Abstract
Missing data commonly occur in large epidemiologic studies. Ignoring incompleteness or handling the data inappropriately may bias study results, reduce power and efficiency, and alter important risk/benefit relationships. Standard ways of dealing with missing values, such as complete case analysis (CCA), are generally inappropriate due to the loss of precision and risk of bias. Multiple imputation by fully conditional specification (FCS MI) is a powerful and statistically valid method for creating imputations in large data sets which include both categorical and continuous variables. It specifies the multivariate imputation model on a variable-by-variable basis and offers a principled yet flexible method of addressing missing data, which is particularly useful for large data sets with complex data structures. However, FCS MI is still rarely used in epidemiology, and few practical resources exist to guide researchers in the implementation of this technique. We demonstrate the application of FCS MI in support of a large epidemiologic study evaluating national blood utilization patterns in a sub-Saharan African country. A number of practical tips and guidelines for implementing FCS MI based on this experience are described.
Keywords: Missing data, multiple imputation, fully conditional specification, complete case analysis, blood utilization
1. INTRODUCTION
Missing data are a pervasive problem in large epidemiologic studies. Incomplete data may arise due to refusal, attrition, measurement errors and miscommunication. Missing data result in a loss of precision and are also a source of bias if observations are not missing completely at random (MCAR) [1–3]. The most widely adopted strategy for dealing with missing data is to omit observations having missing values and perform a complete case analysis (CCA). In certain circumstances (e.g. when there is less than 5% missingness and the missing is MCAR), CCA may be an acceptable approach. In practice, however, these circumstances rarely occur [4]. The cumulative effect of missing data in several variables often leads to exclusion of a substantial proportion of the original sample, which in turn causes a substantial loss of precision and power. CCA may suffer from a loss of information in the incomplete cases and risk of bias if the missing data are not MCAR. More general objections to CCA are that it lacks an underlying statistical rationale and that it is difficult to determine when CCA will yield reasonable results [5]. Other method, like single imputation (SI), simply replaces the missing value with either a mean value or another appropriate value from a similar unit or “neighbor,” to create a ‘complete’ data set [3,4]. SI underestimates the uncertainty introduced by imputation, which may cause the generation of inappropriately small variances and potentially biased estimates. None of these above ad hoc approaches is statistically valid in general and they can lead to serious bias.
Statistical methods for addressing missing values have been actively pursued in recent years, including maximum likelihood (ML) estimation [6], Bayesian estimation [7] and multiple imputation (MI) [8], all of which are based on the assumption that data are missing at random (MAR) [9]. These approaches are especially useful when the data contain many patterns of missing values, or when both categorical and continuous random variables are involved. However, MI is the only technique that is computationally straightforward, versatile, relatively easy to apply, and increasingly available in standard statistical software, including SAS PROC MI and R MICE (Multiple Imputation by Chained Equations) package [10]. For general missing data patterns, there are two major iterative methods for doing multiple imputation: the joint modeling (JM) and the fully conditional specification (FCS) method [11]. Joint modeling is based on the assumption of joint multivariate normality of all variables, which implies that valid imputations may be generated by linear regression equations. It is ill-suited for imputing categorical variables since it assumes normality and linearity [12, 13]. FCS relaxes that assumption and is rapidly emerging as a commonly used method for handling missing data [14–16]. FCS MI specifies the multivariate imputation model on a variable-by-variable basis by a set of conditional densities, one for each incomplete variable. This permits a great deal of flexibility, since an appropriate regression model can be selected for each variable (e.g. linear regression for continuous variables, logistic regression for categorical variables) [10, 17]. Simulation studies provide evidence that FCS MI generally yields estimates that are unbiased and provide appropriate coverage [11, 18]. However, FCS is still rarely used in epidemiology, perhaps in part because relatively little practical guidance is available for implementing and evaluating this method. Only few studies have looked at practical questions about how to implement MI in large data sets used for diverse purposes [19–21].
The present study aims to provide an introduction to FCS MI with a focus on practical aspects and challenges in using this method for dealing with multivariate missing data. We introduce the basic concepts and general methodology, and provide detailed guidance based on our experience with a large epidemiologic study evaluating national blood utilization patterns in Namibia, a country in southern Africa. Studying blood utilization patterns is essential for forecasting and predicting future blood stock requirements. However, broader analyses which evaluate blood utilization at a national level are lacking [22]. To our knowledge, this study is the first multi-year evaluation of national blood component use in an African country [23].
