Abstract
Background
Acute myeloid leukemia (AML) is commonly characterized by several chromosomal abnormalities resulting in the formation of chimeric genes that play various roles in leukemogenesis. Translocations resulting in the ETV6-ABL1 fusion gene are rare in AML and other hematologic malignancies with only thirty-two previously reported cases in the literature, five of which were AML.
Findings
Herein, we report the case of a 73-year-old male with acute myeloid leukemia arising from MDS, negative for PDGFRA and PDGFRB, positive for bone marrow eosinophilia, rash, and marked fluid retention, which improved dramatically with imatinib therapy. Conventional cytogenetics revealed a t(3;9)(p25;q34), t(5;18)(q13;p11.2), and additional material of unknown origin at 12p11.2 in 2 out of 10 metaphases analyzed. Interphase FISH studies showed evidence of ETV6 (12p13) and ABL1 (9q34) rearrangements in 41.3 % and 5.7 % of the cells respectively. FISH studies on previously G-banded metaphases showed colocalization of ABL1 and ETV6 signals to the short arm of chromosome 3 at 3p25 suggesting a possible ETV6-ABL1 fusion. Subtelomeric metaphase FISH studies also showed the presence of a subtelomere 3p signal on the long arm of the derivative 9, and no subtelomere 3p signal on the derivative chromosome 12.
Conclusions
These findings suggest a complex rearrangement involving an insertion of ETV6 into 3p25 followed by a reciprocal translocation involving 3p25 and 9q34, resulting in a possible ETV6-ABL1 fusion. This case highlights the importance of FISH to characterize complex rearrangements in myeloid malignancies, particularly those resulting in clinically significant chimeric genes.
Keywords: Cytogenetics, FISH, AML, MDS, ETV6-ABL1, Imatinib
Introduction
The ETV6-ABL1 fusion gene is uncommon in hematological malignancies, including chronic myeloid leukemia (CML), acute myeloid leukemia (AML), and acute lymphoblastic leukemia (ALL), and has been reported in only thirty-two patients [1]. Six reported cases of hematological malignancies bearing ETV6-ABL1 in the context of complex rearrangements involving additional translocation partners have been reported, resulting in the translocation of the fusion gene to a third derivative chromosome [2–7]. The rarity of this event is due in part to the opposite transcriptional orientation of ETV6 and ABL1 relative to the centromere, which requires at least three separate chromosomal breaks to form a functional fusion gene [8]. The structure and function of the ETV6-ABL1 oncoprotein is very similar to that of the BCR-ABL1 protein with the ETV6 helix loop helix domain (HLH) deregulating the kinase activity of ABL1 leading to activation of a non-receptor tyrosine kinase that initiates downstream pathways affecting growth rate, cellular survival, and independence as well as transforming capacity [1, 3]. Because of the common functional activity with the BCR-ABL1 fusion protein, ETV6-ABL1 positive patients have been observed to respond to therapy with tyrosine kinase inhibitors, albeit at varying degrees and with high likelihood of relapse [1].
Herein, we present the case of a 73-year-old male diagnosed with acute myeloid leukemia. Cytogenetic analysis revealed a t(5;18) and a t(3;9), as well as additional material of unknown origin on the short arm of chromosome 12. Additional interphase and metaphase FISH studies revealed an insertion of ETV6 into 3p and translocation of ABL1 to the same locus on 3p, resulting in a possible ETV6-ABL1 fusion gene. The patient responded transiently to imatinib therapy, but eventually relapsed and expired.
Case presentation
The patient was a 73-year-old male with acute myeloid leukemia (AML) and hypereosinophilia, arising from antecedent myelodysplastic syndrome (MDS). He was initially found to have thrombocytopenia fifteen months prior to transfer during a pre-surgical workup for surgery to treat carpal tunnel syndrome. A bone marrow biopsy performed six months later had findings consistent with myelodysplastic syndrome with fewer than 5 % blasts in the bone marrow. He subsequently received three cycles of decitabine: the first dose was given in February 2015, the second dose was given in May 2015, and the third dose was given in July 2015. Eight months after bone marrow biopsy, he presented to an outside hospital with a fever and was found to have leukocytosis with circulating blasts, and a repeat bone marrow biopsy identified AML, possibly acute eosinophilic leukemia, with 20 % blasts identified in the bone marrow. Broad-spectrum antibiotics were started and the patient was transferred to UCLA for escalation of care. Shortly after transfer, he developed progressive renal failure requiring dialysis. Persistent blasts were treated with azacytidine, but he developed severe pancytopenia. In addition, eosinophilia, rash and marked fluid retention led his clinical team to consider therapy with imatinib, which promptly led to resolution of those findings. A follow-up bone marrow aspiration and biopsy one month later identified a hypercellular marrow showing marked eosinophilia with increased atypical immature forms, markedly reduced myeloid precursors other than the eosinophilic series including increased atypical immature eosinophils, reduced erythropoiesis and megakaryopoiesis, and increased blasts (10-11 % of the marrow elements). The overall marrow histology was consistent with acute myeloid leukemia possibly, acute myelocytic leukemia.
