfluorosulfate probe 4 bearing a biphenyl moiety. The fluorosulfate group is shown in red. (b) GFP fluorescence of the overexpressed and probe-labeled GFP-CRABP2 and FABP5-GFP analyzed by native PAGE. The modification of GFP-CRABP2 by probe 4 caused a significant band-shift of GFP-CRABP2 to higher molecular weight. The corresponding Coomassie-stained gel is shown in Figure S21. (c) In-gel fluorescence and western blot analysis of the modified GFP-CRABP2 and FABP5-GFP after cell lysis and incorporation of rhodamine-N3 using CuAAC Click. The corresponding Coomassie-stained gels are shown in Figure S22.
We next explored mechanistic aspects of the covalent modification of CRABP2 by arylfluorosulfate probe 3. Understanding how and why arylfluorosulfates chemoselectively modify the conserved Tyr residue in the fatty acid-binding site in a given iLBP should provide insight into the future design and application of arylfluorosulfates in a biological context. Since the Arg~Arg~Tyr carboxylic acid binding module is conserved in many iLBPs, we hypothesized that the presence of the proximal Arg residues in the fatty acid binding site would perturb the pKa of the Tyr phenol. We measured the extent of modification of wild-type (WT) CRABP2 by probe 3 (10 µM) at a fixed time point in buffers varying in pH from 4.9 to 8.9, employing SDS-PAGE for conjugate quantification (Figures 3b–c). Modification of Tyr134 in CRABP2 is pH dependent. The phenol exhibits an apparent pKa of ~7.6, rationalizing why the conserved Tyr in iLBPs is reactive at near-neutral pH. This result suggests that Tyr134, which is susceptible to SuFEx reaction, appears to be pKa perturbed by one or both of the flanking Arg residues. To further examine this hypothesis, we overexpressed and purified three CRABP2 mutants–Arg111Leu, Arg132Leu and Tyr134Phe (Figure S19),82–84 and tested their reactivity with probe 3 (Figure 3d). The Tyr to Phe mutation eliminates the nucleophile required for the conjugation reaction, whereas the Arg to Leu mutations eliminate positive charges that we hypothesize lower the pKa of the neighboring Tyr residue, which appears to be necessary but not sufficient for conjugation reactivity. All three mutants appear to fold properly into WT-like conformations at pH 8.0 and even at pH 10.4, as discerned by their circular dichroism spectra (Figures S20–21). As expected, probe 3 was not able to modify the Tyr134Phe CRABP2 mutant at pH 8 (Figure 3d; for longer exposure see Figure S22). Single mutation of either of the Arg residues to Leu greatly impaired the modification of CRABP2 by probe 3 (Figure 3d), even at an elevated pH of 10.4, which is above the pKa of a normal Tyr residue. The inability of probe 3 to react with the phenolate in the Arg-to-Leu mutants suggests that the flanking Arg residues could be catalyzing the sulfur exchange reaction in addition to apparently perturbing the pKa of the phenol in iLBPs. At pH 10.4, probe 3 starts to show a low level of non-specific labeling, as revealed by the appearance of a fluorescent signal for the Tyr134Phe mutant, potentially suggesting that other nucleophilic residues such as lysine or additional tyrosine residues could become reactive (e.g., Tyr51; Figure 3d). Accordingly, the extent of labeling of WT CRABP2 at pH 10.4 buffer is elevated compared to pH 8.0. Overall, it seems that the Arg residues contribute to the chemoselective modification of Tyr134 by probe 3 at neutral pH, by means of perturbing the pKa of Tyr134 and/or by stabilizing the departing fluoride anion.
We hypothesized that the non-covalent equilibrium binding of probe 3 to CRABP2 and the concomitant activation of the fluorosulfate is important for fast conjugation. To measure the labeling kinetics of CRABP2 by probe 3 (Figure 3e), we incubated recombinant CRABP2 (2.5 µM) in pH 8.0 buffer with increasing concentrations of probe 3. The probe 3-CRABP2 conjugation reaction is rate limited by the chemical step (our data indicate that probe binding and disassociation occur more rapidly than conjugate formation) at the selected concentrations of probe 3 (7–100 µM). The observed rates of the emergence of conjugate fluorescence report on the rate of conjugate formation. We calculated the kobs by quantifying the in-gel fluorescence after quenching the reaction at different time points. Steady-state kinetic parameters were obtained from kobs yielding a kinact of 0.106 min−1 and a Ki of 27 µM.65 The rate of probe 3-CRABP2 conjugation is observed to plateau at higher concentrations of probe 3 (Figure 3e), suggesting that the equilibrium binding of probe 3 to CRABP2, in part, facilitates the high efficiency labeling of CRABP2 by arylfluorosulfate 3.
We next sought to test the selectivity of the biphenyl arylfluorosulfate core structure for modification of CRABP2 over other iLPBs in live cells. The fluorescein moiety in probe 3, which impairs cell permeability, was replaced with an ethylene glycol substructure terminating in an alkyne handle. The resulting biphenyl-based arylfluorosulfate probe 4 (Figure 4a) was envisioned to be more cell permeable. The conjugation capacity of probe 4 was evaluated by incubating this arylfluorosulfate (100 µM) with recombinant CRABP2, FABP3, FABP4, or FABP5 (2 µM) at 25 °C in buffer for 1 h before subjecting the samples to LC-ESI-MS analysis. Analogous to the reaction selectivity and efficiency exhibited by probe 3, probe 4 quantitatively modified CRABP2 within 1 h (Figure S23), but did not modify FABP3, FABP4 and FABP5, even when incubated for 24 h (Figure S24). To test the reaction in live cells, we overexpressed an N-terminal GFP fusion of CRABP2 (GFP-CRABP2) or a C-terminal fusion of GFP to FABP5 (FABP5-GFP) in HEK293T cells.62,63 Their expression levels were comparable, as observed by the GFP fluorescence in native PAGE analysis (Figures 4b and S25). Treating the GFP-CRABP2 or FABP5-GFP transfected cells with probe 1 (20 µM) for 2 h resulted in apparent modification of GFP-CRABP2 and to a lesser extent FABP5-GFP, as observed by SDS-PAGE in-gel fluorescence analysis after clicking on a rhodamine fluorophore (Figures 4c and S26). In contrast, treatment with compound 4 resulted in modification of GFP-CRABP2, but not of FABP5-GFP (Figure 4c). The near-complete modification of GFP-CRABP2 with probe 4 is also apparent from a gel-shift of the protein-probe conjugate in the native PAGE (Figures 4b and S25). Rhodamine fluorescence of GFP-CRABP2 modified with probe 4 is much stronger than when treated with probe 1, consistent with probe 4 being a much more efficient modifier of CRABP2 (Figure 4c). These data suggest that probe 4 could be used to efficiently and selectively label CRABP2 with very low background proteome reactivity in vivo. The ≈38 kDa band in Figure 4c is not observed in non-transfected 293T cells or in FABP5-GFP transfected 293T cells, thus it is likely a truncated GFP-CRABP2 product (Figure S27).
