Abstract
Background
The genus Arachis is native to a region that includes Central Brazil and neighboring countries. Little is known about the genetic variability of the Brazilian cultivated peanut (Arachis hypogaea, genome AABB) germplasm collection at the DNA level. The understanding of the genetic diversity of cultivated and wild species of peanut (Arachis spp.) is essential to develop strategies of collection, conservation and use of the germplasm in variety development. The identity of the ancestor progenitor species of cultivated peanut has also been of great interest. Several species have been suggested as putative AA and BB genome donors to allotetraploid A. hypogaea. Microsatellite or SSR (Simple Sequence Repeat) markers are co-dominant, multiallelic, and highly polymorphic genetic markers, appropriate for genetic diversity studies. Microsatellite markers may also, to some extent, support phylogenetic inferences. Here we report the use of a set of microsatellite markers, including newly developed ones, for phylogenetic inferences and the analysis of genetic variation of accessions of A. hypogea and its wild relatives.
Results
A total of 67 new microsatellite markers (mainly TTG motif) were developed for Arachis. Only three of these markers, however, were polymorphic in cultivated peanut. These three new markers plus five other markers characterized previously were evaluated for number of alleles per locus and gene diversity using 60 accessions of A. hypogaea. Genetic relationships among these 60 accessions and a sample of 36 wild accessions representative of section Arachis were estimated using allelic variation observed in a selected set of 12 SSR markers. Results showed that the Brazilian peanut germplasm collection has considerable levels of genetic diversity detected by SSR markers. Similarity groups for A. hypogaea accessions were established, which is a useful criteria for selecting parental plants for crop improvement. Microsatellite marker transferability was up to 76% for species of the section Arachis, but only 45% for species from the other eight Arachis sections tested. A new marker (Ah-041) presented a 100% transferability and could be used to classify the peanut accessions in AA and non-AA genome carriers.
Conclusion
The level of polymorphism observed among accessions of A. hypogaea analyzed with newly developed microsatellite markers was low, corroborating the accumulated data which show that cultivated peanut presents a relatively reduced variation at the DNA level. A selected panel of SSR markers allowed the classification of A. hypogaea accessions into two major groups. The identification of similarity groups will be useful for the selection of parental plants to be used in breeding programs. Marker transferability is relatively high between accessions of section Arachis. The possibility of using microsatellite markers developed for one species in genetic evaluation of other species greatly reduces the cost of the analysis, since the development of microsatellite markers is still expensive and time consuming. The SSR markers developed in this study could be very useful for genetic analysis of wild species of Arachis, including comparative genome mapping, population genetic structure and phylogenetic inferences among species.
Background
The genus Arachis, family Fabaceae, is native to South America, probably from a region including Central Brazil and Paraguay [1]. There are 69 species described in the genus, assembled into nine sections, according to the morphology, geographic distribution and crossability [2]. Some of these species have been used for forage in South America, but the most important species is the cultivated peanut, Arachis hypogaea L. This crop is widely grown in more than 80 countries in the Americas, Asia and Africa [3] and is used for both human consumption and as a source of oil. World production is increasing and has reached 37 million tons (in the shell) and 5.8 million tons of oil, according to FAO estimates [4]. Peanut now ranks fifth among the world oil production crops.
Nearly all Arachis species are diploid, but cultivated peanut is an allotetraploid (genome AABB). It is a member of the section Arachis, which also includes about 25 diploid and one tetraploid wild species (A. monticola) [2]. Arachis hypogaea is classified based on the presence or absence of flowers on the main axis into two subspecies, hypogaea and fastigiata. These two subspecies were further classified into six botanical varieties based on morphology and growth habits [2]. Subspecies hypogaea was divided in two botanical varieties, hypogaea and hirsuta, while ssp. fastigiata in the varieties fastigiata, vulgaris, aequatoriana and peruviana. The identity of the progenitor species of cultivated peanut has been of great interest. Several species have been suggested as putative A and B genome donors [reviewed by [5-7]]. RFLP analysis that included 17 diploid species of the section Arachis and three A. hypogaea accessions suggested a single origin for domesticated peanut and ancestral species related to A. duranensis (A genome) and A. ipaënsis (B genome) as the most likely progenitors of A. hypogaea [5]. In situ hybridization analysis of six diploid species and one A. hypogaea accession, and RAPD and ISSR (Inter-simple sequence repeat) analyses of 13 A. hypogaea accessions and 15 wild species, however, suggested A. villosa (A genome) and A. ipaënsis (B genome) as the progenitors of cultivated peanut [6,7].
Cultivated peanut exhibits a considerable amount of variability for various morphological, physiological, and agronomic traits. However, little variation has been detected at the DNA level using techniques such as RAPDs (Random Amplified Polymorphic DNAs), AFLPs (Amplified Fragment Length Polymorphisms) and RFLPs (Restriction Fragment Length Polymorphisms) [5,8-15]. The low level of variation in cultivated peanut has been attributed to three causes or to combinations of them: (1) barriers to gene flow from related diploid species to domesticated peanut as a consequence of the polyploidization event [16]; (2) recent polyploidization, from one or a few individual(s) of each diploid parental species, combined with self-pollination [8]; or (3) use of few elite breeding lines and little exotic germplasm in breeding programs, resulting in a narrow genetic base [17,18]. Recently, some studies revealed DNA polymorphism in A. hypogaea using SSRs (Simple Sequence Repeats), AFLPs, RAPDs and ISSRs [7,19-23].
Little is known about the genetic variability of the Brazilian A. hypogaea germplasm collection at the DNA level. Knowledge of the genetic variation of peanut accessions is important for their efficient use in breeding programs, for studies on crop evolution, and for conservation purposes. Molecular marker analysis, joined to phenotypic evaluation, is a powerful tool for grouping of genotypes based on genetic distance data and for selection of progenitors that might constitute new breeding populations. In the context of ex situ conservation, it has been demonstrated that molecular markers are very useful for the management of germplasm collections [24]. SSRs are ideal tools for such studies as they are PCR-based markers, genetically defined, typically co-dominant, multiallelic, and uniformly dispersed throughout plant genomes. SSRs have been used in plants for many genetic applications, including the assessment of genetic variability in germplasm collections or pedigree reconstruction [reviewed by [25]]. In Arachis, SSR markers have been recently developed and proved to be useful for accession discrimination and assessment of genetic variation [19,22,23]. Moreover, since little genetic variability has been detected in cultivated peanut, the use of a polymorphic marker, such as SSRs, in addition to distinguishing closely related genotypes, should also be useful for phylogenetic studies, as demonstrated in other crops, such as wheat [26], melon [27], potato [28], and coffee [29].