Material and methods
Conventional cytogenetics
Chromosome analysis was performed using standard cytogenetic techniques on the bone marrow samples from this patient. The karyotypes were prepared using the Applied Imaging CytoVision software (Applied Imaging, Genetix, Santa Clara, CA) and described according to the ISCN 2013 nomenclature [9].
FISH
Fluorescence in situ hybridization (FISH) was performed on interphase nuclei using the Vysis BCR/ABL1/ASS1 Tri-color DF FISH Probe Kit, Vysis LSI BCR/ABL Dual Color, Dual Fusion Probe Kit, and Vysis ETV6 Break Apart FISH Probe Kit from Abbott Molecular (Des Plaines, Illinois 60018) on interphase nuclei. Additionally, metaphase FISH was performed with the TotelVysion 3p, Spectrum Green, TotelVysion 3q Spectrum Orange probes, as well as the previously mentioned probes on previously G-banded metaphases.
Results
Conventional cytogenetics
Conventional cytogenetics revealed a t(3;9)(p25;q34), t(5;18)(q13;p11.2), and additional material of unknown origin at 12p11.2 in 2 out of 10 metaphases analyzed. The remaining 8 metaphases were cytogenetically normal (Fig. 1).
Fig. 1.
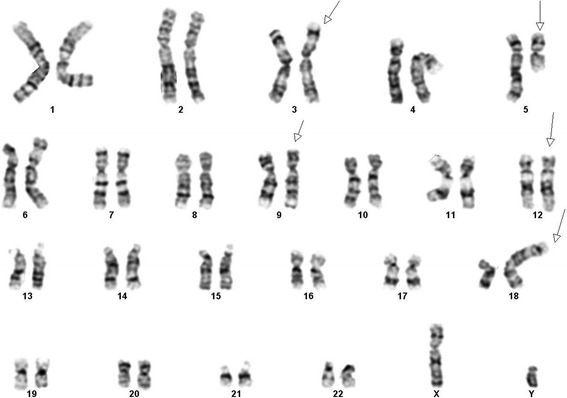
G-banded karyotype showing a three-way rearrangement involving 3, 9, and 12 as well as reciprocal translocation between chromosomes 5 and 18
FISH
Interphase FISH studies confirmed a rearrangement in 41.3 % (124/300) of nuclei examined involving ETV6 using the Vysis ETV6 Break Apart FISH Probe Kit and a rearrangement involving ABL1 in 5.7 % (17/300) nuclei examined using Vysis BCR/ABL1/ASS1 Tri-color DF FISH Probe Kit. These findings were described as (Figs. 2 and 3):
nuc ish(ASS1x2,ABL1x3,BCRx2)[17/300]
nuc ish(ETV6x2)(5'ETV6 sep 3'ETV6x1)[124/300]
Fig. 2.
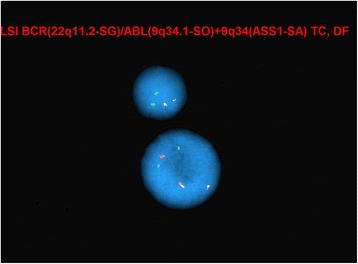
Interphase FISH showing ABL1 rearrangement (evidenced by additional red ABL1 signal)
Fig. 3.
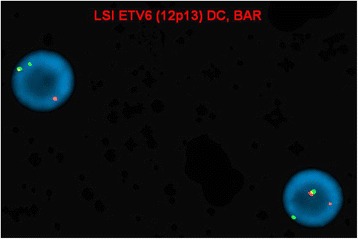
Interphase FISH showing ETV6 rearrangement (evidenced by split red and green ETV6 signals)
Metaphase FISH studies using the same probes on previously G-banded metaphases showed colocalization of ABL1 and ETV6 signals to the short arm of chromosome 3, suggesting the presence of an ETV6-ABL1 fusion gene. Subtelomeric metaphase FISH studies also showed the presence of a subtelomere 3p signal on the derivative 9q, and no subtelomere 3p signals on the derivative 12. In light of conventional cytogenetic findings, the karyotype was conveyed as follows (Figs. 4, 5 and 6):
46,XY,der(3)ins(3;12)(p25;p13p13)t(3;9)(p25;q34),t(5;18)(q13;p11.2),der(9)t(3;9),der(12)ins(3;12)(p25;p13p13)add(12)(p13)[2]/46,XY[8]
Fig. 4.
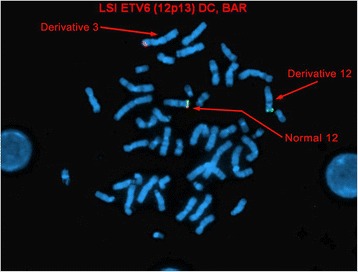
Metaphase FISH showing localization of 5’ ETV6 red signal to the short arm of the derivative chromosome 3. The derivative chromosome 12 only shows the 3’ ETV6 green signal
Fig. 5.
