Abstract
Ligand-dependent activation, γ-secretase-processed cleavage, and RBPj-mediated downstream transcriptional activities of Notch receptors constitute the “canonical” Notch signaling pathway, which is essential for skin organogenesis. However, in Msx2-Cre mice, keratinocyte-specific deletion of the Rbpj gene in utero produced a significantly milder phenotype than either global Notch or γ-secretase loss. Herein, we investigated the underlying mechanisms for this apparent non-canonical signal using mouse genetics. We found no evidence that ligand back signaling contributed to skin organogenesis. The perdurance of RBPj protein did not establish an epigenetic memory of a canonical signal in the youngest epidermal stem cells, and Notch targets were not de-repressed. We provide evidence that γ-secretase-dependent but RBPj-independent NICD activity operating in the first hair follicles is responsible for a delay in follicular destruction, which results in lower serum TSLP levels, milder B-cell Lymphoproliferative Disease, and improved survival in Msx2-Cre+/tg;Rbpjf/f mice. Minimal amounts of NICD were sufficient for rescue, which was not mediated by transcription, suggesting that NICD is acting through a novel mechanism.
Keywords: Notch, hair, skin, non-canonical signaling, RBPj-independent
Introduction
The skin provides a protective barrier, temperature regulation, and communication of gender and age-specific signals. Mouse skin develops from a single layer of ectodermal cells that stratify at embryonic day 13.5 (E13.5) to form suprabasal layers (Byrne et al., 1994; Fuchs and Green, 1980) and undergo induction of appendages such as hair follicles (HF; (Aubin-Houzelstein, 2012; Jamora et al., 2003; Schmidt-Ullrich and Paus, 2005)).
The Notch signaling pathway is a conserved, short-range communication pathway cells use to regulate diverse cellular functions (Artavanis-Tsakonas et al., 1999; Bray, 2006; Kopan and Ilagan, 2009). Loss- and gain-of-function studies suggest that Notch signals are essential for terminal epidermal differentiation and promote exit from the basal layer (Blanpain et al., 2006). Although Notch signaling is not required for initial HF lineage specification, it maintains the follicular architecture (Lin et al., 2011; Lin et al., 2000; Pan et al., 2004; Vauclair et al., 2005; Watt et al., 2008). Notch receptors are activated by a family of ligands (Delta-like (Dll)1, Dll3, Dll4, Jagged1, and Jagged2) expressed on the surface of neighboring cells. Ligand binding unfolds a protective juxtamembrane domain permitting ADAM10 cleavage, ectodomain shedding, and intra-membrane cleavage by the enzyme γ-secretase. This enzyme liberates the Notch Intracellular Domain (NICD), which translocates to the nucleus where it interacts with a DNA-binding partner, RBPj. NICD binding to RBPj allows recruitment of Mastermind-like protein, which recruits p300 and the machinery necessary to activate transcription (Bray and Bernard, 2010; Bray, 2006; Gordon et al., 2008; Kopan and Ilagan, 2009; Kovall and Blacklow, 2010). The signaling axis involving γ-secretase cleavage, nuclear association with RBPj, and transcriptional activation of targets is known as the “canonical’ pathway. Other modes of signaling, collectively known as “non-canonical” signals, (Sanalkumar et al., 2010) have been reported but a biochemical foundation for such alternative pathways is still lacking. At present, non-canonical activity in mammalian tissues were noted when NICD was overexpressed (Chen et al., 2013).
Three of the four Notch paralogs, Notch1–3, co-regulate mouse skin differentiation. Using the Msx2-Cre transgene to create keratinocyte-specific calico loss-of-function mosaics (henceforth, labeled as msx2LOF), we reported that Notch1 performs the majority of this function (Demehri et al., 2008; Demehri et al., 2009b; Liu et al., 2015). Loss of both Notch1 and Notch2 (Notchmsx2LOF) or both Preselin1 and Preselin2 (PSmsx2LOF) genes, resulted in elevated Thymic stromal lymphopoietin (TSLP) expression and secretion, driving a severe B-cell lymphoproliferative disease (BLPD) and inflammation that limit survival to the first month of life (Demehri et al., 2008; Pan et al., 2004). Surprisingly, deletion of the floxed (f) Rbpj alleles with the same Cre line (RBPjmsx2LOF) produced a true null (Sato et al., 2012), yet, displayed significantly milder quantitative phenotypes (Demehri et al., 2008; Demehri et al., 2009b; Demehri et al., 2012). Confounding the interpretation of RBPjmsx2LOF mice, the observation that postnatal deletion of Rbpj gene with K5-CreErt2 (RBPjK5LOF) was phenotypically indistinguishable from N1N2K5LOF (Dumortier et al., 2010).
RBPj proteins can nucleate repressor complexes, which NICD can reverse (Goodfellow et al., 2007; Koelzer and Klein, 2006; Olave et al., 1998; Skalska et al., 2015). Thus, one possible explanation for the differences between RBPjmsx2LOF and the other Notch mutants could have been target derepression at an early critical step (Figure 1a). This possibility was formally ruled out by generating triple mutant mice, demonstrating that RBPj loss in Msx2-Cre+/tg; Psen1f/f;Psen2−/−;Rbpjf/f (PS;RBPjmsx2LOF) or Msx2-Cre+/tg;Notch1f/f;Notch2f/f;Rbpjf/f (Notch;RBPjmsx2LOF) mice did not ameliorate the severe phenotype associated with loss of canonical Notch signaling (Demehri et al., 2008). These observations promoted us to consider the possibility that an RBPj-independent (i.e. non-canonical) Notch function contributed to mouse skin development.
Figure 1. Schema for Notch Signaling and reagents used in this study.
