Stem cell derived retinal pigment epithelium (RPE) holds promise as a cell replacement therapy for macular degeneration. This work demonstrates the role of the cAMP and transforming growth factor β signaling pathways in RPE epithelialization in vitro. Modulation of these pathways by small molecules leads to increased efficiency of RPE generation with a higher yield.
Keywords: Retinal pigment epithelium, Stem cells, Proliferation, cAMP, Transforming growth factor-β
Abstract
Retinal pigment epithelium (RPE) cell integrity is critical to the maintenance of retinal function. Many retinopathies such as age-related macular degeneration (AMD) are caused by the degeneration or malfunction of the RPE cell layer. Replacement of diseased RPE with healthy, stem cell-derived RPE is a potential therapeutic strategy for treating AMD. Human embryonic stem cells (hESCs) differentiated into RPE progeny have the potential to provide an unlimited supply of cells for transplantation, but challenges around scalability and efficiency of the differentiation process still remain. Using hESC-derived RPE as a cellular model, we sought to understand mechanisms that could be modulated to increase RPE yield after differentiation. We show that RPE epithelialization is a density-dependent process, and cells seeded at low density fail to epithelialize. We demonstrate that activation of the cAMP pathway increases proliferation of dissociated RPE in culture, in part through inhibition of transforming growth factor-β (TGF-β) signaling. This results in enhanced uptake of epithelial identity, even in cultures seeded at low density. In line with these findings, targeted manipulation of the TGF-β pathway with small molecules produces an increase in efficiency of RPE re-epithelialization. Taken together, these data highlight mechanisms that promote epithelial fate acquisition in stem cell-derived RPE. Modulation of these pathways has the potential to favorably impact scalability and clinical translation of hESC-derived RPE as a cell therapy.
Significance
Stem cell-derived retinal pigment epithelium (RPE) is currently being evaluated as a cell-replacement therapy for macular degeneration. This work shows that the process of generating RPE in vitro is regulated by the cAMP and transforming growth factor-β signaling pathway. Modulation of these pathways by small molecules, as identified by phenotypic screening, leads to an increased efficiency of generating RPE cells with a higher yield. This can have a potential impact on manufacturing transplantation-ready cells at large scale and is advantageous for clinical studies using this approach in the future.
Introduction
Retinal pigment epithelium (RPE) cells are a highly specialized, polarized cell type situated as a tightly packed monolayer between photoreceptors and the choroid. RPE cells help to maintain homeostasis and photoreceptor function in the retina via a range of processes including metabolism and storage of retinoid, phagocytosis of rod outer segments, scattered light absorption, barrier activity, and ion transport [1]. Loss of RPE function is associated with diseases such as age-related macular degeneration (AMD), retinitis pigmentosa, proliferative vitreoretinopathy (PVR), and diabetic retinopathies, among others, which often result in loss of vision [2]. The prevalence of AMD is on the rise with increasing population longevity [3], which necessitates the development of new therapeutic approaches to treat this disease. A potential treatment for at least some of these ocular pathologies is the replacement of the dysfunctional RPE with healthy RPE, including RPE derived from pluripotent stem cells [4]. Several studies have demonstrated successful derivation of mature RPE from human embryonic stem cells (hESCs) or human induced pluripotent stem cells (hiPSCs) by using a range of differentiation protocols that produce RPE at varying levels of efficiency and time scales [5–7]. Stem cell-derived RPE is functional [8–10] and has a gene expression signature equivalent to primary RPE [11–13]. Furthermore, their transplantation shows long-term protective effects, leading to restored visual function in animal models of retinal dystrophy [14–17]. However, manufacture of transplantation-ready RPE cells with high yield and purity under conditions compatible with good manufacturing practice (GMP) still remains a significant challenge that needs to be overcome to successfully use this approach more widely.
RPE cells are normally quiescent in the adult eye. However, in vitro dissociation and passaging procedures release RPE from cell-cycle arrest and induce proliferation. This is accompanied by an epithelial-mesenchymal transition, in which cells deviate from an epithelial state and acquire a mesenchymal phenotype. This is followed by a Forkhead Box M1 (FOXM1)-dependent mesenchymal-epithelial transition that results in reuptake of epithelial characteristics [18]. The proliferative capacity of RPE allows serial expansion of cells in culture, albeit limited to a few passages, because cells lose key cytological and functional attributes with extended passage [19]. This limits the capacity of expansion of RPE with respect to manufacturing cells at scale for therapy. In this report, we sought to further understand the molecular mechanisms underpinning the acquisition of RPE cell fate and investigate the contribution of cellular proliferation together with cellular density to this process. Manipulation of these mechanisms would potentially allow better control of the differentiation and expansion process, which might favorably impact cell production for clinical applications.
Cyclic AMP (cAMP) is a second messenger used in intracellular signal transduction and can regulate the function of protein kinases, which, in turn, phosphorylate other proteins, including transcription factors such as cAMP response element-binding protein. Activation of cAMP-inducible genes regulates cellular processes such as differentiation and proliferation in a variety of cell types [20]. In RPE cells, cAMP has been shown to promote differentiation and maturation, purportedly via proliferation-independent mechanisms, such as promoting melanosome and pigmentation-related pathways [21–24]. Approaches to increase cAMP (e.g., with the use of vasointestinal peptide, a hormone that raises intracellular cAMP) have been utilized in directing differentiation of embryonic stem cells toward the RPE lineage [9]. Similarly, review of the literature suggests that cAMP signaling controls migratory and fibrotic responses similar to those seen in PVR [25–27]. Therefore, we sought to determine whether cAMP treatment impacted gain of RPE identity during in vitro dissociation and culture of hESC-derived RPE. Interestingly, we observed that treatment with dibutyryl-cAMP (dbcAMP; bucladesine), a cell-permeable synthetic analog of cAMP, or use of molecules such as forskolin that increase intracellular cAMP through activation of adenylyl cyclase, promote reacquisition of RPE fate across a range of seeding densities in a proliferation-dependent manner. Investigation into the underlying molecular pathways further suggested that interplay between cAMP signaling and the transforming growth factor-β (TGF-β) pathway might be responsible for the phenotypic effects observed.
The TGF-β pathway plays a central role in several cellular processes, where it transduces extracellular signals into intracellular transcriptional responses that control cell growth, apoptosis, and differentiation. The TGF-β superfamily of endogenous signaling ligands consists of more than 25 different ligands, including bone morphogenetic proteins (BMPs), growth and differentiation factors, and activins. TGF-β ligands (TGF-β1, -β2, and -β3) recruit TGF-β type II receptors (TGF-βR2, ACTRIIA, or ACTRIIB) with a set of type I receptors (activin receptor-like kinase 4 [ALK4], ALK5, or ALK7). Receptor dimerization together with phosphorylation of SMADs 2/3 due to the receptor serine/threonine kinase activity leads to SMAD nuclear translocation and downstream regulation of transcription resulting in signal propagation [28]. In this report, we show that inhibition of TGF-β signaling, in particular through inhibition of ALK5 kinase activity, with the use of small molecule tools results in an increase in epithelial phenotype of RPE. This suggests that signaling through TGF-β type I receptors is an important determinant of epithelial cell fate.
In summary, this study furthers our understanding of the biology of stem cell-derived RPE and additionally highlights potential points of intervention in pathways that could be targeted to increase efficiency of generating RPE suitable for transplantation.
Materials and Methods
Cell Culture and Manipulations
RPE were generated from the hESC line SHEF1 (obtained from Intercytex Ltd., Manchester, U.K., http://www.intercytex.com; also available in the UK Stem Cell Bank under accession no. R-05-007) by using the spontaneous differentiation method described previously [7]. Pluripotent cells were cultured as feeder-free colonies on hESC-qualified Matrigel (BD Biosciences, San Jose, CA, http://www.bdbiosciences.com) by using mTesR1 medium (StemCell Technologies, Vancouver, BC, Canada, http://www.stemcell.com). RPE foci (passage 0; P0) were excised with a scalpel and dissociated into a single cell suspension by using Accutase (ThermoFisher Scientific, Oakwood Village, OH, https://www.thermofisher.com) and plated onto CellStart (Invitrogen, Carlsbad, CA, http://www.invitrogen.com)-coated surfaces for expansion for a period of 8 weeks. For all experiments, the starting population of cells used were at P1 or P2, such that each round of expansion led to three to four population doublings. For experiments involving culture of cells in transwells, cells at P1 were plated on Vitronectin (AMS Biotechnology, Abingdon, U.K., http://www.amsbio.com)-coated transwells at a density of 362,500 cells per cm2 and cultured for a period of 3 weeks. Quantification of vascular endothelial growth factor (VEGF) and pigment epithelium-derived factor (PEDF) in the cell culture supernatant was performed on the Meso Scale discovery platform by using commercially available kits (MS6000, Human VEGF PEDF duplex assay; Meso Scale Diagnostics, Rockville, MD, https://www.mesoscale.com). Bead phagocytosis assay was performed by using FluoSpheres polystyrene microspheres (1.0 μm of Red; Invitrogen) following a published protocol [29]. Where required, media was supplemented with 10 μM Forskolin (Sigma-Aldrich, St. Louis, MO, http://www.sigmaaldrich.com), 0.5 mM dbcAMP (Sigma-Aldrich), 10 ng/ml TGF-β1, or 10 μg/ml 1D11 (R&D Biosystems, Minneapolis, MN, https://www.rndsystems.com) at the time of seeding. For screening, compounds were obtained from the Pfizer library (Pfizer, Cambridge, U.K., http://www.pfizer.co.uk), solubilized in 100% dimethyl sulfoxide (DMSO), and serially diluted such that final assay and vehicle concentration was 0.1% DMSO. ALK2 inhibitors DMH1 and LDN-212854 were purchased from Tocris Bioscience (Tocris Bioscience, Bristol, U.K., https://www.tocris.com). For all treatments, medium was replenished three times per week.
