Abstract
The chemical nature of the 5′ end of RNA is a key determinant of RNA stability, processing, localization, translation efficiency1,2, and has been proposed to provide a layer of “epitranscriptomic” gene regulation3. Recently it has been shown that some bacterial RNA species carry a 5′-end structure reminiscent of the 5′ 7-methylguanylate “cap” in eukaryotic RNA. In particular, RNA species containing a 5′-end nicotinamide adenine dinucleotide (NAD+) or 3′-desphospho-coenzyme A (dpCoA) have been identified in both Gram-negative and Gram-positive bacteria3–6. It has been proposed that NAD+, reduced NAD+ (NADH), and dpCoA caps are added to RNA after transcription initiation, in a manner analogous to the addition of 7-methylguanylate caps6–8. Here, we show instead that NAD+, NADH, and dpCoA are incorporated into RNA during transcription initiation, by serving as non-canonical initiating nucleotides (NCINs) for de novo transcription initiation by cellular RNA polymerase (RNAP). We further show that both bacterial RNAP and eukaryotic RNAP II incorporate NCIN caps, that promoter DNA sequences at and upstream of the transcription start site determine the efficiency of NCIN capping, that NCIN capping occurs in vivo, and that NCIN capping has functional consequences. We report crystal structures of transcription initiation complexes containing NCIN-capped RNA products. Our results define the mechanism and structural basis of NCIN capping, and suggest that NCIN-mediated “ab initio capping” may occur in all organisms
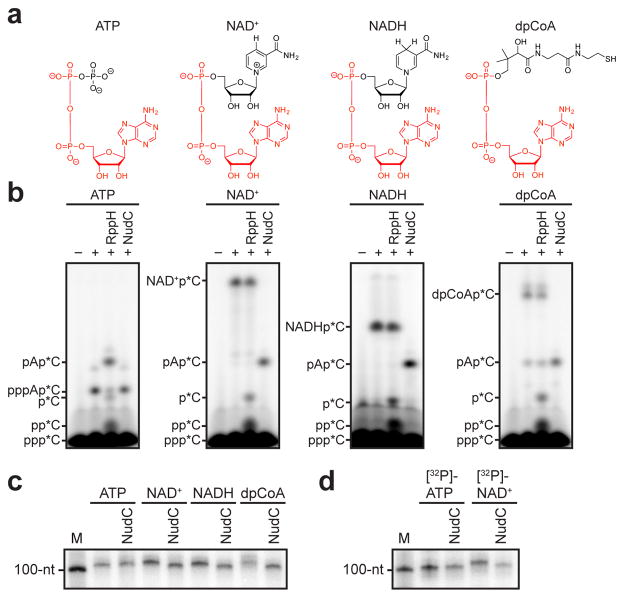
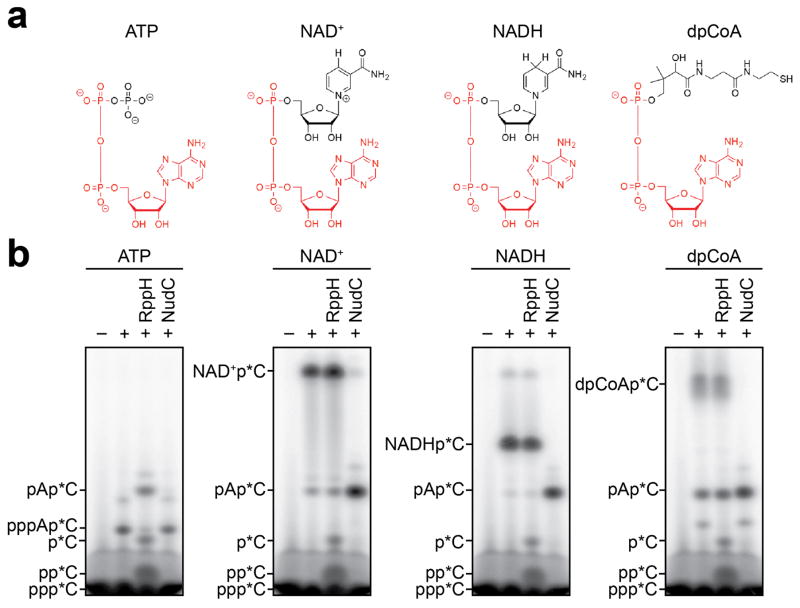
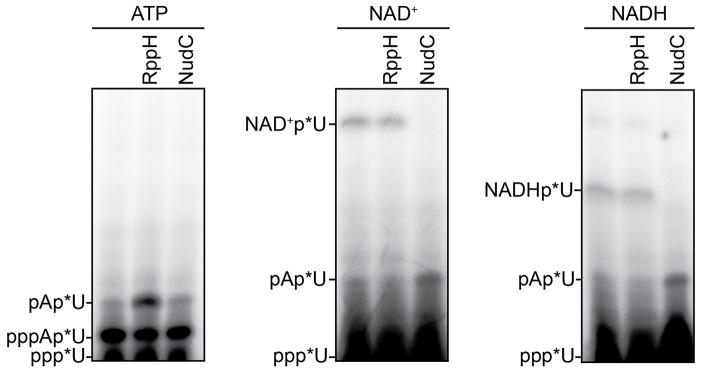
NAD+, NADH, and dpCoA are nucleotides that share an adenosine-diphosphate substructure with adenosine triphosphate (ATP) (Figure 1a, red). Therefore, in principle, NAD+, NADH, and dpCoA could serve in place of ATP as initiating nucleotides in de novo transcription initiation by a cellular RNAP, and thereby be incorporated ab initio into RNA (see Supplementary Discussion). To assess whether NAD+, NADH, and dpCoA serve as initiating nucleotides for a cellular RNAP, we performed in vitro transcription experiments with Escherichia coli RNAP and DNA templates containing promoters for RNAs identified as most-highly NAD+-capped in vivo6 (PgadY and PrnaI; Figure 1b; Extended Data Figure 1b). We performed reactions using ATP, NAD+, NADH, or dpCoA as initiating nucleotide and [α32P]-CTP as extending nucleotide. In each case, we observed efficient formation of an initial RNA product (Figure 1b; Extended Data Figure 1b). RppH, which processes 5′-triphosphate and 5′-diphosphate RNAs to 5′-monophosphate RNAs9, processed the product obtained with ATP, but had no effect on products obtained with NAD+, NADH, and dpCoA (Figure 1b; Extended Data Figure 1b). Conversely, NudC, which processes 5′-capped RNAs to 5′-monophosphate RNAs6, had no effect on the product formed with ATP, but processed products obtained with NAD+, NADH, and dpCoA (Figure 1b; Extended Data Figure 1b). Liquid-chromatography/tandem-mass-spectrometry analysis confirms the detection and structural assignment of the product of NAD+-mediated transcription initiation (Extended Data Figure 2).
Figure 1. De novo transcription initiation by ATP and NCINs.
a. Structures of ATP, NAD+, NADH, and dpCoA. Red, identical atoms. b. Initial RNA products of in vitro transcription reactions with ATP, NAD+, NADH, or dpCoA as initiating nucleotide and [α32P]-CTP as extending nucleotide (E. coli RNAP; PgadY). Products were treated with RppH (processes 5′-triphosphate RNA to 5′-monophosphate RNA and 5′-NTP to 5′-NDP/5′-NMP9,14) or NudC (processes 5′-NAD+/NADH-capped RNA to 5′-monophosphate RNA6) as indicated. c, d. Full-length RNA products of in vitro transcription reactions with ATP, NAD+, NADH, or dpCoA as initiating nucleotide and [α32P]-CTP, GTP, and UTP as extending nucleotides (c), or with [γ32P]-ATP or [α32P]-NAD+ as initiating nucleotide and CTP, GTP, and UTP as extending nucleotides (d) (E. coli RNAP; PrnaI fused to an A-less cassette ). M, 100-nt marker. For gel source data, see Supplementary Figure 1.
To assess whether initial RNA products formed with NCINs can be extended to full-length RNA products, we performed transcription reactions with ATP, NAD+, NADH, or dpCoA as initiating nucleotide and [α32P]-UTP, CTP, and GTP as extending nucleotides on templates containing PrnaI linked to a 100-nt A-less cassette. We observed production of full-length products in all cases, and observed that full-length products with NAD+, NADH, and dpCoA contained NudC-sensitive, capped 5′-ends (Figure 1c). Equivalent results were obtained using [32P]-NAD+, rather than [α32P]-UTP, to detect products (Figure 1d; Extended Data Figure 3).
