Abstract
Staging and pathological grading are useful, but imperfect predictors of recurrence in head and neck squamous cell carcinoma (HNSCC). Accordingly, molecular biomarkers that predict the risk of recurrence are necessary to improve clinical outcomes. The methylation statuses of the promoters of 11 tumor-related genes (p16, RASSF1A, E-cadherin, H-cadherin, MGMT, DAPK, DCC, COL1A2, TAC1, SST, and GALR1) were analyzed in 133 HNSCC cases using quantitative methylation-specific PCR. We detected frequent methylation of p16 (44%), RASSF1A (18%), E-cadherin (53%), H-cadherin (35%), MGMT (35%), DAPK (53%), DCC (42%), COL1A2 (44%), TAC1 (61%), SST (64%), and GALR1 (44%) in HNSCC. Disease-free survival was lower in patients with 6–11 methylated genes than in those with 0–5 methylated genes (log-rank test, P = 0.001). In a multivariate Cox proportional hazards analysis, the methylation of E-cadherin, COL1A2, TAC1, and GALR1 was associated with poor survival, with hazard ratios of 4.474 (95% CI, 1.241–16.124). In a joint analysis of these four genes, patients with 2–4 methylated genes had a significantly lower survival rate than those with 0–1 methylated genes in early-stage HNSCC. Importantly, the methylation of some genes was closely related to poor prognosis in early-stage HNSCC, providing strong evidence that these hypermethylated genes are valuable biomarkers for prognostic evaluation.
Keywords: tumor-suppressor genes, hypermethylation, head and neck cancer, metastases, biomarker
INTRODUCTION
The treatment strategy for patients with HNSCC is generally guided by tumor–node–metastasis (TNM) classification and clinical staging. Only 90% of stage I patients can be cured by surgery or radiotherapy. The fraction of aggressive tumors rises to 30% for stage II patients, 50% at stage III, and 70% at stage IV. These aggressive tumors relapse quickly and progress, ultimately causing death [1–3]. Biomarker development is necessary to improve our understanding of the molecular basis of HNSCC progression and to provide sufficient discriminatory prognostic power for the effective clinical management of this disease [4].
Alterations in epigenetic marks (i.e., hypermethylation events) are useful biomarkers; for example, MGMT epigenetic alterations are useful biomarkers in glioblastomas [5] and GSTP1 alterations are useful in prostate cancers [6]. MGMT predicts the response to DNA-alkylating drugs [7]. GSTP1 is an established biomarker for prostate cancer diagnosis and prognosis [7]. In a previous analysis of HNSCC, there was no observable effect of p16, MGMT, DAPK, or E-cadherin on prognosis for patients with laryngeal and hypopharyngeal cancer [8]. Tan et al. demonstrated that hypermethylated promoters in the surgical margins of HNSCC predict local recurrences and disease-specific deaths based on a panel of three genes (p16, cyclin A1, and DCC) [9]. According to Carvalho et al., salivary DNA promoter hypermethylation analyses facilitate the early diagnosis of HNSCC, and several hypermethylated genes (DAPK, DCC, MINT-31, TIMP-3, p16, MGMT, and CCNA1) have already been identified in salivary rinse samples [10–12]. Although several hypermethylated candidate genes have been identified in HNSCC, few are suitable methylation markers for clinical use.
Therapeutic procedures differ substantially between early- and late-stage HNSCC. Early-stage HNSCC patients receive minimally invasive surgery or irradiation alone, and late-stage patients receive aggressive therapy, such as expanded surgery and/or concomitant chemoradiotherapy [2, 3, 13]. The ability to distinguish between low- and high-risk HNSCCs at an early stage may reduce follow-up costs. We hypothesized that the quantitative methylation-specific PCR (Q-MSP) assay could be used to define patterns of DNA methylation that differentiate low- and high-risk HNSCCs.
We prospectively analyzed 11 genes involved in cell cycle control (p16, galanin receptor 1; GALR1), DNA damage repair (O6-alkylguanine DNA alkyltransferase; MGMT), apoptosis (death-associated protein kinase; DAPK, Ras association domain-containing protein 1; RASSF1A), inflammatory reactions (tachykinin, precursor 1; TAC1), antitumor and antisecretory activity (somatostatin; SST), and tumor cell invasion (E-cadherin, H-cadherin, deleted in colorectal carcinoma [DCC], and collagen alpha-2(I) chain [COL1A2]) in a cohort of clinically well-characterized HNSCC samples. Furthermore, we identified the most powerful combination of hypermethylated genes, and characterized an early stage-specific marker for HNSCC treatment.
RESULTS
Distributions of individual methylated genes
We used Q-MSP to examine promoter methylation of 11 genes in 133 primary HNSCC tumors. One hundred and thirty primary tumors (97.7%) showed hypermethylation of at least one gene in the panel. Thirty-seven (37 of 133; 27.8%) tumors included 0 to 3 hypermethylated genes, 18.0% (24/133) had 4 hypermethylated genes, 38.3% (51/133) had 5 to 7 hypermethylated genes, and 15.8% (21/133) had 8 to 10 hypermethylated genes (Figure 1A). The mean number of methylated genes was 4.95 (range, 0–10). In particular, we detected frequent methylation of p16 (44.4%), RASSF1A (18.0%), E-cadherin (53.6%), H-cadherin (35.3%), MGMT (35.3%), DAPK (53.4%), DCC (42.1%), COL1A2 (44.4%), TAC1 (61.0%), SST (64.0%), and GALR1 (44.4%) in HNSCC (Figure 1B).
Figure 1. Summary of gene promoter hypermethylation in 133 HNSCC samples.
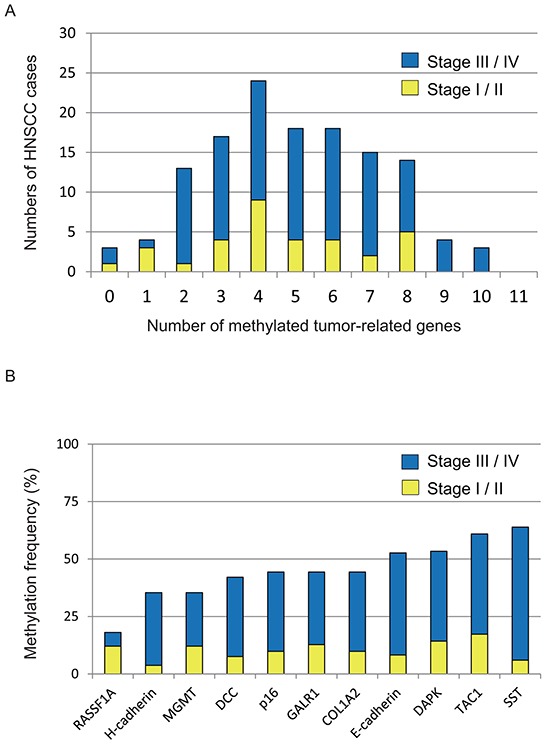
A. Bar graph showing the numbers of tumor-related methylated genes for 33 stage I/II (yellow bars) and 100 stage III/IV (blue bars) samples. B. Bar graph showing methylation frequencies (%) of 11 tumor-related genes in the cohort. Blue bars, samples of stage III/IV cases; yellow bars, samples of stage I/II cases.