RNA Extraction, cDNA Synthesis, and Quantitative Polymerase Chain Reaction
Total RNA was extracted from RPE cells by using the RNEasy Mini or Micro Kit (Qiagen, Hilden, Germany, http://www.qiagen.com) with on-column DNase digestion. cDNA was synthesized by using the High Capacity cDNA Synthesis kit (Applied Biosystems, Foster City, CA, http://www.appliedbiosystems.com). Individual gene expression was assessed by using predesigned TaqMan assays, and the reactions were carried out on the ABI7900 or CFX96 platform. Gene expression in all instances was quantified by the relative quantification method of 2-ΔΔCt and normalized to geometric means of housekeeping genes.
Microarray Analysis
mRNA was hybridized on Illumina HT-12v4 BeadChips (Illumina, San Diego, CA, http://www.illumina.com) according to manufacturer’s instructions. Raw microarray data were log-transformed and quantile-normalized by using the beadarray package in Bioconductor. Differential expression analysis was performed using limma and geneset enrichment analysis with camera. All other analysis was performed with standard Bioconductor/R packages. Standard hierarchical clustering algorithms were used to achieve visual clarity of the heatmaps. The microarray data are available in the ArrayExpress database under accession nos. E-MTAB-854 and E-MTAB-3878.
Immunocytochemistry
Samples were fixed in 4% paraformaldehyde in phosphate-buffered saline (PBS) for 15 minutes, followed by blocking and permeabilization using 0.3% Triton X-100 in PBS and 10% normal donkey serum. Primary antibodies used in this study were mouse anti-premelanosome protein 17 (anti-PMEL17; 1:25; M0634; Dako, Carpinteria, CA, http://www.dako.com), rabbit anti-Ki67 (1:500; VP-K451; Vector Laboratories, Burlingame, CA, http://vectorlabs.com), rabbit anti-zonula occludens 1 (anti-ZO1; 1:100; 187430, ThermoFisher Scientific), mouse anti-α-smooth muscle actin (anti-α-SMA; 1:1,000; Sigma-Aldrich A5228), mouse anti-cellular retinaldehyde-binding protein (anti-CRALBP; 1:200; MA1-813; Affinity Biosciences, Cambridgeshire, U.K., http://www.affbiotech.com), rabbit anti-Mer receptor tyrosine kinase (anti-MERTK; 1:50; ab52968; Abcam, Cambridge, MA, http://www.abcam.com). For 5-ethynyl-2-deoxyuridine (EdU) incorporation, cells were treated with 10 µM EdU 18 hours before fixation. EdU incorporation was measured by using the Click-iT 488 Imaging kit (Thermo Fisher Scientific Life Sciences, Waltham, MA, http://www.thermofisher.com) according to manufacturer’s instructions. Nuclei were counterstained with the nuclear dye 4′,6-diamidino-2-phenylindole (DAPI). For immunocytochemistry of transwell sections, the transwell membrane was fixed and frozen in a cryomold containing M1 embedding matrix, and 10 μm sections were prepared by using a cryostat and were immunostained. Images were captured and analyzed on the ImageXpress platform (Molecular Devices, Sunnyvale, CA, http://www.moleculardevices.com) or on a confocal microscope (Leica Biosystems, Buffalo Grove, IL, http://www.leicabiosystems.com).
Results
RPE Epithelialization Is a Density-Dependent Process
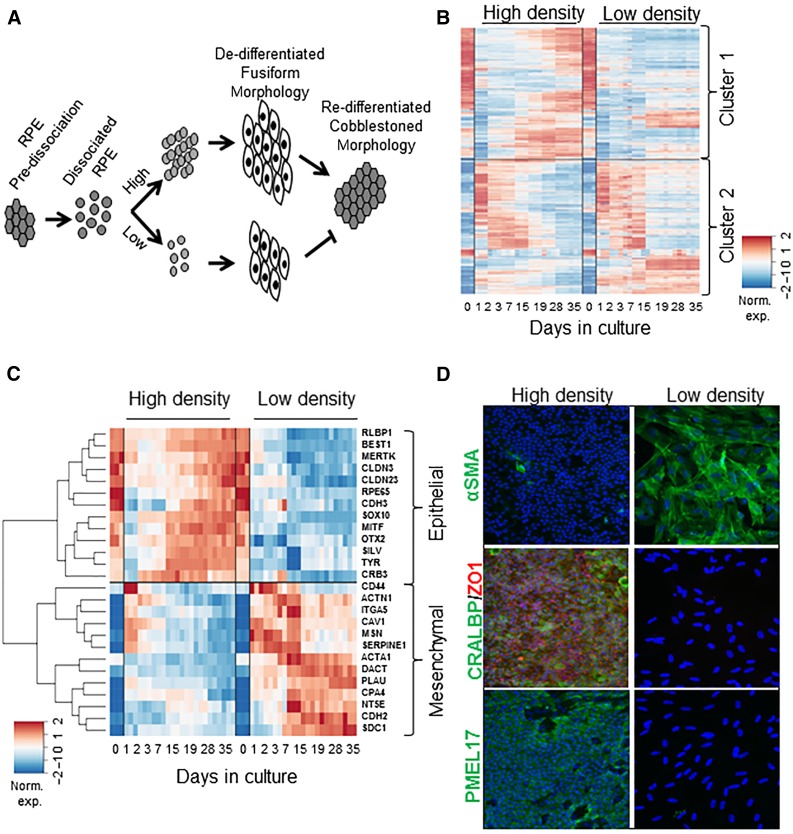
Although RPE are differentiated epithelial cells, they retain the ability to change state into a mesenchymal phenotype when dissociated and cultured. This transitional state then undergoes a mesenchymal-epithelial transition to redifferentiate into an epithelial phenotype (Fig. 1A) [18]. Successful epithelialization is critically reliant on seeding density such that cells seeded at high density successfully re-epithelialize, whereas cells seeded at low density remain mesenchymal. In order to gain further insight into this density-dependent process, we used whole-genome microarrays to analyze a transcriptional profile time course of cells seeded either at high (100,000 cells per cm2) or low (8,000 cells per cm2) density. Strikingly, the expression patterns of most genes clustered into two distinct, symmetrical groups (Fig. 1B). Genes in the first cluster exhibit relatively high expression in the starting RPE sample (day 0; D0), but show an immediate drop in expression after dissociation. These genes then steadily increase in expression toward the starting level in the high-density culture, but remain suppressed in the low-density cultures. The second cluster shows the opposite pattern: initially low in expression, these genes immediately rise after dissociation before decreasing to their D0 level in the high-density culture, whereas their expression tends to remain high in the low-density cultures. Importantly, we note that the transcriptional state of the cells seeded at high density returns toward that of the starting RPE population, whereas cells seeded at low density remain distinct. The list of genes belonging to clusters 1 and 2 can be seen in supplemental online Table 1.
Figure 1.
