Abstract
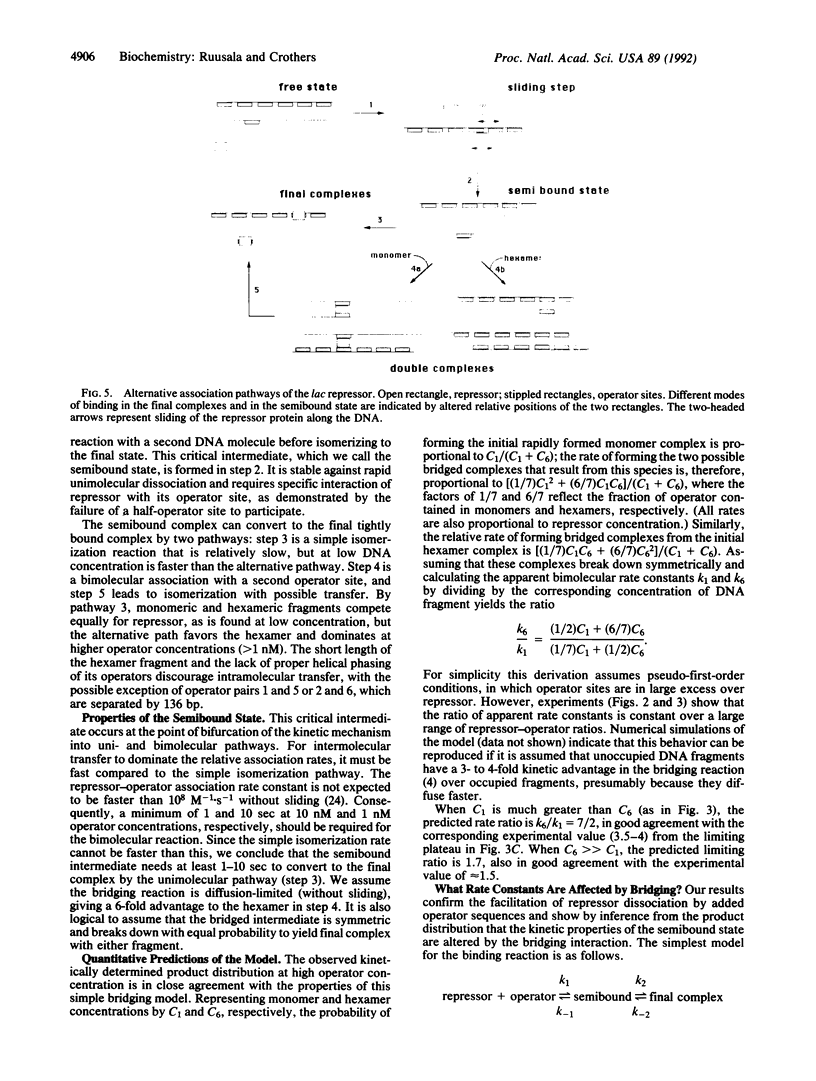
The lac repressor associates with its operator at a rate faster than conventional diffusion allows, either because of one-dimensional diffusion of a captured repressor along the DNA (sliding) or because the tetrameric repressor can be rapidly transferred between DNA sites distant from each other in the primary sequence (direct transfer). We report measurements of relative repressor-operator association rates at physiological operator concentrations. We assay for the ability of DNA targets of equal length (approximately 200 base pairs) containing one or six operator segments to compete for repressor; as the sliding range decreases, the hexameric operator fragment should react up to six times faster than the monomeric operator fragment. We find that the advantage enjoyed by the hexameric fragment varies from little to none at low operator concentration (depending on ionic strength) to more than 3-fold at high concentration. We ascribe this behavior to sliding at low operator concentration and to an increasing contribution of bimolecular direct transfer events as concentration increases. The observations require a "semibound" intermediate state of the protein at operator sites. This species can either undergo a relatively slow (tau greater than 1 sec) unimolecular isomerization to the final complex, or the isomerization can occur in a bridged complex with another operator site, accompanied by transfer to the second operator with probability of 0.5. Bridging alters one or more rate constants in the complex.