Clinicopathological characteristics of primary HNSCC patients
Patient clinical features were used to examine differences in methylation index (MI) with respect to age, gender, alcohol exposure, smoking status, tumor size, lymph-node status, and stage. Based on continuous marker methylation analyses, the MI of 11 tumor-related genes (TRGs) were not correlated with any patient characteristics (Supplementary Figure S2). As summarized in Table 1, we performed a detailed analysis of methylation status for each gene according to clinical characteristics. We found that p16 promoter methylation is inversely associated with age (Fisher's exact tests; P = 0.019). There was an association between methylation of the COL1A2 promoter and gender (P = 0.022). Methylation of TAC1 was significantly correlated with alcohol exposure (P = 0.039). Methylation of GALR1 was significantly correlated with tumor size (P = 0.002) and clinical stage (P = 0.009). Methylation of the promoters of other genes was not associated with age at onset, gender, alcohol exposure, smoking status, tumor site, tumor size, lymph-node status, or clinical stage (Table 1).
Table 1. Distribution of methylation status by selected epidemiologic and clinical characteristics.
| Gene | Methylation status | Characteristics | Age | Gender | Alcohol exposure | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Overall(%) | 70< | 70> | P † | Female | Male | P † | drinker | non drinker | P † | ||
| p16 | Yes | 59 (44.4) | 49 | 10 | 0.019* | 12 | 47 | 1 | 38 | 21 | 1 |
| No | 74 (55.6) | 47 | 27 | 12 | 62 | 49 | 25 | ||||
| RASSF1A | Yes | 24 (18.0) | 16 | 8 | 1 | 4 | 20 | 1 | 15 | 9 | 1 |
| No | 109 (82.0) | 80 | 29 | 20 | 89 | 72 | 37 | ||||
| E-cadherin | Yes | 70 (52.6) | 47 | 23 | 0.182 | 14 | 56 | 0.653 | 46 | 24 | 1 |
| No | 63 (47.4) | 49 | 14 | 10 | 53 | 41 | 22 | ||||
| H-cadherin | Yes | 47 (35.3) | 34 | 13 | 1 | 10 | 37 | 1 | 32 | 15 | 0.705 |
| No | 86 (64.7) | 62 | 24 | 14 | 72 | 55 | 31 | ||||
| MGMT | Yes | 47 (35.3) | 30 | 17 | 1 | 9 | 38 | 1 | 30 | 17 | 1 |
| No | 86 (64.7) | 66 | 20 | 15 | 71 | 57 | 29 | ||||
| DAPK | Yes | 71 (53.4) | 53 | 18 | 0.563 | 17 | 54 | 0.072 | 51 | 20 | 0.104 |
| No | 62 (46.6) | 43 | 19 | 7 | 55 | 36 | 26 | ||||
| DCC | Yes | 56 (42.1) | 39 | 17 | 1 | 10 | 46 | 1 | 37 | 19 | 1 |
| No | 77 (57.9) | 57 | 20 | 14 | 63 | 50 | 27 | ||||
| COL1A2 | Yes | 59 (44.4) | 43 | 16 | 1 | 16 | 43 | 0.022* | 36 | 23 | 1 |
| No | 74 (55.6) | 53 | 21 | 8 | 66 | 51 | 23 | ||||
| TAC1 | Yes | 81 (60.9) | 55 | 26 | 0.234 | 13 | 68 | 1 | 59 | 22 | 0.039* |
| No | 52 (39.1) | 41 | 11 | 11 | 41 | 28 | 24 | ||||
| SST | Yes | 85 (63.9) | 63 | 22 | 1 | 13 | 72 | 1 | 60 | 25 | 1 |
| No | 48 (36.1) | 33 | 15 | 11 | 37 | 27 | 21 | ||||
| GALR1 | Yes | 59 (44.4) | 44 | 15 | 0.7 | 11 | 48 | 1 | 36 | 23 | 1 |
| No | 74 (55.6) | 52 | 22 | 13 | 61 | 51 | 23 | ||||
| Smoking status | Tumor size | Lympho-node status | Stage | ||||||||
| smoker | non smoker | P† | T1-2 | T3-4 | P† | N0 | N+ | P† | I-II | III-IV | P† |
| 40 | 19 | 1 | 28 | 31 | 1 | 27 | 32 | 1 | 16 | 43 | 1 |
| 55 | 19 | 31 | 43 | 32 | 42 | 17 | 57 | ||||
| 17 | 7 | 1 | 12 | 12 | 1 | 9 | 15 | 0.503 | 5 | 19 | 0.795 |
| 78 | 31 | 47 | 62 | 50 | 59 | 28 | 81 | ||||
| 52 | 18 | 0.45 | 33 | 37 | 0.6 | 30 | 40 | 1 | 16 | 54 | 0.688 |
| 43 | 20 | 26 | 37 | 29 | 34 | 17 | 46 | ||||
| 35 | 12 | 0.689 | 23 | 24 | 1 | 18 | 29 | 0.362 | 10 | 37 | 0.535 |
| 60 | 26 | 36 | 50 | 41 | 45 | 23 | 63 | ||||
| 33 | 14 | 1 | 20 | 27 | 0.856 | 20 | 27 | 0.856 | 13 | 34 | 1 |
| 62 | 24 | 39 | 47 | 39 | 47 | 20 | 66 | ||||
| 53 | 18 | 0.443 | 33 | 38 | 0.605 | 31 | 40 | 1 | 17 | 54 | 1 |
| 42 | 20 | 26 | 36 | 28 | 34 | 16 | 46 | ||||
| 38 | 18 | 1 | 25 | 31 | 1 | 26 | 30 | 1 | 13 | 43 | 0.839 |
| 57 | 20 | 34 | 43 | 33 | 44 | 20 | 57 | ||||
| 42 | 17 | 1 | 23 | 36 | 0.295 | 22 | 37 | 0.162 | 11 | 48 | 0.161 |
| 53 | 21 | 36 | 38 | 37 | 37 | 22 | 52 | ||||
| 63 | 18 | 0.051 | 34 | 47 | 1 | 34 | 47 | 1 | 19 | 62 | 1 |
| 32 | 20 | 25 | 27 | 25 | 27 | 14 | 38 | ||||
| 61 | 24 | 1 | 40 | 45 | 0.469 | 36 | 49 | 1 | 23 | 62 | 0.532 |
| 34 | 14 | 19 | 29 | 23 | 25 | 10 | 38 | ||||
| 42 | 17 | 1 | 17 | 42 | 0.002* | 22 | 37 | 0.162 | 8 | 51 | 0.009* |
| 53 | 21 | 42 | 32 | 37 | 37 | 25 | 49 | ||||
Fisher's exact probability test.