The objectives of the present work were: (1) to develop new SSR markers for genetic analysis of cultivated peanut (A. hypogaea), (2) to employ a set of SSR markers to analyze the genetic variation among wild and cultivated peanut accessions of the Brazilian germplasm collection, and (3) to evaluate the cross-species transferability of SSR markers and their usefulness in phylogenetic studies of the genus Arachis.
Results and Discussion
SSR development and screening of markers
Microsatellite enriched genomic libraries of A. hypogaea were constructed in order to develop new SSR markers for the species. Digestion of the A. hypogaea genomic DNA with five different enzymes (AluI, MseI, RsaI, Sau3AI, and Tsp509I) revealed that Tsp509I produced the most adequate profile for library development, with fragments ranging from 200 to 800 bp in size. Four libraries were initially constructed based on trinucleotide repeat motifs (TTG, TGG, ATG, and ATC). Hybridization analysis revealed that the TTG/AAC repeats were more abundant in the peanut genome than the other tested motifs (TTG > TGG > ATG > ATC). This is in agreement with a survey of published DNA sequences in 54 plant species, where the TTG/AAC repeat was one of the most abundant SSRs [30]. Therefore, the TTG library was used for SSR development for A. hypogaea. Screening of 750 clones by anchored PCR showed that 162 had SSRs (21.6%) and these were sequenced. Out of the 162 positive clones sequenced, there were 91 unique (non-redundant) sequences (56.2% of the sequenced clones) but only 67 of them were suitable for primer designing (41.4% of the positive clones). The design of primers for the other 24 unique SSRs identified was not possible due to the occurrence of very short tandem repeats (< 5 units) or low GC content of the regions flanking the SSR. The anchored PCR screening prior to sequencing significantly improved the yield of useful clones. Few false-positive clones were obtained (less than 10% of the sequenced clones). The percentage of primers designed, in relation to the number of clones sequenced (41.4%), indicated that the method used was relatively efficient for the discovery and development of SSR markers for peanut.
The 67 SSR markers (see Additional File 1) were screened for polymorphism on seven samples, including five varieties of A. hypogaea, one accession of A. ipaënsis and one accession of A. duranensis. Of these, 62 markers (92.5%) generated clearly interpretable PCR products, but only three markers (Ah-041, Ah-193 and Ah-558) were polymorphic in cultivated peanut (Table 1). Four other markers (Ah-075, Ah-229, Ah-522 and Ah-657) showed to be invariant for the five A. hypogaea accessions, but were polymorphic in A. ipaënsis and A. duranensis accessions. Since ancestral species related to these two species are considered potential progenitors of the AABB genome, these seven markers were included in the analysis of the genetic relationships between accessions of the Brazilian germplasm collection (see below). The other 55 markers were not polymorphic in A. hypogea accessions and were not considered for further analysis. They have been useful, however, for studies with diploid wild species of peanut currently under development (data not shown).
Table 1.
SSR loci characterization Primer pairs, repeat motifs, range of fragment sizes, total number of alleles (A), and gene diversity (h) estimates based on the analysis of 60 Arachis hypogaea accessions for the eight polymorphic SSR markers.
| Identification | Motif | Size range (bp) | A** | h |
| Ah-041* | (AAC)21 (CAA)10 | 280–292 | 2 | 0.468 |
| Ah-193* | (AAC)16 (GA)24 | 444–462 | 8 | 0.718 |
| Ah-558* | (AAC)9 | 226–244 | 6 | 0.520 |
| Ah4-04 | (GA)19 | 84–108 | 7 (6) | 0.702 |
| Ah4-20 | (GA)19 | 201–215 | 4 (3) | 0.646 |
| Ah4-24 | (ATA)17 | 406–418 | 5 (5) | 0.761 |
| Ah4-26 | (CT)25 imperfect | 156–213 | 15 (8) | 0.743 |
| Lec-1 | (AT)18 | 212–294 | 27 (14) | 0.929 |
**Enclosed in parenthesis, the number of fragments detected by Hopkins et al. [19].
SSR marker characterization
Marker loci duplication
Four (Ah-193, Ah-229, Ah-522, and Ah-657) of the seven newly developed markers produced single fragments both in the tetraploid accessions and in the diploid species belonging to the Arachis section (see Additional File 2), what indicates that each of them amplified alleles at single loci. Two markers (Ah-075 and Ah-558) produced one or two fragments in the diploid species and three (Ah-075) or four (Ah-558) fragments in the tetraploid accessions. These results are consistent with allele amplification on duplicated loci. The presence of the three alleles (150/144/138 bp) amplified by marker Ah-075 in the tetraploid accessions was confirmed by the identification of diploid species homozygous for each of these three alleles (for example, accessions K 30006, Sv 3806, V 6389, V 10309) or heterozygous for the three pairs of alleles (for example, accessions K 30076, K 30097, V 14167). Similar results were observed for marker Ah-558, but one (244 bp) of the six alleles (244/241/235/232/229/226 bp) detected in the tetraploid accessions was not found in any of the tested diploid wild species. The remaining marker (Ah-041) amplified one (292 bp) or two of three alleles (300/292/280 bp) in accessions of cultivated peanut and in the diploid species (Figure 1). This indicates that either the individuals tested are highly heterozygous for this locus or this marker locus is duplicated in the cultivated peanut genome. Since A. hypogaea is an allotetraploid and preferentially an autogamous species, with only 2.5% outcrossing [31], the latter hypothesis is more probable. Five other markers previously developed [19] were also analyzed for marker locus duplication, and two of them (Ah4-04 and Ah4-24) seem to represent a single locus in the genome. Therefore, five out of 12 markers (41,7%) probably amplified duplicated loci in most of the cultivated peanut accessions. This proportion, although not representative, indicates that locus duplication in peanut might be higher than found for other polyploid species, such as wheat [32], apple [33], cassava [34], and sweet potato [35]. The inclusion of hybrids and parental diploid plants in the study contributed to a better understanding of the genetic basis of these marker loci. Markers that amplified three or four alleles in tetraploid accessions amplified consistently only two alleles in the interspecific hybrids. In depth analyses of peanut segregant populations should elucidate the inheritance of these SSR loci.
Figure 1.