2. METHOD
2.1. Blood Transfusion Service of Namibia (NAMBTS) Nationally Representative Census
NAMBTS is the only organization authorized to collect, process and distribute blood and blood components intended for transfusion in Namibia. Clinical and demographic data from 46 transfusion centers were reviewed for a four year period from August 1, 2007 through July 31, 2011. A total of 39,313 blood requests (each representing a transfusion event) were submitted to NAMBTS during the study period [23]. Since these data are primarily used for billing purposes, clinical and patient demographic variables captured on the paper-based blood request form (e.g., diagnosis, age and sex) were sometimes, but not always, entered into a national electronic database. To standardize the analysis, data on diagnoses reported by clinicians were matched to broad diagnostic categories in the WHO International Classification of Disease (ICD-10) system. As shown in Table 1, records were 100% complete for location and date, as well as number and type of blood component ordered. However, 23.2%, 19.6% and 9.9% of records were missing for Diagnosis, Age and Gender, respectively. Due to the cumulative effect of missing data in these variables, 32.4% of the total 39,313 blood requests had at least one missing value. To create a full national census for the four year study period, and to minimize bias due to any systematic differences between complete records and those with missing data, FCS MI was performed to impute estimated values. Based on the imputed datasets obtained by this method, we further analyzed blood utilization patterns stratified by diagnosis, gender and age, and developed a unique portrait of blood use in Namibia.
Table 1.
Frequency Analysis of Missing Variables in Original NAMBTS Data During the Study Period: X Observed; • Missing
| Missing data pattern | 2007/2008 | 2008/2009 | 2009/2010 | 2010/2011 | Grand Total | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diagnosis | Age | Gender | n (events) | % | n (events) | % | n (events) | % | n (events) | % | n (events) | % |
| X | X | X | 5825 | 66.19 | 7060 | 70.86 | 7418 | 70.53 | 6286 | 62.66 | 26589 | 67.63 |
| X | X | • | 67 | 0.76 | 37 | 0.37 | 6 | 0.06 | 17 | 0.17 | 127 | 0.32 |
| X | • | X | 818 | 9.30 | 1031 | 10.35 | 875 | 8.32 | 586 | 5.84 | 3310 | 8.42 |
| X | • | • | 61 | 0.69 | 42 | 0.42 | 16 | 0.15 | 38 | 0.38 | 157 | 0.40 |
| • | X | X | 1080 | 12.27 | 819 | 8.22 | 1048 | 9.96 | 1875 | 18.69 | 4822 | 12.27 |
| • | X | • | 19 | 0.22 | 7 | 0.07 | 9 | 0.09 | 15 | 0.15 | 50 | 0.13 |
| • | • | X | 226 | 2.57 | 169 | 1.70 | 255 | 2.42 | 437 | 4.36 | 1087 | 2.76 |
| • | • | • | 704 | 8.00 | 798 | 8.01 | 891 | 8.47 | 778 | 7.76 | 3171 | 8.07 |
| Grand Total Events | 8800 | 100% | 9963 | 100% | 10518 | 100% | 10032 | 100% | 39313 | 100% | ||
Note: A transfusion event is defined as any patient record in which at least 1 type of blood component is ordered for an individual patient. Total numbers of each type of blood component unit associated with each transfusion event are established and stratified by component type and by year [23].
2.2. Imputation
Assumptions
The risk of bias due to missing data depends on the reasons why data are missing. Three types of missing data are commonly classified: missing completely at random (MCAR), missing at random (MAR), and missing not at random (MNAR). MCAR indicates that the probability of an observation being missing does not depend on the value of any variables under study, which is a fairly strong assumption and tends to be relatively rare. MAR indicates the probability of missing depend only on the subset of complete cases, and is less restrictive than MCAR. Most MI methods, including FCS MI, generally assume that the data is at least MAR, and therefore remains valid if observations are MCAR.
MNAR indicates the probability that a missing value is associated with the missing variable itself and with other variables. It can be difficult to determine whether variables are MNAR, because the information that would confirm that values are MNAR is unobserved. As a result, the decision to treat data as MNAR is often made based on theoretical and/or substantive information, rather than information present in the data itself. Therefore, biases caused by data that are MNAR can be addressed only by sensitivity analyses examining the effect of different assumptions about the missing data mechanism.