Cell-density dependent effects on re-epithelialization in stem cell-derived RPE. (A): Schematic representation of density-dependent RPE culture where RPE seeded at high density undergo proliferation and successfully epithelialize whereas RPE seeded at low density remain mesenchymal. (B): Heatmap of expression profiles of the top 250 expressed genes ranked by the significance of their expression changes over time in high-density (100,000 cells per cm2) and low-density (8,000 cells per cm2) cultures. Raw expression data are mean centered and scaled to unit variance prior to clustering. The genes cluster into two groups (1 and 2) based on the observed expression pattern. Cluster 1 genes show initial downregulation, whereas cluster 2 genes show initial upregulation upon dissociation and culture. Genes from both clusters return to basal levels with time upon high density seeding but not upon low density seeding. (C): Heatmap showing changes in gene expression of a panel of representative epithelial and mesenchymal markers over a timecourse of RPE culture where cells are seeded as in B. (D): Representative images showing immunocytochemistry for epithelial (CRALBP, ZO1, and PMEL17) and mesenchymal markers (α-SMA) at day 42 in cultures seeded as in B. Images have been captured at ×10 magnification. Abbreviations: ACTN1, actinin-α-1; BEST1, bestrophin 1; CDH2, N-cadherin; CDH3, cadherin 3; CLDN23, claudin 23; CLDN3, claudin 3; CPA4, carboxypeptidase A4; CRALBP, cellular retinaldehyde-binding protein; CRB3, crumbs homolog 3; DACT, Dapper homolog; ITGA5, integrin-α-5; MERTK, Mer receptor tyrosine kinase; MITF, microphthalmia-associated transcription factor; MSN, medium spiny neuron; Norm. exp, normalized expression; OTX2, orthodenticle homeobox 2; PLAU, urokinase plasminogen activator; PMEL17, premelanosome protein 17; RLBP1, retinaldehyde-binding protein 1; RPE, retinal pigment epithelium; SDC1, syndecan 1; SERPIN, serine protease inhibitor; SILV, silver homolog; α-SMA, α-smooth muscle actin; SOX10, SRY-box containing gene 10; TYR, tyrosinase; ZO1, zonula occludens 1.
We further examined the expression of epithelial and mesenchymal markers, and, consistent with visual observations, there was an initial upregulation of mesenchymal markers concomitant with a decrease in epithelial and RPE-specific markers upon dissociation. This was followed by a re-expression of epithelial markers in high-density cultures, with time indicating a successful mesenchymal-epithelial transition. In contrast, cells seeded at low density retained high expression of mesenchymal markers and low expression of epithelial markers throughout the time course (Fig. 1C). Immunostaining for representative RPE markers PMEL17, CRALBP, ZO1, and the mesenchymal marker α-SMA was consistent with the mRNA expression pattern, confirming that low-density cultures did not display an epithelial phenotype (Fig. 1D). Taken together, this analysis strongly suggests that an appropriately high seeding density is required for successful epithelialization. In addition, we confirmed that the RPE obtained after re-epithelialization were polarized and functional in terms of phagocytosis and ability to secrete trophic factors such as VEGF and PEDF, which are key features of native RPE. RPE was seeded on transwell inserts and cultured for a period of 3 weeks. Cross-sections of the insert were immunostained for the RPE markers ZO1, MERTK, and PMEL17 (supplemental online Fig. 1A). The expression of ZO1 was restricted to the cell junctions, MERTK to the apical surface, and PMEL17 throughout the cell body, consistent with cell polarization. Furthermore, RPE were capable of phagocytosing fluorescently labeled beads (supplemental online Fig. 1B), which serves as a surrogate measure of ability to phagocytose rod outer segments [30]. Finally, we measured the ability of RPE to secrete VEGF and PEDF. Spent medium was collected from RPE cultured on transwells and compared with ARPE19 cells cultured under identical conditions. We observed an increase in VEGF and PEDF secretion by RPE with time in culture (e.g., at day 21 vs. day 4), indicative of attainment of polarization and functionality upon re-epithelialization (supplemental online Fig. 1C). In contrast, ARPE19 did not secrete detectable levels of PEDF, consistent with the notion that they do not replicate all aspects of native RPE function.
Increasing cAMP Levels Promotes Acquisition of RPE Identity in a Time- and Density-Dependent Manner
We have shown a density-dependent effect on reacquisition of epithelial identity where cells seeded at high density readily differentiate back into a RPE state over a period of time, but cells seeded at a lower density do not. Next, we wanted to understand the pathways underlying RPE epithelialization and identify their putative modulators. Given the literature suggesting effects of cAMP signaling on RPE characteristics [21–25], we selected forskolin (FSK) as a tool to increase intracellular cAMP levels. We found that treatment with 10 μM FSK led to a consistent increase in pigmentation and epithelial characteristics across a range of seeding densities (Fig. 2A; supplemental online Fig. 2). This prompted us to ask whether FSK treatment modulated the gene expression profile of cells during the time course of RPE culture. TaqMan low density arrays (TLDAs) were used to perform quantitative polymerase chain reaction (qPCR) with a panel of 24 genes indicative of epithelial or mesenchymal identity (supplemental online Fig. 3). We performed a principal component analysis on this multivariate data and found that the first principal component correlated with high expression of epithelial genes and low expression of mesenchymal genes. We used the first principal component as a univariate weighted score of RPE identity and therefore used it to measure the effect of FSK at a range of seeding densities (2.5k, 10k, 15k, 20k, 25k, 27.5k, 30k, 38k; where k is ×1,000 cells per cm2) after 63 days in culture. This showed that treatment with FSK led to an increase in “RPE score,” that is, acquisition of the RPE phenotype, at every plating density tested (Fig. 2B).
Figure 2.
Increasing cAMP signaling promotes acquisition of RPE identity across multiple cell densities. (A): Representative brightfield images showing RPE seeded at multiple densities in the presence or absence of 10 μM forskolin treatment at day 63 in culture. Scale bars = 400 µm. (B): The RPE score (main text) of each sample in the presence (blue) and absence (red) of 10 μM FSK at day 63 is plotted against the seeding density (2.5k, 10k, 15k, 20k, 25k, 27.5k, 30k, and 38k). The shaded area represents 95% confidence intervals, and the solid circles represent biological replicates at the given density. Abbreviations: FSK, forskolin; k, ×1,000 cells per cm2; RPE, retinal pigment epithelium.
A limitation of the TLDA-based approach is the comparatively limited set of genes profiled. Because of this, we were limited in our ability to generate unbiased hypotheses about the underlying mechanism of increased epithelialization with increased cAMP levels. We therefore sought to extend these initial qPCR-based observations to global gene expression profiling. In order to generate a comprehensive dataset, we used microarrays to assess the effect of increased cAMP signaling on cells seeded at 10,000; 20,000; and 40,000 cells per cm2 over time (3, 15, and 34 days in culture). The stable, cell-permeant cAMP analog, dbcAMP, was used to enhance intracellular cAMP signaling in this study. Interestingly, principal component analysis showed that at every timepoint sampled, the gene expression signature of cells treated with 0.5 mM dbcAMP was equivalent to cells seeded at double the density in medium alone (Fig. 3A). For example, the expression profile of cells seeded at 10,000 cells per cm2 with dbcAMP was equivalent to that of cells at 20,000 cells per cm2 without dbcAMP. Similarly, the expression profile of cAMP-treated cells at 20,000 cells per cm2 was comparable to cells at 40,000 cells per cm2 without supplementation. A clustered heatmap representation of the top 1% variable genes reiterates the same observation of a “doubling effect” of density in cells exposed to dbcAMP, where cells treated with dbcAMP clustered with cells seeded at twice the density without dbcAMP exposure (Fig. 3B). We next looked at the expression of specific RPE markers and found them to be consistent with the profiles seen with the whole-genome analyses (Fig. 3C). These expression patterns were further confirmed by gene-specific qPCR for representative genes (supplemental online Fig. 4A) together with immunostaining for protein expression (supplemental online Fig. 4B). Additionally, we determined that a dose of at least 0.5 mM dbcAMP was required to see this effect (supplemental online Fig. 4C). In summary, these data show that intracellular elevation of cAMP signaling leads to an increase in acquisition of RPE phenotype.
Figure 3.
Gene expression analysis of dbcAMP-treated retinal pigment epithelium (RPE). (A): Principal component analysis of the microarray gene expression data are shown for samples obtained from three different seeding densities (indicated by the shape of each point: circle, 10,000 cells per cm2; square, 20,000 cells per cm2; diamond, 40,000 cells per cm2) in the presence (blue) or absence (red) of dbcAMP. The data from four time points (day 0, 3, 15, and 34 after seeding) is indicated by the shading of each point and the labeled ellipses such that color intensity increases with increasing time in culture. The day 0 samples are indicated as black points and the proportion of the total variance captured by each principal component is indicated in the axis title. This shows that at each time point tested, there is a “doubling effect,” that is, clustering of samples seeded at half the density in the presence of dbcAMP with samples at double the density but without dbcAMP. (B): Heatmap of gene expression levels for the top 1% most variable genes (rows) observed at day 34 in culture. The seeding density and dbcAMP treatment status each sample is indicated by the labels at the bottom of the heatmap. Expression levels are shown mean centered and scaled to unit variance for each gene. Clustering of samples consistent with the doubling effect can be seen. (C): Heatmap of gene expression levels for selected RPE markers across all timepoints, seeding density and dbcAMP treatment is shown. Abbreviations: BEST1, bestrophin 1; D0, day 0; D3, day 3; D15, day 15; D34, day 34; dbcAMP, dibutyryl-cAMP; k, ×1,000 cells per cm2; MITF, microphthalmia-associated transcription factor; RLBP1, retinaldehyde-binding protein 1; SILV, silver homolog; TYR, tyrosinase.