Full text
PDF
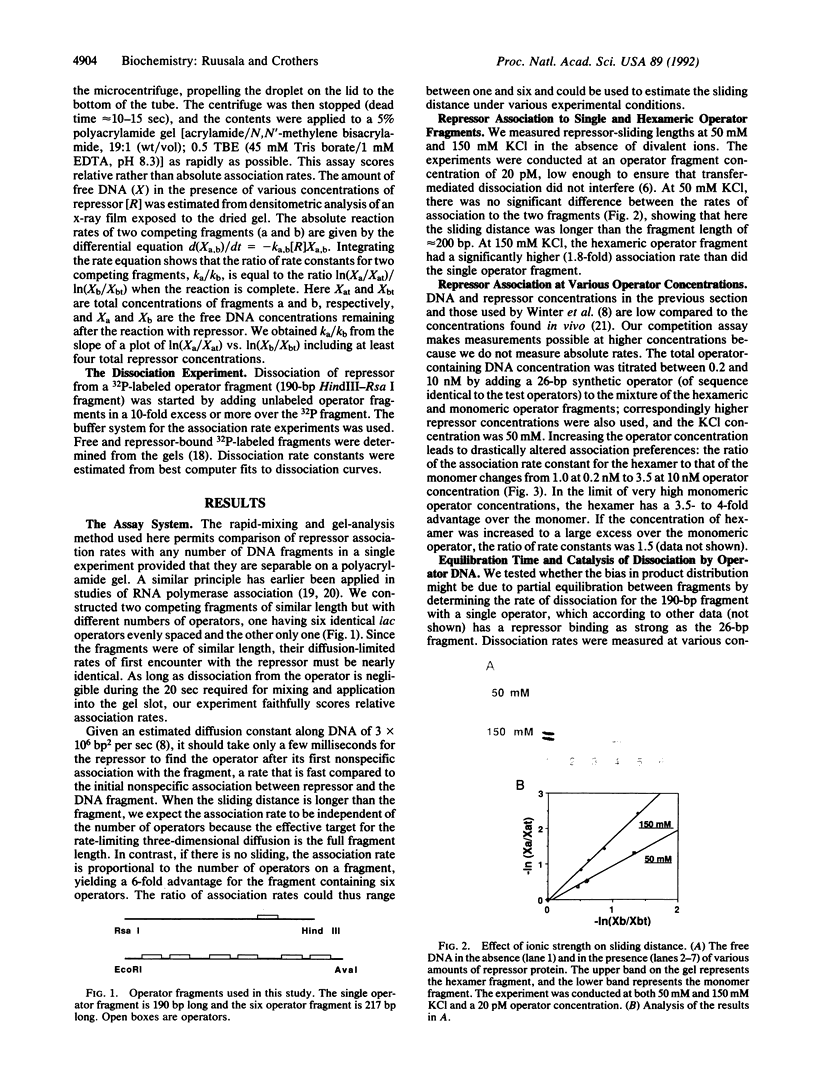
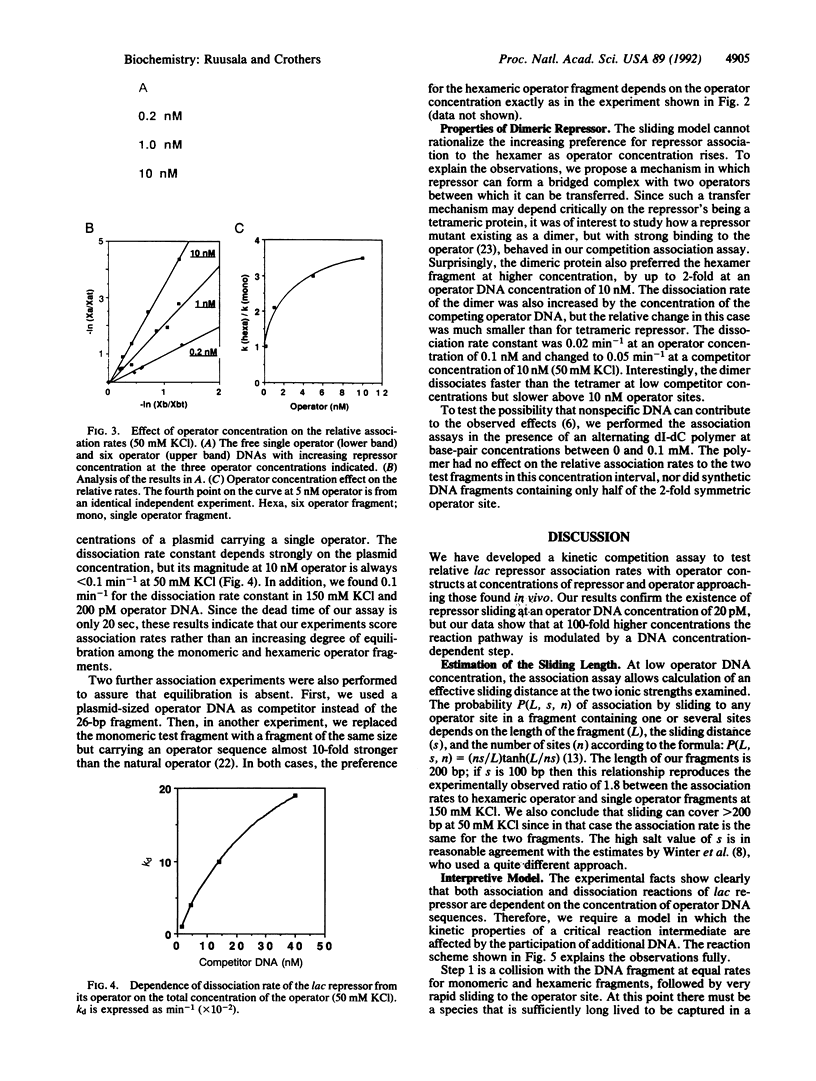


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Berg O. G., Winter R. B., von Hippel P. H. Diffusion-driven mechanisms of protein translocation on nucleic acids. 1. Models and theory. Biochemistry. 1981 Nov 24;20(24):6929–6948. doi: 10.1021/bi00527a028. [DOI] [PubMed] [Google Scholar]
- Bresloff J. L., Crothers D. M. DNA-ethidium reaction kinetics: demonstration of direct ligand transfer between DNA binding sites. J Mol Biol. 1975 Jun 15;95(1):103–123. doi: 10.1016/0022-2836(75)90339-3. [DOI] [PubMed] [Google Scholar]
- Fried M. G., Crothers D. M. Kinetics and mechanism in the reaction of gene regulatory proteins with DNA. J Mol Biol. 1984 Jan 25;172(3):263–282. doi: 10.1016/s0022-2836(84)80026-1. [DOI] [PubMed] [Google Scholar]
- Fried M., Crothers D. M. Equilibria and kinetics of lac repressor-operator interactions by polyacrylamide gel electrophoresis. Nucleic Acids Res. 1981 Dec 11;9(23):6505–6525. doi: 10.1093/nar/9.23.6505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jack W. E., Terry B. J., Modrich P. Involvement of outside DNA sequences in the major kinetic path by which EcoRI endonuclease locates and leaves its recognition sequence. Proc Natl Acad Sci U S A. 1982 Jul;79(13):4010–4014. doi: 10.1073/pnas.79.13.4010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kammerer W., Deuschle U., Gentz R., Bujard H. Functional dissection of Escherichia coli promoters: information in the transcribed region is involved in late steps of the overall process. EMBO J. 1986 Nov;5(11):2995–3000. doi: 10.1002/j.1460-2075.1986.tb04597.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kania J., Müller-Hill B. Construction, isolation and implications of repressor-galactosidase - beta-galactosidase hybrid molecules. Eur J Biochem. 1977 Oct 3;79(2):381–386. doi: 10.1111/j.1432-1033.1977.tb11819.x. [DOI] [PubMed] [Google Scholar]
