Abstract
In animal gonads PIWI-clade Argonaute proteins repress transposons sequence-specifically via bound piRNAs. These are processed from single-stranded precursor RNAs by largely unknown mechanisms. Here we show that primary piRNA biogenesis is a 3′ directed and phased process that, in the Drosophila germline, is initiated by secondary piRNA-guided transcript cleavage. Phasing results from consecutive endo-nucleolytic cleavages catalyzed by Zucchini, implying coupled formation of 3′ and 5′ ends of flanking piRNAs. Unexpectedly, Zucchini also participates in 3′ end formation of secondary piRNAs. Its function can, however, be bypassed by downstream piRNA-guided precursor cleavages coupled to exonucleolytic trimming. Our data uncover an evolutionarily conserved piRNA biogenesis mechanism where Zucchini plays a central role in defining piRNA 5′ and 3′ ends.
The piRNA pathway silences transposable elements (TEs) during animal gametogenesis. It utilizes PIWI-clade Argonaute proteins bound to ~23-30nt piRNAs, which act as sequence specific guides to specify targets via base-pair complementarity (1, 2). piRNAs are processed from single-stranded precursor transcripts via two biogenesis pathways, both of which are initiated by endo-nucleolytic definition of piRNA 5′ ends: 5′ ends of primary piRNAs (mostly loaded into Drosophila Piwi) are generated by Zucchini (3–8). They carry a 5′ uridine (1U) but otherwise appear to be derived randomly from their precursors (9–13). 5′ ends of secondary piRNAs (loaded into Drosophila Aub/AGO3) are specified via piRNA-guided slicing. Here, reciprocal cleavages of complementary transcripts (ping-pong cycle) define piRNA pairs, whose 5′ ends display a 10 nucleotide offset (14, 15). piRNA 3′ ends lack a nucleotide preference and the molecular events underlying their formation are elusive. It is thought that after 5′ end definition longer piRNA intermediates are loaded into PIWI proteins (16) and subsequently trimmed at their 3′ ends to mature piRNAs (17).
How are transcripts selected as piRNA biogenesis substrates? We noticed ectopic piRNA production from mRNAs in fly ovaries mutant for the Rhino-Deadlock-Cutoff complex (18–20). The corresponding piRNA profiles initiate abruptly (e.g. row; Fig. 1A, B) and some of these initiation sites are also apparent in wildtype ovaries (‘GFP control’). Consistent with suggestions that piRNA-guided transcript cleavage initiates 3′ directed piRNA biogenesis (21), we identified a target site in the row transcript with complementarity to a 1360 piRNA (‘trigger-piRNA’; Fig. 1C). The 5′ end of this piRNA maps ten nucleotides away from the 5′ end of the most abundant row piRNA (‘responder-piRNA’). While the 1360 trigger-piRNA occupies Aub and AGO3 equally, the row responder-piRNA resides mostly in Aub (Fig. 1B, C, (22)). In contrast, row piRNAs originating downstream of the responder-piRNA (‘trail-piRNAs’) occupy Piwi and only to a small extent Aub (Fig. 1B). We identified 570 mRNAs with similar characteristics. On average, responder-piRNAs populate predominantly Aub but are also present in AGO3 and Piwi (Fig. 1D). Trail-piRNAs instead are funneled into Piwi (~85%), moderately into Aub (~15%) but not into AGO3. When considering all piRNAs mapping within 200nt downstream of the trigger site, 70% of Aub/AGO3-bound piRNAs, but only 7% of Piwi-bound piRNAs, correspond to the responder-piRNA (Fig. 1D).
Figure 1. Aub/AGO3 mediated slicing triggers phased piRNA biogenesis.
(A) Normalized piRNA populations from Rhino depleted ovaries or from control ovaries mapping to the row mRNA.
(B) As in (A) but piRNAs bound to Aub/AGO3/Piwi (22) are shown individually.
(C) Alignment of the 1360 trigger-piRNA with the row mRNA (triangle: slicer cleavage position).
(D) Metaplots showing profiles of Aub/AGO3/Piwi-bound piRNAs (from Rhino-depleted ovaries) at genic trigger sites (profiles represent the median of normalized values; responder peak: 100%).
