Abstract
Phosphoinositides (PIs) control fundamental cell processes, and inherited defects of PI kinases or phosphatases cause severe human diseases including Lowe syndrome due to mutations in OCRL that encodes a PI(4,5)P2 5-phosphatase. Here we unveil a lysosomal response to the arrival of autophagosomal cargo where OCRL plays a key role. We identify mitochondrial DNA and TLR9 as the cargo and the receptor that triggers and mediates, respectively, this response. This lysosome-cargo response is required to sustain the autophagic flux and involves a local increase in PI(4,5)P2 that is confined in space and time by OCRL. Depleting or inhibiting OCRL leads to an accumulation of lysosomal PI(4,5)P2, an inhibitor of the calcium channel mucolipin-1 that controls autophagosome-lysosome fusion. Hence, autophagosomes accumulate in OCRL-depleted cells and in the kidneys of Lowe syndrome patients. Importantly, boosting the activity of mucolipin-1 with selective agonists restores the autophagic flux in cells from Lowe syndrome patients.
Introduction
Mutations in the OCRL gene, which encodes a PI(4,5)P2 5-phosphatase, cause Oculo Cerebro Renal Lowe syndrome, a rare X-linked syndrome characterized by congenital cataracts, psychomotor disability, and renal Fanconi syndrome due to the inability of proximal tubule kidney cells (PTCs) to reabsorb low molecular proteins and solutes from the ultrafiltrate1, 2. The progression of kidney damage towards end-stage renal failure is a leading cause of death in patients affected by Lowe syndrome, a disease for which no therapy is currently available.
OCRL is a multidomain cytosolic protein that associates with diverse subcellular compartments including clathrin-coated vesicles, early endosomes, the Trans Golgi Network (TGN), and the primary cilium3. The association of OCRL with membranes involves several interactors such as AP2, clathrin, Rab GTPases, APPL1 and Ses proteins3. OCRL has been shown to control endocytic recycling, endosome-to-Golgi transport, early endocytic and phagocytic steps, cytokinesis, and cilium formation3, 4.
The loss of OCRL function leads to an accumulation of PI(4,5)P2 and to uncontrolled actin polymerization on clathrin coated vesicles and early endosomes4, 5. This imbalance in PI(4,5)P2 and actin dynamics impairs the endocytic trafficking of different classes of receptors, including the multiligand receptor megalin that is responsible for low molecular protein reabsorption at the proximal tubule5. While impaired megalin trafficking may explain some signs of the disease, such as low-molecular-weight proteinuria5, many gaps remain in our understanding of the mechanisms linking OCRL mutations to the impairment of proximal tubular cell function, to the progressive deterioration of kidney function, and to the neuropathological signs in Lowe syndrome6, 7. Here, by combining unbiased and targeted approaches, we describe an unanticipated role of OCRL at lysosomes where it is recruited in the course of a response that we find to occur at lysosomes upon arrival of autophagic cargo.
Results
OCRL depletion induces the upregulation of lysosomal genes and morphological alterations of lysosomes
We initially undertook an unbiased approach to uncover how cells respond to the loss of OCRL by analyzing the changes in gene expression caused by the depletion of OCRL in human PTCs, the cells that are primarily compromised in Lowe syndrome (Supplementary Fig. 1a-c and Supplementary Table 1).
We found that one of the gene classes most significantly upregulated after OCRL depletion coded for lysosomal proteins (Fig. 1a, b and Supplementary Fig. 2a). This upregulation was accompanied by the nuclear translocation of the transcription factor TFEB (Supplementary Fig. 2b-d), a master regulator of lysosomal biogenesis that is activated under conditions that require increased degradation efficiency or induce lysosomal dysfunction8. The activation of TFEB and the presence of morphologically abnormal lysosomes in OCRL-depleted cells, in PTCs from Lowe syndrome patients, as well as in a recently described zebrafish model of Lowe syndrome9, 10 (Fig. 1c-e) led us to investigate the role of OCRL at lysosomes.
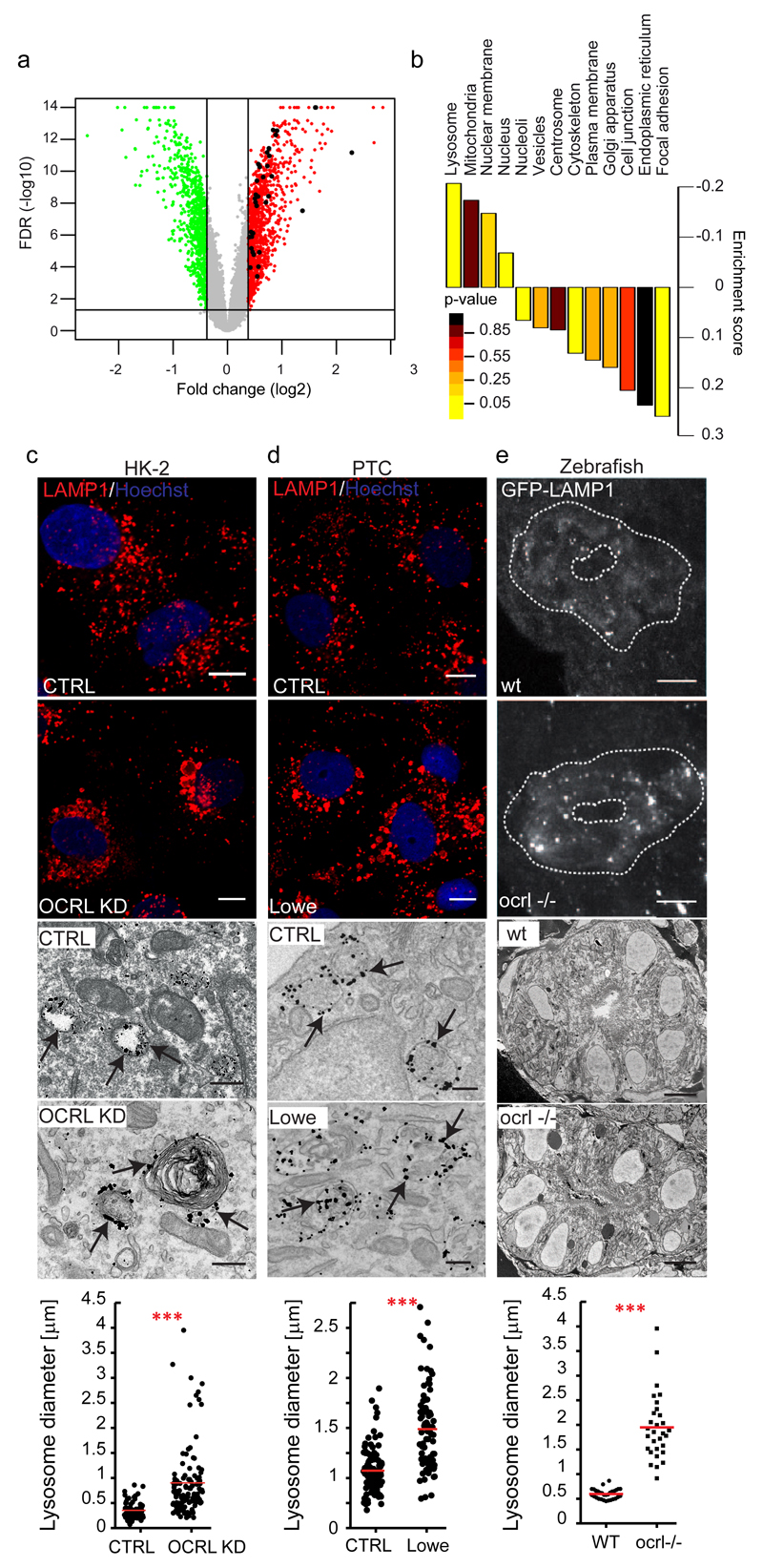
Figure 1. OCRL depletion/mutation induces upregulation of lysosomal genes and morphological changes in lysosomes.
(a) Volcano plot of OCRL-KD gene expression data. Horizontal black line: -log10 of FDR (False Discovery Rate, significance threshold 0.05); vertical black lines: log2 fold change (1.3-fold threshold). 910 genes up-regulated above (red) and 630 genes down-regulated below (green) the threshold are shown. Black dots indicate 24 upregulated (35 probe sets) out of the 194 lysosomal genes annotated in the Lysoplex list50. (b) Gene set enrichment analysis of the data in a. Subcellular compartments, except lysosomes, were defined according to the Human Protein Atlas. Lysosomal genes were defined according to Lysoplex50 and excluded from the other categories. p-values calculated as described in Methods. (c-e) Lysosomal enlargement in OCRL-depleted HK-2 cells, in PTCs of Lowe syndrome patients, and in the proximal pronephric tubule of ocrl-/- zebrafish embryos. (c) Representative images of Control and OCRL-KD cells immunostained with an anti-LAMP1 antibody (upper panels, scale bars, 10 µm) or immunoelectron-microscopy with an anti-LAMP1 antibody (lower panels, scale bars, 250 nm). (d) PTCs obtained from the urine of healthy control subjects (CTRL) and Lowe Syndrome patients were processed as in a. Black arrows indicate LAMP1-positive structures. (e) Confocal transverse sections of zebrafish proximal pronephric tubules from 72 hpf GFP-LAMP1-expressing wild-type (wt) and ocrl-/- mutant embryos9 labelled with an anti-GFP antibody. White dashed lines indicate the outline of pronephric tubules (upper panels). Scale bars, 5 µm. Block face scanning electron microscopy images of transverse sections through proximal pronephric tubules from 72 hpf wild-type (wt) or ocrl-/- embryos (lower panels). Scale bar, 5 µm. Graphs show the morphometric analysis of LAMP1-positive structures in c [n=81 (CTRL) and n=126 (OCRL KD) structures pooled from 3 independent experiments] and d [n=102 (CTRL), n=84 (Lowe) structures pooled from 3 independent experiments]. In e the external circumference of each lysosomal compartment (electron dense oval or spherical membrane-enclosed organelle) was manually traced in each section and the average calculated. n=30 sections (wt); n=30 sections (ocrl-/-) (10 sections analyzed for three kidneys per category). Data are presented as means ± s.d. ***p < 0.001, calculated by two-tailed Student's t-test.
OCRL is recruited to lysosomes in response to autophagosome-lysosome fusion in an AP2- and clathrin-dependent manner
As mentioned above OCRL has been reported to associate with clathrin-coated pits, clathrin-coated vesicles, early endosomes, the TGN, and the primary cilium3. However, this distribution pattern apparently contrasts with the first reported localization of OCRL, which indicated that OCRL associates with lysosomes that were however loaded with sucrose11. We reasoned that this apparent discrepancy might be due to the different loading state of lysosomes and therefore analysed the distribution of OCRL under a condition that induces a lysosomal cargo load such as starvation, during which lysosomes receive autophagosomal cargoes. We confirmed that OCRL associates mainly with early endosomes and the TGN (Supplementary Fig. 1b) and only occasionally colocalizes with the lysosomal marker LAMP1 under steady state conditions (Fig. 2a, b). However, it massively translocates to lysosomes upon induction of autophagy (Fig. 2a, b). The observation that this translocation was diminished when the fusion of autophagosomes with lysosomes was inhibited by depleting key components of the tethering (VPS16) and fusion (STX17) machinery or interfering with microtubule dynamics (vinblastine treatment)12–14 (Fig. 2a, b) indicated that the fusion of autophagosomes with lysosomes triggers the recruitment of OCRL to lysosomes and prompted us to look for other components and for the signalling pathways mediating this response.
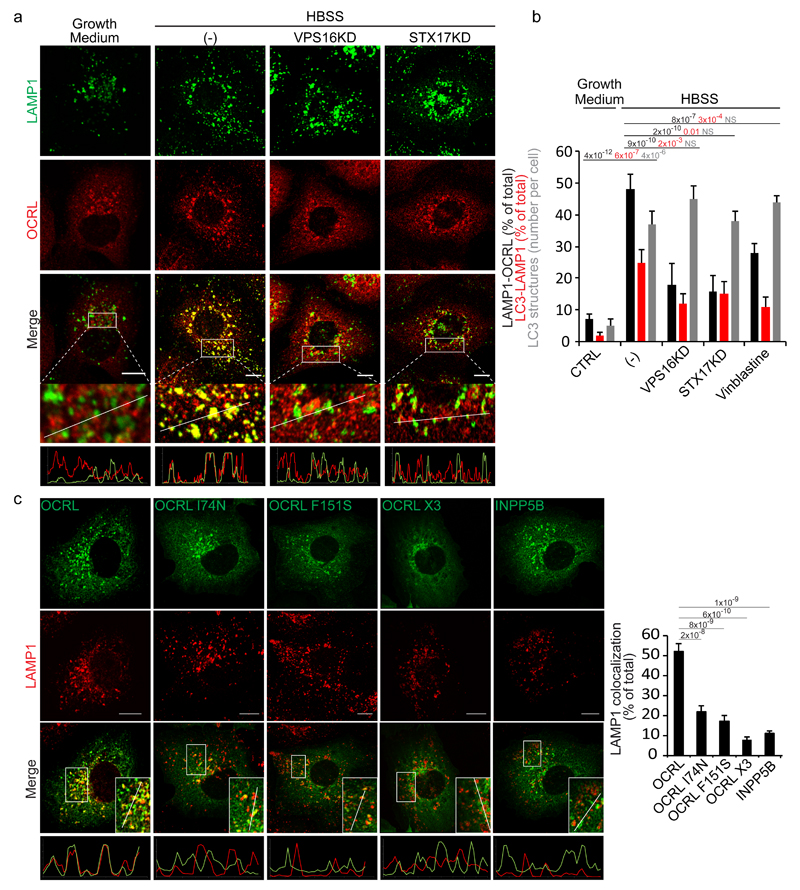
Figure 2. Autophagosome-lysosome fusion induces an AP2 and clathrin-dependent recruitment of OCRL to lysosomes.