Algorithms
The key step of the MI procedure is the specification of the imputation model. Two general approaches for imputing multivariate missing data have emerged: joint modeling (JM) and FCS. JM involves specifying a multivariate distribution for the missing data, and drawing imputation from their conditional distributions by Markov Chain Monte Carlo (MCMC) techniques [10]. This methodology is attractive if the multivariate distribution is a reasonable description of the data. However, in practice, the data often consists of variables with different scales, and quite complex relations between variables may occur that are hard to capture in an explicitly specified joint distribution for the entire data. Instead of drawing the imputations from a pre-specified joint distribution, FCS imputations are generated sequentially by specifying an imputation model for each variable given the other variables. Let Y be the partially observed complete sample, consisting of p variables, from the multivariate distribution P(Y|θ). Further, let Y−j be all variables in the data except Yj, j = 1, …, p. We assume that the multivariate distribution of Y is completely specified by θ, a vector of unknown parameters. The posterior distribution of θ is obtained by iteratively sampling from conditional distributions of the form
The parameters θ1,…, θp are specific to the respective conditional densities and are not necessarily the product of a factorization of the “true’ joint distribution P(Y|θ). FCS starts with an initial imputation and draws imputations by iterating over the conditional densities and sequentially filling in the current draws of each variable. The tth iteration of the Gibbs sampler is
where is the imputed value for the variable j at the tth iteration [24]. After the cycle reaches convergence, the current draws are taken as the first set of imputed values. The cycle is then repeated until the desired number of imputations has been achieved.
FCS MI Using SAS PROC MI
SAS PROC MI performs the imputation stage and can be used with either monotone or arbitrary missing patterns. The FCS statement is a new addition to the PROC MI in SAS version 9.3. This procedure does not start with a specified multivariate posterior distribution of observed data, but instead uses a separate conditional distribution of each imputed variable. It is attractive because of its ability to impute both continuous and categorical variables appropriately. It can also incorporate features such as the specification of upper or lower bounds for variables, and a rounding option for imputed values. The general coding procedure for PROC MI using FCS statement is shown in Supporting Materials.
The discriminant function, logistic regression, regression, and predictive mean matching methods are available in the FCS statement. For continuous variables, the regression (REG) and predictive mean matching (REGPPM) methods can be used to impute missing values. The logistic regression (LOGISTIC) method can be used for variables having binary or ordinal responses, and the discriminant function (DISCRIM) method is used for variables having binary or nominal responses.
Generally the imputation model should include all the variables likely to be used in the subsequent analyses [25]. For the imputation of a particular variable, the model should include variables in the complete-data model, variables that are correlated with the imputed variable, and variables that are associated with the missingness of the imputed variable. The dependent variable(s) must be included in the imputation model. Otherwise the imputed values will not have the same relationship to the dependent variable that the observed values do. Typically m = 4–20 imputations are created, resulting in 4–20 “complete” imputed data sets, though more are computationally feasible and better characterize the variability introduced into the results due to the imputation process [26]. In our case, the FCS statement included a specific modelling approach to impute missing values for both continuous (Age) and categorical variables (Diagnosis and Gender) with arbitrary missing patterns.
Imputation Diagnostics
Once the imputation model has been specified and the initial imputations created, the quality of imputations should be examined. Graphic and numeric diagnostics are commonly used for identifying problematic variables and detecting possible implausible values [27]. Imputations can be checked by using a standard of reasonability: the differences between the observed and imputed values, and the distribution of the completed data as a whole, can be checked to see whether they make sense in the context of the problem being studied. These diagnostics are applied to one randomly selected completed data set constructed by FCS imputations and then repeated with another one to confirm if similar results are obtained. Kernel density estimate plots are used to visually compare the distributions of the observed, imputed and completed values of each variable. When there is large number of variables, it may be difficult to carefully examine graphical summaries of each variable. Numerical summaries that compare differences in means and standard deviations are an additional approach to identifying problematic variables and may be more feasible within the context of large datasets. For numeric diagnostic, nonparametric Kolmogorov-Smirnov (KS) test is used to numerically compare the marginal distribution and test statistically significant differences (p-value). When p-value is less than 0.05, we would reject the hypothesis that there is no significant difference between two empirical distributions. Imputation diagnostics should be used to identify potentially problematic variables. Then information regarding the missingness, along with substantive knowledge, can be used to determine whether the imputations are in fact reasonable or whether the procedure needs to be further modified [19].