P<0.05.
Association between TRG methylation and survival
Table 2 illustrates the overall associations between the methylation status of individual TRGs and disease-free survival (DFS) based on a logistic regression model. After adjusting for age, gender, smoking status, stage, we found that hypermethylation of E-cadherin, COL1A2, TAC1, and GALR1 was associated with significantly reduced survival, with hazard ratios of 2.263 (95% CI, 1.103–4.641), 3.824 (95% CI, 1.794–8.152), 3.216 (95% CI, 1.491–6.937), and 3.125 (95% CI, 1.489–6.557), respectively (Table 2).
Table 2. Methylation status of individual genes and associations with disease-free survival using Logistic regression model.
| Gene | Methylation status | Overall(%) | Recurrence events | Adjusted RR (95% CI)† | |
|---|---|---|---|---|---|
| Positive (N = 67) | Negative (N = 66) | ||||
| p16 | Yes | 59 (44.4) | 32 | 27 | 1.300 (0.632-2.672) |
| No | 74 (55.6) | 35 | 39 | ||
| RASSF1A | Yes | 24 (18.0) | 13 | 11 | 1.214 (0.490-3.009) |
| No | 109 (82.0) | 54 | 55 | ||
| E-cadherin | Yes | 70 (52.6) | 42 | 28 | 2.311 (1.126-4.744)* |
| No | 63 (47.4) | 25 | 38 | ||
| H-cadherin | Yes | 47 (35.3) | 29 | 18 | 1.961 (0.935-4.113) |
| No | 86 (64.7) | 38 | 48 | ||
| MGMT | Yes | 47 (35.3) | 27 | 20 | 1.639 (0.780-3.444) |
| No | 86 (64.7) | 40 | 46 | ||
| DAPK | Yes | 71 (53.4) | 41 | 30 | 1.751 (0.857-3.580) |
| No | 62 (46.6) | 26 | 36 | ||
| DCC | Yes | 56 (42.1) | 31 | 25 | 1.419 (0.701-2.875) |
| No | 77 (57.9) | 36 | 41 | ||
| COL1A2 | Yes | 59 (44.4) | 41 | 18 | 3.824 (1.802-8.117)* |
| No | 74 (55.6) | 26 | 48 | ||
| TAC1 | Yes | 81 (60.9) | 49 | 32 | 3.824 (1.802-7.532)* |
| No | 52 (39.1) | 18 | 34 | ||
| SST | Yes | 85 (63.9) | 45 | 40 | 1.485 (0.712-3.099) |
| No | 48 (36.1) | 22 | 26 | ||
| GALR1 | Yes | 59 (44.4) | 39 | 20 | 3.132 (1.489-6.585)* |
| No | 74 (55.6) | 28 | 46 | ||
Adjusted for age (70 and older vs. <70), gender, smoking status and stage (I, II vs. III, IV).
P<0.05.
Based on log-rank tests, we detected an association between poor survival and the methylation phenotype defined as ≥6 methylated genes (P = 0.001) (Supplementary Table S1). Kaplan-Meier plots indicated that methylation of 11 TRGs in patient samples was related to the duration of DFS. The DFS was lower in patients with 6–11 methylated genes than in the group with 0–5 methylated genes (60.3% versus 16.1%, respectively; log-rank test, P = 0.001) (Figure 2A). Among 59 patients with T1 and T2 tumors, the DFS rate was 26.8% in the group of patients with 6–11 methylated genes and was 67.5% in the 0–5 group (log-rank test, P = 0.038) (Figure 2B). Among 59 patients with N0 lympho-node status, there was no significant association between patients with 6–11 methylated genes and 0 to 5 methylated genes (log-rank test, P = 0.124) (Figure 2D). Among 33 patients with stage I and II patients, no correlation was found between patients with a high (6 to 11) and low (0 to 5) number of methylated genes (log-rank test, P = 0.165) (Figure 2F). In 100 stage III and IV patients, the DFS was statistically significantly worse in the group with a high number of methylated genes (log-rank test, P = 0.007) (Figure 2G).
Figure 2. Kaplan-Meier survival curves for patients with HNSCC according to the methylation status of 11 tumor-related genes.
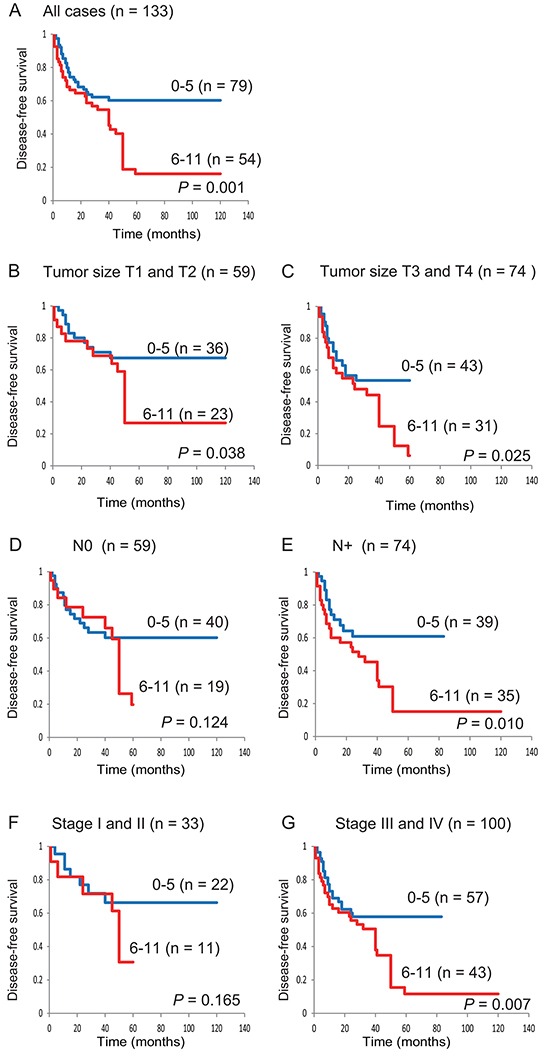
Disease-free survival for A. all 133 HNSCC cases, B. tumor size T1 and T2 cases (n = 59), C. tumor size T3 and T4 cases (n = 74), D. lymph-node status N0 cases (n = 59), E. lymph-node status N+ cases (n = 74), F. stage I and II cases (n = 33), and G. stage III and IV cases (n = 100). Blue line: patients with 0–5 methylated genes; red line: patients with 6–11 methylated genes.
