Abstract
Classical Rett syndrome (RTT) is a neurodevelopmental disorder where most of cases carry MECP2 mutations. Atypical RTT variants involve mutations in CDKL5 and FOXG1. However, a subset of RTT patients remains that do not carry any mutation in the described genes. Whole exome sequencing was carried out in a cohort of 21 female probands with clinical features overlapping with those of RTT, but without mutations in the customarily studied genes. Candidates were functionally validated by assessing the appearance of a neurological phenotype in Caenorhabditis elegans upon disruption of the corresponding ortholog gene. We detected pathogenic variants that accounted for the RTT-like phenotype in 14 (66.6 %) patients. Five patients were carriers of mutations in genes already known to be associated with other syndromic neurodevelopmental disorders. We determined that the other patients harbored mutations in genes that have not previously been linked to RTT or other neurodevelopmental syndromes, such as the ankyrin repeat containing protein ANKRD31 or the neuronal acetylcholine receptor subunit alpha-5 (CHRNA5). Furthermore, worm assays demonstrated that mutations in the studied candidate genes caused locomotion defects. Our findings indicate that mutations in a variety of genes contribute to the development of RTT-like phenotypes.
Electronic supplementary material
The online version of this article (doi:10.1007/s00439-016-1721-3) contains supplementary material, which is available to authorized users.
Introduction
Rett syndrome (RTT, MIM 312750) is a postnatal progressive neurodevelopmental disorder (NDD), originally described in the 1960s by Andreas Rett (Rett 1966), that most frequently manifests itself in girls during early childhood, with an incidence of approximately 1 in 10,000 live births (Chahrour and Zoghbi 2007). RTT patients are asymptomatic during the first 6–18 months of life, but gradually develop severe motor, cognitive, and behavioral abnormalities that persist for life. It is the second most common cause of intellectual disability in females after Down’s syndrome (Chahrour and Zoghbi 2007). Around 90 % of the cases are explained by more than 800 reported mutations in the methyl CpG-binding protein 2 gene (MECP2) (RettBASE: MECP2 Variation Database) (Christodoulou et al. 2003), which is located in the X chromosome and which causes most of the classical or typical forms of RTT (Chahrour and Zoghbi 2007), and it was originally identified as encoding a protein that binds to methylated DNA (Lewis et al. 1992). Individuals affected by atypical or variant RTT present with many of the clinical features of RTT, but do not necessarily have all of the classic characteristics of the disorder (Neul et al. 2010). Approximately 8 % of classic RTT and 42 % of variant RTT patients are MECP2 mutation-negative (Monros et al. 2001; Percy 2008). Some of the latter group have mutations in other genes, such as that of the cyclin-dependent kinase-like 5 (CDKL5), which is described in individuals with an early seizure onset variant of RTT (Kalscheuer et al. 2003) or the forkhead box G1 (FOXG1), which is responsible for the congenital variant of RTT (Ariani et al. 2008). However, there remains a subset of patients with a clinical diagnosis of RTT who are mutation-negative for all the aforementioned genes. Next generation sequencing (NGS) has emerged as a potentially powerful tool for the study of such genetic diseases (Zhu et al. 2015).
Herein, we report the use of a family based exome sequencing approach in a cohort of 20 families with clinical features of RTT, but without mutations in the usually studied genes. We establish the neurological relevance of the newly identified candidate genes by assessing them in Caenorhabditis elegans model.
Materials and methods
Patient samples
A cohort of 19 Spanish parent–child trios and one family with two affected daughters who exhibited clinical features associated with RTT were recruited at Sant Joan de Deu Hospital in Barcelona, Catalonia, Spain. These patients had been diagnosed on the basis of the usual clinical parameters (Monros et al. 2001), and according to the recently revised RettSearch International Consortium criteria and nomenclature (Neul et al. 2010), but were found to be mutation-negative for MECP2, CDKL5 and FOXG1 in the original single-gene screening. The parents were clinically evaluated and it was not observed any evidence of intellectual disability. Genomic DNA from these patients was extracted from peripheral blood leukocytes using standard techniques, and analyzed by exome sequencing at the Cancer Epigenetics and Biology Program (PEBC) in Barcelona, Catalonia, Spain. Ethical approval for the molecular genetic studies was obtained from each institutional review board.
Whole exome sequencing and Sanger validation
Coding regions were captured using the TruSeq DNA Sample Preparation and Exome Enrichment Kit (Illumina, San Diego, California). Paired-end 100 × 2 sequences were sequenced with the Illumina HiScan SQ system at the National Center for Genomic Analysis in Barcelona. We also included the exome sequencing data of an MECP2, a CDKL5 and a JMJD1C (Sáez et al. 2016) RTT-associated family for data processing to improve the de novo single nucleotide variant calling. The complete exome sequencing data of all the studied samples are available from the Sequence Read Archive (http://www.ncbi.nlm.nih.gov/sra) with the ID: SRP073424 (private link for the reviewer until publication: http://www.ncbi.nlm.nih.gov/sra/SRP073424). The overall coverage statistics for each individual of the families, considering the regions captures using Exome Enrichment Kit, and number of reads in the position of the variation is shown in Supplementary Table 1. The identified variants were validated by Sanger sequencing using a BigDye® Terminator v3.1 Cycle Sequencing Kit in an Applied Biosystems 3730/DNA Analyzer. The raw data were analyzed with Codon Code Aligner Software. The primers used for Sanger sequencing are shown in Supplementary Table 2.
Caenorhabditis elegans handling
The techniques used for the culture of Caenorhabditis elegans were essentially as described (Brenner 1974). The worms were backcrossed at least three times to avoid background mutations. The behavior of three sets of ten animals was independently assessed in locomotion assays without food that were performed at 20 °C, as previously described (Sawin et al. 2000).