(E) Heatmap indicating piRNA levels (Σ Aub/AGO3 5′ ends) mapping antisense (≤6 mismatches) to 570 mRNAs with trigger events (position 0: responder-piRNA 5′ end). The binary histograms show the percentage of transcripts with a cloned 5′ end of indicated piRNAs mapping in sense/antisense orientation at nucleotide resolution.
(F) Heatmaps indicating AGO3/Aub/Piwi-bound piRNA 5′ end levels from Rhino depleted ovaries in a window centered on 570 genic responder-piRNAs. The corresponding binary histograms indicate the percentage of transcripts that exhibit a cloned piRNA 5′ end at a given position.
Only 8% of the 570 triggered transcripts harbor fully complementary target sites for Aub/AGO3-bound piRNAs at the responder position (Fig. S1A). However, when allowing up to six mismatches, piRNA 5′ ends exhibit a high likelihood to align precisely ten nucleotides offset from responder-piRNA 5′ ends (Fig. 1E). We could identify trigger-piRNAs in ~75% of the cases (Fig. S1B-G; (22)). Based on this, slicing characteristics of Aub/AGO3 follow known features of Argonaute proteins (23–25). Trigger-piRNAs also conform to rules of heterotypic ping-pong (14, 15) (Fig. S1E).
5′ ends of trail-piRNAs are immediately downstream of responder-piRNA 3′ ends (Fig. 1D) and trail-piRNAs in Piwi and Aub display pronounced phasing (Fig. 1F). The phase is ~27nt, which is precisely one average piRNA length (23-30nt; Fig. 1F). Both phasing accuracy and piRNA levels decrease with increasing distance from the trigger site suggesting that piRNA biogenesis occurs in a 3′ directed, processive fashion. To investigate whether slicing generally initiates directional and phased piRNA biogenesis we analyzed TE-mapping piRNAs from wildtype ovaries. Piwi/Aub/AGO3-bound piRNAs mapping in vicinity of abundant ping-pong piRNAs display patterns virtually identical to the ones described above (Fig. S2A, B). Biogenesis of Piwi-bound piRNAs (by definition primary piRNAs) is therefore largely a consequence of piRNA-guided target cleavage.
To investigate how tight piRNA 3′ end formation is coupled to 5′ end formation of the immediate downstream piRNA we analyzed piRNA length and 3′ end patterns. On average, piRNAs exhibit a broad length distribution (Fig. 2A-C) (14). We noticed that individual piRNAs sharing the same 5′ end often have distinct length profiles and 3′ ends (S3A). We therefore grouped all piRNAs with unambiguous 5′ ends into length cohorts (Fig. 2A-C; S3B; (22)). Length profiles of AGO3-bound piRNAs are the least defined while those bound to Piwi display the most accurate 3′ ends, irrespective of whether they are processed in the germline or soma (Fig. S3C-E). No obvious nucleotide-bias within piRNAs correlates with their distinct length groups. However, uridine (U) residues are enriched immediately downstream of dominant piRNA 3′ ends (Fig. 2D, E; S3F-J). Length cohorts with two dominant species display a downstream U-bias for both prominent 3′ ends (e.g. clusters 5 in Fig. S3H-J). The downstream U-bias is strongest for Piwi-bound and weakest for AGO3-bound piRNAs, where it is only evident for the most accurately defined length populations (Fig. 2E; S3G). As Piwi/Aub-bound piRNAs display a 1U bias and primary piRNAs exhibit strong phasing, the pronounced downstream U-bias implies that piRNA 3′ end formation simultaneously specifies the 5′ end of the downstream piRNA. Indeed, when Piwi-bound piRNAs are aligned at their dominant 3′ ends, neighboring piRNA 5′ ends are highly enriched at the +1 position, irrespective of the tissue (Fig. S4A).