Patients with 2 to 4 methylated genes (in an analysis of E-cadherin, COL1A2, TAC1, and GALR1) had a trend toward worse survival than those with 0 to 1 methylated genes (57.0% versus 37.3%, respectively; log-rank test, P = 0.126) (Figure 3A). For the 4 genes, among 59 patients with T1 and T2 tumor sizes, DFS was slightly lower in the 2–4 methylated genes group than in the 0–1 methylated genes group (39.8% versus 68.5%, respectively; log-rank test, P = 0.109) (Figure 3B). Among 59 patients with N0 lymph-node status, patients with 0 to 1 methylated genes showed significantly better DFS than patients with 2 to 4 methylated genes (71.8% versus 36.6%, respectively; log-rank test, P = 0.029) (Figure 3D). Among 33 stage I and II patients, those with 0 to 1 methylated genes showed significantly better DFS than patients with 2 to 4 hypermethylated genes (74.5% versus 26.7%, respectively; log-rank test, P = 0.035) (Figure 3F). These data indicate that the methylation profiles of E-cadherin, COL1A2, TAC1, and GALR1 are a powerful combination for the prediction of early-stage HNSCC.
Figure 3. Kaplan-Meier survival curves for patients with HNSCC according to E-cadherin, COL1A2, TAC1, and GALR1 methylation status.
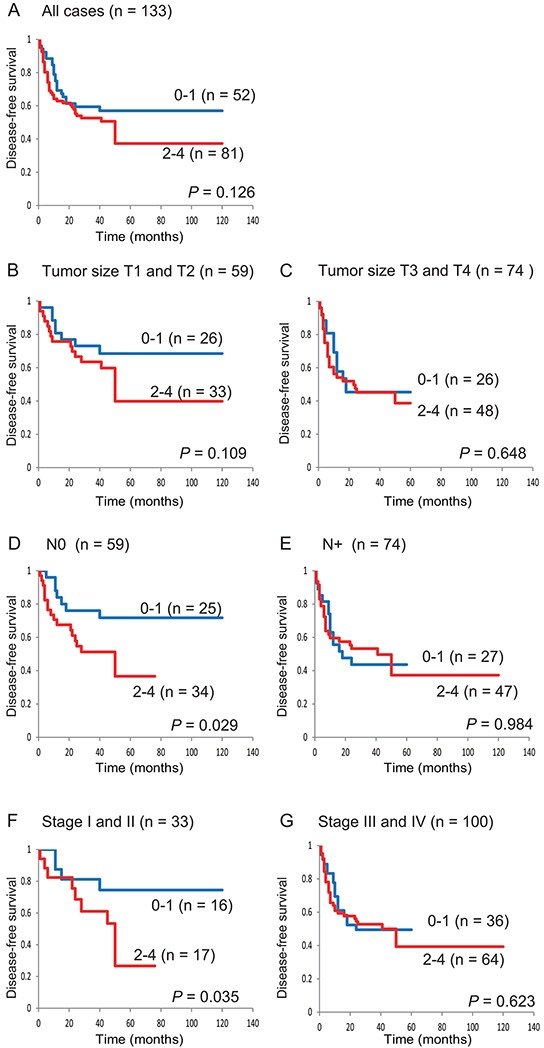
Disease-free survival for A. all 133 HNSCC cases, B. tumor size T1 and T2 cases (n = 59), C. tumor size T3 and T4 cases (n = 74), D. lymph-node status N0 cases (n = 59), E. lymph-node status N+ cases (n = 74), F. stage I and II cases (n = 33), and G. stage III and IV cases (n = 100). Blue line: patients with 0–1 methylated genes; red line: patients with 2–4 methylated genes.
Moreover, based on a multivariate Cox proportional hazard regression of 59 patients with N0 lymph-node status, which included age, gender, smoking status, and tumor stage, the group with methylation of E-cadherin, COL1A2, TAC1, and GALR1 had a 4.474-times greater hazard than the group without methylation (Table 3).
Table 3. The promoter hypermethylation pattern and associations with disease-free survival using Cox proportional hazards model in 59 patients with N0 lymph-node status.
| Gene (Any) | Methylation status | Overall(%) | Recurrence events | Adjusted HR (95% CI)† | |
|---|---|---|---|---|---|
| Positive(N=28) | Negative(N=31) | ||||
| E-cadherin & COL1A2 | Yes | 41 (69.5) | 23 | 18 | 2.359 (0.989-5.629) |
| No | 18 (30.5) | 5 | 13 | ||
| E-cadherin & TAC1 | Yes | 44 (74.6) | 25 | 19 | 1.989 (0.833-4.748) |
| No | 15 (25.4) | 3 | 12 | ||
| E-cadherin & GALR1 | Yes | 36 (61.0) | 20 | 16 | 2.266 (0.994-5.170) |
| No | 23 (39.0) | 8 | 15 | ||
| COL1A2 & TAC1 | Yes | 41 (69.5) | 24 | 17 | 1.202 (0.535-2.701) |
| No | 18 (30.5) | 4 | 14 | ||
| COL1A2 & GALR1 | Yes | 32 (54.2) | 19 | 13 | 5.097 (1.597-16.266)* |
| No | 27 (45.8) | 9 | 18 | ||
| TAC1 & GALR1 | Yes | 43 (72.9) | 24 | 19 | 2.170 (0.910-5.172) |
| No | 16 (27.1) | 4 | 12 | ||
| E-cadherin & COL1A2 & TAC1 | Yes | 50 (84.7) | 27 | 23 | 1.704 (0.681-4.261) |
| No | 9 (15.3) | 1 | 8 | ||
| E-cadherin & COL1A2 & GALR1 | Yes | 43 (72.9) | 23 | 20 | 3.288 (1.286-8.407)* |
| No | 16 (27.1) | 5 | 11 | ||
| E-cadherin & TAC1 & GALR1 | Yes | 47 (79.7) | 26 | 21 | 2.753 (1.048-7.229)* |
| No | 12 (20.3) | 2 | 10 | ||
| COL1A2 & TAC1 & GALR1 | Yes | 46 (78.0) | 24 | 22 | 2.782 (0.997-7.766)* |
| No | 13 (22.0) | 4 | 9 | ||
| E-cadherin & COL1A2 & TAC1 & GALR1 | Yes | 50 (84.7) | 26 | 24 | 4.474 (1.241-16.124)* |
| No | 9 (15.3) | 2 | 7 | ||
Adjusted for age (70 and older vs. <70), gender, smoking status and stage (I, II vs. III, IV).