Increasing cAMP Levels Promotes RPE Proliferation
In order to gain insight into the molecular mechanisms by which cAMP positively modulates RPE fate, we performed gene set enrichment analyses (GSEA) of the dbcAMP-induced expression changes. Besides markers of the RPE phenotype, there was enhanced representation of sets of genes associated with increased proliferation (Fig. 4A; supplemental online Table 2). This suggested that the increased epithelial phenotype could occur as a result of increased proliferation. To test this hypothesis, we assessed proliferation in cells treated with or without dbcAMP, using the incorporation of EdU, a modified thymidine analog, which can be visualized using “Click” chemistry-dependent imaging [31]. We found an increase in proliferating cells upon supplementation with dbcAMP on days 7, 14, and 21 of culture (Fig. 4B). Importantly, with increased time (e.g., at day 56), there was no significant difference in proliferation between cells supplemented with dbcAMP or not (Fig. 4B). This suggests that, although intracellular cAMP elevation promotes proliferation, it is not an uncontrolled feature, and homeostatic RPE mechanisms are in play to achieve quiescence after the initial phase of proliferation upregulation. An increase in proliferation, measured by immunostaining for Ki67, was seen upon treatment of cells with FSK (Fig. 4C), showing that intracellular cAMP elevation by different methods resulted in a similar effect. We also determined the optimum period of dbcAMP exposure required to observe effects on proliferation. Cells were seeded at a density of 38,000 cells per cm2 and cultured for a period of 8 weeks with continuous dbcAMP treatment for 2 weeks, 3 weeks, or the entire duration of 8 weeks. The increase in proliferation resulted in more cells whose nuclei stained positive for the nuclear-specific fluorescent dye DAPI. Using DAPI count as a surrogate for cell number estimation, we observed no difference in cell number between the dbcAMP exposure times tested (Fig. 4D). From these data, we conclude that exposure of cells to dbcAMP for the first 2 weeks after dissociation and seeding is sufficient to result in increased proliferation.
Figure 4.
dbcAMP increases proliferation of retinal pigment epithelium (RPE) cells. (A): Exemplar Gene Ontology terms, derived from comparison of cultures in the presence vs. absence of dbcAMP at Day34 (20,000 cells per cm2 seeding density), alongside their gene set test significance p values (p < .05). (B): Representative images showing EdU incorporation in the presence or absence of dbcAMP in RPE seeded at 38,000 cells per cm2 at different timepoints in culture. The quantification of EdU incorporation is shown below. Bars represent Mean + SD (n = 8). (C): Representative images showing immunocytochemistry for Ki67 in the presence or absence of 10 μM FSK in RPE seeded at 38,000 cells per cm2 at different timepoints in culture. The quantification of images is shown below. Bars represent mean + SD (n = 3). (D): Representative images showing nuclei stained with DAPI in RPE treated seeded at 38,000 cells per cm2 and cultured for a period of 8 weeks with different periods of exposure to dbcAMP Quantification of cell number, measured by DAPI positive nuclei per frame imaged is shown below. Bars represent mean + SD (n = 3). All images have been captured at ×10 magnification. Abbreviations: 2+6, 2-week dbcAMP+ 6-week media; 3+5, 3-week dbcAMP+ 5-week media; 8, 8-week dbcAMP; D, downregulated; D7, day 7; D14, day 14; D21, day 21; D56, day 56; DAPI, 4′,6-diamidino-2-phenylindole; dbcAMP, dibutyryl-cAMP; EdU, 5-ethynyl-2-deoxyuridine; FDR, false discovery rate; FSK, forskolin; GO, Gene Ontology; U, upregulated.
Activation of cAMP Signaling Suppresses the TGF-β Pathway to Promote Successful Epithelialization
To understand mode of cAMP action, causal reasoning analysis [32] of the gene expression dataset was performed to predict molecular regulators of the observed gene expression changes. We compared the gene expression profiles of cultures seeded at a density of 20,000 cells per cm2 in the presence or absence of dbcAMP at day 34 and observed that TGF-β signaling was a key mechanism being suppressed by dbcAMP treatment (p = 7.95 × 10−17). In order to further decipher the interplay between cAMP-TGF-β pathways, we made use of information available in the literature. It is known that activation of the TGF-β pathway leads to downstream activation of SMADs 2/3, which then directly affects gene expression by binding to gene promoters. We identified genes that are directly bound by SMAD3 using a publically available chromatin immunoprecipitation-sequencing (ChIP-Seq) dataset [33]. We then investigated how the expression of these genes changed upon dbcAMP treatment. Interestingly, we observed a significantly decreased expression of SMAD3-responsive genes in RPE cells treated with dbcAMP (p < 1 × 10−10) (Fig. 5A). Taken together, these two lines of evidence support the notion that active cAMP signaling, which promotes proliferation and epithelialization, is associated with inhibition of the TGF-β pathway in RPE.
Figure 5.
Role of TGF-β signaling in retinal pigment epithelium. (A): Histogram showing change in expression of SMAD3-bound genes in dbcAMP versus control cultures at day 34 (20,000 cells per cm2 seeding density). The frequency of genes is plotted on the y axis and the log fold change is plotted on the x axis such that no change in expression is equivalent to a zero log fold change. The leftward shift of the distribution indicates a significant decrease in expression of SMAD3 responsive genes with dbcAMP treatment. (B): Quantitative polymerase chain reaction-based measurement of transcript expression of a panel of epithelial (CRB3, BEST1, and PMEL) and mesenchymal (CDH2, MMP2, and GREM1) markers in cells seeded at 100,000 cells per cm2 and exposed to 10 ng/ml TGF-β1 for 5 days. ATP5B and CYC1 are used as housekeeping genes. Bars represent mean + SD (n = 3). p < .05 (Student’s t test). (C): Representative images showing immunocytochemistry for indicated markers in cells seeded at high density and exposed to TGF-β1 for 5 days. Images have been captured at ×10 magnification. (D): Quantification of C. Abbreviations: BEST1, bestrophin 1; CDH2, N-cadherin; CRB3, crumbs homolog 3; GREM1, gremlin 1; MMP2, matrix metalloproteinase 2; PMEL, premelanosome protein; PMEL17, premelanosome protein 17; α-SMA, α-smooth muscle actin; TGF-β, transforming growth factor-β.
In order to explore the role of TGF-β signaling more deeply, we looked at the expression profile of TGF-β ligands, as well as genes downstream of type 1 and 2 TGF-β receptor activation in high- versus low-density cultures. This demonstrated that high-density cultures, which epithelialize normally, have low-expression TGF-β pathway members (supplemental online Fig. 5). This is in line with the hypothesis that TGF-β signaling is suppressed for successful epithelialization. In order to further verify this observation, we asked whether exogenous activation of the TGF-β pathway in high-density cultures, which would normally have suppressed TGF-β signaling, would lead to a decrease in epithelial phenotype acquisition. Recombinant TGF-β1 was added to cultures seeded at a high density of 100,000 cells per cm2 and maintained in the medium for a period of 5 days. An increase in transcript expression of mesenchymal markers, along with a decrease in expression of epithelial markers as measured by qPCR, was observed, which suggested that activation of TGF-β signaling indeed prevented successful epithelialization (Fig. 5B). This was also verified by immunostaining for expression of epithelial (PMEL17) and mesenchymal (α-SMA) proteins (Fig. 5C, 5D).
We next tested whether the effect on phenotype was a consequence of change in cellular proliferation. Consistent with the effects seen with dbcAMP supplementation, activation of TGF-β signaling by addition of recombinant TGF-β1 protein led to a decrease in proliferation as measured by immunostaining for Ki67 (Fig. 5C, 5D). Taken together, our data support the hypothesis that TGF-β signaling and its associated antiproliferative effect is suppressed in RPE cultures seeded at high density that acquire an epithelial phenotype.
