Abstract
During cell division, accurate chromosome segregation is tightly regulated by Polo-like kinase 1 (PLK1) and opposing activities of Aurora B kinase and protein phosphatase 1 (PP1). However, the regulatory mechanisms underlying the aforementioned hierarchical signaling cascade during mitotic chromosome segregation have remained elusive. Sds22 is a conserved regulator of PP1 activity, but how it regulates PP1 activity in space and time during mitosis remains elusive. Here we show that Sds22 is a novel and cognate substrate of PLK1 in mitosis, and the phosphorylation of Sds22 by PLK1 elicited an inhibition of PP1-mediated dephosphorylation of Aurora B at threonine 232 (Thr232) in a dose-dependent manner. Overexpression of a phosphomimetic mutant of Sds22 causes a dramatic increase in mitotic delay, whereas overexpression of a non-phosphorylatable mutant of Sds22 results in mitotic arrest. Mechanistically, the phosphorylation of Sds22 by PLK1 strengthens the binding of Sds22 to PP1 and inhibits the dephosphorylation of Thr232 of Aurora B to ensure a robust, error-free metaphase-anaphase transition. These findings delineate a conserved signaling hierarchy that orchestrates dynamic protein phosphorylation and dephosphorylation of critical mitotic regulators during chromosome segregation to guard chromosome stability.
Keywords: cell cycle, centromere, cyclin-dependent kinase (CDK), cytoskeleton, kinetochore, phosphatase, phosphorylation, tubulin
Introduction
During mitosis, deregulation of the faithful chromosomal segregation will result in aneuploidy of daughter cells, which further promotes chromosomal instability and tumorigenesis (1). To achieve faithful chromosome segregation during cell division, the kinetochores of replicated chromatids need to attach to opposite poles of the mitotic spindle. Bipolar attachment of spindle microtubules to kinetochores is monitored by the spindle assembly checkpoint (SAC)6 and dynamically regulated by phosphorylation and dephosphorylation to allow correction of improper attachment and stabilization of correct attachments (2–4). Key elements of this regulation are PLK1, Aurora B, and PP1/Sds22, whose opposing activities need to be tightly balanced (5–7).
Aurora B is the catalytic subunit of chromosome passenger complex, which regulates chromosome condensation, correction of erroneous kinetochore-microtubule attachments, activation of the SAC, and cytokinesis (8). During mitotic spindle assembly, centromere-localized Aurora B destabilizes kinetochore-microtubule interactions that are under low mechanical tension (2, 9) by phosphorylating several components of the outer kinetochore, including NDC80/Hec1, KNL-1, and Dsn1 (3, 10). The Aurora B-mediated error correction is important for bipolar attachment of all chromosomes, which satisfies the SAC to initiate anaphase (11, 12).
PLK1 is a critical serine/threonine kinase that regulates multiple stages of mitosis (13, 14). PLK1 controls multiple events in mitosis such as mitotic entry, centrosome maturation, bipolar spindle formation, stable microtubule-kinetochore attachment, cohesion dissociation, chromosome alignment and segregation, and cytokinesis (14). PLK1 is overexpressed in many human tumors and is associated with tumorigenesis (15). It has been recently reported that PLK1 phosphorylates the N terminus of Haspin, leading to its activation and resulting in the phosphorylation of the threonine 3 on histone H3 tail (H3T3) (16, 17). Then H3pT3 is recognized by the chromosome passenger complex, resulting in cluster-mediated autophosphorylation and autoactivation of Aurora B (18, 19). In addition, priming phosphorylation of Survivin Ser(P)20 by PLK1 is critical for orchestrating Aurora B activity in centromeres, which is essential for accurate chromosome segregation and faithful completion of cytokinesis (20).
Phosphatase specificity of PP1 is achieved through association with accessory subunits that control the specificity and selectivity via spatial localization of PP1 related to its cognate substrates. PP1 has multiple binding partners that determine its localization, specificity, and activity of PP1 in cellular dynamics and plasticity during interphase and mitosis (21). In prometaphase and metaphase, PP1 (in mammals isoforms α, β, and γ) localizes to kinetochores (7, 22–25), mostly through binding to the RVXF motif of the outer kinetochore protein KNL1 (7).
Sds22 (also called Sds22p in yeast or PPP1R7) was originally identified in fission yeast using high copy suppression of Schizosaccharomyces pombe dis2 (26). It is a highly conserved, essential protein that physically interacts with PP1 in yeasts and metazoans (27, 28). Sds22 interacts with PP1 through leucine-rich repeats (LRRs), a non-canonical PP1-binding motif composed of an extended helix configuration (29). Sds22 localizes to kinetochores and regulates recruitment of PP1 (23). Upon cellular depletion of Sds22, Aurora B autophosphorylation at Thr232 in the activation loop and unbalanced Aurora B activity is increased during establishment of bipolar attachment (23). During anaphase, Sds22 is required to stabilize kinetochore-spindle attachment (30, 31). However, whether Sds22 localizes to kinetochores remains elusive (7, 32).
Despite the obvious importance of PP1/Sds22 for chromosome segregation, their localization at the kinetochore and their functional relationship to Aurora B remain obscure. Additionally, it was unclear whether and how PLK1, Aurora B, and PP1/Sds22 cooperate at the centromere during mitosis.
In this study, we delineate a novel regulatory mechanism underlying Aurora B kinase activation through the phosphorylation of Sds22 by PLK1. Our biochemical and cellular characterization demonstrates that Sds22 is a substrate of PLK1 in mitosis, which is critical for orchestrating the activity of PP1 and Aurora B at the centromere and accurate chromosome segregation. Our study sheds light on the mechanism of Aurora B regulation through PLK1 and PP1/Sds22 in mitosis.
Results
PLK1 Interacts with and Phosphorylates Sds22
Sds22 is a kinetochore-related protein and plays a key role in faithful mitosis (23, 30–32). Given that, we sought to examine whether Sds22 physically interacts with any other components of the kinetochore. To this end, we carried out a yeast two-hybrid assay to screen Sds22-interacting proteins. Specifically, all candidate kinetochore protein cDNAs were cloned into BD vectors and co-transfected with AD-Sds22, followed by growth selection. As shown in Fig. 1A, co-transfection of AD-Sds22 with BD-PLK1 rescued the transcription of the reporter gene in the yeast strain, whereas no complementary effect was observed between AD-Sds22 and BD-Mad1/Mad2/Sgo1/CENP-H. Thus, our yeast genetic screen suggested that PLK1 is a potential interacting protein physically bound to Sds22.