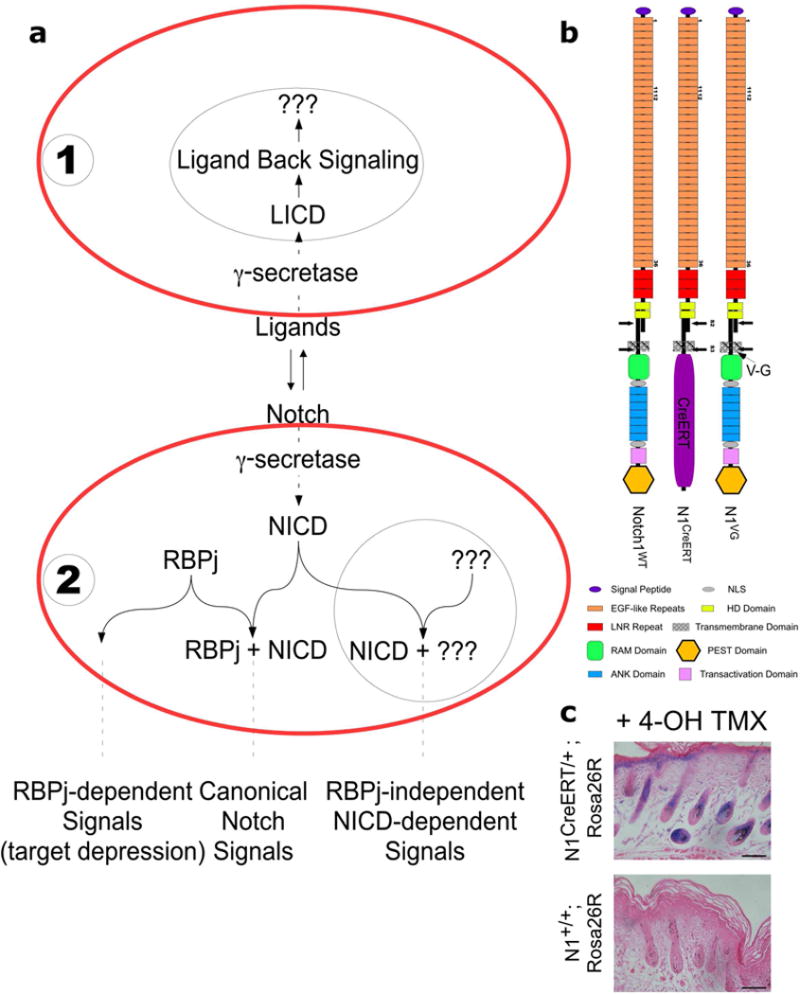
(a). In the canonical Notch signaling pathway, ligand from cell #1 induces cleavage and RBPj dependent signal in cell #2. However, Ligand Back Signaling can occur concomitantly in cell #1 (black circle). RBPj-dependent activity, and γ-secretase dependent, RBPj-independent NICD signals (black circle) are also potentially functioning in cell #2. (b). Diagram of Notch1wt, N1CreERT and N1VG receptors. (c). N1CreERT receptor is activated by ligands, and thus can trigger LBS. X-gal staining of 4-OHT tamoxifen-injected N1CreERT Rosa26R/Rosa26R skin. Bars, 100 μm.
Herein, we used genetic analyses to provide an explanation for the differences between the phenotypes reported in msx2LOF mice. We ruled out epigenetic preservation of a canonical signal in the youngest epidermal stem cells. We also ruled out ligand back signaling as a contributor to skin development (Figure 1a (LaVoie and Selkoe, 2003)). Genetic dissection of the phenotype provided evidence that γ-secretase dependent but RBPj-independent NICD activity is responsible for the milder RBPjmsx2LOF phenotype in Notch;RBPjmsx2LOF background. Moreover, we show that this RBPj-independent function of Notch signaling is apparent only in the first cycle HF keratinocytes, explaining why only RBPjmsx2LOF, but not RBPjK5LOF mice, display the milder phenotype.
RESULTS
RBPj perdurance does not explain the milder phenotype
The differences between RBPjmsx2LOF and RBPjK5LOF may reflect perdurance of RBPj during a critical window in skin development creating an “epigenetic memory” of a canonical signal in Rbpjdel epidermal stem cells (Tadeu and Horsley, 2013). Alternatively, an RBPj-independent function of Notch is operating during fetal and early postnatal skin development. We monitored the decay rate of RBPj protein in Rbpjf/f primary keratinocytes grown in vitro following adeno-Cre-mediated deletion of Rbpj gene (FigureS1b,c,d). The RBPj protein could be detected up to 90 hours post adeno-Cre treatment (FigureS1d). Since Msx2-Cre is expressed at embryonic day 9.5 (E9.5) (Sun et al., 2000), canonical Notch signals could be present at the beginning of epidermal stratification (E13.5) in the RBPjmsx2LOF embryos (FigureS1a) if RBPj half-life in vitro and in vivo were similar. If an early signal was preserved by epigenetic memory, E18.5 RBPjmsx2LOF epidermis would resemble the wild-type at the transcriptional level. To test this hypothesis, we profiled the epidermal transcriptome at E18.5. Unsupervised hierarchical clustering analyses could not distinguish between the epidermal RBPjmsx2LOF and PSmsx2LOF transcriptomes, which were significantly different from the wild-type epidermis (FigureS1e). Likewise, pathway analysis could not identify any pathway signature that could differentiate between RBPjmsx2LOF and PSmsx2LOF (TableS1). These results are inconsistent with the hypothesis that perdurance of a canonical Notch signal created an epigenetic memory during a critical window prior to stratification.
To ask if the phenotypic differences between RBPjmsx2LOF and PSmsx2LOF mice could reflect differences in post-transcriptional or post-translational modifications, we performed a liquid chromatography-mass spectrometry (LC-MS/MS) analysis to identify differences in the epidermal protein quotient between newborn RBPjmsx2LOF and PSmsx2LOF mice. Epidermal samples from three newborn RBPjmsx2LOF, PSmsx2LOF, and wild-type littermates were collected. Peptides prepared by Lys-C digestion were analyzed by LC-MS/MS and raw data analyses were conducted as described in the supplemental methods. Annotated peptides were compared between the samples and to the pooled wild-type epidermal peptides as control. Unsupervised hierarchical clustering analysis clustered all replicates from the same genotype together, confirming the precision of the surgical procedure removing the mutant skin and avoiding contamination by wild-type cells (FigureS2a). Again, RBPjmsx2LOF epidermal peptides clustered with the PSmsx2LOF epidermal peptides and were clearly distinguishable from the wild-type proteome (FigureS2a). Abundant peptides shared between RBPjmsx2LOF and wild-type epidermis but absent in PSmsx2LOF did not enrich for any particular protein or post-translational modification including phosphorylation, sumoylation, palmitoylation, or ubiquitination (TableS2). In summary, statistical analyses of the transcriptome and the proteome data confirmed that the RBPjmsx2LOF epidermis was indistinguishable from the PSmsx2LOF epidermis at birth.
Finally, appendicular keratinocytes also experienced deletion of floxed alleles. To explore the possibility that the differences between the RBPjmsx2LOF and PSmsx2LOF phenotypes reflected differences in appendicular keratinocytes, we performed microarray analysis on RNA samples isolated from total skin of RBPjmsx2LOF, PSmsx2LOF, PS;RBPjmsx2LOF and wild-type littermates at P0. Previous analyses had shown that lifespan, WBC counts, and serum TSLP levels of PS;RBPjmsx2LOF and PSmsx2LOF (Msx2-Cre+/tg;Psen1f/f;Psen2−/−;Rbpj+/f) differ significantly from RBPjmsx2LOF (Msx2-Cre+/tg;Psen1+/f;Psen2−/−;Rbpjf/f) born to the same parents (Demehri et al., 2008) eliminating genetic background as a deriver of the phenotypic differences. Unsupervised hierarchical clustering analyses could not distinguish between the transcriptomes of mice with mutant genotypes, which were all clearly distinguishable from unaffected compound heterozygous littermates (FigureS2b and TableS3).
