Abstract
Differentiation in the dimorphic bacterium Caulobacter crescentus results from a sequence of discontinuous, stage-specific events that leads to the production of a stalked cell and a new motile swarmer cell after each asymmetric cell division. As reported previously, pseudoreversion analysis of mutations in the pleiotropic developmental gene pleC identified three cell division genes: divJ, divK, and divL. We show here that one of these genes, divJ, encodes a predicted protein of 596 residues with an extensive hydrophobic N-terminal region and a C-terminal domain containing all of the invariant residues found in the family of bacterial histidine protein kinases. Our results also show that divJ is discontinuously transcribed early in the swarmer cell cycle during a period that coincides with the G1 to S transition. We propose that the DivJ protein is one member of a signal transduction pathway regulating the cell cycle and differentiation in Caulobacter and that protein modification by phosphorylation may play a central role in coupling developmental events to progress through the cell division cycle.
Full text
PDF



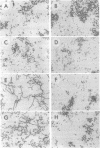
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aricò B., Scarlato V., Monack D. M., Falkow S., Rappuoli R. Structural and genetic analysis of the bvg locus in Bordetella species. Mol Microbiol. 1991 Oct;5(10):2481–2491. doi: 10.1111/j.1365-2958.1991.tb02093.x. [DOI] [PubMed] [Google Scholar]
- Aricó B., Miller J. F., Roy C., Stibitz S., Monack D., Falkow S., Gross R., Rappuoli R. Sequences required for expression of Bordetella pertussis virulence factors share homology with prokaryotic signal transduction proteins. Proc Natl Acad Sci U S A. 1989 Sep;86(17):6671–6675. doi: 10.1073/pnas.86.17.6671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bourret R. B., Borkovich K. A., Simon M. I. Signal transduction pathways involving protein phosphorylation in prokaryotes. Annu Rev Biochem. 1991;60:401–441. doi: 10.1146/annurev.bi.60.070191.002153. [DOI] [PubMed] [Google Scholar]
- Burbulys D., Trach K. A., Hoch J. A. Initiation of sporulation in B. subtilis is controlled by a multicomponent phosphorelay. Cell. 1991 Feb 8;64(3):545–552. doi: 10.1016/0092-8674(91)90238-t. [DOI] [PubMed] [Google Scholar]
- Ely B., Croft R. H., Gerardot C. J. Genetic mapping of genes required for motility in Caulobacter crescentus. Genetics. 1984 Nov;108(3):523–532. doi: 10.1093/genetics/108.3.523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huguenel E. D., Newton A. Localization of surface structures during procaryotic differentiation: role of cell division in Caulobacter crescentus. Differentiation. 1982;21(2):71–78. doi: 10.1111/j.1432-0436.1982.tb01199.x. [DOI] [PubMed] [Google Scholar]
- Iuchi S., Matsuda Z., Fujiwara T., Lin E. C. The arcB gene of Escherichia coli encodes a sensor-regulator protein for anaerobic repression of the arc modulon. Mol Microbiol. 1990 May;4(5):715–727. doi: 10.1111/j.1365-2958.1990.tb00642.x. [DOI] [PubMed] [Google Scholar]
- Kyte J., Doolittle R. F. A simple method for displaying the hydropathic character of a protein. J Mol Biol. 1982 May 5;157(1):105–132. doi: 10.1016/0022-2836(82)90515-0. [DOI] [PubMed] [Google Scholar]
- Lipman D. J., Pearson W. R. Rapid and sensitive protein similarity searches. Science. 1985 Mar 22;227(4693):1435–1441. doi: 10.1126/science.2983426. [DOI] [PubMed] [Google Scholar]