In an attempt to gain structural insights into how probe 4 modifies CRABP2 with high efficiency and selectivity, we purified a tag-less version of CRABP2 and formed its conjugate with probe 4 (Figure S28) for crystallography experiments. We crystallized this conjugate and determined its structure at 1.75 Å resolution (Table S9). The diarylsulfate linkage between Tyr134 and the remaining substructure of probe 4 was clearly visible in the electron density map (Figures 5a and S29). The electron density for most of the PEG and the terminal alkyne could not be observed or reliably modeled, suggesting that the PEG substructure is extended towards the outer edge of the ligand-binding pocket and is solvent-exposed and flexible in the CRABP2-probe 4 conjugate. The biphenyl substructure binds within the spacious ligand- binding pocket of CRABP2 in at least two different conformations (Figure 5a; see Figures S29 and S30 for the conjugate structure showing both modeled conformations of the remaining substructure of Probe 4). The biphenyl substructure of probe 4 is within van der Waals distance of many hydrophobic side chains lining the ligand-binding pocket of CRABP2, suggesting the importance of the hydrophobic effect in the equilibrium binding of probe 4 to CRABP2 (Figures 5a and S30). Structural alignment of the protein backbone of the CRABP2-probe 4 conjugate (Figure S30) in comparison to the RA-bound CRABP2 structure of Geiger and coworkers (Figure 5b, PDBID: 2FR3)69 revealed that the protein moieties align very well (RMSD = 0.44 Å), except for slight differences in the conformation of the Thr56-Arg59 β-turn. The probe 4-derived substructure of the conjugate occupies almost the same space as that of RA bound to CRABP2 (cf. Figures 5a and 5b or inspect Figure S31, which is a composite of Figures S30 and 5b. The outer ring of probe 4 and the adjacent PEG moiety are in a position where CRABP2 has a significantly larger pocket than FABP4 and FABP5 (Figure S32), which might be important in accommodating the bulky trimethylcyclohexene ring component of retinoic acid (Figures 5b and S31). We hypothesize that before the irreversible step of probe 4 reacting with CRABP2 occurs, that the biphenyl substructure of probe 4 binds in a position closer to the outer edge of the ligand binding pocket and, therefore, could require the whole extended binding pocket. This is consistent with the fact that the B-values of the biphenyl and PEG substructures of the conjugate are greater than for the protein, suggesting a sub-optimal interaction between 4 and CRABP2 after conjugate formation. Overall, the structural results confirmed the presence of a diarylsulfate ester connection derived from the reaction between probe 4 and Tyr134 of CRABP2. These results also demonstrate that structural complementarity between the arylfluorosulfate probe and CRABP2 was sufficient to achieve the high efficiency labeling and notable in vivo selectivity. In the structure of the CRABP2-arylfluorosulfate conjugate, the Arg111 and Arg132 side chains both coordinate to the diarylsulfate ester moiety, either directly (Arg132) or through two bridging water molecules (Arg111) (Figure S33). Thus, both side chains could facilitate or catalyze conjugate formation through hydrogen-bonding to the departing fluoride anion.
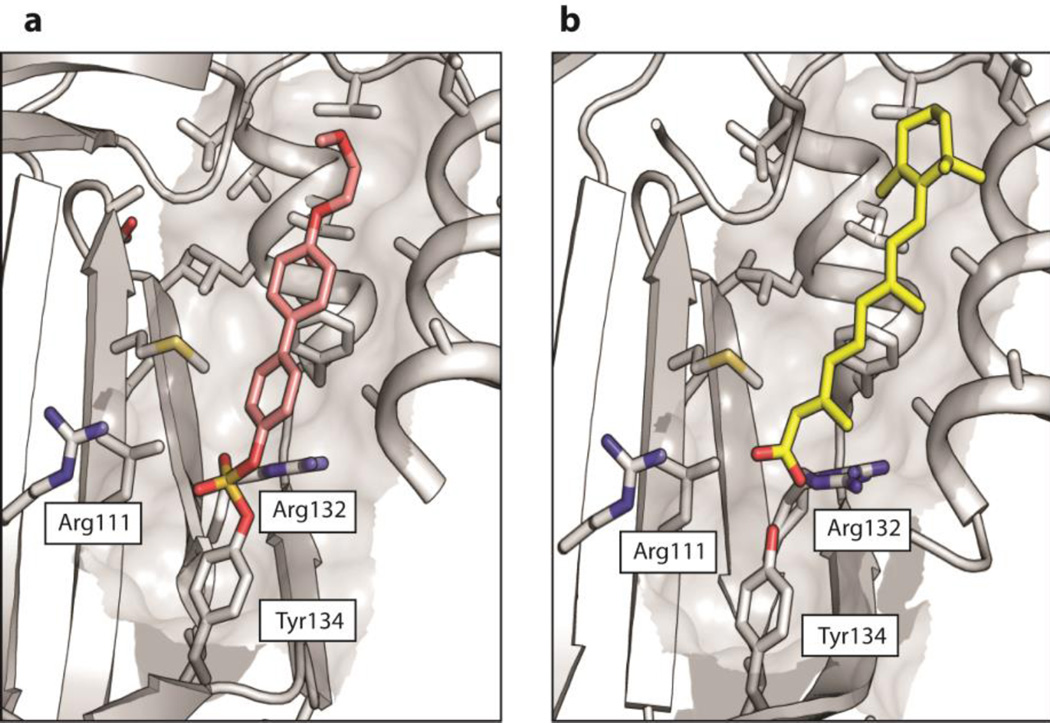
Figure 5.