Ligand Back Signaling does not explain the differences in phenotype
In the absence of RBPj protein, Notch ligands are still able to interact with Notch receptors (Figure 1a). It has been previously shown that like Notch receptors, DSL ligands also undergo proteolytic cleavages by ADAM metalloproteases and γ-secretase following receptor-ligand interaction releasing the Ligand Intracellular Domain (LICD) (Bland et al., 2003; Bordonaro et al., 2011; Hiratochi et al., 2007; Ikeuchi and Sisodia, 2003; LaVoie and Selkoe, 2003; Qi et al., 1999; Six et al., 2003; Zolkiewska, 2008). If these endoproteolytic fragments can survive the N-end rule degradation machinery, they would be generated in RBPjmsx2LOF, but not in PSmsx2LOF. This provides an alternative explanation for the milder RBPjmsx2LOF phenotype if they acted without altering the transcriptome by P0 or altered the transcriptome postnatally. Because either Msx2-Cre+/tg;Notch1+/f;Notch2f/f;Rbpjf/f or Msx2-Cre+/tg;Notch1f/f;Notch2+/f;Rbpjf/f display the RBPjmsx2LOF phenotype (data not shown), a single extracellular domain (ECD) from either receptor should be sufficient to mediate LBS. Note that absence of evidence for accumulation of LICD in skin cells cannot be used to argue it has no role there.
Since Notch1 plays a major role in the skin we decided to restore the N1ECD without introduction of the N1ICD by using NIP1::CreErt2 (Pellegrinet et al., 2011). This allele of Notch1 contains the entire extracellular and transmembrane domains, but the intracellular domain has been replaced with a tamoxifen-inducible Cre-recombinase (CreErt2) (Figure 1b). Ligand binding triggers proteolytic cleavages of the Notch1-CreERT protein (N1CreERT), releasing the inactive CreERT molecule instead of N1ICD. As expected, ligand-receptor engagement can promote recombination of floxed reporter alleles in the presence of tamoxifen (Figure 1c and (Pellegrinet et al., 2011)), and N1CreERT receptors induce LICD accumulation in the ligand-presenting cells to the same degree as the wild type receptor (FigureS1f,g). To ask if the LBS contributed to the RBPjmsx2LOF phenotype, we generated mice with the genotype Msx2-Cre+/tg;Notch1f/CreErt2;Notch2f/f (NotchCre/msx2LOF). Histological sections from NotchCre/msx2LOF and RBPjmsx2LOF mice (Figure 2a) revealed that NotchCre/msx2LOF skin had higher number of cystic HFs with a disrupted morphology and fewer adipocytes relative to the RBPjmsx2LOF skin (Pan et al., 2004). Lifespan, WBC counts, and serum TSLP levels of NotchCre/msx2LOF mice were indistinguishable from those of Notchmsx2LOF mice (Figure 2b). In conclusion, the data is inconsistent with a role for ligand back signaling in mammalian skin development. Postnatal events that ameliorate the Notch phenotype in RBPjmsx2LOF must rely on a different mechanism.
Figure 2. The RBPjmsx2LOF phenotype does not reflect Ligand Back Signaling.
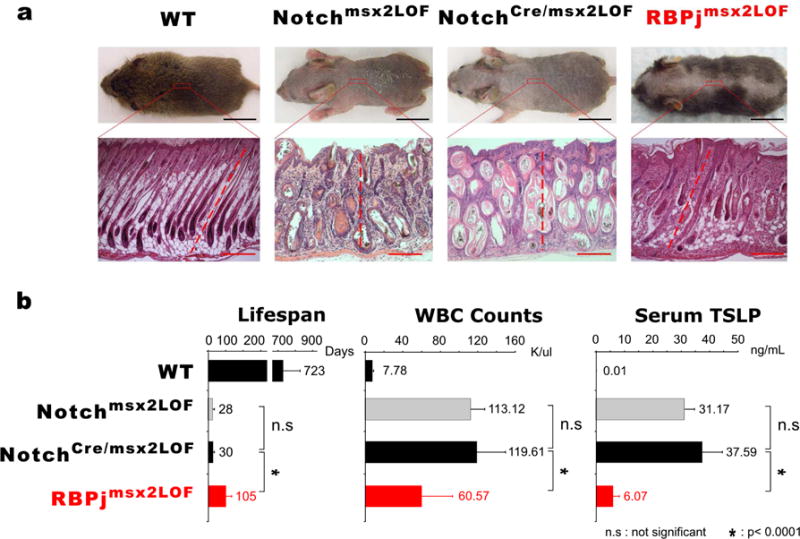
(a). Samples were collected at P9. Hair phenotypes and histological sections indicate that NotchCre/msx2LOF mice resembled Notchmsx2LOF mice, but not RBPjmsx2LOF mice. Black Bars, 1cm; Red Bars, 100 μm. (b). Lifespan, WBC counts and Serum TSLP level analyses. WBC and TSLP measured within first two weeks animals’ life.
γ-secretase dependent N1ICD drives the milder phenotype in NotchVG;RBPjmsx2LOF mice
NICD is generated in RBPjmsx2LOF keratinocytes but not in Notchmsx2LOF or PSmsx2LOF cells. To determine whether NICD contributed to the mild RBPjmsx2LOF phenotype, we incorporated a hypomorphic Notch1 allele, N1VG (Huppert et al., 2005; Huppert et al., 2000), which carries the V1744G substitution in the transmembrane domain (Figure 1b). The V1744G substitution substantially reduced the half-life of the N1ICD (Blat et al., 2002; Huppert et al., 2000; Tagami et al., 2008). Mice homozygous for this hypomorphic allele (N1VG/VG) cannot survive (Huppert et al., 2000) but across other alleles N1VG provides some activity (Liu et al., 2015). To test whether the residual N1ICD production in Notch;RBPjmsx2LOF can reproduce the RBPjmsx2LOF phenotype, we replaced one null allele of Notch1 with N1VG (Msx2-Cre+/tg; Notch1VG/f;Notch2f/f;Rbpjf/f; or NotchVG;RBPjmsx2LOF). In this genetic background, N1VGICD will not be able to restore canonical Notch signals (since RBPj is missing), and its inherent instability would alleviate the concern that it will accumulate to non-physiological levels in the absence of RBPj.
Notch;RBPjmsx2LOF mice are phenotypically indistinguishable from Notchmsx2LOF (Demehri et al., 2008). We compared the hair phenotypes, the lifespan, WBC counts, and serum TSLP levels of Notchmsx2LOF, Notch;RBPjmsx2LOF, RBPjmsx2LOF and NotchVG;RBPjmsx2LOF mice. Although no canonical signals can be generated, NotchVG;RBPjmsx2LOF mice resembled RBPjmsx2LOF mice, not Notchmsx2LOF mice (Figure 3). Histological analysis confirmed that NotchVG;RBPjmsx2LOF skin displayed fewer cystic HFs, increased adipocyte numbers, and a milder dismorphology when compared to Notchmsx2LOF or Notch;RBPjmsx2LOF skin (Figure 3a). Analysis of lifespan, WBC counts, and serum TSLP levels reveal that NotchVG;RBPjmsx2LOF animals were similar to the RBPjmsx2LOF animals (Figure 3b), indicating that the RBPjmsx2LOF phenotype is driven by N1ICD (or N2ICD in Msx2-Cre+/tg;Notch1f/f;Notch2+/f;Rbpjf/f mice, data not shown). Collectively, these results indicate that the presence of NICD molecules, even in limiting amounts, somehow ameliorates the Notchmsx2LOF phenotype in a non-canonical (RBPj-independent) manner after birth.