FCS MI Analysis
The m imputations are intended to represent a plausible range of values that approximate the missing value, had it not been missing. The variability of values within this range allows the uncertainty in the imputation process to be quantified and integrated into the analysis. Each of the m “complete” data sets is analyzed using a standard analytic method that will estimate the quantities of scientific interest. Results on each data set will vary due to the difference in values during the multiple imputations. Then the estimates from the imputed data sets are combined or pooled to generate a single set of estimates. The overall estimate is the average of the estimates. The variance of that overall estimate is a function of variance within each imputed data set and the variance across the data sets:
The strength of MI is that any analysis model can be applied to the imputed data sets. In SAS, the command PROC MIANALYZE is used to combine results across datasets automatically. This make the analysis of imputed datasets no more complicated than running a single regression in a single dataset. In our case, multinomial logistic regression model was applied to each imputed data set to compute the conditional proportions (p) and 95% Confidence Interval (CI) for three blood unit types (RBC, Platelet and FFP). Finally the results were combined using PROC MIANALYZE to give the valid estimates. FCS MI results in statistically valid estimates with confidence intervals that account for the uncertainty caused by the missing data as well as the sampling error of the estimates using CCA [14].
Comparison of FCS MI and CCA
MI is widely advocated as an improvement over CCA. However, it is often implemented without adequate consideration of whether it offers any advantage over CCA for the research question of interest, or whether potential gains may be offset by bias from a poorly fitting imputation model, particularly as the amount of missing data increases. For these reasons, it has been recommended to carry out a CCA in parallel when using MI for handling missingness [5]. Where CCA and MI analysis give different results, the analyst should attempt to understand why, and this should be reported in publications.
3. RESULTS
Table 1 shows the frequency analysis of three missing variables (Diagnosis, Age and Gender) in original NAMBTS data sample. Due to the cumulative effect of missing data in three variables, 32.4% of total blood requests had at least one missing value. FCS MI was then performed to handle missing data and create a full four year NAMBTS national census. The imputation model included all the variables likely to be used in the subsequent analyses to ensure that all of the information in the large dataset was used. The imputation number was chosen as 20 and finally 20 complete data sets were obtained.
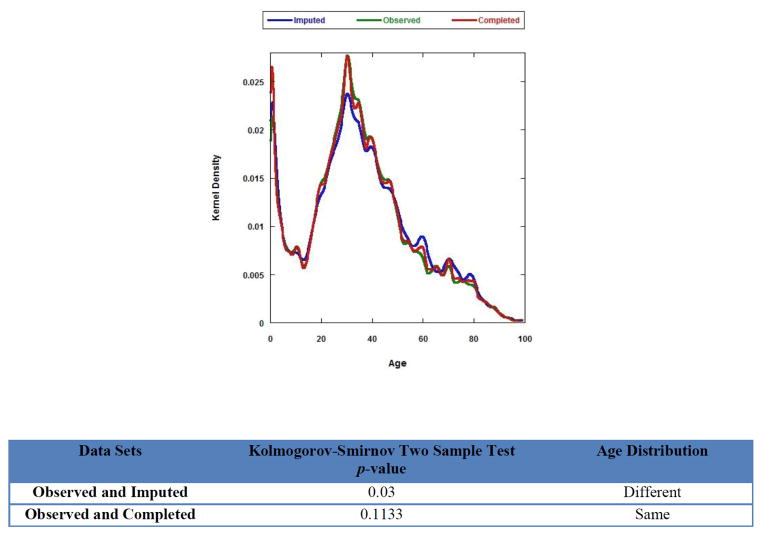
To examine the quality of imputation, one imputed data set was selected randomly for imputation diagnostics, and the missing variable Age was used as an example for imputation diagnostics. As shown in Figure 1, the shape of the Age distribution of the imputed values (blue line) differed from that of the observed values (green line), while the distribution between the observed (green line) and the completed data (red line) was quite similar. Numeric diagnostics were further applied to numerically compare the empirical distributions of Age in the observed, imputed and completed data and test the statistically significant differences (p-value). We could identify quickly the differences of Age distribution among three different data sets. Figure 2 shows a simple example if we choose a ‘good’ or ‘bad’ imputation model, what will happen in imputation diagnostics. Since the Age distribution is not normal, REGPMM model is chosen as an appropriate model (blue line) for imputing Age, instead of using REG model (red line) which assumes normality. For imputing Diagnosis and Gender which both had nominal responses, thus the discriminant function (DISCRIM) method was used to impute these categorical variables.
Figure 1.
Imputation diagnostics (Graphic and Numeric).