(a) Crystal structure of the probe 4-CRABP2 covalent conjugate at 1.75 Å resolution. The protein moiety is shown in gray and the protein-conjugated probe 4 is shown in pink (only one of two alternative conformations shown for clarity). See Figure S25 for a depiction with both conformations shown. Protein side chains within 5 Å of the probe 4 are depicted in stick format. The side chains of the Arg~Arg~Tyr module are indicated. (b) Analogous view of CRABP2 bound to retinoic acid (RA), shown in yellow. PDB:2FR3.
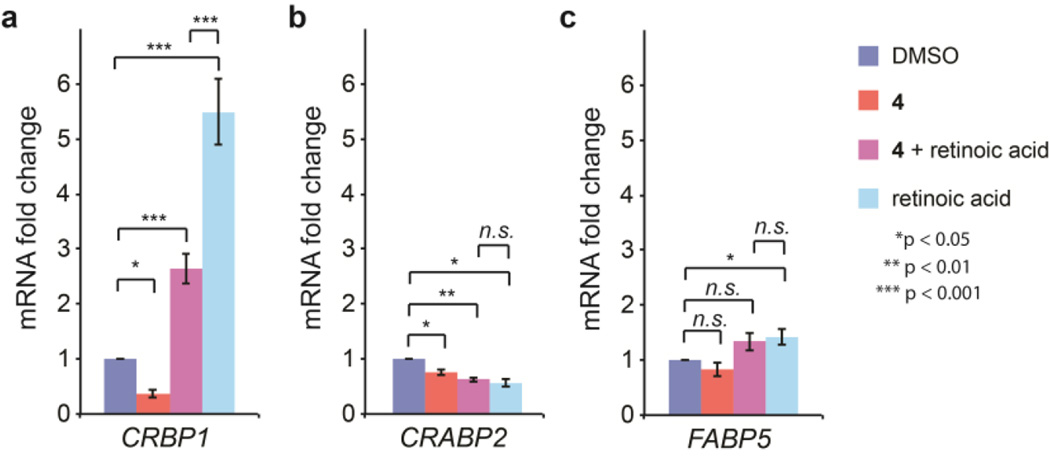
A critical function of certain iLBP family members62,69,85–87 is to transport RA across the nuclear membrane to deliver it to distinct RA receptors (RAR), which are also transcription factors.88–92 These receptors initiate diverse cellular processes, such as differentiation, cell cycle arrest, apoptosis, or proliferation, through transcriptional activation. For instance, CRABP2 delivers RA to RARα, whose transcriptional program leads to apoptosis.93 In contrast, FABP5 carries RA or long-chain fatty acids to peroxisome proliferator-activated receptor (PPAR) β/δ, and its subsequent transcriptional response leads to proliferation.94 We tested whether arylfluorosulfate 4, which selectively modifies CRABP2, could directly inhibit CRABP2 delivery of RA to RARα, which in MCF-7 mammary carcinoma cells has a growth inhibitory and proapoptotic role.63,87 As a readout, we used qPCR to quantify CRBP1 mRNA – a target gene downstream of CRABP2-mediated RARα-RA transcriptional reprogramming.95 RA treatment of MCF-7 cells led to a 5.5-fold induction of CRBP1 mRNA (Figure 6a). Pretreatment with compound 4 attenuated this induction by 2-fold (Figure 6a), demonstrating inhibition of RA signaling. Interestingly, treatment with 4 in the absence of exogenous RA resulted in a repression of CRBP1 transcription, suggesting that the fluorosulfate could also inhibit basal RA signaling. In contrast, transcription of FABP5 and CRABP2 (genes that are not transcriptionally regulated by RARα and therefore serve as controls) were only minimally changed by treatment with 4 or RA (Figures 6b–c), demonstrating the lack of a general transcriptional effect by arylfluorosulfates. These results show that treatment with 4 attenuated RARα transcriptional reprogramming mediated by CRABP2•RA delivery to RARα. Future experiments will scrutinize the selectivity of the RA-associated transcriptional reprogramming mediated by 4 and its dependency on CRABP2.
Figure 6.
Inhibition of CRABP2/RARα target gene induction by probe 4. (a) mRNA transcript levels of the RA signaling target gene CRBP1 as measured by qPCR in MCF-7 cells. MCF-7 cells in charcoal-treated FBS were pretreated with 20 µM 4 or DMSO for 4 h and subsequently treated with 100µM RA or EtOH for 24 h. (b-c) mRNA transcript levels of CRABP2 (b) and FABP5 (c) in MCF-7 cells treated under the same conditions as in (a). Error bars show SEM (n = 4). *p < 0.05, **p < 0.01, ***p < 0.001.
DISCUSSION
Only a few dozen chemical abstract service entries exist for arylfluorosulfates over the past 70 years, largely because of the difficulty associated with their preparation. Arylfluorosulfates are now easily prepared from phenols and SO2F2.1 Arylfluorosulfates appear to be well-suited for targeting specific proteins within the human proteome. From the results presented herein, the tyrosine side chain hydroxyl group appears to be a favored reactant, when found in the proper protein context. Even though the exact pKa of the Tyr residue in the Arg~Arg~Tyr carboxylic acid binding motif within iLBPs has not been accurately measured, it is unlikely that pKa perturbation alone is enough to mediate Tyr reactivity based on the limited number of conjugates formed in cells treated with the various arylfluorosulfate probes and given that a shift to higher pH did not increase conjugate formation for CRABP2 Arg111 or 132Leu. In addition to pKa perturbation, we hypothesize that the surrounding protein environment is essential to help catalyze the SuFEx reaction. We hypothesize that hydrogen bonding from a reaction-susceptible protein to the departing fluoride ion in the transition state and/or having the appropriate electrostatic field effects in a reaction susceptible protein and/or a complementary amphipathic environment in a reaction-susceptible protein is critical for arylfluorosulfate reactivity.1,25–37 Comparing structurally analogous aryl sulfonyl fluorides and arylfluorosulfates, the latter are dramatically less reactive, yet are reactive enough to potentially become very selective covalent probes.