Screen for Small Molecules That Positively Modulate Epithelialization
A key requirement for a successful RPE cell-replacement therapy is the scalable and efficient generation of RPE with a differentiated epithelial phenotype. In this context, identification of small molecules that could promote consistent acquisition of an epithelial phenotype, even at low densities, would be particularly advantageous. Therefore, we sought to leverage and extend our initial observations and look for small-molecule tools that would promote epithelial phenotype acquisition at low density, a condition in which RPE would not normally epithelialize. We generated a bespoke library consisting of 1,000 small molecules that were known agonists or antagonists of proteins within pathways likely to affect RPE fate based on hypotheses posed by expression data or published literature indications. This included signaling pathways such as BMP, Wnt, TGF-β, etc. FSK was used as a positive control compound known to promote re-epithelialization as shown by our dbcAMP data. We established a high-content immunocytochemistry-based assay for PMEL17, a marker of mature pigmented RPE, in a 384-well format to allow compound screening at sufficiently high throughput with modest cell requirements. Primary screening was performed by using compounds at a concentration of 10 μM. A positive hit was defined as a molecule that increased percent positive PMEL17 expression to greater than 3 standard deviations above the DMSO vehicle control. Effect on DAPI measurement, which is indicative of cell number, was also quantified. Data generated by using these parameters allowed us to identify whether positive hits functioned potentially through a proliferative mode of action. Compounds that led to a significant decrease in DAPI compared with vehicle control with no effect on PMEL17 were considered toxic and ruled out from additional analysis. Data deconvolution demonstrated significant effects on PMEL17 caused by multiple inhibitors of the type 1 TGF-β receptors with a preponderance of compounds with activity against ALK5 kinase. These compounds appeared to promote epithelialization through increasing cell proliferation (i.e., having an effect of increased DAPI count concomitant with increased percent of PMEL17). However, we did not find any mechanisms other than TGF-β signaling in this investigation.
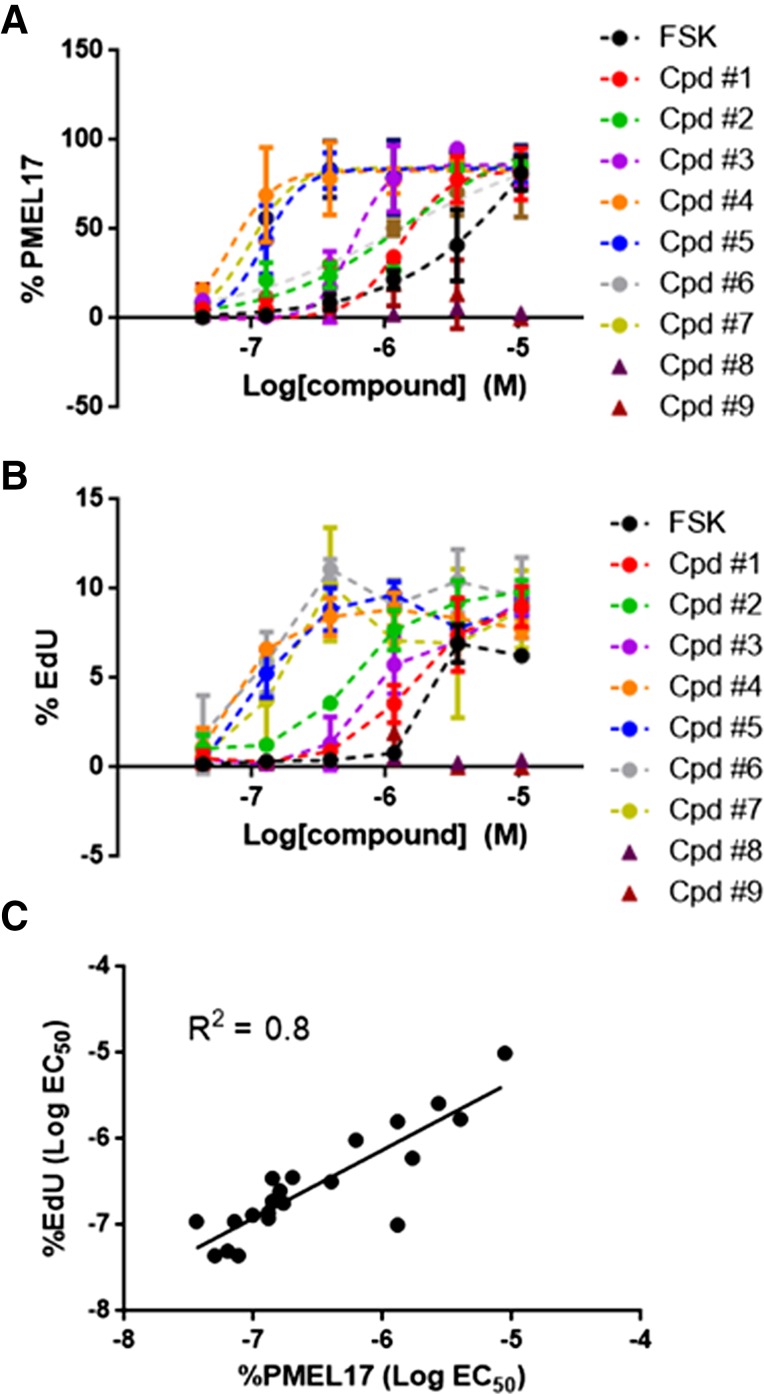
In order to verify primary hits, we extended our investigation to a broader set of ALK5 inhibitors and generated dose-response curves to measure impact on RPE phenotype by immunocytochemistry for PMEL17 (Fig. 6A). We included a further assessment of cellular proliferation by quantifying the level of incorporation of EdU measured by imaging and determined effect of compound dose on this output (Fig. 6B). We determined EC50 values for both measures and noted a close correlation (R2 = 0.8) between EC50 for percent PMEL17-positive and percent EdU-positive cells for all ALK5 inhibitors tested (Fig. 6C), supporting the notion that ALK5 inhibition promotes cellular proliferation and attainment of RPE phenotype. The observed EC50 values for both PMEL17 and EdU were broadly in the same rank order as the published potencies against ALK5 for the compounds tested (supplemental online Fig. 6). However, this is limited by the fact that the ALK5 potencies have not been remeasured in parallel, instead relying on literature values across different assay formats, which can introduce variability. We were also able to rule out off-target effects due to particular compound chemotypes because a consistent effect on RPE phenotype was observed with ALK5 inhibitor compounds from different chemical series. In addition, it is noteworthy that inhibitors of ALK2, a receptor for BMP ligands that signal through a distinct arm in the TGF-β cascade, had no effect on RPE phenotype, indicating specificity of the ALK5 mechanism (compound 8 [DMH1] and compound 9 [LDN-212854] in Fig. 6A, 6B).
Figure 6.
Effect of activin receptor-like kinase 5 (ALK5) inhibitors on retinal pigment epithelium phenotype and proliferation. Cells were seeded at a density of 15,000 cells per cm2 and treated with compounds for 14 days. (A, B): Immunocytochemistry was performed, and dose-response curves were generated measuring percentage of DAPI-positive cells staining positive for PMEL17 (A) and EdU (B). Forskolin was used as the positive control. (C): Correlation between EC50 for both measures for all ALK5 inhibitors screened is shown. Abbreviations: Cpd, compound; EC50, half maximal effective concentration; EdU, 5-ethynyl-2-deoxyuridine; FSK, forskolin; M, molar; PMEL17, premelanosome protein 17; R2, coefficient of determination.
To confirm that the output measure of immunostaining for PMEL17 was indicative of a true increase in epithelial RPE characteristics, we performed qPCR to measure expression of a panel of epithelial and mesenchymal markers. Compounds 6 and 7 (EC50 = 1.27 and 0.1 µM for PMEL17, respectively) were chosen as representative examples for this broader analysis. RPE were seeded at a density of 15,000 cells per cm2 and treated with compounds at three concentrations (10, 1, and 0.1 µM). A consistent increase in several RPE markers (BEST1, LRAT, MERTK, RPE65, and RLBP1) was seen, along with a decrease in a marker indicative of a mesenchymal phenotype (GREM1) (supplemental online Fig. 7). Measurement of these multiple endpoints provided further support to the notion that ALK5 inhibitors promote acquisition of the RPE phenotype.
Validation of TGF-β Mechanism
Following on from compound screening, we wanted to independently verify the TGF-β mechanism through an orthogonal, non-chemical-based interrogation. To enable this, we selected 1D11, a well-characterized monoclonal pan-TGF-β-neutralizing antibody [34], to inhibit endogenous TGF-β signaling. RPE were seeded at low density of 15,000 cells per cm2, where TGF-β signaling should be active, and treated with neutralizing antibody for a period of 14 days. An increase in epithelial markers with an accompanying decrease in mesenchymal markers was observed at both the transcript (Fig. 7A) and protein (Fig. 7B) level, measured by qPCR and immunocytochemistry, respectively, indicating enhanced uptake of epithelial characteristics. We also observed an increase in proliferation as measured by imaging of EdU incorporation, confirming that suppression of TGF-β signaling led to enhanced proliferation (Fig. 7B). Taken together, our data are consistent with the hypothesis that TGF-β signaling serves to impede epithelial phenotype acquisition in RPE and that its inhibition can lead to enhanced uptake of epithelial identity.