- Kao-Huang Y., Revzin A., Butler A. P., O'Conner P., Noble D. W., von Hippel P. H. Nonspecific DNA binding of genome-regulating proteins as a biological control mechanism: measurement of DNA-bound Escherichia coli lac repressor in vivo. Proc Natl Acad Sci U S A. 1977 Oct;74(10):4228–4232. doi: 10.1073/pnas.74.10.4228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim J. G., Takeda Y., Matthews B. W., Anderson W. F. Kinetic studies on Cro repressor-operator DNA interaction. J Mol Biol. 1987 Jul 5;196(1):149–158. doi: 10.1016/0022-2836(87)90517-1. [DOI] [PubMed] [Google Scholar]
- Krämer H., Niemöller M., Amouyal M., Revet B., von Wilcken-Bergmann B., Müller-Hill B. lac repressor forms loops with linear DNA carrying two suitably spaced lac operators. EMBO J. 1987 May;6(5):1481–1491. doi: 10.1002/j.1460-2075.1987.tb02390.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lehming N., Sartorius J., Oehler S., von Wilcken-Bergmann B., Müller-Hill B. Recognition helices of lac and lambda repressor are oriented in opposite directions and recognize similar DNA sequences. Proc Natl Acad Sci U S A. 1988 Nov;85(21):7947–7951. doi: 10.1073/pnas.85.21.7947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin S. Y., Riggs A. D. Lac repressor binding to non-operator DNA: detailed studies and a comparison of eequilibrium and rate competition methods. J Mol Biol. 1972 Dec 30;72(3):671–690. doi: 10.1016/0022-2836(72)90184-2. [DOI] [PubMed] [Google Scholar]
- Müller-Hill B., Crapo L., Gilbert W. Mutants that make more lac repressor. Proc Natl Acad Sci U S A. 1968 Apr;59(4):1259–1264. doi: 10.1073/pnas.59.4.1259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oehler S., Eismann E. R., Krämer H., Müller-Hill B. The three operators of the lac operon cooperate in repression. EMBO J. 1990 Apr;9(4):973–979. doi: 10.1002/j.1460-2075.1990.tb08199.