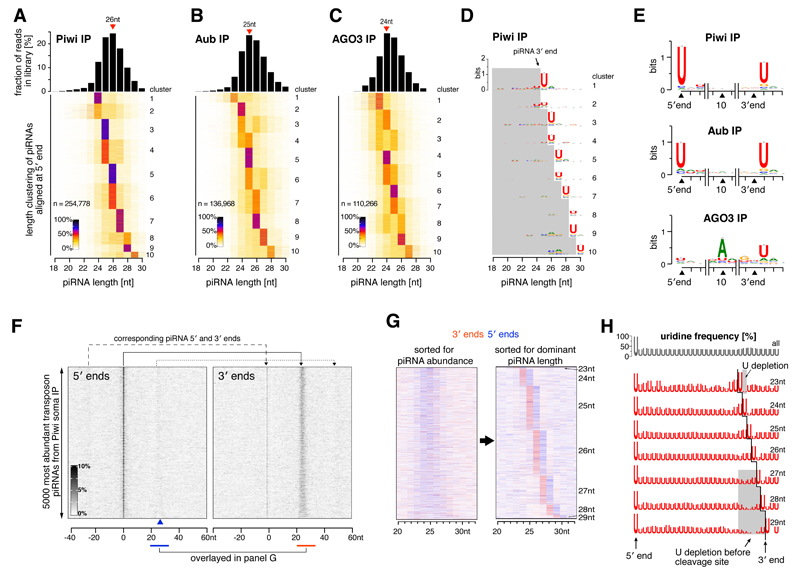
Figure 2. Biogenesis of flanking primary piRNAs is coupled.
(A-C) Top: piRNA length histograms; bottom: heatmaps showing length groups for piRNAs in Piwi/Aub/AGO3 (5′ ends with similar length profiles were grouped using K-means clustering).
(D) Sequence logos indicating nucleotide biases for piRNA length clusters defined in (A) (grey box: piRNA body; for Aub/AGO3 see Fig. S3).
(E) Sequence logos indicating nucleotide biases for Piwi/Aub/AGO3-bound piRNAs at 5′ ends, position 10 and 3′ ends (only 3′ ends where ≥50% of piRNAs terminate were considered).
(F) Heatmaps displaying somatic Piwi-bound piRNA 5′ or 3′ ends in a window around the most abundant TE piRNAs (sum of each line scaled to 100%; sorted according to piRNA level at position 0).
(G) Heatmaps displaying 5′ or 3′ end counts of piRNAs mapping to positions 20-32 downstream of major piRNA 5′ ends (detail from (F)). The left plot is sorted as in (F), the right plot is resorted for dominant piRNA length species (see also Fig. S4B).
(H) Sequence logos indicating uridine frequency along piRNA sequences of different length cohorts (defined in S4B; black line: dominant piRNA 3′ ends).
To investigate 3′ end formation further we focused on somatic Piwi-bound piRNAs. We selected the 5,000 most abundant TE antisense piRNA 5′ ends, sorted them for their abundance and displayed 5′ and 3′ ends of all piRNAs mapping in their immediate vicinity (Fig. 2F). 3′ ends of neighboring piRNAs are enriched precisely one nucleotide upstream of selected 5′ ends (position 0; dashed arrow). 5′ ends of downstream piRNAs instead display a fuzzy enrichment around +27nt (blue arrow head). However, resorting the heatmap according to increasing length of the dominant piRNA species for each 5′ end at position 0 resolves the strong enrichment of 5′ ends precisely 1nt downstream of dominant 3′ ends (Fig. 2G; S4B).
The obtained piRNA length cohorts (23-29nt; Fig. 2G; S4B) display a strong downstream U-bias (Fig. 2H). U-residues are depleted upstream of 3′ ends of unusually long piRNA cohorts (27-29mers) and an inverse trend is seen for the 23mer piRNA cohort. This suggests that 3′ ends of Piwi-bound piRNAs are defined by an endonuclease that cleaves immediately upstream of a U-residue and 23-30nt downstream of the piRNA 5′ end. Asymmetrically distributed U-residues within the cleavage window force generation of atypically long or short piRNAs.