Figure 7.
Inhibition of TGF-β signaling promotes retinal pigment epithelium phenotype. (A): Quantitative polymerase chain reaction-based measurement of transcript expression of a panel of epithelial (BEST1, PMEL, LRAT, MERTK, RPE65, and RLBP1) and mesenchymal (GREM1) markers in RPE seeded at 5,000 cells per cm2 and treated with 10 μg/ml anti-TGF-β antibody (1D11) for a period of 14 days. Data are normalized to expression of vehicle control. HPRT1 is used as a housekeeping gene. Bars represent mean + SD (n = 3). p < .05 (Student’s t test). (B): Representative images showing immunocytochemistry for indicated markers along with their quantification in cells seeded at 5,000 cells per cm2 and exposed to 10 µg/ml anti-TGF-β for 14 days. Images have been captured at ×10 magnification. Abbreviations: BEST1, bestrophin 1; EdU, 5-ethynyl-2-deoxyuridine; GREM1, gremlin 1; LRAT, lecithin retinol acyltransferase; MERTK, Mer receptor tyrosine kinase; PMEL17, premelanosome protein 17; RLBP1, retinaldehyde-binding protein 1; TGF-β, transforming growth factor-β.
Discussion
Stem cells provide a potentially unlimited source of cells for regenerative therapy in a range of human disorders, including retinal diseases such as age-related macular degeneration. AMD is the leading cause of blindness in people over the age of 65 in the developed world. It is a progressive disease caused by death and dysfunction of the retinal pigment epithelium, which in turn leads to photoreceptor loss. The specific loss of the RPE monolayer, combined with other advantages of targeting this particular tissue (immune privileged niche, ease of accessibility, separation from systemic circulation, well established methods for diagnosis, and monitoring progression) suggests AMD as a prime candidate for exploring replacement of the damaged RPE monolayer with stem cell-derived healthy RPE as a therapy [35–38].
The RPE monolayer, formed in the early embryo, is a terminally differentiated cell sheet that develops with a center to periphery gradient such that only the peripheral cells retain low levels of proliferation, whereas the remainder remains nonproliferative throughout life [39, 40]. However, in culture, their normal quiescence can be released, resulting in a re-entry into the cell cycle accompanied by proliferation. This results in transdifferentiation, in which cells deviate from an epithelial state and instead express hallmarks of a fusiform, mesenchymal state. This is followed by a mesenchymal-epithelial transition regulated by the transcription factor FOXM1, which results in re-epithelialization [18]. However, this process is density-dependent, where cells seeded at an appropriately high density successfully acquire an epithelial phenotype. Cultures seeded at low density fail to attain the epithelial state and instead continue to remain mesenchymal. Taking advantage of the critical seeding density-dependent nature of the culture, we wanted to gain further understanding of the signaling pathways that control the transition and acquisition of the epithelial phenotype. This would allow us to interrogate pathways of interest and hence promote epithelial fate achievement and increased RPE yield.
In the first instance, we have shown that manipulating cAMP levels significantly impacts the reattainment of the RPE phenotype in a time- and density-dependent manner. Published literature suggested that cAMP exerts direct effects on RPE phenotypes, such as pigmentation and phagocytosis, while inhibiting RPE proliferation [25, 26]. We show that elevating cAMP levels in dissociated RPE cells through FSK treatment or use of the cell-permeant cAMP analog dbcAMP increases acquisition of RPE phenotype through increasing proliferation. Remarkably, global gene expression profiling suggests that dbcAMP supplementation promotes RPE epithelialization to mimic a doubling of seeding density compared with cells without supplementation. This is of particular relevance in terms of RPE yield while manufacturing cells for a clinical application, for example, culturing transplantation-ready cells under GMP-compatible conditions for a clinical trial. In such a case, modulation of cAMP signaling through the use of dbcAMP could equate to a 100% increase in cell yield and consequently affect cost and feasibility of the cell-replacement therapy approach.
It is important to appreciate that, instead of uncontrolled proliferation, our data are consistent with the interpretation that in vitro RPE culture is a homeostatic system in which cell contact signals the cells to terminate proliferation and achieve quiescence [40]. This is distinct from contexts where increased proliferation could lead to potentially detrimental impacts, for example, when RPE cell monolayers are not anchored and cells at the edge of a monolayer could proliferate and acquire a mesenchymal identity without these critical inhibitory signals from other RPE cells. An ocular pathology where this occurs is PVR, where proliferation of cell types including RPE, together with the appearance of a mesenchymal, migratory RPE state, results in the formation of a fibrotic, contractile scar tissue, which inhibits vision [41]. Interestingly, one of the key pathways that has been implicated in PVR formation and progression is TGF-β signaling [42, 43]. TGF-β acts as a potent driver of fibrosis progression through the induction of the epithelial-mesenchymal transition, in which epithelial cells acquire mesenchymal phenotypes, leading to enhanced motility and invasion [44].
Given the precedence of TGF-β in pathogenesis of PVR, we were intrigued that analysis of gene expression upon cAMP elevation identified a potential role of this pathway in epithelial fate acquisition. We chose to investigate this in the context of a phenotypic screening approach in which RPE seeded at low density, which would normally not epithelialize, was challenged with a focused library of small molecules targeting signaling pathways of interest, including, but not limited to, TGF-β signaling. We then looked for those molecules that could promote acquisition of the epithelial phenotype at low density and retrospectively used their known target activities to inform on the signaling pathway that could be involved. To our knowledge, this study provides the first example of use of hESC-derived RPE cells for high-throughput phenotypic screening, successfully miniaturized to a 384-well format, and highlights that this clinically applied cell type is amenable to such interrogations. This approach led us to confirm the role of the TGF-β pathway, specifically signaling through the ALK5 receptor, as being key to RPE epithelial fate acquisition. TGF-β signaling controls a plethora of cellular responses and plays an important role in animal development [28, 45, 46]. Several studies in diverse cell types show that activation of TGF-β signaling leads to inhibition of cell proliferation and an increase in migration and uptake of mesenchymal characteristics [47, 48]. Polarized, functional RPE express and secrete low levels of TGF-β ligands to maintain the normal physiology of the subretinal space. Pathological conditions such as PVR lead to loss of polarity, which manifests in a high level of TGF-β secretion and loss of epithelial integrity and function [49, 50]. Our results suggest that inhibition of the TGF-β pathway via ALK5 signaling with small molecules effectively promotes the attainment of the epithelial state of RPE in a proliferation-dependent manner, which potentially allows more favorable cell-cell contacts and promotes epithelialization in a manner akin to that of cAMP. Additionally, use of multiple compound chemotypes, as well as recapitulating the same results with an orthogonal, antibody-based neutralization method, further lends support to the key role of TGF-β signaling. Based on our data, which show that TGF-β signaling is downregulated in high-density cultures, we propose that use of ALK5 inhibitors will promote proliferation and yield when used in cultures at low density, where this signaling pathway is active. This would translate to a bona fide increase in RPE yield because these cultures would normally not re-epithelialize in the absence of these compounds.
Further work is needed to fully characterize the activity of molecules and to understand their potency and specificity for the various TGF-β receptor subtypes in a head-to-head comparison. It also remains to be tested how cAMP elevation mechanistically inhibits the TGF-β pathway to potentiate RPE re-epithelialization. It is possible that increased cAMP signaling leads to protein kinase A activation, which in turn antagonizes the ras/mitogen-activated protein kinase kinase/extracellular signal-regulated kinase (ras/MEK/ERK ) signaling cascade, leading to attenuation of TGF-β signaling, as has been described in other cellular models [51–55]. This hypothesis requires further investigation to be validated. Another area that warrants further exploration is to test whether additional pathways that control TGF-β signaling (e.g., the Hippo and Crumbs pathway, which are key sensors of cell density [56] and hence may be relevant to cell contact dependent RPE epithelialization) also play a role in this process. This will help to fully comprehend how proliferation-dependent cell-cell contacts signal the initiation of the epithelialization program and how a network of signaling pathways feeds into it.
Conclusion
To transition to a successful and efficacious cell replacement therapy, stem cell-derived RPE need to be produced at scale with high yield and under GMP conditions. Presence of transdifferentiated RPE exhibiting mesenchymal characteristics that develop upon seeding cultures at low density can potentially compromise the functionality of the cell population. Given the intricate link between seeding density and epithelial fate outcome, the use of small molecule tools, for example, dbcAMP or inhibitors of TGF-β signaling as demonstrated in this report, could support large-scale manufacturing of functional RPE with high yield, which would be advantageous for successful clinical translation and therapeutic output.