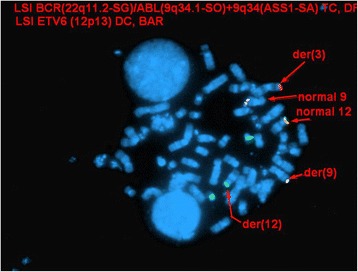
Metaphase FISH showing colocalization of ABL1 (red signal) and 5’ ETV6 (red signal) signals to the short arm of chromosome 3 using the BCR/ABL1/ASS1 Tri-color DF FISH Probe Kit and ETV6 Break Apart FISH Probe Kit
Fig. 6.
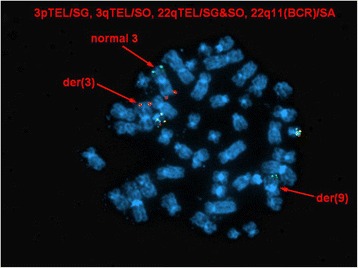
Metaphase FISH showing localization of subtelomere 3p signal on the derivative chromosome 9 and no subtelomeric 3 signals on the derivative chromosome 12
Discussion
This case highlights the formation of a potential ETV6-ABL1 fusion gene as a result of a complex, three-way rearrangement. The conventional and molecular cytogenetic findings in this case suggest an insertion of ETV6 in the short arm of the derivative chromosome 3 followed by a reciprocal translocation involving the same derivative chromosome 3 and ABL1 (9q34), resulting in the potential fusion gene. The breakpoint on chromosome 3 at which the aforementioned rearrangements occurred - 3p25 - harbors ANKRD28 (3p25.1), which has been implicated in AML in the context of t(3;11)(p25;p15) involving NUP98 (11p15) [10]. In other studies, 3p25 was found to be the most frequently deleted chromosomal band on 3p in AML [11]. Given this information, the involvement of 3p25 in this three-way rearrangement may result in deregulation of particular target genes relevant to leukemogenesis in this region. The translocation that occurred concomitant to the three-way rearrangement - t(5;18)(q13;p11.2) - has not been observed in AML and has not been associated with any clinical or hematopathologic features.
In total, there have been thirty-two reported cases of ETV6-ABL1 fusion gene in numerous hematologic malignancies including eleven cases of acute lymphoblastic leukemia, five cases of acute myeloid leukemia, and sixteen cases of myeloproliferative neoplasms (including CML) (Table 1). The rarity of this chromosomal rearrangement is thought to be due in part to the opposite transcriptional orientation of two genes relative to the centromere, which requires at least three break-and-join events for an in-frame fusion transcript to be formed. The rearrangement is often not detected using conventional cytogenetic techniques because of its cryptic nature due to the similar G-banding pattern of the distal long arm of chromosome 9 and the distal short arm of chromosome 12 [12]. Additionally, it has been observed that commercially available ABL1 FISH probes may not detect aberrations in the gene in this context, suggesting that the abnormality may remain undetected in a number of cases. In interphase cells, for example, the resulting ABL1 signal can be disproportionately small and can potentially be considered as noise and disregarded [1]. Thus, the rarity of the ETV6-ABL1 fusion is not only due to the multi-step mechanism required for its formation, but also because of technological limitations of FISH probes and molecular cytogenetic analysis.
Table 1.
Reported cases of ETV6-ABL1 fusion in acute lymphoblastic leukemia (ALL), acute myeloid leukemia (AML), and myeloproliferative neoplasm (MPN), including chronic myelogenous leukemia (CML)
| Reference | Age/Sex | Karyotype | FISH | Transcript |
|---|---|---|---|---|
| Acute lymphoblastic leukemia (ALL) | ||||
| [23] | 22 mo/F | N/A | N/A | Type A |
| [24] | 4 yo/M | 47,XXYc,del(6)(q15q23)[35]/47,XXYc[7] | Intact ETV6 (5’ and 3’) on derivative 9 | Type A and B |
| [25] | 30 yo/M | 45,XY,der(1)t(1;9)(q41;q?),der(9)t(9;13)(q?;q12.3),del(9p21.1 ~ p23),-13,der(16)t(16;22)(p13.3;q11.2) | ABL1 signal on apparently normal chromosome 12 | Type A and B |
| [26] | 8 yo/F | N/A | N/A | Type A and B |
| [7] | 8 yo/M | 46,XY, der(1)inv(1) (p11p34.2)t(1;9)(p11;p21)del(1)(q41),der(9)t(9;12) (q34.3;p13.3),der(9)t(1;9)(p11;p21),der(12)t(1;9;12) (q41?;q34.3;p13.3)[14]/46,XY)[5] | Normal signal patterns | Type A and B |