To experimentally test the phased biogenesis of primary piRNAs we generated transgenic flies expressing a piRNA biogenesis reporter harboring a piRNA target site (Fig. 3A). Without a target site, no piRNAs are produced (Fig. 3B). Insertion of a single complementary target site for an Aub/AGO3 piRNA represses GFP expression (Fig. S5), triggers generation of a ping-pong responder-piRNA and forces 3′ directed, phased biogenesis of trail-piRNAs (Fig. 3C). Trail-piRNA levels and phasing accuracy decrease with increasing distance from the trigger site but are detectable for several hundred nucleotides. To probe the impact of U-residues on biogenesis patterns, we constructed a sensor with four Us in regular 26nt intervals downstream of the trigger site. This increases levels and phasing accuracy of reporter derived piRNAs (Fig. 3D). Spacing of single Us in decreasing intervals of 28 to 23nt dictates piRNA 5′ ends but also defines piRNA 3′ ends precisely one nucleotide upstream (Fig. 3E). These findings reinforce the notion that 3′ and 5′ ends of adjacent Piwi-bound piRNAs are formed via a single endonucleolytic cleavage upstream of a U residue.
Figure 3. Primary piRNA biogenesis is continuous and guided by uridine residues.
(A) Cartoon of the piRNA biogenesis reporter with target site for an Aub/AGO3-bound piRNA.
(B, C) Normalized small RNA 5′ end profiles from reporter lacking a piRNA target site (B) or from reporter with target site for an Aub/AGO3-bound piRNA (C; grey: antisense reads; red: sense reads; blue bars: U residues; numbers: normalized piRNA counts; histograms: length profile and 1U bias of trail-piRNAs).
(D) Similar to (C), but reporter contains 4 Us every 26nt.
(E) Similar to (C), but reporter contains single Us in indicated intervals.
The endonuclease Zucchini generates primary piRNA 5′ ends (3–8) and is therefore also the prime candidate for their 3′ end formation. A direct test of this is precluded as primary piRNAs depend on Zucchini. We therefore asked whether Zucchini is involved in 3′ end formation of secondary piRNAs whose 5′ ends are formed via slicing. Many Aub/AGO3-bound secondary piRNAs are generated in Zucchini-depleted ovaries (26) and we used their 5′ ends as anchor points (Fig. S6A). In wildtype ovaries, piRNAs mapping to these 5′ ends display the characteristic nucleotide bias at the first position (1U for Aub piRNAs) and the tenth position (10A for AGO3 piRNAs; Fig. 4A). They also exhibit a downstream U-bias. In Zucchini depleted ovaries, the length of these piRNAs and therefore their 3′ ends are altered and the downstream U-bias is lost, implicating Zucchini in 3′ end formation (Fig. 4A, S6B, C). The downstream U-bias is therefore most likely a fingerprint of Zucchini, making it the central nuclease in piRNA biogenesis. The less severe impact on AGO3-bound piRNAs and their weaker downstream U-bias suggests that many AGO3-bound piRNAs are Zucchini-independent or are resected at their 3′ ends in wildtype ovaries.
Figure 4. Zucchini is involved in 3′ end formation of Drosophila and mouse piRNAs.
(A) Sequence logos for nucleotides at 5′ end, 10th position and 3′ end of Aub-bound or AGO3-bound piRNAs in control or Zucchini depleted ovaries.
(B) Normalized wildtype small RNA 5′ end profile for reporter (similar to Fig. 3D) with single target site for a Zucchini-independent piRNA.
(C) As in (B) but piRNAs isolated from Zucchini depleted ovaries.
(D, E) As in (C) but reporter contains target sites for two (D) or three (E) Zucchini-independent piRNAs.
(F) Heatmaps showing length cohorts of Mili-bound piRNAs from Tdrkh heterozygous mouse testes.
(G) Sequence logos showing nucleotide composition around dominant piRNA 3′ ends for length cohorts defined in (G); for individual length clusters see Fig. S6.
(H) Frequency of Mili-bound piRNA 5′ ends around aligned dominant piRNA 3′ ends from Tdrkh heterozygous testes.
(I-K) As in panels F-H but from Tdrkh mutant testes.