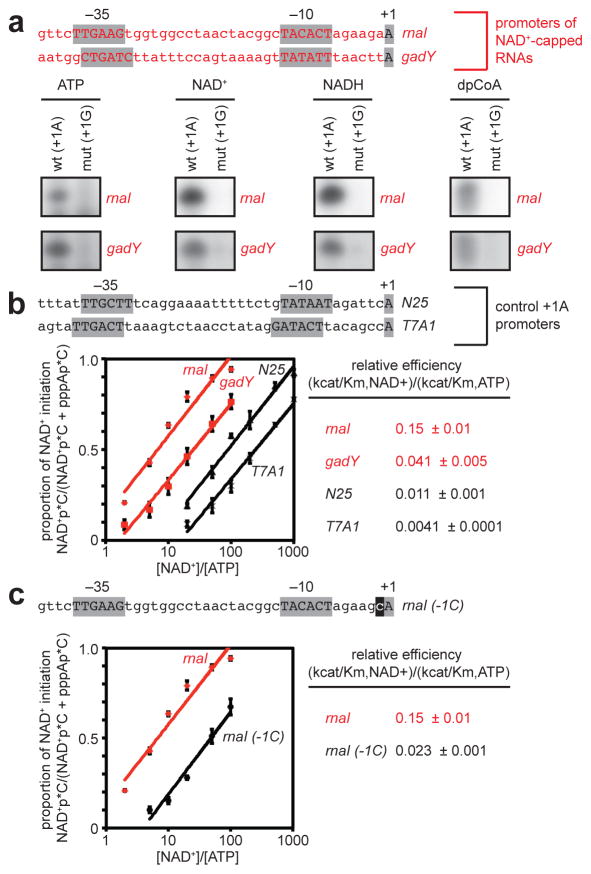
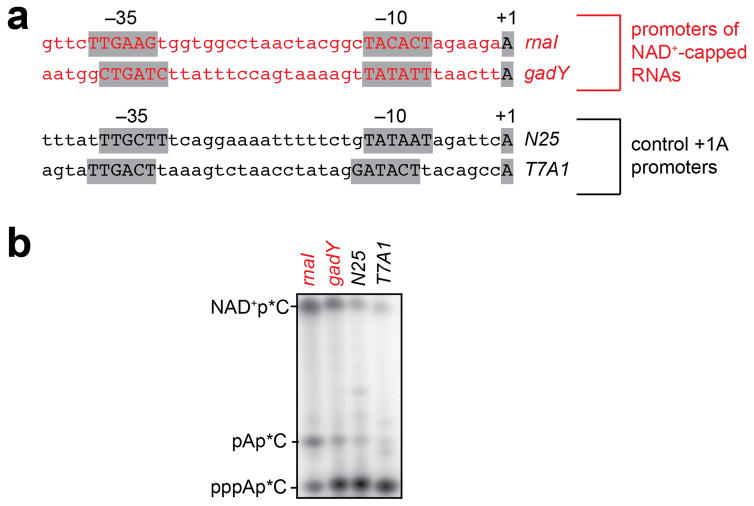
Because NAD+, NADH, and dpCoA have the same Watson-Crick base pairing preferences as ATP (Figure 1a), NCIN-mediated initiation should occur only for RNAs transcribed from promoters that contain A:T at the transcription start site, +1 (+1A promoters). Consistent with this inference, we find that NCIN-mediated initiation occurs only with +1A promoters, and not with corresponding +1G promoter derivatives (Figure 2a).
Figure 2. Promoter-sequence effects on efficiency of NCIN-mediated transcription initiation.
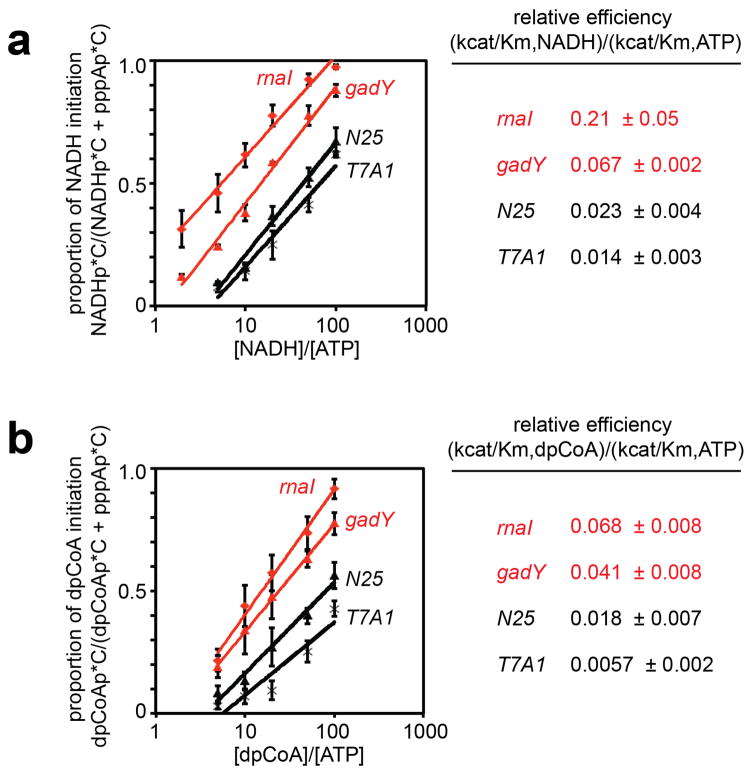
a. NCIN capping requires A+1. Top, promoters of NAD+-capped RNAs (promoter elements and start sites in gray). Bottom, initial RNA products of in vitro transcription reactions with ATP, NAD+, NADH, or dpCoA as initiating nucleotide and [α32P]-CTP as extending nucleotide [E. coli RNAP; wt (+1A), PrnaI (upper) or PgadY (lower); mut (+1G), +1G derivative of PrnaI (upper) or PgadY (lower)]. b. Promoter sequence determinants in addition to A+1 affect NCIN capping. Top, control +1A promoters. Bottom left, dependence of NAD+ capping on [NAD+]/[ATP] ratio (mean±SEM of 4 determinations). Bottom right, relative efficiencies of NAD+ capping. c. Promoter position −1 affects NCIN capping. Top, PrnaI (-1C) (-1 in black). Other features as in b. (mean±SEM of 3 determinations). For gel source data, see Supplementary Figure 1.
In principle, other aspects of promoter sequence, in addition to +1A at the transcription start site, could affect NCIN-mediated initiation. Consistent with this inference, we find that relative efficiencies of NAD+-mediated initiation and ATP-mediated initiation, (kcat/Km,NAD+)/(kcat/Km,ATP), differ over a one- to two-order-of-magnitude range at different +1A promoters, with PrnaI and PgadY exhibiting high relative efficiencies and PN25 and PT7A1 promoters exhibiting low relative efficiencies (Figures 2a–b; Extended Data Figures 4–5). Substitution of the PrnaI promoter position immediately upstream of the transcription start site, −1, results in an order-of-magnitude reduction in relative efficiency (Figure 2c), indicating that position −1 is a key determinant of NAD+-mediated initiation.
Having obtained results establishing that NAD+, NADH, and dpCoA function as NCINs for transcription initiation in vitro and that promoter sequence determines the efficiency of NCIN-mediated inititation in vitro, we hypothesized that NCIN-mediated inititation also may occur in vivo and be responsible for the generation of NAD+-, NADH-, and dpCoA-capped RNAs in vivo. Consistent with this hypothesis, the promoters for full-length RNAs identified as enriched for NAD+-capping in vivo all are +1A promoters6, the class of promoters competent for NAD+-mediated initiation in vitro (Figure 2a). Further consistent with this hypothesis, the promoters identified as most-highly enriched for NAD+ capping in vivo are PrnaI and PgadY6, the two promoters with highest efficiencies of NAD+-mediated initiation in vitro (Figure 2b).
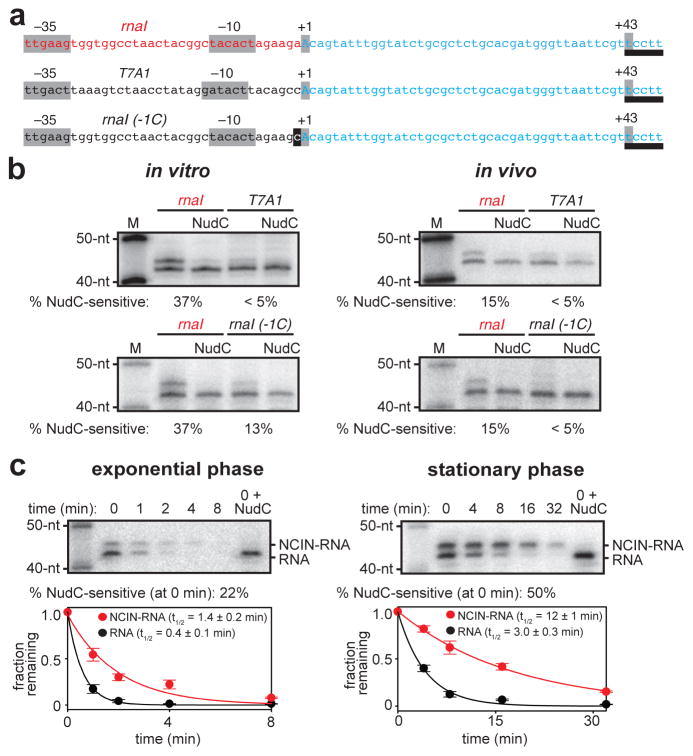
To show directly that NCIN-mediated inititation occurs in vivo, we assessed whether the promoter-sequence dependence for NCIN-mediated initiation observed in vitro is observed in vivo. We fused promoters observed in vitro to have high (PrnaI) and low (PT7A1, PrnaI (-1C)) efficiencies of NAD+-mediated initiation (Figures 2b–c) to transcribed regions directing synthesis of the same RNA (reference RNA; Figure 3a) and compared yields of NudC-sensitive, capped RNA products (Figure 3b). We found that the promoter with high efficiency of NAD+-mediated initiation in vitro yielded high levels of NudC-sensitive products in vivo (PrnaI; Figure 3b), whereas the promoters with low efficiency of NAD+-mediated initiation in vitro yielded low, undetectable, levels of NudC-sensitive products in vivo (PT7A1, PrnaI (-1C); Figure 3b). We conclude that NCIN-mediated initiation occurs in vivo, and that most, or all, NCIN-capped RNA products in vivo arise from NCIN-mediated initiation.