FIGURE 1.
PLK1 interacts with and phosphorylates Sds22. A, yeast cells were co-transformed with an Sds22 prey construct and the indicated kinetochore protein bait constructs (PLK1, Mad2, Mad1, Sgo1, and CENP-H). An example of such an experiment in which cells were selected on supplemented minimal plates lacking adenine, tryptophan, leucine, and histidine is presented. B, aliquots of HEK293T cells were transiently transfected to express Sds22 with wild type PLK1 (PLK1WT) and kinase-deficient mutant of PLK1 (PLK1KD). 36 h after transfection, the cells were lysed and precipitated with FLAG-agarose. The beads were washed and boiled in 1× SDS-PAGE sample buffer prior to electrophoresis. Western blotting analyses confirmed the co-precipitation of PLK1 with Sds22. C, schematic illustration of human PLK1 functional domain. D, the co-precipitation assay revealed that Sds22 binds to PLK1 full-length (PLK-FL), kinase domain (PLK1-N), and PBD (PLK1-C). Note that the PBD domain showed higher binding efficiency than the kinase domain. E, PLK1 immunoprecipitates from nocodazole-synchronized HeLa cells were immunoblotted with antibodies against PLK1, Sds22, and BubR1. Note that PLK1 immunoprecipitation brought down Sds22 and BubR1. F, Sds22 is a substrate of PLK1 in vivo. HeLa cells transfected with FLAG-Sds22 were arrested at prometaphase using nocodazole. The mitotic cells were collected and divided into two groups: one was treated with DMSO, and the other was treated with the PLK1 inhibitor BI 2536 (10 nm) for 30 min in the presence of 20 μm MG132. The cells were lysed after being pelleted and washed with PBS. The cell lysates clarified by centrifugation were incubated with FLAG M2 beads at 4 °C with an end to end mixing for 3 h. After incubation, the FLAG M2 beads immunoprecipitated with FLAG-Sds22 proteins were washed and then boiled in 1× SDS-PAGE sample buffer. The bound FLAG-Sds22 proteins were separated on an SDS-PAGE for CBB staining. The indicated two FLAG-Sds22 bands were cut from the SDS-PAGE gel for phosphorylation sites identification using two-dimensional LC/MS techniques. G, Sds22 is a substrate of PLK1 in vitro. Bacterial expression of recombinant MBP-Sds22 fusion proteins were phosphorylated in vitro using [γ-32P]ATP and active PLK1 as described under “Materials and Methods.” The samples were separated by SDS-PAGE. Left panel, CBB-stained gel of samples of MBP-Sds22WT plus PLK1, and MBP-Sds225A plus PLK1. Right panel, the same gel was dried and subsequently exposed with x-ray film. Note that there was dramatic incorporation of 32P into wild type Sds22 but little into the mutant Sds22 or MBP.
To examine whether PLK1 forms a cognate complex with Sds22 in cells, we carried out an immunoprecipitation assay using lysates from HEK293T cells transiently transfected to express FLAG-PLK1 (WT and kinase-deficient mutant (KD)) and Sds22-GFP (Fig. 1B). Western blotting analyses demonstrated that FLAG-PLK1 (WT and KD) interacts with Sds22-GFP but not GFP (Fig. 1B). To identify which domain of PLK1 physically binds to Sds22, we carried out an immunoprecipitation assay using lysates from HEK293T cells transiently transfected to express FLAG-Sds22 and GFP-PLK1 and its deletion mutants, as illustrated in Fig. 1C. Western blotting analyses validated that both the PLK1 kinase domain (PLK1-N) and Polo-binding domain (PBD; PLK1-C) physically interact with Sds22 (Fig. 1D). Immunoprecipitation of mitotic cell lysates with an anti-PLK1 mouse antibody pulled down endogenous Sds22 (Fig. 1E, middle panel, lane 3). Sds22 was not precipitated by nonspecific mouse IgG (Fig. 1E, middle panel, lane 2), demonstrating the specific interaction between Sds22 and PLK1.
The interaction between PLK1 and Sds22 prompted us to examine whether Sds22 is a substrate of PLK1. Our computational analyses suggested that Ser24, Ser27, Ser44, Ser47, and Thr277 of Sds22 are potential substrates of PLK1 (33), which are conserved among eukaryotic Sds22 proteins. Recent quantitative phosphoproteomics revealed that Ser24, Ser27, Ser44, and Ser47 of Sds22 are physiologically phosphorylated in different context (34–37). To uncover the PLK1-elicited phosphorylation sites of Sds22 in mitosis, we carried out mass spectrometric identification of potential phosphorylation sites of Sds22 isolated from mitotic HeLa cells in the presence and absence of a selective PLK1 inhibitor BI 2536 (20). Specifically, aliquots of HeLa cells were transiently transfected to express FLAG-Sds22 for 24 h followed by nocodazole synchronization (100 ng/ml for 18 h). Mitotic HeLa cells were collected by shake-off followed by incubation with 10 nm BI 2536 or DMSO for 30 min in the presence of 20 μm MG132. Cell lysates were then generated and clarified for immunoprecipitation as described (20). As shown in Fig. 1F, FLAG-Sds22 proteins were immunoisolated from DMSO- and BI 2536-treated HeLa cells. Sds22 protein bands, from DMSO- and BI 2536-treated cells, were removed for in-gel digestion followed by mass spectrometric identification (12). Our mass spectrometric analyses have identified seven phosphorylation sites in mitotic HeLa cells in which five sites are responsive to BI 2536 treatment. These five sites are Ser24, Ser27, Ser44, Ser47, and Thr277 as listed in supplemental Fig. S1. The respective phosphopeptides identified by mass spectrometer and phosphorylation sites were summarized in supplemental Table S1.
To test whether Ser24, Ser27, Ser44, Ser47, and Thr277 are substrates of PLK1, we performed in vitro phosphorylation on recombinant MBP-Sds22 fusion proteins, including both wild type and non-phosphorylatable Sds22 mutants in which Ser24, Ser27, Ser44, Ser47, and Thr277 were replaced by alanines (Sds225A). As shown in Fig. 1G, incubation of MBP-tagged Sds22 wild type (Sds22WT) and non-phosphorylatable mutant (Sds225A) with [γ-32P]ATP and PLK1 resulted in the incorporation of 32P into Sds22WT, but not into Sds225A mutant and MBP. This PLK1-mediated phosphorylation is specific, because incubation of Sds22 with [γ-32P]ATP in the absence of PLK1 resulted in undetectable incorporation of radioactivity into the wild type protein (data not shown). Thus, we conclude that Ser24, Ser27, Ser44, Ser47, and Thr277 of Sds22 are bona fide substrates of PLK1 during mitosis.