To experimentally test Zucchini’s involvement in 3′ end formation of secondary piRNAs, we constructed a biogenesis reporter with a single target site for a Zucchini-independent piRNA. In wildtype ovaries this reporter gives rise to a prominent ping-pong responder and to trail-piRNAs (Fig. 4B). As expected, loss of Zucchini ablates trail-piRNAs (Fig. 4C; S2C). But although trigger-piRNA levels and reporter silencing were unaffected, levels of the responder-piRNA also dropped >30-fold (Fig. 4C, S6D). Zucchini is therefore centrally involved in 3′ end formation of ping-pong piRNAs. The piRNA target site in this reporter is from the F-element, one of the TEs that maintain high levels of secondary piRNAs in Zucchini-deficient ovaries (Fig. S6E). How then are F-element piRNAs generated independently of Zucchini? The central difference between the F-element and the reporter is that the latter contains only a single Aub/AGO3 target site. We added a downstream target site for a second Zucchini-independent piRNA, which would generate a ~50nt long piRNA intermediate. This leads to recovery of the first but not the second responder-piRNA in Zucchini depleted ovaries (Fig. 4D, S6F). Introducing a target site for a third Zucchini independent piRNA leads to recovery of the first two responder-piRNAs (Fig. 4E, S6G). A downstream slicer cleavage can therefore bypass Zucchini’s role in piRNA 3′ end formation. This must be coupled to exonucleolytic trimming, consistent with the existence of an activity that resects intermediates to mature piRNAs (17).
To uncover an involvement of mouse Zucchini (MitoPLD; (8, 27)) in piRNA 3′ end formation, we searched for a downstream U-bias in published piRNA datasets. Although mouse piRNA populations group into well-defined length cohorts, they lack a downstream U-bias (Fig. 4F, G; S7A; (28)). Similarly, piRNA 3′ ends and adjacent 5′ ends display no coupling signature (Fig. 4H), suggesting fundamental differences in piRNA 3′ end formation between flies and mouse. However, Mili-bound primary piRNAs are extensively trimmed at their 3′ ends (28). While mature Mili-bound piRNAs are ~24-28nt long, Mili associates with ~30-40nt piRNA intermediates harboring mature 5′ ends in Tdrkh/Papi mutants (Fig. S7B, C). This suggests that Tdrkh recruits an exonuclease to PIWI-proteins to facilitate piRNA precursor trimming (28, 29). Trimming might therefore occlude the downstream U-bias defined by MitoPLD. Indeed, Mili-bound piRNA intermediates from Tdrkh mutants display a strong downstream U-bias and exhibit strong 3′/5′ coupling (Fig. 4I-K; S7D). We conclude that coupled biogenesis of neighboring piRNAs is conserved between Drosophila and mouse.
Altogether, a model for piRNA biogenesis can be drawn (Fig. S8). A critical first step is the specification of piRNA precursors, leading to the endo-nucleolytic definition of a piRNA 5′ end. In the fly germline, the dominating process is piRNA-guided target slicing, which specifies the 5′ end of a responder-piRNA. 3′ end formation of this piRNA can occur via a second slicer cleavage event, which liberates a piRNA intermediate for exonucleolytic trimming. Alternatively, 3′ end formation is catalyzed by Zucchini, which cleaves the precursor upstream of a U-residue. Zucchini-mediated 3′ end formation promotes phased and 3′ directed primary piRNA biogenesis. In flies, Zucchini cleavage products seem directly compatible with Piwi binding. In mouse, MitoPLD cleavage products are too long, making 3′ end trimming essential. How primary piRNA biogenesis initiates in Drosophila ovarian somatic cells or adult mouse testes remains elusive.
Supplementary Material
Acknowledgments
We thank J. Gokcezade for fly injections, Harvard TRiP and Bloomington stock centers for flies, D. Jurczak for bioinformatics support and the CSF NGS unit for Illumina sequencing. This work was supported by the Austrian Academy of Sciences, the European Community (ERC # 260711EU), the Austrian Science Fund (Y 510-B12) and by SNF and HFSP fellowships to FM. Sequencing datasets are deposited at GEO (GSE64802, GSE55842).
