Abstract
Cancer pharmacogenomics is an evolving landscape and has the potential to significantly impact cancer care and precision medicine. Harnessing and understanding the genetic code of both the patient (germline) and the tumor (somatic) provides the opportunity for personalized dose and therapy selection for cancer patients. While germline DNA is useful in understanding the pharmacokinetic and pharmacodynamic disposition of a drug, somatic DNA is particularly useful in identifying drug targets and predicting drug response. Molecular profiling of somatic DNA has resulted in the current breadth of targeted therapies available, expanding the armamentarium to battle cancer. This review provides an update on cancer pharmacogenomics and genomics-based medicine, challenges in applying pharmacogenomics to the clinical setting, and patient perspectives on the use of pharmacogenomics to personalize cancer therapy.
Keywords: oncology, personalized, pharmacogenetics, germline, somatic, DNA, biomarker
Introduction
The 21st century has been a major milestone in the advancement of cancer therapies. With the unveiling of the first human genome in 2003 to the discovery of thousands of cancer biomarkers through efforts such as The Cancer Genome Atlas, these practice-changing initiatives have resulted in the development of dozens of next-generation targeted therapies, which have improved survival for cancer patients.1 The importance of genomics-based medicine was further highlighted by President Obama in January 2015 through the Precision Medicine Initiative, launched to advance pharmacogenomics, identify new targets for treatment and disease prevention, and to lay the scientific foundation for precision medicine for many diseases, including cancer.2
The transition from development of standard cytotoxic chemotherapies to highly targeted agents and immunotherapies has resulted in the current breadth of treatment options available.3 Further, increased numbers of targeted therapies are receiving accelerated drug approval alongside companion diagnostic assays, which are critical in identifying predictive biomarkers that allow for a personalized approach to therapy selection.4 A highly focused attack on targetable driver mutations has not only resulted in superior response rates and overall survival (OS) compared to traditional, nontargeted chemotherapy but has also allowed for more rapid time to drug approval, ensuring timely access of life-prolonging drugs to cancer patients in dire need of more options. More recently, there has been a transition in cancer treatment to increased utility of immunotherapies and various combinations of cytotoxic chemotherapies/targeted agents with immunotherapies.5 This ever expanding armamentarium of cancer therapies means greater reliance on biomarkers to help clinicians decide which therapies a particular patient may benefit from.
Pharmacogenetics – the study of how genes influences drug response – provides the opportunity to stratify patients into those likely to respond or not respond to therapy, or those likely to experience or not experience toxicity.6 The term “pharmacogenomics” is commonly used in the literature to define the broader field of genomics and genome-wide associations with drug response. Genomics cancer research provides the ability to analyze both the patient’s (germline) and the tumor’s (somatic) DNA. While germline DNA is readily obtained via a blood sample or buccal swab, somatic DNA is primarily obtained via tumor biopsy and is therefore a more invasive collection procedure and is subject to sample selection. Clinically relevant (inherited) germline variations in the host may be valuable in determining the pharmacokinetic disposition of drugs and drug response (in addition to identification of disease-susceptibility genetic variants), while somatic mutations found within the tumor are acquired and are particularly useful in assessing the pharmacodynamic effects of a drug and ultimately tumor response.4
Novel next-generation sequencing methods with lower costs and reduced turnaround time have provided the ability to screen for hundreds to thousands of mutations within germline and somatic DNA. These multiplex methods are likely to replace single-gene testing, given the excessive costs with performing multiple single-gene tests and the clinical utility of having access to larger panels up front to guide decision-making downstream.7 As a result, our expanding knowledge of cancer genomics has resulted in a transition from characterizing tumors solely by anatomical location and histology to consideration of molecular profiles. This is evident by several tumor-agnostic, biomarker-driven clinical trials, such as NCI-MATCH, American Society of Clinical Oncology’s TAPUR, Southwest Oncology Group’s LUNG-MAP, and many others.8
This review aims to summarize updates in cancer pharmacogenomics and genomics-based medicine, including the use of both germline and somatic DNA to personalize cancer therapy, challenges in applying clinical pharmacogenomics to standard practice, and patient perspectives on the use of pharmacogenomics to guide cancer therapy.
Biomarkers for determining toxicity and treatment response
In 1998, the National Institutes of Health Biomarkers Definitions Working Group defined a biomarker as “a characteristic that is objectively measured and evaluated as an indicator of normal biological processes, pathogenic processes, or pharmacologic responses to a therapeutic intervention”.9 A pharmacogenetic or pharmacogenomic biomarker is defined as any “genetic or genomic marker that is associated with drug response”.3
Biomarkers can be categorized into two broad types: prognostic and predictive. A prognostic biomarker is a marker, or measurable trait, that provides information on the likely course of cancer, including aggressiveness of disease, regardless of treatment. Widely recognized examples include gene expression arrays such as the 70-gene profile MammaPrint™10 or 21-gene profile Oncotype Dx11 for estrogen/progesterone receptor-positive, lymph node-negative breast cancer and microsatellite instability (MSI) in colorectal cancer patients.12,13 MammaPrint™ (Agendia, Inc., Irvine, CA, USA) and Oncotype Dx assist in determining the risk of breast cancer recurrence in women with early stage breast cancer and provide guidance as to which high-risk patients may require additional chemotherapy.11 While MSI and mutations within DNA repair genes can result in increased risk of developing colorectal cancer (eg, Lynch Syndrome), MSI-high (MSI-H) colorectal tumors also indicate a favorable prognosis compared to microsatellite stable/low-frequency MSI (MSS/MSI-L) tumors, independently of chemotherapy, in local and advanced colorectal cancer.12,13 More recently, data suggests that MSI-H colorectal cancers harbor a large mutational load, which may be predictive of response to immunotherapy agents both within and beyond colorectal cancer, suggesting MSI status can also serve as a predictive biomarker to drugs such as nivolumab and pembrolizumab.14 Biomarkers such as BRAF mutations are considered a poor prognostic biomarker conferring aggressive disease in colorectal cancer, while also considered a positive predictive biomarker in other diseases, such as melanoma, conferring increased response to BRAF inhibitors.15,16
A predictive biomarker is a marker, or measurable trait, that can be used to identify patients most likely to benefit from treatment and/or those predisposed to toxicity.3 Examples of clinically relevant germline and somatic predictive biomarkers for drug response/toxicity are discussed in the next section of the paper and are the primary focus of this review. Notably, some biomarkers may be characterized as both prognostic and predictive within the same tumor type, such as overexpression of HER2 in breast cancer, which without chemotherapy is considered a poor prognostic biomarker resulting in an aggressive phenotype; however, with the development of therapies targeting HER2 (eg, trastuzumab), this biomarker is also considered a positive predictive biomarker for therapy response. Table 1 provides a summary of cancer therapies with pharmacogenomic information in the Food and Drug Administration (FDA)-approved drug label, and Figure 1 provides an illustration of clinically relevant somatic mutations and drug targets in cancer.
Table 1.
Summary of oncology pharmacogenomic biomarkers in FDA drug labeling
| Disease | Biomarker | Therapy | Frequency |
|---|---|---|---|
| Breast | HER2 | Trastuzumab, lapatinib, pertuzumab, ado-trastuzumab emtansine | 20% |
| ESR1 | Exemestane, letrozole, anastrozole, fulvestrant, tamoxifen, | 60% | |
| Colorectal | KRAS | Cetuximab, panitumumab | 35%–40% |
| EGFR | Cetuximab, panitumumab | 35%–45% | |
| DPYD | 5-Fluorouracil, capecitabine | <5% | |
| UGT1A1 | Irinotecan | 30% | |
| Lung | ALK | Crizotinib, ceritinib | 5%–7% |
| EGFR | Erlotinib, gefitinib, afatinib, osimertinib | 15%–20% | |
| Melanoma | BRAF | Vemurafenib, dabrafenib, trametinib | 50%–60% |
| Acute promyelocytic leukemia | PML-RARα | Arsenic trioxide, tretinion | >95% |
| Chronic myeloid leukemia | BCR-ABL | Imatinib, dasatinib, nilotinib, bosutinib, ponatinib, omacetaxine mepesuccinate | >95% |
| UGT1A1 | Nilotinib | 30% | |
| Cutaneous T-cell lymphoma | CD-25/IL2RA | Denileukin diftitox | 75% |
| Chronic lymphocytic leukemia (CLL) | del(17p) | Ibrutinib | 3%–8% at diagnosis; up to 30% in refractory CLL |
| CD20/MS4A1 | Obinutuzumab, rituximab | 25% | |
| Acute lymphocytic leukemia | TPMT | 6-Mercaptopurine, thioguanine | <5% |
| Non-Hodgkin’s lymphoma | CD20/MS4A1 | Rituximab, tositumomab | >90% |
Note: Data from http://www.fda.gov/drugs/scienceresearch/researchareas/pharmacogenetics/ucm083378.htm.87
Abbreviations: DPYD, dihydropyrimidine dehydrogenase; UGT1A1, uridinediphosphate glucuronosyl transferase 1A1; ALK, anaplastic lymphoma kinase; CD, cluster of differentiation; del(17p), deletion 17p; TPMT, thiopurine-S-methyltransferase; FDA, Food and Drug Administration.