Figure 3. NCIN-mediated transcription initiation in vivo.
a. Templates having rnaI, T7A1, and rnaI (-1C) promoters fused to identical transcribed regions (promoter elements, start sites, and position of RNA 3′-end in gray; DNA that directs synthesis of reference RNA in blue; site for MazF-mt3 endoribonuclease used to generate RNA products having uniform RNA 3′-ends, underlined). b. NCIN capping in vitro (left; 1 mM NAD+, 200 μM ATP, CTP, UTP, and GTP) and in vivo (right; RNA isolated from cells, treated with MazF-mt3 or MazF-mt3 plus NudC, and detected by hybridization). M, markers (40-nt, 50-nt). c. Effects of NCIN capping on RNA stability in vivo (rnaI template; left, exponential-phase cells; right, stationary-phase cells; times, minutes after addition of RNA-synthesis inhibitor rifampin; half-life values are the mean±SEM of 3 determinations for exponential phase and 5 determinations for stationary phase). For gel source data, see Supplementary Figure 1.
To assess whether NCIN capping has functional consequences in vivo, we measured stabilities of NCIN capped and uncapped RNA in vivo, using a ΔNudC strain to eliminate NudC processing. We observe that NCIN capping results in a large, ~3- to 4-fold, increase in RNA stability (Figure 3c), demonstrating a functional consequence of NCIN capping in vivo. We further observe that levels of NCIN capping are ~2-fold higher in stationary-phase vs. exponential-phase (Figure 3c), demonstrating a growth-phase-dependence in NCIN capping.
All cellular RNAPs are members of a protein family having conserved structures and mechanisms10–12. To assess whether eukaryotic RNAP II, like bacterial RNAP, can use NCINs as intiating nucleotides, we performed in vitro transcription experiments with Saccharomyces cerevisiae RNAP II using ATP, NAD+, or NADH as initiating nucleotide and [α32P]-UTP as extending nucleotide (Extended Data Figure 6). The results matched those for bacterial RNAP: initiation occurred with NAD+ and NADH, and products were insensitive to RppH and sensitive to NudC. We conclude that eukaryotic RNAP II can initiate transcription with NAD+ and NADH, and we suggest that ab initio capping with NAD+ and NADH also may occur in eukaryotes.
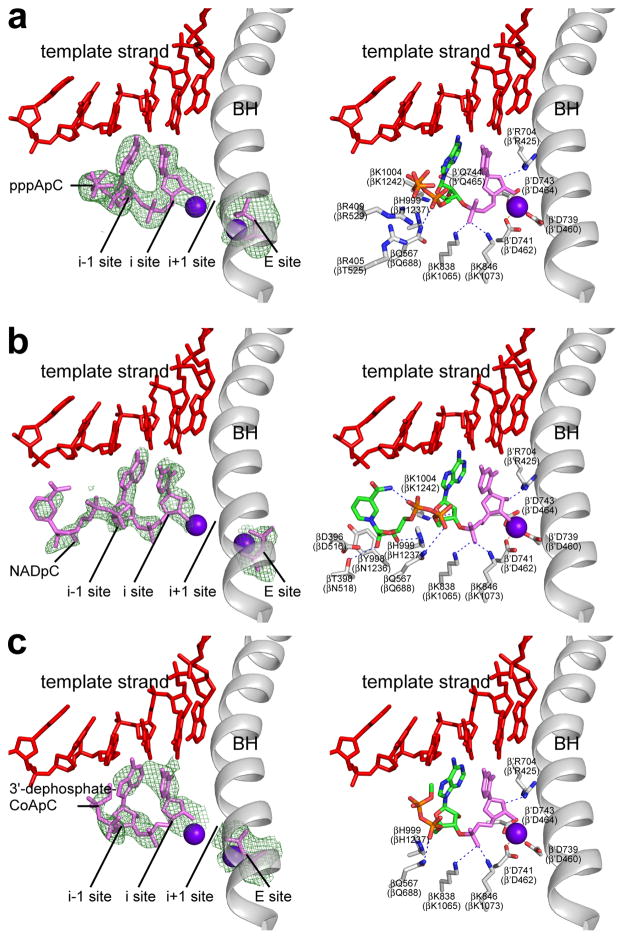
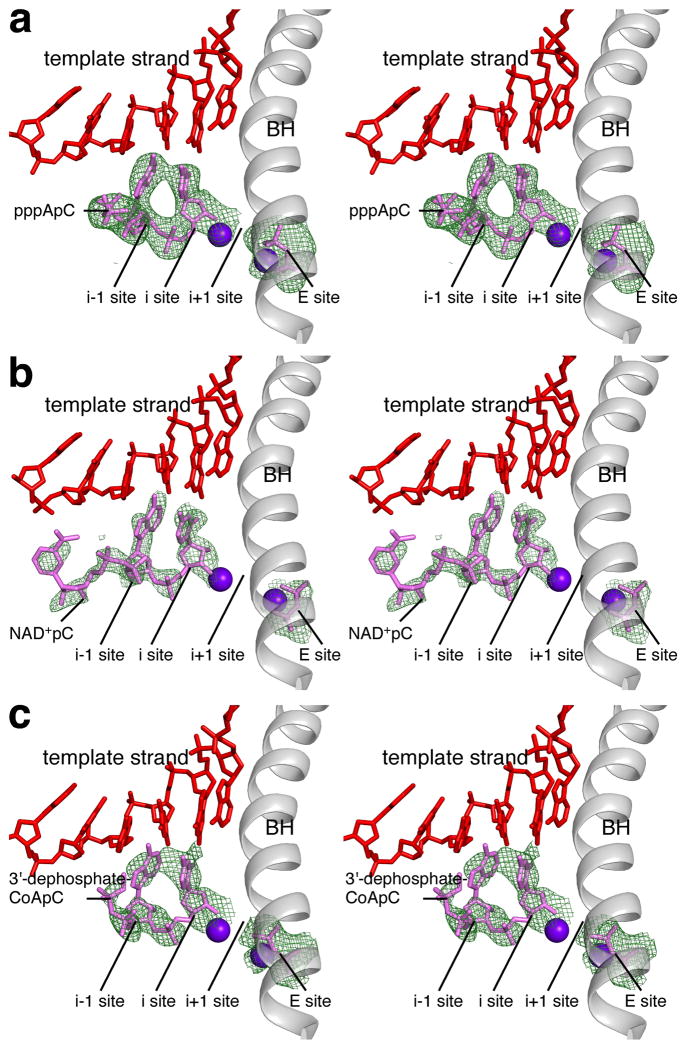
To define the structural basis of transcription initiation with ATP, NAD+, and dpCoA as initiating nucleotides we determined crystal structures of product complexes obtained by soaking crystals of Thermus thermophilus RNAP-promoter open complex13, RPo, with ATP and CTP, NAD+ and CTP, and dpCoA and CTP (Figure 4; Extended Data Figure 7; Extended Data Table 1). The results indicate that, in each case, the initiating entity--ATP, NAD+, or dpCoA--and an extending nucleotide--CTP--were able to react in crystallo to form an initial RNA product--pppApC, NAD+pC, or dpCoApC, respectively--and that RNAP was able to translocate in crystallo by 1 bp relative to the nucleic-acid scaffold, yielding a complex in a post-translocated state poised for RNA extension (Figure 4). The results for ATP provide the first structural description of an initial product complex for a cellular RNAP. The results for NAD+ and dpCoA show graphically that NAD+ and dpCoA serve as initiating nucleotides in transcription initiation and define the interactions that the initial RNA products make with DNA and RNAP (see Supplementary Discussion).
Figure 4. Structural basis of NCIN-mediated transcription initiation.
a–c. Crystal structures of RPo-pppApC, RPo-NAD+pC, and RPo-dpCoApC. Left, electron density and atomic model for initial RNA product. Green mesh, Fo-Fc omit map (contoured at 2.5σ in a–b and 2.2σ in c); red, DNA; pink, RNA product and diphosphate in “E site” (see15–17); violet spheres, Mg2+(I) and Mg2+(II); gray, RNAP bridge helix. Right, contacts between RPo and initial RNA product. Green and orange, carbon and phosphorus atoms derived from initiating nucleotide; pink, atoms derived from extending nucleotide; red, DNA atoms and non-DNA oxygen atoms; blue, nitrogen atoms; gray sticks, RNAP carbon atoms; violet sphere, Mg2+(I); gray ribbon, RNAP bridge helix.
Our results establish that NAD+, NADH, and dpCoA function as non-canonical initiating nucleotides (NCINs) for de novo transcription initiation and demonstrate the occurrence of ab initio, as opposed to post-transcription-initiation, capping of cellular RNAs.
METHODS
Proteins
E. coli RNAP core enzyme was prepared from BL21(DE3) cells transformed with plasmid pVS10 (gift of I. Artsimovitch) as described in18. σ70 was prepared from BL21(DE3) cells transformed with plasmid pσ70-His (gift of J. Roberts) as described in19,20. RppH was purchased from NEB. NudC was purified from BL21(DE3) cells from pET NudC-His, which was prepared as described6. Briefly, nudC was amplified by PCR from E. coli strain MG1655 using oligonucleotides JB221 and JB222. The PCR product was digested with XbaI and NotI and inserted into pET28c digested with XbaI and NotI. T. thermophilus RNAP holoenzyme was prepared as described13. Saccharomyces cerevisiae RNAP II was prepared as described21. MazF-mt3 protein, prepared as described22, was a gift from N. Woychik (Rutgers University, Piscataway, NJ).