Phosphorylation of Sds22 Is Critical for Accurate Mitotic Progression
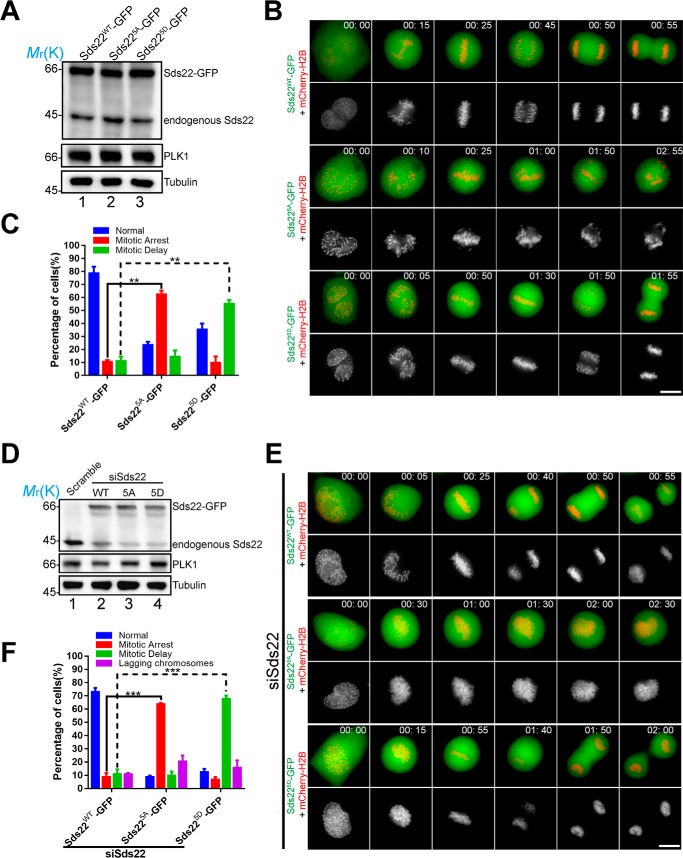
To examine the role of PLK1-mediated phosphorylation of Sds22 in mitosis, we generated non-phosphorylatable and phosphomimetic Sds22 mutants tagged with GFP, which were transiently transfected into HeLa cells together with mCherry-H2B. Comparison of expression levels of exogenous and endogenous Sds22 were shown in Fig. 2A. Real time analyses of transfected HeLa cell preparations showed that non-phosphorylatable and phosphomimetic mutants of Sds22 (Sds225A and Sds225D) interfere with mitotic progression, compared with the exogenously expression of Sds22WT (Fig. 2B). Expression of Sds225A-GFP resulted in a typical mitotic arrest phenotype (the chromosomes did not align along the equatorial plane more than 1 h from nuclear envelope breakdown (NEBD)) (Fig. 2, B, middle panel, and C), and expression of Sds225D-GFP resulted in a typical mitotic delay phenotype in which misaligned chromosome finally congressed to metaphase plate but failed to segregate (Fig. 2B, lower panel). Our statistical analyses show that alternation of PLK1-elicited Sds22 phosphorylation perturbed accurate chromosome segregation (Fig. 2C; **, p < 0.01).
FIGURE 2.
Phosphorylation of Sds22 is critical for mitotic progression. A, the expression levels of Sds22WT-GFP, Sds225A-GFP, and Sds225D-GFP in HeLa cells co-transfected with mCherry-H2B. Cells were collected at 48 h after transfection and analyzed by Western blots using indicated antibodies. Tubulin served as loading control. B, live cell imaging of chromosome movements in HeLa cells expressing wild type and mutant Sds22. Aliquots of HeLa cells were transiently transfected to express various GFP-tagged Sds22 plasmids with mCherry-H2B. 36 h after transfection; images were acquired at the indicated time points. Note that expression of Sds225A-GFP resulted in mitotic arrest phenotype, and expression of Sds225D-GFP resulted in mitotic delay phenotype. Scale bar, 10 μm. C, statistical analyses of mitotic phenotypes as in B. Error bars represent means ± S.E.; >20 cells of each category from three independent preparations. Student's t test was used to calculate p value for comparison. **, p < 0.01. D, the expression levels of Sds22WT-GFP and its mutants in endogenous Sds22-depleted HeLa cells. Cells were collected at 48 h after transfection and analyzed by Western blots using indicated antibodies. Tubulin served as loading control. E, live cell imaging of chromosome movements in Sds22 depleted HeLa cells expressing wild type and mutant Sds22 proteins. Note that in Sds22-depleted HeLa cells, expression of Sds225A-GFP resulted in mitotic arrest phenotype, and expression of Sds225D-GFP resulted in mitotic delay phenotype. Scale bar, 10 μm. F, statistical analyses of mitotic phenotypes seen in live cell imaging as in E. Error bars represent means ± S.E.; >20 cells of each category from three independent preparations. Student's t test was used to calculate p value for comparison. ***, p < 0.001.
To minimize the interference of the endogenous Sds22 with exogenously expressed Sds22 proteins in mitosis, we transiently transfected Sds22 siRNA together with GFP-tagged Sds22 and mCherry-H2B into HeLa cells. Expression levels of exogenous Sds22 were shown in Fig. 2D. Analyses of endogenous Sds22 protein level showed the knockdown efficiency of Sds22 siRNA (Fig. 2D). Real time analyses of the transfected HeLa cells showed that compared with the exogenous expression of Sds22-GFP showing a normal mitotic phenotype (Fig. 2, E, top panel, and F), expression of Sds225A-GFP resulted in a typical mitotic arrest phenotype (the chromosomes did not align along the equatorial plane more than 1 h from NEBD) (Fig. 2, E, middle panel from 00:30 to 02:30, and F), and expression of Sds225D-GFP resulted in a typical mitotic delay phenotype (after more than 30 min, the aligned chromosomes along the equatorial plane could not separate) (Fig. 2, E, bottom panel from 00:55 to 01:40, and F). Thus, we conclude that PLK1-mediated phosphorylation of Sds22 is critical for mitotic progression.