Figure 3. γ-secretase dependent NICD reproduced the RBPjmsx2LOF phenotype in NotchVG;RBPjmsx2LOF mice.
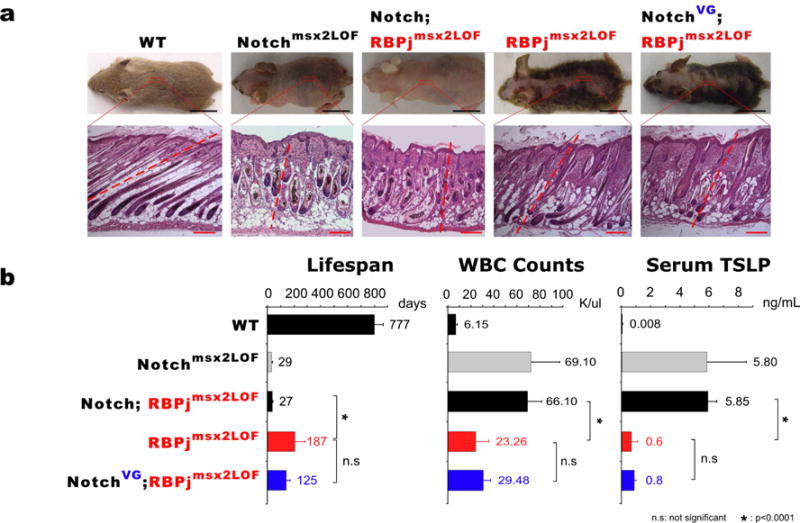
(a). Samples were collected at P9. Hair phenotypes and histological sections suggest that NICD can rescue the phenotype in Notch;RBPjmsx2LOF mice in a non-canonical manner after birth. Black Bars, 1cm; Red Bars, 100 μm. (b). Lifespan, WBC counts and Serum TSLP level analyses. WBC and TSLP measured within first two weeks animals’ life.
Non-canonical NICD activity in HFs result in reduced TSLP production and amelioration of the phenotypes
High serum TSLP levels during the first two weeks of life induce uncontrolled expansion of immature B cells, which drive mortality in Notchmsx2LOF and PSmsx2LOF mice (Demehri et al., 2008; Demehri et al., 2012). These and additional unpublished observations (RK, SD) indicate TSLP levels are the key driver of the phenotypes monitored herein. TSLP mRNA levels on total RNA isolated from E18.5, P0, P6, P9, and P11 wild-type, Notchmsx2LOF, PSmsx2LOF, and RBPjmsx2LOF epidermis were compared by qRT-PCR. RBPjmsx2LOF epidermis contained the highest levels of TSLP mRNA at E18.5 (Figure 4a); however, epidermal expression of TSLP mRNA was similar in all Notch pathway mutants after birth (Figure 4a).
Figure 4. Lower TSLP mRNA expression in RBPjmsx2LOF skin compared to Notchmsx2LOF and PSmsx2LOF skin.
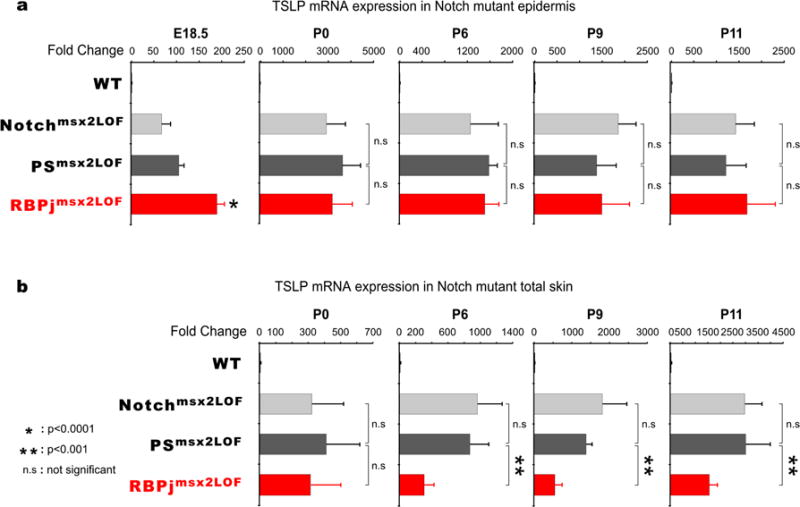
(a). Real-time qPCR analyses of TSLP mRNA expression in wild-type, Notchmsx2LOF, PSmsx2LOF and RBPjmsx2LOF epidermis from E18.5, P0, P6, P9, and P11 mice. Error bars indicate standard deviation. (b). RBPjmsx2LOF total skin has significantly reduced TSLP mRNA expression compared to Notchmsx2LOF and PSmsx2LOF total skin. Real-time qPCR analyses of TSLP mRNA in wild-type, Notchmsx2LOF, PSmsx2LOF and RBPjmsx2LOF dorsal skin from P0, P6, P9, and P11 mice. Error bars indicate standard deviation.
Next, we compared TSLP mRNA levels in dorsal skin including appendicular keratinocytes at P0, P6, P9 and P11. At birth, all mice contained similar levels of TSLP mRNA in total skin samples (Figure 4b). However, starting at P6 TSLP mRNA levels in dorsal skin of Notchmsx2LOF and PSmsx2LOF mice reached a significantly higher level than RBPjmsx2LOF (~950, ~850 vs. ~300 fold over wild type, respectively; Figure 4b). Since epidermal TSLP mRNA levels at that age were similar among all mutants, these findings suggest that RBPjmsx2LOF appendicular keratinocytes produced less TSLP than those with other Notch pathway mutants. The timing of appendicular TSLP expression corresponded to the onset of follicular destruction (Pan et al., 2004).
To determine whether the differences in TSLP expression mirrored the number of disrupted HFs, we histologically analyzed skin samples from E18.5 to P11. HFs display morphological abnormalities at birth in all Notch pathway mutants (Blanpain et al., 2006; Pan et al., 2004) and conversion to epidermal cysts begins around P8 (Pan et al., 2004). To quantify the degree of HF to cyst formation in different mutants, we adapted a rank order-based scoring system, defining normal hair as 1, and a keratin cyst lacking dermal papilla as 10 (Figure 5a, legend). Three blinded observers independently assigned a rank score to each HF within 10 different 3×0.5mm areas of dorsal skin from P9 mice. We calculated the average rank score and standard deviation for each genotype. Whereas the majority of HFs in Notchmsx2LOF and PSmsx2LOF mice converted to epidermal cysts, most of the HFs in RBPjmsx2LOF skin maintained a recognizable hair shaft (Figure 5a). Accordingly, the number of TSLP producing cells within the HF of mutant mice was significant higher in Notchmsx2LOF and PSmsx2LOF relative to RBPjmsx2LOF HF (Figure 5b).
Figure 5. RBPjmsx2LOF P9 hair follicles have better morphology than Notchmsx2LOF or PSmsx2LOF follicles.