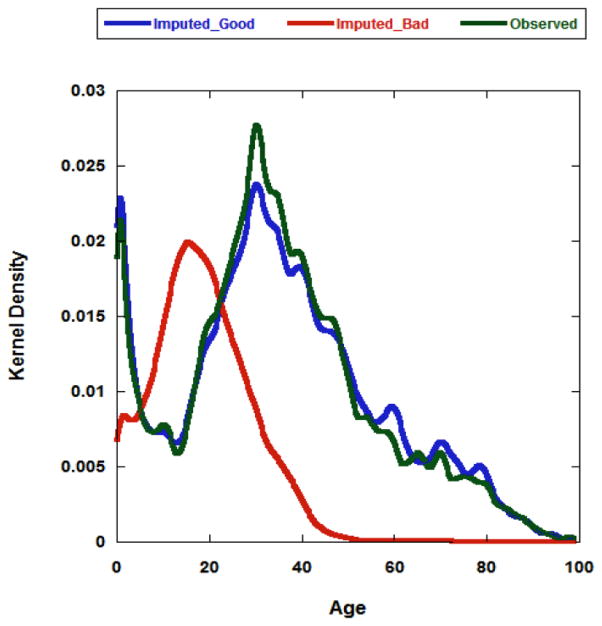
Figure 2.
Comparison of imputation models: Blue line represents the Age distribution of imputed data by an appropriate REGPMM model (Imputed_Good); Red line represents the Age distribution of imputed data by an inappropriate REG model (Imputation_Bad); Green line represents the Age distribution of the observed data set.
To evaluate the blood utilization pattern, the conditional proportion (p) of each blood unit type (RBC, Platelet and FFP) was computed to develop a unique portrait of blood use in Namibia. Table 2 showed the counts of each type of blood component unit associated with each transfusion event, stratified by component types. A total of 39,313 events accounted for 91,389 blood component units. 91,389 and 60,632 total blood component units were counted from FCI MS and CCA respectively. Table 3 (Supporting Tables 4, 5) showed the total number of RBC (FFP and platelet) units requested during the study period were further stratified by diagnosis, age and gender (Supporting Tables 4 and 5 shown in Supporting Materials). The predominant four ICD categories associated with three blood component units were listed for comparison. For example, Table 3 showed the top four diagnoses in the “Diseases of the blood and blood-forming organs and certain disorders involving the immune mechanism (D50-D89)”, “Infectious Disease (A00-B99)”, “Pregnancy (O00-O99)”, and “Gastrointestinal (K20-K93)” accounted for 38.9%, 14.8%, 11.1% and 6.1% of RBC units issued, respectively. The remaining 30% of units issued were associated with 15 other ICD categories, none of which individually accounted for more than 5% of all units, and six of which accounted for <1% of all units. These 15 other ICD categories were classified as “All others”. Studying blood utilization patterns is essential for forecasting and predicting future blood stock requirements and it may help set realistic national blood collection goals [23].
Table 2.
Comparison of Total Blood Component Unit Counts by FCS MI and CCA
| FCS MI | CCA | ||||
|---|---|---|---|---|---|
| Component Type | n (units)* | % | 95% CI | n (units) | % |
| RBC | 78,660 | 86.1 | (85.8, 86.3) | 52,284 | 86.2 |
| FFP | 9,751 | 10.7 | (10.5, 10.9) | 6,082 | 10.0 |
| Platelets | 2,978 | 3.3 | (3.1, 3.4) | 2,266 | 3.7 |
| Total Units | 91,389 | 100.0 | – | 60,632 | 100.0 |
Mean value from 20 imputed data sets.
Table 3.