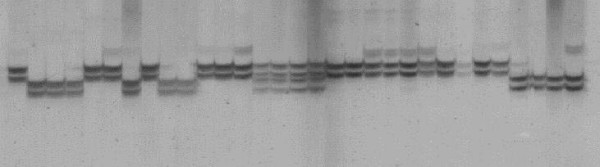
Microsatellite polymorphism (marker Ah-041) in Arachis species visualized in silver-stained denaturing polyacrilamide gel.Samples are: (1) A. duranensis-V14167; (2) A. ipaënsis; (3) A. magna-V13760; (4) A. batizocoi-K9484; (5) A. cardenasii; (6) A. stenosperma-V10229; (7) A. magna-K30097; (8) A. helodes; (9) A. hoehnei; (10) A. batizocoi-K9484m; (11) A. villosa; (12) A. microsperma; (13) A. simpsonii; (14) A. monticola; (15) A. hypogaea fastigiata fastigiata (16) A. hypogaea fastigiata vulgaris; (17) A. hypogaea fastigiata peruviana; (18) A. hypogaea hypogaea hypogaea; (19) A. hypogaea hypogaea hypogaea, Xingu type;(20) A. stenosperma-V10309; (21–23) Hybrids K7988 × V10309; (24) A. duranensis-K7988; (25) A. diogoi; (26) A. duranensis-G10038; (27) A. kempf-mercadoi; (28) Arachis sp. 6389; (29) A. cruziana; (30) A. praecox; (31) A. valida.
Marker allelic diversity
Allelic diversity for a sample of 60 A. hypogaea accessions (Additional File 2) was estimated for the three newly developed polymorphic markers (Ah-041, Ah-193, Ah-558) and five other SSR markers (Ah4-04, Ah4-20, Ah4-24, Ah4-26 e Lec-1), previously developed for cultivated peanut [19] (Table 1). Marker Ah6-125 [19] was not included in the analysis since it was found to be invariant for the 60 accessions of cultivated peanut. The number of alleles detected ranged from 2 to 27 at each of the eight polymorphic loci, with an average of 8.4 alleles per locus. Gene diversity values ranged from 0.468 to 0.929, with an average value of 0.683. For polyploid organisms, gene diversity is an estimate of the probability that two randomly chosen genes from a population are different and it is not related to the expected proportion of heterozygous, as for diploid organisms in Hardy-Weinberg equilibrium [36]. The values obtained in the present study are probably overestimated, since some of the markers amplified alleles on duplicated loci. Thus, they cannot be compared to those obtained for diploid species, but they are similar to values found for other polyploid species such as cassava [34], potato [37] and wheat [38]. Moreover, average h values estimated by RFLP markers were considerably lower for A. hypogaea [10]. These results demonstrate the usefulness of microsatellite markers for genetic diversity analysis of cultivated peanut.
Marker Ah-041 and genome specific alleles
One of the markers developed in the present study (Ah-041) presented interesting banding patterns (Figure 1) on AA and non-AA genomes of diploid wild species. Genome AA is characterized by the presence of a pair of chromosomes (called AA) that are considerably smaller than the other chromosomes [39], while genome BB lacks this small chromosome pair. All AA genome species, previously characterized by cytological analysis, showed a 292 bp allele when genotyped at the Ah-041 locus, while all BB genome species and A. glandulifera (DD genome) showed a 280 bp allele. Arachis hypogaea accessions showed both alleles (292/280) or they had only the 292 bp allele. Arachis monticola, the other tetraploid species of section Arachis, had also both 292 and 280 bp alleles. Accessions of five AA genome species (A. stenosperma, A. villosa, A. microsperma, A. simpsonii and A. kempff-mercadoi), one BB genome species (A. valida) and A. glandulifera showed a 300 bp band. Marker Ah-041 allowed allele detection on all the 114 accessions tested, with a 100% transferability in the genus. Species from the sections Erectoides, Extranervosae, and Triseminatae had different alleles (277 and 274 bp). Allele 280 bp was also found in species from the sections Trierectoides, Heteranthae, Procumbentes, and Rhizomatosae. Only section Arachis species showed the allele 292 bp (AA genome). Although allelic segregation and genome mapping of this marker locus has not yet been examined in controlled crosses, our results suggest that the 292 bp allele is specific to AA genome species. If so, this marker may be valuable for genetic studies of peanut, including phylogenetic inferences in the genus Arachis and studies about genome origin (see below). It should be noticed that the absolute transferability of marker Ah-041 to all Arachis species suggests that this SSR sequence is positioned in or near coding regions. However, search on the National Center for Biotechnology Information (NCBI) database http://www.ncbi.nlm.nih.gov/BLAST/ did not find any significantly complete or partially homologous sequence.
Transferability of SSR markers
Twelve SSR markers developed for Arachis hypogaea (Ah-041, Ah-193, Ah-558, Ah-075, Ah-229, Ah-522, Ah-657, Ah4-04, Ah4-20, Ah4-24, Ah4-26 and Lec-1) were tested on 54 accessions of wild species of Arachis, representing the nine sections of the genus. The transferability of the twelve markers was up to 76% for species of the section Arachis, but ranged from 23% (Triseminatae) to 62% (Procumbentes), with an average of 45%, for species of the other eight Arachis sections (see Additional File 3). Transferability of SSR markers between related species is a consequence of the homology of flanking regions of the microsatellites and the size of the region between the primer pair amenable to amplification by PCR. Other studies have demonstrated the conservation of SSR sequences in plants, as reviewed by Gupta and Varshney [25]. The possibility of using microsatellite markers developed for one species in genetic evaluation of other species greatly reduces the cost of the analysis, since the development of microsatellite markers is still expensive and time consuming. The SSR markers developed in this study could be very useful for genetic analysis of wild species of Arachis, including comparative genome mapping, population genetic structure and phylogenetic inferences among species.
Relationships between Arachis spp. accessions
Genetic similarities among accessions were estimated by shared allele distance in pairwise comparisons of 60 A. hypogaea accessions and 36 other accessions of wild species belonging to section Arachis (accessions 1 to 96 – Additional File 2), using the same set of 12 markers described above. Among the 60 A. hypogaea accessions, the average genetic distance was 0.336. Some accessions, such as Tatu, Tatu2 e Pd2622, shared all the alleles detected with the 12 SSR markers.