(a) Fusion of autophagosomes with lysosomes recruits OCRL to lysosomes. HK-2 cells were incubated in growth medium, or in HBSS for 3 hours (-), or in HBSS for 3 hours after impairment of autophagosome-lysosome fusion by siRNA-mediated KD of the HOPS component VPS1612 or of the autophagosomal SNARE STX1714. Cells were stained for OCRL and LAMP1 (a lysosomal marker). The lower panels are enlargements of the boxed areas and fluorescence intensity profiles in the green and red channels of the regions underneath the white lines. Scale bars, 10 µm. (b) Quantification of OCRL association with lysosomes under conditions described in a, or after the addition of vinblastine (20 μM) that also inhibits autophagosome-lysosome fusion13. The percentage of LAMP1 structures that are positive for OCRL, the percentage of LC3 structures that are positive for LAMP1 (as a measure of the arrival of autophagosomal cargo to lysosomes), and the average number of LC3 structures per cell are reported. Values are means ± s.d. of n=450 cells pooled from 3 independent experiments. NS (not significant). (c) Representative images and quantification of the colocalization of wt OCRL, the AP2 (OCRL-F151S), clathrin(OCRL-I74N) or AP2-clathrin triple (OCRL-X3) binding mutants, and INPP5B with LAMP1. Insets, enlargements of the boxed areas; lower panels, fluorescence intensity profiles in the green and red channels of the regions underneath the white lines. Scale bars, 10 µm. Data represent the percentage of total LAMP1 structures positive for each OCRL form or for INPP5B. Means ± s.d. n=200 cells pooled from 3 independent experiments; n=100 OCRL-X3 transfected cells pooled from 3 independent experiments. p-values calculated by One-way ANOVA with Tukey's post hoc test.
We first explored the involvement of clathrin and AP2, two main recruiters of OCRL to cellular membranes15, 16, and found that mutations in the AP2- and/or clathrin-binding sites of OCRL impaired the translocation of OCRL to lysosomes16 (Fig. 2c). Confirming the importance of these two binding sites, INPP5B, a PI(4,5)P2 5-phosphatase that is highly similar to OCRL but lacks both the AP2- and clathrin-interacting motifs16, is not recruited to lysosomes in response to the delivery of autophagic cargo (Fig. 2c).
Autophagosome-lysosome fusion triggers an increase in lysosome PI(4,5)P2 and the recruitment of AP2 and clathrin to lysosomes
Prompted by the observation that AP2 and clathrin were required for the association of OCRL with lysosomes in response to autophagosome-lysosome fusion, we followed their localization during the course of the autophagy process. AP2 and clathrin have been reported to associate with lysosomes17 and to be required for lysosome reformation at the end of autophagy upon mTOR signalling reactivation17–19. We found that AP2 and clathrin are in fact recruited to lysosomes early during the autophagy process at a stage when mTOR is still inactive, and in a manner that is concomitant with and dependent on the fusion of autophagosomes with lysosomes and a local increase in PI(4,5)P2 produced by PI4KIIIβ, a kinase that preserves lysosomal identity20, and the kinases PIP5K1α and β (Fig. 3a-c, Supplementary Fig. 3a, b). To search for signalling pathways mediating the lysosomal response to autophagosome fusion, we first considered the lysosome-based mTOR signalling21. However, multiple lines of evidence indicated that the local increase in lysosomal PI(4,5)P2 and the recruitment of OCRL are not dependent on mTOR activation or inactivation: first, OCRL is more associated with lysosomes (as compared to steady state) under conditions in which mTOR is either silent or fully active (Fig. 3b and Supplementary Fig. 3c); second, the overall time course of OCRL translocation to lysosomes does not parallel that of mTOR inactivation-reactivation during nutrient deprivation-induced autophagy followed by nutrient re-addition (Fig. 3b); third, it is not starvation (and thus mTOR inactivation) per se but the arrival of autophagosomal cargo to lysosomes that induces this response (Fig. 3c); fourth, OCRL translocation can be seen using agents that induce autophagy and inactivate mTOR (torin 1) or induce autophagy without affecting mTOR activity (Tat-Beclin-1)22 (Supplementary Fig. 3d, e).
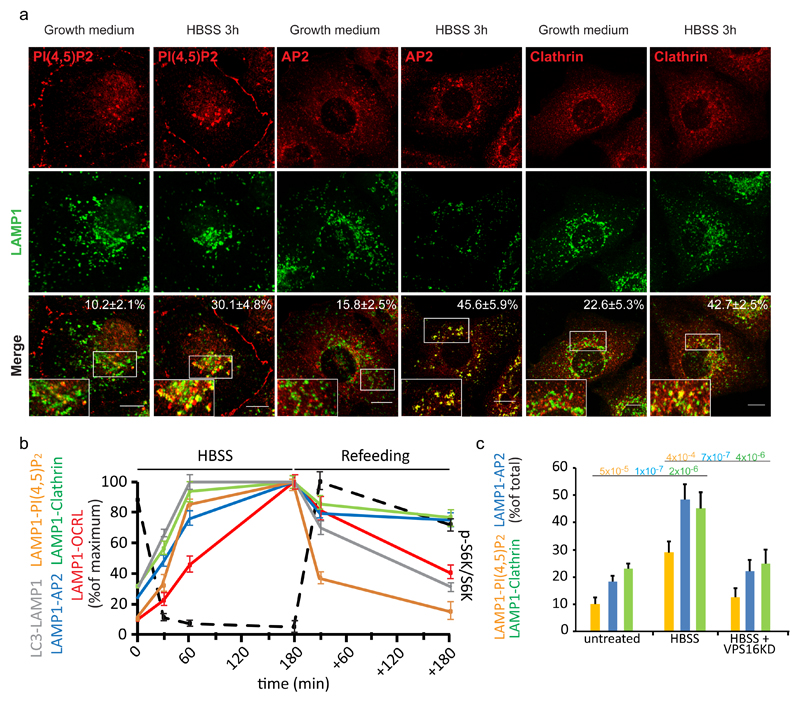
Figure 3. Autophagosome-lysosome fusion induces a local increase in PI(4,5)P2 and AP2 and clathrin recruitment to lysosomes.
(a) Representative images and quantification of the recruitment of the indicated proteins and of lysosomal PI(4,5)P2 levels during starvation-induced autophagy in HK-2 cells. Cells were incubated in growth medium or in HBSS for 3 hours, fixed and stained with an anti-LAMP1 antibody together with an anti-AP2, anti-clathrin or anti-PI(4,5)P2 antibody. Scale bars, 10 μm. Insets, enlargements of the boxed areas. Means ± s.d., n=450 cells per condition pooled from 6 independent experiments. (b) Quantification of the arrival of autophagic cargo to lysosomes (LC3-LAMP1), of the level of lysosomal PI(4,5)P2 (assessed by anti-PI(4,5)P2 antibody staining, LAMP1-PI(4,5)P2), and of the recruitment of AP2 (LAMP1-AP2), clathrin (LAMP1-clathrin), and OCRL (LAMP1-OCRL) to lysosomes at 30, 60 and 180 min of HBSS treatment and 30 min and 180 min after complete medium re-addition (Refeeding). At the same time points mTOR activity status was analyzed (black dotted line) as the ratio of phospho-S6 kinase to total S6 kinase (a representative blot is reported in Supplementary Fig. 3c and an unprocessed scan of the blot is shown in Supplementary Fig. 9). Data are expressed as percentage of the maximum value. Mean values ± s.d. n=450 cells per condition pooled from 3 independent experiments. (c) CTRL (mock-treated) cells and VPS16-KD cells (to impair autophagosome-lysosome fusion) were incubated in growth medium or in HBSS for 3 hours, fixed and stained as above. The graph reports the percentage of total LAMP1 structures that were positive for PI(4,5)P2, AP2, and clathrin. Means ± s.d. of n=450 cells per condition pooled from 3 independent experiments. p-values calculated by one-way ANOVA with Tukey's post-hoc test. Statistic source data can be found in Supplementary Table 2.
TLR9 mediates the lysosomal response to autophagosome fusion
We therefore considered the involvement of other endolysosome-based signalling pathways, such as members of the family of toll like receptors (TLRs) that are present in immune as well as in non-immune cells (including kidney proximal tubule cells23–25) and recognize both exogenous and endogenous macromolecules26–28. We depleted TLR family members that can signal from endolysosomes (i.e. TLR3, TLR4, TLR7, TLR9) and found that the response to autophagosome-lysosome fusion was blunted – in terms of PI(4,5)P2 generation, AP2, clathrin and OCRL recruitment – by selectively depleting or pharmacologically inhibiting TLR9 (Fig. 4a-c). TLR9 resides mainly in the ER at steady state (with only a small fraction of receptors present in the endolysosomal compartment) but upon arrival of TLR9 ligands into lysosomes it translocates to lysosomes via mechanisms that are only partially understood and that involve COPII-dependent ER export, the chaperone UNC93B1, and the small GTPase ARF329, 30. In line with the notion that TLR9 activation is accompanied by its translocation to lysosomes, we found that TLR9 translocates from the ER to lysosomes upon autophagy induction (Fig. 4d and Supplementary Fig. 4a) in a fashion that is dependent on autophagosome-lysosome fusion and on the presence of the TLR9 chaperone UNC93B1 (Fig. 4d).
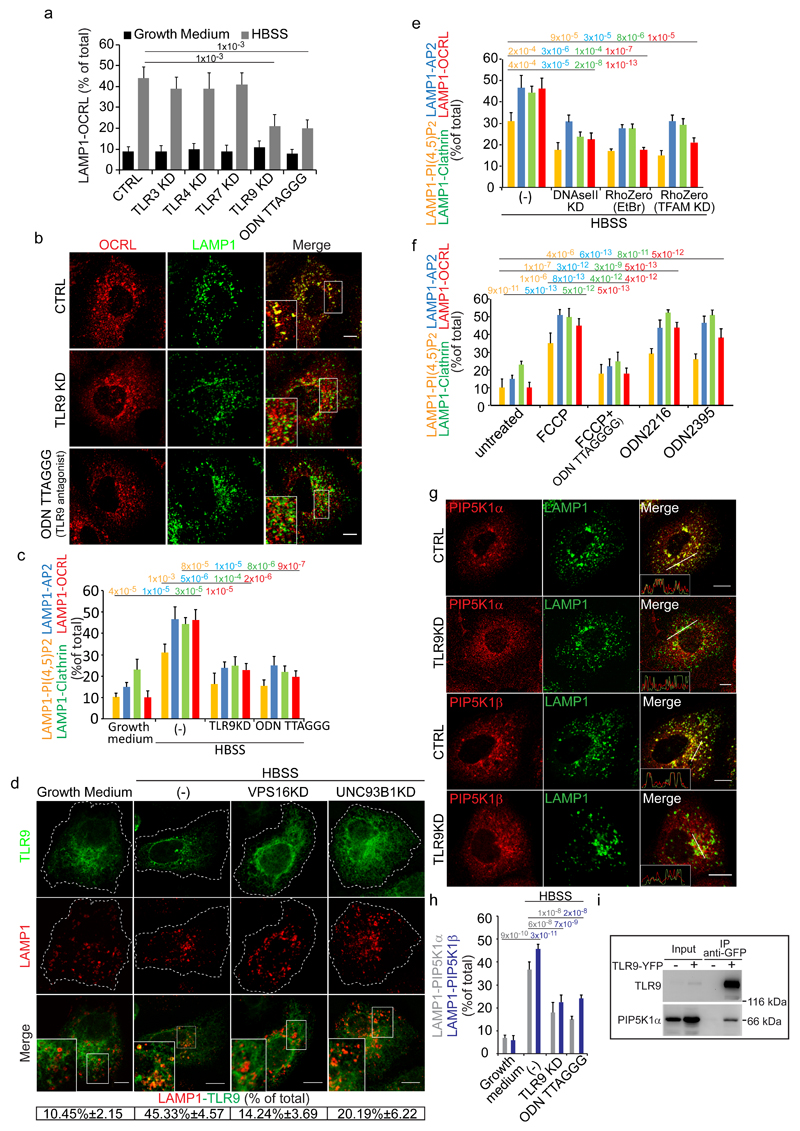
Figure 4. TLR9 stimulated by mitochondrial DNA released into lysosomes by autophagosomes mediates the lysosome cargo response.
(a) Colocalization of LAMP1 and OCRL in control cells, cells KD for the indicated TLRs or treated with the TLR9 antagonist ODN TTAGGG (0.5 μM). Mean values ± s.d. n=100 cells per condition pooled from 3 independent experiments. (b) OCRL association with lysosomes in CTRL, TLR9-KD and ODN TTAGGG-treated starved cells. Insets, enlargements of the boxed areas. (c) Effects of TLR9 depletion/inhibition on the lysosomal cargo response Mean values ± s.d. n=200 cells per condition pooled from 3 independent experiments. (d) Colocalization of TLR9-YFP with LAMP1 under the indicated conditions. Insets, enlargements of the boxed areas. Mean values ± s.d. n=100 cells per condition pooled from 3 independent experiments. p < 0.01 for starved vs. fed CTRL cells and starved CTRL cells vs. starved VPS16KD and UNC93B1KD cells. (e) Colocalization of PI(4,5)P2 and the indicated proteins with LAMP1 in starved CTRL cells (-), or after DNAseII KD, or after mtDNA depletion (RhoZero) obtained as specified in Supplementary Fig. 4b. Mean values ± s.d. n=200 cells per condition pooled from 3 independent experiments. (f) HK-2 cells untreated or treated with FCCP (10 µM) alone or with TLR9 antagonist (ODNTTAGGG), or treated with TLR9 agonists (ODN2216 and ODN2395) and stained for PI(4,5)P2 and the indicated proteins. Mean values ± s.d. n=450 cells per condition pooled from 3 independent experiments. (g) CTRL and TLR9-KD starved cells stained for LAMP1 and for PIP5K1α or PIP5K1β. Insets, fluorescence intensity profiles in the green and red channels of the regions underneath the white lines. (h) Quantification of PIP5K1α or PIP5K1β colocalization with LAMP1. Mean values ± s.d. n=100 cells per condition pooled from 3 independent experiments. (i) Cell lysates from non-transfected or TLR9-YFP transfected cells were immunoprecipitated with anti-GFP antibody and analyzed by Western blot. Results representative of five independent experiments. An unprocessed scan of the blot is in Supplementary Fig. 9. p-values are indicated, calculated by One-way ANOVA with Tukey's post-hoc test. Scale bars, 10 μm. Statistic source data in Supplementary Table 2.