Furthermore, the arylfluorosulfate functional group is less reactive than well-established electrophiles that have been used in protein profiling probes, such as fluorophosphonates,18,19 vinyl sulfones,20–22 phenylsulfonate esters,58,96,97 epoxides,98,99 and α,β-unsaturated carbonyls (e.g. acrylamides).23,24 The higher reactivity of these functional groups means that they cannot, in general, distinguish between members of enzyme families,100 but are nevertheless very useful for functional annotation of poorly characterized enzyme families, discovery of novel enzyme inhibitors as therapeutics, and for developing biomarkers.16,19,52,55,58,60,98,100–102 In contrast, arylfluorosulfates (e.g., 4) seem to be able to react selectively with one member of a protein family. The lower reactivity of arylfluorosulfates means that they rely more on secondary criteria, i.e. relatively high affinity equilibrium binding to those proteins (reflected by Ki or KD) containing an appropriate arylfluorosulfate-activating environment in their binding site, in addition to having an appropriate proximal nucleophile.
Numerous FDA-approved drugs function by forming a covalent conjugate with their targets; examples include beta-lactam antibiotics, aspirin, omeprazole, the blockbuster drug clopidogrel, and the more recently approved anti-cancer drugs afatinib and ibrutinib.102–104 While the irreversible mode of action for many covalent drugs was discovered after their clinical use had already become widespread, there is a growing interest in combining activity-based protein profiling with medicinal chemistry to tune the reversible binding and reactivity of covalent drugs for their protein target, while minimizing off-target reactivity.101,102,105–108 For this strategy to succeed, the employed electrophiles must be of low intrinsic reactivity to avoid potential off-target reactivity. Successful recent examples include the targeting of cysteines in kinases by acrylamides.109 Arylfluorosulfates seem to have potential in this regard.
The non-enzyme family of iLBPs targeted by the simple arylfluorosulfates studied herein comprises FABPs, CRABPs and cytoplasmic retinol binding proteins (CRBPs).47–50,52,62,76,110 CRABP2 is an important modulator of RA-dependent signaling, which drives some cancers.62,63,85,87,111 CRABP2 transports RA from the cytoplasm to the nucleus, facilitating RA binding to the nuclear hormone receptor RARα and possibly others.62,63,87 FABP5 is one of the most ubiquitously expressed FABPs and binds a wide array of ligands including fatty acids, fatty acid metabolites, retinoic acid and numerous synthetic drugs and probes.49 FABP5-mediated signaling through the nuclear hormone receptor PPARβ/δ has been found to be involved in a range of pathologies, including metabolic syndrome, atherosclerosis and cancer.85,86,112,113 Interestingly, CRABP2 and FABP5 appear to compete for RA delivery from the cytoplasm to their respective nuclear hormone receptors, mediating transcriptional programs that control the fate of cancer cells.63 Another well-studied iLBP, FABP4, has been suggested to be important for the development of metabolic syndrome, through its distinct actions in adipocytes and macrophages related to metabolic and inflammatory responses.48,114–118 Inhibitors of FABP4 have proven to be useful for the treatment of diabetes and atherosclerosis in mouse models.48,114,119–122 Consequently, arylfluorosulfate-based pharmacological agents that modify selective iLBP function(s) may offer therapeutic opportunities for these diseases. Arylfluorosulfate probes that selectively target a given iLBP should better illuminate the function of that family member, including their role in nuclear hormone receptor transcriptional reprogramming of cells.89–92 Further studies of the mechanistic details associated with covalent labeling of iLBPs by arylfluorosulfate probes will likely provide insight into the design of arylfluorosulfate probes exhibiting high reaction efficiency and high selectivity for covalently modifying a specific protein within living cells.
CONCLUSIONS
We presented simple arylfluorosulfates to the proteome of HeLa cells and then used the CuAAC click reaction to attach a biotin affinity chromatography handle to identify proteins that were covalently linked to the sulfur center of the probe using mass spectrometry. We found that arylfluorosulfate probes could modify iLBPs with low background labeling of the proteome. The probes modify iLBPs at a tyrosine that is part of a conserved carboxylic acid binding motif comprising two Arg residues and a Tyr. The proximal arginine residues appear to lower the pKa of the Tyr residue and may help catalyze the sulfur exchange reaction by enabling fluoride anion departure. We anticipate that optimizing the structure of arylfluorosulfate probes for stronger reversible binding to the target proteins will enhance the chemical modification efficiency and specificity for a given protein, as exemplified by the selective modification of CRABP2 by 3 and 4. Efforts underway to use structurally distinct arylfluorosulfates to target other proteins comprising the cellular proteome will create powerful and valuable chemical tools for studying and perturbing protein function in living systems. The greatest opportunity may be to use an activatable arylfluorosulfate electrophile to covalently target a genetically validated protein to address an unmet medical need.45,101,102,105–108
Supplementary Material
Acknowledgments
This work was supported by the Skaggs Institute for Chemical Biology, the Lita Annenberg Hazen Foundation, and by National Institutes of Health grants GM117154 (K.B.S.), DK106582 (J.W.K.), CA087660 (B.F.C.), DK099810 (E.S.), and AI117675 (I.A.W.). L.P. was supported by a grant from the Leukemia and Lymphoma Society. D.E.M. was supported by a grant from the Hewitt Foundation. Portions of this research were carried out at the Stanford Synchrotron Radiation Lightsource, a Directorate of SLAC National Accelerator Laboratory and an Office of Science User Facility operated for the U.S. Department of Energy Office of Science by Stanford University. The SSRL Structural Molecular Biology Program is supported by the DOE Office of Biological and Environmental Research and by the National Institutes of Health, National Institute of General Medical Sciences (including P41GM103393) and the National Center for Research Resources (P41RR001209).
Footnotes
ASSOCIATED CONTENT
Supporting information. Supplementary Figures and Tables are supplied as Supporting Information. This material is available free of charge via the Internet at http://pubs.acs.org.
The structure of the CRABP2-probe 4 conjugate has been deposited in the Protein Data Bank under accession code 5HZQ.
The authors declare no competing financial interests.