| [7] | 5 yo/M | 46,XY[20] | Normal signal patterns | Type A and B |
| [7] | 33 yo/F | 46,XX,t(8;9;12)(p12;q34;p13)[22] | ABL1 signal on 12p | Type A and B |
| [13] | 25 yo/F | 46,XX,del(9)(p22),der(10)t(9;10)(q22; p15)[12]/46,XX[8] | N/A | Type A and B |
| [27] | 58 yo/M | 46,XY,t(9;12)(q34;p13)[2]/45,XY, −2,−14,+17[2]/46,XY[16] | N/A | Type A and B |
| [27] | 49 yo/F | 46,XY,t(9;12)(q34;p13)[10] | N/A | Type A and B |
| [12] | 30 yo/F | 47,XX,+5[19]/46,XX[1] | Extra ETV6 and ABL1 signals in interphase nuclei | Type A and B |
| Acute myeloid leukemia (AML) | ||||
| [3] | 81 yo/M | t(9;12;14)(q34;p13;q22) in the context of complex karyotype | ETV6 signal on derivative 14 with concomitant deletion of other ETV6 allele | Type B |
| [4] | 29 yo/F | 46,XY,t(8;12)(p21;p13)[15] | 5’ ETV6 and ABL1 signals on derivative 8 | Type A |
| [4] | 48 yo/F | 46,XY,t(9;12)(q34;p13)[18/20]/51,XY,+8,+9,t(9;12)(q34;p13),+12,+14,+17[2] | 5’ ETV6 signal on derivative 9 | Type B |
| [14] | 38 yo/M | 49,XY,+11,t(9;12)(q34;p1?), +der(12)t(9;12),+19,der(22)t(1;22)(q21;q11) | ETV6-ABL1 fusion signals on 2 homologues of chromosome 12 | Type A and B |
| [13] | 52 yo/M | 46,XY[20] | ABL1 signal on apparently normal chromosome 12 | Type B |
| Myeloproliferative neoplasm (MPN), including chronic myeloid leukemia (CML) | ||||
| [28] | 49 yo/M | N/A | N/A | Type B |
| [2] | 32 yo/M | t(12;14)(p12;q11-13)[24]/46,XY[1] | 5’ ETV6 on apparently normal chromosome 9 | Type B |
| [24] | 59 yo/M | 46,XY,del(6)(p21),?t(9;12)(q34;p12)[16] | 5’ ETV6 on derivative 9q | Type A and B |
| [29] | 44 yo/F | 46,XX,t(9;12)(q34;p13)[16]/46,XX[4] | 5’ ETV6 on derivative 9q | N/A |
| [30] | 53 yo/M | 46,XY[20] | Normal signal patterns | Type A and B |
| [20] | 36 yo/M | 45,XY,-7,t(9;12)(q34;q13)[2]/45,idem,t(12;13)(p12;q13)[10]/46,XY[13] | ABL1 signal on derivative 12 | Type A |
| [6] | 72 yo/M | 46,XY,t(12;17) (p11.2;p11.2) | ABL1 and ETV6 signals on derivative 17 | Type B |
| [31] | 65 yo/F | 46,XX,t(5;9)(q13;q34) | 3’ ABL1 on 12p | Type B |
| [32] | 57 yo/M | 46,XY | 3’ ABL1 on 12p | N/A |
| [18] | 24 yo/F | 46,XX | ETV6 signal on apparently normal chromosome 9 | Type A and B |
| [33] | 61 yo/F | 46,XX,add(9)(q34) | ETV6 signal on derivative chromosome 9 | N/A |
| [21] | 79 yo/M | 46,XY | ABL1 signal on apparently normal chromosome 12 | N/A |
| [22] | 36 yo/M | 46,XY,t(9;12)(q34;p13) | Extra ABL1 signal (interphase FISH) | Type B |
| [1] | 46/F | 46,XY,t(9;12)(q34;p13)[10] | Extra ETV6 and ABL1 signals (interphase FISH) | Type B |
| [5] | N/A | Complex rearrangement involving chromosomes 6, 9, and 12 | Suggestive of ETV6-ABL1 fusion | N/A |
| [34] | 31 yo/M | 46,XY,der(9)t(9;12)(q34;p13), del(12)(p13)[1] | ETV6 signal on derivative 9 | Type A and B |
Age/sex of patient, karyotype, salient FISH findings, and type of ETV6-ABL1 fusion transcript detected are provided
To date, five cases of AML bearing a possible ETV6-ABL1 fusion have been reported (Table 1). Two of the cases were designated M1, two were M6, and one was not reported. One of the cases showed a straightforward t(9;12)(q34;p13) with concurrent FISH studies showing the 3’ ABL1 signal on the 12p, albeit in the context of a complex karyotype [4]. Two of the cases showed rearrangements involving a third chromosome - Golub et al. reported a case of AML (M6) with a t(9;12;14)(q34;p13;q22) without additional FISH studies, and La Starza et al. reported a case of AML (M1) with a t(8;12)(p21;p13) that showed colocalization of 5’ ETV6 and 3’ ABL1 signals on 8p21 by FISH [3, 4]. The two remaining cases showed normal karyotypes by conventional cytogenetics, but showed 3’ ABL1 signals on 12p [13, 14]. Among all of these cases, reported secondary abnormalities observed included gains of chromosomes 8, 9, 11, 12, 14, 17, and 19, as well as a t(1;22) [4, 14].