Supplementary Material
Acknowledgments
We thank Maria Isabel Rosa, Juliette Steer, Anna Wilbrey, and Julie Kerby for help with RPE foci dissection and for initial workup of RPE culture; Mary-Ann Madsen and Christos Michaeledes for initial work with forskolin; Lawrence Welch for help with qPCR; Anna Ashton and members of the Pfizer Drug Safety Research and Development group for optimizing methods for RPE characterization; and Nicole E. Bodycombe for help with deconvolution of screening data.
Author Contributions
P.C.: conception and design, collection and/or assembly of data, data analysis and interpretation, manuscript writing, final approval of manuscript; A.G., R.I.S., and R.M.O.: data analysis and interpretation; E.I.: collection and/or assembly of data; P.J.W.: financial support, final approval of manuscript; M.B.: conception and design, data analysis and interpretation, final approval of manuscript; C.L.B.: conception and design, manuscript writing, final approval of manuscript.
Disclosure of Potential Conflicts of Interest
P.C., A.G., E.I., R.I.S., R.M.O., P.J.W., M.B., and C.L.B. are employees of Pfizer Ltd. P.C. and P.J.W. are co-inventors on a patent titled, “Method for Producing Retinal Pigment Epithelium Cells” (U.S. Pub. No. US2015/0159134 A1).
References
- 1.Strauss O. The retinal pigment epithelium in visual function. Physiol Rev. 2005;85:845–881. doi: 10.1152/physrev.00021.2004. [DOI] [PubMed] [Google Scholar]
- 2.da Cruz L, Chen FK, Ahmado A, et al. RPE transplantation and its role in retinal disease. Prog Retin Eye Res. 2007;26:598–635. doi: 10.1016/j.preteyeres.2007.07.001. [DOI] [PubMed] [Google Scholar]
- 3.Lindekleiv H, Erke MG. Projected prevalence of age-related macular degeneration in Scandinavia 2012-2040. Acta Ophthalmol. 2013;91:307–311. doi: 10.1111/j.1755-3768.2012.02399.x. [DOI] [PubMed] [Google Scholar]
- 4.Nazari H, Zhang L, Zhu D, et al. Stem cell based therapies for age-related macular degeneration: The promises and the challenges. Prog Retin Eye Res. 2015;48:1–39. doi: 10.1016/j.preteyeres.2015.06.004. [DOI] [PubMed] [Google Scholar]
- 5.Leach LL, Clegg DO. Concise review: Making stem cells retinal: Methods for deriving retinal pigment epithelium and implications for patients with ocular isease. Stem Cells. 2015;33:2363–2373. doi: 10.1002/stem.2010. [DOI] [PubMed] [Google Scholar]
- 6.Maruotti J, Sripathi SR, Bharti K, et al. Small-molecule-directed, efficient generation of retinal pigment epithelium from human pluripotent stem cells. Proc Natl Acad Sci USA. 2015;112:10950–10955. doi: 10.1073/pnas.1422818112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Vugler A, Carr AJ, Lawrence J, et al. Elucidating the phenomenon of HESC-derived RPE: Anatomy of cell genesis, expansion and retinal transplantation. Exp Neurol. 2008;214:347–361. doi: 10.1016/j.expneurol.2008.09.007. [DOI] [PubMed] [Google Scholar]
- 8.Maeda T, Lee MJ, Palczewska G, et al. Retinal pigmented epithelial cells obtained from human induced pluripotent stem cells possess functional visual cycle enzymes in vitro and in vivo. J Biol Chem. 2013;288:34484–34493. doi: 10.1074/jbc.M113.518571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Buchholz DE, Hikita ST, Rowland TJ, et al. Derivation of functional retinal pigmented epithelium from induced pluripotent stem cells. Stem Cells. 2009;27:2427–2434. doi: 10.1002/stem.189. [DOI] [PubMed] [Google Scholar]
- 10.Idelson M, Alper R, Obolensky A, et al. Directed differentiation of human embryonic stem cells into functional retinal pigment epithelium cells. Cell Stem Cell. 2009;5:396–408. doi: 10.1016/j.stem.2009.07.002. [DOI] [PubMed] [Google Scholar]
- 11.Strunnikova NV, Maminishkis A, Barb JJ, et al. Transcriptome analysis and molecular signature of human retinal pigment epithelium. Hum Mol Genet. 2010;19:2468–2486. doi: 10.1093/hmg/ddq129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Liao JL, Yu J, Huang K, et al. Molecular signature of primary retinal pigment epithelium and stem-cell-derived RPE cells. Hum Mol Genet. 2010;19:4229–4238. doi: 10.1093/hmg/ddq341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Klimanskaya I, Hipp J, Rezai KA, et al. Derivation and comparative assessment of retinal pigment epithelium from human embryonic stem cells using transcriptomics. Cloning Stem Cells. 2004;6:217–245. doi: 10.1089/clo.2004.6.217. [DOI] [PubMed] [Google Scholar]
- 14.Guha S, Baltazar GC, Tu LA, et al. Stimulation of the D5 dopamine receptor acidifies the lysosomal pH of retinal pigmented epithelial cells and decreases accumulation of autofluorescent photoreceptor debris. J Neurochem. 2012;122:823–833. doi: 10.1111/j.1471-4159.2012.07804.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Lu B, Malcuit C, Wang S, et al. Long-term safety and function of RPE from human embryonic stem cells in preclinical models of macular degeneration. Stem Cells. 2009;27:2126–2135. doi: 10.1002/stem.149. [DOI] [PubMed] [Google Scholar]
- 16.Lund RD, Wang S, Klimanskaya I, et al. Human embryonic stem cell-derived cells rescue visual function in dystrophic RCS rats. Cloning Stem Cells. 2006;8:189–199. doi: 10.1089/clo.2006.8.189. [DOI] [PubMed] [Google Scholar]
- 17.Plaza Reyes A, Petrus-Reurer S, Antonsson L, et al. Xeno-free and defined human embryonic stem cell-derived retinal pigment epithelial cells functionally integrate in a large-eyed preclinical model. Stem Cell Rep. 2016;6:9–17. doi: 10.1016/j.stemcr.2015.11.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Choudhary P, Dodsworth BT, Sidders B, et al. A FOXM1 dependent mesenchymal-epithelial transition in retinal pigment epithelium cells. PLoS One. 2015;10:e0130379. doi: 10.1371/journal.pone.0130379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Croze RH, Buchholz DE, Radeke MJ, et al. ROCK inhibition extends passage of pluripotent stem cell-derived retinal pigmented epithelium. Stem Cells Translational Medicine. 2014;3:1066–1078. doi: 10.5966/sctm.2014-0079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Della Fazia MA, Servillo G, Sassone-Corsi P. Cyclic AMP signalling and cellular proliferation: Regulation of CREB and CREM. FEBS Lett. 1997;410:22–24. doi: 10.1016/s0014-5793(97)00445-6. [DOI] [PubMed] [Google Scholar]
- 21.Kokkinaki M, Abu-Asab M, Gunawardena N, et al. Klotho regulates retinal pigment epithelial functions and protects against oxidative stress. J Neurosci. 2013;33:16346–16359. doi: 10.1523/JNEUROSCI.0402-13.2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kojima A, Nakahama K, Ohno-Matsui K, et al. Connexin 43 contributes to differentiation of retinal pigment epithelial cells via cyclic AMP signaling. Biochem Biophys Res Commun. 2008;366:532–538. doi: 10.1016/j.bbrc.2007.11.159. [DOI] [PubMed] [Google Scholar]
- 23.Koh SM. VIP enhances the differentiation of retinal pigment epithelium in culture: From cAMP and pp60(c-src) to melanogenesis and development of fluid transport capacity. Prog Retin Eye Res. 2000;19:669–688. doi: 10.1016/s1350-9462(00)00010-0. [DOI] [PubMed] [Google Scholar]
- 24.Hecquet C, Lefevre G, Valtink M, et al. cAMP inhibits the proliferation of retinal pigmented epithelial cells through the inhibition of ERK1/2 in a PKA-independent manner. Oncogene. 2002;21:6101–6112. doi: 10.1038/sj.onc.1205765. [DOI] [PubMed] [Google Scholar]
- 25.Hackett S, Friedman Z, Campochiaro PA. Cyclic 3′,5′-adenosine monophosphate modulates retinal pigment epithelial cell migration in vitro. Arch Ophthalmol. 1986;104:1688–1692. doi: 10.1001/archopht.1986.01050230126047. [DOI] [PubMed] [Google Scholar]