Results
Clinical criteria for selecting RTT trios
The 21 patients (derived from the 20 families studied) included in this study fulfilled the recently revised clinical criteria for the diagnosis of RTT following the usual clinical parameters (Monros et al. 2001), and the RettSearch International Consortium criteria and nomenclature (Neul et al. 2010). Specifically, all patients presented stereotypic hand movements, 90.5 % of them (19/21) showed microcephaly and also presented onset of the first signs of the disease before the age of 12 months. 66.7 % of patients (14/21) acquired motor skills, while a further seven (33.3 %), who had a more severe phenotype, never walked. Language skills were progressively lost in 28.6 % of the patients and 71.4 % of them (15/21) never acquired them. Additionally, important episodes of epilepsy were experienced by 81.0 % of the patients (17/21), and 57.1 % of them (12/21) manifested apneas and/or hyperventilation.
Bioinformatic process for filtering and selecting pathogenic variants
Before their inclusion in this study, patients underwent an extensive clinical and genetic work-up to detect genetic alterations in MECP2, CDKL5, and FOXG1. However, no molecular diagnosis could be established. We performed whole exome sequencing (WES) on the 61 individuals (20 pairs of healthy parents and 21 affected daughters) separately by subjecting whole blood derived genomic DNA to exome enrichment and sequencing. We focused our analysis on de novo single nucleotide variants (SNVs) due to their known relevance in autism and mental retardation-related diseases (Vissers et al. 2010). On average, WES gave rise to 419,045 variants, including SNVs and indels, of which 19,951 non-synonymous variants per family (4.7 %) were predicted to have a functional impact on the genomic sequence. To select variants that had not previously been described in the healthy population, we filtered out the variants with an allele frequency of 1 % or higher (the classic definition of a polymorphism) formerly observed in the Single Nucleotide Polymorphism database (dbSNP) and the 1000 Genomes Pilot Project data. Afterwards, to focus on de novo inheritance, patients’ variants were filtered first against variants found in their own parents and then against a pool of controls comprising all the healthy parents included in the study. Following this process, we achieved an average of 106 SNVs per family, which corresponded to 81 mutated genes per family. De novo candidate variants were selected on the basis of the quality of the alignments, damage score predictors and the conservation level of each of the genes during evolution. The complete exome sequencing data of all the studied samples are available from the Sequence Read Archive (http://www.ncbi.nlm.nih.gov/sra).
The global yield of genomic analysis following the bioinformatic process described herein enabled 22 coding de novo mutations to be identified in 66.7 % (14 of 21) of Rett-like patients: 20 SNVs and 2 indels. The identified variants and their de novo status were confirmed by conventional Sanger sequencing. Illustrative samples are shown in Fig. 1. Interestingly, in seven (33.3 %) of the studied RTT probands, exome sequencing did not detect any genetic change relative to their respective parents. The clinical characteristics of these seven patients without obvious pathogenic variants are summarized in Table 1. In one of the families, there were two affected children, and an analysis of potentially relevant recessive variants was performed. For the recessive analysis, and following the same criteria to define a variant as deleterious, we selected the variants with homozygous recessive genotype, and then at the gene level, we also selected the genes presenting more than one heterozygotic variant in the same gene (compound heterozygosity). We did not find any candidate gene consistent with the phenotype of the family with the two affected sisters.
Fig. 1.

Sanger sequencing validation of the de novo variants identified by exome sequencing. Illustrative examples for ZNF620 (c.856C > T p.Gly286Ser), GABBR2 (c.1699G > A p.Ala567Ser) and PDLIM7 (c.1222C > T p.Asp408Asn) are shown
Table 1.
Clinical summary of patients without exome candidates
| Proband | Age (years) | Onset of signs | Microcephaly | Sitting alone | Ambulation | Respiratory function | Epilepsy | Hand use | Stereotypies | Language | Total score |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 15 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 8 |
| 2 | 28 | 3 | 1 | 1 | 0 | 1 | 2 | 2 | 3 | 2 | 15 |
| 5.1 | 7 | 3 | 1 | 3 | 4 | 1 | 0 | 3 | 2 | 2 | 19 |
| 5.2 | 5 | 3 | 1 | 1 | 2 | 0 | 0 | 2 | 3 | 2 | 14 |
| 7 | 16 | 2 | 1 | 0 | 0 | 0 | 2 | 1 | 2 | 1 | 9 |
| 15 | 8 | 3 | 0 | 1 | 2 | 0 | 2 | 1 | 1 | 1 | 10 |
| 18 | 3.5 | 3 | 1 | 1 | 1 | 0 | 0 | 1 | 3 | 0 | 10 |
Clinical scores of our series of patients according to Pineda scale. Severity classification ranges from 0 to 4 as follows: age of onset of first signs (1: >24 months; 2: 12–24 months; 3: 0–12 months), microcephaly (0: absent; 1: present), sitting alone (1: acquired < 8 months; 2: seat and mantains; 3: seat and lost), ambulation (0: acquired < 18 months, 1: acquired < 30 months; 2: acquired > 30 months; 3: lost acquisition; 4: never acquired), respiratory function (0: no dysfunction; 1: hyperventilation and/or apnea), epilepsy (0: absent; 1: present and controlled; 2: uncontrolled or early epilepsy), hands use (0: acquired and conserved; 1: acquired and partially conserved; 2: acquired and lost; 3: never acquired), onset of stereotypies (1: > 10 years, 2: > 36 months; 3: 18-36 months) and languages (0: preserved and propositive; 1: lost; 2: never acquired). The total score is the sum of the scores of each clinical feature
Variants in genes previously associated with neurodevelopmental disorders
Of the 22 identified coding de novo mutations in the assessed RTT-like patients, five (22.7 %) occurred in genes previously associated with neurodevelopmental disorders that presented a clinicopathological phenotype with features coinciding with those of Rett syndrome (Table 2). In particular, we identified four mutations in genes such as HCN1 (Nava et al. 2014) and GRIN2B (Endele et al. 2010; Lemke et al. 2014), which are associated with early infantile epileptic encephalopathy; SLC6A1, which is associated with epilepsy and myoclonic-atonic seizures (Carvill et al. 2015); TCF4, which is associated with Pitt–Hopkins syndrome (Sweatt 2013); and SCN1A, which is associated with Dravet syndrome (Brunklaus and Zuberi 2014) (Table 2). The clinical characteristics of these five patients with variants in genes previously associated with neurodevelopmental phenotypes are summarized in Table 3. A comparison of the clinical features of our RTT-like patients, where we have identified mutations in candidate genes previously associated with other neurodevelopmental disorders, with those observed for these diseases is summarized in Table 4.