P<0.05.
DISCUSSION
We found that aberrant patterns of promoter methylation in primary tumors are indicators of an increased risk of recurrence in patients with stage I and II HNSCC. Patients with clinical stage I and II (T1-2N0) oral squamous cell carcinoma usually undergo partial glossectomy alone. However, approximately 25% of these patients develop delayed neck metastasis, which may lead to an unfavorable course [14]. Similarly, the outcomes of patients with T1-T2N0 larynx and hypopharynx cancer who are initially treated with radiotherapy or minimally invasive surgery followed by conservative surgery for radiation failure are unclear [15]. Biomarker discovery for early-stage HNSCC is crucial to improve patient outcomes. Using surgical tissues from a pilot cohort of 33 stage I/II HNSCC patients, we identified markers that may be suitable prognostic indicators for local recurrence and poor survival.
The methylation of E-cadherin, COL1A2, TAC1, and GALR1 in primary early-stage HNSCC indicated metastatic risk in regional lymph nodes and distant organs. The methylation of p16 and H-cadherin is associated with the early recurrence of stage I non-small cell lung carcinoma [4]. We speculate that the detection of early hypermethylation events in HNSCC tumors diagnosed as tumor-free by conventional imaging analysis can be used to identify subjects at risk of recurrence.
E-cadherin promoter methylation has been detected in many tumor types [16]. Low E-cadherin expression is associated with an increased risk of late cervical metastasis in stage I and II oral cancer patients [17], and the overexpression of SIP1 and downregulation of E-cadherin predict delayed neck metastasis in stage I and II oral tongue carcinoma after partial glossectomy [18]. However, the observed methylation levels of E-cadherin vary among studies with respect to cancer type and survival.
COL1A2 is a fibrillar collagen found in most connective tissues, and is the main component of the organic part of bones. COL1A2 inactivation contributes to increased proliferation and migration activity of bladder cancer and osteosarcoma cells [19, 20]. Aberrant COL1A2 promoter methylation has been detected in various cancer types, such as breast carcinoma [21], medulloblastoma [22], and melanoma [23, 24]. Our data also suggested that hypermethylation of COL1A2 is associated with improved survival in patients with HNSCC [25].
TAC1 encodes substance P, which has proliferative and anti-apoptotic effects via the activation of the ERK1/2 and nuclear factor-κB pathways, and neurokinin A, which has antiproliferative properties [3]. Hypermethylation of TAC1 has been described in esophageal [26], gastric [27], colon [28], and breast cancer [29]. DFS is correlated with TAC1 methylation (log-rank test, P = 0.002), but not with TACR1 methylation in HNSCC [30].
GALR1 inhibits HNSCC cell proliferation via ERK1/2-mediated effects on cell cycle control proteins, such as p27, p57, and cyclin D1 [31]. We also found that GALR1 methylation is associated with a significantly worse survival rate in HNSCC patients [32]. Doufekas et al. have recently shown that GALR1 methylation in vaginal swabs from women with postmenopausal bleeding indicates endometrial malignancy with high sensitivity and specificity [33]. Guo et al. demonstrated that the methylation profiles of GALR1, AGTR1, SLC5A8, ZMYND10, and NTSR1 are a powerful combination for non-small cell lung cancer prediction [34].
Accordingly, many genes have been reported as individual biomarkers for prognosis in HNSCC. However, combined hypermethylation patterns increased predictive power for early-stage HNSCC. The current method to assess the risk of recurrence in patients with early-stage HNSCC is imprecise; half of such tumors recur after curative surgery. The correlation between short survival times and the number of methylated genes in the regional lymph nodes supports the presence of micrometastases at those sites. We identified a combination of hypermethylated genes that increase the predictive ability for early-stage HNSCC. Simultaneous analyses of the methylation status of multiple tumor suppressor genes are important for predictions of tumorigenesis, biological behavior, and the development of future targeted therapy.
In conclusion, the methylation profiles of E-cadherin, COL1A2, TAC1, and GALR1 were the most powerful combination for predicting early-stage HNSCC. This demonstrates that molecular stratification may predict cancer progression. These findings can benefit HNSCC screening and surveillance algorithms. Although our study was retrospective, was conducted at a single institution, and the number of patients was small, it serves as a platform to establish optimal therapeutic strategies for early-stage HNSCC.
MATERIALS AND METHODS
Tumor samples
A total of 133 primary HNSCC samples in an original cohort were obtained during surgery in the Department of Otolaryngology–Head and Neck Surgery, Hamamatsu University School of Medicine between 1977 and 2011. Clinical information, including age, sex, tumor site, smoking status, alcohol exposure, tumor size, lymph node status, stage, and recurrence events were obtained from clinical records. The male to female ratio was 109:24, and the mean age was 64.1 years (range 39–90). The primary tumors were located in the oral cavity (n = 45), hypopharynx (n = 31), larynx (n = 25), oropharynx (n = 22), and paranasal cavity (n = 10). The patients were stage I/II (n = 33) and stage III/IV (n = 100). Thirty-six matched pairs of head and neck tumor tissues and adjacent normal mucosal tissues were obtained from the surgical specimens.