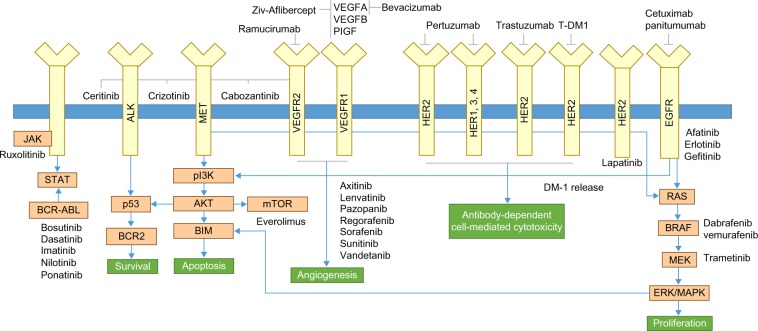
Figure 1.
Summary of somatic cancer biomarkers and targeted therapies.
Notes: This figure depicts examples of key signaling pathways and downstream effects of mutations within somatic biomarkers, and their respective targeted therapies.
Abbreviations: ALK, anaplastic lymphoma kinase; JAK, Janus kinase; mTOR, mammalian target of rapamycin; p13K, phosphoinositide 3-kinase; TDM-l, ado-trastuzumab emtansine; VEGFR, vascular endothelial growth factor receptor.
Examples of germline and somatic predictive biomarkers
ALK and EGFR in NSCLC
The abnormal fusion gene, EML4-ALK, resulting in constitutive activation of anaplastic lymphoma kinase (ALK), occurs in ~5% of all non-small-cell lung cancer (NSCLC) cases. The translocation activates various growth stimulatory signaling pathways, including RAS-MEK-ERK, janus kinase 3 (JAK3)-STAT3, and PI3K-AKT pathways.17
Initial Phase I trials demonstrated an overall response rate (ORR) of 60.8% in patients with ALK-positive NSCLC and treated with the targeted tyrosine kinase inhibitor (TKI), crizotinib, which competitively inhibits ALK.17 In contrast, the ORR in similar patients historically treated with standard chemotherapy was 15%–20%, resulting in accelerated FDA approval in 2011 alongside a companion diagnostic test to detect ALK positivity (Vysis ALK Break Apart fluorescence in-situ hybridization [FISH] Probe Kit). The confirmatory randomized Phase III trial comparing crizotinib versus docetaxel/pemetrexed in ALK-positive NSCLC clearly demonstrated improved ORR (65% vs 20%; P<0.05) and median progression-free survival (PFS) (7.7 vs 3.0 months; P<0.05) in patients randomized to receive crizotinib.18
Similar to most highly targeted therapies, the benefit of crizotinib is short-lived due to acquired resistance. Common mechanisms of resistance include EGFR, ROS1, KIT, MET, or secondary resistant ALK translocations not previously identified in the pretreatment tumor.19 Ceritinib, a highly potent second-generation ALK inhibitor, received accelerated FDA approval in 2014 for the treatment of ALK-positive metastatic NSCLC in patients progressing on or intolerant to crizotinib. A Phase I study identified ORRs of 58% and 56% in crizotinib naïve and resistant cases, respectively, suggesting strong activity in crizotinib-resistant tumors.20 In December 2015, another ALK inhibitor, alectinib, received accelerated FDA approval in patients with ALK-positive metastatic NSCLC who have progressed on or are intolerant to crizotinib, based on an ORR of 48% in patients progressing on crizotinib. Importantly, an ORR of 61% was identified in patients with brain metastases, lasting an average of 9.1 months.21
As evident by the approvals of crizotinib, ceritinib, and alectinib, the historically prolonged drug development paradigm is shifting to more rapid accelerated approval of exceedingly effective targeted therapies, even prior to the completion of confirmatory Phase III randomized trials. For example, only 4 years lapsed from the time of discovering ALK as a major oncogenic driver in NSCLC to the approval of the first targeted therapy (2007 to 2011),22 versus 13 years that lapsed from the time of discovering HER2 as an oncogenic driver in breast cancer to the approval of trastuzumab (1985 to 1998), the first HER2-targeted agent.23
Activating mutations in the EGFR gene also occurs in ~20% of NSCLC cases and results in constitutive signaling via the PI3K-AKT and MAPK pathways.24 Deletions in exon 19 and a missense mutation at exon 21 (arginine to leucine substitution [L858R]) account for >90% of all EGFR mutations and are considered a positive predictive biomarker for small-molecule TKIs targeting the EGFR domain (erlotinib, gefitinib, afatinib).25 In a biomarker clinical trial, Zhou et al randomized patients with EGFR-mutated NSCLC and evaluated first-line erlotinib versus standard platinum-based chemotherapy.26 Median PFS in erlotinib-treated patients was significantly greater than that noted in chemotherapy-treated patients (13.1 vs 4.6 months, hazard ratio [HR] 0.16, 95% confidence interval [CI] 0.10–0.26; P<0.0001). The ORR was 83% and 36% for erlotinib and chemotherapy-treated patients, respectively, unequivocally demonstrating the superiority of EGFR inhibitors versus standard chemotherapy in this population.26
A companion diagnostic test for erlotinib (cobas EGFR Mutation Test) was FDA-approved in 2013 along with expanded approval for first-line use in patients with EGFR-mutated metastatic NSCLC. Afatinib, a second-generation EGFR inhibitor, received FDA approval in 2013 for the first-line treatment of patients with EGFR-mutated NSCLC. Afatinib’s irreversible binding mechanism of action results in superior activity in resistant tumors that have progressed on erlotinib or gefitinib.27 Median PFS was significantly longer (11.1 months) in patients with EGFR-mutated NSCLC who received afatinib versus standard chemotherapy (cisplatin and pemetrexed) (6.9 months).28
The two primary mechanisms of resistance include MET amplification, which activates the MAPK signal transduction pathway, and a secondary point mutation in EGFR (T790M) that blocks the capacity for erlotinib to bind to and inhibit EGFR.29 The T790M mutation, occurring in approximately two-thirds of resistant cases, results in stronger affinity for ATP, reducing the ability of erlotinib, an ATP-competitive reversible EGFR inhibitor, to bind to the EGFR tyrosine kinase domain.30 Osimertinib (Tagrisso), a novel inhibitor of the EGFR (T790M) mutation, has resulted in ORRs >60% in patients who have progressed on prior EGFR inhibitor therapy and who express the resistant mutation.31 As a result, osimertinib received accelerated FDA approval in 2015, while another T790M inhibitor, rociletinib, is also currently being evaluated by the FDA for approval.
BRAF in melanoma
Approximately 50% of cutaneous metastatic melanoma cases carry mutations in BRAF, 90% of which results in glutamic acid substituted for valine at codon 600, termed V600E. Activating BRAF mutations result in constitutive activation and downstream signaling of the MAPK pathway, promoting cell proliferation and tumor growth. Vemurafenib, a potent inhibitor of BRAF V600E, received accelerated FDA approval in 2011 alongside a companion diagnostic test, cobas 4800 BRAF V600 Mutation Test.3 In the pivotal Phase III trial, 675 treatment-naïve patients with metastatic melanoma expressing the BRAF mutation were randomly assigned to receive dacarbazine or vemurafenib. The investigators identified a 63% reduced risk of death (HR 0.37; P<0.001) and a 74% reduced risk of disease progression in the vemurafenib arm compared to dacarbazine (HR 0.26; P<0.001).15
As noted with pervious targeted therapies, resistance eventually ensues in most patients with a mean time to diagnosis of first lesion at ~3 months and a median PFS of 6–8 months. The most commonly described resistance mechanisms include RAS and MEK mutations resulting in paradoxical activation of the MAPK pathway. Combining a BRAF inhibitor with a MEK inhibitor has been a logical approach to combat BRAF inhibitor resistance.32 In a Phase III trial, investigators randomly assigned 704 patients with BRAF V600E-positive metastatic melanoma to receive either a combination of dabrafenib (BRAF inhibitor) and trametinib (MEK inhibitor) or vemurafenib as first-line therapy. Results demonstrated an OS rate at 12 months of 72% in the combination group and 65% in the vemurafenib group (HR 0.69, 95% CI 0.53–0.89; P=0.005). The median PFS was 11.4 and 7.3 months in the combination and vemurafenib group, respectively (HR 0.56, 95% CI 0.46–0.69; P<0.001).33 Although dabrafenib and trametinib received approval as monotherapy in 2013, more recently (2015), these drugs received FDA approval to be used in combination to treat patients with BRAF-positive unresectable or metastatic melanoma.