TLR9 is activated by mitochondrial DNA released into lysosomes by autophagosomes
The ligands for TLR9 are unmethylated CpG motifs that are found in bacterial DNA and in mitochondrial DNA (mtDNA)31. Indeed, the activation of TLR9 in response to mitochondrial damage and increased autophagy has been reported recently in non-immune cells such as cardiomyocytes32 and hepatocytes33. We thus hypothesized that the autophagic cargo that triggers the TLR9-mediated lysosomal response might be mtDNA released into lysosomes by autophagosomes. In support of our hypothesis, depleting mitochondrial DNA (by two independent means, Supplementary Fig. 4b, c) or depleting DNaseII (the lysosomal DNase required for TLR9 activation34) inhibited the lysosomal response (Fig. 4e, Supplementary Fig. 4d), while inducing mitophagy with FCCP (a mitochondrial uncoupler that induces mitochondrial damage35–37) or activating TLR9 with synthetic agonists mimicking unmethylated CpG motifs elicited the lysosomal response in terms of an increase in lysosomal PI(4,5)P2 and the recruitment of AP2, clathrin and OCRL (Fig. 4f).
We investigated the signalling cascade triggered by the activation of TLR9 in response to mtDNA and found that known components of the TLR9 signalling and trafficking pathway – such as TIRAP, which also translocated to lysosomes (Supplementary Fig. 4e), MyD88, IRAK4 and UNC93B1 – were involved (Supplementary Fig. 4f). We next asked how the activation of TLR9 could lead to the local increase in PI(4,5)P2. Although a link between the activation of TLR and PI(4,5)P2 generation has been previously reported, the underlying mechanisms have remained unexplored so far38. We found that the activation of TLR9 is accompanied by and required for the translocation of PIP5Kα and β to lysosomes (Fig. 4g, h) and that TLR9 belongs to a molecular complex containing also PIP5Kα, since the two proteins can be co-immunoprecipitated (Fig. 4i).
The TLR9-mediated lysosomal cargo response sustains the autophagic flux
Altogether, the above data unmasked a regulatory circuit in lysosomes that is activated by autophagic cargo and prompted us to ask what might be the purpose(s) of this circuit and what is the specific role of OCRL in it. We found that this circuit controls local (i.e. lysosomal) and transcriptional responses. Locally, it is required for lysosomal homeostasis since lysosomes enlarge and accumulate internal membranes upon depletion or inhibition of TLR9 (Fig. 5a, b) and for efficient autophagic flux (Fig. 5c, d) since the depletion or pharmacological inhibition of TLR9 led to the accumulation of autophagosomes (Fig. 5d). This accumulation is accompanied by impaired autophagosome-lysosome fusion (as assessed by following the yellow-to-red conversion of the mRFP-GFP tandem-LC3 reporter39 Fig. 5e) possibly induced by defective recycling of the autophagic SNARE syntaxin1714 from autolysosomes (Supplementary Fig. 4g). Concomitantly with this local lysosomal response, TLR9 stimulation by mtDNA triggers the canonical TLR9-dependent signalling cascade involving the activation and nuclear translocation of the transcription factor NF-kB and an increase in the transcription of proinflammatory cytokine and Interferon beta genes (Fig. 5f, g). This indicates that the mtDNA released into lysosomes during autophagy has the potential to trigger an inflammatory response. We envisage that this response is usually self-limiting due to the complete degradation of oligonucleotides containing unmethylated CpG motifs by lysosomal exonucleases, nucelotidases and phosphatases but might become sustained under conditions of prolonged stress and/or impaired degradative capacity, such as during lysosomal storage disorders, which, in fact, are often accompanied by chronic inflammation40.
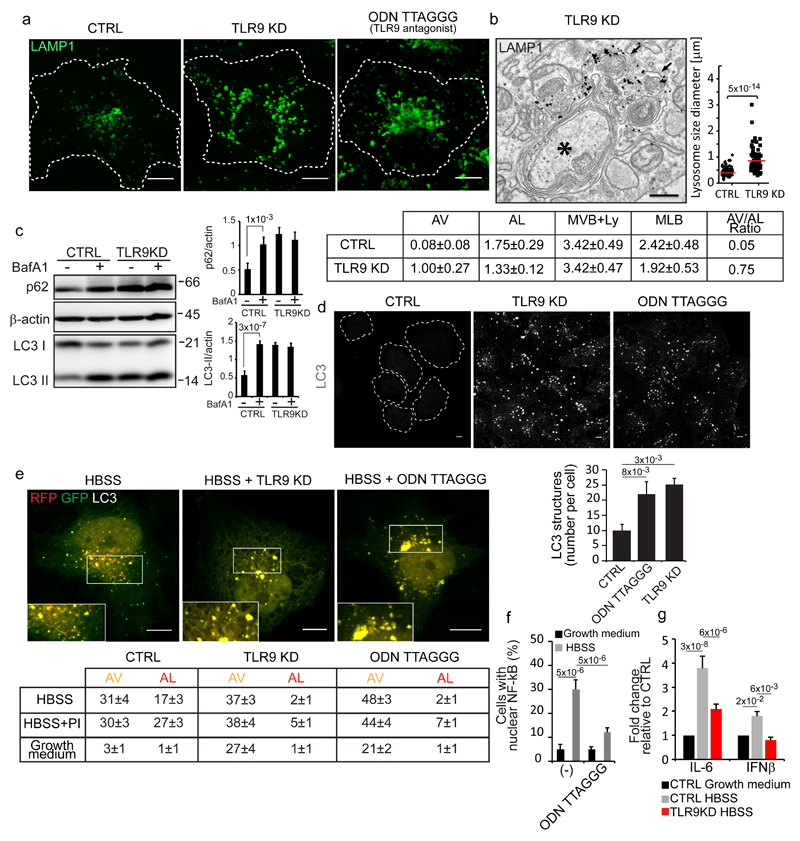
Figure 5. TLR9 is required for lysosomal homeostasis and for efficient autophagic flux.
(a) Control, TLR9-KD and TLR9 antagonist-treated cells were stained for LAMP1. (b) Representative image of autophagosomes (black asterisk) and lysosomes (labelled with anti-LAMP1 antibodies, black arrows) in TLR9-KD cells. The graph shows the size distribution of LAMP1-positive structures, n=85 (CTRL) and n=80 (TLR9-KD) structures pooled from 3 independent experiments. The table reports the morphometric analysis: Autophagosomes (AV), Autolysosomes (AL); Multivesicular bodies and electron-dense lysosomes (MVB+Ly); Multilamellar bodies (MLB). p ≤ 0.01 for AV and MLB in TLR9-KD relative to CTRL cells. Mean values ± s.e.m. (c) Cell lysates were probed with the indicated antibodies. BafA1 (Bafilomycin A1). β-actin was used as a loading control. Graphs show p62/actin and LC3II/actin ratios. Mean values ± s.d. n=3 lysates per condition pooled from 3 independent experiments. Unprocessed scans of the blots are shown in Supplementary Fig. 9. (d) Control (CTRL), TLR9-KD, and TLR9-antagonist treated cells were fixed and immunostained with an anti-LC3 antibody. The number of LC3-positive structures per cell is reported in the graph. Mean values ± s.d. n=450 cells per condition pooled from 3 independent experiments. (e) Cells expressing mRFP-GFP tandem-tagged LC3 were incubated in growth medium, in HBSS for 3 hours with or without a protease inhibitor cocktail (PI) or the TLR9 antagonist ODN TTAGGG (0.5 μM) as indicated.. Insets are enlargements of the boxed areas. The table reports the means (± s.d.) of AV and AL. n=60 cells per condition pooled from 3 independent experiments. p < 0.001 for TLR9-KD and ODNTTAGGG-treated cells vs. CTRL cells. (f) NF-kB nuclear localization. Mean values ± s.d. n=400 cells per condition pooled from 3 independent experiments. (g) qPCR analysis of IL-6 and IFN-β mRNA levels in the indicated conditions. Mean values ± s.d. n=3 RNAs per condition pooled from 3 independent experiments. p-values are indicated, calculated by One-way ANOVA with Tukey's post hoc test except in b where Student's t-test was used. Statistic source data can be found in Supplementary Table 2. Scale bars, 10 μm in a, d, e and 500 nm in b.
OCRL controls the autophagy flux through its 5-phosphatase activity
We then explored the role of OCRL in the lysosomal cargo response and its relevance for the manifestations of Lowe syndrome. We found that OCRL-depleted cells have a higher number of mature autophagosomes under normal growth conditions41 (Fig. 6a-c, Supplementary Fig. 5a). This elevated number of autophagosomes is due to delayed basal autophagic flux through lysosomes as assessed by the increase in the autophagy substrate p6242 (Supplementary Fig. 5b). Although TFEB activation might in part contribute to the increase in basal p62 levels in OCRL-depleted cells, the inability of Bafilomycin to induce a further increase in p62 levels indicates that this increase is mainly due to impaired autophagic flux to lysosomes, as also testified by the increase in LC3II protein levels (Supplementary Fig. 5b). The impaired autophagic flux is caused by an impairment of autophagosome-lysosome fusion as assessed by the lower colocalization of autophagosomal and lysosomal markers and the impaired yellow-to-red conversion of the tandem fluorescent LC3 probe (Fig. 6d, e and Supplementary Fig. 5c, d). Importantly, a striking accumulation of autophagosomes was also observed in proximal tubules from kidney biopsies of Lowe syndrome patients (Fig. 6f, Supplementary Fig. 6a, b) highlighting the physiopathological relevance of the lysosomal role of OCRL, prompting us to clarify the molecular mechanisms underlying this role. These involve the catalytic activity of OCRL since YU142670, an OCRL inhibitor43, also induced an accumulation of autophagosomes (Fig. 6a, b) while the autophagic flux could be restored by expressing wt but not catalytically-inactive OCRL in OCRL-defective PTCs or in PTCs from Lowe syndrome patients (Fig. 6a, b, g). Of note, we detected an excess of lysosomal PI(4,5)P2 in OCRL-depleted cells (Fig. 7a, b and Supplementary Fig. 5e, f) which could be rescued by re-expressing wt, but not catalytically inactive, OCRL (Supplementary Fig. 5f) or by lowering PI(4,5)P2 production through the depletion of PIP5K α or β (Fig. 7b). Notably, these two independent PI(4,5)P2-reducing approaches were both effective in rescuing the autophagy flux in OCRL-depleted cells (Fig. 6a, b and Fig. 7c) indicating that the excess lysosomal PI(4,5)P2 had a causative role in impairing autophagosome-lysosome fusion in OCRL-depleted cells.
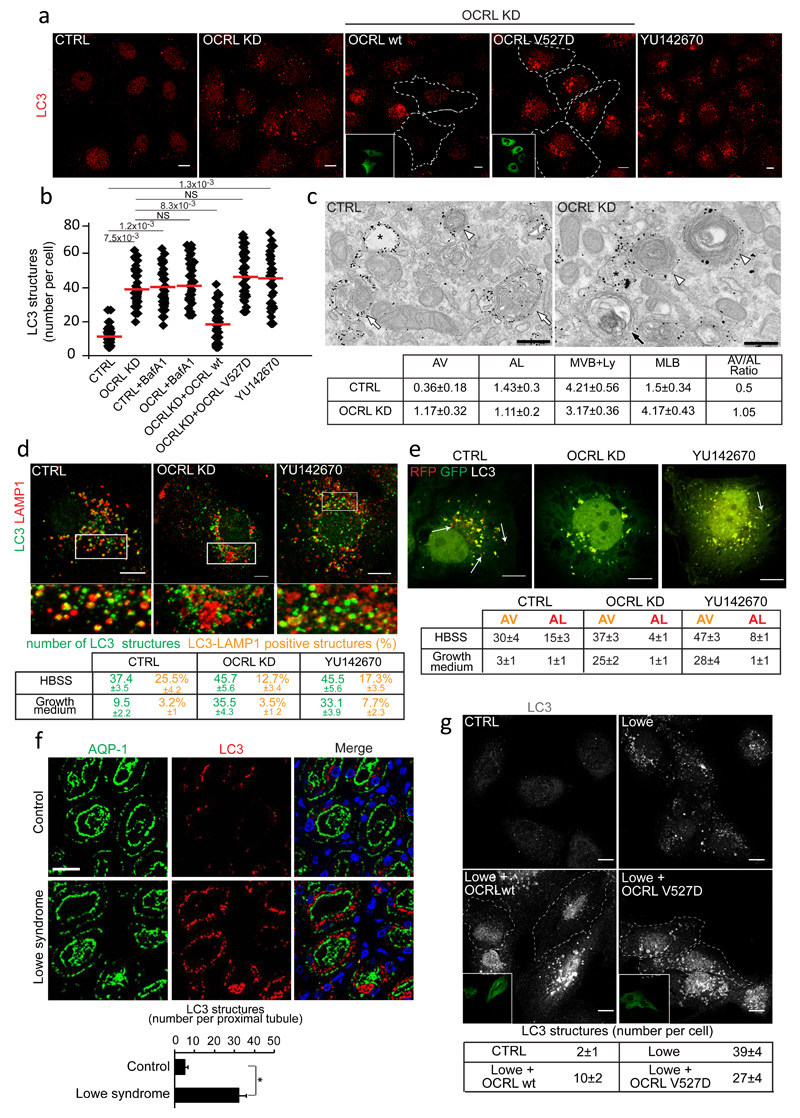
Figure 6. Autophagosomes accumulate in OCRL-depleted cells and PTCs and kidneys from Lowe syndrome patients.