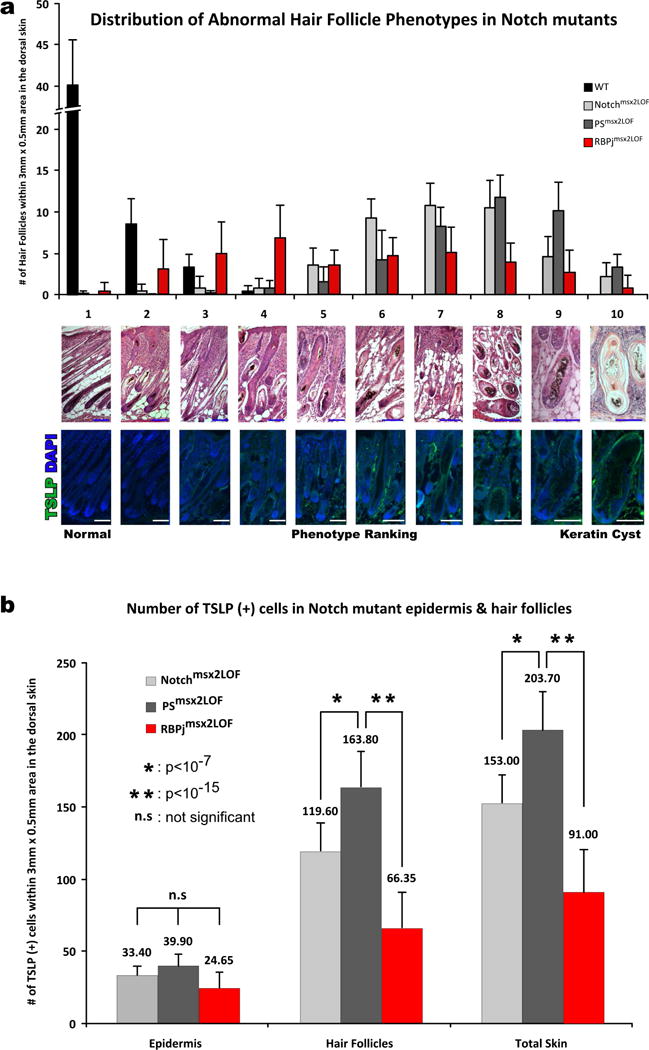
Three blinded observers within 10 random 3×0.5mm2 dorsal skin areas (a) ranked H&E stained hair follicles based on morphology, and (b) counted TSLP (+) cells after TSLP IHC. The average distribution of scores is shown. Follicles with a score <4 were contained within a single plane whereas hair follicles with score ≥5 have tortured orientation. Hair follicles with scores 5–7 did not form a cyst but lost the typical hair follicle shape. Follicles scoring 8 or higher converted to cysts. Error bars indicate standard deviation. TSLP immunofluorescence (Green) is shown in lower panels. Blue and white bars, 100 μm.
No transcriptional signature in distinguished RBPjmsx2LOF appendages
These analyses support the hypothesis that the TSLP level mirrors the degree of HF disruption and drive the severity of phenotypes; somehow, the presence of NICD prevents HF disruption in the first cycle. Since mice from all genetic backgrounds experience a similar disruption in the skin differentiation program at birth, we focused the analysis on post-partum skin. We asked if NICD presence protects the RBPjmsx2LOF HF by altering gene expression via association with other DNA binding proteins at P3, just before the conversion to TSLP-producing keratin cysts. We isolated HF bulbs from freshly frozen P3 skin sections using Laser Capture Microdissection (LCM; details in Sup. Methods, FigureS3a). ~100 hair follicles from Notchmsx2LOF, PSmsx2LOF, RBPjmsx2LOF and wild-type samples were pooled into 3 biological replicates for each genotype and subjected to RNA isolation followed by RNA-Seq. A total of 2047 genes were differentially expressed (≥1.5 fold) in three or more biological replicates of Notch mutant hair follicles compared to wild-type controls (p-value<0.05; FigureS3b). Unsupervised hierarchical clustering analysis failed to distinguish between the mutants (FigureS3d). Although RBPjmsx2LOF HF transcripts were not differentiated from other pathway mutants, gene Ontology (GO) and Pathway Enrichment analysis revealed that members of the Retinoic Acid (RA) signaling pathway, RA binding proteins 1, 4 and 7 (RBP1, RBP4, and RBP7) and CRABP1 were differentially expressed (TableS4). RBP4 is produced in the liver and secreted into the circulation where it contributes to internalization of Retinol into cells; therefore its mRNA levels in the skin are unlikely to play a meaningful role. Western Blot analysis determined that RBP1 and RBP7 were significantly reduced in Notchmsx2LOF and PSmsx2LOF skin compared to RBPjmsx2LOF skin (FigureS3c); however, we could not establish if the reduction in RBP1 and RBP7 proteins significantly contribute to the increased fraction of follicular cysts.
Discussion
Outcomes dependent on canonical Notch signals should respond the same way to perturbations such as removing the receptors (no NICD), γ-secretase (no NICD release), or RBPj (no binding of NICD to target enhancers). When keratinocyte-specific Msx2-Cre mediated deletion of Rbpj in-utero produced a significantly milder phenotype than loss of Notch or γ-secretase, we sought mechanistic insight. Because postnatal, tamoxifen-induced deletion of Rbpj with K5-Cre produced identical phenotypes to loss of Notch1 and 2, and because the double mutants lacking RBPj and Notch or RBPj and γ-secretase in the Msx2-Cre strain did not display the milder RBPj phenotype, we ruled out a mechanism based on de-repression of key targets (Koelzer and Klein, 2006; Skalska et al., 2015). Transcriptome and proteome analysis supported this conclusion. Likewise, phenotypic analysis coupled with transcriptome and proteome analysis dispensed the hypotheses of an “epigenetic memory” or ligand back signaling as the mechanism explaining the milder RBPj phenotype. However, we demonstrated that restoring sub-physiological levels of N1ICD in NotchVG;RBPjmsx2LOF mice was sufficient to ameliorate the phenotype seen in Notch;RBPjmsx2LOF. Combined, these experiments confirmed that NICD and γ-secretase were required for, a yet to be defined, RBPj-independent function of Notch (uncleaved Notch is present in the PSmsx2LOF mice but it does not function), but do not identify the cell type in which the activity is needed or the nature of this activity.
Hair follicles form in a process that starts in utero, enter a growth phase (anagen) before birth, and are dismantled (catagen) during a short window at P15-P17. After catagen ends the HF enter a short resting period (telogen) and re-enter Anagen synchronously around P28 (Muller-Rover et al., 2001). A key difference between RBPjmsx2LOF and RBPjK5LOF is that HFs develop in the absence of Notch signaling in the former, whereas the first anagen proceeds almost entirely with active Notch signaling in the latter. This difference proved to be the key to explaining the differences between these two lines. Several experiments suggest it is here that NICD controls the outcome.