Table 2.
List of patients with variants found in genes previously associated with neurodevelopmental phenotypes
| Proband | Gene | Protein | NM number | Variant: genomic coordinates | cDNA change | Protein change | Gene-disease association |
|---|---|---|---|---|---|---|---|
| 4 | HCN1 | Hyperpolarization Activated Cyclic Nucleotide Gated Potassium Channel 1 | NM_021072.3 | 5:45396665 | c.1159G > T | p.Ala387Ser | Early infantile epileptic encephalopathy 24 |
| 8 | SCN1A | Sodium Channel Protein Type I Subunit Alpha | NM_001165963.1 | 2:166866266 | c.3965C > G | p.Arg1322Thr | Dravet syndrome |
| 10 | TCF4 | Transcription Factor 4 | NM_001243236.1 | 18:52901827 | c.958delC | p.Gln320Ser_fs8X | Pitt–Hopkins syndrome |
| 11 | GRIN2B | Glutamate receptor ionotropic, NMDA 2B | NM_000834.3 | 12:13764782 | c.1657C > A | p.Pro553Thr | Autosomal Dominant Mental Retardation 6; Early infantile epileptic encephalopathy 27 |
| 17 | SLC6A1 | Solute Carrier Family 6 Member 1 | NM_003042.3 | 3:11067528 | c.919G > A | p.Gly307Arg | Myoclonic-atonic epilepsy and schizophrenia |
Table 3.
Clinical summary of the patients with variants in genes previously associated with neurodevelopmental phenotypes
| Proband | Gene variant | Age (years) | Onset of the signs | Microcephaly | Sitting alone | Ambulation | Respiratory function | Epilepsy | Hand use | Stereotypies | Language | Total score |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 4 | HCN1 | 24 | 3 | 1 | 1 | 3 | 1 | 2 | 3 | 1 | 2 | 16 |
| 8 | SCN1A, MGRN1, BTBD9 | 7 | 3 | 1 | 1 | 4 | 1 | 2 | 1 | 3 | 2 | 18 |
| 10 | TCF4 | 16 | 3 | 1 | 1 | 2 | 1 | 2 | 1 | 1 | 2 | 14 |
| 11 | GRIN2B, SEMA6B | 3 | 2 | 0 | 1 | 3 | 0 | 1 | 1 | 1 | 2 | 10 |
| 17 | SLC6A1 | 36 | 3 | 0 | 1 | 1 | 0 | 1 | 1 | 1 | 0 | 7 |
Clinical scores of our series of patients according to Pineda scale. Severity classification ranges from 0 to 4 as follows: age of onset of first signs (1: > 24 months; 2: 12–24 months; 3: 0–12 months), microcephaly (0: absent; 1: present), sitting alone (1: acquired < 8 months; 2: seat and mantains; 3: seat and lost), ambulation (0: acquired < 18 months, 1: acquired < 30 months; 2: acquired > 30 months; 3: lost acquisition; 4: never acquired), respiratory function (0: no dysfunction; 1: hyperventilation and/or apnea), epilepsy (0: absent; 1: present and controlled; 2: uncontrolled or early epilepsy), hands use (0: acquired and conserved; 1: acquired and partially conserved; 2: acquired and lost; 3: never acquired), onset of stereotypies (1: > 10 years, 2: > 36 months; 3: 18-36 months) and languages (0: preserved and propositive; 1: lost; 2: never acquired). The total score is the sum of the scores of each clinical feature
Table 4.
Comparison of the clinical features of RTT-like patients from who we have identified mutations in candidate genes previously associated with other neurodevelopmental disorders with those observed for these diseases
| Disease | Rett | Atypical Rett | Pitt–Hopkins | Dravet | EEIE27 | MAE | EEIE24 | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GENE | MECP2 | CDKL5 | TCF4 | SCN1A | GRIN2B | SLC6A1 | HCN1 | |||||||
| OMIM/Patient | 312,750 | 6 | 308,350 | 16 | 602,272 | 10 | 182,390 | 8 | 616,139 | 11 | 616,421 | 17 | 615,871 | 4 |
| Onset age | 6–18 m | 12–24 m | 1–3 m | 0–12 m | 6–12 m | 0–12 m | 2 days–7 m | 0–12 m | 0–24 m | 12–24 m | 0–24 m | 0–12 m | 0–24 m | 0–12 m |
| Microcephaly | Yes | Yes | Yes | Yes | Yes | Yes | ± | Yes | ± | No | ± | No | NA | Yes |
| Hypotonia | Yes | Yes | Yes | Yes | Yes | Yes | No | Yes | ± | Yes | ± | Yes | NA | Yes |
| Epilepsy | 80 % | Yes | Yes | Yes | Yes | Yes | Yes | Yes | ± | Yes | Yes | Yes | Yes | Yes |
| Respiratory dysfunction | 80 % | Yes | Yes | Yes | Yes | Yes | Yes | Yes | No | No | No | No | NA | Yes |
| Expressive language dysfunction | Yes | Yes | Yes | Yes | Yes | Yes | ± | Yes | Yes | Yes | No | No | NA | Yes |
| Preserved use of hands | No | No | ± | No | No | No | Yes | No | ± | No | ± | No | NA | No |
| Stereotypies | Yes | Yes | Yes | Yes | Yes | Yes | ± | Yes | ± | Yes | ± | Yes | NA | Yes |
| Inheritance | XL | XL | AD | AD | AD | AD | AD | |||||||
EEIE epileptic encephalopathy, early infantile, MAE, myoclonic-atonic epilepsy, m months, XL X linkage, AD autosomal dominant, NA unavailable
Variants in genes previously not associated with neurodevelopmental disorders
Of the 22 identified coding de novo variants in the RTT-like patients assessed here, 17 (77.3 %) occurred in genes that had not previously been associated with neurodevelopmental disorders (Table 5). However, two of these variants were associated with non-neurodevelopmental disorders: a BTBD9 variant linked to restless leg syndrome (Kemlink et al. 2009), and an ATP8B1 SNV associated with familial cholestasis (Klomp et al. 2004), respectively. Interestingly, the BTBD9 variant was detected in the same patient that carried the SCN1A variant associated with Dravet syndrome (Table 2). The other 15 potentially pathogenic variants identified occurred in genes that had not been linked to any genetic disorder of any type. However, there was an enrichment of genes with a potential role in neuronal biology and functionality, such as the gamma-aminobutyric type B receptor subunit 2 (GABBR2), the neuronal acetylcholine receptor subunit alpha-5 (CHRNA5), the Huntington-associated protein 1 (HAP1), the axon guider semaphorin 6B, the ankyrin repeat containing proteins ANKRD31 and AGAP6, and the neuronal voltage-gated calcium channel CACNA1 (Table 5). Proband 14 was a particularly interesting case in which four potential pathogenic variants were present, affecting zinc finger (ZNF620), a nucleolar complex (NOC3L), G patch domain (GPATCH2) and GRAM domain (GRAMD1A)-related proteins (Table 5). The clinical characteristics of these patients with variants in genes previously not associated with neurodevelopmental disorders are summarized in Table 6.