(a) LC3 in cells with or without the OCRL inhibitor YU14267043 and OCRL-KD cells with or without siRNA-resistant wild-type or catalytically inactive (V527D) OCRL. Insets, transfected cells. (b) Quantification of LC3-structures in cells described in a. Red line, mean ± s.d. n=200 cells per condition pooled from 4 independent experiments. NS (not significant). (c) Ultrastructure and quantification of AV (black arrow), AL (white arrows), MVB+Ly (white arrowheads), MLB in Control and OCRL-KD cells labelled for LAMP1. p≤0.05 for AV and MLB in OCRL-KD vs. CTRL cells. Mean ± s.e.m. n=14 (CTRL), n=18 (OCRL-KD) fields pooled from 3 independent experiments. (d) LC3 and LAMP1 in CTRL, OCRL-KD and YU142670-treated cells. Lower panels, enlargements of the boxed areas. Means ± s.d. of LC3 and LAMP1 colocalization (mean ±s.d). n=120 cells per condition pooled from 3 independent experiments. p<0.001 for LC3 structures in OCRL-KD and YU142670-treated vs. fed CTRL and for LC3-LAMP1 colocalization in starved CTRL vs fed CTRL. p<0.05 for LC3-LAMP1 colocalization in starved CTRL vs. starved OCRL-KD. (e) mRFP-GFP-LC3 in control, OCRL-KD and YU142670-treated cells. Arrows, AL. Quantification of AV and AL. Mean ± s.d. n=150 cells per condition pooled from 3 independent experiments. p<0.001 for AV and AL in OCRL-KD and YU142670-treated cells vs. CTRL cells. (f) Autophagosomes accumulate in aquaporin (AQP1)-positive proximal tubules. DAPI (blue), nuclei. Means ± s.e.m., n=50 proximal tubules pooled from 10 fields per biopsy from three Lowe syndrome patients and four control subjects. (g) LC3 in PTCs from healthy subjects or Lowe Syndrome patients transfected or not with wt-OCRL or V527D-OCRL. Insets, transfected cells outlined in the main image. Mean ± s.d. of LC3-positive structures per cell, n=150 cells per condition pooled from 3 independent experiments. p<0.001 for Lowe vs. CTRL and for Lowe+OCRL wt vs. Lowe. p<0.01 for Lowe+OCRL V527D vs. Lowe. p values in b, d-g calculated by One-way ANOVA with Tukey's post-hoc test, and by Student's t-test in c. Statistic source data in Supplementary Table 2. Scale bars, 10μm in a, d, g: 250nm in c: 50μm in f.
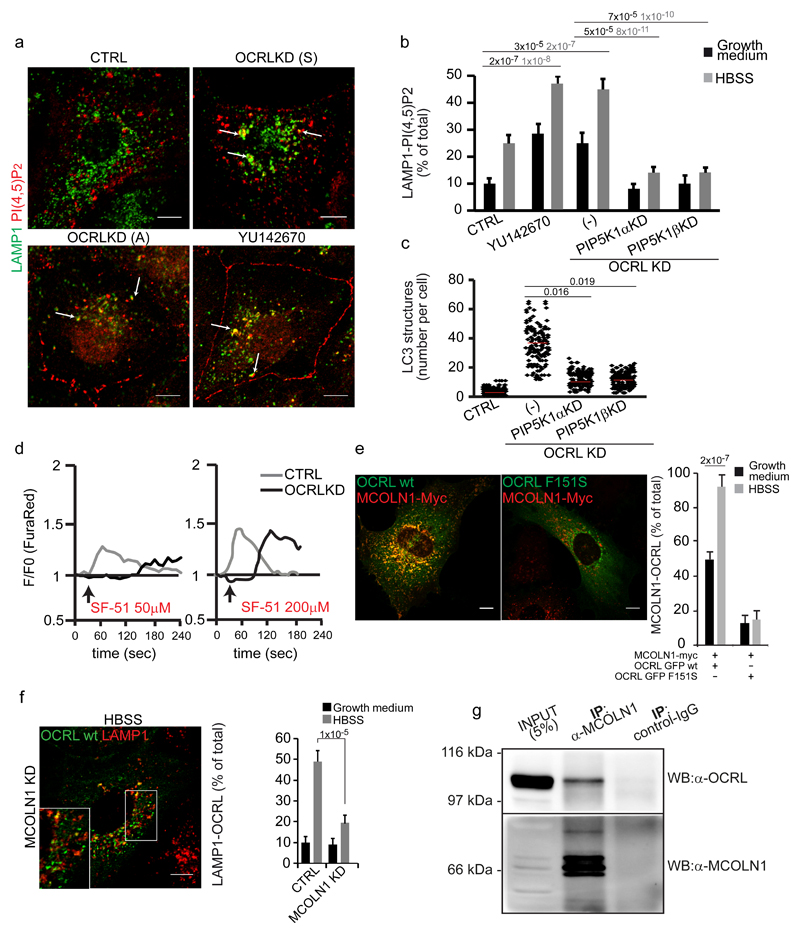
Figure 7. Autophagy flux is impaired due to PI(4,5)P2-mediated inhibition of MCOLN1 activity in OCRL-depleted cells.
(a) Mock (CTRL), YU142670-treated and OCRL-KD (Sigma (S) or Ambion (A) siRNA pools) cells were incubated in HBSS for 3 hours and labeled with anti-LAMP1 (green) and anti-PI(4,5)P2 (red) antibodies. White arrows indicate structures positive for both markers. (b) Quantification of PI(4,5)P2-LAMP1 colocalizing structures in control, YU142670-treated, OCRL-KD (-), OCRL-KD plus PIP5K1ß-KD, and OCRL-KD plus PIP5K1α-KD cells. Mean values ± s.d. n=200 cells per condition pooled from 4 independent experiments. (c) Quantification of LC3-positive structures in the cells described in b. Red line, mean values. (d) OCRL depletion impairs MCOLN1-dependent calcium release. Control or OCRL-KD cells loaded with the Ca2+-sensitive dye FuraRed were treated with 50 or 200 µM of the MCOLN1 agonist SF-5147. The ratio of fluorescence excited at 457 and 488 nm was measured and normalized against F0 (time-point before drug addition, black arrow). Plots correspond to the average of the response of n=20 cells from a single experiment, that was independently repeated 3 times. (e) Representative images and quantification of MCOLN1 and OCRL colocalization in cells transfected with wild type OCRL or the AP2-defective binding mutant (OCRL-F151S) along with MCOLN1-Myc. Right, colocalization of OCRL with MCOLN1. Mean values ± s.d. n=100 cells per condition pooled from 3 independent experiments. (f) MCOLN1-KD cells transfected with GFP-OCRL were treated with HBSS for 3 hours and stained for LAMP1 (red). Right, quantification of OCRL colocalization with LAMP1. Mean values ± s.d. n=100 cells per condition pooled from 3 independent experiments. (g) Cell lysates from HK-2 cells were immunoprecipitated using an anti-MCOLN1 antibody and analyzed by Western blot (WB) with anti-MCOLN1 or anti-OCRL antibodies. The results are representative of five independent experiments. An unprocessed scan of the blot is shown in Supplementary Fig. 9. Parallel samples were analyzed by LC-MS/MS and peptides significantly matching human OCRL (Q01968-2) were found in anti-MCOLN1 immunoprecipitates but not in control-IgG immunoprecipitates. p-values calculated by One-way ANOVA with Tukey's post hoc test. Statistic source data can be found in Supplementary Table 2. Scale bars, 10 μm.
OCRL is required to preserve the activity of MCOLN1
We searched for the target(s) of the inhibitory effect of PI(4,5)P2 on autophagosome-lysosome fusion. One interesting candidate was mucolipin-1 (MCOLN1, also referred as TRPML1), a lysosomal calcium channel required for lysosome fusion with autophagosomes whose activity is stimulated by PI(3,5)P2 but inhibited by PI(4,5)P244–46. Indeed, we found that MCOLN1-dependent calcium release was compromised – i.e. no response at low concentrations and slower response at high concentrations of MCOLN1 agonist – in OCRL-KD cells47 as compared to control cells (Fig. 7d). We also obtained independent lines of evidence indicating that MCOLN1 and OCRL interact: firstly, the overexpression of MCOLN1 recruits wt but not the AP2-defective binding mutant of OCRL to lysosomes (Fig. 7e); secondly, MCOLN1 is required for OCRL translocation to lysosomes (Fig. 7f); thirdly, the two proteins can be co-immunoprecipitated (Fig. 7g). Thus, our results indicate that both MCOLN1 (Fig. 7f) and AP2 (Fig. 2c) are required for the lysosomal recruitment of OCRL, but that neither of them on its own is sufficient since AP2-defective OCRL mutants are no longer recruited to lysosomes even in cells overexpressing MCOLN1 (Fig. 7e) and AP2-competent OCRL cannot be recruited to lysosomes in the absence of MCOLN1 (Fig. 7f). This combined requirement indicates that a mechanism of coincidence detection drives and calibrates in time and space the recruitment of OCRL to lysosomes ensuring that OCRL is recruited to lysosomal regions/domains containing both MCOLN1 and AP2.
Boosting MCOLN1 activity rescues the autophagic flux in Lowe syndrome patient PTCs
Since OCRL depletion did not have an impact on MCOLN1 localization (Supplementary Fig. 7a), we concluded that the MCOLN1 dysfunction in OCRL-depleted cells was due to reduced activity caused by an excess of PI(4,5)P2. Hence, we reasoned that boosting the activity of MCOLN1 in OCRL-depleted cells might rescue lysosomal function. To this end we used two independent approaches: the overexpression of MCOLN1 (the wt form and MCOLN1-3A, a mutant MCOLN1 unable to bind PI(4,5)P244) or the use of a MCOLN1 activator at a concentration that is effective in inducing lysosomal calcium release in OCRL-depleted cells (Fig. 7d). We observed that both the overexpression of MCOLN1, more potently in its PI(4,5)P2-insensitive form (MCOLN1-3A), and treatment with the MCOLN1 activator SF-51 (which we further characterized for its specificity, Supplementary Fig. 7b) rescued the defects in autophagic flux in OCRL-depleted cells (Fig. 8a, b). Importantly, the MCOLN1 activator also rescued the impaired autophagic flux in PTCs from Lowe syndrome patients (Fig. 8c, d), thus indicating MCOLN1 as a possible drug target for the treatment of Lowe syndrome.
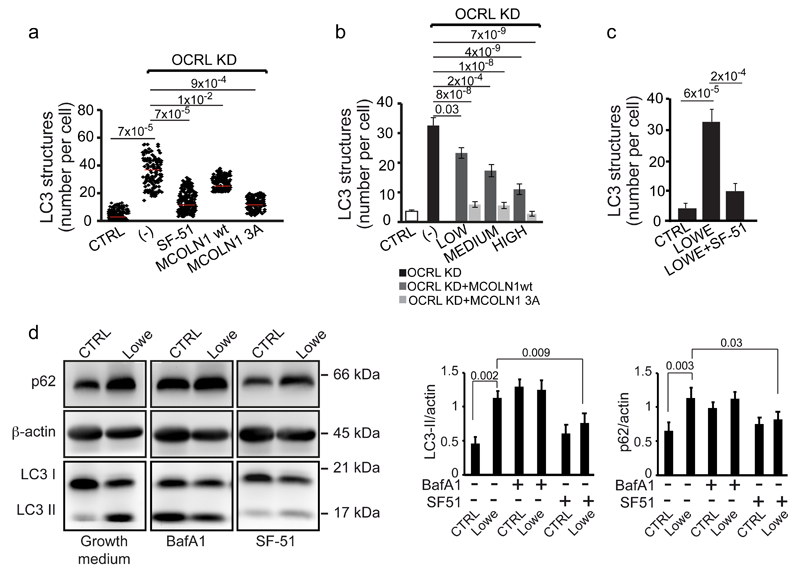
Figure 8. Autophagy flux is rescued by boosting MCOLN1 activity in OCRL-depleted cells and in Lowe syndrome patient PTCs.
(a) Quantification of LC3-positive structures in control, OCRL KD cells (-), OCRL-KD cells incubated in serum-free medium with 200 μM of the MCOLN1 agonist SF-51, or transfected with wt or a mutant form (MCOLN1-3A) of MCOLN1 that is PI(4,5)P2-insensitive44. Red line indicates mean values ± s.d. n=200 cells per condition pooled from 4 independent experiments. (b) Quantification of LC3-positive structures in control, OCRL KD (-), or OCRL-KD cells expressing different levels (low, medium and high) of wt MCOLN1 or MCOLN1-3A. Values are means ± s.d., n=75 cells per condition pooled from 5 independent experiments. (c) Quantification of the number of LC3-positive structures in CTRL PTCs, Lowe Syndrome PTCs, and Lowe Syndrome PTCs treated with 200 μM of the MCOLN1 agonist SF-51 for 2 hours. Means ± s.d. n=102 (CTRL), n=84 (Lowe), n=106 (Lowe + SF-51) cells pooled from 3 independent experiments. p-values calculated by One-way ANOVA with Tukey's post hoc test. Statistic source data can be found in Supplementary Table 2. (d) Cell lysates (50 μg/sample) from CTRL and Lowe syndrome PTCs incubated in growth medium or treated with 100 nM Bafilomycin A1 (Baf A1) or with 200 μM of the MCOLN1 agonist SF-51 for 2 hours were analyzed by SDS-PAGE and immunodetected with the indicated antibodies. β-actin was used as a loading control. A representative blot is shown and an unprocessed scan of the blot is shown in Supplementary Fig. 9. The data in both graphs on the right are mean values ± s.d., n=6 lysates per condition pooled from 6 independent experiments. P-values calculated by Student's t-test. Statistic source data can be found in Supplementary Table 2.