Phosphorylation of Sds22 Promotes Aurora B Kinase Activity
Given that the Sds22 mutants caused mitotic arrest and mitotic delay phenotypes and the major difference between unaligned and aligned chromosomes is the phosphorylation of a series of outer kinetochore proteins by Aurora B in a tension-regulated manner (38), we therefore asked whether the Sds22 mutants influenced the Aurora B activity. Indeed, compared with Sds225A-GFP, overexpression of Sds22WT-GFP and Sds225D-GFP exhibits higher intensity of CENP-A Ser(P)7 (a marker of Aurora B activity) staining on the kinetochores, indicating that Sds225A decreases Aurora B activity (Fig. 3, A and B).
FIGURE 3.
Phosphorylation of Sds22 promotes Aurora B kinase activity. A, phosphorylation of a kinetochore Aurora B substrate CENP-A is decreased by expression of Sds225A. Compared with Sds225A-GFP, overexpression of Sds22WT-GFP and Sds225D-GFP produces stronger CENP-A Ser(P)7 staining on the kinetochores. Scale bar, 10 μm. B, statistical analyses of CENP-A Ser(P)7 immunofluorescence intensity at the kinetochores as in A indicate that Sds225A decreases Aurora B activity. Error bars represent means ± S.E.; >20 cells of each categories from three independent preparations. Student's t test was used to calculate p value for comparison. *, p < 0.05; **, p < 0.01. C, phosphorylation of CENP-A at Ser7 (a kinetochore substrate of Aurora B) is increased by Sds225D expression in PLK1-depleted HeLa cells. Compared with Sds225D-GFP, overexpression of Sds22WT-GFP and Sds225A-GFP produces weaker CENP-A Ser(P)7 staining on the kinetochores. Scale bar, 10 μm. D, statistical analyses of CENP-A Ser(P)7 immunofluorescence intensity at the kinetochores as in C indicatethat Sds225D expression increases Aurora B activity in PLK1-depleted HeLa cells. Error bars represent means ± S.E.; >20 cells of each categories from three different independent preparations. Student's t test was used to calculate p value for comparison. *, p < 0.05; ***, p < 0.001. E, the expression levels of Sds22WT-GFP and its mutants in endogenous PLK1-depleted HeLa cells. Cells were collected at 48 h after transfection and analyzed by Western blotting analyses using indicated antibodies. Tubulin served as loading control. F, phosphorylation of CENP-A at Ser7 is decreased by expression of Sds225A in Sds22-depleted HeLa cells. Compared with Sds225A-GFP, in Sds22 depleted HeLa cells, overexpression of Sds22WT-GFP and Sds225D-GFP produces stronger CENP-A Ser(P)7 staining on the kinetochores. Scale bar, 10 μm. G, statistical analyses of CENP-A Ser(P)7 immunofluorescence intensity at the kinetochores as in F indicate that in Sds22-depleted HeLa cells, Sds225A decreases Aurora B activity (Error bars represent means ± S.E.; >20 cells of each categories from three different preparations). Student's t test was used to calculate p value for comparison. **, p < 0.01; ***, p < 0.001. H, phosphorylation of CENP-A at Ser7 is decreased by kinetochore targeted Sds225A in Sds22-depleted HeLa cells. Compared with Hec1-mCherry-Sds225A, in Sds22-depleted HeLa cells, overexpression of Hec1-mCherry-Sds22WT and Hec1-mCherry-Sds225D produced stronger CENP-A Ser(P)7 staining on the kinetochores. Scale bar, 10 μm. I, statistical analyses of CENP-A Ser(P)7 immunofluorescence intensity at the kinetochores as in H indicate that in Sds22-depleted HeLa cells, kinetochore targeted Sds225A decreases Aurora B activity. Error bars represent means ± S.E.; >20 cells of each categories from three different preparations. Student's t test was used to calculate p value for comparison. ***, p < 0.001. J, the expression levels of Hec1-mCherry-Sds22WT and its mutants in endogenous Sds22-depleted HeLa cells. Cells were collected at 48 h after transfection and analyzed by Western blots using indicated antibodies. Tubulin served as loading control. ACA, anti-centromere antibody.
To exclude the effects of the endogenous PLK1 and Sds22 in mitosis, we transiently transfected GFP-Sds22 wild type and mutants in PLK1-depleted and Sds22-depleted HeLa cells. Quantitative analyses showed that compared with Sds225D-GFP, in PLK1 shRNA-transfected HeLa cells, overexpression of Sds22-GFP and Sds225A-GFP produced weaker CENP-A Ser(P)7 staining on the kinetochores, implying that Sds225D increases Aurora B activity (Fig. 3, C and D). Expression protein levels of wild type, non-phosphorylatable and phosphomimetic Sds22 were comparative (Fig. 3E). Compared with Sds225A-GFP, in Sds22-depleted HeLa cells, overexpression of Sds22WT-GFP and Sds225D-GFP produced stronger CENP-A Ser(P)7 staining on the kinetochores, indicating that expression of non-phosphorylatable Sds22 suppresses Aurora B activity (Fig. 3, F and G).
The kinetochore localization of Sds22 remains controversial (7, 32), live cell imaging and immunofluorescence results showed that the kinetochore localization of Sds22 was not obvious (Figs. 2, B and E, and 3, A, C, and F). To validate that the Sds22 function seen above is kinetochore-centric, we engineered kinetochore-localized Sds22 by fusing wild type Sds22 and its mutants to Hec1-mCherry (39, 40). To confirm that phosphomimetic Sds22 promotes Aurora B activity, aliquots of HeLa cells were transiently transfected to express Hec1-mCherry-Sds22 in the presence of Sds22 siRNA. As shown in Fig. 3H, all three Hec1-mCheery-Sds22 proteins were localized to kinetochore. Consistent with overexpression experiments, Aurora B kinase activity is highest in Hec1-mCherry-Sds225D-expressing cells judged by the fluorescence intensity of Ser(P)7-CENP-A (Fig. 3G). The fluorescence intensity of Ser(P)7-CENP-A is minimized in Hec1-mCherry-Sds225A-expressing cells. Quantitative analyses of normalized Ser(P)7-CENP-A level showed that, compared with Hec1-mCherry-Sds225A, in Sds22-depleted HeLa cells, overexpression of Hec1-mCherry-Sds22WT and Hec1-mCherry-Sds225D produced higher levels of Aurora B activity at the kinetochore marked by CENP-A Ser(P)7 staining, suggesting that kinetochore targeted Sds225A decreases Aurora B activity (Fig. 3I; p < 0.001). Western blotting analyses confirmed that expression protein levels of three Hec1-mCherry-Sds22 constructs were comparable, whereas the endogenous levels in the Hec1-mCherry-Sds22-expressed cells were suppressed (Fig. 3J).