In vitro transcription assays with E. coli RNAP: linear templates
Linear transcription templates were synthesized by PCR using Phusion HF Polymerase master mix (Thermo) and oligonucleotides listed in Supplementary Table 1. PCR reactions contained 5 nM of the indicated template oligo, and 0.5 μM of the indicated forward and reverse primers. PCR products were purified using Qiagen PCR-clean up columns prior to use in transcription reactions.
-
Sequence of gadY (−65 to +35) template used in Figures 1b, 2a–b, and Extended Data Figures 4 and 5:
AGCGTATAGCTTATGTTTATAAAAAAATGGCTGATCTTATTTCCAGTAAAAGTTATATTTAACTTACTGAGAGCACAAAGTTTCCCGTGCCAACAGGGAG (prepared using oligos JB224, JB230, JB231)
-
Sequence of gadY (+1G) (−65 to +35) template used in Figure 2a:
AGCGTATAGCTTATGTTTATAAAAAAATGGCTGATCTTATTTCCAGTAAAAGTTATATTTAACTTGCTGAGAGCACAAAGTTTCCCGTGCCAACAGGGAG (prepared using oligos JB244, JB230, JB231)
-
Sequence of rnaI (−65 to +35) template used in Figures 2a–c and Extended Data Figures 1b, 4, and 5:
CGAGGTATGTAGGCGGTGCTACAGAGTTCTTGAAGTGGTGGCCTAACTACGGCTACACTAGAAGAACTGTATTTGGTATCTGCGCTCTGCTGAAGCCAGTT (prepared using oligos JB288, JB281, JB269)
-
Sequence of mutant rnaI (+1G) (−65 to +35) template used in Figure 2a:
CGAGGTATGTAGGCGGTGCTACAGAGTTCTTGAAGTGGTGGCCTAACTACGGCTACACTAGAAGAGCTGTATTTGGTATCTGCGCTCTGCTGAAGCCAGTT (prepared using oligos JB287, JB281, JB269)
-
Sequence of mutant rnaI (-1C) (−65 to +35) template used in Figure 2c:
CGAGGTATGTAGGCGGTGCTACAGAGTTCTTGAAGTGGTGGCCTAACTACGGCTACACTAGAAGCACTGTATTTGGTATCTGCGCTCTGCTGAAGCCAGTT (prepared using oligos JB367, JB281, JB269)
-
Sequence of rnaI (−65 to +112) A-less cassette template used in Figures 1c–d and Extended Data Figure 3:
CGAGGTATGTAGGCGGTGCTACAGAGTTCTTGAAGTGGTGGCCTAACTACGGCTACACTAGAAGAACTTGTTTTGGTGTCTGCGCTCCTCCTTGCCTGTTTCCTCGGTTCTTTGTGTTGGTTGCTCTGTGTTCCTTCGTTTTTCCGCCCTGCTTGGCGGTTTTTTCGTTTTCT GTGC (prepared using oligos JB288, JB281, JB251)
-
Sequence of N25 (−65 to +35) template used in Figure 2b and Extended Data Figures 4 and 5:
ATCCGTCGAGGGAAATCATAAAAAATTTATTTGCTTTCAGGAAAATTTTTCTGTATAATAGATTCACTAATTTGAGAGAGGAGTTTAAATATGGCTGGTT (prepared using oligos JB232, JB233, JB234)
-
Sequence of T7A1 (−65 to +35) template used in Figure 2b and Extended Data Figures 4 and 5:
GATTAATTTAAAATTTATCAAAAAGAGTATTGACTTAAAGTCTAACCTATAGGATACTTACAGCCACTGAGAGGGACACGGCGAATAGCCATCCCAACGA (prepared using oligos JB235, JB228, JB229)
Promoter −35 elements, −10 elements and transcription start sites are in bold and underlined.
In vitro transcription assays with E. coli RNAP: abortive initiation assays
10 nM of linear template was mixed with 50 nM core RNAP and 250 nM σ70 in 60 μl transcription buffer (10 mM Tris-HCl pH 8.0, 40 mM KCl, 10 mM MgCl2, 0.1 mM EDTA, 1 mM DTT, 0.1 mg/ml BSA, 2% glycerol) and incubated at 37 °C for 15 min to form open complexes. The indicated initiating nucleotide [0.2 mM ATP (Roche), 1 mM NAD+ (Roche), 1 mM NADH (Roche) or 1 mM dpCoA (Sigma-Aldrich)] was added along with 6 μCi [α32P]-CTP (Perkin Elmer; 3000 Ci/mmol) and reactions were incubated at 37 °C for 10 min to allow for product formation. For the NAD+/ATP competition assays shown in Figure 2b–c, 1 mM NAD+ was present in each reaction along with increasing concentrations of ATP. The data in Extended Data Figure 4b shows initial products formed in reactions with 1 mM NAD+ and 50 μM ATP. For the NADH/ATP competition assays shown in Extended Data Figure 5a, 1 mM NADH was present in each reaction along with increasing concentrations of ATP. For the dpCoA/ATP competition assays shown in Extended Data Figure 5b, 1 mM dpCoA was present in each reaction along with increasing concentrations of ATP.
For experiments shown in Figure 2a–c and Extended Data Figures 4–5, reactions were stopped by addition of an equal volume of gel loading buffer [90% formamide, 100 mM Tris-HCl, pH 8.0, 18 mM EDTA, 0.025% xylene cyanol, 0.025% bromophenol blue]. For the reactions shown in Figure 1b and Extended Data Figure 1b the samples were passed through a Nanosep Centrifugal Device (10 kDa molecular weight cutoff; Pall Corporation), the flow through was collected, mixed with 1 U RppH (NEB), 400 nM NudC or NudC storage buffer (50 mM Tris-HCl, pH 7.9, 0.5M NaCl, 50 mM KCl, 1 mM MgCl2, 7 mM TCEP, 50% glycerol), incubated at 37 °C for 30 min, and mixed with an equal volume of gel loading buffer. Samples were run on 15% TBE-urea polyacrylamide gels (UreaGel system, National Diagnostics). Autoradiography of gels was performed using storage phosphor screens and a Typhoon 9400 variable mode imager (GE Life Science) and quantified using ImageQuant software. We note that the presence of contaminating AMP in the dpCoA stock (Extended Data Figure 8) leads to the generation of pApC products in reactions performed with dpCoA.
For the competition assays in Figure 2b–c and Extended Data Figure 5, (NCIN)pC and pppApC products were quantified, the fraction of (NCIN)pC products was plotted vs. [NCIN]/[ATP], and the value of (kcat/Km,NCIN)/(kcat/Km, ATP) was calculated as the value of [NCIN]/[ATP] for which the fraction of (NCIN)pC products was 0.5. Values were obtained from averaging results of three (Figure 2c, Extended Data Figure 5) or four (Figure 2b) technical replicates.
In vitro transcription assays with E. coli RNAP: run-off transcription assays
For reactions shown in Figure 1c, open complexes formed as described above were mixed with the indicated initiating nucleotide [0.2 mM ATP, 1 mM NAD+, 1 mM NADH or 1 mM dpCoA] along with 6 μCi [α32P]-CTP (Perkin Elmer; 3000 Ci/mmol), 200 μM CTP, 200 μM UTP, and 200 μM GTP, and reactions were incubated at 37 °C for 10 min to allow for product formation. For the reactions shown in Figure 1d and Extended Data Figure 4, open complexes formed as described above were mixed with 10 μCi [γ32P]-ATP (Perkin Elmer; 6000 Ci/mmol), or 20 μCi [α32P]-NAD+ (Perkin Elmer; 800 Ci/mmol) along with 200 μM CTP, 200 μM UTP and 200 μM GTP. Reactions were incubated at 37 °C for 10 min to allow for product formation, and stopped by addition of 100 μl stop solution (0.6 M Tris-HCl, pH 8.0, 18 mM EDTA, 0.1 mg/ml glycogen). Samples were extracted with acid phenol:chloroform and RNA transcripts were recovered by ethanol precipitation, resuspended in transcription buffer and divided into 10 μl aliquots. For the experiment in Figure 1d, 400 nM NudC was added to one aliquot while NudC storage buffer was added to the other for mock treatment. For the experiment in Extended Data Figure 3, 0.25 U alkaline phosphatase (Thermo Fisher) was added to one aliquot, 400 nM NudC was added to one aliquot, both 0.25 U alkaline phosphatase and 400 nM NudC was added to one aliquot, and NudC storage buffer was added to another aliquot for mock treatment. Reactions were incubated at 37 °C for 30 min and stopped by addition of an equal volume of gel loading buffer and analyzed by gel electrophoresis on 15% TBE-urea polyacrylamide gels as described above. Reactions were loaded alongside an Ambion Decade Marker System (Life Technologies) to enable an estimation of the approximate size of the full-length transcripts generated.