Table 5.
List of patients with variants in new candidate disease genes
| Proband | Gene | Protein | Function | NM number | Variant: genomic coordinates | cDNA change | Protein change | ExAC | SIFT | Polyphen2 | PROVEAN | Mutation Taster2 | Conservation |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 3 | AGAP6 | ArfGAP with GTPase domain, ankyrin repeat and PH domain 6 | Putative GTPase-activating protein | NM_001077665.2 | 10:51748528 | c.53insC | p.Asp18Ala_fs10X | Not present | NA | NA | B | P | 405 |
| 8 | MGRN1 | Mahogunin RING Finger Protein 1 | E3 ubiquitin-protein ligase | NM_001142290.2 | 16:4723583 | c.880C > T | p.Arg294Cys | 0.000077 | P | P | P | P | 573 |
| 8 | BTBD9 | BTB (POZ) Domain-Containing 9 | Putative protein–protein interactor | NM_001099272.1 | 6:38256093 | c.1409C > T | p.Ala470Val | Not present | B | P | P | P | 512 |
| 11 | SEMA6B | Semaphorin-6B | Role in axon guidance | NM_032108.3 | 19:4555540 | c.508G > A | p.Gly170Ser | Not present | P | P | P | P | 510 |
| 12 | VASH2 | Vasohibin 2 | Angiogenesis inhibitor | NM_001301056.1 | 1:213161902 | c.1044A > C | p.Glu348Asp | Not present | B | B | B | B | 473 |
| 13 | CHRNA5 | Neuronal acetylcholine receptor subunit alpha-5 | Excitator of neuronal activity | NM_000745.3 | 15:78882481 | c.748C > A | p.Pro250Thr | Not present | B | P | P | P | 519 |
| 14 | ZNF620 | Zinc Finger Protein 620 | Transcriptional regulator | NM_175888.3 | 3:40557941 | c.856G > A | p.Gly286Ser | Not present | P | P | P | P | 317 |
| 14 | GRAMD1A | GRAM Domain-Containing 1A | Not described | NM_020895.3 | 19:35506764 | c.1106G > A | p.Arg369His | Not present | P | P | P | P | 358 |
| 14 | NOC3L | Nucleolar complex protein 3 homolog | Regulator of adipogenesis | NM_022451.10 | 10:96097586 | c.2137G > A | p.Ala713Thr | Not present | B | B | B | B | 0 |
| 14 | GPATCH2 | G patch domain-containing protein 2 | Regulator of cell proliferation | NM_018040.3 | 1:217784371 | c.878G > A | p.Gly293Asp | Not present | B | P | P | P | 304 |
| 19 | GABBR2 | Gamma-aminobutyric acid type B receptor subunit 2 | Inhibitor of neuronal activity | NM_005458.7 | 9:101133817 | c.1699G > A | p.Ala567Thr | Not present | P | P | P | P | 412 |
| 19 | ATP8B1 | Phospholipid-transporting ATPase IC | Aminophospholipid translocator | NM_005603.4 | 18:55328507 | c.2606C > T | p.Thr869Ile | Not present | P | P | P | P | 361 |
| 20 | HAP1 | Huntingtin-Associated Protein 1 | Vesicular transporter | NM_177977.2 | 17:39890655 | c.232G > A | p.Ala78Thre | Not present | P | B | B | B | 0 |
| 21 | PDLIM7 | PDZ and LIM domain protein 7 | Scaffold protein | NM_005451.4 | 5:176910933 | c.1222G > A | p.Asp408Asn | Not present | P | P | B | P | 515 |
| 21 | SRRM3 | Serine/Arginine Repetitive Matrix 3 | Splicing activator | NM_001291831.1 | 7:75890878 | c.655C > G | p.Ser218Cys | Not present | P | P | P | P | 491 |
| 22 | ANKRD31 | Ankyrin Repeat Domain 31 | Not described | NM_001164443.1 | 5:74518166 | c.196A > T | p.Ile66Phe | Not present | P | P | B | B | 401 |
| 23 | CACNA1I | Voltage-Gated Calcium Channel Subunit Alpha 1I | Calcium signaling in neurons | NM_021096.3 | 22:40066855 | c.4435C > T | p.Leu1479Phe | Not present | B | P | B | P | 695 |
ExAC, frequency of the identified variants in the exome aggregation consortium. Four in silico prediction tools of functional mutation impact were used: ‘Sorting Tolerant From Intolerant’ (SIFT), ‘Polymorphism Phenotyping v2’ (Polyphen2); ‘Protein Variation Effect Analyzer’ (PROVEAN) and Mutation Taster2. The output results were classified as: likely pathogenic (P), likely benign (B) and not available (NA). Conservation scores refer to the conservation level of the nucleotide at the position of the identified variant between 46 species of vertebrates based on PhastCons. It ranges from 0 to 1000: the highest, the more conserved during evolution
Table 6.