DPYD and 5-FU
5-Fluorouracil (5-FU), an antimetabolite chemotherapeutic agent, has been the mainstay therapy of several gastrointestinal-related cancers, including colon, rectal, and gastric. Dihydropyrimidine dehydrogenase (DPD) is the rate-limiting enzyme responsible for the metabolism of ~80%–85% of 5-FU to the inactive 5,6-dihydrofluorouracil.3 To date, over 30 genetic variants in DPYD, the gene coding for DPD, may result in deficient DPD enzyme activity. The most common variant is a splice site mutation (IV14+1G>A [also referred to as DPYD*2A]) and is estimated to confer deficient DPD enzyme activity in 2%–3% of patients, ultimately increasing the risk of 5-FU-induced toxicity.34,35
In one of the largest genotyping studies conducted in 5-FU-treated colorectal cancer patients (n=2,594), investigators observed that the incidence of grade 3 or greater 5FU-related toxicities in DPYD*2A, I560S, and D949V carriers were 22/25 (88.0%), 2/4 (50.0%), and 22/27 (81.5%), respectively. In a multivariate model, DPYD*2A (odds ratio [OR] 15.21; P<0.001) and D949V (OR 9.10; P<0.001) were significantly associated with grade 3 or greater toxicities.36 In a large systematic review of DPYD and 5-FU toxicity (n=7,365 patients from eight studies), investigators used individual patient data to study the association of DPYD*2A, c.2846A>T, c.1679T>G, c.1236G.A/HapB3, and c.1601G>A with 5-FU-related toxicities. DPYD c.1679T>G (adjusted relative risk [RR] 4.40; P<0.0001) and c.1236G>A/HapB3 (adjusted RR 1.59; P<0.0001) were significantly associated with 5-FU-related toxicity, including gastrointestinal and hematological toxicity. DPYD*2A and c.2846A>T were also associated with severe toxicity (adjusted RR 2.85; P<0.0001 and 3.02; P<0.0001, respectively).37 In a prospective study, 2,038 patients treated with 5-FU were genotyped for DPYD*2A, of whom 22 carried at least one variant allele and were treated with approximately half the normal dose. The risk of grade ≥3 toxicity was reduced from 73% in historical controls (DPYD*2A carriers receiving standard full dose 5-FU) to 28% by genotype-guided dosing, while drug-induced death was reduced from 10% to 0%, respectively. Further, genotype-guided dosing demonstrated a similar incidence of grade ≥3 toxicity compared with wild-type patients receiving the standard dose (23%) and similar systemic drug exposure, suggesting adequate treatment at reduced doses. Although marginal, average treatment cost per patient was lower for screening (US$3,767) than for non-screening (US$3,828).38
Estimating DPD deficiency by quantifying DPD enzyme activity in peripheral blood mononuclear cells may be challenging, as standardized thresholds for deficient activity are not readily available or validated and different laboratories use different cutoffs. Further, there are cost and resource implications given the minor allele frequency is <1.0%, resulting in a high number of patients needed to screen to prove a beneficial impact on reducing toxicity. The FDA label includes a warning for increased risk of toxicity in patients with deficient DPD activity; however, the definition of what constitutes “deficient” is not defined, nor does the FDA mandate genetic testing for DPYD prior to administration of 5-FU. The Clinical Pharmacogenetics Implementation Consortium (CPIC) guidelines provide dosing recommendations of fluoropyrimidines by DPYD genotype/phenotype (https://www.pharmgkb.org/guideline/PA166109594): patients who are homozygous variant or who have complete DPD deficiency (~0.2% of patients) should receive an alternative drug, while patients who are heterozygous variant or who have intermediate enzyme activity (~3%–5% of the population) should initiate 5-FU with at least a 50% dose reduction followed by titration based on toxicity or pharmacokinetic levels.39
UGT1A1 and irinotecan
Irinotecan, a topoisomerase I inhibitor, has activity in many cancers, including gastrointestinal and lung carcinomas. Irinotecan is a prodrug and requires bioactivation via carboxylesterase enzymes to the active metabolite, SN-38, which is then glucuronidated and inactivated to SN-38G via uridine diphosphate glucuronosyl transferase 1A1 (UGT1A1).40 Polymorphic UGT1A1 variants lead to a reduction in gene expression and enzyme activity, resulting in supratherapeutic SN-38 levels and an increased risk of severe toxicity, primarily neutropenia.41 The UGT1A1*28 allele is the most common and found in ~39% of Caucasians, 43% of Africans, and 16% of Asians.41,42
A meta-analysis demonstrated a significant increase in toxicity for homozygous variant patients (UGT1A1*28/*28) at medium (150–250 mg/m2) and high doses (>250 mg/m2) (OR 3.2 and 27.8, respectively).43 Importantly, the standard irinotecan dose currently administered to metastatic colorectal cancer patients receiving FOLFIRI (5-FU, leucovorin, irinotecan) is 180 mg/m2 every 2 weeks, without genotyping. The FDA label states that the UGT1A1*28/*28 genotype is a risk factor for severe neutropenia and recommends consideration of an initial dose reduction; however, no specific dosing guidelines are provided. CPIC guidelines currently do not provide dosing recommendations based on UGT1A1 genotype for irinotecan, while the Dutch Pharmacogenetics Working Group44 suggests an initial dose reduction by 30% in *28/*28 patients if the starting recommended dose is >250 mg/m2 (https://www.pharmgkb.org/guideline/PA166104951).
More recently, there has been a shift from focusing on the increased risk of toxicity in UGT1A1*28/*28 patients to the potential under-dosing of *1/*1 and *1/*28 patients who are receiving standard irinotecan doses. In fact, a genotype-driven dose escalation study identified irinotecan maximum tolerated doses (MTD) to be 370 and 310 mg/m2 for *1/*1 and *1/*28 patients, respectively, suggesting higher doses than the standard 180 mg/m2 are tolerable in wild-type and heterozygous patients.45 A similar genotype-driven Phase I study demonstrated that the MTD in *1/*1, *1/*28, and *28/*28 patients was 390, 340, and 150 mg/m2, respectively.46 Interestingly, both studies suggests a dose–response relationship, where the ORR in patients treated at higher doses was significantly greater than in patients treated at lower doses (65%–67% vs 24%–25%, respectively).45,46 Further, the second study demonstrated a significant difference in median time to disease progression (16 vs 7 months, respectively; P=0.003).46 Given these Phase I studies were not powered to detect differences in response or survival, investigators are currently conducting a Phase II clinical trial to investigate whether higher irinotecan doses stratified by genotype results in prolonged PFS and OS (ClinicalTrials.gov; NCT02138617).
BCR-ABL and CML
Drug development of highly potent targeted therapies in hematologic malignancies was made promising by milestone discoveries that identified the Philadelphia chromosome, formed by a translocation between chromosomes 9 and 22 fusing the breakpoint cluster region (BCR) with the c-ABL oncogene, ultimately responsible for the development of chronic myeloid leukemia (CML).47 The BCR-ABL translocation occurs in > 95% of CML cases and results in constitutive activation of signal transduction pathways associated with cell proliferation and tumor growth.48 Imatinib was the first TKI discovered in a high-throughput screening assay for agents that inhibit the translocation and approved in 2001 to treat BCR-ABL-positive CML.