Discussion
Here we have unveiled a heretofore unknown ability of lysosomes to sense the arrival of autophagic cargo and at the same time a pathophysiological pathway in Lowe syndrome (Supplementary Fig. 8). The mtDNA released by autophagosomes into lysosomes stimulates TLR9 which activates its known downstream effectors, such as the transcription factor NF-kB (Supplementary Fig. 8), but also induces a local increase in PI(4,5)P2. The observation that OCRL, a 5-phosphatase, is recruited almost simultaneously with PIP5Ks demonstrates the need for a strict spatial and temporal restriction of PI(4,5)P2 levels in lysosomes. The local increase in PI(4,5)P2 is needed for the recruitment of AP2 and clathrin. We envisage that AP2-mediated clathrin recruitment controls distinct budding events in lysosomes in distinct stages of autophagy. During ongoing autophagy (this report) it may mediate the recycling of components, such as syntaxin 17, needed to sustain an efficient autophagic flux, while at the end of autophagy, as shown by Rong et al.19, it mediates autophagic lysosome reformation to regenerate primary lysosomes that have been consumed through their fusion with autophagosomes. PI(4,5)P2, which is required for these budding events, has to be nevertheless restricted to limited domains in the lysosome since it can act as an endogenous inhibitor of the calcium channel MCOLN144, 48. This is the task of OCRL, which interacts with MCOLN1 and which ensures PI(4,5)P2-free microdomains around MCOLN1 (Supplementary Fig. 8) . Indeed, the uncontrolled increase in lysosomal PI(4,5)P2 that results from the loss of function of OCRL inhibits MCOLN1 activity and, consequently, the autophagic flux, inducing the accumulation of autophagosomes that is observed in cells and, notably, in the kidney of Lowe syndrome patients. The lysosome dysfunction and the impaired autophagic flux are bound to play a pivotal role in the progression of renal damage, a so far poorly understood process, and, possibly, also in the generation of central nervous system-related symptoms and of neuropathological signs in Lowe syndrome patients6, 7, 49. Finally, and most importantly, MCOLN1 emerges as an interesting drug target since MCOLN1 agonists are able to restore the autophagosomal flux in cells from Lowe syndrome patients.
Methods
Antibodies, cDNAs, and reagents
All chemical reagents were of analytical grade or higher and purchased from Sigma-Aldrich unless otherwise specified. The anti-OCRL, -PI4KIIIβ and -GST polyclonal antibodies have been described in5. The other primary and secondary antibodies used in this study are listed in Supplementary Table 3. Other reagents: Quant-iT™PicoGreen®, FuraRed, Hanks’ Balanced Salt Solution with Calcium and Magnesium (HBSS) 1X, OPTI-MEM (Life Technologies); SF-51, YU14267043 (STK001646 and STK883823, respectively) (Vitas-M Laboratory); agarose-linked anti-GFP antibody (Vector Laboratories); Bafilomycin, vinblastine, FCCP, ethidium bromide, uridine, Protein A/G Sepharose (Sigma-Aldrich). Sodium pyruvate (Euroclone); ODN 2216, ODN 2395, ODN TTAGGG (InvivoGen); Torin-1 (Tocris Bioscience); Tat-beclin 1 peptide, digitonin (Merck-Millipore). The GST-tagged PH domain of PLCδ was prepared as described in Vicinanza et al.5. Tandem fluorescent mRFP-eGFP-LC3 was kindly provided by A. Fraldi (TIGEM, Pozzuoli, Italy). TLR9-YFP (plasmid #13642) and TIRAP-GFP (plasmid #52739) (Addgene). The full-length WT and mutant form (V527D) of OCRL were described previously5.
Mutations in the clathrin-binding domains (LIDIA, I74N and LIDLE, deltaLIDLE) and in the AP-2 binding motif (FxDxF, F151S) of OCRL were generated using the QuikChange mutagenesis system (Agilent Technologies) following the manufacturer’s protocol using the primers listed in Supplementary Table 4. The GFP-OCRL-X316 triple mutant was generated by sequential mutagenesis of LIDIA, LIDLE and FxDxF domains.
Full-length INPP5B (NM_005540) was obtained from HeLa cell cDNA using the primers reported in Supplementary Table 4 and cloned using the restriction sites SalI/SmaI into the pEGFP-C1 expression vector (Clontech).
Myc-DDK-MCOLN1 (NM_020533) was purchased from Origene Technologies. The MCOLN1[R42A;R43A;R44A] mutant (reported in the main text and figures as MCOLN1-3A) was generated using the primers reported in Supplementary Table 4.
Cell culture, transfection and treatments
HK-2 cells were grown as previously described5. HK-2 cells were transiently transfected using TransIT-LT1 (Mirus Bio LLC), unless otherwise specified, according to the manufacturer’s instructions, and incubated for 18-24 hours before fixation. The HK-2 cell line was checked for mycoplasma contamination by a PCR-based method. The HK-2 cell line was not found in the database of cell lines that are currently known to be cross-contaminated or misidentified by ICLAC and NCBI Biosample. The cell lines used in this study were not authenticated. All cell-based experiments were repeated at least three times.
Proximal tubular cells from Lowe syndrome patients
The procedure for Proximal Tubular Cell (PTC) isolation and immortalization from the urine of control healthy subjects and from Lowe syndrome patients is reported in5. PTCs were transfected using JetPEI (Polyplus transfection) according to the manufacturer’s instructions.
siRNA treatment
siRNA sequences used in this study are listed in Supplementary Table 1. siRNAs against TFEB, described in8, were kindly provided by C. Settembre (TIGEM, Pozzuoli, Italy).
HK-2 cells were transfected with siRNAs for 96 hours using DharmaFECT 4 (Dharmacon) according to the manufacturer’s instructions. The siRNA duplexes were used at 50 pmol for OCRL, TLRs, TIRAP, IRAK4, MyD88, DNaseII, UNC93B1 and TFAM and at 25 pmol for PIP5Ks, MCOLN1, VPS16, STX17, RAB7 and TFEB. Mock-treated or non-targeting siRNA-treated HK-2 cells are referred to as controls (CTRL). OCRL KD was performed using a pool of two OCRL siRNAs from Sigma (N1, N5, used for all experiments unless otherwise stated) and a pool of three OCRL siRNAs from Ambion, described in the text as OCRL KD (A).
Quantitative real-time PCR
Real-time quantitative PCR (qRT-PCR) was carried out with the LightCycler 480 SYBR Green I mix (Roche) using the Light Cycler 480 II detection system (Roche) with the following conditions: 95ºC, 10 min; (95ºC, 10 s; 60ºC, 10 s; 72ºC, 15 s) x 45 cycles. For expression studies the qRT-PCR results were normalized against an internal control (β-actin and HPRT1).
For mitochondrial gene expression analysis and for the evaluation of mtDNA depletion, qRT-PCR was performed as previously described51. The sequences of primers used in this study are listed in Supplementary Table 4.
Microarray Experiments
The Affymetrix Gene-Chip (HG-U113A) hybridization experiments were performed in triplicate at the Coriell Genotyping and Microarray Center, Coriell Institute for Medical Research, Camden, NJ, USA, on total RNA extracted from control and OCRL-depleted cells. To identify downstream transcriptional effects of OCRL-siRNA treatment, microarray data were pre-processed using the Bioconductor package Affy52 and normalized with the RMA method53. Differentially expressed genes between conditions (KD vs. control) were identified using a Bayesian T-test54. For each p-value, the Benjamini-Hochberg procedure was used to calculate the False Discovery Rate (FDR) to avoid the problem of multiple testing.
Gene Set Enrichment Analysis was performed using the freely available software GSEA v2.0 from the Broad Institute55. The goal of GSEA is to determine whether members of a given gene set tend to occur toward the top (or bottom) of a ranked gene list. In order to run the GSEA algorithm, RMA-normalized microarray data were used. The GSEA algorithm collapsed the probe sets into gene symbols (~13,300 genes) and ranked the genes in the OCRL-KD expression dataset based on the fold change after OCRL KD. GSEA analysis for subcellular distribution was conducted using Human Protein Atlas for all but lysosomal genes which were defined according to Lysoplex50 and excluded from the other categories. To evaluate the significance of the enrichment score the hypergeometric distribution was calculated that describes the probability of k successes in n draws (up- and down-regulated genes for each enriched category).
CLEAR analysis
We selected the genes corresponding to the top 250 upregulated probes in the OCRL-KD expression profile. The proximal promoter sequences (1000 bp upstream and 200 downstream of the transcriptional start site) for each gene were obtained from the UCSC genome browser55 and searched with the matchPWM function in the Bioconductor package Biostrings56 for matches with the CLEAR motif57.
Mitochondrial DNA (mtDNA) depletion and visualization
Mitochondrial DNA depletion was achieved by siRNA knock down of TFAM or by incubating HK-2 cells in complete medium with Ethidium Bromide (100ng/mL), uridine (50mg/mL) and sodium pyruvate (110mg/mL) for 4 days to reduce mtDNA levels. Evaluation of depletion was performed using Quant-iT™PicoGreen® and qRT-PCR as previously described51, 58.
Immunoprecipitation and Western blot
HK-2 cells were plated in 150 mm plates until they reached 95% confluency. Cells were then washed 3 times with ice-cold 1X PBS, scraped, and proteins were extracted in ice-cold lysis buffer (25 mM TRIS-HCl, pH 7.4, 100 mM NaCl, 5 mM EDTA and 1% NP-40 supplemented with phosphatase and protease inhibitor cocktails, Roche), centrifuged for 15 min at 16,000 x g at 4°C, and the supernatants were collected. Cell supernatants (1 mg) were incubated for 16 hours at 4°C with 1 μg of the appropriate antibody or with 15 μL of agarose-linked anti-GFP antibody (for TLR9-YFP immunoprecipitation). Samples were then incubated with Protein A/G Sepharose for 2 hours at 4°C (except for TLR9-YFP immunoprecipitation). Immunoprecipitates were collected by centrifugation at 5,000 x g at 4°C, washed five times in lysis buffer and twice in lysis buffer without NP-40 and the proteins were eluted with Laemmli sample buffer (60 mM TRIS-HCl pH 6.8, 2% SDS, 10% glycerol, 5% β-mercaptoethanol, 0.01% bromophenol blue) and denatured at 95°C for 5 min. Samples were analysed by SDS-PAGE (8%) under reducing conditions and transferred to nitrocellulose. The membranes were then incubated with the appropriate antibodies. Enhanced chemiluminescence reagent (Euroclone) was used for protein detection. Western blot quantitative analysis (p62/β-actin and LC3/β-actin ratio and p-S6K/S6K ratio) was performed using ImageJ band analysis59.
Confocal fluorescence microscopy, super-resolution microscopy, image processing, and colocalization analysis
HK-2 cells and PTCs were grown to subconfluence on glass coverslips and immunofluorescence microscopy and quantitative image analysis were performed as described5, except for OCRL-LAMP1 co-labelling where a different permeabilization step was used [i.e. 5 min with 20 µM digitonin in Buffer A (20 mM PIPES pH 6.8, 137 mM NaCl, 2.7 mM KCl)]. The experiments were repeated at least three times and representative images are shown. The level of colocalization (i.e. LAMP1-OCRL, LAMP1-TLR9, LAMP1-TIRAP, LAMP1-LC3, LAMP1-AP2, LAMP1-Clathrin, LAMP1-TLR9-YFP, LAMP1-TIRAP, LAMP1-PI4KIIIβ, LAMP1-PIP5K1α, LAMP1-PIP5K1β, LAMP1-PI(4,5)P2, MCOLN1-OCRL, MCOLN1-LAMP1) was calculated by acquiring confocal serial sections from about 50-100 cells per experimental condition per experiment, exported in TIFF format, and processed as previously described5. Briefly confocal images of at least 50-100 cells per condition were acquired at the same laser power and photomultiplier gain. Images were then processed using Image J software. Single channels from each image were converted into 8-bit grayscale images and thresholded in order to subtract background. The Image J “Analyze Particles” plugin was then used to identify and count the total number of the structures (with an area above 0.10 μm2) in channel 1 (i.e. LAMP1). The structures in channel 2 (i.e. OCRL) were used to build a mask that was then overlapped with the LAMP1 structures to subtract the structures containing both markers. The remaining structures, positive only for LAMP1, were counted and by difference the number of structures containing both LAMP1 and OCRL was calculated and expressed as % of the total number of LAMP1-positive structures. Note that this quantitative analysis procedure does not use merged images and is not affected by the fluorescence intensity.
For conventional confocal microscopy a confocal laser microscope (Zeiss LSM 700 confocal microscope systems; Carl Zeiss, Gottingen, Germany) with a 63 x 1.4 NA oil immersion objective was used. Super-resolution microscopy was performed with a Zeiss LSM 880 (Carl Zeiss, Gottingen, Germany), Airyscan-equipped confocal microscope with a 63X 1.4 NA oil immersion objective. After acquisition images were processed with the Airyscan processing tool available on the Zen software provided by Zeiss, with a processing threshold set at 3.9.
PI(4,5)P2 detection
Control (mock) and OCRL siRNA-treated HK-2 cells were grown to 70-80% density on glass coverslips and PI(4,5)P2 staining was performed according to60. Briefly cells were fixed for 15 min at room temperature by addition of 1 volume of pre-warmed 4% PFA (paraformaldehyde in PBS) to the growth medium (2% PFA final concentration). Cells were then permeabilized for 5 min with 20 µM digitonin in Buffer A. Cells were then blocked for 45-60 min with buffer A containing 5% FBS (Fetal Bovine Serum) and 50 mM NH4Cl. Recombinant GST-PH-PLCδ (0.2 mg/ml) or anti-PI(4,5)P2 antibody were added at this stage in buffer A for 90 min at room temperature. After three washes with buffer A, anti-GST (in the case of GST-PH-PLCδ) and anti-LAMP1 antibodies were applied for 1h at rt. After three washes secondary fluorophore-labelled antibodies were added. Cells were post-fixed for 5 min with 2% PFA, washed twice in 1X PBS containing 50 mM NH4Cl, washed once in milliQ water and analysed using a confocal laser microscope (Zeiss LSM 700 confocal microscope systems; Carl Zeiss, Gottingen, Germany) with a 63 x 1.4 NA oil immersion objective. Quantitative analysis of PI(4,5)P2-LAMP1 colocalization was performed as previously reported5.
Autophagy induction by Torin 1 and Tat-beclin 1 peptide
HK-2 cells were treated for 2 hours with 250 nM of the specific mTOR inhibitor Torin 1 (Tocris bioscience) added to complete growth medium. HK-2 cells were washed twice with PBS and once with Optimem and then incubated for two hours with 20 μM of the Tat-beclin 1 peptide22 dissolved in acidified OPTI-MEM (9 mM HCl).