It has been recently proposed that PLK1 is critical for orchestrating Aurora B activity in centromere through Haspin-H3T3-Survivin signaling axis (16–20, 41). To examine whether the phosphorylation of Sds22 by PLK1 retains Aurora B kinase activity, we sought to monitor Aurora B activity at the centromere using FRET-based sensors that report quantitative changes in substrate phosphorylation. We adapted a sensor design in which changes in intramolecular FRET between cyan and yellow fluorescent proteins (CFP-YFP) depends on changes in phosphorylation of an Aurora B substrate peptide that is conserved among members of the kinesin-13 family (20). To mimic the localization of endogenous Aurora B substrate (8), the sensor was targeted to centromere (CENP-B fusion; Fig. 4, B and C). To validate the sensor response to changes in Aurora B activity, we imaged mitotic cells after kinase inhibition (experimental scheme in Fig. 4A). The FRET/CFP emission ratio of the sensor increased after treatment with a selective Aurora B inhibitor Hesperadin (Fig. 4B), consistent with the previous reports (20, 38).
FIGURE 4.
Phosphorylation of Sds22 by PLK1 is essential for Aurora B kinase activity. A, schematic illustration of experimental design. B, HeLa cells expressing the centromere-targeted Aurora B kinase sensor were imaged at metaphase. Aurora B kinase sensor is responsive to Aurora B inhibitor Hesperadin. PLK1 inhibitor BI 2536 also modulates Aurora B kinase activity in mitosis. Scale bar, 10 μm. C, HeLa cells expressing the centromere-targeted Aurora B kinase sensor and mCherry-Sds22WT and its mutants were imaged at metaphase. Aurora B kinase sensor is responsive to mCherry-Sds225A. Scale bar, 10 μm. D, statistical analyses of the FRET/CFP emission ratio as in B and C indicate that mCherry-Sds225A decreases Aurora B activity. Error bars represent means ± S.E.; >20 cells of each categories from three different preparations. Student's t test was used to calculate p value for comparison. *, p < 0.05; **, p < 0.01. E, Western blotting analysis of the level of Ser(P)7-CENP-A indicates that expression of Sds225A suppresses Aurora B kinase activity. F, statistical analyses of band intensity of CENP-A Ser(P)7 as in E indicate that expression of Sds225A decreases Aurora B activity. Western blot signal intensities were quantified using ImageJ software. The average protein band intensities were measured, and the background intensities were subtracted. The CENP-A Ser(P)7 intensities were then normalized against CENP-A values to account for any variations in immunoblot acquisition. Error bars represent means ± S.E. from three different preparations. Student's t test was used to calculate p value for comparison. *, p < 0.05; **, p < 0.01; ***, p < 0.001.
Given that PLK1-mediated phosphorylation of Sds22 increases Aurora B activity, we next examined whether inhibition of PLK1 reduces Aurora B activity. As shown in Fig. 4 (B and D), the addition of PLK1 kinase inhibitor BI 2536 resulted in a dramatic reduction of phosphorylation of the sensor in centromere, supporting the notion that PLK1 kinase is critical for activating Aurora B at the centromere. To examine whether PLK1 phosphorylation toward Sds22 is important for Aurora B activity, we measured centromeric Aurora B activity in cells expressing the wild type Sds22 and its phospho-mutants. As predicted, expression of mCherry-Sds225A resulted in a reduction of centromeric Aurora B kinase activity judging by an increased FRET/CFP emission ratio (Fig. 4C). In contrast, expression of mCherry-Sds225D decreased the FRET/CFP emission ratio, demonstrating that centromeric Aurora B kinase activity is a function of PLK1-phosphorylated Sds22 (Fig. 4, C and D). Importantly, this Sds225D-elicited Aurora B kinase activity is minimized by Hesperadin treatment (Fig. 4C, bottom panel). Our statistical analyses of the FRET/CFP emission ratio, from more than 20 cells per categories, indicate that overexpression of mCherry-Sds225A decreases Aurora B activity at the kinetochore.
We next examined the phosphorylation of Aurora B endogenous substrates using phospho-specific antibodies against Ser(P)7-CENP-A in cells expressing the wild type Sds22-GFP and its mutants. As shown in Fig. 4E, the band intensity of Ser(P)7-CENP-A is minimal under treatment with Hesperadin. Quantitative analyses indicated that expression of non-phosphorylatable Sds225A-GFP resulted in 50% reduction of Aurora B activity compared with that of phospho-mimicking Sds225A-GFP-expressing cells judged by the level of Ser(P)7-CENP-A (Fig. 4F). Thus, these data demonstrate that PLK1-mediated phosphorylation of Sds22 is an important factor for sustaining Aurora B activity during the process of chromosome alignment in mitosis.
Phosphorylation of Sds22 Inhibits PP1 Activity
Recent studies suggest that the binding of Sds22 to PP1 at the kinetochore negatively regulates PP1-mediated dephosphorylation of Aurora B, affecting Aurora B activity (32). Conversely, our data suggest that PLK1-mediated phosphorylation of Sds22 increases Aurora B activity. To examine whether PLK1-mediated phosphorylation of Sds22 influences its binding to PP1, MBP pulldown assays were performed using MBP-Sds22 as an affinity matrix to isolate GST-PP1γ. As shown in Fig. 5A, compared with MBP-Sds225A, MBP-Sds22WT, and MBP-Sds225D exhibited higher binding efficiency with GST-PP1γ. To determine whether it is the same in cells, anti-FLAG immunoprecipitation assays were performed. Anti-GFP immunoblots confirmed that, compared with Sds225A-GFP, Sds22WT-GFP and Sds225D-GFP showed higher binding efficiency with FLAG-PP1γ (Fig. 5B).
FIGURE 5.