The large randomized Phase III trial, IRIS, included 5 years of follow-up in over 550 patients with BCR-ABL-positive CML treated with either imatinib at 400 mg/day or interferon alfa plus Ara-C.49 Patients randomized to receive imatinib had significantly improved rates of complete hematologic response (97% vs 56%, P<0.001), major and complete cytogenetic responses (85% and 74% vs 22% and 8%, respectively, P < 0.001), and discontinuation of assigned therapy due to intolerance (3% vs 31%), compared to patients who received interferon alfa plus Ara-C.49
Due to emerging resistance with initial imatinib therapy, subsequent TKIs with somewhat different mechanisms of action were developed over the past several years, including dasatinib, nilotinib, bosutinib, and ponatinib. Slight differences in mechanism allow for second- and third-generation TKIs to be used after progression on prior TKI therapy. Imatinib, nilotinib, and ponatinib all bind to the inactive conformation of the c-ABL kinase, preventing the conformational switch to the active form, while dasatinib inhibits both the active and inactive conformations. Nilotinib has a higher affinity for c-ABL kinase than imatinib, resulting in greater selectivity and potency.50 Approximately, 10%–15% of CML patients will present with resistance to imatinib and ~20%–25% will develop resistance over time (eg, 1–5 years).51 The T315I point mutation occurring in the c-ABL oncogene resulting in steric hindrance represents the highest magnitude of resistance as current TKIs require threonine at position 315 in order to bind to their targets.52 Ponatinib features a carbon–carbon triple bond, which limits steric hindrance, ultimately overcoming initial resistance.53
TPMT and 6-mercaptopurine
6-Mercaptopurine (6-MP) is one of the mainstay treatments for acute lymphocytic leukemia (ALL). It inhibits the formation and synthesis of purine ribonucleotides through the incorporation of thioguanine nucleotide analogs, resulting in cell death. Thiopurine-S-methyltransferase (TPMT) catalyzes the S-methylation and inactivation of thiopurine analogs. Approximately 5%–10% of the population carries at least one variant TPMT allele conferring reduced TPMT enzyme activity, while ~0.5% are homozygous resulting in no TPMT enzyme activity.54 TPMT*2 and *3 alleles account for over 90% of altered TPMT activity. The risk of myelosuppression, the dose-limiting toxicity of 6-MP, is significantly greater in patients who have increased thioguanine nucleotides secondary to defective TPMT activity.
One of the pivotal trials investigating TPMT genotype and phenotype with 6-MP tolerability enrolled 180 children with ALL treated with a 6-MP-based regimen. The investigators identified an inverse relationship between thioguanine nucleotide concentration and TPMT enzyme activity. Importantly, the percentage of wild-type, heterozygous, and homozygous-deficient patients who were able to tolerate the full 6-MP dose throughout therapy was 84%, 65%, and 7%, respectively (P<0.001),55 suggesting that TPMT is a critical marker of tolerating and completing 6-MP treatment. Interestingly, a larger study of 814 children with ALL identified that patients carrying one variant TPMT allele had significantly fewer rates of positive minimal residual disease (9%) versus patients who were homozygous wild-type (23%) (P=0.02), suggesting TPMT genotype influences drug exposure and resulting drug response, in addition to tolerability.56 In a genotype-guided dosing study, investigators identified the OR of severe myelosuppression in TPMT heterozygous children with ALL and receiving standard 6-MP dosages (ie, no dose adjustment for genotype) was four times that of wild-type children receiving standard 6-MP doses, whereas the OR for TPMT heterozygous children receiving a preemptive 30%–70% dose reduction was 1.30, similar to that seen in wild-type children receiving standard 6-MP doses.57
CPIC guidelines suggest an initial 90% dose reduction in homozygous variant patients and a 30%–70% dose reduction in heterozygous patients. Based on the FDA label for 6-MP, genetic testing prior to treatment is recommended, but not required, and no dose adjustments are provided in the drug label.58 Importantly, inter-patient tolerability of 6-MP is not completely explained by TPMT genotype alone, as a considerable number of patients with normal TPMT genotype still experience severe dose-limiting myelosuppression, compromising adequate treatment with 6-MP. Yang et al at St Jude Children’s Research Hospital identified a novel SNP in the gene NUDT15, which was significantly (and independently) associated with 6-MP tolerability (rs116855232; P=8.8×10−9).59 In vitro, NUDT15 was demonstrated to act as a protectant by eliminating oxidized purine nucleoside triphosphates to prevent incorporation into DNA. Importantly, the NUDT15 polymorphism was relatively common in East Asians (9.8%) and Hispanics (3.9%), but infrequent (0.2%) in Europeans, emphasizing the critical role ethnic diversity plays in pharmacogenomics.
Challenges in applying clinical pharmacogenomics
Frequency of mutations and cost-effectiveness
Utilizing genomic data to stratify patients according to targetable lesions can permit the use of smaller numbers of patients needed to treat (NNT) in a given clinical trial and increase statistical power for establishing effectiveness. Unfortunately, there is often the need to screen large numbers of patients to identify those with the genetic marker of interest. While some mutations occur at a relatively high frequency (ie, BCR-ABL in >95% of CML cases,48 C-KIT in 85% of gastrointestinal stromal tumors,60 BRAF in 50% of metastatic melanomas15), others occur at very low frequencies (ie, ALK in ~5% of NSCLCs, DPYD in 1% of the general population). The higher the NNT, the more difficult it is to prove cost-effectiveness of testing. In fact, a cost-effectiveness analysis demonstrated that ALK testing in stage IV nonsquamos NSCLC with subsequent crizotinib treatment for ALK-positive NSCLC was not cost-effective secondary to high drug costs and low biomarker frequency;61 however, given significantly enhanced response with targeted therapy in ALK-positive NSCLC, testing remains routine and standard of care.
Single gene testing is likely to be substituted by multiplex genomic characterization using next-generation sequencing technology, particularly as the cost of these technologies continue to decline.62 Obtaining a large genomic panel up front, in the same manner a physician orders a blood or chemistry panel, will allow for decisions to be made downstream, eliminating the barrier of turnaround time. Additionally, the more results available from a single run the more cost-effective the process will be. For example, testing a NSCLC patient for EGFR, ALK, and ROS as single, individual assays may cost just as much, if not more, than running a targeted whole exome panel of hundreds to thousands of genes, which may inevitably provide additional information about other treatment options.
Quantification of the economic impact of genomic-driven medicine as well as cost-effectiveness analyses of pharmacogenomic profiling is becoming more important. Despite the potential clinical utility of next-generation sequencing-based diagnostic tests, laboratories and providers continue to struggle to get reimbursed for such tests.63 The reimbursement landscape is difficult to navigate and places test developers in a tough situation as they must demonstrate clinical utility and cost-effectiveness to receive coverage, but demonstrating such utility requires running the test in the first place. As such, many test developers are offering the test at significantly reduced prices in hopes that this will generate enough data to convince payers to provide coverage for such tests down the road. Managing big data and tracking clinical outcomes for patients undergoing sequencing will be key to demonstrate clinical utility and cost-effectiveness.
Identifying an actionable target
Distinguishing between functional “driver mutations” and random, nonfunctional “passenger mutations” yields a challenge in selecting targeted therapies for intervention.64 Sequencing of matched primary and metastatic tumors has characterized patterns of “Darwinian-like” evolution, with acquisition of driver mutations that promote tumor progression.65,66 Since each pathway contains multiple genes, there are various combinations of driver mutations that can disrupt a pathway important for the dissemination of cancer.67 Common adult epithelial cancers (eg, breast, colorectal, prostate) often undergo five to seven rate-limiting events,68,69 and experimental studies have shown engineered changes in the function of at least five genes is required for transformation into malignant cancerous cells.70 Factors that may help distinguish between driver and passenger mutations include whether the mutation is silent or non-silent, the location or frequency of the mutation, and the depth of sequencing required to detect the mutation.