A dendrogram based on the neighbor joining method was constructed for the 96 accessions of the section Arachis (Figure 2). Considering initially the A. hypogaea accessions, two main clusters were evident. Group I contained all 32 fastigiata/fastigiata, all four fastigiata/vulgaris accessions, the only hypogaea/hirsuta accession (Mf 1538) and one accession not identified to the subspecies level (Wi 632). The two fastigiata/aequatoriana accessions included in the study (Mf 1678 and Mf1640) were also clustered in this group. The two fastigiata/peruviana accessions (Mf 1560 e Sv 429) formed a separate subgroup within Group I. Another subgroup was formed by a fastigiata/aequatoriana accession (Mf 1640), two variant forms of hypogaea/hypogaea accessions (Nambiquarae, accession As 436 and Malhado, accession V 12577) and V 12549, identified as "ssp hypogaea Xingu type", because of its morphological differences from the Virginia type. All 45 accessions of Group I had the alleles 292 bp and 280 bp at the Ah-041 locus. The other group (Group II) contained solely the 15 Virginia type (hypogaea/hypogaea) accessions examined in the study. The accessions of this group had only the 292 bp allele (Figure 1). Polymorphism at the Ah-041 locus could be caused by mutation at the primer binding site in the Group II accessions, preventing the detection of the 280 bp PCR product, or by an insertion/deletion event. If the Ah-041 locus is duplicated in allotetraploid peanut, these results could indicate that cultivated peanut did not have a single origin since Group I accessions could be derived from hybridization of AA and non-AA ancestral diploid species followed by genome duplication, while Group II accessions could be derived from straight genome duplication of an AA diploid species. Multiple origin hypotheses, based on cytological, crossing and isozyme studies, have been previously suggested to explain the cultivated peanut origin [40-42].
Figure 2.
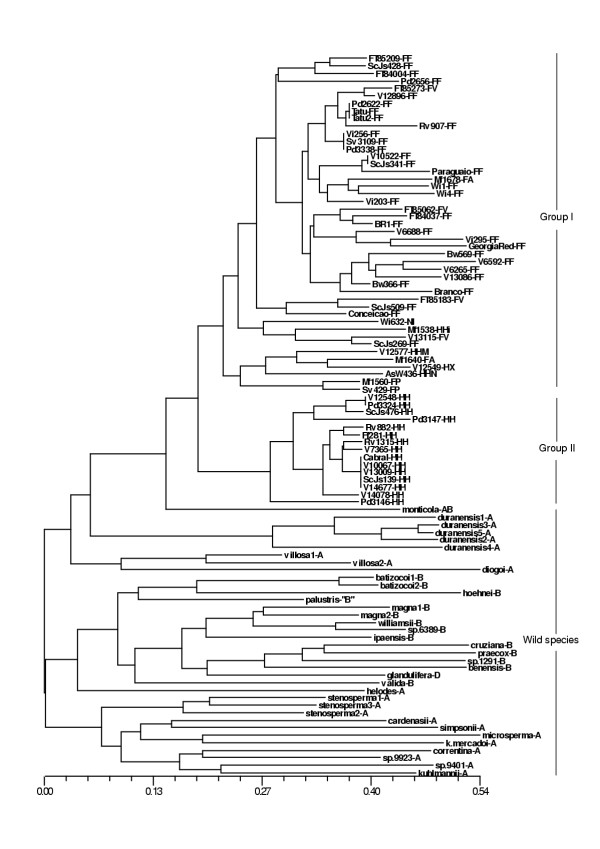
Dendrogram based on allele sharing genetic distances of 60 A. hypogaea and 36 Arachis wild species generated by the neighbor joining method.The letters, after each A. hypogaea accession number, refer to the subspecies, varieties, and types: FF-fastigiata/fastigiata; FV-fastigiata/vulgaris; FA-fastigiata/aequatoriana; FP-fastigiata/peruviana; HH-hypogaea/hypogaea; HHi-hypogaea/hirsuta; HHN-hypogaea/hypogaea Nambiquarae type; HHM-hypogaea/hypogaea "Malhado" type; HX-hypogaea Xingu type. Ni, corresponds to a non-identified accession to the subspecies level. The letters after the wild species accession names refer to the genome type (A or B), according to the patterns obtained with marker Ah-041. The D genome of A. glandulifera was defined by Stalker, 1991.
The taxonomic status of A. hypogaea ssp. fastigiata var. peruviana is controversial. Studies based on AFLP [21] as well as RAPD and ISSR [7] markers have shown that accessions of this variety have greater similarity to ssp. hypogaea rather than ssp. fastigiata, to which they are currently thought to belong [2]. In the present study, the two fastigiata/peruviana accessions grouped with the other fastigiata accessions and not with the 15 hypogaea/hypogaea accessions clustered in Group II. The data, therefore, corroborates the current classification of this botanical variety. However, the classification of hypogaea/hirsuta and the three variant forms of hypogaea/hypogaea (Nambiquarae, Malhado and Xingu types) remains controversial, since these four accessions did not cluster in Group II.
The SSR markers used in the present study detected genetic variability in A. hypogaea accessions, despite the known narrow genetic base of cultivated peanut. In the two main clusters (Group I and Group II), some similarity groups, genetically distant from each other, were formed. These groups were not related to collection sites, seed color, or any other known trait. However, the establishment of similarity groups, jointly with morphologic and agronomic data, can be used in the selection and crossing of superior plants, allowing for a more efficient use of the existing genetic variability conserved in the collection.
Considering the wild species, the analysis showed that, in general, the species containing the same genome clustered together. However, accessions of A. duranensis clustered with A. hypogaea and A. monticola accessions (Figure 2). This result corroborates the hypothesis that an ancestral species related to A. duranensis or A. duranensis itself could be the donor of the AA genome to A. hypogaea. In the group composed of BB genome species there is a cluster of four species (A. ipaënsis, A. magna, A. williamsii and Arachis sp. 6389). It has recently been noticed that plants of these four species are morphologically very similar and produce fertile hybrids. A revision of their taxonomic status is currently under evaluation by JFM Valls and CE Simpson.
The genetic distance analysis based on SSR marker polymorphism supports previous estimates of high genetic similarity between A. monticola accessions and A. hypogaea accessions (Figure 2). Only one (Ah-522) out of the 12 SSR loci included in the present study showed alleles detected in A. hypogaea not observed in A. monticola. Other studies with molecular data have also shown the little differentiation between these two taxa [8,9,11,42]. Moreover, hybrids between A. monticola and A. hypogaea are fertile [2]. These facts have led some researchers to suggest that A. monticola and A. hypogaea should not be considered as separate species [11]. Our results, based on SSR markers and only one accession of A. monticola, are certainly insufficient to define the taxonomic status of A. monticola. However, they indicated that this species could be directly related to the allotetraploid ancestral progenitor of A. hypogaea, as suggested by several studies [6,7,43].