Phosphorylation of Sds22 inhibits PP1 activity. A, in vitro pulldown assay of PP1γ with Sds22WT and its mutants. The gel was stained with CBB (upper panel). Western blotting analysis with an anti-GST antibody (lower panel) confirmed the interaction between PP1γ with Sds22. Note that compared with MBP-Sds225A, MBP-Sds22WT, and MBP-Sds225D showed higher binding efficiency with GST-PP1γ (lane 10). B, association of PP1γ with Sds22. Aliquots of HEK293T cells were transiently transfected to express various GFP-tagged Sds22 plasmids with FLAG-PP1γ. 36 h after the transfection, the cells were lysed and precipitated with FLAG-M2 agarose beads as described under “Materials and Methods.” Note that Sds225A-GFP association with PP1γ is weak. C, Sds22 inhibits PP1-mediated dephosphorylation of Aurora B Thr(P)232 in a dose-dependent manner. PP1 specifically dephosphorylates Aurora B Thr(P)232. Recombinant, autophosphorylated Aurora B was incubated either alone or with PP1 plus purified Sds22 at indicated concentrations. λ-Phosphatase (λ-PPase) was used as a positive control. Phosphorylation of Aurora B Thr(P)232 was detected by Western blot with phospho-specific antibodies. Equal loading was monitored by GST blot. D, enzymatic kinetics of PP1-mediated dephosphorylation of Aurora B Thr(P)232 in response to increased concentrations of Sds22. PP1 concentration is 40 nm. The Michaelis-Menten data of PP1 activity at various concentrations of Sds22 were shown. Error bars represent means ± S.E. from three different preparations. E, Lineweaver-Burk plot of Lineweaver-Burk analysis (D) is an efficient method of linearizing substrate velocity data so as to determine the kinetic constants Km and Vmax. Based on Lineweaver-Burk analysis, the calculated Km was derived for PP1/Sds22WT (0.04 ± 0.003), PP1/Sds225A (0.013 ± 0.006), and PP1/Sds225D (0.03 ± 0.002). The kinetic analyses indicate that phosphorylation of Sds22 inhibits PP1-mediated dephosphorylation of Thr(P)232 of Aurora B. F, a proposed working model accounting for hierarchy signaling regulation of Aurora B by PP1/Sds22 and PLK1 during mitosis. See text for details.
To confirm whether there is a direct link between PLK1-mediated phosphorylation of Sds22 and PP1, we reconstituted PP1γ-mediated dephosphorylation of Aurora B in vitro. We purified recombinant GST-tagged Aurora B, which is active and autophosphorylated at Thr232 (42, 43). Aurora B was incubated with purified PP1 either alone or with increasing concentrations of Sds22, as described previously (32). Incubation with PP1 alone led to a notable decrease of Thr(P)232 signal, demonstrating that PP1 directly dephosphorylates Aurora B at Thr232 (Fig. 5C), consistent with a previous report (32). Importantly, addition of Sds22WT and Sds225D significantly blocked Aurora B dephosphorylation in a dose-dependent manner, whereas Sds225A did not exhibit regulatory effect (Fig. 5C). The kinetics data were plotted in the Michaelis-Menten format with velocity versus concentration of casein (Fig. 5D) and in the double reciprocal format with 1/velocity versus 1/concentration of Aurora B (Fig. 5E). On the basis of a Lineweaver-Burk analysis, the calculated Km was 0.013 ± 0.002 for 5A and 0.03 ± 0.002 for the 5D mutant. The enzymatic analyses indicate that pPLK1-mediated phosphorylation of Sds22 inhibits PP1-mediated dephosphorylation of Aurora B at Thr232.
Discussion
Proper chromosomal oscillation and kinetochore microtubules dynamics ensures precise chromosome bi-orientation and faithful chromosome segregation in mitosis. Our studies revealed that PLK1 physically interacts with Sds22, both in vitro and in vivo, and that Sds22 is a cognate substrate of PLK1 in mitosis. The PLK1-mediated phosphorylation of Sds22 promotes the kinase activity of Aurora B in vivo. Importantly, the dynamic regulation of Sds22 phosphorylation by PLK1 is essential for accurate chromosome segregation as perturbation of Sds22 phosphorylation dynamics causes severe defects in chromosome segregation during mitosis. Our study provides a novel insight into a better understanding of PLK1-Sds22/PP1-Aurora B feedback loop in regulating microtubule dynamics and chromosome attachment error correction during cell division as illustrated in our working model Fig. 5F.
Functional PP1 holoenzyme consists of a catalytic subunit and a regulatory subunit. Early studies suggested that most regulatory subunits contain the sequence motif RVX(F/W). More recently, systematic analysis of the docking peptides that combined biochemistry with molecular dynamics gave rise to a refined consensus sequence of (H/K/R)(A/C/H/K/M/N/Q/R/S/T/V)(V)(C/H/K/N/Q/R/S/T)(F/W) (44). However, Sds22 binds to PP1 through a non-canonical PP1-binding motif LRR rather than the RVX(F/W) motif (29). It would be of great interest in the future to delineate the structure-activity relationship of Sds22-PP1 at the kinetochore and the PLK1-elicited regulation described here.
It should be noticed that most of the phosphorylation sites of Sds22 (Ser24, Ser27, Ser44, and Ser47) by PLK1 lie outside of the LRR, the PP1-binding motif. Definitely, further work should be done to investigate whether PLK1-mediated phosphorylation of Sds22 promotes its binding to PP1 through conformational change wherein PP1-binding motif was exposed after phosphorylation.
It is noteworthy that PLK1-Sds22 interaction may also orchestrate the anaphase events, because perturbation of Sds22 resulted in aberrant polar relaxation and chromosome instability phenotypes (30, 31). Because numerous studies argued that Sds22 remains on kinetochores throughout anaphase when PLK1 has relocated to central spindle and midzone, it would be of central importance to identify the elements responsible for Sds22 distribution at the kinetochore during chromosome segregation during cell division and the spatiotemporal dynamics of PP1-Sds22 interaction during kinetochore attachment error correction process.
Taken together, we propose that dynamic phospho-regulation of Sds22 by PLK1 establishes sustaining Aurora B activity for kinetochore attachment error correction via minimizing PP1-elicited dephosphorylation of Aurora B at Thr232. It is likely that multiple mitotic kinase cascades coordinate to orchestrate faithful chromosome segregation in mitosis. The PLK1-Sds22/PP1-Aurora B axis established here is a core of this dynamic molecular society that links accurate mitotic progression to chromosome stability during cell division.
Materials and Methods
Plasmids
To generate GFP-tagged Sds22 and mutants, PCR-amplified cDNA was cloned into pEGFP-N3 vector (Clontech). GFP-tagged PLK1 full-length and truncations were inserted into pEGFP-C2 vector (Clontech). The bacterially expressed constructs of Sds22 were cloned into pMal-C2 vector (New England Biolabs). Bacterially expressed constructs of PP1γ and Aurora B were cloned into pGEX-6P-1 (GE Healthcare). To generate FLAG-tagged protein, PLK1, PP1γ, and Sds22 cDNA were inserted into the p3×FLAG-myc-CMV-24 vector (Sigma-Aldrich). mCherry-H2B fusion protein was cloned into pcDNA3.1-B vector. Hec1-mCherry-Sds22 was created by inserting the human Sds22 at the C terminus of mCherry and Hec1 at the N terminus of mCherry in vector pCDNA3.1, according to the procedures described previously (39). All plasmid constructs were sequenced for verification.