The ETV6-ABL1 fusion includes a helix loop helix (HLH) domain of ETV6 and tyrosine kinase domain of ABL1, and each domain is necessary for constitutive phosphorylation to occur [4]. Millon et al. found that mice transplanted with ETV6-ABL1-positive hematopoietic stem cells developed CML-like myeloproliferative disease, and that the TEL pointed homology oligomerization domain was essential to ETV6-ABL1-driven leukemogenesis [15]. It is well known that the ABL1 kinase has altered catalytic specificity in human leukemia. Co-immunoprecipitation studies show that the ETV6-ABL1 fusion protein tends to form complexes with CrkL in Ba/F3 cells, and this interaction phosphorylates CrkL and possibly CrkII [16]. However, it is known that the in vitro studies tend to have a wider range of substrates than the cellular forms [16]. Further analysis of Crk and CrkL adaptor proteins show that they play an essential role in integrating signals from a wide variety of sources such as apoptotic cells, extracellular matrix molecules, and growth factors, and there is mounting evidence to indicate that these proteins are associated with human diseases including susceptibility to pathogens and cancer [17].
ETV6 along with five other genes, BCR, ZMIZ, EML, and Nup21 form chimeric transcripts with ABL1. There must be a joining of the 3’ sequence of ABL1 with the 5’ end of the partner genes, and most of these genes are associated with a wide spectrum hematologic malignancies. Despite this heterogeneity, there is likely a common pluripotent stem cell that gives rise to similar transduction pathways and transforming activity [1]. Due to diagnostic, prognostic, and treatment-related implications, these cases further underscore the use of FISH along with routine chromosome analysis to properly characterize rare, albeit clinically significant fusion genes.
Eosinophilia is a recurrent morphologic finding in ETV6-ABL1-positive myeloid malignancies [4]. Of the five known AML ETV6-ABL1 positive cases, three out of the five were reported to have an increased abnormal eosinophil count, consistent with the findings in the present case. Another finding common to most of the patients was leukocytosis and out of those three cases, two had both leukocytosis and eosinophilia [3, 4, 13, 14]. Each patient was treated with chemotherapy including, cytosine-arabinoside, idarubicin, etoposide, mitoxantrone, and cytarabine. The two patients treated with imatinib responded transiently with resolved fluid retention, eosinophilia, and leukocytosis; however, full remission was not achieved [14]. There was only one patient who achieved full cytogenetic and hematological remission 20 months after undergoing allogeneic hematopoietic stem cell transplantation, which suggests its effectiveness in the treatment of ETV6-ABL1-positive AML patients with eosinophilia and leukocytosis [4].
Although there is limited information about the pathogenesis of myeloid neoplasms positive for ETV6-ABL1, chronic myeloid leukemia (CML) positive for BCR-ABL1 has been well studied and the molecular mechanisms of leukemogenesis and courses of clinical management are established. Tyrosine kinase inhibitors (TKI) such as imatinib are effective agents for inhibiting the constitutively activated BCR-ABL1 tyrosine kinase in CML [18]. Similarly, ETV6-ABL1 is also known to constitutively activate the ABL1 tyrosine kinase, leading to cell cycle deregulation and leukemogenesis [19]. Due to the similar molecular pathogenesis of BCR-ABL1 and ETV6-ABL1 driven leukemogenesis, TKIs have also been considered in the treatment of patients bearing the ETV6-ABL1 fusion. Six out of eleven CML patients positive for ETV6-ABL1 reported in the literature were treated with imatinib: three patients showed a transient favorable response followed by relapse, one patient showed significantly decreased levels of leukemic clones, and two patients treated with 400 mg/day during the chronic phase achieved complete remission [1, 6, 18, 20–22]. Of the three that relapsed, Gancheva et al. reported a case in which the patient was administered an additional TKI, nilotinib, and the patient was able to sustain a positive response following the relapse [1]. Perna et al. did further analysis on another patient who achieved complete remission post-treatment which showed that the ETV6-ABL1 transcript became undetectable, the white blood count normalized, and expression of C-MYC, ID1, BCL-XL, and NUP-98 had decreased significantly [22]. All in all, the molecular targets of ETV6-ABL1 and BCR-ABL1 have significant overlaps that warrant further investigation to elucidate the effectiveness of TKIs on ETV6-ABL1 positive hematologic malignancies.
All in all, this is the sixth reported case of AML bearing the ETV6-ABL1 fusion gene and provides additional insight into the pathogenesis of this subset of malignancies. It is particularly important to utilize complimentary cytogenetic methodologies - namely conventional cytogenetics and FISH - in order to elucidate cryptic abnormalities, which occur more frequently in this context, and to properly characterize karyotypic changes. Additionally, screening using RT-PCR as well as other methodologies has proven useful when cytogenetic analysis is unavailable or yields negative results and in the context of broad molecular screening to identify previously unreported cases. Finally, the consideration of tyrosine kinase inhibitors, particularly second-generation ones, in the treatment of ETV6-ABL1-positive hematological malignancies has shown varying responses, and further investigation of its utility and clinical efficacy is warranted.