- Minnich S. A., Newton A. Promoter mapping and cell cycle regulation of flagellin gene transcription in Caulobacter crescentus. Proc Natl Acad Sci U S A. 1987 Mar;84(5):1142–1146. doi: 10.1073/pnas.84.5.1142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mullin D. A., Newton A. Ntr-like promoters and upstream regulatory sequence ftr are required for transcription of a developmentally regulated Caulobacter crescentus flagellar gene. J Bacteriol. 1989 Jun;171(6):3218–3227. doi: 10.1128/jb.171.6.3218-3227.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Newton A., Ohta N. Regulation of the cell division cycle and differentiation in bacteria. Annu Rev Microbiol. 1990;44:689–719. doi: 10.1146/annurev.mi.44.100190.003353. [DOI] [PubMed] [Google Scholar]
- O'Neill E. A., Hynes R. H., Bender R. A. Recombination deficient mutant of Caulobacter crescentus. Mol Gen Genet. 1985;198(2):275–278. doi: 10.1007/BF00383006. [DOI] [PubMed] [Google Scholar]
- Ohta N., Chen L. S., Mullin D. A., Newton A. Timing of flagellar gene expression in the Caulobacter cell cycle is determined by a transcriptional cascade of positive regulatory genes. J Bacteriol. 1991 Feb;173(4):1514–1522. doi: 10.1128/jb.173.4.1514-1522.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohta N., Masurekar M., Newton A. Cloning and cell cycle-dependent expression of DNA replication gene dnaC from Caulobacter crescentus. J Bacteriol. 1990 Dec;172(12):7027–7034. doi: 10.1128/jb.172.12.7027-7034.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohta N., Swanson E., Ely B., Newton A. Physical mapping and complementation analysis of transposon Tn5 mutations in Caulobacter crescentus: organization of transcriptional units in the hook gene cluster. J Bacteriol. 1984 Jun;158(3):897–904. doi: 10.1128/jb.158.3.897-904.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perego M., Cole S. P., Burbulys D., Trach K., Hoch J. A. Characterization of the gene for a protein kinase which phosphorylates the sporulation-regulatory proteins Spo0A and Spo0F of Bacillus subtilis. J Bacteriol. 1989 Nov;171(11):6187–6196. doi: 10.1128/jb.171.11.6187-6196.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seki T., Yoshikawa H., Takahashi H., Saito H. Nucleotide sequence of the Bacillus subtilis phoR gene. J Bacteriol. 1988 Dec;170(12):5935–5938. doi: 10.1128/jb.170.12.5935-5938.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sheffery M., Newton A. Regulation of periodic protein synthesis in the cell cycle: control of initiation and termination of flagellar gene expression. Cell. 1981 Apr;24(1):49–57. doi: 10.1016/0092-8674(81)90500-6. [DOI] [PubMed] [Google Scholar]
- Sommer J. M., Newton A. Pseudoreversion analysis indicates a direct role of cell division genes in polar morphogenesis and differentiation in Caulobacter crescentus. Genetics. 1991 Nov;129(3):623–630. doi: 10.1093/genetics/129.3.623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sommer J. M., Newton A. Sequential regulation of developmental events during polar morphogenesis in Caulobacter crescentus: assembly of pili on swarmer cells requires cell separation. J Bacteriol. 1988 Jan;170(1):409–415. doi: 10.1128/jb.170.1.409-415.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sommer J. M., Newton A. Turning off flagellum rotation requires the pleiotropic gene pleD: pleA, pleC, and pleD define two morphogenic pathways in Caulobacter crescentus. J Bacteriol. 1989 Jan;171(1):392–401. doi: 10.1128/jb.171.1.392-401.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stock J. B., Ninfa A. J., Stock A. M. Protein phosphorylation and regulation of adaptive responses in bacteria. Microbiol Rev. 1989 Dec;53(4):450–490. doi: 10.1128/mr.53.4.450-490.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wanner B. L. Is cross regulation by phosphorylation of two-component response regulator proteins important in bacteria? J Bacteriol. 1992 Apr;174(7):2053–2058. doi: 10.1128/jb.174.7.2053-2058.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]