In vitro transcription assays with E. coli RNAP: transcription assays with negatively supercoiled DNA templates
-
Plasmid pJB89 contains sequence for the PrnaI-reference RNA fusion inserted between the XbaI and SalI sites of plasmid pSG28923. The sequence of the insert is:
TCTAGACGAGGTATGTAGGCGGTGCTACAGAGTTCTTGAAGTGGTGGCCTAACTACGGCTACACTAGAAGAACAGTATTTGGTATCTGCGCTCTGCACGATGGGTTAATTCGTTCCTTGGATCCGAATAGCCATCCCAATCGAACAGGCCTGCTGGTAATCGCAGGCCTTTTTATTTGGATGTCGAC
-
Plasmid pJB91 contains sequence for the PT7A1-reference RNA fusion inserted between the XbaI and SalI sites of plasmid pSG28923. The sequence of the insert is:
TCTAGAGATTAATTTAAAATTTATCAAAAAGAGTATTGACTTAAAGTCTAACCTATAGGATACTTACAGCCACAGTATTTGGTATCTGCGCTCTGCACGATGGGTTAATTCGTTCCTTGGATCCGAATAGCCATCCCAATCGAACAGGCCTGCTGGTAATCGCAGGCCTTTTTATTTGGATGTCGAC
-
Plasmid pJB95 contains sequence for the PrnaI (-1C)-reference RNA fusion inserted between the XbaI and SalI sites of plasmid pSG28923. The sequence of the insert is:
TCTAGACGAGGTATGTAGGCGGTGCTACAGAGTTCTTGAAGTGGTGGCCTAACTACGGCTACACTAGAAGCACAGTATTTGGTATCTGCGCTCTGCACGATGGGTTAATTCGTTCCTTGGATCCGAATAGCCATCCCAATCGAACAGGCCTGCTGGTAATCGCAGGCCTTTTTATTTGGATGTCGAC
Promoter −35 elements and −10 elements are in bold and underlined. DNA that directs synthesis of the reference RNA is colored blue. Recognition site for MazF-mt3 (UCCUU; 22) is underlined. Reference RNA includes sequence of the tR2 terminator.
For in vitro reactions shown in Figure 3b, open complexes were formed using 10 nM of plasmid (pJB89, pJB91, or pJB95) and 50 nM core RNAP and 250 nM σ70 in 30 μl transcription buffer (see above). Transcription was carried out for 10 min at 37 °C using 1 mM NAD+, 200 μM ATP, UTP, CTP and GTP. Reactions were stopped by the addition of 120 μl of stop solution and samples were extracted with acid phenol:chloroform and ethanol precipitated. The resulting RNAs were resuspended in 10 mM Tris, pH 8.0, supplemented with 2 μg of RNA isolated from wild-type MG1655 cells and treated with 2 μM MazF-mt3 for 1 hr at 37 °C. Reactions were divided into two, 9 μl aliquots and 1 μl 10× transcription buffer was added to each. 1 μM NudC was added to one of these aliquots and NudC storage buffer was added to the other. Reactions were incubated at 37 °C for 90 min and stopped by addition of an equal volume of gel loading buffer.
Analysis of RNA products generated in vivo
For the experiments of Figure 3b, E. coli MG1655 cells containing plasmids pJB89, pJB91, or pJB95 were shaken at 220 RPM at 37 °C in 25 ml LB (10 g Bacto-tryptone, 5 g Bacto-yeast extract and 2.5 g NaCl per liter) containing 25 μg/ml chloramphenicol in 125 ml flasks (Bellco). When cell density reached an OD600 ~0.6, 2 mL of the cell suspension was centrifuged (30 sec, 10,000 × g at room temperature) to collect cells, supernatants were removed, and cell pellets were rapidly frozen on dry ice and stored at −80 °C.
For the RNA stability experiments of Figure 3c, strain MG1655 ΔnudC::Kan was constructed by P1 transduction of the ΔnudC::Kan cassette from JW554824 into MG1655. MG1655 ΔnudC::Kan cells containing plasmids pJB89, pJB91, or pJB95 were grown as described above to an OD600 ~0.6 (exponential phase) or an OD600 ~3.5 (stationary phase). 2 mL of the cell suspension was collected as described above (the RNA derived from these cells was used as the 0 min time point). Rifampicin was added to a final concentration of 2 mg/mL to the remainder of the culture. 2 mL aliquots were taken at the indicated times and cells were collected as described above. Frozen pellets were resuspended in 1 ml of TRI Reagent solution (Molecular Research Center). Samples were incubated at 70 °C for 10 min and then centrifuged (10 min, 21,000 × g, 4 °C) to remove insoluble material. The supernatant was transferred to a fresh tube and 200 μl of chloroform was added, samples were mixed by vortexing, and then centrifuged (10 min, 21,000 × g, 4 °C). The aqueous phase was transferred to a fresh tube, extracted with acid phenol:chloroform, and RNA transcripts were recovered by ethanol precipitation and resuspended in 10 mM Tris, pH 8.0.
5 μg of isolated RNA was treated with 2 μM MazF-mt3 in 10 mM Tris, pH 8.0 (total volume, 25 μl) and incubated for 60 min at 37 °C. Reactions were stopped by addition of an equal volume of gel loading buffer. In cases where RNA products were treated with NudC, reactions were divided into into two aliquots and 10× transcription buffer was added to each to a final concentration of 1×. 1 μM NudC was added to one of these aliquots and NudC storage buffer was added to the other. Reactions were incubated at 37 °C for 45 min and stopped by addition of an equal volume of gel loading buffer.
Detection of RNA products generated in vitro or in vivo by hybridization
In Figure 3b–c, RNA products were detected by a procedure comprising electrophoresis on 10% 7.5 M urea slab gels (equilibrated and run in 1× TBE), transfer to and UV cross-linking to a membrane (Nytran supercharge nylon membrane, GE Healthcare Life Sciences), hybridization with a 32P-labeled “locked-nucleic-acid” probe complementary to positions +31 to +42 of RNA-standard (Sequence, agCaaAttAacCc; LNA bases capitalized; purchased from Exiquon, Woburn, MA; 32P-labeled using T4 polynucleotide kinase), high stringency washing (procedure as described in25), and storage-phosphor imaging. Bands were quantified using ImageQuant software. The % of RNA products in the upper band was determined for each sample as: 100 * (upper band/(upper band + lower band)). Values of %NudC-sensitive products represent the reduction in the % of RNA products in the upper band upon treatment with NudC. Values shown represent the average of three technical replicates (in vitro) or three biological replicates (in vivo).
For the experiments in Figure 3c, values of half-life were determined by fitting data to a single-exponential decay function (mean±SEM of three biological replicates for exponential phase and five biological replicates for stationary phase).
In vitro transcription assays with E. coli RNAP: liquid-chromatography/tandem-mass spectrometry (LC/MS/MS) detection of NAD+pC
Open complexes formed as described above were mixed with 2 mM NAD+ and 10 μM CTP, incubated at 37 °C for 1 hr, passed through a Nanosep Centrifugal Device, and the flow through was analyzed by LC/MS/MS. (Control samples in which NAD+, CTP, or RNAP were not present in the reactions were also prepared.) LC/MS/MS analyses were performed in negative ion mode with a Finnigan LTQ mass spectrometer equipped with an electrospray ionization (ESI) interface coupled with a Finnigan Surveyor HPLC system. The flow rate from the Finnigan Surveyor pump was 0.3 ml/min. The autosampler temperature was 5 °C. The liquid chromatography method used a YMC ODS-A 5 μm particle size 120Å pore size column, 3.0 mm × 100 mm. The samples were separated using a gradient mobile phase consisting of 5 mM ammonium formate buffered to pH 7.9 in water (A) and methanol (B). The gradient condition was: 0–5 min, 100% A; 5–15 min, 100–30% A; 15–23 min, 30–0% A; 23–28 min, 0% A; 28–40 min, 100% A. Column temperature was 25°C. The temperature of the heated capillary was 200 °C. Fragmentation was activated by collision induced dissociation (CID) with a collision energy of 20–25%. The search for NAD+pC was conducted by isolating its anion [(M–2H), m/z 967] at isolation width 3 using full scan mode, and analyzing its fragmentation spectrum (collision energy 20%) from m/z = 210 to m/z = 1000. The characteristic fragment m/z = 845 (loss of nicotinamide) was used to identify NAD+pC. The instrument control, data acquisition, and data analysis were performed by Xcalibur software. Ammonium formate (99%), water (HPLC grade) and methanol (HPLC grade) used in these LC/MS/MS experiments were purchased from Sigma-Aldrich.
In vitro transcription assays with Saccharomyces cerevisiae RNAP II
Bubble templates for initiation were utilized following the approach in26 by mixing 2.5 nmol of oligonucleotide CKO1639 and 2.5 nmol of oligonucleotide CKO1621 (see Supplementary Table 1) in 100 μl of annealing buffer (40 mM Tris, pH 8.0, 50 mM NaCl, 1 mM EDTA), incubating at 95 °C for 3 min, and slowly cooling to 23 °C (1 °C/min) to allow annealing. The oligo mixture was run on a 2.5% agarose gel (Apex, Genesee) equilibrated and run in 1× TBE [90 mM Tris base (BioRad), 90 mM boric acid (Calbiochem), 2 mM EDTA (JT Baker)], the gel was stained with ethidium bromide and the band corresponding to the bubble template was excised from the gel, crushed, and incubated with 500 μl elution buffer (500 mM NH4OAc, 10 mM Mg(OAc)2, 1 mM EDTA, 0.1% SDS). Gel debris was removed using a Spin-X column (Costar) and nucleic acids were isolated by ethanol precipitation and re-suspended in nuclease-free water (Ambion) to a concentration of 0.5 μg/μl. 500 ng of the template was mixed with M280 magnetic streptavidin beads (Invitrogen) in 1× TE (from Ambion components) supplemented with 1 M NaCl (EMD/Calbiochem) and incubated at 25 °C for 30 min to allow binding. Templates bound to the beads were washed three times using magnetic capture to separate beads in 1× TE supplemented with 1 M NaCl, once with 1× TE, and then equilibrated into 1× transcription buffer [TB: 20 mM Tris, pH 8.0 (Ambion), 40 mM KCl (Ambion), 10 mM MgCl2 (Ambion), 10% glycerol (Macron/Avantor), 0.25 mg/ml BSA (Ambion), 1 mM DTT (Gold Biotechnology)].