Abbreviations
ALL, acute lymphoblastic leukemia; AML, acute myeloid leukemia; CML, chronic myelogenous leukemia; FISH, fluorescence in situ hybridization; HLH, helix loop helix; MDS, myelodysplastic syndrome; MPD, myeloproliferative disorder; RT-PCR, reverse transcriptase-polymerase chain reaction; TKI, tyrosine kinase inhibitor
Acknowledgements
Thank you to the UCLA Clinical Cytogenetics Laboratory.
Funding
This work was supported by the UCLA Clinical and Molecular Cytogenetics Laboratory in the UCLA Department of Pathology and Laboratory Medicine.
Availability of data and materials
All data utilized to complete this manuscript is provided in the manuscript – there are no supporting materials.
Authors’ contributions
CAT, KS, and DSS contributed equally and led drafting, conducted survey of relevant literature, and edited and revised all drafts. GS and MS provided clinical presentation/findings, revised the manuscript, and provided numerous comments. All authors read and approved the final manuscript.
Competing interests
The authors declare that they have no competing interests.
Consent for publication
Written informed consent was obtained from the patient.
Ethics approval and consent to participate
The experiments with human samples included in this work were performed in accordance with the Declaration of Helsinki. No ethics committee approval was required for these experiments. Written informed consent was obtained from the patient.
Contributor Information
Carlos A. Tirado, Email: carlostirado@hotmail.com
Ken Siangchin, Email: Ksiangchin30@gmail.com.
David S. Shabsovich, Email: dshabsovich@gmail.com
Maryam Sharifian, Email: msharifian@mednet.ucla.edu.
Gary Schiller, Email: gschiller@mednet.ucla.edu.
References
- 1.Gancheva K, Virchis A, Howard-Reeves J, Cross NC, Brazma D, Grace C, Kotzampaltiris P, Partheniou F, Nacheva E. Myeloproliferative Neoplasm with ETV6-ABL1 Fusion: A Case Report and Literature Review. Mol Cytogenet. 2013;6(1):39. doi: 10.1186/1755-8166-6-39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Andreasson P, Johansson B, Carlsson M, Jarlsfelt I, Fioretos T, Mitelman F, Höglund M. BCR/ABL-negative chronic myeloid leukemia with ETV6/ABL fusion. Genes Chromosomes Cancer. 1997;20(3):299–304. doi: 10.1002/(SICI)1098-2264(199711)20:3<299::AID-GCC11>3.0.CO;2-K. [DOI] [PubMed] [Google Scholar]
- 3.Golub TR, Goga A, Barker GF, Afar DE, Mclaughlin J, Bohlander SK, Rowley JD, Witte ON, Gilliland DG. Oligomerization of the ABL Tyrosine Kinase by the Ets Protein TEL in Human Leukemia. Mol Cell Biol. 1996;16(8):4107–116. doi: 10.1128/MCB.16.8.4107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.La Starza R, Trubia M, Testoni N, Ottaviani E, Belloni E, Crescenzi B, Martelli M, Flandrin G, Pelicci PG, Mecucci C. Clonal eosinophils are a morphologic hallmark of ETV6/ABL1 positive acute myeloid leukemia. Haematologica. 2002;87(8):789–94. [PubMed] [Google Scholar]
- 5.Pelly T, Harvey Y, Brookwell R, Wordsworth H, Keng TB. A case report of an ETV6-ABL1 [t(6;9;12)] positive myeloproliferative neoplasm. Pathology. 2012;45(1):S95.