REFERENCES
- 1.Dong J, Krasnova L, Finn MG, Sharpless KB. Angew Chem Int Ed Engl. 2014;53:9430. doi: 10.1002/anie.201309399. [DOI] [PubMed] [Google Scholar]
- 2.Baker BR. Annu Rev Pharmacol. 1970;10:35. doi: 10.1146/annurev.pa.10.040170.000343. [DOI] [PubMed] [Google Scholar]
- 3.Baker BR. Ann N Y Acad Sci. 1971;186:214. doi: 10.1111/j.1749-6632.1971.tb31146.x. [DOI] [PubMed] [Google Scholar]
- 4.Baker BR, Erickson EH. J Med Chem. 1968;11:245. doi: 10.1021/jm00308a011. [DOI] [PubMed] [Google Scholar]
- 5.Baker BR, Hurlbut JA. J Med Chem. 1968;11:233. doi: 10.1021/jm00308a009. [DOI] [PubMed] [Google Scholar]
- 6.Baker BR, Hurlbut JA. J Med Chem. 1968;11:241. doi: 10.1021/jm00308a010. [DOI] [PubMed] [Google Scholar]
- 7.Baker BR, Kozma JA. J Med Chem. 1968;11:652. doi: 10.1021/jm00310a005. [DOI] [PubMed] [Google Scholar]
- 8.Baker BR, Kozma JA. J Med Chem. 1968;11:656. doi: 10.1021/jm00310a005. [DOI] [PubMed] [Google Scholar]
- 9.Baker BR, Lourens GJ. J Med Chem. 1967;10:1113. doi: 10.1021/jm00318a029. [DOI] [PubMed] [Google Scholar]
- 10.Baker BR, Lourens GJ. J Med Chem. 1968;11:38. doi: 10.1021/jm00307a007. [DOI] [PubMed] [Google Scholar]
- 11.Baker BR, Lourens GJ. J Med Chem. 1968;11:677. doi: 10.1021/jm00310a009. [DOI] [PubMed] [Google Scholar]
- 12.Baker BR, Meyer RB. J Med Chem. 1968;11:489. doi: 10.1021/jm00309a018. [DOI] [PubMed] [Google Scholar]
- 13.Baker BR, Wood WF. J Med Chem. 1968;11:650. doi: 10.1021/jm00310a006. [DOI] [PubMed] [Google Scholar]
- 14.Hett EC, Xu H, Geoghegan KF, Gopalsamy A, Kyne RE, Jr, Menard CA, Narayanan A, Parikh MD, Liu S, Roberts L, Robinson RP, Tones MA, Jones LH. ACS Chem Biol. 2015;10:1094. doi: 10.1021/cb5009475. [DOI] [PubMed] [Google Scholar]
- 15.Narayanan A, Jones LH. Chem Sci. 2015;6:2650. doi: 10.1039/c5sc00408j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gu C, Shannon DA, Colby T, Wang Z, Shabab M, Kumari S, Villamor JG, McLaughlin CJ, Weerapana E, Kaiser M, Cravatt BF, van der Hoorn RA. Chem Biol. 2013;20:541. doi: 10.1016/j.chembiol.2013.01.016. [DOI] [PubMed] [Google Scholar]
- 17.Grimster NP, Connelly S, Baranczak A, Dong J, Krasnova LB, Sharpless KB, Powers ET, Wilson IA, Kelly JW. J Am Chem Soc. 2013;135:5656. doi: 10.1021/ja311729d. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Webb EC. Biochem J. 1948;42:96. doi: 10.1042/bj0420096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Cravatt BF, Sorensen EJ. Curr Opin Chem Biol. 2000;4:663. doi: 10.1016/s1367-5931(00)00147-2. [DOI] [PubMed] [Google Scholar]
- 20.Bogyo M, McMaster JS, Gaczynska M, Tortorella D, Goldberg AL, Ploegh H. Proc Natl Acad Sci U S A. 1997;94:6629. doi: 10.1073/pnas.94.13.6629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Palmer JT, Rasnick D, Klaus JL, Bromme D. J Med Chem. 1995;38:3193. doi: 10.1021/jm00017a002. [DOI] [PubMed] [Google Scholar]
- 22.Roush WR, Gwaltney SL, Cheng JM, Scheidt KA, McKerrow JH, Hansell E. J Am Chem Soc. 1998;120:10994. [Google Scholar]
- 23.Fry DW, Bridges AJ, Denny WA, Doherty A, Greis KD, Hicks JL, Hook KE, Keller PR, Leopold WR, Loo JA, McNamara DJ, Nelson JM, Sherwood V, Smaill JB, Trumpp-Kallmeyer S, Dobrusin EM. Proc Natl Acad Sci U S A. 1998;95:12022. doi: 10.1073/pnas.95.20.12022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Schirmer A, Kennedy J, Murli S, Reid R, Santi DV. Proc Natl Acad Sci U S A. 2006;103:4234. doi: 10.1073/pnas.0600445103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Dong J, Sharpless KB, Kwisnek L, Oakdale JS, Fokin VV. Angew Chem Int Ed Engl. 2014;53:9466. doi: 10.1002/anie.201403758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Bourne Y, Sharpless KB, Taylor P, Marchot P. J Am Chem Soc. 2016;138:1611. doi: 10.1021/jacs.5b11384. [DOI] [PubMed] [Google Scholar]
- 27.Fried SD, Bagchi S, Boxer SG. Science. 2014;346:1510. doi: 10.1126/science.1259802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Shi Y, Acerson MJ, Shuford KL, Shaw BF. ACS Chem Neurosci. 2015;6:1696. doi: 10.1021/acschemneuro.5b00146. [DOI] [PubMed] [Google Scholar]
- 29.Menger FM. Chem Soc Rev. 1972;1:229. [Google Scholar]
- 30.Jung Y, Marcus RA. J Am Chem Soc. 2007;129:5492. doi: 10.1021/ja068120f. [DOI] [PubMed] [Google Scholar]
- 31.Beattie JK, McErlean CSP, Phippen CBW. Chemistry. 2010;16:8972. doi: 10.1002/chem.201001705. [DOI] [PubMed] [Google Scholar]
- 32.Ganti G, Mccammon JA. J Mol Graphics. 1986;4:200. [Google Scholar]
- 33.Guo D, Zhu D, Zhou X, Zheng B. Langmuir. 2015;31:13759. doi: 10.1021/acs.langmuir.5b04031. [DOI] [PubMed] [Google Scholar]
- 34.Klijn JE, Engberts JB. Nature. 2005;435:746. doi: 10.1038/435746a. [DOI] [PubMed] [Google Scholar]
- 35.McFearin CL, Richmond GL. J Phys Chem C. 2009;113:21162. [Google Scholar]