2.5 ug purified Saccharomyces cerevisiae RNAP II was added to each template in 50 μl 1× TB, and incubated with rotation at 25 °C for 20 min. Complexes were washed following magnetic capture twice with 200 μl 1× TB, and resuspended in 22 μl 2× TB containing 0.4 U/μl RNase Inhibitor (NEB). 7.5 μl of this mixture was aliquoted for reactions (4 reactions per bubble template).
5 μl of “initiator”/[α32P]-UTP mixes were added to give final concentrations of: 1 mM ATP (GE Lifesciences), 2.5 μM [α32P]-UTP (from 800 Ci/mmol [12.5 μM], Perkin Elmer); 1 mM NAD+, 2.5 μM [α32P]-UTP (from 800 Ci/mmol [12.5 μM], Perkin Elmer); or 1 mM NADH, 2.5 μM [α32P]-UTP (from 800 Ci/mmol [12.5 μM], Perkin Elmer). Reactions were incubated at 25 °C for 2 hr then passed through a Nanosep Centrifugal Device (10 kD molecular weight cutoff). The flow through was collected, mixed with 0.5 μl (2.5 U) RppH (NEB), 0.5 μl NudC (400 nM final) or 0.5 μl NudC storage buffer, incubated at 37 °C for 30 min, and mixed with an equal volume of urea gel loading buffer [10 M urea (Calbiochem), 0.5 M EDTA, pH 8.0 (JT Baker), 1× TBE]. Samples were run on 25% 19:1 acrylamide:bisacrylamide gel [from 40% 19:1 solution (BioRad), 7 M urea (Calbiochem), 1× TBE (90 mM Tris base; BioRad), 90 mM boric acid (Calbiochem), 2 mM EDTA (JT Baker)]. Autoradiography of gels was performed using storage phosphor screens and a Pharos (Bio-Rad) imager. Radiolabeled products were visualized using ImageLab (Bio-Rad) software.
Structure determination: RPo-pppApC
Crystals of T. thermophilus RPo were prepared using a derivative of the consensus-bacterial promoter nucleic-acid scaffold used for analysis of RPo in13 having A in place of C at position −1 of the DNA template strand, and were grown and handled essentially as described13. (T. thermophilus RNAP is a bacterial RNAP that is homologous in structure, sequence, and mechanism to E. coli RNAP, but, empirically, yielding crystals of RNAP that diffract to higher resolution than E. coli RNAP13,27.) Crystallization drops contained 1 μl RPo in 20 mM Tris-HCl, pH 7.7, 100 mM NaCl, and 1% glycerol, and 1 μl reservoir buffer (RB; 100 mM Tris-HCl, pH 8.4, 200 mM KCl, 50 mM MgCl2, and 9.5% PEG4000), and were equilibrated against 400 μl RB in a vapor-diffusion hanging-drop tray. Rod-like crystals appeared in 1 day, and were used to micro-seed hanging drops using the same conditions. ATP and CTP (Sigma-Aldrich) were soaked into RPo crystals by addition of 0.2 μl 40 mM ATP and 40 mM CTP in 60% (v/v) RB to the crystallization drop, and incubated for 2 min at 22 °C. Crystals were transferred into reservoir solutions containing 2 mM ATP, 2 mM CTP, and 17.5% (v/v) (2R, 3R)-(-)-2,3-butanediol (Sigma-Aldrich) and were flash-cooled with liquid nitrogen.
Diffraction data were collected at BNL beamline X29A (temperature, 100 K; wavelength, 1.08 Å), processed and scaled using HKL200028. Structure factors were converted using the French-Wilson algorithm in Phenix29 and were subjected to anisotropy correction using the UCLA MBI Diffraction Anisotropy server30 (http://services.mbi.ucla.edu/anisoscale/). The structure was solved by molecular replacement with Molrep31 using one RNAP molecule from the structure of T. thermophilus RPo [PDB 4G7H]13 as the search model. Early-stage refinement included rigid-body refinement of each RNAP molecule, followed by rigid-body refinement of each subunit of RNAP molecules. Cycles of iterative model building with Coot32 and refinement with Phenix33 were performed. Atomic models of the DNA nontemplate strand, the DNA template strand, and RNA were built into mFo-DFc omit maps, and subsequent cycles of refinement and model building were performed. The final crystallographic model of RPo-pppApC, refined to Ramachandran statistics of 97.84% favoured, 2.10% allowed, and 0.06% outliers, has been deposited in the PDB with accession code 5D4C.
Structure determination: RPo-NAD+pC
NAD+ (Sigma-Aldrich) and CTP were soaked into RPo crystals (prepared as described above) by addition of 0.2 μl 30 mM NAD and 40 mM CTP in 50% (v/v) RB to the crystallization drop, and incubated for 2 hr at 22 °C. Crystals were transferred into reservoir solutions containing 1.5 mM NAD, 2 mM CTP, and 17.5% (v/v) (2R, 3R)-(-)-2,3-butanediol and were flash-cooled with liquid nitrogen.
Diffraction data were collected at APS beamline 19-ID-D (temperature, 100 K; wavelength, 0.98 Å), processed and scaled using HKL200028, and subjected to anisotropic correction using the UCLA MBI Diffraction Anisotropy server30 (http://services.mbi.ucla.edu/anisoscale/). The structure was solved and refined using procedures analogous to those described above for RPo-pppApC. The final crystallographic model of RPo-NAD+pC, refined to Ramachandran statistics of 97.81% favoured, 2.15% allowed, and 0.04% outliers, has been deposited in the PDB with accession code 5D4D.
Structure determination: RPo-dpCoApC
dpCoA (Sigma-Aldrich) and CTP were soaked into RPo crystals (prepared as described above) by addition of 0.2 μl 60 mM 3′-dephosphate-CoA and 40 mM CTP in 50% (v/v) RB to the crystallization drop, and incubated for 2 hr at 22 °C. Crystals were transferred into reservoir solutions containing 3 mM 3′-dephosphate-CoA, 2 mM CTP, and 17.5% (v/v) (2R, 3R)-(-)-2,3-butanediol and were flash-cooled with liquid nitrogen.
Diffraction data were collected at APS beamline 19-ID-D (temperature, 100 K; wavelength, 0.98 Å), processed and scaled using HKL200028, and subjected to anisotropic correction using the UCLA MBI Diffraction Anisotropy server30 (http://services.mbi.ucla.edu/anisoscale/). The structure was solved and refined using procedures analogous to those described above for RPo-pppApC. The final crystallographic model of dpCoApC, refined to Ramachandran statistics of 97.54% favoured, 2.43% allowed, and 0.03% outliers, has been deposited in the PDB with accession code 5D4E.
Analysis of dpCoA by liquid chromatography (LC)-UV
AMP (Sigma-Aldrich) and dpCoA were diluted to 1 mM in water (HPLC grade) before injection. LC-UV analyses were performed using a Waters 2960 HPLC system coupled with a Waters 2996 photodiode array detector. The flow rate from the Finnigan Surveyor pump was 0.3 ml/min. The autosampler temperature was 5 °C. The LC method used an YMC ODS-A 5 μm particle size 120Å pore size column, 3.0 mm × 100 mm. The samples were separated using a gradient mobile phase consisting of 25 mM triethylammonium formate buffered to pH 3.0 in water (A) and methanol (B). The gradient condition was: 0–5 min, 100% A; 5–15 min, 100–30% A; 15–23 min, 30–0% A; 23–28 min, 0% A; 28–40 min, 100% A. Column temperature was 25 °C. Detector wave length was 260 nm. The instrument control, data acquisition, and data analysis were performed by MassLynx software. Triethylammonium acetate buffer (2.0 M) and formic acid (>95%) were purchased from Sigma-Aldrich.
Extended Data
Extended Data Figure 1. De novo transcription initiation by ATP and NCINs.
a. Structures of ATP, NAD+, NADH, and dpCoA. Red, identical atoms. b. Initial RNA products of in vitro transcription reactions with ATP, NAD+, NADH, or dpCoA as initiating nucleotide and [α32P]-CTP as extending nucleotide (E. coli RNAP; PrnaI; see analogous data for PgadY in Figure 1b ). Products were treated with RppH (processes 5′-triphosphate RNA to 5′-monophosphate RNA and 5′-NTP to 5′-NDP/5′-NMP9,14) or NudC (processes 5′-NAD+/NADH-capped RNA to 5′-monophosphate RNA6) as indicated. For gel source data, see Supplementary Figure 1.