Intra-tumoral heterogeneity provides an additional challenge to tumor sequencing and therapy selection as cancers are rarely homogenous and contain large areas with a variable ratio of normal and cancerous cells.71 Gerlinger et al demonstrated that, among all somatic mutations found on multiregion sequencing in renal cell carcinomas and associated metastatic sites, 63%–69% were heterogenous and not detectable in every region. The investigators concluded that a single tumor biopsy specimen identifies only a minority of genetic aberrations present within an entire tumor, ultimately resulting in sampling bias;66 however, it is difficult to ascertain whether or not these results are generalizable to other tumor types. Methods to overcome this barrier include deeper sequencing (of up to 1,000-fold), measuring the effect of treatment at the biopsy site (ie, pretreatment and on-treatment biopsy to assess target inhibition), determine the disease prognosis at multiple biopsy sites or new lesions, determine the biopsy sites that most likely account for disease prognosis, profiling of circulating tumor cells (CTCs), or re-biopsy the primary tumor at disease progression or at the metastatic site.72 CTCs and associated circulating free DNA (cfDNA) provide a unique opportunity to identify driver mutations from noninvasive blood specimens. Tumors shed CTCs and cfDNA into the bloodstream and it is thought that these cells contribute to tumor metastases. cfDNA is typically easier to isolate and more abundant that CTCs themselves, and they tend to represent all tumors in the body and all cells in the tumor, helping to alleviate the issue of tumor heterogeneity.73
Finally, the treatment paradigm for a patient may be driven by the context in which the mutation is found (eg, “x” number of genes/pathways involved) rather than the mutational status of one single gene. For example, multi-pathway inhibition is likely necessary in a variety of tumors prone to resistance or feedback upregulation (ie, BRAF plus MEK inhibitor in metastatic melanoma74 or dual HER2 inhibition in breast cancer).53,54
Big data and integration into health care systems
A globally linked database that integrates genomic data and clinical outcomes will be essential as genomics-based medicine continues its assimilation into health care systems.71 Challenges to integrating genomics-based medicine into health care systems include tumor and biologic heterogeneity, limited access to novel targeted therapies, limited eligibility criteria for enrollment of heterogenous populations in clinical trials, and the paucity of evidence-based medicine to guide clinical decisions, that is, lack of large randomized controlled clinical trials. Leveraging health information technology, translational medicine, patient-centered outcomes, and comparative effectiveness research with real-time high-quality data will allow matching of individual patient characteristics and genomic data to provide the evidence needed.
Rapid-learning health care, part of American Society of Clinical Oncology’s CancerLinQ initiative, is a term used to describe the process of discovery through extension of patient care. CancerLinQ is considered a “quality measurement and reporting system through which oncologists can harness the depth and power of their patients’ clinical records and big data to improve the care they deliver”.75 Data acquired through patient care and clinical research are integrated into a databank, while the health care system “learns” by continuously collecting and analyzing data in a strategic manner, ultimately generating novel hypotheses for investigation and allowing clinicians and researchers to access real-time clinical decision support tools. This method ensures that the intrinsic health care system drives the process of discovery based on real-time clinical practice.76
A secure electronic medical record (EMR) that is responsive to the needs of clinicians paralleled with rapid-learning health care using population-based clinical data will assist in narrowing the gap between research and clinical practice. A survey of health care professionals using ten different EMR systems demonstrated that only 4% of respondents reported that their systems offered any decision support regarding genetic test results.77 The Electronic Medical Records and Genomics (eMERGE) Network was created to develop, disseminate, and apply research that combines DNA biobanks with EMRs for large-scale, high-throughput genetic research. A major goal of this network is to identify how EMRs can serve as a resource for large complex genomic analysis of disease and therapeutic outcomes across the general patient population base. The network also focuses on social and ethical issues, including privacy and confidentiality, as a major concern by patients is the unauthorized use and access to their genetic information.78
Patient-focused perspectives
The integration of pharmacogenomic testing into routine clinical practice will depend not only on clinician acceptance but also on patient adoption and understanding of the risks and benefits. Ethical concerns, including violation of confidentiality, stigmatization, and social pressure to accept pharmacogenomic testing, are among the many challenges that patients face in the adoption of pharmacogenomic testing. Importantly, these concerns may differ based on the genomic test itself, that is, whether the test involves disease susceptibility variants, drug metabolism/pharmacokinetic variants, tumor targets, etc.
A survey study of 328 patients and 378 general practitioners was conducted to determine attitudes toward pharmacogenomic testing. Investigators identified that most patients (96%) and physicians (52%) “appreciated the availability of tests”; however, ~30% of the patients “worried about potential unfavorable test results” (35%) and “violation of privacy” (36%). Female patients were more likely to have a “fearful attitude” (OR 2.85, 95% CI 1.58–5.12), while younger patients were more likely to be “hopeful about the usefulness of pharmacogenomic testing” (OR 2.12, 95% CI 1.01–4.46). General practitioners concerns were primarily related to the potential for patients to be pressured into testing (72%) or the impact of private health insurance agencies obtaining this information (61%).79 Another study sought to explore perceptions of pharmacogenomic testing among individuals with differing parental status or educational exposure to pharmacogenomics and to investigate parents’ views between testing for themselves and testing for their children. The investigators identified that adequate explanation of pharmacogenomics before testing appeared to be the most critical issue to the respondents. Respondents with more knowledge about pharmacogenomics were more comfortable with testing, and when testing was for their child, parents valued their own understanding more than their child’s assent, suggesting education plays a critical role in patient acceptance.80 Blanchette et al conducted a survey study to describe patients’ knowledge, attitudes, and expectations toward genomic testing in cancer. The investigators identified that 76% of patients were at least interested in learning more about pharmacogenomic testing, and 64% reported they believed testing would significantly improve their cancer care. Further, the median score on a 12-item questionnaire to assess knowledge of cancer pharmacogenomics was 8 out of 12, which was significantly associated with education level (P<0.0001). At the time of the survey, approximately half of all patients reported having “sufficient knowledge to make an informed decision to pursue testing”, whereas 34% of patients indicated a need for more formal genetic counseling and guidance.81
Despite the FDA and other consortia’s efforts toward providing information for health professionals and patients about the impact of genetic variation on drug response, there is substantial debate about the clinical utility and safety of pharmacogenomic testing, due in part to the lack of evidence. Studies have reported that the general public is relatively supportive of pharmacogenomic testing, and both physician and patient demand may outweigh concerns regarding the lack of robust evidence.82 Importantly, patients with advanced cancer seem to be motivated to participate in pharmacogenomic testing for cancer therapy; however, this will require further education to understand the clinical utility of testing, differences in germline versus somatic variants, and laws providing protection of patients’ rights (eg, Genetic Information Nondiscrimination Act). Further, surveys of clinicians have demonstrated that the large majority believe pharmacogenomics is important to medication management; however, few have utilized pharmacogenomics, primarily due to lack of knowledge and lack of robust clinical evidence.83 Addressing these concerns will be critical to advance the field of pharmacogenomics and genomics-based medicine in the future.
Conclusion and future directions
Over 140 FDA-approved drugs require pharmacogenomic information on the drug label, resulting in approximately one-quarter of all outpatients who receive at least one drug which is (are) vulnerable to drug/gene interactions.84 Anticancer drugs are one of many classes of medications greatly impacted by pharmacogenomics given their distinct mechanisms of action, narrow therapeutic indices, and potential to analyze two sets of DNA (germline and somatic). Cancer biomarkers can be used as a diagnostic tool for the early detection of cancers, as a prognostic tool to estimate the course and aggressiveness of the disease, and as a predictive tool to estimate an individual’s response to therapy, including drug efficacy and toxicity. Understanding the intricate molecular profile of tumors will help to discover novel driver mutations, provide a larger breadth of targeted therapy options, and allow for better patient stratification in biomarker-driven clinical trials.
Cancer sequencing efforts may capture germline information from matched normal tissue or blood samples, which may be informative for drug/dose selection or disease susceptibility, and somatic mutations, which primarily drive selection of targeted cancer therapies.7 These efforts will generate tremendous amounts of data, and clinicians must be prepared to interpret and utilize this information to optimize cancer therapeutics. A major focus on bioinformatics to readily retrieve actionable information and evidence-based guidelines to translate results into prescribing decisions will be key in the advancement of molecular profiling and selection of targeted therapies.