- 36.Narayan S, Muldoon J, Finn MG, Fokin VV, Kolb HC, Sharpless KB. Angew Chem Int Ed Engl. 2005;44:3275. doi: 10.1002/anie.200462883. [DOI] [PubMed] [Google Scholar]
- 37.Scatena LF, Richmond GL. Chem Phys Lett. 2004;383:491. [Google Scholar]
- 38.Richmond GL. Chem Rev. 2002;102:2693. doi: 10.1021/cr0006876. [DOI] [PubMed] [Google Scholar]
- 39.Sharpless KB, Kolb HC. Abstr Pap Am Chem S. 1999;217:U95. [Google Scholar]
- 40.Kolb HC, Finn MG, Sharpless KB. Angew Chem Int Ed Engl. 2001;40:2004. doi: 10.1002/1521-3773(20010601)40:11<2004::AID-ANIE2004>3.0.CO;2-5. [DOI] [PubMed] [Google Scholar]
- 41.Wang Q, Chan TR, Hilgraf R, Fokin VV, Sharpless KB, Finn MG. J Am Chem Soc. 2003;125:3192. doi: 10.1021/ja021381e. [DOI] [PubMed] [Google Scholar]
- 42.Deutsch DG, Lin S, Hill WA, Morse KL, Salehani D, Arreaza G, Omeir RL, Makriyannis A. Biochem Biophys Res Commun. 1997;231:217. doi: 10.1006/bbrc.1997.6072. [DOI] [PubMed] [Google Scholar]
- 43.Powers JC, Tanaka T, Harper JW, Minematsu Y, Barker L, Lincoln D, Crumley KV, Fraki JE, Schechter NM, Lazarus GG, et al. Biochemistry. 1985;24:2048. doi: 10.1021/bi00329a037. [DOI] [PubMed] [Google Scholar]
- 44.Shannon DA, Gu C, McLaughlin CJ, Kaiser M, van der Hoorn RA, Weerapana E. Chembiochem. 2012;13:2327. doi: 10.1002/cbic.201200531. [DOI] [PubMed] [Google Scholar]
- 45.Shannon DA, Weerapana E. Curr Opin Chem Biol. 2015;24:18. doi: 10.1016/j.cbpa.2014.10.021. [DOI] [PubMed] [Google Scholar]
- 46.Baranczak A, Liu Y, Connelly S, Du WG, Greiner ER, Genereux JC, Wiseman RL, Eisele YS, Bradbury NC, Dong J, Noodleman L, Sharpless KB, Wilson IA, Encalada SE, Kelly JW. J Am Chem Soc. 2015;137:7404. doi: 10.1021/jacs.5b03042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Bernlohr DA, Simpson MA, Hertzel AV, Banaszak LJ. Annu Rev Nutr. 1997;17:277. doi: 10.1146/annurev.nutr.17.1.277. [DOI] [PubMed] [Google Scholar]
- 48.Furuhashi M, Hotamisligil GS. Nat Rev Drug Discov. 2008;7:489. doi: 10.1038/nrd2589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Smathers RL, Petersen DR. Hum Genomics. 2011;5:170. doi: 10.1186/1479-7364-5-3-170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Banaszak L, Winter N, Xu Z, Bernlohr DA, Cowan S, Jones TA. Adv Protein Chem. 1994;45:89. doi: 10.1016/s0065-3233(08)60639-7. [DOI] [PubMed] [Google Scholar]
- 51.Maeda K, Cao H, Kono K, Gorgun CZ, Furuhashi M, Uysal KT, Cao Q, Atsumi G, Malone H, Krishnan B, Minokoshi Y, Kahn BB, Parker RA, Hotamisligil GS. Cell Metab. 2005;1:107. doi: 10.1016/j.cmet.2004.12.008. [DOI] [PubMed] [Google Scholar]
- 52.Hulce JJ, Cognetta AB, Niphakis MJ, Tully SE, Cravatt BF. Nat Methods. 2013;10:259. doi: 10.1038/nmeth.2368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Tornøe CW, Christensen C, Meldal M. J Org Chem. 2002;67:3057. doi: 10.1021/jo011148j. [DOI] [PubMed] [Google Scholar]
- 54.Rostovtsev VV, Green LG, Fokin VV, Sharpless KB. Angew Chem Int Ed Engl. 2002;41:2596. doi: 10.1002/1521-3773(20020715)41:14<2596::AID-ANIE2596>3.0.CO;2-4. [DOI] [PubMed] [Google Scholar]
- 55.Speers AE, Adam GC, Cravatt BF. J Am Chem Soc. 2003;125:4686. doi: 10.1021/ja034490h. [DOI] [PubMed] [Google Scholar]
- 56.Hong V, Presolski SI, Ma C, Finn MG. Angew Chem Int Ed Engl. 2009;48:9879. doi: 10.1002/anie.200905087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Besanceney-Webler C, Jiang H, Zheng T, Feng L, Soriano del Amo D, Wang W, Klivansky LM, Marlow FL, Liu Y, Wu P. Angew Chem Int Ed Engl. 2011;50:8051. doi: 10.1002/anie.201101817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Speers AE, Cravatt BF. Chem Biol. 2004;11:535. doi: 10.1016/j.chembiol.2004.03.012. [DOI] [PubMed] [Google Scholar]
- 59.Presolski SI, Hong VP, Finn MG. Curr Protoc Chem Biol. 2011;3:153. doi: 10.1002/9780470559277.ch110148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Weerapana E, Wang C, Simon GM, Richter F, Khare S, Dillon MB, Bachovchin DA, Mowen K, Baker D, Cravatt BF. Nature. 2010;468:790. doi: 10.1038/nature09472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Klock HE, Lesley SA. Methods Mol Biol. 2009;498:91. doi: 10.1007/978-1-59745-196-3_6. [DOI] [PubMed] [Google Scholar]
- 62.Budhu AS, Noy N. Mol Cell Biol. 2002;22:2632. doi: 10.1128/MCB.22.8.2632-2641.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Schug TT, Berry DC, Shaw NS, Travis SN, Noy N. Cell. 2007;129:723. doi: 10.1016/j.cell.2007.02.050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Schneider CA, Rasband WS, Eliceiri KW. Nat Methods. 2012;9:671. doi: 10.1038/nmeth.2089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Liu Y, Zhang X, Tan YL, Bhabha G, Ekiert DC, Kipnis Y, Bjelic S, Baker D, Kelly JW. J Am Chem Soc. 2014;136:13102. doi: 10.1021/ja5056356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Evans P. Acta Crystallogr D Biol Crystallogr. 2006;62:72. doi: 10.1107/S0907444905036693. [DOI] [PubMed] [Google Scholar]