First, histological analysis of HFs, TSLP protein and mRNA quantification point out that the differences between RBPjmsx2LOF and other Notch mutants appear in disrupted HF. The majority of NICD-containing RBPjmsx2LOF HFs are mildly perturbed, express significantly less TSLP mRNA, and have fewer TSLP producing cells. This, in turn, leads to lower serum TSLP levels in newborn RBPjmsx2LOF mice, which experience a milder BLPD. This is key to survival as evident from the observation that controlling BLPD with irradiation prolonged the lifespan of Notchmsx2LOF and PSmsx2LOF animals (Demehri et al., 2008; Demehri et al., 2012). Because TSLP cannot induce BLPD after P7 (Demehri et al., 2008), affected K5-Cre pups do not develop B-LPD and all survive much longer relative to Msx2-Cre mice. Our analysis predicts that careful examination of the HF in the second hair cycle will reveal some delay in RBPjK5LOF follicular destruction, but eventually all the second cycle follicles convert to cysts and thus TSLP levels become similar among all mutants.
In conclusion, we characterized a perplexing phenotype observed only when Rbpj gene was deleted in-utero, and defined an NICD-dependent, RBP-independent function in the anagen follicle. While we are unable to provide a compelling biochemical explanation of how NICD delivers this non-canonical function, we offer two speculations. The inner root sheet (IRS) is the most affected cell type in Notch1 mutants (Pan et al., 2004), and its absence promoted accelerated catagen (Lee et al., 2007; Vauclair et al., 2005). Thus, we speculate that NICD could have a cytoplasmic role that promotes IRS function. Second, RNA-Seq from affected HF suggested that retinoid binding proteins (RBP1 and RBP7) were somehow dependent on NICD, but not on RBPj; however, we could not link RA levels to the RBPjmsx2LOF phenotype.
Material and Methods
Generation of mice
Mice were maintained on mixed genetic background and housed at the Washington University in St. Louis (WUSTL) or Cincinnati Children’s Hospital Medical Center (CCHMC) animal facilities. Experiments were performed in accordance with relevant institutional and national guidelines and regulations approved by the respective Animal Studies Committees. Animals not generated as outlined previously (Demehri et al., 2008; Demehri et al., 2009a; Demehri et al., 2009b; Demehri et al., 2012; Demehri et al., 2014; Pan et al., 2004) are: NotchCre/msx2LOF (Msx2-Cre+/tg;Notch1f/CreErt2;Notch2f/f) generated by crossing Notch1+/CreErt2 females with Msx2-Cre+/tg;Notch1+/f;Notch2f/f males to obtain Msx2-Cre+/tg; Notch1f/CreErt2;Notch+/f males, which were then crossed with Notch1f/f;Notch2f/f females. NotchVG;RBPjmsx2LOF (Msx2-Cre+/tg;Notch1f/VG;Notch2f/f;Rbpjf/f) mice were generated by crossing Notch1+/VG females with Msx2-Cre+/tg;Notch1+/f;Notch2+/f;Rbpj+/f males to obtain Msx2-Cre+/tg;Notch1VG/f;Notch2+/f;Rbpj+/f males, which then crossed with Notch1f/f;Notch2f/f;Rbpjf/f females.
Husbandry of Notch mutants
To improve survival, non-control littermates were culled. Severely affected individuals were left with their mothers for their entire life span and were observed daily for loss of appetite, weight loss, weakness, and other signs of morbidity. Moribund animals were euthanized.
Measurement of WBC counts
Blood samples were collected through the mandibular vein. White Blood Cell (WBC) counts were performed with Hemavet 950 analyzer (Drew Scientific). If necessary, blood smears were counted by the pathologist.
Serum TSLP measurements
Total serum TSLP levels were determined by using Quantikine mouse TSLP ELISA kits (R&D Systems) and/or Mouse TSLP ELISA MAX™ Deluxe (BioLegend) kits according to the manufacturer’s instructions.
Histology, H&E Staining, Immunohistochemistry and X-Gal Staining
Skin samples were fixed in 4% paraformaldehyde (PFA) in PBS overnight (o/n), washed 3×15 min with PBS followed by 15 min washes with 30% Ethanol (EtOH), 50% EtOH, and 70% EtOH. Samples were stored in 70% ETOH at 4°C or dehydrated in 100% ETOH, equilibrated in Xylene, and embedded in paraffin blocks for sectioning at 8 μm thickness. A standard H&E staining protocol was used for histology assessment. Immunostaining on paraffin-embedded tissue samples were performed with anti-TSLP (R&D Systems) biotinylated antibody. Sections stained with X-Gal for β-galactosidase activity were processed according to as described previously (Kopan et al., 2002).
RNA isolation
Epidermis was isolated by incubating skin in 5mg/mL dispase for 3–4 hours at 4°C. Total RNA was isolated from epidermal samples using QIAzol (Qiagen) and RNeasy kit (Qiagen) according to the manufacturer’s instructions. After RNase-free DNase I (Invitrogen, USA) treatment RNA dissolved in 30 μl of RNAse-free water was stored in −80°C. RNA quality control was determined by gel electrophoresis. Reverse transcription was performed with 1 μg of total RNA, random primers and SuperScript III RNase H-Reverse Transcriptase (Invitrogen) in 20 μl.
Quantitative RT-PCR for TSLP transcript
Expression levels of TSLP transcripts were examined by real-time qPCR with a 7500 Real-Time PCR System using SYBR Green dye (Applied Biosystems), iQ SYBR Green Supermix (Bio-Rad) or fluorescent reporter probes. Primer sequences are provided in supplemental information. We either performed absolute quantification with a standard curve or relative quantification of expression according to the “ΔΔC methods” (Livak and Schmittgen, 2001) using Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) as a reference. The specificity of the PCRs were checked by recording a melting curve and by sequencing the amplicons.
Statistics
Student t-test was used to determine the significance in graphs. P-value below 0.05 was accepted as a significant difference. For RNA-seq, the Audic Claverie Test was used for statistical significant difference (P < 0.05) with Benjamini Hochberg FDR multiple testing correction.
Acknowledgments
We thank the Washington University School of Medicine (WUSM) Genome Technology Access Center (GTAC) for microarray-transcriptome analysis. We also thank James Malone and the Alvin J. Siteman Cancer Center at WSUM and Barnes-Jewish Hospital the Proteomics Core analyses. The Siteman Cancer Center and GTAC are supported in part by an NCI Cancer Center Support Grant #P30 CA91842; GTAC is also supported by ICTS/CTSA Grant# UL1 TR000448 from the National Center for Research Resources (NCRR). We thank Hung-Chi Liang and Shawn Smith for total RNA amplification, David Fletcher and CCHMC DNA sequencing core for RNA-seq analysis. We thank Dr. Gail Martin (Msx2-Cre+/tg mice), Dr. Tom Gridley (Notch2f/f mice), Dr. Jie Shen (Psen1f/f) mice, and Dr. Tasuku Honjo (Rbpjf/f) mice. This work was supported by grant from the National Institutes of Health (R01-GM055479-18 to R.K).
Abbreviations
- NICD
Notch Intracellular Domain
- LICD
Ligand Intracellular Domain
- ECD
Extracellular Domain
- RBPj
Recombining binding protein Jk
- TSLP
Thymic Stromal Lymphopoietin
- HF
Hair Follicle
- msx2LOF
Msx2-Cre+/tg mediated loss-of-function
- BLPD
B-cell lymphoproliferative disease
- WBC
White blood cell
- f
floxed
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
The authors state no conflict of interest.