Clinical summary of patients with variants in new candidate disease genes
| Proband | Gene variant | Age (years) | Onset of the signs | Microcephaly | Sitting alone | Ambulation | Respiratory function | Epilepsy | Hand use | Stereotypies | Language | Total score |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 3 | AGAP6 | 14 | 3 | 1 | 2 | 4 | 1 | 1 | 3 | 2 | 2 | 19 |
| 8 | SCN1A, MGRN1, BTBD9 | 7 | 3 | 1 | 1 | 4 | 1 | 2 | 1 | 3 | 2 | 18 |
| 11 | GRIN2B, SEMA6B | 3 | 2 | 0 | 1 | 3 | 0 | 1 | 1 | 1 | 2 | 10 |
| 12 | VASH2 | 11 | 3 | 1 | 1 | 4 | 1 | 0 | 2 | 2 | 1 | 15 |
| 13 | CHRNA5 | 10 | 3 | 1 | 2 | 4 | 1 | 1 | 2 | 3 | 2 | 19 |
| 14 | ZNF620, GRAMD1A, NOC3L, GPATCH2 | 2 | 3 | 1 | 1 | 2 | 0 | 1 | 3 | 2 | 2 | 15 |
| 19 | GABBR2, ATP8B1 | 2 | 3 | 1 | 1 | 4 | 0 | 0 | 3 | 2 | 2 | 16 |
| 20 | HAP1 | 24 | 3 | 1 | 1 | 1 | 1 | 0 | 2 | 2 | 1 | 12 |
| 21 | PDLIM7, SRRM3 | 5 | 3 | 1 | 3 | 4 | 0 | 1 | 2 | 1 | 2 | 17 |
| 22 | ANKRD31 | 17 | 3 | 0 | 1 | 2 | 0 | 1 | 2 | 3 | 2 | 14 |
| 23 | CACNA1I | 1/8 | 3 | 1 | 1 | 2 | 1 | 0 | 3 | 3 | 2 | 16 |
Clinical scores of our series of patients according to Pineda scale. Severity classification ranges from 0 to 4 as follows: age of onset of first signs (1: >24 months; 2: 12–24 months; 3: 0–12 months), microcephaly (0: absent; 1: present), sitting alone (1: acquired < 8 months; 2: seat and mantains; 3: seat and lost), ambulation (0: acquired < 18 months, 1: acquired < 30 months; 2: acquired > 30 months; 3: lost acquisition; 4: never acquired), respiratory function (0: no dysfunction; 1: hyperventilation and/or apnea), epilepsy (0: absent; 1: present and controlled; 2: uncontrolled or early epilepsy), hands use (0: acquired and conserved; 1: acquired and partially conserved; 2: acquired and lost; 3: never acquired), onset of stereotypies (1: > 10 years, 2: > 36 months; 3: 18–36 months) and languages (0: preserved and propositive; 1: lost; 2: never acquired). The total score is the sum of the scores of each clinical feature
Neurological phenotype of candidate genes in C. elegans
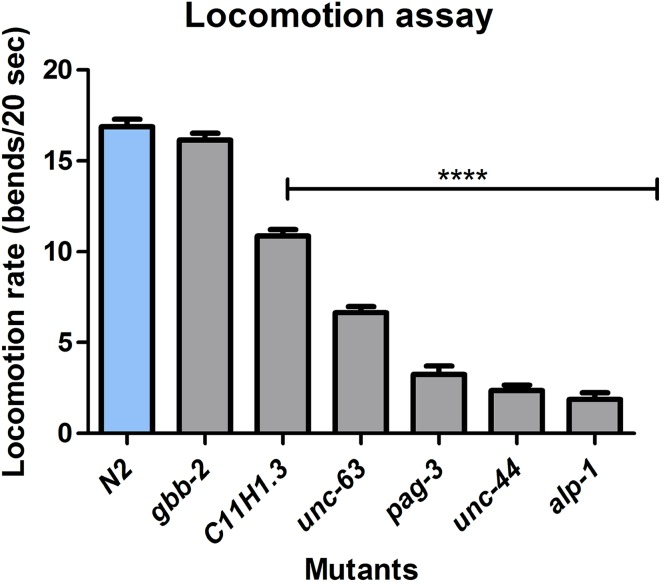
To demonstrate a neurological effect for a loss of function of the detected genes that had not previously been associated with neurodevelopmental disorders (Table 5), we used the model organism C. elegans to confirm the genotype-phenotype correlation. We obtained all the available C. elegans mutants that carry deleterious mutations in the orthologous genes to those human genes with potentially pathogenic mutations in the patients. In this model, backcrossing is a commonly used procedure to obtain a specific mutant strain without any secondary mutations from its genetic composition. Under these conditions, we were able to test six available mutant strains that were backcrossed at least three times to prove that any observed phenotype was really associated to specific mutations in the orthologous genes. To this end, we studied the C. elegans mutants carrying deleterious mutations in the gene orthologs of the human genes PDLIM7, ANKRD31, ZNF620, CHRNA5, MGRN1 and GABBR2 described in Table 7. Considering that the loss of normal movement and coordination is one of the clearest signs shown by Rett patients, we performed a locomotion assay of the nematodes as previously described (Sawin et al. 2000), using the wild-type N2 strain as a control (Supplementary Video 1). We observed that in 83.3 % (5 of 6) of the cases the mutation of the ortholog of the human exome sequencing identified genes in C. elegans exhibited a locomotion defective phenotype (Fig. 2). The most severe phenotypes were represented by alp-1, unc-44 and pag-3, with mutations in the orthologs of PDZ and LIM domain protein 7 (PDLIM7), ankyrin repeat containing protein ANKRD31 and the zinc protein ZNF620, respectively (Fig. 2 and Supplementary Videos 2, 3 and 4). The case of alp-1 was particularly interesting, because mutant worms were not only thinner than usual and completely locomotion defective, but they exhibited transitory spasms. Significant defects, such as slower locomotion and uncoordinated movement, were also observed in the mutants of unc-63 and C11H1.3, the C. elegans orthologs of the genes coding for the neuronal acetylcholine receptor subunit alpha-5 (CHRNA5) and mahogunin RING finger protein 1 (MGRN1), respectively. Although we did not find a clear locomotion defect in the gbb-2 mutant (the ortholog of GABBR2) (Fig. 2), it occurs in the gbb-1;gbb-2 double mutant (Dittman and Kaplan 2008), gbb-1 being the C. elegans ortholog of GABBR1 (gamma-aminobutyric acid type B receptor subunit 1). The clinical picture of the particular RTT cases with mutations in the genes studied in C. elegans is shown in Table 6.