- 67.French S, Wilson K. Acta Crystallogr A. 1978;34:517. [Google Scholar]
- 68.McCoy AJ, Grosse-Kunstleve RW, Adams PD, Winn MD, Storoni LC, Read RJ. J Appl Crystallogr. 2007;40:658. doi: 10.1107/S0021889807021206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Vaezeslami S, Mathes E, Vasileiou C, Borhan B, Geiger JH. J Mol Biol. 2006;363:687. doi: 10.1016/j.jmb.2006.08.059. [DOI] [PubMed] [Google Scholar]
- 70.Winn MD, Ballard CC, Cowtan KD, Dodson EJ, Emsley P, Evans PR, Keegan RM, Krissinel EB, Leslie AG, McCoy A, McNicholas SJ, Murshudov GN, Pannu NS, Potterton EA, Powell HR, Read RJ, Vagin A, Wilson KS. Acta Crystallogr D Biol Crystallogr. 2011;67:235. doi: 10.1107/S0907444910045749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Winn MD, Murshudov GN, Papiz MZ. Methods Enzymol. 2003;374:300. doi: 10.1016/S0076-6879(03)74014-2. [DOI] [PubMed] [Google Scholar]
- 72.Vagin AA, Steiner RA, Lebedev AA, Potterton L, McNicholas S, Long F, Murshudov GN. Acta Crystallogr D Biol Crystallogr. 2004;60:2184. doi: 10.1107/S0907444904023510. [DOI] [PubMed] [Google Scholar]
- 73.Smart OS, Womack TO, Sharff A, Flensburg C, Keller P, Paciorek W, Vonrhein C, Bricogne G. Cambridge, United Kingdom: Global Phasing Ltd; 2011. http://www.globalphasing.com. [Google Scholar]
- 74.Ong SE, Blagoev B, Kratchmarova I, Kristensen DB, Steen H, Pandey A, Mann M. Mol Cell Proteomics. 2002;1:376. doi: 10.1074/mcp.m200025-mcp200. [DOI] [PubMed] [Google Scholar]
- 75.Wisniewski JR, Ostasiewicz P, Dus K, Zielinska DF, Gnad F, Mann M. Mol Syst Biol. 2012;8:611. doi: 10.1038/msb.2012.44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Storch J, Corsico B. Annu Rev Nutr. 2008;28:73. doi: 10.1146/annurev.nutr.27.061406.093710. [DOI] [PubMed] [Google Scholar]
- 77.Armstrong EH, Goswami D, Griffin PR, Noy N, Ortlund EA. J Biol Chem. 2014;289:14941. doi: 10.1074/jbc.M113.514646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Gonzalez JM, Fisher SZ. Acta Crystallogr F Struct Biol Commun. 2015;71:163. doi: 10.1107/S2053230X14027897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Tsai CY, Peh MT, Feng W, Dymock BW, Moore PK. PLoS One. 2015;10:e0119511/1. doi: 10.1371/journal.pone.0119511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Yates JR, 3rd, Eng JK, McCormack AL, Schieltz D. Anal Chem. 1995;67:1426. doi: 10.1021/ac00104a020. [DOI] [PubMed] [Google Scholar]
- 81.Chen WDJ, Li S, Liu Y, Wang Y, Yoon L, Wu P, Sharpless KB, Kelly JW. Angew Chem Int Ed Engl. 2016;55:1835. doi: 10.1002/anie.201509016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Vasileiou C, Wang W, Jia X, Lee KS, Watson CT, Geiger JH, Borhan B. Proteins. 2009;77:812. doi: 10.1002/prot.22495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Berbasova T, Nosrati M, Vasileiou C, Wang W, Lee KS, Yapici I, Geiger JH, Borhan B. J Am Chem Soc. 2013;135:16111. doi: 10.1021/ja404900k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Yapici I, Lee KS, Berbasova T, Nosrati M, Jia X, Vasileiou C, Wang W, Santos EM, Geiger JH, Borhan B. J Am Chem Soc. 2015;137:1073. doi: 10.1021/ja506376j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Chen NN, Li Y, Wu ML, Liu ZL, Fu YS, Kong QY, Chen XY, Li H, Liu J. Exp Dermatol. 2012;21:13. doi: 10.1111/j.1600-0625.2011.01392.x. [DOI] [PubMed] [Google Scholar]
- 86.Liu RZ, Graham K, Glubrecht DD, Germain DR, Mackey JR, Godbout R. Am J Pathol. 2011;178:997. doi: 10.1016/j.ajpath.2010.11.075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Sessler RJ, Noy N. Mol Cell. 2005;18:343. doi: 10.1016/j.molcel.2005.03.026. [DOI] [PubMed] [Google Scholar]
- 88.Braissant O, Foufelle F, Scotto C, Dauca M, Wahli W. Endocrinology. 1996;137:354. doi: 10.1210/endo.137.1.8536636. [DOI] [PubMed] [Google Scholar]
- 89.Mangelsdorf DJ, Thummel C, Beato M, Herrlich P, Schuetz G, Umesono K, Blumberg B, Kastner P, Mark M, et a. Cell. 1995;83:835. doi: 10.1016/0092-8674(95)90199-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Rosen ED, Sarraf P, Troy AE, Bradwin G, Moore K, Milstone DS, Spiegelman BM, Mortensen RM. Mol Cell. 1999;4:611. doi: 10.1016/s1097-2765(00)80211-7. [DOI] [PubMed] [Google Scholar]
- 91.Spiegelman BM. Diabetes. 1998;47:507. doi: 10.2337/diabetes.47.4.507. [DOI] [PubMed] [Google Scholar]
- 92.Evans RM. Science. 1988;240:889. doi: 10.1126/science.3283939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Donato LJ, Noy N. Cancer Res. 2005;65:8193. doi: 10.1158/0008-5472.CAN-05-1177. [DOI] [PubMed] [Google Scholar]
- 94.Levi L, Wang Z, Doud MK, Hazen SL, Noy N. Nat Commun. 2015;6:8794. doi: 10.1038/ncomms9794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Corlazzoli F, Rossetti S, Bistulfi G, Ren M, Sacchi N. PLoS One. 2009;4:e4305. doi: 10.1371/journal.pone.0004305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Hagenstein MC, Sewald N. J Biotechnol. 2006;124:56. doi: 10.1016/j.jbiotec.2005.12.005. [DOI] [PubMed] [Google Scholar]