RBC Utilization by Diagnosis (ICD Category), Age and Gender: A. FCS MI; B. CCA
| A. FCS MI | Male | Female | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ICD category | 0–14 years | 15–49 years | 50+ years | 0–14 years | 15–49 years | 50+ years | Totals | |||||||
| % | 95% CI | % | 95% CI | % | 95% CI | % | 95% CI | % | 95% CI | % | 95% CI | n (units) | % | |
| D. Diseases of the blood and blood-forming organs and certain disorders involving the immune mechanism (D50-D89) | 2.3 | (2.1,2.4) | 9.2 | (8.9,9.4) | 4.5 | (4.3,4.7) | 1.9 | (1.7,2) | 16.3 | (15.9,16.7) | 4.8 | (4.6,5.1) | 30,616 | 38.9 |
| A/B. Infectious disease (A00-B99) | 0.6 | (0.5,0.6) | 4.2 | (4,4.4) | 1.6 | (1.4,1.7) | 0.5 | (0.5,0.6) | 6.4 | (6.1,6.7) | 1.5 | (1.3,1.7) | 11,648 | 14.8 |
| O. Pregnancy (O00-O99) | 0 | (0.0) | 0 | (0,0) | 0 | (0,0) | 0.3 | (0.2,0.4) | 9.9 | (9.6,10.2) | 0.8 | (0.5,1.2) | 8,702 | 11.1 |
| K. Gastrointestinal (K20-K93) | 0.1 | (0,0.1) | 1.8 | (1.7,1.9) | 1.6 | (1.5,1.8) | 0.1 | (0,0.1) | 1.2 | (1.1,1.4) | 1.3 | (1.1,1.4) | 4,796 | 0.1 |
| All others | 1.8 | (1.7,1.9) | 6.9 | (6.6,7.2) | 4.8 | (4.6,5.1) | 1.3 | (1.2,1.4) | 9.2 | (8.9,9.5) | 5.1 | (4.7,5.6) | 22,898 | 29.1 |
| 78,660 | 100.0 | |||||||||||||
| B. CCA | Male | Female | ||||||||||||
| ICD category | 0–14 years | 15–49 years | 50+ years | 0–14 years | 15–49 years | 50+ years | Totals | |||||||
| % | % | % | % | % | % | n (units) | % | |||||||
| D. Diseases of the blood and blood-forming organs and certain disorders involving the immune mechanism (D50-D89) | 2.3 | 9.0 | 4.3 | 1.7 | 16.5 | 4.4 | 19,984 | 38.2 | ||||||
| A/B. Infectious disease (A00-B99) | 0.5 | 4.6 | 1.5 | 0.4 | 6.7 | 1.2 | 7,790 | 14.8 | ||||||
| O. Pregnancy (O00-O99) | 0 | 0 | 0 | 0.0 | 12.0 | 0.1 | 6,348 | 12.1 | ||||||
| K. Gastrointestinal (K20-K93) | 0.1 | 2.0 | 1.7 | 0.1 | 1.1 | 1.2 | 3,205 | 6.1 | ||||||
| All Others | 2.0 | 7.2 | 5.1 | 1.3 | 9.0 | 4.0 | 14,9572 | 28.6 | ||||||
| 52,284 | 100.0 | |||||||||||||
4. DISCUSSION
To obtain valid inferences or statistical estimates of interest from imputed data, imputation should preserve the structure in the data, as well as any uncertainty about this structure, and account for any reasons related to the process that generated the missing data. FCS MI specifies the multivariate imputation model on a variable-by-variable basis by a set of conditional densities, one for each incomplete variable. It is particularly appealing in settings in which a number of variables have missing data, some of which are continuous and some of which are categorical [26, 28].
Checking of imputation models is necessary because it can identify model defects and facilitate model improvement. Some deviations of the observed and imputed data can be expected under MAR (Figure 1). But that is not necessarily a problem because the distributions should be similar only if the data are MCAR. In fact, these differences may be indicative of the bias that imputation is trying to address. However, we can see dramatic distribution differences between observed and imputed data if the ‘bad’ REG model is chosen for imputing Age (Figure 2). This large difference in imputation diagnostics is a sign for a potential problem, meaning further assessment of the imputation model is required. The best practice may be to repeat the analysis under different imputation models to see if, and how, changes in the imputation model result in changes in the final results. Choosing a good imputation model is important since the quality of the imputation model will influence the quality of the final results.
Once the imputation model has been specified and the initial imputations created, imputation diagnostics are commonly used for identifying problematic variables and detecting possible implausible values. More complex imputation diagnostics method can be found [27], in which residuals from regression models were used to determine which differences in distribution were reasonable. Methods for addressing imputation diagnostics are an area of on-going statistical research. Further research is needed to incorporate the imputation diagnostics directly into common MI software packages [29].
Comparing results from FCS MI and CCA may provide clues about the nature of the data. It also provides reassurance if inference from the two are similar, but may highlight issues with one or both approaches if results differ substantially. As shown in Table 3 (Supporting Tables 4, 5), FCS MI and CCA were conducted to estimate the conditional proportions for three blood unit uses (RBC, Platelet and FFP) stratified by Diagnosis, Age and Gender. While resorting to complete cases is simple, CCA suffers from a loss of information in the incomplete cases and risk of bias if the missing data is not MCAR. FCS MI may reduce bias in estimates while accounting for the uncertainty in the imputation process, preserving study power and holding less restrictive but more plausible MAR assumption.