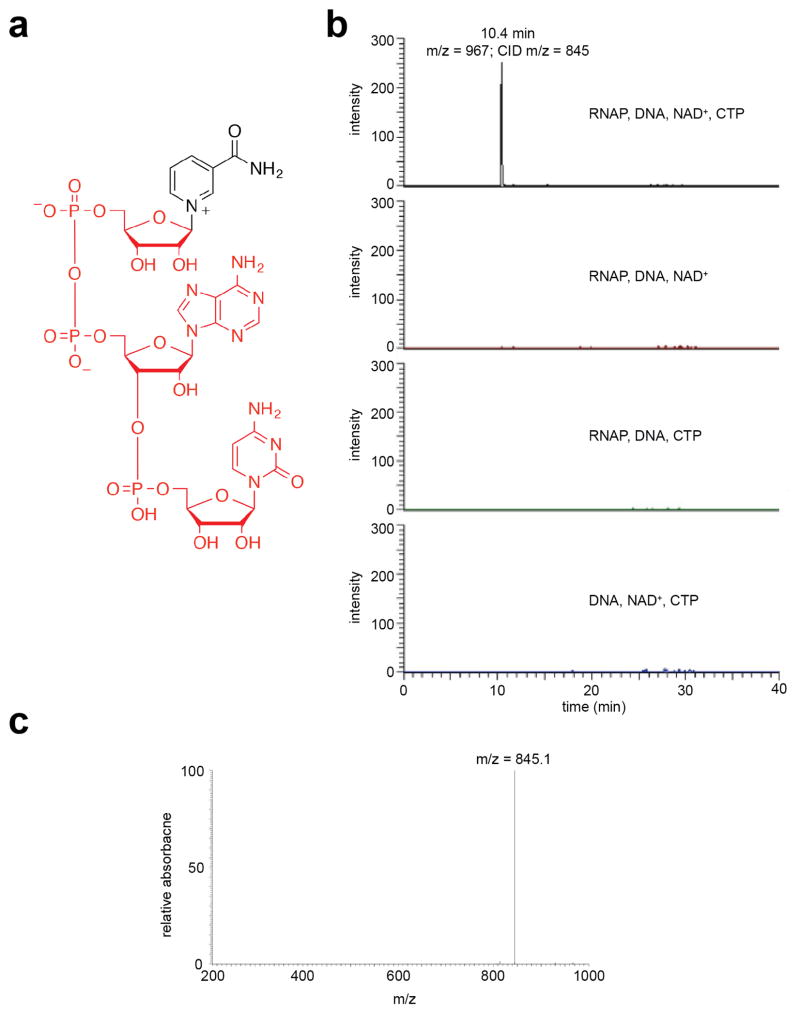
Extended Data Figure 2. LC/MS/MS analysis of initial RNA products of in vitro transcription reactions with NAD+ as initiating nucleotide and CTP as an extending nucleotide.
a. Structure of NAD+pC (red, atoms corresponding to CID-generated fragment ion). b. Extracted ion chromatogram (signal derived from detection of parent ion of m/z = 967 and CID fragment of m/z = 845 corresponding to NAD+pC minus nicotinamide). Reactions contained the indicated components. c. Mass spectrum of CID fragment.
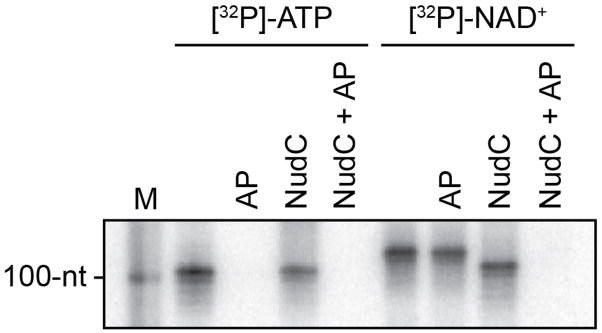
Extended Data Figure 3. Sensitivity of full-length RNA products to alkaline phosphatase treatment.
Full-length RNA products of in vitro transcription reactions with [γ32P]-ATP or [α32P]-NAD+ as initiating nucleotide and CTP, GTP, and UTP as extending nucleotides (E. coli RNAP; PrnaI fused to an A-less cassette ). Products were treated with alkaline phosphatase (AP; processes 5′ phosphates) or NudC (processes 5′-NAD+/NADH-capped RNA to 5′-monophosphate RNA6) as indicated. Results indicate that full-length RNA products generated in reactions with [α32P]-NAD+ as initiating nucleotide are not sensitive to AP until they are processed by NudC. M, 100-nt marker. For gel source data, see Supplementary Figure 1.
Extended Data Figure 4. Promoter-sequence effects on efficiency of NCIN-mediated transcription initiation: NAD+.
a. Templates having rnaI, gadY, N25, and T7A1 promoters used in the assays. b. Representative raw data from experiments of Figure 2b. Initial RNA products of in vitro transcription reactions performed in the presence of 50 μM ATP and 1 mM NAD+ as initiating nucleotides and [α32P]-CTP as extending nucleotide (E. coli RNAP; PrnaI, PgadY, PN25, or PT7A1). (We note that contaminating AMP in the NAD+ stock results in production of pAp*C.) For gel source data, see Supplementary Figure 1.
Extended Data Figure 5. Promoter-sequence effects on efficiency of NCIN-mediated transcription initiation: NADH and dpCoA.
a. Left, dependence of NADH capping on [NADH]/[ATP] ratio (mean±SEM of 3 determinations). Right, relative efficiencies of NADH capping. (E. coli RNAP; PrnaI, PgadY, PN25, or PT7A1). b. Left, dependence of dpCoA-capping on [dpCoA]/[ATP] ratio (mean±SEM of 3 determinations). Right, relative efficiencies of dpCoA capping. (E. coli RNAP; PrnaI, PgadY, PN25, or PT7A1).
Extended Data Figure 6. NCIN-mediated de novo transcrption initiation by eukaryotic RNAP II.
Initial RNA products of in vitro transcription reactions with ATP, NAD+, or NADH as initiating nucleotide and [α32P]-UTP as extending nucleotide. Reactions were performed with yeast RNAP II and an artificial-bubble transcription initiation template. Products were treated with RppH or NudC as indicated. For gel source data, see Supplementary Figure 1.
Extended Data Figure 7. Structural basis of NCIN-mediated transcription initiation: stereoviews.
a–c. Crystal structures of RPo-pppApC, RPo-NAD+pC, and RPo-dpCoApC. Stereoviews of density and fit for initial RNA product. Green mesh, Fo-Fc omit map (contoured at 2.5σ in A–B and 2.2σ in c); red, DNA; pink, RNA product and diphosphate in “E site” (see15–17); violet spheres, Mg2+(I) and Mg2+(II); gray, RNAP bridge helix.
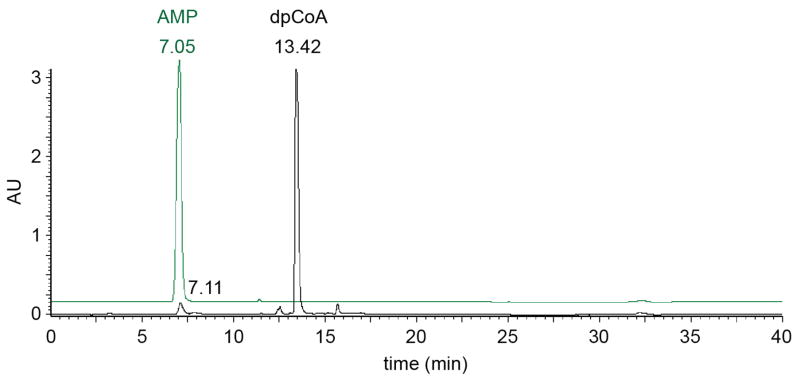
Extended Data Figure 8. AMP content of dpCoA stock.
HPLC chromatogram of dpCoA stock (Sigma-Aldrich, lot SLBJ2886V; 50 nmol). Green: HPLC chromatogram of AMP (20 nmol). Comparison of chromatograms indicates that the dpCoA stock contains ~2% AMP. The observation that the dpCoA stock contains ~2% AMP in the dpCoA stock accounts for the formation of pApC in reactions performed with dpCoA (Figure 1b).
Extended Data Table 1.
Crystallographic refinement and statistics.
| RPo-pppApC | RPo-NAD+pC | RPo-dpCoApC | |
|---|---|---|---|
| Data collection | |||
| Space group | P2(1) | P2(1) | P2(1) |
| Cell dimensions | |||
| a, b, c (Å) | 186.0, 103.6, 297.4 | 185.0, 103.5, 296.3 | 185.8, 103.9, 297.1 |
| α, β, γ (°) | 90.0, 98.3, 90.0 | 90.0, 98.3, 90.0 | 90.0, 98.5, 90.0 |
| Resolution (Å) | 50.00–3.30(3.36–3.30)* | 50.00–3.00(3.05–3.00)* | 40.00–3.10(3.15–3.10)* |
| Rsym or Rmerge | 0.185(0.954) | 0.119(0.920) | 0.120(0.840) |
| I/σI | 8.7(1.7) | 10.6(1.5) | 10.4(1.4) |
| Completeness (%) | 0.961(0.875) | 0.979(0.934) | 0.989(0.960) |
| Redundancy | 5.4(5.0) | 4.0(3.7) | 3.8(3.5) |
| Refinement | |||
| Resolution (Å) | 50.00–3.30 | 50.00–3.00 | 40.00–3.10 |
| No. reflections | 154105 | 218773 | 187101 |
| Rwork/Rfree | 0.211/0.258 | 0.201/0.254 | 0.210/0.247 |
| No. atoms | |||
| Protein | 55006 | 54790 | 55297 |
| Ligand/ion | 1536 | 1664 | 1651 |
| Water | 58 | 895 | 775 |
| B-factors | |||
| Protein | 48.0 | 31.2 | 49.1 |
| Ligand/ion | 68.0 | 65.3 | 80.4 |
| Water | 12.9 | 11.7 | 27.4 |
| R.m.s deviations | |||
| Bond lengths (Å) | 0.004 | 0.008 | 0.004 |
| Bond angles (°) | 0.79 | 0.95 | 0.69 |
Highest resolution shell is shown in parenthesis.