Conclusions
The present study showed the utility of microsatellite markers for the detection of polymorphisms among cultivated peanut accessions and for the genetic relationship analysis between A. hypogaea accessions and wild species of section Arachis. New SSR markers were developed for genetic analysis of peanut, but the percentage of markers that detected polymorphism among accessions of A. hypogaea was low, corroborating the notion that cultivated peanut presents a relatively reduced variation at the DNA level. However, a set of informative SSR markers detected considerable levels of genetic variability in the Brazilian Peanut Germplasm Collection. Based on this data, cultivated peanut was classified into two major groups, one composed of solely the Virginia type (hypogaea/hypogaea) accessions and another defined by fastigiata/fastigiata, fastigiata/vulgaris, hypogaea/hirsuta, fastigiata/aequatoriana, fastigiata/peruviana and two variant forms of hypogaea/hypogaea accessions. The identification of similarity groups could be useful for the selection of parental plants to be used in the breeding programs. Marker transferability was up to 76% for species of the section Arachis, but ranged from 23% (Triseminatae) to 62% (Procumbentes), with an average of 45%, for species of the other eight Arachis sections. The possibility of using microsatellite markers developed for one species in genetic evaluation of other species greatly reduces the cost of the analysis, since the development of microsatellite markers is still expensive and time consuming. The SSR markers developed in this study could be very useful for genetic analysis of wild species of Arachis, including comparative genome mapping, population genetic structure and phylogenetic inferences among species. A marker (Ah-041) was identified that allows the discrimination of AA from non-AA genome accessions of peanut.
Methods
Plant material
A single A. hypogaea plant was used for DNA library construction (accession UF 91108). Five A. hypogaea samples (accessions 3274, 3294, 3310, 3324, and UF 91108), one A. ipaënsis (accession 3309) and one A. duranensis (accession 4454) were used to test the effectiveness of SSR markers to detect polymorphism at the DNA level. These plants were obtained from USDA-ARS Plant Genetic Resources Conservation Unit (Griffin, GA) peanut collection.
A total of 114 accessions of wild and cultivated peanut were included in the study (Additional File 2). Ninety-six accessions belonging to section Arachis were used for genetic diversity analysis. Of these, 60 are A. hypogaea accessions, 47 of which collected in Brazil, representing both A. hypogaea ssp. hypogaea and ssp. fastigiata, and their six varieties. The other 36 accessions belong to 27 wild species, including representatives of four new taxa that are being described by JFM Valls and CE Simpson. The wild species include tetraploid A. monticola (AABB), 13 diploid (2n = 20) species showing the small chromosome pair [44] which are associated to the AA genome of A. hypogaea, the diploid A. glandulifera, for which a DD genome has been suggested [45], two diploids with 2n = 18 [46] and ten species with 20 chromosomes lacking the small chromosome pair typical of the AA genome, which are classified as BB genome species (Additional File 2). Other 18 accessions of wild species belonging to eight Arachis sections were included in the analysis to test the transferability of SSR markers. Three hybrid plants of the A. stenosperma (V 10309) × A. duranensis (K 7988) cross and one hybrid plant of the A. appressipila (G 10002) × A. repens (Nc 1579) cross were also used to verify the heritability of alleles in interspecific crosses. The 114 accessions and the four hybrid plants were obtained from the Brazilian Peanut Germplasm Collection, maintained at Embrapa Genetic Resources and Biotechnology – CENARGEN (Brasília-DF, Brazil). All plants were grown from seed or cuttings (sections Rhizomatosae and Caulorrhizae) under greenhouse conditions at CENARGEN prior to DNA extraction.
DNA extraction
Total genomic DNA used for library construction was extracted and purified on cesium chloride gradients following standard protocols [47]. For the other samples, total genomic DNA was extracted according to published protocol [48], with some modifications: Proteinase K (20 mg/ml) was added to the extraction buffer and an additional step of polysaccharide precipitation with 2 M NaCl was included.
Library construction
Total genomic DNA libraries enriched for trinucleotide repeats were constructed following a standard protocol with minor modifications [49]. Peanut total genomic DNA was digested with five different enzymes (AluI, MseI, RsaI, Sau3AI, and Tsp509I) to identify which one produced the most adequate fragment profile for library development. Fragments ranging in size from 200 to 800 bp were transferred to S&S NA 45 DEAE cellulose membranes, after electrophoresis in 1.5% low-melting agarose gel. DNA was resuspended in TE buffer and ligated to Tsp509 I adapters. Four genomic libraries enriched for trinucleotide repeats were constructed. Fragments were selected by hybridization with biotinylated trinucleotide repeats ((TTG)6, (TGG)6, (ATG)6, (ATC)6) and recovered by magnetic beads linked to streptavidine. Enriched fractions were amplified by PCR, using a primer complementary to the adapters. Amplification products were purified using Wizard PCR Preps DNA purification system (Promega, Madison, WI), cloned into λ ZAP II phagemid vector (Stratagene) and then transformed into E. coli strain XL1-Blue. Transformed cells were grown on LB-Amp plates (50 μg/ml ampicillin) at 37°C, until plaques were between 0.5 and 1.0 mm in diameter. Individual clones were picked up from the plates and phage were eluted in SM buffer (500 μl) with a drop of chloroform. Clone DNAs were then amplified by PCR using primers complementary to the vector DNA flanking the insert (T3 or SK and T7 or KS primers, Stratagene, CA), and electrophoresed in 1.5% agarose gels in 0.5X TBE buffer to determine insert sizes. To determine the position of the SSRs relative to the vector cloning site, an anchored PCR strategy was performed for each positive clone [49], using T3 (or SK) and T7 (or KS) primers separately plus another primer complementary to the target SSR.
DNA sequencing and primer design
Clones containing repeats that were not directly adjacent to the cloning site were amplified, treated with ExoI and shrimp alkaline phosphatase enzymes, and PCR products were sequenced on an ABI 377 automated DNA sequencer (Applied Biosystems, CA, USA), using fluorescent dye-terminator chemistry. Redundant sequences were identified using the software Sequencher (Gene Codes Corporation). Primers complementary to unique DNA sequences flanking the SSRs were designed using the computer program Primer 3 (Whitehead Institute of Biomedical Research – http://www-genome.wi.mit.edu/cgi-bin/primer/primer3_www.cgi). Primer design parameters were as follows: (i) Tm ranged from 60°C to 65°C; (ii) 1°C difference in Tm between primer pairs; (iii) GC content ranging from 45% to 55%, (iv) absence of complementarity between primers, (v) default values for the other parameters.