- 97.Uttamchandani M, Li J, Sun H, Yao SQ. Chembiochem. 2008;9:667. doi: 10.1002/cbic.200700755. [DOI] [PubMed] [Google Scholar]
- 98.Weerapana E, Simon GM, Cravatt BF. Nat Chem Biol. 2008;4:405. doi: 10.1038/nchembio.91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Liu Y, Tan YL, Zhang X, Bhabha G, Ekiert DC, Genereux JC, Cho Y, Kipnis Y, Bjelic S, Baker D, Kelly JW. Proc Natl Acad Sci U S A. 2014;111:4449. doi: 10.1073/pnas.1323268111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Evans MJ, Cravatt BF. Chem Rev. 2006;106:3279. doi: 10.1021/cr050288g. [DOI] [PubMed] [Google Scholar]
- 101.Leung D, Hardouin C, Boger DL, Cravatt BF. Nat Biotechnol. 2003;21:687. doi: 10.1038/nbt826. [DOI] [PubMed] [Google Scholar]
- 102.Singh J, Petter RC, Baillie TA, Whitty A. Nat Rev Drug Discov. 2011;10:307. doi: 10.1038/nrd3410. [DOI] [PubMed] [Google Scholar]
- 103.Modjtahedi H, Cho BC, Michel MC, Solca F. Naunyn-Schmiedeberg's Arch Pharmacol. 2014;387:505. doi: 10.1007/s00210-014-0967-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Byrd JC, Furman RR, Coutre SE, Flinn IW, Burger JA, Blum KA, Grant B, Sharman JP, Coleman M, Wierda WG, Jones JA, Zhao W, Heerema NA, Johnson AJ, Sukbuntherng J, Chang BY, Clow F, Hedrick E, Buggy JJ, James DF, O'Brien S. N Engl J Med. 2013;369:32. doi: 10.1056/NEJMoa1215637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Kalgutkar AS, Dalvie DK. Expert Opin Drug Discovery. 2012;7:561. doi: 10.1517/17460441.2012.688744. [DOI] [PubMed] [Google Scholar]
- 106.Potashman MH, Duggan ME. J Med Chem. 2009;52:1231. doi: 10.1021/jm8008597. [DOI] [PubMed] [Google Scholar]
- 107.Sanderson K. Nat Rev Drug Discov. 2013;12:649. doi: 10.1038/nrd4103. [DOI] [PubMed] [Google Scholar]
- 108.Singh J, Petter RC, Kluge AF. Curr Opin Chem Biol. 2010;14:475. doi: 10.1016/j.cbpa.2010.06.168. [DOI] [PubMed] [Google Scholar]
- 109.Fry DW. Anti-Cancer Drug Des. 2000;15:3. [PubMed] [Google Scholar]
- 110.Storch J, McDermott L. J Lipid Res. 2009;50(Suppl):S126. doi: 10.1194/jlr.R800084-JLR200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Tagore R, Thomas HR, Homan EA, Munawar A, Saghatelian A. J Am Chem Soc. 2008;130:14111. doi: 10.1021/ja806463c. [DOI] [PubMed] [Google Scholar]
- 112.Levi L, Lobo G, Doud MK, von Lintig J, Seachrist D, Tochtrop GP, Noy N. Cancer Res. 2013;73:4770. doi: 10.1158/0008-5472.CAN-13-0384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Gally F, Chu HW, Bowler RP. PLoS One. 2013;8:e51784. doi: 10.1371/journal.pone.0051784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Furuhashi M, Tuncman G, Gorgun CZ, Makowski L, Atsumi G, Vaillancourt E, Kono K, Babaev VR, Fazio S, Linton MF, Sulsky R, Robl JA, Parker RA, Hotamisligil GS. Nature. 2007;447:959. doi: 10.1038/nature05844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Tuncman G, Erbay E, Hom X, De Vivo I, Campos H, Rimm EB, Hotamisligil GS. Proc Natl Acad Sci U S A. 2006;103:6970. doi: 10.1073/pnas.0602178103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Ishimura S, Furuhashi M, Watanabe Y, Hoshina K, Fuseya T, Mita T, Okazaki Y, Koyama M, Tanaka M, Akasaka H, Ohnishi H, Yoshida H, Saitoh S, Miura T. PLoS One. 2013;8:e81318. doi: 10.1371/journal.pone.0081318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Cao H, Gerhold K, Mayers JR, Wiest MM, Watkins SM, Hotamisligil GS. Cell. 2008;134:933. doi: 10.1016/j.cell.2008.07.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Berger WT, Ralph BP, Kaczocha M, Sun J, Balius TE, Rizzo RC, Haj-Dahmane S, Ojima I, Deutsch DG. PLoS One. 2012;7:e50968. doi: 10.1371/journal.pone.0050968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Lan H, Cheng CC, Kowalski TJ, Pang L, Shan L, Chuang CC, Jackson J, Rojas-Triana A, Bober L, Liu L, Voigt J, Orth P, Yang X, Shipps GW, Jr, Hedrick JA. J Lipid Res. 2011;52:646. doi: 10.1194/jlr.M012757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Krusinova E, Pelikanova T. Diabetes Res Clin Pract. 2008;82(Suppl 2):S127. doi: 10.1016/j.diabres.2008.09.023. [DOI] [PubMed] [Google Scholar]
- 121.Hertzel AV, Hellberg K, Reynolds JM, Kruse AC, Juhlmann BE, Smith AJ, Sanders MA, Ohlendorf DH, Suttles J, Bernlohr DA. J Med Chem. 2009;52:6024. doi: 10.1021/jm900720m. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Burak MF, Inouye KE, White A, Lee A, Tuncman G, Calay ES, Sekiya M, Tirosh A, Eguchi K, Birrane G, Lightwood D, Howells L, Odede G, Hailu H, West S, Garlish R, Neale H, Doyle C, Moore A, Hotamisligil GS. Sci Transl Med. 2015;7:319ra205. doi: 10.1126/scitranslmed.aac6336. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.