Supplementary Material
Acknowledgments
We thank N. Woychik for MazF-mt3 protein. Work was supported by NSF grant CHE-1361462 (J.K.L.), Welch Foundation Grant A-1763 (C.D.K.), Czech Science Foundation 15-05228S (L.K., N.P.), and NIH grants NIEHS P30 ES005022, GM097260 (C.D.K.), GM041376 (R.H.E.), GM088343 (B.E.N.), GM096454 (B.E.N.), and GM115910 (B.E.N.).
Footnotes
Author Contributions
L.K., J.K.L., C.D.K., R.H.E., and B.E.N. designed experiments; J.G.B. (Figures 1, 2, 3, E1, E3, E4, E5), Y.Z. (Figures 4, E7), Y.T. (Figures E2, E8), N.P. (Figures 1, E1), I.B. (Figures 1, E1), L.G. (Figures E2, E8), and M.L. (Figures E2, E8) performed experiments; B.B. provided resources to perform experiments; J.G.B., Y.Z., Y.T., N.P., I.B., L.G., L.K., J.K.L., C.D.K., R.H.E., and B.E.N. performed data analysis; R.H.E. and B.E.N. wrote the paper.
The coordinates and structure factors of RPo-pppApC, RPo-NAD+pC, and RPo-dpCoApC have been deposited in the Protein Data Bank under accession numbers 5D4C, 5D4D, and 5D4E, respectively.
References
- 1.Topisirovic I, Svitkin Y, Sonenberg N, Shatkin A. Cap and cap-binding proteins in the control of gene expression. RNA. 2011;2:277–298. doi: 10.1002/wrna.52. [DOI] [PubMed] [Google Scholar]
- 2.Hui M, Foley P, Belasco J. Messenger RNA degradation in bacterial cells. Annu Rev Genet. 2014;48:537–559. doi: 10.1146/annurev-genet-120213-092340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Jaschke A, Hofer K, Nubel G, Frindert J. Cap-like structures in bacterial RNA and epitranscriptomic modification. Curr Opin Microbiol. 2016;30:44–49. doi: 10.1016/j.mib.2015.12.009. [DOI] [PubMed] [Google Scholar]
- 4.Kowtoniuk W, Shen Y, Heemstra J, Agarwal I, Liu D. A chemical screen for biological small molecule-RNA conjugates reveals CoA-linked RNA. Proc Natl Acad Sci USA. 2009;106:7768–7773. doi: 10.1073/pnas.0900528106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chen Y, Kowtoniuk W, Agarwal I, Shen Y, Liu D. LC/MS analysis of cellular RNA reveals NAD-linked RNA. Nature Chem Biol. 2009;5:879–881. doi: 10.1038/nchembio.235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Cahova H, Winz ML, Hofer K, Nubel G, Jaschke A. NAD captureSeq indicates NAD as a bacterial cap for a subset of regulatory RNAs. Nature. 2015;519:374–377. doi: 10.1038/nature14020. [DOI] [PubMed] [Google Scholar]
- 7.Shuman S. RNA capping: progress and prospects. RNA. 2015;21:735–737. doi: 10.1261/rna.049973.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Luciano D, Belasco J. NAD in RNA: unconventional headgear. Trends Biochem Sci. 2015;40:245–247. doi: 10.1016/j.tibs.2015.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Deana A, Celesnik H, Belasco J. The bacterial enzyme RppH triggers messenger RNA degradation by 5′ pyrophosphate removal. Nature. 2008;451:355–358. doi: 10.1038/nature06475. [DOI] [PubMed] [Google Scholar]
- 10.Ebright R. RNA polymerase: structural similarities between bacterial RNA polymerase and eukaryotic RNA polymerase II. J Mol Biol. 2000;304:687–698. doi: 10.1006/jmbi.2000.4309. [DOI] [PubMed] [Google Scholar]
- 11.Cramer P. Multisubunit RNA polymerases. Curr Opin Struc Biol. 2002;12:89–97. doi: 10.1016/s0959-440x(02)00294-4. [DOI] [PubMed] [Google Scholar]
- 12.Decker K, Hinton D. Transcription regulation at the core: similarities among bacterial, archaeal, and eukaryotic RNA polymerases. Annu Rev Microbiol. 2013;67:113–139. doi: 10.1146/annurev-micro-092412-155756. [DOI] [PubMed] [Google Scholar]
- 13.Zhang Y, et al. Structural basis of transcription initiation. Science. 2012;338:1076–1080. doi: 10.1126/science.1227786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Xu W, Jones CR, Dunn CA, Bessman MJ. Gene ytkD of Bacillus subtilis encodes an atypical nucleoside triphosphatase member of the Nudix hydrolase superfamily. J Bacteriol. 2004;186:8380–8384. doi: 10.1128/JB.186.24.8380-8384.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Westover K, Bushnell D, Kornberg R. Structural basis of transcription: nucleotide selection by rotation in the RNA polymerase II active center. Cell. 2004;119:481–489. doi: 10.1016/j.cell.2004.10.016. [DOI] [PubMed] [Google Scholar]
- 16.Zhang Y, et al. GE23077 binds to the RNA polymerase ‘i’ and ‘i+1’ sites and prevents the binding of initiating nucleotides. eLife. 2014;3:e02450. doi: 10.7554/eLife.02450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Basu R, et al. Structural basis of transcription initiation by bacterial RNA polymerase holoenzyme. J Biol Chem. 2014;289:24549–24559. doi: 10.1074/jbc.M114.584037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Artsimovitch I, Svetlov V, Murakami K, Landick R. Co-overexpression of Escherichia coli RNA polymerase subunits allows isolation and analysis of mutant enzymes lacking lineage-specific sequence insertions. J Biol Chem. 2003;278:12344–12355. doi: 10.1074/jbc.M211214200. [DOI] [PubMed] [Google Scholar]
- 19.Marr M, Roberts J. Promoter recognition as measured by binding of polymerase to nontemplate strand oligonucleotide. Science. 1997;276:1258–1260. doi: 10.1126/science.276.5316.1258. [DOI] [PubMed] [Google Scholar]
- 20.Perdue S, Roberts J. A backtrack-inducing sequence is an essential component of Escherichia coli σ70-dependent promoter-proximal pausing. Mol Microbiol. 2010;78:636–650. doi: 10.1111/j.1365-2958.2010.07347.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kaplan C, Larsson K, Kornberg R. The RNA polymerase II trigger loop functions in substrate selection and is directly targeted by α-amanitin. Mol Cell. 2008;30:547–556. doi: 10.1016/j.molcel.2008.04.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Schifano J, et al. An RNA-seq method for defining endoribonuclease cleavage specificity identifies dual rRNA substrates for toxin MazF-mt3. Nat Comm. 2014;5:3538. doi: 10.1038/ncomms4538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Vvedenskaya I, et al. Massively Systematic Transcript End Readout, “MASTER”: Transcription Start Site Selection, Transcriptional Slippage, and Transcript Yields. Mol Cell. 2015;60:953–965. doi: 10.1016/j.molcel.2015.10.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Baba T, et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Sys Biol. 2006;2 doi: 10.1038/msb4100050. 2006 0008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Goldman S, Nair N, Wells C, Nickels B, Hochschild A. The primary σ factor in Escherichia coli can access the transcription elongation complex from solution in vivo. eLife. 2015;4 doi: 10.7554/eLife.10514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Cabart P, Jin H, Li L, Kaplan C. Activation and reactivation of the RNA polymerase II trigger loop for intrinsic RNA cleavage and catalysis. Transcription. 2014;5:e28869. doi: 10.4161/trns.28869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Murakami K. Structural biology of bacterial RNA polymerase. Biomolecules. 2015;5:848–864. doi: 10.3390/biom5020848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Otwinowski Z, Minor W. Processing of X-ray diffraction data collected in oscillation mode. Meth Enzymol. 1997;276:307–326. doi: 10.1016/S0076-6879(97)76066-X. [DOI] [PubMed] [Google Scholar]
- 29.French S, Wilson K. On the treatment of negative intensity observations. Acta Crystallographica Section A. 1978;34:517–525. [Google Scholar]
- 30.Strong M, et al. Toward the structural genomics of complexes: crystal structure of a PE/PPE protein complex from Mycobacterium tuberculosis. Proc Natl Acad Sci USA. 2006;103:8060–8065. doi: 10.1073/pnas.0602606103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Vagin A, Teplyakov A. MOLREP: an Automated Program for Molecular Replacement. J Appl Crystallogr. 1997;30:1022–1025. [Google Scholar]
- 32.Emsley P, Lohkamp B, Scott W, Cowtan K. Features and development of Coot. Acta Crystallographica Section D, Biological Crystallography. 2010;66:486–501. doi: 10.1107/S0907444910007493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Adams PD, et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallographica Section D Biological Crystallography. 2010;66:213–221. doi: 10.1107/S0907444909052925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Malygin A, Shemyakin M. Adenosine, NAD and FAD can initiate template-dependent RNA synthesis catalyzed by Escherichia coli RNA polymerase. FEBS Lett. 1979;102:51–54. doi: 10.1016/0014-5793(79)80926-6. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.