- 6.Tirado CA, Sebastian S, Moore JO, Gong JZ, Goodman BK. Molecular and Cytogenetic Characterization of a Novel Rearrangement Involving Chromosomes 9, 12, and 17 Resulting in ETV6 (TEL) and ABL Fusion. Cancer Genet Cytogenet. 2005;157(1):74–77. doi: 10.1016/j.cancergencyto.2004.06.001. [DOI] [PubMed] [Google Scholar]
- 7.Zuna J, Zaliova M, Muzikova K, Meyer C, Lizcova L, Zemanova Z, Brezinova J, Votava F, Marschalek R, Stary J, Trka J. Acute leukemias with ETV6/ABL1 (TEL/ABL) fusion: poor prognosis and prenatal origin. Genes Chromosomes Cancer. 2010;49(10):873–84. doi: 10.1002/gcc.20796. [DOI] [PubMed] [Google Scholar]
- 8.De Braekeleer E, Douet-Guilbert N, Rowe D, Bown N, Morel F, Berthou C, Férec C, De Braekeleer M. ABL1 fusion genes in hematological malignancies: a review. Eur J Haematol. 2011;86(5):361–71. doi: 10.1111/j.1600-0609.2011.01586.x. [DOI] [PubMed] [Google Scholar]
- 9.Shaffer LG, McGowan-Jordan J, Schmid M. An International System for Human Cytogenetic Nomenclature. Basel: International Standing Committee on Human Cytogenetic Nomenclature, Karger; 2013. [Google Scholar]
- 10.Ishikawa M, Yagasaki F, Okamura D, Maeda T, Sugahara Y, Jinnai I, Bessho M. A novel gene, ANKRD28 on 3p25, is fused with NUP98 on 11p15 in a cryptic 3-way translocation of t(3;5;11)(p25;q35;p15) in an adult patient with myelodysplastic syndrome/acute myelogenous leukemia. Int J Hematol. 2007;86(3):238–45. doi: 10.1007/BF03006927. [DOI] [PubMed] [Google Scholar]
- 11.Johansson B, Billström R, Kristoffersson U, Åkerman M, Garwicz S, Ahlgren T, Malm C, Mitelman F. Deletion of Chromosome Arm 3p in Hematologic Malignancies. Leukemia. 1997;11(8):1207–213. doi: 10.1038/sj.leu.2400718. [DOI] [PubMed] [Google Scholar]
- 12.Song JS, Shin SY, Lee ST, Kim HJ, Kim SH. A Cryptic ETV6/ABL1 Rearrangement Represents a Unique Fluorescence In Situ Hybridization Signal Pattern in a Patient with B Acute Lymphoblastic Leukemia. Ann Lab Med. 2014;34(6):475. doi: 10.3343/alm.2014.34.6.475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Park J, Kim M, Lim J, Kim Y, Han K, Kim JS, Lee S, Kim HJ, Min WS. Variant of ETV6/ABL1 Gene Is Associated with Leukemia Phenotype. Acta Haematologica Acta Haematol. 2013;129(2):78–82. doi: 10.1159/000342490. [DOI] [PubMed] [Google Scholar]
- 14.O'Brien SG, Vieira SA, Connors S, Bown N, Chang J, Capdeville R, Melo JV. Transient response to imatinib mesylate (STI571) in a patient with the ETV6-ABL t(9;12) translocation. Blood. 2002;99(9):3465–7. doi: 10.1182/blood.V99.9.3465. [DOI] [PubMed] [Google Scholar]
- 15.Million RP, Aster J, Gilliland DG, Van Etten RA. The Tel-Abl (ETV6-Abl) tyrosine kinase, product of complex (9;12) translocations in human leukemia, induces distinct myeloproliferative disease in mice. Blood. 2002;99(12):4568–77. doi: 10.1182/blood-2001-12-0244. [DOI] [PubMed] [Google Scholar]
- 16.Voss J, Posern G, Hannemann JR, Wiedemann LM, Turhan AG, Poirel H, Bernard OA, Adermann K, Kardinal C, Feller SM. The Leukaemic Oncoproteins Bcr-Abl and Tel-Abl (ETV6/Abl) Have Altered Substrate Preferences and Activate Similar Intracellular Signalling Pathways. Oncogene. 2000;19(13):1684–690. doi: 10.1038/sj.onc.1203467. [DOI] [PubMed] [Google Scholar]
- 17.Birge RB, Kalodimos C, Inagaki F, Tanaka S. Crk and CrkL Adaptor Proteins: Networks for Physiological and Pathological Signaling. Cell Commun Signal. 2009;7(1):13. doi: 10.1186/1478-811X-7-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kawamata N, Dashti A, Lu D, Miller B, Koeffler HP, Schreck R, Moore S, Ogawa S. Chronic Phase of ETV6-ABL1 Positive CML Responds to Imatinib. Genes Chromosomes Cancer. 2008;47(10):919–21. doi: 10.1002/gcc.20593. [DOI] [PubMed] [Google Scholar]
- 19.Greuber EK, Smith-Pearson P, Wang J, Pendergast AM. Role of ABL family kinases in cancer: from leukaemia to solid tumours. Nat Rev Cancer. 2013;13(8):559–71. doi: 10.1038/nrc3563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Barbouti A, Ahlgren T, Johansson B, Höglund M, Lassen C, Turesson I, Mitelman F, Fioretos T. Clinical and Genetic Studies of ETV6/ABL1-positive Chronic Myeloid Leukaemia in Blast Crisis Treated with Imatinib Mesylate. Br J Haematol. 2003;122(1):85–93. doi: 10.1046/j.1365-2141.2003.04391.x. [DOI] [PubMed] [Google Scholar]