Author Contributions
M.T. and R.K. conceived and designed the experiments and wrote the manuscript. M.T. performed the experiments. R.R.T designed and supervised the proteomic analysis and described the method.
Microarray and RNA-Seq data are deposited in GEO under accession numbers GSE77813 (E18.5 epidermis microarray), GSE77887 (P0 total skin microarray), and GSE77997 (P3 HF RNA-Seq).
References
- Artavanis-Tsakonas S, Rand MD, Lake RJ. Notch signaling: cell fate control and signal integration in development. Science. 1999;284:770–6. doi: 10.1126/science.284.5415.770. [DOI] [PubMed] [Google Scholar]
- Aubin-Houzelstein G. Notch signaling and the developing hair follicle. Advances in experimental medicine and biology. 2012;727:142–60. doi: 10.1007/978-1-4614-0899-4_11. [DOI] [PubMed] [Google Scholar]
- Bland CE, Kimberly P, Rand MD. Notch-induced proteolysis and nuclear localization of the Delta ligand. The Journal of biological chemistry. 2003;278:13607–10. doi: 10.1074/jbc.C300016200. [DOI] [PubMed] [Google Scholar]
- Blanpain C, Lowry WE, Pasolli HA, Fuchs E. Canonical notch signaling functions as a commitment switch in the epidermal lineage. Genes Dev. 2006;20:3022–35. doi: 10.1101/gad.1477606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blat Y, Meredith JE, Wang Q, Bradley JD, Thompson LA, Olson RE, et al. Mutations at the P1(′) position of Notch1 decrease intracellular domain stability rather than cleavage by gamma-secretase. Biochem Biophys Res Commun. 2002;299:569–73. doi: 10.1016/s0006-291x(02)02705-5. [DOI] [PubMed] [Google Scholar]
- Bordonaro M, Tewari S, Atamna W, Lazarova DL. The Notch ligand Delta-like 1 integrates inputs from TGFbeta/Activin and Wnt pathways. Experimental cell research. 2011;317:1368–81. doi: 10.1016/j.yexcr.2011.03.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bray S, Bernard F. Notch targets and their regulation. Curr Top Dev Biol. 2010;92:253–75. doi: 10.1016/S0070-2153(10)92008-5. [DOI] [PubMed] [Google Scholar]
- Bray SJ. Notch signalling: a simple pathway becomes complex. Nature reviews Molecular cell biology. 2006;7:678–89. doi: 10.1038/nrm2009. [DOI] [PubMed] [Google Scholar]
- Byrne C, Tainsky M, Fuchs E. Programming gene expression in developing epidermis. Development. 1994;120:2369–83. doi: 10.1242/dev.120.9.2369. [DOI] [PubMed] [Google Scholar]
- Chen S, Tao J, Bae Y, Jiang MM, Bertin T, Chen Y, et al. Notch gain of function inhibits chondrocyte differentiation via Rbpj-dependent suppression of Sox9. J Bone Miner Res. 2013;28:649–59. doi: 10.1002/jbmr.1770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demehri S, Liu Z, Lee J, Lin MH, Crosby SD, Roberts CJ, et al. Notch-deficient skin induces a lethal systemic B-lymphoproliferative disorder by secreting TSLP, a sentinel for epidermal integrity. PLoS biology. 2008;6:e123. doi: 10.1371/journal.pbio.0060123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demehri S, Morimoto M, Holtzman MJ, Kopan R. Skin-derived TSLP triggers progression from epidermal-barrier defects to asthma. PLoS Biol. 2009a;7:e1000067. doi: 10.1371/journal.pbio.67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demehri S, Turkoz A, Kopan R. Epidermal Notch1 loss promotes skin tumorigenesis by impacting the stromal microenvironment. Cancer cell. 2009b;16:55–66. doi: 10.1016/j.ccr.2009.05.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demehri S, Turkoz A, Manivasagam S, Yockey LJ, Turkoz M, Kopan R. Elevated Epidermal Thymic Stromal Lymphopoietin Levels Establish An Anti-Tumor Environment In The Skin. Cancer Cell. 2012;4:494–505. doi: 10.1016/j.ccr.2012.08.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demehri S, Yockey LJ, Visness CM, Jaffee KF, Turkoz A, Wood RA, et al. Circulating TSLP associates with decreased wheezing in non-atopic preschool children: Data from the URECA birth cohort. Clin Exp Allergy. 2014 doi: 10.1111/cea.12270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dumortier A, Durham AD, Di Piazza M, Vauclair S, Koch U, Ferrand G, et al. Atopic dermatitis-like disease and associated lethal myeloproliferative disorder arise from loss of Notch signaling in the murine skin. PloS one. 2010;5:e9258. doi: 10.1371/journal.pone.0009258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuchs E, Green H. Changes in keratin gene expression during terminal differentiation of the keratinocyte. Cell. 1980;19:1033–42. doi: 10.1016/0092-8674(80)90094-x. [DOI] [PubMed] [Google Scholar]
- Goodfellow H, Krejci A, Moshkin Y, Verrijzer CP, Karch F, Bray SJ. Gene-specific targeting of the histone chaperone asf1 to mediate silencing. Dev Cell. 2007;13:593–600. doi: 10.1016/j.devcel.2007.08.021. [DOI] [PubMed] [Google Scholar]
- Gordon WR, Arnett KL, Blacklow SC. The molecular logic of Notch signaling–a structural and biochemical perspective. Journal of cell science. 2008;121:3109–19. doi: 10.1242/jcs.035683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hiratochi M, Nagase H, Kuramochi Y, Koh CS, Ohkawara T, Nakayama K. The Delta intracellular domain mediates TGF-beta/Activin signaling through binding to Smads and has an important bi-directional function in the Notch-Delta signaling pathway. Nucleic acids research. 2007;35:912–22. doi: 10.1093/nar/gkl1128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huppert S, Ilagan MXG, De Strooper B, Kopan R. Analysis of Notch function in presomitic mesoderm suggests a g-secretase-independent role for presenilins in somite differentiation. Dev Cell. 2005;8:677–88. doi: 10.1016/j.devcel.2005.02.019. [DOI] [PubMed] [Google Scholar]
- Huppert S, Le A, Schroeter EH, Mumm JS, Saxena MT, Milner LA, et al. Embryonic Lethality in Mice Homozygous for a Processing Deficient Allele of Notch1. Nature. 2000;405:966–70. doi: 10.1038/35016111. [DOI] [PubMed] [Google Scholar]
- Ikeuchi T, Sisodia SS. The Notch ligands, Delta1 and Jagged2, are substrates for presenilin-dependent “gamma-secretase” cleavage. The Journal of biological chemistry. 2003;278:7751–4. doi: 10.1074/jbc.C200711200. [DOI] [PubMed] [Google Scholar]
- Jamora C, DasGupta R, Kocieniewski P, Fuchs E. Links between signal transduction, transcription and adhesion in epithelial bud development. Nature. 2003;422:317–22. doi: 10.1038/nature01458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koelzer S, Klein T. Regulation of expression of Vg and establishment of the dorsoventral compartment boundary in the wing imaginal disc by Suppressor of Hairless. Developmental biology. 2006;289:77–90. doi: 10.1016/j.ydbio.2005.10.008. [DOI] [PubMed] [Google Scholar]