Although attractive, FCS MI is not without drawbacks [17]. First, FCS MI is based on the assumption of MAR. For missing data which is MNAR, new methods generating MIs under MNAR model will be required for handing such kind of missing data [30, 31], which is out of the interest of this study. Another way is to preclude MNAR data to MAR by changing the study design. For example, when MNAR attrition is anticipated, we could ask one more question at each occasion of measurement for each participant, “How likely are you to drop out this study before next session?” Collecting this additional covariate and including it in the imputation model will effectively convert an MNAR situation to MAR [3]. Thus, FCS MI can still be used. Second, for FCS MI, each conditional density has to be specified separately, so substantial modeling effort can be needed for data sets with many variables. Third, FCS MI lacks the theoretical justification of some other well developed imputation approaches like MCMC. Relatively little is known about the quality of the resulting imputations because the implied joint distributions may not exist theoretically and that convergence criteria are ambiguous [11].
In conclusion, FCS MI is a powerful and statistically valid method for creating imputations in large data sets with complex data structures. This paper provides a detailed guidance for using FCS MI method to deal with multivariate missing data in large data sets, with the aim of helping researchers to implement and use this method for their own data.
Supplementary Material
Acknowledgments
The authors thank Dr. Steven J. Gutreuter and Dr. John P. Pitman of the CDC Division of Global HIV/AIDS for contributions on the statistical methods. The authors also thank Prof. Yichuan Zhao of Georgia State University and Prof. Xu Zhang of University of Mississippi Medical Center for their helpful discussion and comments. This project has been supported by the President’s Emergency Plan for AIDS Relief (PEPFAR) through the Centers for Disease Control and Prevention.
Footnotes
DISCLAIMER
This manuscript has been gone through and cleared by CDC eClearance system. The findings and conclusions in this manuscript are those of the author(s) and do not necessarily represent the official position of the Centers for Disease Control and Prevention.
The supporting materials can be downloaded from the journal website along with the article.
References
- 1.Little RJ, Rubin DB. Statistical analysis of missing data. 2. Hoboken: John Wiley & Sons; 2002. http://dx.doi.org/10.1002/9781119013563. [Google Scholar]
- 2.He YL. Circ Cardiovasc Qual Outcomes. 2010;3:98–105. doi: 10.1161/CIRCOUTCOMES.109.875658. http://dx.doi.org/10.1161/CIRCOUTCOMES.109.875658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Pigott TD. Educ Res Eval. 2001;7(4):353–83. http://dx.doi.org/10.1076/edre.7.4.353.8937. [Google Scholar]
- 4.Graham JW. Annu Rev Psychol. 2009;60:549–76. doi: 10.1146/annurev.psych.58.110405.085530. http://dx.doi.org/10.1146/annurev.psych.58.110405.085530. [DOI] [PubMed] [Google Scholar]
- 5.White IR, Carlin JB. Statist Med. 2010;29:2920–31. doi: 10.1002/sim.3944. http://dx.doi.org/10.1002/sim.3944. [DOI] [PubMed] [Google Scholar]
- 6.Enders CK. Struct Equ Modelling. 2001;8:128–41. http://dx.doi.org/10.1207/S15328007SEM0801_7. [Google Scholar]
- 7.Oba S, Sato M, Takemasa I, Monden M, Matsubara K, Ishii S. Bioinformatics. 2003;19:2088–96. doi: 10.1093/bioinformatics/btg287. http://dx.doi.org/10.1093/bioinformatics/btg287. [DOI] [PubMed] [Google Scholar]
- 8.Patrician PA. Res Nurs Health. 2002;25:76–84. doi: 10.1002/nur.10015. http://dx.doi.org/10.1002/nur.10015. [DOI] [PubMed] [Google Scholar]
- 9.Newgard CD, Haukoos JS. Acad Emerg Med. 2007;14:669–78. doi: 10.1197/j.aem.2006.11.038. [DOI] [PubMed] [Google Scholar]