Bisulfite modification and quantitative methylation-specific PCR analysis
Genomic DNA was obtained from tumor and normal mucosal tissues using the QIAamp DNA Mini Kit (Qiagen, Courtaboeuf, France). DNA was subjected to bisulfite treatment, as described previously. [32] The bisulfite-modified DNA was used as a template for fluorescence-based real-time PCR. [35] The amplifications were performed using the TaKaRa Thermal Cycler Dice™ Real Time System TP800 (TaKaRa, Tokyo, Japan). The Q-MSP primers for methylated DNA were Q-MSP-ACTB-F (5′-TGGTGATGGAGGAGGTTTAGAAGT-3′) and Q-MSP-ACTB-R (5′-AACCAATAAAACCTACTCCTCCCTTAA-3′). A standard curve was generated using serial dilutions of universal methylated DNAs (EpiScope™ Methylated HCT116 gDNA; TaKaRa, Tokyo, Japan). The normalized methylation value (NMV) was defined as follows: NMV = (TRGs-S/TRGs-FM)/(ACTB-S/ACTB-FM), where TRGs-S and TRGs-FM represent TRG methylation levels in the sample and universally methylated DNAs, respectively, and ACTB-S and ACTB-FM correspond to b-actin in the sample and universally methylated DNAs, respectively. To analyze the methylation status of p16 [36], RASSF1A [36], CDH1 [36], CDH13 [37], MGMT [38], DAPK [38], DCC [39], COL1A2 [25], TAC1 [30], SST [40], and GALR1 [32], primers and conditions were used as previously described.
Data analysis and statistics
A receiver-operator characteristic (ROC) curve analysis was performed using the NMVs for the 36 HNSCC and 36 adjacent normal mucosal tissues by StatMate IV (ATMS Co., Ltd., Tokyo, Japan). Using this approach, the area under the ROC curve indicated the optimal sensitivity and specificity levels at which to distinguish normal tissues from HNSCC tissues, and NMV thresholds were calculated for the TRGs. The cutoff value determined from this ROC curve was applied to determine TRG methylation frequency (Supplementary Figure S1). To determine the overall methylation rate in individual samples, the methylation index (MI) was used. [41, 42] The MI for each sample was defined as the ratio of the number of methylated genes to the number of genes tested (i.e., 11). The selected MI was defined as the number of methylated genes relative to the total number of genes tested (for E-cadherin, COL1A2, TAC1, and GALR1).
For the frequency analysis in the contingency tables, the associations between variables and methylation status were analyzed statistically using Fisher's exact tests. The disease-free interval was measured from the date of treatment to the date when locoregional recurrence or distant metastasis was diagnosed. DFS probabilities were estimated by the Kaplan-Meier method, and the log-rank test was applied to assess the significance of differences among actuarial survival curves. Multivariate logistic-regression analysis considering age (70 and older vs. <70), sex, smoking status, stage (I, II vs. III, IV), and methylated genes was used to identify the predictive value of the prognostic factors. Cox's proportional hazards regression analysis, which included age (70 and older vs. less than 70), sex, smoking status, stage (I, II vs. III, IV), and any methylated genes, was used to identify the multivariate predictive value of the prognostic factors [43, 44]. A significant difference was identified when the probability was less than 0.05. Statistical analyses were implemented in StatMate IV (ATMS Co., Ltd., Tokyo, Japan).
SUPPLEMENTARY FIGURES AND TABLES
Acknowledgments
The authors would like to thank Ms. Yuko Mohri for providing excellent technical support.
Footnotes
CONFLICTS OF INTEREST
None declared.
GRANT SUPPORT
A Grant-in-Aid for Scientific Research (No. 23592524, No. 24592594 and No. 25861485) from the Ministry of Education, Culture, Sports, Science, and Technology of Japan.
REFERENCES
- 1.Forastiere A, Koch W, Trotti A, Sidransky D. Head and neck cancer. N Engl J Med. 2001;345:1890–1900. doi: 10.1056/NEJMra001375. [DOI] [PubMed] [Google Scholar]
- 2.Choong N, Vokes E. Expanding role of the medical oncologist in the management of head and neck cancer. CA Cancer J Clin. 2008;58:32–53. doi: 10.3322/CA.2007.0004. [DOI] [PubMed] [Google Scholar]
- 3.Kanazawa T, Misawa K, Misawa Y, Uehara T, Fukushima H, Kusaka G, Maruta M, Carey TE. G-protein-coupled receptors: Next generation therapeutic targets in head and neck cancer? Toxins (Basel) 2015;7:2959–2984. doi: 10.3390/toxins7082959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Brock MV, Hooker CM, Ota-Machida E, Han Y, Guo M, Ames S, Glockner S, Piantadosi S, Gabrielson E, Pridham G, Pelosky K, Belinsky SA, Yang SC, et al. DNA methylation markers and early recurrence in stage i lung cancer. N Engl J Med. 2008;358:1118–1128. doi: 10.1056/NEJMoa0706550. [DOI] [PubMed] [Google Scholar]
- 5.Esteller M, Garcia-Foncillas J, Andion E, Goodman SN, Hidalgo OF, Vanaclocha V, Baylin SB, Herman JG. Inactivation of the DNA-repair gene mgmt and the clinical response of gliomas to alkylating agents. N Engl J Med. 2000;343:1350–1354. doi: 10.1056/NEJM200011093431901. [DOI] [PubMed] [Google Scholar]
- 6.Esteller M, Corn PG, Urena JM, Gabrielson E, Baylin SB, Herman JG. Inactivation of glutathione s-transferase p1 gene by promoter hypermethylation in human neoplasia. Cancer Res. 1998;58:4515–4518. [PubMed] [Google Scholar]
- 7.Heyn H, Esteller M. DNA methylation profiling in the clinic: Applications and challenges. Nat Rev Genet. 2012;13:679–692. doi: 10.1038/nrg3270. [DOI] [PubMed] [Google Scholar]
- 8.Dikshit RP, Gillio-Tos A, Brennan P, De Marco L, Fiano V, Martinez-Penuela JM, Boffetta P, Merletti F. Hypermethylation, risk factors, clinical characteristics, and survival in 235 patients with laryngeal and hypopharyngeal cancers. Cancer. 2007;110:1745–1751. doi: 10.1002/cncr.22975. [DOI] [PubMed] [Google Scholar]