- Kopan R, Ilagan MX. The canonical Notch signaling pathway: unfolding the activation mechanism. Cell. 2009;137:216–33. doi: 10.1016/j.cell.2009.03.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kopan R, Lee J, Lin MH, Syder AJ, Kesterson J, Crutchfield N, et al. Genetic mosaic analysis indicates that the bulb region of coat hair follicles contains a resident population of several active multipotent epithelial lineage progenitors. Developmental biology. 2002;242:44–57. doi: 10.1006/dbio.2001.0516. [DOI] [PubMed] [Google Scholar]
- Kovall RA, Blacklow SC. Mechanistic insights into Notch receptor signaling from structural and biochemical studies. Curr Top Dev Biol. 2010;92:31–71. doi: 10.1016/S0070-2153(10)92002-4. [DOI] [PubMed] [Google Scholar]
- LaVoie MJ, Selkoe DJ. The Notch ligands, Jagged and Delta, are sequentially processed by alpha-secretase and presenilin/gamma-secretase and release signaling fragments. The Journal of biological chemistry. 2003;278:34427–37. doi: 10.1074/jbc.M302659200. [DOI] [PubMed] [Google Scholar]
- Lee J, Basak JM, Demehri S, Kopan R. Bi-compartmental communication contributes to the opposite proliferative behavior of Notch1-deficient hair follicle and epidermal keratinocytes. Development. 2007;134:2795–806. doi: 10.1242/dev.02868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin HY, Kao CH, Lin KM, Kaartinen V, Yang LT. Notch signaling regulates late-stage epidermal differentiation and maintains postnatal hair cycle homeostasis. PloS one. 2011;6:e15842. doi: 10.1371/journal.pone.0015842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin MH, Leimeister C, Gessler M, Kopan R. Activation of the Notch pathway in the hair cortex leads to aberrant differentiation of the adjacent hair-shaft layers. Development. 2000;127:2421–32. doi: 10.1242/dev.127.11.2421. [DOI] [PubMed] [Google Scholar]
- Liu Z, Brunskill E, Varnum-Finney B, Zhang C, Zhang A, Jay YP, et al. The intracellular Domains of Notch1 and 2 Are Functionally Equivalent During Development and Carcinogenesis. Development. 2015 doi: 10.1242/dev.125492. in press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25:402–8. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
- Muller-Rover S, Handjiski B, van der Veen C, Eichmuller S, Foitzik K, McKay IA, et al. A comprehensive guide for the accurate classification of murine hair follicles in distinct hair cycle stages [Review] Journal of Investigative Dermatology. 2001;117:3–15. doi: 10.1046/j.0022-202x.2001.01377.x. [DOI] [PubMed] [Google Scholar]
- Olave I, Reinberg D, Vales LD. The Mammalian Transcriptional Repressor Rbp (Cbf1) Targets Tfiid and Tfiia to Prevent Activated Transcription. Genes & Development. 1998;12:1621–37. doi: 10.1101/gad.12.11.1621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pan Y, Lin M, Tian X, Cheng H, Gridley T, Shen J, et al. g-Secretase functions through Notch signaling to maintain skin appendages but is not required for their patterning or initial morphogenesis. Dev Cell. 2004;7:731–43. doi: 10.1016/j.devcel.2004.09.014. [DOI] [PubMed] [Google Scholar]
- Pellegrinet L, Rodilla V, Liu Z, Chen S, Koch U, Espinosa L, et al. Dll1- and dll4-mediated notch signaling are required for homeostasis of intestinal stem cells. Gastroenterology. 2011;140:1230–40. e1–7. doi: 10.1053/j.gastro.2011.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qi H, Rand MD, Wu X, Sestan N, Wang W, Rakic P, et al. Processing of the notch ligand delta by the metalloprotease Kuzbanian. Science. 1999;283:91–4. doi: 10.1126/science.283.5398.91. [DOI] [PubMed] [Google Scholar]
- Sanalkumar R, Dhanesh SB, James J. Non-canonical activation of Notch signaling/target genes in vertebrates. Cellular and molecular life sciences: CMLS. 2010;67:2957–68. doi: 10.1007/s00018-010-0391-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sato C, Turkoz M, Dearborn JT, Wozniak DF, Kopan R, Hass MR. Loss of RBPj does not cause neurodegeneration or memory impairments but disrupts olfactory function in aged mice. PLoS ONE. 2012;7:e48180. doi: 10.1371/journal.pone.0048180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt-Ullrich R, Paus R. Molecular principles of hair follicle induction and morphogenesis. BioEssays: news and reviews in molecular, cellular and developmental biology. 2005;27:247–61. doi: 10.1002/bies.20184. [DOI] [PubMed] [Google Scholar]
- Six E, Ndiaye D, Laabi Y, Brou C, Gupta-Rossi N, Israel A, et al. The Notch ligand Delta1 is sequentially cleaved by an ADAM protease and gamma-secretase. Proceedings of the National Academy of Sciences of the United States of America. 2003;100:7638–43. doi: 10.1073/pnas.1230693100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skalska L, Stojnic R, Li J, Fischer B, Cerda-Moya G, Sakai H, et al. Chromatin signatures at Notch-regulated enhancers reveal large-scale changes in H3K56ac upon activation. EMBO J. 2015 doi: 10.15252/embj.201489923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun X, Lewandoski M, Meyers EN, Liu YH, Maxson RE, Jr, Martin GR. Conditional inactivation of Fgf4 reveals complexity of signalling during limb bud development. Nature genetics. 2000;25:83–6. doi: 10.1038/75644. [DOI] [PubMed] [Google Scholar]
- Tadeu AM, Horsley V. Notch signaling represses p63 expression in the developing surface ectoderm. Development. 2013;140:3777–86. doi: 10.1242/dev.093948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tagami S, Okochi M, Yanagida K, Ikuta A, Fukumori A, Matsumoto N, et al. Regulation of Notch Signaling by Dynamic Changes in the Precision in S3 Cleavage of Notch-1. Mol Cell Biol. 2008;28:165–76. doi: 10.1128/MCB.00863-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vauclair S, Nicolas M, Barrandon Y, Radtke F. Notch1 is essential for postnatal hair follicle development and homeostasis. Developmental biology. 2005;284:184–93. doi: 10.1016/j.ydbio.2005.05.018. [DOI] [PubMed] [Google Scholar]
- Watt FM, Estrach S, Ambler CA. Epidermal Notch signalling: differentiation, cancer and adhesion. Current opinion in cell biology. 2008;20:171–9. doi: 10.1016/j.ceb.2008.01.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zolkiewska A. ADAM proteases: ligand processing and modulation of the Notch pathway. Cellular and molecular life sciences: CMLS. 2008;65:2056–68. doi: 10.1007/s00018-008-7586-4. [DOI] [PMC free article] [PubMed] [Google Scholar]