Quantitative analysis of LC3 structures
CellProfiler™, a free, open-source image analysis software (www.cellprofiler.org), was used for quantitative analysis of confocal microscopy images by applying specific pipelines described in61. The pipeline “Speckle Counting” was used to identify smaller objects (LC3-positive structures) surrounding larger objects (nuclei) and to perform per-object aggregate measurements (such as the number of LC3-positive structures per nucleus). Briefly, the images were converted in grayscale using the module “ColorToGray”. Then the “ObjectIdentification” module was used to identify nuclei (called primary objects) based on Hoechst staining. Secondary objects (LC3-positive structures) were detected and counted using the module “IdentifySecondaryObjects” by expanding the area surrounding the primary object by 20 pixels.
Human kidney biopsies
Human kidney biopsies were obtained from archived samples of three patients with clinical diagnosis of Lowe syndrome, corroborated in one case (shown in Fig. 6f) with the genetic diagnosis of mutation in OCRL, two patients affected by glomerular (and not tubular) disease, and two controls (non-transplanted, normal human kidney). Informed consent was obtained and the use of the human biopsy samples was approved by the Ethical Review Board of Saint-Luc Academic Hospital (Brussels, Belgium) and the EURenOmics consortium (FP7, 2007-2013, ga# 305608).
Immunofluorescence on human kidney samples
Paraffin blocks of human kidney samples were sectioned into consecutive slices with a thickness of 6 μm using a Leica RM2255 rotary microtome (Thermo-Fisher Scientific) on Superfrost Plus glass slides (12-550-15, Thermo-Fisher Scientific). Before staining, slides were deparaffinized in changes of CitriSolv (22-143-975, Thermo-Fisher Scientific,) and 70% isopropanol. Antigen retrieval was accomplished by incubating in sodium citrate buffer (1.8% 0.1M citric acid, 8.2% 0.1M sodium citrate, in distilled water, pH 6.0) in a rice cooker for 30 minutes. The slides were blocked with PBS blocking buffer (1% BSA, 0.2% non-fat dry milk in PBS) for 30 min and stained with primary antibodies specific for LC3B (Nanotools) or LAMP1 (Abcam) and AQP1 (Merck-Millipore) diluted in blocking buffer overnight at 4°C. The anti-LC3B antibody recognizes both the soluble (LC3-I) and the membrane-bound form (LC3-II) of LC3B62. After two washes in 0.1% Tween 20 (v/v in PBS), the slides were incubated with the corresponding fluorophore-conjugated secondary antibodies (Life Technologies) diluted in blocking buffer at room temperature for 1 hour and counterstained with 10 μM Hoechst 33342 (Life Technologies,). The slides were subsequently mounted in Prolong Gold Anti-fade reagent (Life Technologies) and images were acquired on a Leica SP5 confocal laser scanning microscope (Center for Microscopy and Image Analysis, University of Zurich) equipped with a Leica APO 63x NA 1.4 oil immersion objective and then processed with Adobe Photoshop (Adobe System) software.
Quantitative image analysis was performed by random selection of ~10 visual fields that included 5-6 proximal tubules that were scanned for each slide, using the same setting parameters (i.e. pinhole, laser power, and offset gain and detector amplification below pixel saturation). The number of LC3- or LAMP1-positive structures per proximal tubule (AQP1-positive) was quantified by means of Quantitation Module of Volocity software (PerkinElmer Life Science, I40250).
Zebrafish strains and husbandry
Zebrafish were maintained at The University of Manchester Biological Services Unit in accordance with the policies of UK Animal Act 1986. Wild Type adults and larvae were of the AB strain. The ocrl1-/- mutant line (ZDB-GENO-120531-1) was previously described9, 10.
The only procedure involving animals was the breeding of transgenics, which is subject to local ethical review and performed under a Home Office license. No statistical method was used to predetermine sample size; the experiments were not randomized, and the Investigators were not blinded to allocation during experiments and outcome assessment. Experiments were performed on larvae at 3 days post-fertilisation, at which point they are not classified as animal experiments.
Electron Microscopy
Cells for pre-embedding immunoelectron microscopy were fixed, permeabilized, and labelled as described previously63. From each sample, thin 65 nm sections were cut using a Leica EM UC7 ultramicrotome. EM images were acquired from thin sections using a FEI Tecnai-12 electron microscope (FEI) equipped with a VELETTA CCD digital camera (Soft Imaging Systems GmbH, Munster, Germany). We used the following criteria for morphological identification: Lysosomes (Ly): LAMP1-positive single-membrane-bound bodies containing electron dense material; Multivesicular bodies (MVB): LAMP1-positive single-membrane-bound bodies containing more than 9 intralumenal vesicles; Multilamellar bodies (MLB): LAMP1-positive single-membrane-bound bodies containing myelin-like figures; Autophagic vacuoles (AV): double-membrane-bound vacuoles containing sequestered material (only cytoplasm, cytoplasm and/or ER, cytoplasm and/or mitochondria, ER and/or mitochondria); Autolysosomes (AL): LAMP1-positive single-membrane-bound bodies containing electron dense cytoplasmic material and/or organelles. The number of autophagosomes was then calculated using iTEM software (Olympus SYS, Germany). Zebrafish samples were processed for EM as described9.
Dissection of autophagosome-lysosome fusion by tandem fluorescent-LC3
Control (mock), OCRL- and TLR9- siRNA-treated HK-2 cells were grown to 70% confluency and transiently transfected with a plasmid encoding an mRFP-GFP-tagged LC339. After transfection, cells were kept in growth medium for 16 hours and then incubation was continued in normal growth medium or the cells were treated with HBSS for 3 hours. CTRL cells were also treated for 3 hours with 25 μM of OCRL inhibitor (YU142670) or 0.5 μM of TLR9 antagonist (ODN TTAGGG). Where reported, cells were also treated with protease inhibitors cocktail39 (PI: Leupeptin, E64D and Pepstatin, Sigma-Aldrich) for the indicated time points. After fixation with 4% PFA, cells were washed three times with 1X PBS containing 50 mM NH4Cl, mounted and imaged using a Zeiss LSM 710 confocal microscope (equipped with a 63 X 1.4 NA oil immersion objective).
Calcium Measurements by Confocal Imaging using FuraRed
FuraRed-loaded cells were analysed according to the previously described protocol64. Briefly, mock HK-2 cells or OCRL-silenced HK-2 cells were loaded with 5 μM FuraRed-AM for 1 hour at 37°C in calcium imaging buffer. Pseudocolor ratiometric images (458 and 488 nm excitation, 610 nm emission) were collected using a Carl Zeiss microscope (LSM 710) equipped with a variable filter wheel. FuraRed ratios (emission at F458/F488) were used to represent changes in cytosolic Ca2+ levels in basal conditions or after specific treatments. The number of cells that mobilized calcium was quantified in a minimum of three wells and compared with control cells transfected with scramble siRNAs. More than 20 cells in triplicate were used in the different experiments.
Statistics and Reproducibility
For experiments involving living animals no statistical method was used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Immunofluorescence experiments were repeated independently three times and at least 50 cells were analysed from a single experiment. Representative images are shown in Fig.1c, 1d, 1e, 2a, 2c, 3a, 4b, 4d, 4g, 5a, 5d, 5e, 6a, 6d, 6e, 6f, 6g, 7a, 7e, 7f; and in Supplementary Fig. 2b, 3b, 3d, 4b, 4d, 4e, 4g, 5a, 5c, 5e, 5f, 6a, 6b and 7a.
Western blotting and immunoprecipitation experiments were repeated at least three times and representative blots are shown in Fig. 4i, 5c, 7g and in Supplementary Fig 1a, 3a, 5b and 7c. Uncropped scans of the blots are shown in Supplementary Fig. 8.
Most data are presented as the means ± standard deviation (s.d.) unless otherwise specified. Statistical comparisons were made using Student's t-test when comparing two groups. One-way ANOVA with Tukey’s post-hoc tests were performed for comparisons among more than two groups relative to a single factor (treatment or siRNA-induced knock down). Two-way ANOVA and Tukey’s post-hoc tests were performed for comparisons among more than two groups relative to two factors (treatment and siRNA-induced knock down). These analyses were performed in R65 [http://www.R-project.org/] environment. For all analyses a p-value < 0.05 was considered to be statistically significant. For the quantification of LC3- and LAMP1-positive structures in kidney biopsies, the analysis of variance (ANOVA) was also used. A P value < 0.05 was considered to be statistically significant. The results of the statistical analysis, comprising the p values of the whole experimental data sets and the p values deriving from post hoc tests are reported in Supplementary Table 2.
Data availability
Microarray data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE54284. Source data for Fig. 2b, 2c, 3a, 3c, 4a, 4c, 4e, 4f, 4h, 5b, 5c, 5d, 5e, 5f, 5g, 6b, 6c, 6d, 6e, 6g, 7b, 7c, 7e, 7f, 8b, 8c and 8d and for Supplementary Fig. 2b, 3a, 3b, 3e, 4e, 4f, 4g, 5a, 5f, 7b are available in Supplementary Table 2: Statistic source data. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Supplementary legends
Supplementary Figure 1
Effects of siRNA treatment on protein and mRNA levels.
(a) HK-2 cells were treated with non-targeting siRNA (CTRL) or with Sigma Aldrich (S) or Ambion (A) OCRL siRNA pools for 96 hours. Cell lysates (50 μg/sample) were analyzed by SDS-PAGE and immunoblotted using an anti-OCRL antibody or an anti-β-actin antibody as a loading control. A representative blot is shown. An unprocessed scan of the blot is shown in Supplementary Fig. 9. The graph reports the results of the quantitative analysis of OCRL levels from n=10 independent experiments for CTRL (mock) and OCRLKD (S) and n=5 independent experiments for CTRL (non-targeting) and OCRLKD (A). The data are expressed as the level of OCRL normalized for β-actin as a % of the CTRL. Mean values ± s.d. (b) Representative images of CTRL (non-targeting siRNA) and OCRL-KD HK-2 cells stained for OCRL, TGN46 (a Trans Golgi Network marker) and EEA1 (an Early Endosome marker). Insets represent OCRL-TGN46 and OCRL-EEA1 merged enlargements of the boxed areas. Scale bars, 10 μm. (c) HK-2 cells were treated with the indicated siRNAs (Listed in Supplementary Table 1). Total RNA (1 μg/sample) was analyzed by qRT–PCR using specific primers (Listed in Supplementary Table 4). Expression levels in siRNA-treated cells are reported relative to the expression of the corresponding gene in mock-treated cells (CTRL) normalized to β-actin. All qPCR reactions were carried out in triplicate and the data represent mean values ± s.d. from n=5 independent experiments. Statistical significance calculated by Student's t-test.
Supplementary Figure 2
OCRL depletion induces upregulation of lysosomal genes and TFEB nuclear translocation. (a) Fold change expression of the indicated lysosomal genes (selected amongst those most upregulated in the microarray) in OCRL-KD (Ambion siRNAs pool) cells relative to non-targeting siRNA-treated cells (CTRL) as assessed by quantitative RT-PCR carried out in triplicate and the data represent mean values ± s.d. from n=6 independent experiments. p values calculated using Student’s t test (unpaired). (b) CTRL, OCRL-KD (treated either with Sigma (S) or Ambion (A) pools) and YU142670-treated HK-2 cells were stained with an anti-TFEB antibody (green). Mean values ± s.d. n=150 cells pooled from 3 independent experiments. Numbers on the images report the percentage of cells with nuclear TFEB. p < 1.6x10-7 for CTRL vs. OCRLKD (S), OCRLKD (A) and YU142670-treated cells calculated by One-way ANOVA with Tukey's post-hoc test. Scale bars, 10 µm. Statistic source data for Supplementary Fig. 2b can be found in Supplementary Table 2. (c) The position of the TFEB binding site (CLEAR motif, blue boxes) relative to the transcriptional start site (0) in the promoters of the indicated upregulated lysosomal genes in OCRL-KD cells, assessed as described in Methods. The color intensity is proportional to the match score (the darker the higher). (d) Residual upregulation of the indicated lysosomal genes in HK-2 cells knocked down for both OCRL and TFEB relative to OCRL-KD cells as assessed by quantitative RT-PCR carried out in triplicate. The data represent mean values ± s.d. from n=6 independent experiments. p values were calculated by Student's t-test. NS, not significant. Notice the inverse correlation between the presence of a CLEAR motif and the extent of residual upregulation upon TFEB depletion.
Supplementary Figure 3
OCRL, AP2 and clathrin recruitment to lysosomes during autophagy requires the synthesis of PI(4,5)P2 and is independent of mTOR activation or inactivation. (a) Representative image of the colocalization of PI4KIIIβ with LAMP1 in HBSS. Scale bar, 10 μm. The graph reports quantification of PI4KIIIβ with LAMP1 colocalization in growth medium and HBSS. Mean values ± s.d. n=200 cells pooled from 3 independent experiments. (b) Quantification of PI(4,5)P2, AP2, clathrin and OCRL colocalization with LAMP1 during starvation (HBSS for 3 hours) after impairing PI(4,5)P2 synthesis (PI4KIIIβ-KD, PIP5K1α-KD and PIP5K1β-KD). Mean values ± s.d. n=200 cells per condition pooled from 3 independent experiments. (c) Cell lysates (50 μg) from HK-2 cells treated with HBSS and then re-fed by addition of complete growth medium for the indicated time points were analyzed by SDS-PAGE and immunoblotted using an anti-p70, an anti-phospho-p70 (p-p70), or an anti-GAPDH antibody as a loading control. A representative blot is shown and an unprocessed scan of the blot is shown in Supplementary Fig. 9. (d) Starvation-independent autophagy stimuli induce an increase in PI(4,5)P2 and recruitment of AP2, clathrin and OCRL to lysosomes. HK-2 cells were treated with the mTOR inhibitor Torin-1, with Tat-beclin 1 peptide (see Methods), or left untreated (Growth medium) and then stained for OCRL and LAMP1. The insets are enlargements of the boxed areas. Scale bars, 10 μm. (e) Quantification of PI(4,5)P2, AP2, clathrin and OCRL colocalization with LAMP1 and of the average number of autophagosomes (LC3 structures) per cell in untreated cells and in cells treated with Torin-1 and Tat-beclin 1 peptide. Mean values ± s.d. n=100 cells per condition pooled from 3 independent experiments. p values calculated by one-way ANOVA with Tukey's post-hoc test. Statistic source data for Supplementary Fig. 3 can be found in Supplementary Table 2.