- 26.Smith-Thomas LC, Richardson PS, Rennie IG, et al. Influence of pigment content, intracellular calcium and cyclic AMP on the ability of human retinal pigment epithelial cells to contract collagen gels. Curr Eye Res. 2000;21:518–529. [PubMed] [Google Scholar]
- 27.Huang W, Wang L, Yuan M, et al. Adrenomedullin affects two signal transduction pathways and the migration in retinal pigment epithelial cells. Invest Ophthalmol Vis Sci. 2004;45:1507–1513. doi: 10.1167/iovs.03-0731. [DOI] [PubMed] [Google Scholar]
- 28.Massagué J. TGFβ signalling in context. Nat Rev Mol Cell Biol. 2012;13:616–630. doi: 10.1038/nrm3434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Carr AJ, Vugler A, Lawrence J, et al. Molecular characterization and functional analysis of phagocytosis by human embryonic stem cell-derived RPE cells using a novel human retinal assay. Mol Vis. 2009;15:283–295. [PMC free article] [PubMed] [Google Scholar]
- 30.Osakada F, Ikeda H, Mandai M, et al. Toward the generation of rod and cone photoreceptors from mouse, monkey and human embryonic stem cells [published correction appears in Nat Biotechnol 2008;26:352] Nat Biotechnol. 2008;26:215–224. doi: 10.1038/nbt1384. [DOI] [PubMed] [Google Scholar]
- 31.Chehrehasa F, Meedeniya AC, Dwyer P, et al. EdU, a new thymidine analogue for labelling proliferating cells in the nervous system. J Neurosci Methods. 2009;177:122–130. doi: 10.1016/j.jneumeth.2008.10.006. [DOI] [PubMed] [Google Scholar]
- 32.Chindelevitch L, Ziemek D, Enayetallah A, et al. Causal reasoning on biological networks: Interpreting transcriptional changes. Bioinformatics. 2012;28:1114–1121. doi: 10.1093/bioinformatics/bts090. [DOI] [PubMed] [Google Scholar]
- 33.Zhang Y, Handley D, Kaplan T, et al. High throughput determination of TGFβ1/SMAD3 targets in A549 lung epithelial cells. PLoS One. 2011;6:e20319. doi: 10.1371/journal.pone.0020319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Dasch JR, Pace DR, Waegell W, et al. Monoclonal antibodies recognizing transforming growth factor-beta. Bioactivity neutralization and transforming growth factor beta 2 affinity purification. J Immunol. 1989;142:1536–1541. [PubMed] [Google Scholar]
- 35.Brandl C, Grassmann F, Riolfi J, et al. Tapping stem cells to target AMD: Challenges and prospects. J Clin Med. 2015;4:282–303. doi: 10.3390/jcm4020282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Schwartz SD, Hubschman JP, Heilwell G, et al. Embryonic stem cell trials for macular degeneration: A preliminary report. Lancet. 2012;379:713–720. doi: 10.1016/S0140-6736(12)60028-2. [DOI] [PubMed] [Google Scholar]
- 37.Schwartz SD, Regillo CD, Lam BL, et al. Human embryonic stem cell-derived retinal pigment epithelium in patients with age-related macular degeneration and Stargardt’s macular dystrophy: Follow-up of two open-label phase 1/2 studies. Lancet. 2015;385:509–516. doi: 10.1016/S0140-6736(14)61376-3. [DOI] [PubMed] [Google Scholar]
- 38.Kamao H, Mandai M, Okamoto S, et al. Characterization of human induced pluripotent stem cell-derived retinal pigment epithelium cell sheets aiming for clinical application. Stem Cell Rep. 2014;2:205–218. doi: 10.1016/j.stemcr.2013.12.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Al-Hussaini H, Kam JH, Vugler A, et al. Mature retinal pigment epithelium cells are retained in the cell cycle and proliferate in vivo. Mol Vis. 2008;14:1784–1791. [PMC free article] [PubMed] [Google Scholar]
- 40.Kokkinopoulos I, Shahabi G, Colman A, et al. Mature peripheral RPE cells have an intrinsic capacity to proliferate; a potential regulatory mechanism for age-related cell loss. PLoS One. 2011;6:e18921. doi: 10.1371/journal.pone.0018921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Pastor JC, Rojas J, Pastor-Idoate S, et al. Proliferative vitreoretinopathy: A new concept of disease pathogenesis and practical consequences. Prog Retin Eye Res. 2016;51:125–155. doi: 10.1016/j.preteyeres.2015.07.005. [DOI] [PubMed] [Google Scholar]
- 42.Nassar K, Grisanti S, Tura A, et al. A TGF-β receptor 1 inhibitor for prevention of proliferative vitreoretinopathy. Exp Eye Res. 2014;123:72–86. doi: 10.1016/j.exer.2014.04.006. [DOI] [PubMed] [Google Scholar]
- 43.Hoerster R, Muether PS, Vierkotten S, et al. Upregulation of TGF-ss1 in experimental proliferative vitreoretinopathy is accompanied by epithelial to mesenchymal transition. Graefes Arch Clin Exp Ophthalmol. 2014;252:11–16. doi: 10.1007/s00417-013-2377-5. [DOI] [PubMed] [Google Scholar]
- 44.Chen X, Xiao W, Wang W, et al. The complex interplay between ERK1/2, TGFβ/Smad, and Jagged/Notch signaling pathways in the regulation of epithelial-mesenchymal transition in retinal pigment epithelium cells. PLoS One. 2014;9:e96365. doi: 10.1371/journal.pone.0096365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Derynck R, Muthusamy BP, Saeteurn KY. Signaling pathway cooperation in TGF-β-induced epithelial-mesenchymal transition. Curr Opin Cell Biol. 2014;31:56–66. doi: 10.1016/j.ceb.2014.09.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Leask A, Abraham DJ. TGF-beta signaling and the fibrotic response. FASEB J. 2004;18:816–827. doi: 10.1096/fj.03-1273rev. [DOI] [PubMed] [Google Scholar]
- 47.Dunfield LD, Nachtigal MW. Inhibition of the antiproliferative effect of TGFbeta by EGF in primary human ovarian cancer cells. Oncogene. 2003;22:4745–4751. doi: 10.1038/sj.onc.1206617. [DOI] [PubMed] [Google Scholar]
- 48.Takehara K, LeRoy EC, Grotendorst GR. TGF-beta inhibition of endothelial cell proliferation: Alteration of EGF binding and EGF-induced growth-regulatory (competence) gene expression. Cell. 1987;49:415–422. doi: 10.1016/0092-8674(87)90294-7. [DOI] [PubMed] [Google Scholar]
- 49.Hirsch L, Nazari H, Sreekumar PG, et al. TGF-β2 secretion from RPE decreases with polarization and becomes apically oriented. Cytokine. 2015;71:394–396. doi: 10.1016/j.cyto.2014.11.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Holtkamp GM, De Vos AF, Peek R, et al. Analysis of the secretion pattern of monocyte chemotactic protein-1 (MCP-1) and transforming growth factor-beta 2 (TGF-beta2) by human retinal pigment epithelial cells. Clin Exp Immunol. 1999;118:35–40. doi: 10.1046/j.1365-2249.1999.01016.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Davis MI, Ronesi J, Lovinger DM. A predominant role for inhibition of the adenylate cyclase/protein kinase A pathway in ERK activation by cannabinoid receptor 1 in N1E-115 neuroblastoma cells. J Biol Chem. 2003;278:48973–48980. doi: 10.1074/jbc.M305697200. [DOI] [PubMed] [Google Scholar]
- 52.Cook SJ, McCormick F. Inhibition by cAMP of Ras-dependent activation of Raf. Science. 1993;262:1069–1072. doi: 10.1126/science.7694367. [DOI] [PubMed] [Google Scholar]
- 53.Dumaz N, Marais R. Integrating signals between cAMP and the RAS/RAF/MEK/ERK signalling pathways. Based on the anniversary prize of the Gesellschaft für Biochemie und Molekularbiologie Lecture delivered on 5 July 2003 at the Special FEBS Meeting in Brussels. FEBS J. 2005;272:3491–3504. doi: 10.1111/j.1742-4658.2005.04763.x. [DOI] [PubMed] [Google Scholar]
- 54.Xie L, Law BK, Chytil AM, et al. Activation of the Erk pathway is required for TGF-beta1-induced EMT in vitro. Neoplasia. 2004;6:603–610. doi: 10.1593/neo.04241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Mulder KM. Role of Ras and Mapks in TGFbeta signaling. Cytokine Growth Factor Rev. 2000;11:23–35. doi: 10.1016/s1359-6101(99)00026-x. [DOI] [PubMed] [Google Scholar]
- 56.Varelas X, Samavarchi-Tehrani P, Narimatsu M, et al. The Crumbs complex couples cell density sensing to Hippo-dependent control of the TGF-β-SMAD pathway. Dev Cell. 2010;19:831–844. doi: 10.1016/j.devcel.2010.11.012. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.