Primer screening and PCR amplification of SSR loci
Primer pairs were initially screened for polymorphisms in a set of five cultivated and two wild peanut accessions (3274, 3294, 3310, 3324, UF 91108, 3309, 4454). PCRs were performed in 13 μl volumes, containing 1X PCR buffer (10 mM Tris-HCl pH8.3, 50 mM KCl, 1.5 mM MgCl2), 0.2 mM of each dNTP, 1 unit Taq DNA polymerase, DMSO 50% (1.3 μl), 5 pmol of each primer and 10 ng of genomic DNA. Amplifications were performed using either a 9600 System (Applied Biosystems, CA, USA) or a PTC-100 (MJ Research, MA, USA) thermal cycler, with the following conditions: 96°C for 2 min (1 cycle), 94°C for 1 min, 55–66°C for 1 min, 72°C for 1 min (30 cycles); and 72°C for 7 min (1 cycle). The annealing temperature was optimized for each primer pair to produce clear DNA band amplification, without spurious fragments. PCR products were resolved on 3.5% Metaphor agarose (FMC Bioproducts, ME, USA) gels stained with ethidium bromide. For genotype determination, the amplified products were separated on 4% denaturing polyacrilamide gels stained with silver nitrate [50]. Fragment sizes were estimated by comparison with a 10-bp DNA ladder standard (Gibco/BRL, MD, USA).
Data analysis and transferability of SSR loci
Eight selected SSR markers were characterized for number of alleles per locus and gene diversity [51] using 60 accessions of Arachis hypogaea (accessions 1 to 60 – Additional File 2). The markers included three developed in the present study and five other SSR markers developed for A. hypogaea [19]. Gene diversity (h) at a marker locus was estimated according to the formula: h = 1 - Σ(pi)2, where pi2 is the frequency of the ith allele at this locus [51], using the GDA software [52]. For estimates of genetic distance, 60 accessions of cultivated peanut and 36 accessions from 27 wild species of section Arachis were analyzed (accessions 61 to 96 – Additional File 2). A total of 12 markers were used in this analysis (Ah-041, Ah-193, Ah-558, Ah-075, Ah-229, Ah-522, Ah-657, Ah4-04, Ah4-20, Ah4-24, Ah4-26 e Lec-1). Genetic distances among wild and cultivated peanut accessions based on microsatellite marker polymorphism were estimated by shared allele distance in pairwise comparisons. The estimates were based on the sum of the proportion of common alleles between two peanut accessions examined across loci (PS) divided by twice the number of tested loci [53,54]. Genetic distances were obtained by the parameter [-ln (PS)] using the Genetic Distance Calculator [55]. The diagonal matrix was then submitted to cluster analysis using the neighbor joining method and a genetic distance dendrogram built using the software NTSYS 2.1 [56]. Transferability of the 13 SSR markers (the 12 described above plus marker Ah6-125) was tested using the 36 accessions of section Arachis (accessions 61 to 96 – Additional File 2) and another 18 wild species accessions belonging to the remaining Arachis sections (accessions 97 to 114 – Additional File 2). PCR amplification followed the same protocol used for A. hypogaea. PCR products were visualized on silver-stained 4% denaturing polyacrilamide gels.
Authors' contributions
MM participated in the SSR markers development, carried out the analysis of the Arachis accessions and drafted the manuscript. MH participated in the development of the SSR markers. SM and SK coordinated the development of the SSR markers. JV participated in the conception of the project and provided the germplasm. MF participated in the conception of the project, in the data analysis and reviewed the manuscript. All authors read and approved the manuscript.
Supplementary Material
List of the 67 microsatellite markers for genetic analysis of Arachis spp. Marker sequences, repeat motifs, and annealing temperature (Ta) for the 67 newly developed SSR markers
Arachis spp. accessions analyzed in this study. Numbers in parenthesis, after the specific names, correspond to numbers included in the dendrogram (Figure 2). *Genome types of Arachis hypogaea accessions were defined according to the genotypes obtained with marker Ah-041.
Transferability of SSR markers developed for A. hypogaea across 45 wild species of Arachis. This file contains a table that lists the markers that amplified visible products for each of the 49 accessions of the Arachis wild species included in the study.
Acknowledgments
Acknowledgements
The authors thank Dr. R. E. Dean for computational assistance and L. Zhang and T. M. Jenkins for their technical laboratory assistance. We also thank M.R. Bertozo for generously providing the DNA samples of some wild species of Arachis. Development of SSR marker research supported by grants from the USDA. Characterization of SSR markers supported by Embrapa Recursos Genéticos e Biotecnologia, Brazil.
Contributor Information
Marcio de Carvalho Moretzsohn, Email: marciocm@cenargen.embrapa.br.
Mark S Hopkins, Email: Jshceh@aol.com.
Sharon E Mitchell, Email: sem30@cornell.edu.
Stephen Kresovich, Email: sk20@cornell.edu.
Jose Francisco Montenegro Valls, Email: valls@cenargen.embrapa.br.
Marcio Elias Ferreira, Email: ferreira@cenargen.embrapa.br.