A concerted effort must be made by cancer centers to adopt and implement genomics-based cancer medicine as the current standard practice.85 As cost of genomic sequencing decreases to less than US$1,000 and turnaround time decreases to less than 2 weeks, an increasing number of patients will have their tumors sequenced, allowing for more personalized assignment of targeted therapies for each patient. Cancer centers are beginning to see a shift from singleplex testing to multiplex genomic sequencing. Arguably, the threshold required to necessitate single-gene tests differs greatly compared to the threshold when whole genome or exome information is readily available from a large genomic panel. Experts may debate it is unethical to ignore this retrievable information given the increasing number of phenotypes that exist to predict drug response. Generation of recommendations to translate results into actionable prescribing decisions, however, is often controlled by the level of evidence required to warrant implementation.7
For a genetic test to be adopted into clinical practice, it must provide reliable, predictive, and actionable information that would have otherwise been unknown.85 Before clinical implementation is warranted, robust evidence from randomized controlled clinical trials are often needed; however, reliance on prospective randomized trials as the sole method to justify implementation is unrealistic, and the delay associated with developing, conducting, and interpreting results could potentially deprive patients of life-saving or life-extending therapies.85
The future of genomic cancer medicine should focus on specimen acquisition of both germline and tumor DNA from early and later phase clinical trials with prospectively collected efficacy and toxicity data. This information will be vital in the discovery and validation of pharmacogenomic associations. Subsequently, genes that have passed replication and validation should be assessed for clinical implementation through proof-of-concept and -efficacy, biomarker-driven clinical trials, which may reduce the required sample size, resources, and time needed to justify clinical uptake. A shift toward large retrospective case-control validation and replication studies and Phase II biomarker-driven clinical trials may allow for a more efficient and rapid method of translation from bench to bedside. As our knowledge of cancer at the molecular level continues to expand, clinicians must understand the therapeutic implications of these pathways and the challenges involved with clinical implementation of pharmacogenomics, including the availability of tests and interpretation of results in clinical practice.
Acknowledgments
The author would like to acknowledge and thank Jeryl Villadolid, PharmD, Levine Cancer Institute, for her assistance in developing Figure 1.
Footnotes
Disclosure
The author reports no conflicts of interest in this work.
References
- 1.Wheeler DA, Wang L. From human genome to cancer genome: the first decade. Genome Res. 2013;23(7):1054–1062. doi: 10.1101/gr.157602.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.President Obama’s Precision Medicine Initiative [webpage on the Internet] Washington, DC: The White House; 2015. [Accessed May 5, 2016]. Available from: https://www.whitehouse.gov/the-press-office/2015/01/30/fact-sheet-president-obama-s-precision-medicine-initiative. [Google Scholar]
- 3.Patel JN. Application of genotype-guided cancer therapy in solid tumors. Pharmacogenomics. 2014;15(1):79–93. doi: 10.2217/pgs.13.227. [DOI] [PubMed] [Google Scholar]
- 4.Patel JN, Mandock K, McLeod HL. Clinically relevant cancer biomarkers and pharmacogenetic assays. J Oncol Pharm Pract. 2014;20(1):65–72. doi: 10.1177/1078155212473862. [DOI] [PubMed] [Google Scholar]
- 5.Couzin-Frankel J. Breakthrough of the year 2013. Cancer immunotherapy. Science. 2013;342(6165):1432–1433. doi: 10.1126/science.342.6165.1432. [DOI] [PubMed] [Google Scholar]
- 6.Wang L, McLeod HL, Weinshilboum RM. Genomics and drug response. N Engl J Med. 2011;364(12):1144–1153. doi: 10.1056/NEJMra1010600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Gillis NK, Patel JN, Innocenti F. Clinical implementation of germ line cancer pharmacogenetic variants during the next-generation sequencing era. Clin Pharmacol Ther. 2014;95(3):269–280. doi: 10.1038/clpt.2013.214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Redig AJ, Janne PA. Basket trials and the evolution of clinical trial design in an era of genomic medicine. J Clin Oncol. 2015;33(9):975–977. doi: 10.1200/JCO.2014.59.8433. [DOI] [PubMed] [Google Scholar]
- 9.Biomarkers Definitions Working Group Biomarkers and surrogate end-points: preferred definitions and conceptual framework. Clin Pharmacol Ther. 2001;69(3):89–95. doi: 10.1067/mcp.2001.113989. [DOI] [PubMed] [Google Scholar]
- 10.van ‘t Veer LJ, Dai H, van de Vijver MJ, et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature. 2002;415(6871):530–536. doi: 10.1038/415530a. [DOI] [PubMed] [Google Scholar]
- 11.Harris LN, Ismaila N, McShane LM, et al. Use of Biomarkers to Guide Decisions on Adjuvant Systemic Therapy for Women With Early-Stage Invasive Breast Cancer: American Society of Clinical Oncology Clinical Practice Guideline. J Clin Oncol. 2016;34(10):1134–1150. doi: 10.1200/JCO.2015.65.2289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Tejpar S, Saridaki Z, Delorenzi M, Bosman F, Roth AD. Microsatellite instability, prognosis and drug sensitivity of stage II and III colorectal cancer: more complexity to the puzzle. J Natl Cancer Inst. 2011;103(11):841–844. doi: 10.1093/jnci/djr170. [DOI] [PubMed] [Google Scholar]
- 13.Popat S, Hubner R, Houlston RS. Systematic review of microsatellite instability and colorectal cancer prognosis. J Clin Oncol. 2005;23(3):609–618. doi: 10.1200/JCO.2005.01.086. [DOI] [PubMed] [Google Scholar]
- 14.Le DT, Uram JN, Wang H, et al. PD-1 blockade in tumors with mismatch repair deficiency. J Clin Oncol. 2015;33(Suppl) doi: 10.1056/NEJMoa1500596. abstr LBA100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Chapman PB, Hauschild A, Robert C, et al. Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N Engl J Med. 2011;364(26):2507–2516. doi: 10.1056/NEJMoa1103782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Tran B, Kopetz S, Tie J, et al. Impact of BRAF mutation and microsatellite instability on the pattern of metastatic spread and prognosis in metastatic colorectal cancer. Cancer. 2011;117(20):4623–4632. doi: 10.1002/cncr.26086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kwak EL, Bang YJ, Camidge DR, et al. Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N Engl J Med. 2010;363(18):1693–1703. doi: 10.1056/NEJMoa1006448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Shaw AT, Kim DW, Nakagawa K, et al. Crizotinib versus chemotherapy in advanced ALK-positive lung cancer. N Engl J Med. 2013;368(25):2385–2394. doi: 10.1056/NEJMoa1214886. [DOI] [PubMed] [Google Scholar]
- 19.Katayama R, Shaw AT, Khan TM, et al. Mechanisms of acquired crizotinib resistance in ALK-rearranged lung Cancers. Sci Transl Med. 2012;4(120):120ra117. doi: 10.1126/scitranslmed.3003316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Shaw AT, Kim DW, Mehra R, et al. Ceritinib in ALK-rearranged non-small-cell lung cancer. N Engl J Med. 2014;370(13):1189–1197. doi: 10.1056/NEJMoa1311107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Shaw AT, Gandhi L, Gadgeel S, et al. Alectinib in ALK-positive, crizotinib-resistant, non-small-cell lung cancer: a single-group, multicentre, phase 2 trial. Lancet Oncol. 2016;17(2):234–242. doi: 10.1016/S1470-2045(15)00488-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Soda M, Choi YL, Enomoto M, et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature. 2007;448(7153):561–566. doi: 10.1038/nature05945. [DOI] [PubMed] [Google Scholar]