Supplementary Figure 4
Inhibiting TLR9 or depleting mtDNA inhibits the lysosomal recruitment of OCRL and impairs the recycling of the autophagosomal SNARE STX17. (a) IEM of fed or starved TLR9-YFP expressing HK-2 cells and stained with an anti-GFP antibody (scale bar, 200 nm). Arrows, TLR9 in the ER (white), in Multivesicular bodies/Lysosomes (black), or in autolysosomes (red). (b) Control, TFAM-KD and EtBr treated51, 58 HK-2 cells stained with PicoGreen58 to visualize mitochondrial DNA. Scale bars, 10 μm. (c) Expression of the indicated nuclear (α-SMA) and mitochondrial (ND-6, D-loop, CoxI and CoxII) genes evaluated by qPCR in EtBr-treated cells vs. control cells. All qPCR reactions were carried out in triplicate and the data represent mean values ± s.d. from n=3 independent experiments. (d) OCRL (red) association with lysosomes upon starvation in CTRL, TFAM-KD, and HK-2 cells treated with EtBr. Scale bars, 10 μm. Insets, enlargements of the boxed areas. (e) Fed or starved HK-2 cells expressing TIRAP-GFP stained for LAMP1. Insets, enlargements of the boxed areas. Scale bars, 10 µm. Graph, colocalization of TIRAP with LAMP1. Mean ± s.d. n=100 cells per condition pooled from 3 independent experiments. (f) Fed or starved HK-2 cells KD for adaptors (MyD88 and TIRAP) or transducers (IRAK4) of TLR9 signalling and for UNC93B1. LAMP1-OCRL colocalization. Mean values ± s.d. n=300 cells per condition pooled from 3 independent experiments. (g) Representative images of control or TLR9-KD HK-2 cells expressing the STX17-GFP incubated in HBSS for 3 hours. Insets, enlargements of the boxed areas. Scale bars, 10 µm. Right graph, quantification of LAMP1-STX17 colocalization. n=75 cells per condition pooled from 3 independent experiments. The data show that STX17 is enriched in LAMP1-positive compartments in TLR9-depleted cells as compared to control cells. The LAMP1 structures containing STX17 are, by definition, autolysosomes: since the total number of autolysosomes is lower in TLR9-KD cells (Figure 5b, e), then the fraction of autolysosomes still containing STX17 is markedly increased upon TLR9 deletion, suggesting that recycling of STX17 from autolysosomes is under control of TLR9. The p values are indicated, calculated using Student's t-test in c, e, and by one-way ANOVA with Tukey's post-hoc test in f, g. Statistic source data in Supplementary Table 2.
Supplementary Figure 5
HK-2 cells depleted of OCRL accumulate mature autophagosomes following impairment of autophagosome/lysosome fusion. (a) Representative images of CTRL and OCRL-KD cells incubated in growth medium or in HBSS for 3 hours, stained for ATG16L1 (an early and transient autophagosomal marker). Scale bars, 10 µm. The graph reports the quantification of the ATG16L1-positive structures, means ± s.d. n=150 cells pooled from 3 independent experiments. (b) Immunoblot analysis of cell lysates (50 μg/sample) from CTRL, OCRL-KD (Sigma (S) or Ambion (A) siRNA pools) and YU142670-treated cells with or without 100 nM Bafilomycin A1 (Baf A1) for 2 hours with the indicated antibodies. β-actin was used as a loading control. A representative blot is shown and an unprocessed scan of the blot is shown in Supplementary Fig. 9. Mean values ± s.d. n=3 lysates per condition pooled from 3 independent experiments. (c) HK2 cells were depleted of OCRL using Ambion siRNA pool and treated as described in Figure 6d. Mean values ± s.d. n=120 cells per condition pooled from 3 independent experiments. (d) HK2 cells were depleted of OCRL using Ambion siRNA pool and treated as described in Figure 6e. The arrow indicates an autolysosome. Mean values ± s.d. n=150 cells per condition pooled from 3 independent experiments. (e) CTRL and OCRL-KD cells were incubated in HBSS for 3 hours and labeled with an anti-LAMP1 (green) antibody together with an anti-GST antibody to detect recombinant GST-PHPLCδ (a probe for PI(4,5)P2)5. White arrows indicate structures positive for both markers. Scale bars, 10 μm. (f) Representative images and quantification of PI(4,5)P2-LAMP1 colocalization in OCRL-KD cells expressing an siRNA-resistant wild-type (GFP-OCRL wt) or catalytically inactive (GFP-OCRL V527D) OCRL and stained with antibodies against PI(4,5)P2 (red) and LAMP1 (green). Transfected cells are outlined. Scale bars, 10 μm. Mean values ± s.d. n=300 cells pooled from 3 independent experiments. p values are indicated, calculated by One-way ANOVA with Tukey's post-hoc test in a, c, d, f, and by Student's t-test in b; NS (not significant). Statistic source data for Supplementary Fig. 5 can be found in Supplementary Table 2.
Supplementary Figure 6
Accumulation of autophagosomes and expansion of lysosomal compartments in kidneys of Lowe syndrome patients. (a) Representative confocal micrographs of proximal tubules from two patients with Lowe syndrome and one control immunostained with an anti-LC3 (red) and anti-AQP1 (green, a proximal tubule marker) antibody. Quantification of the LC3-positive structures is given in Figure 6f. Nuclei were counterstained with DAPI (blue). Scale bar, 50 μm. (b) Representative confocal micrographs of proximal tubules from two patients with Lowe syndrome and one control immunostained with an anti-LAMP1 (red) and anti-AQP1 (green) antibody. Nuclei were counterstained with DAPI (blue). Scale bar, 50 μm. The graph reports the quantification of LAMP1-positive structures in AQP1-positive proximal tubules in human kidney biopsies from three Lowe syndrome patients and four controls. Data are means ± s.e.m., n=50 proximal tubules pooled from 10 fields per biopsy. *p < 0.0025 calculated calculated using one-way ANOVA/post-hoc tests.
Supplementary Figure 7
The MCOLN1 agonist SF-51 specifically reverts autophagosome accumulation in OCRL-depleted cells and in PTCs from Lowe syndrome patients. (a) OCRL depletion does not affect MCOLN1 localization at lysosomes. Mock-treated (CTRL) and OCRL-KD HK-2 cells transiently expressing MCOLN1-myc were immunostained with anti-myc (green) and anti-LAMP1 (red) antibodies. Numbers indicate the percentage of LAMP1-MCOLN1 colocalization. Means ± s.d., n=150 transfected cells per condition pooled from 3 independent experiments. Scale bar, 10 μm. (b) SF-5147 does not rescue the autophagosome accumulation caused by depleting components of the autophagosome-lysosome tethering/fusion machinery. Mock-treated (CTRL), OCRL-KD, MCOLN1-KD, RAB7-KD and VPS16-KD HK-2 cells incubated in growth medium with or without 200 μM of the mucolipin-1 agonist SF-51 for 2 hours were immunostained for LC3 (red) and LAMP1 (green). The graph reports the quantification of the number of LC3-positive structures per cell. Means ± s.d. n=300 cells per condition pooled from 3 independent experiments. p values are indicated calculated by One-way ANOVA with Tukey's post hoc test. Scale bar, 10 μm.
Supplementary Figure 8
Working model depicting the lysosome cargo response. At steady state OCRL associates with clathrin-coated pits (CCP), clathrin-coated vesicles (CCV), early endosomes and the Trans Golgi Network (TGN) (and only marginally with lysosomes) and the phosphoinositides that are mainly present in lysosomes are PI3P and PI(3,5)P2. When autophagosomes (AV) fuse with lysosomes (LY) during starvation, the mtDNA released into lysosomes activates TLR9, which translocates from the endoplasmic reticulum (ER) to autolysosomes (AL). TIRAP, a known component of the TLR9 signalling pathway, also translocates to AL. The activation of TLR9 in AL induces the recruitment of PI(4,5)P2-producing enzymes (PIP5Kα and β) that leads to an increase in PI(4,5)P2. This increase is restricted by the simultaneous recruitment of OCRL, a PI(4,5)P2 5-phosphatase. The lysosomal translocation of OCRL is driven by the clathrin adaptor AP2 and clathrin, which associate with lysosomes in a PI(4,5)P2-dependent manner. The fine spatial control of PI(4,5)P2 in lysosomes is required to preserve the activity of MCOLN1 (required for autophagosome-lysosome fusion) which is inhibited by PI(4,5)P2. OCRL, which interacts with MCOLN1, ensures PI(4,5)P2-free microdomains around MCOLN1.
Supplementary Figure 9
Unprocessed scans of original blots shown in figures and supplementary figures.
The uncropped and unprocessed scans of original blots shown in this study are reported.
Supplementary Table 1
siRNA oligonucleotides used in this study.
Names of the genes, RNA sequences, company names and catalog numbers (if available) of the siRNAs used in this study are reported.
Supplementary Table 2
Statistic source data.
The data used to derive the statistics of all the relevant experiments shown in main and supplementary figures are reported.
Supplementary Table 3
Primary and secondary antibodies used in this study.
Names, catalog numbers, clone numbers (only for monoclonal antibodies), company names, working dilution and application in which the antibodies were used in this study are described.
Supplementary Table 4
Sequence of oligonucleotides used in this study.
Names and DNA sequences of qPCR, mutagenesis and cloning primers used in this study are reported.
Acknowledgements
We thank S. Aydin (UCL Brussels), N. Demoulin (UCL Brussels), M-C. Gubler (INSERM Paris), G. Montini (Univ. Bologna), U. Querfeld (Charité Berlin), B. Rudolph (Charité Berlin) and F. Schaeffer (Univ. Heidelberg) for providing renal biopsies from patients with Lowe syndrome and appropriate controls, Cathal Wilson, Francesco Emma, Carmine Settembre, Nicola Brunetti Pierri, and Rossella Venditti for insightful discussions, Graciana Diez-Roux for critical reading of the manuscript, the TIGEM Bioinformatics Core and Mario Failli for the analysis of the gene expression profiles, the Advanced Microscopy and Imaging Core of TIGEM for the EM analysis, and the Central Proteomics Facility, Sir William Dunn Pathology School, Oxford University. MADM acknowledges the support of Telethon (grant TGM11CB1), Italian Association for Cancer Research (AIRC, grant IG2013_14761), European Research Council Advanced Investigator grant no. 670881 (SYSMET), Associazione Italiana Sindrome di Lowe (AISLO), Lowe Syndrome Association, USA (LSA) and Programma Operativo Nazionale (PON) 01_00862. MGDL received a Fellowship from AIRC. Elena Levtchenko was supported by the fund for Scientific Research, Flanders (F.W.O. Vlaanderen) grant 1801110N. FO was supported by a grant from the UK Lowe Syndrome Trust (NoMU/ML/1010) awarded to ML. OD was supported by the Fonds National de la Recherche Scientifique and the Fonds de la Recherche Scientifique Médicale (Brussels, Belgium); the European Community's Seventh Framework Programme under grant agreement n° 305608 (EURenOmics); the Cystinosis Research Foundation (Irvine, CA, USA); the Rare Disease Initiative Zürich (RADIZ), and the Swiss National Science Foundation project grant 310030_146490 (OD and AL).
Footnotes
Author Contributions
M.A.D.M. supervised the entire project; M.A.D.M. and L.S wrote the manuscript with comments from all co-authors; M.G.D.L and L.S designed and conducted the experiments of immuno-localization, induction of autophagy, Western blots, and immunoprecipitation with the help of V.M, M.S. and G.D.T.; M.V. and A.D.C performed the initial experiments on PTCs from Lowe syndrome patients. M.S. and G.D.T designed the strategy and produced plasmid vectors for the different constructs used in the study; M.S and G.D.T prepared recombinant proteins and anti-OCRL antibodies; E.P. designed and conducted the experiments for electron microscopy; D.M. performed the calcium measurements, F.O., T.S. and M.L. performed the study in the OCRL -/- zebrafish model, E.L. isolated the PTCs from the urine of Lowe syndrome patients, A.L. performed the study of immunostaining in kidney biopsies under the supervision of O.L.; A.C, M.M and D.D.B. performed the analysis of the gene expression profiles; A.B. provided the knowledge and material needed for the study of autophagy and TFEB.
Author Information:
The Authors declare no competing interest.