References
- Gregory WC, Krapovickas A, Gregory MP. Structure, variation, evolution, and classification in Arachis. In: Summerfield RJ, Bunting AH, editor. Advances in Legume Science. Kew, England: Royal Botanical Gardens; 1980. pp. 469–481. [Google Scholar]
- Krapovickas A, Gregory WC. Taxonomía del género Arachis (Leguminosae) Bonplandia. 1994;8:1–186. [Google Scholar]
- Singh U, Singh B. Tropical grain legumes as important human foods. Econ Bot. 1992;46:310–321. [Google Scholar]
- FAO (2004) http://faostat.fao.org/faostat/collections?subset=agriculture
- Kochert G, Stalker HT, Gimenes M, Galgaro L, Lopes CR, Moore K. RFLP and cytogenetic evidence on the origin and evolution of allotetraploid domesticated peanut, Arachis hypogaea (Leguminosae) Am J Bot. 1996;83:1282–1291. [Google Scholar]
- Raina SN, Mukai Y. Genomic in situ hybridization in Arachis (Fabaceae) identifies the diploid wild progenitors of cultivated (A. hypogaea) and related wild (A. monticola) peanut species. Pl Syst Evol. 1999;214:251–262. [Google Scholar]
- Raina SN, Rani V, Kojima T, Ogihara Y, Singh KP, Devarumath RM. RAPD and ISSR fingerprints as useful genetic markers for analysis of genetic diversity, varietal identification, and phylogenetic relationships in peanut (Arachis hypogaea) cultivars and wild species. Genome. 2001;44:763–772. doi: 10.1139/gen-44-5-763. [DOI] [PubMed] [Google Scholar]
- Halward TM, Stalker HT, Larue EA, Kochert G. Genetic variation detectable with molecular markers among unadapted germ-plasm resources of cultivated peanut and related wild species. Genome. 1991;34:1013–1020. [Google Scholar]
- Kochert G, Halward T, Branch WD, Simpson CE. RFLP variability in peanut (Arachis hypogaea) cultivars and wild species. Theor Appl Genet. 1991;81:565–570. doi: 10.1007/BF00226719. [DOI] [PubMed] [Google Scholar]
- Paik-Ro OG, Smith RL, Knauft DA. Restriction fragment length polymorphism evaluation of six peanut species within the Arachis section. Theor Appl Genet. 1992;84:201–208. doi: 10.1007/BF00224001. [DOI] [PubMed] [Google Scholar]
- Hilu KW, Stalker HT. Genetic relationships between peanut and wild species of Arachis sect. Arachis (Fabaceae): evidence from RAPDs. Pl Syst Evol. 1995;198:167–178. [Google Scholar]
- He G, Prakash CS. Identification of polymorphic DNA markers in cultivated peanut (Arachis hypogaea L.) Euphytica. 1997;97:143–149. doi: 10.1023/A:1002949813052. [DOI] [Google Scholar]
- Subramanian V, Gurtu S, Nageswara Rao RC, Nigam SN. Identification of DNA polymorphism in cultivated groundnut using random amplified polymorphic DNA (RAPD) assay. Genome. 2000;43:656–660. doi: 10.1139/gen-43-4-656. [DOI] [PubMed] [Google Scholar]
- Gimenes MA, Lopes CR, Valls JFM. Genetic relationships among Arachis species based on AFLP. Genet Mol Biol. 2002;25:349–353. [Google Scholar]
- Herselman L. Genetic variation among Southern African cultivated peanut (A. hypogaea L.) genotypes as revealed by AFLP analysis. Euphytica. 2003;133:319–327. doi: 10.1023/A:1025769212187. [DOI] [Google Scholar]
- Young ND, Weeden NF, Kochert G. Genome mapping in legumes (Family Fabaceae) In: Paterson AH, editor. Genome Mapping in Plants. Austin, TX: Landes Biomedical Press; 1996. pp. 212–227. [Google Scholar]
- Knauft DA, Gorbet DW. Genetic diversity among peanut cultivars. Crop Sci. 1989;29:1417–1422. [Google Scholar]
- Isleib TG, Wynne JC. Use of plant introductions in peanut improvement. In: Shands HL, editor. Use of Plant Introductions in Cultivar Development. Vol. 2. Madison: Crop Science Society of America; 1992. pp. 75–116. [Google Scholar]
- Hopkins MS, Casa AM, Wang T, Mitchell SE, Dean R, Kochert GD, Kresovich S. Discovery and Characterization of Polymorphic Simple Sequence Repeats (SSRs) in Peanut. Crop Sci. 1999;39:1243–1247. [Google Scholar]
- Dwivedi SL, Gurtu S, Chandra S, Yuejin W, Nigam SN. Assessment of genetic diversity among selected groundnut germplasm. I: RAPD analysis. Plant Breeding. 2001;120:345–349. doi: 10.1046/j.1439-0523.2001.00613.x. [DOI] [Google Scholar]
- He G, Prakash CS. Evaluation of genetic relationships among botanical varieties of cultivated peanut (Arachis hypogaea L.) using AFLP markers. Genet Resour Crop Evol. 2001;48:347–352. doi: 10.1023/A:1012019600318. [DOI] [Google Scholar]
- He G, Meng R, Newman M, Gao G, Pittman RN, Prakash CS. Microsatellites as DNA markers in cultivated peanut (A. hypogaea L.) BMC Plant Biology. 2003;3:3. doi: 10.1186/1471-2229-3-3. http://www.biomedcentral.com/1471-2229/3/3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferguson ME, Burow MD, Schulze SR, Bramel PJ, Paterson AH, Kresovich S, Mitchell S. Microsatellite identification and characterization in peanut (A. hypogaea L.) Theor Appl Genet. 2004;108:1064–1070. doi: 10.1007/s00122-003-1535-2. [DOI] [PubMed] [Google Scholar]
- Westman AL, Kresovich S. Use of molecular markers techniques for description of plant genetic variation. In: Callow JA, Ford-Lloyd BV, Newbury HJ, editor. Plant Genetic Resources: Conservation and Use. Wallingford, UK: CAB International; 1997. pp. 9–48. [Google Scholar]
- Gupta PK, Varshney RK. The development and use of microsatellite markers for genetic analysis and plant breeding with emphasis on bread wheat. Euphytica. 2000;113:163–185. doi: 10.1023/A:1003910819967. [DOI] [Google Scholar]
- Lelley T, Stachel M, Grausgruber H, Vollmann J. Analysis of relationships between Aegilops tauschii and the D genome of wheat utilizing microsatellites. Genome. 2000;43:661–668. doi: 10.1139/gen-43-4-661. [DOI] [PubMed] [Google Scholar]
- Danin-Poleg Y, Reis N, Tzuri G, Katzir N. Development and characterization of microsatellite markers in Cucumis . Theor Appl Genet. 2001;102:61–72. doi: 10.1007/s001220051618. [DOI] [Google Scholar]
- Ashkenazi V, Chani E, Lavi U, Levy D, Hillel J, Veilleux RE. Development of microsatellite markers in potato and their use in phylogenetic and fingerprinting analyses. Genome. 2001;44:50–62. doi: 10.1139/gen-44-1-50. [DOI] [PubMed] [Google Scholar]
- Anthony F, Combe MC, Astorga C, Bertrand B, Graziosi G, Lashermes P. The origin of cultivated Coffea arabica L. varieties revealed by AFLP and SSR markers. Theor Appl Genet. 2002;104:894–900. doi: 10.1007/s00122-001-0798-8. [DOI] [PubMed] [Google Scholar]