Table 7.
Phenotype in Caenorhabditis elegans
| Human gene | Ortholog in C.elegans | Similarity (%) | Identity (%) | Mutation in C.elegans | Locomotion phenotype | Neurological phenotypes | Other phenotypes |
|---|---|---|---|---|---|---|---|
| GABBR2 | gbb-2 | 53 | 34 | Deletion | normal | Hypersensitivity to aldicarb | – |
| MGRN1 | C11H1.3 | 58 | 41 | Deletion | locomotion defective | – | – |
| CHRNA5 | unc-63 | 58 | 40 | Deletion | locomotion defective | Uncoordinated locomotion with strong levamisole resistance | – |
| ZNF620 | pag-3 | 65 | 47 | Deletion | locomotion defective | Altered neurosecretion and up-regulation of DCV (Dense Core Vesicles) components | – |
| ANKRD31 | unc-44 | 59 | 39 | Deletion | locomotion defective | Asymmetric dynamics of axonal and dendritic microtubules defects | – |
| PDLIM7 | alp-1 | 65 | 47 | Deletion | locomotion defective | – | Defects in actin filament organization in muscle cells |
Fig. 2.
Locomotion assay in Caenorhabditis elegans. Functional validation of mutations was performed by measuring the locomotion rate, expressed in average of measuring, in C. elegans. Each mutant strain was compared to a wild-type N2 control strain by measuring worm body bends during 20 s in three independent sets of experiments. Locomotion rates of mutants, represented by C11H1.3 (MGRN1), unc-63 (CHRNA5), pag-3 (ZNF620), unc-44 (ANKRD31) and alp-1 (PDLIM7) are significantly lower compared to that of the N2 control strain (p < 0.0001), on the contrary gbb-2 (GABBR2) mutant move similarly. Standard error of the mean (SEM) values is shown. p values obtained according to Student’s t test. ***p < 0.0001
Discussion
Our results indicate that the existence of de novo variants in genes with potential neurological functionalities, such as neuronal receptors (GABBR2 and CHRNA5), axon guiders (SEMA6B), synaptic ionic channels (CACNA1I) and others, contribute to the development of RTT-like clinical phenotypes in the context of wild-type sequences for standard Rett genes such as MECP2 and FOXG1. These patients share most of the clinicopathological features of classic RTT syndrome, such as stereotypic hand movements, relative microcephaly, and onset of the disease after the age of 12 months. Thus, exome sequencing is a powerful tool for genetically characterizing these enigmatic cases. In this regard, once a new candidate gene has been identified, it is now possible to design specific sequencing strategies for the molecular screening of this particular target in larger populations of patients with intellectual disability. The strategy based on exome sequencing patients who have RTT features, but no known mutations in the usual genes, has recently been used in other smaller series of patients (Grillo et al. 2013; Okamoto et al. 2015; Hara et al. 2015; Olson et al. 2015; Lopes et al. 2016). Most importantly, our study and the aforementioned previous reports strengthen the concept that a mutational heterogeneous profile hitting shared neurological signaling pathways contributes to RTT-like syndromes. Examples of confluence in the same molecular crossroads include the gamma-aminobutyric type B receptor subunit 2 (GABBR2) de novo variant, described here, and the formerly identified variant in the gamma-aminobutyric acid receptor delta gene (GABRD) (Hara et al. 2015). Interestingly, a second RTT-like patient has been identified as being a carrier of a de novo GABBR2 variant (Lopes et al. 2016), highlighting the likelihood that this gene and pathway contribute to the clinical entity. Another example of similarly targeted genes in RTT-like patients is that of the proteins containing ankyrin-repeats that are involved in postsynaptic density (Durand et al. 2007). This study has revealed de novo variants in the ankyrin repeat containing proteins AGAP6 and ANKRD31 in RTT-like patients, and the presence of de novo variant of the SH3 and multiple ankyrin repeat domain3 protein (SHANK3) (Hara et al. 2015) and ankyrin-3 (ANK3) (Grillo et al. 2013) has been reported in two RTT-like patients. A final example of the convergence of cellular pathways to provide a common RTT-like phenotype is represented by the disruption of the ionic channels. We found the existence of a voltage-gated calcium channel subunit alpha 11 (CANA1I) de novo variant in an RTT-like patient. Additionally, the presence of de novo variants in the calcium release channel RYR1 (Grillo et al. 2013) and the sodium voltage-gated channel alpha subunit 2 (SCN2A) (Baasch et al. 2014) in two other RTT-like probands have been reported. It is also intriguing that in our study a variant in HAP was found, whereas in similar series heterozygous variants in huntingtin (HTT) have been described (Lopes et al. 2016; Rodan et al. 2016), further reinforcing the links between Huntington’s disease and Rett syndrome (Roux et al. 2012). Another interesting case is provided by TCF4, which is associated with Pitt–Hopkins syndrome (Sweatt 2013), where in addition to our study, others have found mutations in RTT-like patients (Lopes et al. 2016). This observation could be of interest for clinicians due to phenotypic similitudes such as intellectual disability, stereotypic movement, apneas and seizures (Marangi et al. 2012).
Our findings also suggest that a substantial degree of clinical overlap can exist between the features associated with RTT and those of other neurodevelopmental disorders. Our exome sequencing effort indicated that probands originally diagnosed as RTT-like patients were, in fact, carriers of well-known pathogenic de novo mutations linked to Dravet Syndrome (SCN1A), myoclonic-atonic epilepsy (SCLC6A1), or early infantile epileptic encephalopathies 24 (HCN1) and 27 (GRIN2B). The purely clinical classification of these patients, without a thorough genetic study, can be difficult because some of these patients are composites that carry at least two pathogenic variants. For example, in our cases, the Dravet syndrome patient also had a de novo variant in BTBD9 associated with the development of restless leg syndrome. In addition, among the newly identified candidate genes associated with RTT-like features, a few of these patients simultaneously carried two de novo variants (e.g., probands 8, 19 and 21), further complicating the tasks of correctly diagnosing and managing these individuals.