References
- 1.Attree O, et al. The Lowe's oculocerebrorenal syndrome gene encodes a protein highly homologous to inositol polyphosphate-5-phosphatase. Nature. 1992;358:239–242. doi: 10.1038/358239a0. [DOI] [PubMed] [Google Scholar]
- 2.Staiano L, De Leo MG, Persico M, De Matteis MA. Mendelian disorders of PI metabolizing enzymes. Biochim Biophys Acta. 2015;1851:867–881. doi: 10.1016/j.bbalip.2014.12.001. [DOI] [PubMed] [Google Scholar]
- 3.Mehta ZB, Pietka G, Lowe M. The cellular and physiological functions of the Lowe syndrome protein OCRL1. Traffic. 2014;15:471–487. doi: 10.1111/tra.12160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Nandez R, et al. A role of OCRL in clathrin-coated pit dynamics and uncoating revealed by studies of Lowe syndrome cells. Elife. 2014;3:e02975. doi: 10.7554/eLife.02975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Vicinanza M, et al. OCRL controls trafficking through early endosomes via PtdIns4,5P(2)-dependent regulation of endosomal actin. EMBO J. 2011;30:4970–4985. doi: 10.1038/emboj.2011.354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Carvalho-Neto A, Ono SE, Cardoso Gde M, Santos ML, Celidonio I. Oculocerebrorenal syndrome of Lowe: magnetic resonance imaging findings in the first six years of life. Arq Neuropsiquiatr. 2009;67:305–307. doi: 10.1590/s0004-282x2009000200027. [DOI] [PubMed] [Google Scholar]
- 7.Allmendinger AM, Desai NS, Burke AT, Viswanadhan N, Prabhu S. Neuroimaging and renal ultrasound manifestations of Oculocerebrorenal syndrome of Lowe. J Radiol Case Rep. 2014;8:1–7. doi: 10.3941/jrcr.v8i10.1740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Settembre C, et al. TFEB links autophagy to lysosomal biogenesis. Science. 2011;332:1429–1433. doi: 10.1126/science.1204592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Oltrabella F et al. The Lowe syndrome protein OCRL1 is required for endocytosis in the zebrafish pronephric tubule. PLoS Genet. 2015;11:e1005058. doi: 10.1371/journal.pgen.1005058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ramirez IB, et al. Impaired neural development in a zebrafish model for Lowe syndrome. Hum Mol Genet. 2012;21:1744–1759. doi: 10.1093/hmg/ddr608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zhang X, Hartz PA, Philip E, Racusen LC, Majerus PW. Cell lines from kidney proximal tubules of a patient with Lowe syndrome lack OCRL inositol polyphosphate 5-phosphatase and accumulate phosphatidylinositol 4,5-bisphosphate. J Biol Chem. 1998;273:1574–1582. doi: 10.1074/jbc.273.3.1574. [DOI] [PubMed] [Google Scholar]
- 12.Jiang P, et al. The HOPS complex mediates autophagosome-lysosome fusion through interaction with syntaxin 17. Mol Biol Cell. 2014;25:1327–1337. doi: 10.1091/mbc.E13-08-0447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kochl R, Hu XW, Chan EY, Tooze SA. Microtubules facilitate autophagosome formation and fusion of autophagosomes with endosomes. Traffic. 2006;7:129–145. doi: 10.1111/j.1600-0854.2005.00368.x. [DOI] [PubMed] [Google Scholar]
- 14.Itakura E, Mizushima N. Syntaxin 17: the autophagosomal SNARE. Autophagy. 2013;9:917–919. doi: 10.4161/auto.24109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Choudhury R, Noakes CJ, McKenzie E, Kox C, Lowe M. Differential clathrin binding and subcellular localization of OCRL1 splice isoforms. J Biol Chem. 2009;284:9965–9973. doi: 10.1074/jbc.M807442200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Mao Y, et al. A PH domain within OCRL bridges clathrin-mediated membrane trafficking to phosphoinositide metabolism. EMBO J. 2009;28:1831–1842. doi: 10.1038/emboj.2009.155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Traub LM, et al. AP-2-containing clathrin coats assemble on mature lysosomes. J Cell Biol. 1996;135:1801–1814. doi: 10.1083/jcb.135.6.1801. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Yu L, et al. Termination of autophagy and reformation of lysosomes regulated by mTOR. Nature. 2010;465:942–946. doi: 10.1038/nature09076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Rong Y, et al. Clathrin and phosphatidylinositol-4,5-bisphosphate regulate autophagic lysosome reformation. Nat Cell Biol. 2012;14:924–934. doi: 10.1038/ncb2557. [DOI] [PubMed] [Google Scholar]
- 20.Sridhar S, et al. The lipid kinase PI4KIIIbeta preserves lysosomal identity. EMBO J. 2013;32:324–339. doi: 10.1038/emboj.2012.341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zoncu R, et al. mTORC1 senses lysosomal amino acids through an inside-out mechanism that requires the vacuolar H(+)-ATPase. Science. 2011;334:678–683. doi: 10.1126/science.1207056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Shoji-Kawata S, et al. Identification of a candidate therapeutic autophagy-inducing peptide. Nature. 2013;494:201–206. doi: 10.1038/nature11866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Mudaliar H, et al. The role of Toll-like receptor proteins (TLR) 2 and 4 in mediating inflammation in proximal tubules. Am J Physiol Renal Physiol. 2013;305:F143–154. doi: 10.1152/ajprenal.00398.2012. [DOI] [PubMed] [Google Scholar]
- 24.Benigni A, et al. Involvement of renal tubular Toll-like receptor 9 in the development of tubulointerstitial injury in systemic lupus. Arthritis Rheum. 2007;56:1569–1578. doi: 10.1002/art.22524. [DOI] [PubMed] [Google Scholar]
- 25.Li X, et al. The role of Toll-like receptor (TLR) 2 and 9 in renal ischemia and reperfusion injury. Urology. 2013;81:1379 e15–20. doi: 10.1016/j.urology.2013.02.016. [DOI] [PubMed] [Google Scholar]
- 26.Brencicova E, Diebold SS. Nucleic acids and endosomal pattern recognition: how to tell friend from foe? Front Cell Infect Microbiol. 2013;3:37. doi: 10.3389/fcimb.2013.00037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.O'Neill LA, Golenbock D, Bowie AG. The history of Toll-like receptors - redefining innate immunity. Nat Rev Immunol. 2013;13:453–460. doi: 10.1038/nri3446. [DOI] [PubMed] [Google Scholar]
- 28.Zhang Q, et al. Circulating mitochondrial DAMPs cause inflammatory responses to injury. Nature. 2010;464:104–107. doi: 10.1038/nature08780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lee BL, et al. UNC93B1 mediates differential trafficking of endosomal TLRs. Elife. 2013;2:e00291. doi: 10.7554/eLife.00291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wu JY, Kuo CC. ADP-Ribosylation Factor 3 Mediates Cytidine-Phosphate-Guanosine Oligodeoxynucleotide-Induced Responses by Regulating Toll-Like Receptor 9 Trafficking. J Innate Immun. 2015;7:623–636. doi: 10.1159/000430785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zhang JZ, Liu Z, Liu J, Ren JX, Sun TS. Mitochondrial DNA induces inflammation and increases TLR9/NF-kappaB expression in lung tissue. Int J Mol Med. 2014;33:817–824. doi: 10.3892/ijmm.2014.1650. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 32.Oka T, et al. Mitochondrial DNA that escapes from autophagy causes inflammation and heart failure. Nature. 2012;485:251–255. doi: 10.1038/nature10992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Carchman EH, et al. Experimental sepsis-induced mitochondrial biogenesis is dependent on autophagy, TLR4, and TLR9 signaling in liver. FASEB J. 2013;27:4703–4711. doi: 10.1096/fj.13-229476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Pawaria S, et al. Cutting Edge: DNase II deficiency prevents activation of autoreactive B cells by double-stranded DNA endogenous ligands. J Immunol. 2015;194:1403–1407. doi: 10.4049/jimmunol.1402893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Chen G, et al. A regulatory signaling loop comprising the PGAM5 phosphatase and CK2 controls receptor-mediated mitophagy. Mol Cell. 2014;54:362–377. doi: 10.1016/j.molcel.2014.02.034. [DOI] [PubMed] [Google Scholar]
- 36.Ding WX, et al. Nix is critical to two distinct phases of mitophagy, reactive oxygen species-mediated autophagy induction and Parkin-ubiquitin-p62-mediated mitochondrial priming. J Biol Chem. 2010;285:27879–27890. doi: 10.1074/jbc.M110.119537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Narendra D, Tanaka A, Suen DF, Youle RJ. Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J Cell Biol. 2008;183:795–803. doi: 10.1083/jcb.200809125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Nguyen TT, et al. Phosphatidylinositol 4-phosphate 5-kinase alpha facilitates Toll-like receptor 4-mediated microglial inflammation through regulation of the Toll/interleukin-1 receptor domain-containing adaptor protein (TIRAP) location. J Biol Chem. 2013;288:5645–5659. doi: 10.1074/jbc.M112.410126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kimura S, Noda T, Yoshimori T. Dissection of the autophagosome maturation process by a novel reporter protein, tandem fluorescent-tagged LC3. Autophagy. 2007;3:452–460. doi: 10.4161/auto.4451. [DOI] [PubMed] [Google Scholar]
- 40.Platt N, et al. Immune dysfunction in Niemann-Pick disease type C. J Neurochem. 2015;136 Suppl 1:74–80. doi: 10.1111/jnc.13138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ravikumar B, Moreau K, Jahreiss L, Puri C, Rubinsztein DC. Plasma membrane contributes to the formation of pre-autophagosomal structures. Nat Cell Biol. 2010;12:747–757. doi: 10.1038/ncb2078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Klionsky DJ, Abdalla FC, Abeliovich H, Abraham RT, et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy. 2012;8:445–544. doi: 10.4161/auto.19496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Pirruccello M, et al. Identification of inhibitors of inositol 5-phosphatases through multiple screening strategies. ACS Chem Biol. 2014;9:1359–1368. doi: 10.1021/cb500161z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Zhang X, Li X, Xu H. Phosphoinositide isoforms determine compartment-specific ion channel activity. Proc Natl Acad Sci USA. 2012;109:11384–11389. doi: 10.1073/pnas.1202194109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Vergarajauregui S, Connelly PS, Daniels MP, Puertollano R. Autophagic dysfunction in mucolipidosis type IV patients. Hum Mol Genet. 2008;17:2723–2737. doi: 10.1093/hmg/ddn174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Abe K, Puertollano R. Role of TRP channels in the regulation of the endosomal pathway. Physiology (Bethesda) 2011;26:14–22. doi: 10.1152/physiol.00048.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Grimm C, et al. Small molecule activators of TRPML3. Chem Biol. 2010;17:135–148. doi: 10.1016/j.chembiol.2009.12.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Xu H, Ren D. Lysosomal physiology. Annu Rev Physiol. 2015;77:57–80. doi: 10.1146/annurev-physiol-021014-071649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Nikoletopoulou V, Papandreou ME, Tavernarakis N. Autophagy in the physiology and pathology of the central nervous system. Cell Death Differ. 2015;22:398–407. doi: 10.1038/cdd.2014.204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Di Fruscio G, et al. Lysoplex: An efficient toolkit to detect DNA sequence variations in the autophagy-lysosomal pathway. Autophagy. 2015;11:928–938. doi: 10.1080/15548627.2015.1043077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Yuan Y, et al. Mitochondrial dysfunction accounts for aldosterone-induced epithelial-to-mesenchymal transition of renal proximal tubular epithelial cells. Free Radic Biol Med. 2012;53:30–43. doi: 10.1016/j.freeradbiomed.2012.03.015. [DOI] [PubMed] [Google Scholar]
- 52.Gautier L, Cope L, Bolstad BM, Irizarry RA. affy--analysis of Affymetrix GeneChip data at the probe level. Bioinformatics. 2004;20:307–315. doi: 10.1093/bioinformatics/btg405. [DOI] [PubMed] [Google Scholar]
- 53.Irizarry RA, et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics. 2003;4:249–264. doi: 10.1093/biostatistics/4.2.249. [DOI] [PubMed] [Google Scholar]
- 54.Baldi P, Long AD. A Bayesian framework for the analysis of microarray expression data: regularized t -test and statistical inferences of gene changes. Bioinformatics. 2001;17:509–519. doi: 10.1093/bioinformatics/17.6.509. [DOI] [PubMed] [Google Scholar]
- 55.Kuhn RM, Haussler D, Kent WJ. The UCSC genome browser and associated tools. Brief Bioinform. 2013;14:144–161. doi: 10.1093/bib/bbs038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Wasserman WW, Sandelin A. Applied bioinformatics for the identification of regulatory elements. Nat Rev Genet. 2004;5:276–287. doi: 10.1038/nrg1315. [DOI] [PubMed] [Google Scholar]
- 57.Sardiello M, et al. A gene network regulating lysosomal biogenesis and function. Science. 2009;325:473–477. doi: 10.1126/science.1174447. [DOI] [PubMed] [Google Scholar]
- 58.Ashley N, Harris D, Poulton J. Detection of mitochondrial DNA depletion in living human cells using PicoGreen staining. Exp Cell Res. 2005;303:432–446. doi: 10.1016/j.yexcr.2004.10.013. [DOI] [PubMed] [Google Scholar]
- 59.Schneider CA, Rasband WS, Eliceiri KW. NIH Image to ImageJ: 25 years of image analysis. Nat Methods. 2012;9:671–675. doi: 10.1038/nmeth.2089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Hammond GR, Schiavo G, Irvine RF. Immunocytochemical techniques reveal multiple, distinct cellular pools of PtdIns4P and PtdIns(4,5)P(2) Biochem J. 2009;422:23–35. doi: 10.1042/BJ20090428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Carpenter AE, et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 2006;7:R100. doi: 10.1186/gb-2006-7-10-r100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Ladoire S, et al. Immunohistochemical detection of cytoplasmic LC3 puncta in human cancer specimens. Autophagy. 2012;8:1175–1184. doi: 10.4161/auto.20353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Polishchuk EV, Polishchuk RS, Luini A. Correlative light-electron microscopy as a tool to study in vivo dynamics and ultrastructure of intracellular structures. Methods Mol Biol. 2013;931:413–422. doi: 10.1007/978-1-62703-056-4_20. [DOI] [PubMed] [Google Scholar]
- 64.Luciani A, et al. Defective CFTR induces aggresome formation and lung inflammation in cystic fibrosis through ROS-mediated autophagy inhibition. Nat Cell Biol. 2010;12:863–875. doi: 10.1038/ncb2090. [DOI] [PubMed] [Google Scholar]
- 65.R_Core_Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing; Vienna, Austria: 2014. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Microarray data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE54284. Source data for Fig. 2b, 2c, 3a, 3c, 4a, 4c, 4e, 4f, 4h, 5b, 5c, 5d, 5e, 5f, 5g, 6b, 6c, 6d, 6e, 6g, 7b, 7c, 7e, 7f, 8b, 8c and 8d and for Supplementary Fig. 2b, 3a, 3b, 3e, 4e, 4f, 4g, 5a, 5f, 7b are available in Supplementary Table 2: Statistic source data. All other data supporting the findings of this study are available from the corresponding author on reasonable request.