- Wang Z, Weber JL, Zhong G, Tanksley SD. Survey of plant short tandem DNA repeats. Theor Appl Genet. 1994;88:1–6. doi: 10.1007/BF00222386. [DOI] [PubMed] [Google Scholar]
- Norden AV. Peanuts-cultures and uses. Am Peanut Res Ed Assoc, Stillwater, Oklahoma; 1973. Breeding of the cultivated peanut (Arachis hypogaea L.) pp. 175–208. [Google Scholar]
- Bryan GJ, Collins AJ, Stephenson P, Orry A, Smith JB, Gale MD. Isolation and characterization of microsatellites from hexaploid bread wheat. Theor Appl Genet. 1997;94:557–563. doi: 10.1007/s001220050451. [DOI] [Google Scholar]
- Guilford P, Prakash S, Zhu JM, Rikkerink E, Gardiner S, Basset H, Foster R. Microsatellites in Malus × domestica (apple): abundance, polymorphism and cultivar identification. Theor Appl Genet. 1997;94:249–254. doi: 10.1007/s001220050407. [DOI] [Google Scholar]
- Chavarriaga-Aguirre P, Maya MM, Bonierbale MW, Kresovich S, Fregene MA, Tohme J, Kochert G. Microsatellites in cassava (Manihot esculenta Crantz): discovery, inheritance and variability. Theor Appl Genet. 1998;97:493–501. doi: 10.1007/s001220050922. [DOI] [Google Scholar]
- Buteler MI, Jarret RL, LaBonte DR. Sequence characterization of microsatellites in diploid and polyploid Ipomoea . Theor Appl Genet. 1999;99:123–132. doi: 10.1007/s001220051216. [DOI] [Google Scholar]
- Nei M. Molecular Evolutionary Genetics. New York: Columbia University Press; 1987. p. 512. [Google Scholar]
- Provan J, Powell W, Waugh R. Microsatellite analysis of relationships within cultivated potato (Solanum tuberosum) Theor Appl Genet. 1996;92:1078–1084. doi: 10.1007/s001220050233. [DOI] [PubMed] [Google Scholar]
- Prasad M, Varshney RK, Roy JK, Balyan HS, Gupta PK. The use of microsatellites for detecting DNA polymorphism, genotype identification and genetic diversity in wheat. Theor Appl Genet. 2000;100:584–592. [Google Scholar]
- Husted L. Cytological studies of the peanut Arachis. I. Chromosome number and morphology. Cytologia. 1933;5:109–117. [Google Scholar]
- Singh AK, Moss JP. Utilization of wild relatives in genetic improvement of Arachis hypogaea L. 2.Chromosome complements of species in the section Arachis . Theor Appl Genet. 1982;61:305–314. doi: 10.1007/BF00272846. [DOI] [PubMed] [Google Scholar]
- Singh AK. Putative genome donors of Arachis hypogaea (Fabaceae), evidence from crosses with synthetic amphidiploids. Plant Syst Evol. 1988;160:143–151. [Google Scholar]
- Lu J, Pickersgill B. Isozyme variation and species relationships in peanut and its wild relatives (Arachis L. – Leguminosae) Theor Appl Genet. 1993;85:550–560. doi: 10.1007/BF00220913. [DOI] [PubMed] [Google Scholar]
- Gregory WC, Gregory MP. Groundnut. In: Simmonds NW, editor. Evolution of Crop Plants. London: Longman; 1976. pp. 151–154. [Google Scholar]
- Fernandez A, Krapovickas A. Cromosomas y evolución en Arachis (Leguminosae) Bonplandia. 1994;8:187–220. [Google Scholar]
- Stalker HT. A new species in section Arachis of peanuts with D genome. Am J Bot. 1991;78:630–637. [Google Scholar]
- Lavia GI. Karyotypes of Arachis palustris and A.praecox (Section Arachis), two species with basic chromosme number x = 9. Cytologia. 1998;63:177–181. [Google Scholar]
- Reichardt M, Rogers S. Preparation of genomic DNA from plant tissue. In: Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K, editor. Current protocols in molecular biology. New York: John Wiley & Sons; 1997. [DOI] [PubMed] [Google Scholar]
- Grattapaglia D, Sederoff R. Genetic linkage maps of Eucalyptus grandis and Eucalyptus urophylla using a pseudo-testcross: Mapping strategy and RAPD markers. Genetics. 1994;137:1121–1137. doi: 10.1093/genetics/137.4.1121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rafalski JA, Vogel JM, Morgante M, Powel W, Andre C, Tingey SV. Generating and using DNA markers in plants. In: Birren B, Lai E, editor. Analysis of non-mammalian genomes – a practical guide. New York: Academic Press; 1996. pp. 75–134. [Google Scholar]
- Bassam BJ, Caetano-Anolles G, Gresshoff PM. Fast and sensitive silver staining of DNA in polyacrilamide gels. Analyt Biochem. 1991;196:80–83. doi: 10.1016/0003-2697(91)90120-i. [DOI] [PubMed] [Google Scholar]
- Nei M. Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA. 1973;70:3321–3323. doi: 10.1073/pnas.70.12.3321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis PO, Zaykin D. Genetic Data Analysis: Computer program for the analysis of allelic data. Version 1.0 (d16c) 2001. http://lewis.eeb.uconn.edu/lewishome/software.html Free program distributed by the authors over the Internet.
- Bowcock AM, Ruiz-Linhares A, Tomfohrde J, Minch E, Kidd JR, Cavalli-Sforza LL. High resolution of human evolutionary trees with polymorphic microsatellites. Nature. 1994;368:455–457. doi: 10.1038/368455a0. [DOI] [PubMed] [Google Scholar]
- Goldstein DB, Ruiz Linares A, Cavalli-Sforza LL, Feldman MW. Genetic absolute dating based on microsatellite and the origin of modern humans. Proc Natl Acad Sci USA. 1995;92:6723–6727. doi: 10.1073/pnas.92.15.6723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Genetic Distance Calculator http://www2.biology.ualberta.ca/jbrzusto/sharedst.php
- Rohlf FJ. NTSYS-PC: numerical taxonomy and multivariate analysis system – version 2.0. New York: Exeter Software; 1993. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
List of the 67 microsatellite markers for genetic analysis of Arachis spp. Marker sequences, repeat motifs, and annealing temperature (Ta) for the 67 newly developed SSR markers
Arachis spp. accessions analyzed in this study. Numbers in parenthesis, after the specific names, correspond to numbers included in the dendrogram (Figure 2). *Genome types of Arachis hypogaea accessions were defined according to the genotypes obtained with marker Ah-041.
Transferability of SSR markers developed for A. hypogaea across 45 wild species of Arachis. This file contains a table that lists the markers that amplified visible products for each of the 49 accessions of the Arachis wild species included in the study.