Finally, the studies performed in C. elegans validate the functional relevance for nervous system function of the newly proposed candidate genes. Future studies would be necessary to assess the role of the specific variants identified, such as rescuing the defects with the expression of normal cDNAs versus cDNAs containing the mutation, ideally using cDNAs of human origin to prove similar function of the gene in the two species. It is also relevant to mention that for some of the newly reported mutated genes in our RTT-like patients, there are mice models targeting the described loci that show neurological phenotypes such as BTBD9 (motor restlessness and sleep disturbances) (DeAndrade et al. 2012), MGRN1 (spongiform neurodegeneration) (He et al. 2003), SEMA6B (aberrant mossy fibers) (Tawarayama et al. 2010), CHRNA5 (alterations in the habenulo-interpeduncular pathway) (Fowler et al. 2011), GABBR2 (anxiety and depression-related behavior) (Mombereau et al. 2005) and HAP1 (depressive-like behavior and reduced hippocampal neurogenesis) (Chan et al. 2002; Xiang et al. 2015).
Conclusions
Overall, this study demonstrates the genetic mutational diversity that underlies the clinical diagnosis of patients with clinical features that resemble RTT cases. Once the recognized MECP2, CDKL5 and FOXG1 mutations have been discarded, exome sequencing emerges as a very useful strategy for the more accurate classification of these patients. The de novo variants identified by this approach can modify the first diagnostic orientation towards another neurodevelopmental disorder, or pinpoint new genes involved in the onset of RTT-like features. Interestingly, most of these new targets are involved in the same functional networks associated with correct neuronal functionality. Further research is required to understand the role of these proteins in the occurrence of neurodevelopmental diseases. Additional functional experiments, such as the C. elegans assays used in this study, would be extremely helpful for this purpose.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Acknowledgments
The research leading to these results has received funding from the People Programme (Marie Curie Actions) of the European Union’s Seventh Framework Programme FP7/2012 under REA Grant agreement PITN-GA-2012-316758 of the EPITRAIN project and PITN-GA-2009–238242 of DISCHROM; the E-RARE EuroRETT network (Carlos III Health Institute project PI071327); the Foundation Lejeune (France); the Cellex Foundation; the Botín Foundation; the Finestrelles Foundation; the Catalan Association for Rett Syndrome; and the Health and Science Departments of the Catalan government (Generalitat de Catalunya). M.E. is an ICREA Research Professor.
Compliance with ethical statement
Conflict of interest
The authors declare that there is no conflict of interest associated with this manuscript.
Contributor Information
Paola Jurado, Email: pjurado@idibell.cat.
Judith Armstrong, Email: jarmstrong@hsjdbcn.org.
Manel Esteller, Email: mesteller@idibell.cat.
References
- Ariani F, et al. FOXG1 is responsible for the congenital variant of Rett syndrome. Am J Hum Genet. 2008;83:89–93. doi: 10.1016/j.ajhg.2008.05.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baasch AL, et al. Exome sequencing identifies a de novo SCN2A mutation in a patient with intractable seizures, severe intellectual disability, optic atrophy, muscular hypotonia, and brain abnormalities. Epilepsia. 2014;55:e25–e29. doi: 10.1111/epi.12554. [DOI] [PubMed] [Google Scholar]
- Brenner S. The genetics of Caenorhabditis elegans. Genetics. 1974;77:71–94. doi: 10.1093/genetics/77.1.71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brunklaus A, Zuberi SM. Dravet syndrome–from epileptic encephalopathy to channelopathy. Epilepsia. 2014;55:979–984. doi: 10.1111/epi.12652. [DOI] [PubMed] [Google Scholar]
- Carvill GL, et al. Mutations in the GABA transporter SLC6A1 cause epilepsy with myoclonic-atonic seizures. Am J Hum Genet. 2015;96:808–815. doi: 10.1016/j.ajhg.2015.02.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chahrour M, Zoghbi HY. The story of Rett syndrome: from clinic to neurobiology. Neuron. 2007;56:422–437. doi: 10.1016/j.neuron.2007.10.001. [DOI] [PubMed] [Google Scholar]
- Chan EY, et al. Targeted disruption of Huntingtin-associated protein-1 (Hap1) results in postnatal death due to depressed feeding behavior. Hum Mol Genet. 2002;11:945–959. doi: 10.1093/hmg/11.8.945. [DOI] [PubMed] [Google Scholar]
- Christodoulou J, Grimm A, Maher T, Bennetts B. RettBASE: the IRSA MECP2 Variation Database—a new mutation database in evolution. Hum Mutat. 2003;21:466–472. doi: 10.1002/humu.10194. [DOI] [PubMed] [Google Scholar]
- DeAndrade MP, et al. Motor restlessness, sleep disturbances, thermal sensory alterations and elevated serum iron levels in Btbd9 mutant mice. Hum Mol Genet. 2012;21:3984–3992. doi: 10.1093/hmg/dds221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dittman JS, Kaplan JM. Behavioral impact of neurotransmitter-activated G-protein-coupled receptors: muscarinic and GABAB receptors regulate Caenorhabditis elegans locomotion. J Neurosci. 2008;28:7104–7112. doi: 10.1523/JNEUROSCI.0378-08.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Durand CM, et al. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet. 2007;39:25–27. doi: 10.1038/ng1933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Endele S, et al. Mutations in GRIN2A and GRIN2B encoding regulatory subunits of NMDA receptors cause variable neurodevelopmental phenotypes. Nat Genet. 2010;42:1021–1026. doi: 10.1038/ng.677. [DOI] [PubMed] [Google Scholar]