Yeast Two-hybrid Assay
Yeast two-hybrid assays were performed as described previously (45). Briefly, Sds22 cDNA was inserted into the pGADT7 vector to create a fusion with amino acids 768–881 of the GAL4 AD. The resultant AD-Sds22 was transformed into strain AH109 along with different recombinant plasmids expressing a GAL4 DNA-binding domain (DNA-BD) with different kinetochore proteins, respectively. The co-transformed yeast cells were grown on synthetic defined plates with X-α-Gal but lacking Leu, Trp, His, and Ade.
Antibodies and siRNAs
Primary antibodies used in this study were: Anti-Sds22 goat polyclonal antibody (sc-162164; Santa Cruz Biotechnology), anti-PLK1 mouse monoclonal antibody (377100; Life Technologies), and anti-centromere antibody (HCT-0100; ImmunoVision). Anti-GST tag (2625), anti-Ser(P)7 CENP-A (2187), and anti-CENP-A (2186) antibodies were purchased from Cell Signaling Technology. Anti-BubR1 antibody (612503) was purchased from BD Biosciences. Anti-FLAG antibody (F1804), anti-tubulin antibody (DM1A, T9026), and anti-GFP antibody (G6539) were purchased from Sigma-Aldrich. For all Western blots, the signals were detected using horseradish peroxidase-conjugated goat anti-mouse, sheep anti-goat, or goat anti-rabbit antibodies (Jackson ImmunoResearch). FITC- and rhodamine-conjugated secondary antibodies were also obtained commercially (Jackson ImmunoResearch).
Sds22 siRNA (synthesized by Shanghai GenePharma Co., Ltd) targeting to the 3′-UTR (5′-AUCGUCAGUGCCUUUAUUG-3′) of Sds22 gene was used. PLK1 shRNA targeting the 3′-UTR (5′-CCGGAGCTGCATCATCCTTGCAGGTCTCGAGACCTGCAAGGATGATGCAGCTTTTTT-3′) of PLK1 gene was purchased from Sigma-Aldrich (TRCN0000011006).
Cell Culture, Transfection, and Drug Treatments
HeLa and HEK293T cells, from American Type Culture Collection, were cultured and maintained in advanced DMEM (Invitrogen) with 10% (v/v) FBS (HyClone), 2 mm glutamine, and 100 units/ml penicillin plus 100 μg/ml streptomycin (Invitrogen) at 37 °C with 8% CO2. HeLa cells were transfected with plasmids, siRNA (usually 100 nm), or shRNA using Lipofectamine 2000 (Invitrogen). For mitotic synchronization, the cells were blocked at G1/S phase in 2 mm thymidine (Sigma-Aldrich) for 16 h, followed by release in fresh medium for 7–9 h. In some specific experiments, the cells were treated with indicated inhibitors for another 1 or 2 h, wherein Hesperadin was used at 100 nm, and BI 2536 was used at 10 nm, and MG132 was used at 20 μm.
Immunofluorescence Microscopy and Data Analyses
Cells grown on coverslips were fixed with 3.7% formaldehyde (Sigma-Aldrich) in 60 mm PIPES, 25 mm HEPES, pH 6.9, 10 mm EGTA, 2 mm MgCl2, 4 m glycerol, and PBS at 37 °C for 10 min. After formaldehyde fixation, the cells were subsequently permeabilized with 0.1% Triton X-100 in PBS at room temperature for 10 min. Following a 30-min block in 1% BSA and incubation with primary antibodies at room temperature for 1 h, the cells were incubated with secondary antibodies for another 1 h. After staining with DAPI (Sigma-Aldrich) for 1 min, coverslips were mounted in Vectashield mounting medium (Vector). The images were acquired using Olympus 60×/1.42 Plan APO N objective on an Olympus IX71 microscope (Applied Precision Inc.). Deconvoluted images from each focal plane were projected into a single stack montage using SoftWorx software (Applied Precision).
Fluorescence Intensity Quantification
For fluorescence intensity quantification of the level of kinetochore-associated protein, the kinetochore staining signals were measured with ImageJ software (National Institutes of Health), according to the procedures described previously (46). In brief, the average pixel intensities from at least 50 kinetochore pairs from five cells were measured, and background pixel intensities were subtracted. The pixel intensities at each kinetochore pair were then normalized against anti-centromere antibody pixel values to account for any variations in staining or image acquisition.
FRET Assay
Live fluorescence ratio imaging was collected using a Deltavision wide field deconvolution microscope system (Applied Precision Inc.) built on an Olympus IX-71 inverted microscope base equipped with Photometric Coolsnap HQ2 CCD camera as described previously (20), using 60×/NA1.42 PlanApo oil immersion lens (Olympus). For live cell imaging with FRET sensors, CFP was excited at 436 nm, whereas CFP and YFP (FRET channel) emissions were acquired at 470 and 535 nm, respectively. For YFP channel, YFP was excited at 500 nm, and YFP emission was acquired at 535 nm. The FRET emission ratio (FRET/CFP) was calculated by dividing CFPex/CFPem (CFP) to CFPex/YFPem (FRET) using SoftWoRx (Applied Precision Inc.). For statistical analyses, individual centromere/kinetochore was defined automatically from confocal image stacks, and FRET emission ratio on each centromere/kinetochore was measured as previously described (47). The ratios were normalized by dividing by the maximum value of FRET emission ratio for each experiment. According to the design of the FRET-based sensors, a higher FRET emission ratio indicates a lower kinase activity.
Live Cell Microscopy and Data Analyses
Live cell imaging was performed as described previously (48, 49). In brief, the cells were cultured in MatTek glass-bottomed dishes (MatTek) in CO2-independent medium (Gibco) containing 10% (v/v) FBS and 2 mm glutamine in a sealed chamber heated to 37 °C and observed under a DeltaVision deconvolution microscope (Applied Precision). Images were acquired from NEBD with 3- or 5-min intervals and presented in Photoshop and Illustrator (Adobe).
Immunoprecipitation and MBP Pulldown Assay
For immunoprecipitation, HEK293T cells were transfected with indicated plasmids, collected, and lysed in a lysis buffer (50 mm Tris-HCl, pH 7.4, 150 mm NaCl; 1 mm EDTA) containing 0.1% Triton X-100 in the presence of mammalian protease inhibitors mixture (P8340; Sigma-Aldrich). After clarification by centrifugation, cell lysates were incubated with FLAG M2 beads (F2426; Sigma-Aldrich) at 4 °C rotating for 3 h. In immunoprecipitation of endogenous PLK1, HeLa cell lysates were incubated with IgG or PLK1 antibody at 4 °C rotating for 4 h followed by extended incubation with protein A/G microbeads for another 1 h. After incubation, the beads were washed three times with lysis buffer containing 0.1% Triton X-100, washed one time with lysis buffer, and then boiled in 1× SDS-PAGE sample buffer. The bound proteins were separated on an SDS-PAGE gel and transferred onto nitrocellulose membrane for Western blotting analyses.