References
- 1.Malone CD, Hannon GJ. Cell. 2009;136:656. doi: 10.1016/j.cell.2009.01.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Siomi MC, Sato K, Pezic D, Aravin AA. Nat Rev Mol Cell Biol. 2011;12:246. doi: 10.1038/nrm3089. [DOI] [PubMed] [Google Scholar]
- 3.Pane A, Wehr K, Schupbach T. Developmental cell. 2007;12:851. doi: 10.1016/j.devcel.2007.03.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ipsaro JJ, Haase AD, Knott SR, Joshua-Tor L, Hannon GJ. Nature. 2012;491:279. doi: 10.1038/nature11502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Nishimasu H, et al. Nature. 2012;491:284. doi: 10.1038/nature11509. [DOI] [PubMed] [Google Scholar]
- 6.Saito K, et al. Genes Dev. 2010;24:2493. doi: 10.1101/gad.1989510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Olivieri D, Sykora MM, Sachidanandam R, Mechtler K, Brennecke J. EMBO J. 2010;29:3301. doi: 10.1038/emboj.2010.212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Watanabe T, et al. Dev Cell. 2011;20:364. doi: 10.1016/j.devcel.2011.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Malone CD, et al. Cell. 2009;137:522. doi: 10.1016/j.cell.2009.03.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lau NC, et al. Genome Research. 2009 [Google Scholar]
- 11.Aravin A, et al. Nature. 2006;442:203. doi: 10.1038/nature04916. [DOI] [PubMed] [Google Scholar]
- 12.Girard A, Sachidanandam R, Hannon GJ, Carmell MA. Nature. 2006;442:199. doi: 10.1038/nature04917. [DOI] [PubMed] [Google Scholar]
- 13.Li XZ, et al. Mol Cell. 2013;50:67. doi: 10.1016/j.molcel.2013.02.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Brennecke J, et al. Cell. 2007;128:1089. doi: 10.1016/j.cell.2007.01.043. [DOI] [PubMed] [Google Scholar]
- 15.Gunawardane LS, et al. Science. 2007;315:1587. doi: 10.1126/science.1140494. [DOI] [PubMed] [Google Scholar]
- 16.Vourekas A, et al. Nat Struct Mol Biol. 2012;19:773. doi: 10.1038/nsmb.2347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kawaoka S, Izumi N, Katsuma S, Tomari Y. Mol Cell. 2011;43:1015. doi: 10.1016/j.molcel.2011.07.029. [DOI] [PubMed] [Google Scholar]
- 18.Klattenhoff C, et al. Cell. 2009;138:1137. doi: 10.1016/j.cell.2009.07.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhang Z, et al. Cell. 2014;157:1353. doi: 10.1016/j.cell.2014.04.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Mohn F, Sienski G, Handler D, Brennecke J. Cell. 2014;157:1364. doi: 10.1016/j.cell.2014.04.031. [DOI] [PubMed] [Google Scholar]
- 21.Shpiz S, Ryazansky S, Olovnikov I, Abramov Y, Kalmykova A. PLoS Genetics. 2014;10:e1004138. doi: 10.1371/journal.pgen.1004138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.see supplementary online material
- 23.Swarts DC, et al. Nature Struct & Mol Biol. 2014;21:743. doi: 10.1038/nsmb.2879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wee LM, Flores-Jasso CF, Salomon WE, Zamore PD. Cell. 2012;151:1055. doi: 10.1016/j.cell.2012.10.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Reuter M, et al. Nature. 2011;480:264. doi: 10.1038/nature10672. [DOI] [PubMed] [Google Scholar]
- 26.Olivieri D, Senti KA, Subramanian S, Sachidanandam R, Brennecke J. Mol Cell. 2012;47:954. doi: 10.1016/j.molcel.2012.07.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Huang H, et al. Dev Cell. 2011;20:376. doi: 10.1016/j.devcel.2011.01.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Saxe JP, Chen M, Zhao H, Lin H. EMBO J. 2013;32:1869. doi: 10.1038/emboj.2013.121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Honda S, et al. RNA. 2013;19:1405. doi: 10.1261/rna.040428.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.