- 21.Kelly JC, Shahbazi N, Scheerle J, Jahn J, Suchen S, Christacos NC, Mowrey PN, Witt MH, Hostetter A, Meloni-Ehrig AM. Insertion (12;9)(p13;q34q34): A Cryptic Rearrangement Involving ABL1/ETV6 Fusion in a Patient with Philadelphia-negative Chronic Myeloid Leukemia. Cancer Genet Cytogenet. 2009;192(1):36–39. doi: 10.1016/j.cancergencyto.2009.02.012. [DOI] [PubMed] [Google Scholar]
- 22.Perna F, Abdel-Wahab O, Levine RL, Jhanwar SC, Imada K, Nimer SD. ETV6-ABL1-positive "chronic Myeloid Leukemia": Clinical and Molecular Response to Tyrosine Kinase Inhibition. Haematologica. 2010;96(2):342–43. doi: 10.3324/haematol.2010.036673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Papadopoulos P, Ridge SA, Boucher CA, Stocking C, Wiedemann LM. The novel activation of ABL by fusion to an ets-related gene, TEL. Cancer Res. 1995;55(1):34–8. [PubMed] [Google Scholar]
- 24.Van Limbergen H, Beverloo HB, van Drunen E, Janssens A, Hählen K, Poppe B, Van Roy N, Marynen P, De Paepe A, Slater R, Speleman F. Molecular cytogenetic and clinical findings in ETV6/ABL1-positive leukemia. Genes Chromosomes Cancer. 2001;30(3):274–82. doi: 10.1002/1098-2264(2000)9999:9999<1::AID-GCC1089>3.0.CO;2-1. [DOI] [PubMed] [Google Scholar]
- 25.Baeumler J, Szuhai K, Falkenburg JH, van Schie ML, Ottmann OG, Nijmeijer BA. Establishment and cytogenetic characterization of a human acute lymphoblastic leukemia cell line (ALL-VG) with ETV6/ABL1 rearrangement. Cancer Genet Cytogenet. 2008;185(1):37–42. doi: 10.1016/j.cancergencyto.2008.05.001. [DOI] [PubMed] [Google Scholar]
- 26.Malone A, Langabeer S, O'Marcaigh A, Storey L, Bacon CL, Smith OP. A doctor(s) dilemma: ETV6-ABL1 positive acute lymphoblastic leukaemia. Br J Haematol. 2010;151(1):101–2. doi: 10.1111/j.1365-2141.2010.08323.x. [DOI] [PubMed] [Google Scholar]
- 27.Zhou MH, Gao L, Jing Y, Xu YY, Ding Y, Wang N, Wang W, Li MY, Han XP, Sun JZ, Wang LL, Yu L. Detection of ETV6 gene rearrangements in adult acute lymphoblastic leukemia. Ann Hematol. 2012;91(8):1235–43. doi: 10.1007/s00277-012-1431-4. [DOI] [PubMed] [Google Scholar]
- 28.Brunel V, Sainty D, Carbuccia N, Mazzicconacci M, Fernandez F, Simonetti J, Gabert J, Debreuil P, Lafage-Pochitaloff M, Birg F. A TEL/ABL fusion gene on chromosome 12p13 in a case of Ph-. BCR- atypical CML. Leukemia. 1996;10:2003. [Google Scholar]
- 29.Keung YK, Beaty M, Steward W, Jackle B, Pettnati M. Chronic myelocytic leukemia with eosinophilia, t(9;12)(q34;p13), and ETV6-ABL1 gene rearrangement: case report and review of the literature. Cancer Genet Cytogenet. 2002;138(2):139–42. doi: 10.1016/S0165-4608(02)00609-X. [DOI] [PubMed] [Google Scholar]
- 30.Lin H, Guo JQ, Andreeff M, Arlinghaus RB. Detection of dual TEL-ABL transcripts and a Tel-Abl protein containing phosphotyrosine in a chronic myeloid leukemia patient. Leukemia. 2002;16(2):294–7. doi: 10.1038/sj.leu.2402353. [DOI] [PubMed] [Google Scholar]
- 31.Meyer-Monard S, Mühlematter D, Streit A, Chase AJ, Gratwohl A, Cross NC, Jotterand M, Tichelli A. Broad molecular screening of an unclassifiable myeloproliferative disorder reveals an unexpected ETV6/ABL1 fusion transcript. Leukemia. 2005;19(6):1096–9. doi: 10.1038/sj.leu.2403697. [DOI] [PubMed] [Google Scholar]
- 32.Mozziconacci MJ, Sainty D, Chabannon C. A fifteen-year cytogenetic remission following interferon treatment in a patient with an indolent ETV6-ABL positive myeloproliferative syndrome. Am J Hematol. 2007;82(7):688–9. doi: 10.1002/ajh.20873. [DOI] [PubMed] [Google Scholar]
- 33.Nand R, Bryke C, Kroft SH, Divgi A, Bredeson C, Atallah E. Myeloproliferative disorder with eosinophilia and ETV6-ABL gene rearrangement: efficacy of second-generation tyrosine kinase inhibitors. Leuk Res. 2009;33(8):1144–6. doi: 10.1016/j.leukres.2009.03.011. [DOI] [PubMed] [Google Scholar]
- 34.Yamamoto K, Yakushijin K, Nakamachi Y, Miyata Y, Sanada Y, Tanaka Y, Okamura A, Kawano S, Hayashi Y, Matsuoka H, Minami H. Extramedullary T-lymphoid blast crisis of an ETV6/ABL1-positive myeloproliferative neoplasm with t(9;12)(q34;p13) and t(7;14)(p13;q11.2) Ann Hematol. 2014;93(8):1435–8. doi: 10.1007/s00277-013-1975-y. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All data utilized to complete this manuscript is provided in the manuscript – there are no supporting materials.