- 10.Buuren SV, Groothuis-Oudshoorn CG. J Stat Softw. 2011;45:1–67. [Google Scholar]
- 11.Buuren SV, Brand JP, Groothuis-Oudshoorn CG, Rubin DB. J Stat Comput Sim. 2006;76:1049–64. http://dx.doi.org/10.1080/10629360600810434. [Google Scholar]
- 12.Azur MJ, Stuart EA, Frangakis C, Leaf PJ. Int J Mehtods Psychiatr Rec. 2011;20:40–9. doi: 10.1002/mpr.329. http://dx.doi.org/10.1002/mpr.329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Bernaards CA, Belin TR, Schafer JL. Statist Med. 2007;26:1368–82. doi: 10.1002/sim.2619. http://dx.doi.org/10.1002/sim.2619. [DOI] [PubMed] [Google Scholar]
- 14.Schafer Joseph L, Graham John W. Psychol Methods. 2002;7:147–77. http://dx.doi.org/10.1037/1082-989X.7.2.147. [PubMed] [Google Scholar]
- 15.Sterne JA, White IR, Carlin JB, Spratt M, Royston P, Kenward MG, Wood AM, Carpenter JR. BMJ. 2009;339:157–60. doi: 10.1136/bmj.b2393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Horton NJ, Kleinman KP. Am Stat. 2007;61:79–90. doi: 10.1198/000313007X172556. http://dx.doi.org/10.1198/000313007X172556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lee KJ, Carlin JB. Am J Epidemiol. 2010;171:624–32. doi: 10.1093/aje/kwp425. http://dx.doi.org/10.1093/aje/kwp425. [DOI] [PubMed] [Google Scholar]
- 18.Lee KJ, Carlin JB. Emerg Themes Epidemiol. 2012;9:1–10. doi: 10.1186/1742-7622-9-3. http://dx.doi.org/10.1186/1742-7622-9-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Stuart EA, Azur M, Frangakis C, Leaf P. Am J Epidemiol. 2009;169:1133–9. doi: 10.1093/aje/kwp026. http://dx.doi.org/10.1093/aje/kwp026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.He Y, Zaslavsky AM, Landrum MB, Harrington DP, Catalano P. Stat Methods Med Res. 2010;19:653–70. doi: 10.1177/0962280208101273. http://dx.doi.org/10.1177/0962280208101273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Schenker N, Raghunathan TE, Chiu PL, Makuc DM, Zhang GY, Cohen AJ. J Amer Statist Assoc. 2006;101:924–33. http://dx.doi.org/10.1198/016214505000001375. [Google Scholar]
- 22.Meza BPL, Lohrke B, Wilkinson R, Pitman JP, Shiraishi RW, Lowrance DW, Kuehnert MJ, Mataranyika M, Basavaraju SV. Blood Transfus. 2014;12(3):352–61. doi: 10.2450/2013.0143-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Pitman JP, Wilkinson R, Liu Y, Finckenstein B, Sibinga CS, Lowrance DW, Marfin AA, Postma M, Mataranyika M, Basavaraju SV. Transfus Med Rev. 2015;29:45–51. doi: 10.1016/j.tmrv.2014.11.003. http://dx.doi.org/10.1016/j.tmrv.2014.11.003. [DOI] [PubMed] [Google Scholar]
- 24.Carlin BP, Louis TA. Bayesian methods for data analysis. 3. New York, NY: Springer Verlag; 2008. [Google Scholar]
- 25.Glynn RJ, Laird NM, Rubin DB. J Amer Statist Assoc. 1993;88:984–93. http://dx.doi.org/10.1080/01621459.1993.10476366. [Google Scholar]
- 26.Buuren SV. Stat Methods Med Res. 2007;16:219–42. doi: 10.1177/0962280206074463. http://dx.doi.org/10.1177/0962280206074463. [DOI] [PubMed] [Google Scholar]
- 27.Abayomi K, Gelman A, Levy M. Appl Statist. 2008;57:273–91. http://dx.doi.org/10.1111/j.1467-9876.2007.00613.x. [Google Scholar]
- 28.Bartlett JW, Seaman SR, White IR, Carpenter JR. Multiple imputation of covariates by fully conditional specification: Accommodating the substantive model. Stat Meth Med Res. 2014 doi: 10.1177/0962280214521348. http://dx.doi.org/10.1177/0962280214521348. [DOI] [PMC free article] [PubMed]
- 29.Yucel RM. J STAT SOFTW. 2011;45:1–7. doi: 10.18637/jss.v045.i01. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Dziuraa JD, Posta LA, Zhao Q, Fu ZX, Peduzzi P. Yale J Biol Med. 2013;86:343–58. [PMC free article] [PubMed] [Google Scholar]
- 31.Héraud-Bousquet V, Larsen C, Carpenter J, Desenclos JC, Strat YL. BMC Med Res Methodol. 2012;12:1–11. doi: 10.1186/1471-2288-12-73. http://dx.doi.org/10.1186/1471-2288-12-73. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.