- Fowler CD, Lu Q, Johnson PM, Marks MJ, Kenny PJ. Habenular α5 nicotinic receptor subunit signalling controls nicotine intake. Nature. 2011;471:597–601. doi: 10.1038/nature09797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grillo E, et al. Revealing the complexity of a monogenic disease: Rett syndrome exome sequencing. PLoS One. 2013;8:e56599. doi: 10.1371/journal.pone.0056599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hara M, Ohba C, Yamashita Y, Saitsu H, Matsumoto N, Matsuishi T. De novo SHANK3 mutation causes Rett syndrome-like phenotype in a female patient. Am J Med Genet A. 2015;167:1593–1596. doi: 10.1002/ajmg.a.36775. [DOI] [PubMed] [Google Scholar]
- He L, et al. Spongiform degeneration in mahoganoid mutant mice. Science. 2003;299:710–712. doi: 10.1126/science.1079694. [DOI] [PubMed] [Google Scholar]
- Kalscheuer VM, et al. Disruption of the serine/threonine kinase 9 gene causes severe X-linked infantile spasms and mental retardation. Am J Hum Genet. 2003;72:1401–1411. doi: 10.1086/375538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kemlink D, et al. Replication of restless legs syndrome loci in three European populations. J Med Genet. 2009;46:315–318. doi: 10.1136/jmg.2008.062992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klomp LW, et al. Characterization of mutations in ATP8B1 associated with hereditary cholestasis. Hepatology. 2004;40:27–38. doi: 10.1002/hep.20285. [DOI] [PubMed] [Google Scholar]
- Lemke JR, et al. GRIN2B mutations in West syndrome and intellectual disability with focal epilepsy. Ann Neurol. 2014;75:147–154. doi: 10.1002/ana.24073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis JD, Meehan RR, Henzel WJ, Maurer-Fogy I, Jeppesen P, Klein F, Bird A. Purification, sequence, and cellular localization of a novel chromosomal protein that binds to methylated DNA. Cell. 1992;69:905–914. doi: 10.1016/0092-8674(92)90610-O. [DOI] [PubMed] [Google Scholar]
- Lopes F, et al. Identification of novel genetic causes of Rett syndrome-like phenotypes. J Med Genet. 2016;53:190–199. doi: 10.1136/jmedgenet-2015-103568. [DOI] [PubMed] [Google Scholar]
- Marangi G, et al. Proposal of a clinical score for the molecular test for Pitt-Hopkins syndrome. Am J Med Genet A. 2012;158A:1604–1611. doi: 10.1002/ajmg.a.35419. [DOI] [PubMed] [Google Scholar]
- Mombereau C, Kaupmann K, Gassmann M, Bettler B, van der Putten H, Cryan JF. Altered anxiety and depression-related behaviour in mice lacking GABAB(2) receptor subunits. NeuroReport. 2005;16:307–310. doi: 10.1097/00001756-200502280-00021. [DOI] [PubMed] [Google Scholar]
- Monrós E, Armstrong J, Aibar E, Poo P, Canós I, Pineda M. Rett syndrome in Spain: mutation analysis and clinical correlations. Brain Dev. 2001;23:S251–S253. doi: 10.1016/S0387-7604(01)00374-6. [DOI] [PubMed] [Google Scholar]
- Nava C, et al. De novo mutations in HCN1 cause early infantile epileptic encephalopathy. Nat Genet. 2014;46:640–645. doi: 10.1038/ng.2952. [DOI] [PubMed] [Google Scholar]
- Neul JL, et al. Rett syndrome: revised diagnostic criteria and nomenclature. Ann Neurol. 2010;68:944–950. doi: 10.1002/ana.22124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okamoto N, et al. Targeted next-generation sequencing in the diagnosis of neurodevelopmental disorders. Clin Genet. 2015;88:288–292. doi: 10.1111/cge.12492. [DOI] [PubMed] [Google Scholar]
- Olson HE, et al. Mutations in epilepsy and intellectual disability genes in patients with features of Rett syndrome. Am J Med Genet A. 2015;167A:2017–2025. doi: 10.1002/ajmg.a.37132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Percy AK. Rett syndrome: recent research progress. J Child Neurol. 2008;23:543–549. doi: 10.1177/0883073807309786. [DOI] [PubMed] [Google Scholar]
- Rett A. On an unusual brain atrophy syndrome in hyperammonemia in childhood. Wien Med Wochenschr. 1966;116:723–772. [PubMed] [Google Scholar]
- Rodan LH, et al. A novel neurodevelopmental disorder associated with compound heterozygous variants in the huntingtin gene. Eur J Hum Genet. 2016 doi: 10.1038/ejhg.2016.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roux JC, Zala D, Panayotis N, Borges-Correia A, Saudou F, Villard L. Modification of Mecp2 dosage alters axonal transport through the Huntingtin/Hap1 pathway. Neurobiol Dis. 2012;45:786–795. doi: 10.1016/j.nbd.2011.11.002. [DOI] [PubMed] [Google Scholar]
- Sáez MA, et al. Mutations in JMJD1C are involved in Rett syndrome and intellectual disability. Genet Med. 2016;18:378–385. doi: 10.1038/gim.2015.100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawin ER, Ranganathan R, Horvitz HR. C. elegans locomotory rate is modulated by the environment through a dopaminergic pathway and by experience through a serotonergic pathway. Neuron. 2000;26:619–631. doi: 10.1016/S0896-6273(00)81199-X. [DOI] [PubMed] [Google Scholar]
- Sweatt JD. Pitt-Hopkins syndrome: intellectual disability due to loss of TCF4-regulated gene transcription. Exp Mol Med. 2013;45:e21. doi: 10.1038/emm.2013.32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tawarayama H, Yoshida Y, Suto F, Mitchell KJ, Fujisawa H. Roles of semaphorin-6B and plexin-A2 in lamina-restricted projection of hippocampal mossy fibers. J Neurosci. 2010;30:7049–7060. doi: 10.1523/JNEUROSCI.0073-10.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vissers LE, et al. A de novo paradigm for mental retardation. Nat Genet. 2010;42:1109–1112. doi: 10.1038/ng.712. [DOI] [PubMed] [Google Scholar]
- Xiang J, Yan S, Li SH, Li XJ. Postnatal loss of hap1 reduces hippocampal neurogenesis and causes adult depressive-like behavior in mice. PLoS Genet. 2015;11:e1005175. doi: 10.1371/journal.pgen.1005175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu X, et al. Whole-exome sequencing in undiagnosed genetic disorders: interpreting 119 trios. Genet Med. 2015;17:774–781. doi: 10.1038/gim.2014.191. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.