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ricchetti M., Metzger W., Heumann H. One-dimensional diffusion of Escherichia coli DNA-dependent RNA polymerase: a mechanism to facilitate promoter location. Proc Natl Acad Sci U S A. 1988 Jul;85(13):4610–4614. doi: 10.1073/pnas.85.13.4610. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richter P. H., Eigen M. Diffusion controlled reaction rates in spheroidal geometry. Application to repressor--operator association and membrane bound enzymes. Biophys Chem. 1974 Oct;2(3):255–263. doi: 10.1016/0301-4622(74)80050-5. [DOI] [PubMed] [Google Scholar]
- Riggs A. D., Bourgeois S., Cohn M. The lac repressor-operator interaction. 3. Kinetic studies. J Mol Biol. 1970 Nov 14;53(3):401–417. doi: 10.1016/0022-2836(70)90074-4. [DOI] [PubMed] [Google Scholar]
- Sadler J. R., Tecklenburg M., Betz J. L. Plasmids containing many tandem copies of a synthetic lactose operator. Gene. 1980 Feb;8(3):279–300. doi: 10.1016/0378-1119(80)90005-0. [DOI] [PubMed] [Google Scholar]
- Simons A., Tils D., von Wilcken-Bergmann B., Müller-Hill B. Possible ideal lac operator: Escherichia coli lac operator-like sequences from eukaryotic genomes lack the central G X C pair. Proc Natl Acad Sci U S A. 1984 Mar;81(6):1624–1628. doi: 10.1073/pnas.81.6.1624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winter R. B., Berg O. G., von Hippel P. H. Diffusion-driven mechanisms of protein translocation on nucleic acids. 3. The Escherichia coli lac repressor--operator interaction: kinetic measurements and conclusions. Biochemistry. 1981 Nov 24;20(24):6961–6977. doi: 10.1021/bi00527a030. [DOI] [PubMed] [Google Scholar]
- von Hippel P. H., Berg O. G. Facilitated target location in biological systems. J Biol Chem. 1989 Jan 15;264(2):675–678. [PubMed] [Google Scholar]