- 23.King CR, Kraus MH, Aaronson SA. Amplification of a novel v-erbB-related gene in a human mammary carcinoma. Science. 1985;229(4717):974–976. doi: 10.1126/science.2992089. [DOI] [PubMed] [Google Scholar]
- 24.Yarden Y, Shilo BZ. SnapShot: EGFR signaling pathway. Cell. 2007;131(5):1018. doi: 10.1016/j.cell.2007.11.013. [DOI] [PubMed] [Google Scholar]
- 25.Lynch TJ, Bell DW, Sordella R, et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med. 2004;350(21):2129–2139. doi: 10.1056/NEJMoa040938. [DOI] [PubMed] [Google Scholar]
- 26.Zhou C, Wu YL, Chen G, et al. Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): a multicentre, open-label, randomised, phase 3 study. Lancet Oncol. 2011;12(8):735–742. doi: 10.1016/S1470-2045(11)70184-X. [DOI] [PubMed] [Google Scholar]
- 27.Miller VA, Hirsh V, Cadranel J, et al. Afatinib versus placebo for patients with advanced, metastatic non-small-cell lung cancer after failure of erlotinib, gefitinib, or both, and one or two lines of chemotherapy (LUX-Lung 1): a phase 2b/3 randomised trial. Lancet Oncol. 2012;13(5):528–538. doi: 10.1016/S1470-2045(12)70087-6. [DOI] [PubMed] [Google Scholar]
- 28.Sequist LV, Yang JC, Yamamoto N, et al. Phase III study of afatinib or cisplatin plus pemetrexed in patients with metastatic lung adenocarcinoma with EGFR mutations. J Clin Oncol. 2013;31(27):3327–3334. doi: 10.1200/JCO.2012.44.2806. [DOI] [PubMed] [Google Scholar]
- 29.Califano R, Morgillo F, De Mello RA, Mountzios G. Role of mesenchymal-epithelial transition amplification in resistance to anti-epidermal growth factor receptor agents. Ann Transl Med. 2015;3(6):81. doi: 10.3978/j.issn.2305-5839.2015.03.44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Yun CH, Mengwasser KE, Toms AV, et al. The T790M mutation in EGFR kinase causes drug resistance by increasing the affinity for ATP. Proc Natl Acad Sci U S A. 2008;105(6):2070–2075. doi: 10.1073/pnas.0709662105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Janne PA, Yang JC, Kim DW, et al. AZD9291 in EGFR inhibitor-resistant non-small-cell lung cancer. N Engl J Med. 2015;372(18):1689–1699. doi: 10.1056/NEJMoa1411817. [DOI] [PubMed] [Google Scholar]
- 32.Sun C, Wang L, Huang S, et al. Reversible and adaptive resistance to BRAF(V600E) inhibition in melanoma. Nature. 2014;508(7494):118–122. doi: 10.1038/nature13121. [DOI] [PubMed] [Google Scholar]
- 33.Robert C, Karaszewska B, Schachter J, et al. Improved overall survival in melanoma with combined dabrafenib and trametinib. N Engl J Med. 2015;372(1):30–39. doi: 10.1056/NEJMoa1412690. [DOI] [PubMed] [Google Scholar]
- 34.Patel JN, Fuchs CS, Owzar K, Chen Z, McLeod HL. Gastric cancer pharmacogenetics: progress or old tripe? Pharmacogenomics. 2013;14(9):1053–1064. doi: 10.2217/pgs.13.88. [DOI] [PubMed] [Google Scholar]
- 35.Van Kuilenburg AB, Meinsma R, Zoetekouw L, Van Gennip AH. High prevalence of the IVS14 + 1G>A mutation in the dihydropyrimidine dehydrogenase gene of patients with severe 5-fluorouracil-associated toxicity. Pharmacogenetics. 2002;12(7):555–558. doi: 10.1097/00008571-200210000-00007. [DOI] [PubMed] [Google Scholar]
- 36.Lee AM, Shi Q, Pavey E, et al. DPYD variants as predictors of 5- fluorouracil toxicity in adjuvant colon cancer treatment (NCCTG N0147) J Natl Cancer Inst. 2014;106(12):dju298. doi: 10.1093/jnci/dju298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Meulendijks D, Henricks LM, Sonke GS, et al. Clinical relevance of DPYD variants c.1679T>G, c.1236G>A/HapB3, and c.1601G>A as predictors of severe fluoropyrimidine-associated toxicity: a systematic review and meta-analysis of individual patient data. Lancet Oncol. 2015;16(16):1639–1650. doi: 10.1016/S1470-2045(15)00286-7. [DOI] [PubMed] [Google Scholar]
- 38.Deenen MJ, Meulendijks D, Cats A, et al. Upfront Genotyping of DPYD*2A to Individualize Fluoropyrimidine Therapy: A Safety and Cost Analysis. J Clin Oncol. 2016;34(3):227–234. doi: 10.1200/JCO.2015.63.1325. [DOI] [PubMed] [Google Scholar]
- 39.Caudle KE, Thorn CF, Klein TE, et al. Clinical Pharmacogenetics Implementation Consortium guidelines for dihydropyrimidine dehydrogenase genotype and fluoropyrimidine dosing. Clin Pharmacol Ther. 2013;94(6):640–645. doi: 10.1038/clpt.2013.172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Mathijssen RH, van Alphen RJ, Verweij J, et al. Clinical pharmacokinetics and metabolism of irinotecan (CPT-11) Clin Cancer Res. 2001;7(8):2182–2194. [PubMed] [Google Scholar]
- 41.Innocenti F, Ratain MJ. Irinotecan treatment in cancer patients with UGT1A1 polymorphisms. Oncology (Williston Park) 2003;17(5 Suppl 5):52–55. [PubMed] [Google Scholar]
- 42.Patel JN. Cancer pharmacogenomics: implications on ethnic diversity and drug response. Pharmacogenet Genomics. 2015;25(5):223–230. doi: 10.1097/FPC.0000000000000134. [DOI] [PubMed] [Google Scholar]
- 43.Hoskins JM, Goldberg RM, Qu P, Ibrahim JG, McLeod HL. UGT1A1*28 genotype and irinotecan-induced neutropenia: dose matters. J Natl Cancer Inst. 2007;99(17):1290–1295. doi: 10.1093/jnci/djm115. [DOI] [PubMed] [Google Scholar]
- 44.Swen JJ, Nijenhuis M, de Boer A, et al. Pharmacogenetics: from bench to byte – an update of guidelines. Clin Pharmacol Ther. 2011;89(5):662–673. doi: 10.1038/clpt.2011.34. [DOI] [PubMed] [Google Scholar]
- 45.Toffoli G, Cecchin E, Gasparini G, et al. Genotype-driven phase I study of irinotecan administered in combination with fluorouracil/leucovorin in patients with metastatic colorectal cancer. J Clin Oncol. 2010;28(5):866–871. doi: 10.1200/JCO.2009.23.6125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Marcuello E, Paez D, Pare L, et al. A genotype-directed phase I-IV dose-finding study of irinotecan in combination with fluorouracil/leucovorin as first-line treatment in advanced colorectal cancer. Br J Cancer. 2011;105(1):53–57. doi: 10.1038/bjc.2011.206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Nowell PC, Hungerford DA. Chromosome studies on normal and leukemic human leukocytes. J Natl Cancer Inst. 1960;25:85–109. [PubMed] [Google Scholar]
- 48.de Klein A, van Kessel AG, Grosveld G, et al. A cellular oncogene is translocated to the Philadelphia chromosome in chronic myelocytic leukaemia. Nature. 1982;300(5894):765–767. doi: 10.1038/300765a0. [DOI] [PubMed] [Google Scholar]
- 49.Druker BJ, Guilhot F, O’Brien SG, et al. Five-year follow-up of patients receiving imatinib for chronic myeloid leukemia. N Engl J Med. 2006;355(23):2408–2417. doi: 10.1056/NEJMoa062867. [DOI] [PubMed] [Google Scholar]
- 50.Bixby D, Talpaz M. Mechanisms of resistance to tyrosine kinase inhibitors in chronic myeloid leukemia and recent therapeutic strategies to overcome resistance. Hematology Am Soc Hematol Educ Program. 2009:461–476. doi: 10.1182/asheducation-2009.1.461. [DOI] [PubMed] [Google Scholar]
- 51.Zhang WW, Cortes JE, Yao H, et al. Predictors of primary imatinib resistance in chronic myelogenous leukemia are distinct from those in secondary imatinib resistance. J Clin Oncol. 2009;27(22):3642–3649. doi: 10.1200/JCO.2008.19.4076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Gorre ME, Mohammed M, Ellwood K, et al. Clinical resistance to STI-571 cancer therapy caused by BCR-ABL gene mutation or amplification. Science. 2001;293(5531):876–880. doi: 10.1126/science.1062538. [DOI] [PubMed] [Google Scholar]
- 53.Lierman E, Smits S, Cools J, Dewaele B, Debiec-Rychter M, Vandenberghe P. Ponatinib is active against imatinib-resistant mutants of FIP1L1-PDGFRA and KIT, and against FGFR1-derived fusion kinases. Leukemia. 2012;26(7):1693–1695. doi: 10.1038/leu.2012.8. [DOI] [PubMed] [Google Scholar]