- 9.Tan HK, Saulnier P, Auperin A, Lacroix L, Casiraghi O, Janot F, Fouret P, Temam S. Quantitative methylation analyses of resection margins predict local recurrences and disease-specific deaths in patients with head and neck squamous cell carcinomas. Br J Cancer. 2008;99:357–363. doi: 10.1038/sj.bjc.6604478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Rettori MM, de Carvalho AC, Bomfim Longo AL, de Oliveira CZ, Kowalski LP, Carvalho AL, Vettore AL. Prognostic significance of timp3 hypermethylation in post-treatment salivary rinse from head and neck squamous cell carcinoma patients. Carcinogenesis. 2013;34:20–27. doi: 10.1093/carcin/bgs311. [DOI] [PubMed] [Google Scholar]
- 11.Carvalho AL, Henrique R, Jeronimo C, Nayak CS, Reddy AN, Hoque MO, Chang S, Brait M, Jiang WW, Kim MM, Claybourne Q, Goldenberg D, Khan Z, et al. Detection of promoter hypermethylation in salivary rinses as a biomarker for head and neck squamous cell carcinoma surveillance. Clinical cancer research. 2011;17:4782–4789. doi: 10.1158/1078-0432.CCR-11-0324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Rettori MM, de Carvalho AC, Longo AL, de Oliveira CZ, Kowalski LP, Carvalho AL, Vettore AL. Timp3 and ccna1 hypermethylation in hnscc is associated with an increased incidence of second primary tumors. Journal of translational medicine. 2013;11:316. doi: 10.1186/1479-5876-11-316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Georges P, Rajagopalan K, Leon C, Singh P, Ahmad N, Nader K, Kubicek GJ. Chemotherapy advances in locally advanced head and neck cancer. World J Clin Oncol. 2014;5:966–972. doi: 10.5306/wjco.v5.i5.966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Okamoto M, Nishimine M, Kishi M, Kirita T, Sugimura M, Nakamura M, Konishi N. Prediction of delayed neck metastasis in patients with stage i/ii squamous cell carcinoma of the tongue. J Oral Pathol Med. 2002;31:227–233. doi: 10.1034/j.1600-0714.2002.310406.x. [DOI] [PubMed] [Google Scholar]
- 15.Kofler B, Laban S, Busch CJ, Lorincz B, Knecht R. New treatment strategies for hpv-positive head and neck cancer. Eur Arch Otorhinolaryngol. 2014;271:1861–1867. doi: 10.1007/s00405-013-2603-0. [DOI] [PubMed] [Google Scholar]
- 16.Feinberg AP, Tycko B. The history of cancer epigenetics. Nat Rev Cancer. 2004;4:143–153. doi: 10.1038/nrc1279. [DOI] [PubMed] [Google Scholar]
- 17.Lim SC, Zhang S, Ishii G, Endoh Y, Kodama K, Miyamoto S, Hayashi R, Ebihara S, Cho JS, Ochiai A. Predictive markers for late cervical metastasis in stage i and ii invasive squamous cell carcinoma of the oral tongue. Clinical cancer research. 2004;10:166–172. doi: 10.1158/1078-0432.ccr-0533-3. [DOI] [PubMed] [Google Scholar]
- 18.Sakamoto K, Imanishi Y, Tomita T, Shimoda M, Kameyama K, Shibata K, Sakai N, Ozawa H, Shigetomi S, Fujii R, Fujii M, Ogawa K. Overexpression of sip1 and downregulation of e-cadherin predict delayed neck metastasis in stage i/ii oral tongue squamous cell carcinoma after partial glossectomy. Ann Surg Oncol. 2012;19:612–619. doi: 10.1245/s10434-011-2052-1. [DOI] [PubMed] [Google Scholar]
- 19.Mori K, Enokida H, Kagara I, Kawakami K, Chiyomaru T, Tatarano S, Kawahara K, Nishiyama K, Seki N, Nakagawa M. Cpg hypermethylation of collagen type i alpha 2 contributes to proliferation and migration activity of human bladder cancer. Int J Oncol. 2009;34:1593–1602. doi: 10.3892/ijo_00000289. [DOI] [PubMed] [Google Scholar]
- 20.Lauvrak SU, Munthe E, Kresse SH, Stratford EW, Namlos HM, Meza-Zepeda LA, Myklebost O. Functional characterisation of osteosarcoma cell lines and identification of mrnas and mirnas associated with aggressive cancer phenotypes. Br J Cancer. 2013;109:2228–2236. doi: 10.1038/bjc.2013.549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Loss LA, Sadanandam A, Durinck S, Nautiyal S, Flaucher D, Carlton VE, Moorhead M, Lu Y, Gray JW, Faham M, Spellman P, Parvin B. Prediction of epigenetically regulated genes in breast cancer cell lines. BMC Bioinformatics. 2010;11:305. doi: 10.1186/1471-2105-11-305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Anderton JA, Lindsey JC, Lusher ME, Gilbertson RJ, Bailey S, Ellison DW, Clifford SC. Global analysis of the medulloblastoma epigenome identifies disease-subgroup-specific inactivation of col1a2. Neuro Oncol. 2008;10:981–994. doi: 10.1215/15228517-2008-048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Koga Y, Pelizzola M, Cheng E, Krauthammer M, Sznol M, Ariyan S, Narayan D, Molinaro AM, Halaban R, Weissman SM. Genome-wide screen of promoter methylation identifies novel markers in melanoma. Genome Res. 2009;19:1462–1470. doi: 10.1101/gr.091447.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Bonazzi VF, Nancarrow DJ, Stark MS, Moser RJ, Boyle GM, Aoude LG, Schmidt C, Hayward NK. Cross-platform array screening identifies col1a2, thbs1, tnfrsf10d and uchl1 as genes frequently silenced by methylation in melanoma. PLoS One. 2011;6:e26121. doi: 10.1371/journal.pone.0026121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Misawa K, Kanazawa T, Misawa Y, Imai A, Endo S, Hakamada K, Mineta H. Hypermethylation of collagen alpha2 (i) gene (col1a2) is an independent predictor of survival in head and neck cancer. Cancer Biomark. 2011;10:135–144. doi: 10.3233/CBM-2012-0242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Jin Z, Olaru A, Yang J, Sato F, Cheng Y, Kan T, Mori Y, Mantzur C, Paun B, Hamilton JP, Ito T, Wang S, David S, et al. Hypermethylation of tachykinin-1 is a potential biomarker in human esophageal cancer. Clinical cancer research. 2007;13:6293–6300. doi: 10.1158/1078-0432.CCR-07-0818. [DOI] [PubMed] [Google Scholar]