- 54.McLeod HL, Krynetski EY, Relling MV, Evans WE. Genetic polymorphism of thiopurine methyltransferase and its clinical relevance for childhood acute lymphoblastic leukemia. Leukemia. 2000;14(4):567–572. doi: 10.1038/sj.leu.2401723. [DOI] [PubMed] [Google Scholar]
- 55.Relling MV, Hancock ML, Rivera GK, et al. Mercaptopurine therapy intolerance and heterozygosity at the thiopurine S-methyltransferase gene locus. J Natl Cancer Inst. 1999;91(23):2001–2008. doi: 10.1093/jnci/91.23.2001. [DOI] [PubMed] [Google Scholar]
- 56.Stanulla M, Schaeffeler E, Flohr T, et al. Thiopurine methyltransferase (TPMT) genotype and early treatment response to mercaptopurine in childhood acute lymphoblastic leukemia. JAMA. 2005;293(12):1485–1489. doi: 10.1001/jama.293.12.1485. [DOI] [PubMed] [Google Scholar]
- 57.Stocco G, Cheok MH, Crews KR, et al. Genetic polymorphism of inosine triphosphate pyrophosphatase is a determinant of mercaptopurine metabolism and toxicity during treatment for acute lymphoblastic leukemia. Clin Pharmacol Ther. 2009;85(2):164–172. doi: 10.1038/clpt.2008.154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Food and Drug Administration . Purinethol. Silver Spring, MD: Food and Drug Administration; 2003. [Accessed February 17, 2016]. Available from: http://www.accessdata.fda.gov/drugsatfda_docs/label/2004/09053s024lbl.pdf. [Google Scholar]
- 59.Yang JJ, Landier W, Yang W, et al. Inherited NUDT15 variant is a genetic determinant of mercaptopurine intolerance in children with acute lymphoblastic leukemia. J Clin Oncol. 2015;33(11):1235–1242. doi: 10.1200/JCO.2014.59.4671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Heinrich MC, Corless CL, Demetri GD, et al. Kinase mutations and imatinib response in patients with metastatic gastrointestinal stromal tumor. J Clin Oncol. 2003;21(23):4342–4349. doi: 10.1200/JCO.2003.04.190. [DOI] [PubMed] [Google Scholar]
- 61.Djalalov S, Beca J, Hoch JS, et al. Cost efectiveness of EML4-ALK fusion testing and first-line crizotinib treatment for patients with advanced ALK-positive non-small-cell lung cancer. J Clin Oncol. 2014;32(10):1012–1019. doi: 10.1200/JCO.2013.53.1186. [DOI] [PubMed] [Google Scholar]
- 62.Buettner R, Wolf J, Thomas RK. Lessons learned from lung cancer genomics: the emerging concept of individualized diagnostics and treatment. J Clin Oncol. 2013;31(15):1858–1865. doi: 10.1200/JCO.2012.45.9867. [DOI] [PubMed] [Google Scholar]
- 63.Deverka PA, Dreyfus JC. Clinical integration of next generation sequencing: coverage and reimbursement challenges. J Law Med Ethics. 2014;42(Suppl 1):22–41. doi: 10.1111/jlme.12160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Stratton MR, Campbell PJ, Futreal PA. The cancer genome. Nature. 2009;458(7239):719–724. doi: 10.1038/nature07943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Campbell PJ, Yachida S, Mudie LJ, et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature. 2010;467(7319):1109–1113. doi: 10.1038/nature09460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Gerlinger M, Rowan AJ, Horswell S, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med. 2012;366(10):883–892. doi: 10.1056/NEJMoa1113205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Vandin F, Upfal E, Raphael BJ. De novo discovery of mutated driver pathways in cancer. Genome Res. 2012;22(2):375–385. doi: 10.1101/gr.120477.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Miller DG. On the nature of susceptibility to cancer. The presidential address. Cancer. 1980;46(6):1307–1318. doi: 10.1002/1097-0142(19800915)46:6<1307::aid-cncr2820460602>3.0.co;2-6. [DOI] [PubMed] [Google Scholar]
- 69.Mendelsohn J. Personalizing oncology: perspectives and prospects. J Clin Oncol. 2013;31(15):1904–1911. doi: 10.1200/JCO.2012.45.3605. [DOI] [PubMed] [Google Scholar]
- 70.Schinzel AC, Hahn WC. Oncogenic transformation and experimental models of human cancer. Front Biosci. 2008;13:71–84. doi: 10.2741/2661. [DOI] [PubMed] [Google Scholar]
- 71.Macconaill LE, Van Hummelen P, Meyerson M, Hahn WC. Clinical implementation of comprehensive strategies to characterize cancer genomes: opportunities and challenges. Cancer Discov. 2011;1(4):297. doi: 10.1158/2159-8290.CD-11-0110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Meric-Bernstam F, Farhangfar C, Mendelsohn J, Mills GB. Building a personalized medicine infrastructure at a major cancer center. J Clin Oncol. 2013;31(15):1849–1857. doi: 10.1200/JCO.2012.45.3043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Alix-Panabieres C, Pantel K. Circulating tumor cells: liquid biopsy of cancer. Clin Chem. 2013;59(1):110–118. doi: 10.1373/clinchem.2012.194258. [DOI] [PubMed] [Google Scholar]
- 74.Flaherty KT, Infante JR, Daud A, et al. Combined BRAF and MEK inhibition in melanoma with BRAF V600 mutations. N Engl J Med. 2012;367(18):1694–1703. doi: 10.1056/NEJMoa1210093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Schilsky RL, Michels DL, Kearbey AH, Yu PP, Hudis CA. Building a rapid learning health care system for oncology: the regulatory framework of CancerLinQ. J Clin Oncol. 2014;32(22):2373–2379. doi: 10.1200/JCO.2014.56.2124. [DOI] [PubMed] [Google Scholar]
- 76.Abernethy AP, Etheredge LM, Ganz PA, et al. Rapid-learning system for cancer care. J Clin Oncol. 2010;28(27):4268–4274. doi: 10.1200/JCO.2010.28.5478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Scheuner MT, de Vries H, Kim B, Meili RC, Olmstead SH, Teleki S. Are electronic health records ready for genomic medicine? Genet Med. 2009;11(7):510–517. doi: 10.1097/GIM.0b013e3181a53331. [DOI] [PubMed] [Google Scholar]
- 78.Kohane IS. Using electronic health records to drive discovery in disease genomics. Nat Rev Genet. 2011;12(6):417–428. doi: 10.1038/nrg2999. [DOI] [PubMed] [Google Scholar]
- 79.Rogausch A, Prause D, Schallenberg A, Brockmoller J, Himmel W. Patients’ and physicians’ perspectives on pharmacogenetic testing. Pharmacogenomics. 2006;7(1):49–59. doi: 10.2217/14622416.7.1.49. [DOI] [PubMed] [Google Scholar]
- 80.Zhang SC, Bruce C, Hayden M, Rieder MJ. Public perceptions of pharmacogenetics. Pediatrics. 2014;133(5):e1258–e1267. doi: 10.1542/peds.2013-1416. [DOI] [PubMed] [Google Scholar]
- 81.Blanchette PS, Spreafico A, Miller FA, et al. Genomic testing in cancer: patient knowledge, attitudes, and expectations. Cancer. 2014;120(19):3066–3073. doi: 10.1002/cncr.28807. [DOI] [PubMed] [Google Scholar]
- 82.Haga SB, LaPointe NM. The potential impact of pharmacogenetic testing on medication adherence. Pharmacogenomics J. 2013;13(6):481–483. doi: 10.1038/tpj.2013.33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Stanek EJ, Sanders CL, Taber KA, et al. Adoption of pharmacogenomic testing by US physicians: results of a nationwide survey. Clin Pharmacol Ther. 2012;91(3):450–458. doi: 10.1038/clpt.2011.306. [DOI] [PubMed] [Google Scholar]
- 84.Frueh FW, Amur S, Mummaneni P, et al. Pharmacogenomic biomarker information in drug labels approved by the United States food and drug administration: prevalence of related drug use. Pharmacotherapy. 2008;28(8):992–998. doi: 10.1592/phco.28.8.992. [DOI] [PubMed] [Google Scholar]
- 85.McLeod HL. Cancer pharmacogenomics: early promise, but concerted effort needed. Science. 2013;339(6127):1563–1566. doi: 10.1126/science.1234139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Patel JN, Villadolid J. Cancer Drug Delivery: Pharmacogenetics, biomarkers and targeted therapies. In: Kesharwani Rajesh Kumar., editor. Handbook of Research on Novel Approaches for Drug Delivery. Hershey, PA: IGI Global; 2015. in press. [Google Scholar]
- 87.Table of Pharmacogenomic Biomarkers in Drug Labeling [webpage on the Internet] Silver Spring, MD: US Food and Drug Administration; 2015. [Accessed May 5, 2016]. Available from: http://www.fda.gov/drugs/scienceresearch/researchareas/pharmacogenetics/ucm083378.htm. [Google Scholar]