- 27.David S, Kan T, Cheng Y, Agarwal R, Jin Z, Mori Y. Aberrant silencing of the endocrine peptide gene tachykinin-1 in gastric cancer. Biochem Biophys Res Commun. 2009;378:605–609. doi: 10.1016/j.bbrc.2008.11.078. [DOI] [PubMed] [Google Scholar]
- 28.Mori Y, Cai K, Cheng Y, Wang S, Paun B, Hamilton JP, Jin Z, Sato F, Berki AT, Kan T, Ito T, Mantzur C, Abraham JM, et al. A genome-wide search identifies epigenetic silencing of somatostatin, tachykinin-1, and 5 other genes in colon cancer. Gastroenterology. 2006;131:797–808. doi: 10.1053/j.gastro.2006.06.006. [DOI] [PubMed] [Google Scholar]
- 29.Jeschke J, Van Neste L, Glockner SC, Dhir M, Calmon MF, Deregowski V, Van Criekinge W, Vlassenbroeck I, Koch A, Chan TA, Cope L, Hooker CM, Schuebel KE, et al. Biomarkers for detection and prognosis of breast cancer identified by a functional hypermethylome screen. Epigenetics. 2012;7:701–709. doi: 10.4161/epi.20445. [DOI] [PubMed] [Google Scholar]
- 30.Misawa K, Kanazawa T, Misawa Y, Imai A, Uehara T, Mochizuki D, Endo S, Takahashi G, Mineta H. Frequent promoter hypermethylation of tachykinin-1 and tachykinin receptor type 1 is a potential biomarker for head and neck cancer. J Cancer Res Clin Oncol. 2013;139:879–89. doi: 10.1007/s00432-013-1393-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kanazawa T, Iwashita T, Kommareddi P, Nair T, Misawa K, Misawa Y, Ueda Y, Tono T, Carey TE. Galanin and galanin receptor type 1 suppress proliferation in squamous carcinoma cells: Activation of the extracellular signal regulated kinase pathway and induction of cyclin-dependent kinase inhibitors. Oncogene. 2007;26:5762–5771. doi: 10.1038/sj.onc.1210384. [DOI] [PubMed] [Google Scholar]
- 32.Misawa K, Ueda Y, Kanazawa T, Misawa Y, Jang I, Brenner JC, Ogawa T, Takebayashi S, Grenman RA, Herman JG, Mineta H, Carey TE. Epigenetic inactivation of galanin receptor 1 in head and neck cancer. Clinical cancer research. 2008;14:7604–7613. doi: 10.1158/1078-0432.CCR-07-4673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Doufekas K, Hadwin R, Kandimalla R, Jones A, Mould T, Crowe S, Olaitan A, Macdonald N, Fiegl H, Wik E, Salvesen HB, Widschwendter M. Galr1 methylation in vaginal swabs is highly accurate in identifying women with endometrial cancer. Int J Gynecol Cancer. 2013;23:1050–1055. doi: 10.1097/IGC.0b013e3182959103. [DOI] [PubMed] [Google Scholar]
- 34.Guo S, Yan F, Xu J, Bao Y, Zhu J, Wang X, Wu J, Li Y, Pu W, Liu Y, Jiang Z, Ma Y, Chen X, et al. Identification and validation of the methylation biomarkers of non-small cell lung cancer (nsclc) Clin Epigenetics. 2015;7:3. doi: 10.1186/s13148-014-0035-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Misawa Y, Misawa K, Kanazawa T, Uehara T, Endo S, Mochizuki D, Yamatodani T, Carey TE, Mineta H. Tumor suppressor activity and inactivation of galanin receptor type 2 by aberrant promoter methylation in head and neck cancer. Cancer. 2014;120:205–213. doi: 10.1002/cncr.28411. [DOI] [PubMed] [Google Scholar]
- 36.Kristensen LS, Mikeska T, Krypuy M, Dobrovic A. Sensitive melting analysis after real time- methylation specific pcr (smart-msp): High-throughput and probe-free quantitative DNA methylation detection. Nucleic Acids Res. 2008;36:e42. doi: 10.1093/nar/gkn113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Jin Z, Cheng Y, Olaru A, Kan T, Yang J, Paun B, Ito T, Hamilton JP, David S, Agarwal R, Selaru FM, Sato F, Abraham JM, et al. Promoter hypermethylation of cdh13 is a common, early event in human esophageal adenocarcinogenesis and correlates with clinical risk factors. Int J Cancer. 2008;123:2331–2336. doi: 10.1002/ijc.23804. [DOI] [PubMed] [Google Scholar]
- 38.Martone T, Gillio-Tos A, De Marco L, Fiano V, Maule M, Cavalot A, Garzaro M, Merletti F, Cortesina G. Association between hypermethylated tumor and paired surgical margins in head and neck squamous cell carcinomas. Clinical cancer research. 2007;13:5089–5094. doi: 10.1158/1078-0432.CCR-07-0119. [DOI] [PubMed] [Google Scholar]
- 39.Lee EJ, Lee BB, Han J, Cho EY, Shim YM, Park J, Kim DH. Cpg island hypermethylation of e-cadherin (cdh1) and integrin alpha4 is associated with recurrence of early stage esophageal squamous cell carcinoma. Int J Cancer. 2008;123:2073–2079. doi: 10.1002/ijc.23598. [DOI] [PubMed] [Google Scholar]
- 40.Misawa K, Misawa Y, Kondo H, Mochizuki D, Imai A, Fukushima H, Uehara T, Kanazawa T, Mineta H. Aberrant methylation inactivates somatostatin and somatostatin receptor type 1 in head and neck squamous cell carcinoma. PLoS One. 2015;10:e0118588. doi: 10.1371/journal.pone.0118588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Gu J, Berman D, Lu C, Wistuba II, Roth JA, Frazier M, Spitz MR, Wu X. Aberrant promoter methylation profile and association with survival in patients with non-small cell lung cancer. Clinical cancer research. 2006;12:7329–7338. doi: 10.1158/1078-0432.CCR-06-0894. [DOI] [PubMed] [Google Scholar]
- 42.Toyooka S, Maruyama R, Toyooka KO, McLerran D, Feng Z, Fukuyama Y, Virmani AK, Zochbauer-Muller S, Tsukuda K, Sugio K, Shimizu N, Shimizu K, Lee H, et al. Smoke exposure, histologic type and geography-related differences in the methylation profiles of non-small cell lung cancer. Int J Cancer. 2003;103:153–160. doi: 10.1002/ijc.10787. [DOI] [PubMed] [Google Scholar]
- 43.Katz MH. Multivariable analysis: A practical guide for clinicians and public health researchers. Cambridge University Press, Cambridge. 2011 [Google Scholar]
- 44.Herman JG, Baylin SB. Gene silencing in cancer in association with promoter hypermethylation. N Engl J Med. 2003;349:2042–2054. doi: 10.1056/NEJMra023075. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