Pulldown assays were carried out as described previously (45). Briefly, the GST-tagged proteins in the soluble fraction were purified from bacteria by glutathione-agarose chromatography and then eluted by GST elution buffer (50 mm Tris-HCl, 10 mm reduced glutathione, pH 8.0), whereas MBP-tagged proteins were purified using amylose beads. MBP-tagged protein-bound Sepharose beads were incubated with purified GST-tagged proteins for 1 h at 4 °C. After incubation, the beads were washed three times with PBS containing 0.1% Triton X-100 and one time with PBS and then boiled in 1× SDS-PAGE sample buffer. The bound proteins were separated by SDS-PAGE followed by Coomassie Brilliant Blue (CBB) staining or Western blotting analyses using GST antibody.
In Vivo Phosphorylation
HeLa cells were transfected with FLAG-Sds22 for 24 h. For mitotic synchronization, the cells were arrested at prometaphase in 100 ng/ml nocodazole for 18 h. The mitotic cells were collected and divided into two groups: one was treated with DMSO, and the other was treated with BI 2536 (10 nm) for 30 min in the presence of 20 μm MG132. After being pelleted and washed one time with PBS, the cells were lysed in a lysis buffer (50 mm Tris-HCl, pH 7.4, 150 mm NaCl, 1 mm EDTA) containing 0.1% Triton X-100 in the presence of mammalian protease inhibitors mixture (P8340; Sigma-Aldrich). After clarification by centrifugation, cell lysates were incubated with FLAG M2 beads (F2426; Sigma-Aldrich) at 4 °C rotating for 3 h. After incubation, the FLAG M2 beads immunoprecipitated with proteins were washed three times with lysis buffer containing 0.1% Triton X-100 and one time with lysis buffer and then boiled in 1× SDS-PAGE sample buffer. The bound proteins were separated by SDS-PAGE for CBB staining. The target FLAG-Sds22 bands were cut from the SDS-PAGE gel for phosphorylation site identification using two-dimensional LC/MS techniques (ProteomeX-LTQ 10954; Thermo Fisher Scientific).
In Vitro Phosphorylation
In vitro phosphorylation assays were carried out as described previously (49). The PLK1 kinase (7728) was purchased from Cell Signaling Technology. The kinase reactions were performed in 40 μl of 1× kinase buffer (25 mm HEPES, pH 7.2, 50 mm NaCl, 2 mm EGTA, 5 mm MgSO4, 1 mm DTT, 0.01% Brij35) containing 200 ng of PLK1 kinase, 1 mg of MBP-tagged proteins, 5 μCi of [γ-32P]ATP, and 50 μm ATP. The mixtures were incubated at 30 °C for 30 min, and the reactions were stopped with 5× SDS-PAGE sample buffer. Proteins were separated by SDS-PAGE and detected by subsequent autoradiography.
In Vitro Dephosphorylation Assays
PP1 (P0754S) was purchased from New England Biolabs (Beverly, MA). FLAG-Sds22 was expressed in HEK293T cells, affinity-purified using FLAG M2 beads (F2426; Sigma-Aldrich), and eluted using 3×FLAG Peptide (F4799; Sigma-Aldrich) to generate full-length Sds22 protein. Autophosphorylated GST-Aurora B was purified as described (32). 5 pmol of autophosphorylated GST-Aurora B was incubated with 0.5 pmol of PP1 without or with the indicated concentration of Sds22 in 1× NE buffer for protein metallophosphatases (P0754S; New England Biolabs), supplemented with 1 mm MnCl2 and 1 mm protease inhibitors at 30 °C for 15 min, according to the procedures described previously (32). λ-Phosphatase was used at 40 pmol as a positive control. The reactions were stopped with 5× SDS-PAGE sample buffer. Proteins were separated by SDS-PAGE, analyzed by Western blots with Aurora B Thr(P)232 and GST antibodies, and visualized with ECL reagent (Thermo Scientific Pierce) on a LAS400 imaging system (GE Healthcare). Signal intensities were quantified using ImageJ software (National Institutes of Health), according to the procedures described previously (45). GST-Aurora B dephosphorylation was determined by measuring the dephosphorylation in the presence of PP1/Sds22 and subtracted from the experiments in the absence of PP1/Sds22. The data were fit to the Michaelis-Menten equation to derive apparent Vmax and Km with the GraphPad Prism software as described previously (50).
Statistical Analyses
To determine significant differences between means, unpaired Student's t tests assuming unequal variance were performed. Differences were considered significant when p was < 0.05.
Author Contributions
H. D., M. W., and X. L. conceived the project. H. D., M. W., W. P., J. Z., S. A., D. W., and M. Y. designed and performed most biochemical experiments, cell biological characterization, and data analyses. C. W., D. W., F. W., H.-N. G., R. T., and H. Z. performed phosphorylation sites identification using mass spectrometric analyses of mitotic samples. X. G. performed computational analysis for phosphorylation sites using GPS software. W. W. and Y. C. designed and constructed the Aurora B sensor. H. D., G. B., P. H., Z. D., H.-N. G., X. L., and X. Y. wrote the manuscript, and all authors have read and approved the manuscript.
Supplementary Material
Acknowledgments
We thank the members of our groups for insightful discussions during the course of this study.
This work was supported by Chinese Natural Science Foundation Grants 31430054, 31320103904, 91313303, 31621002, 31501095, 31671405, 31371363, and 31601097; Chinese 973 Project Grants 2012CB917204, 2012CB945002, and 2002CB713700; Anhui Province Key Project Grant 08040102005; Chinese Academy of Sciences Center of Excellence Grant 2015HSC-UE010; and Anhui Provincial Natural Science Foundation Grant 1508085SMC213. This work was also supported in part by National Institutes of Health Grants DK056292 and CA164133. The authors declare that they have no conflicts of interest with the contents of this article. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.

This article contains supplemental Table S1 and Fig. S1.
- SAC
- spindle assembly checkpoint
- PLK1
- Polo-like kinase 1
- PP1
- protein phosphatase 1
- H3T3
- threonine 3 on histone H3 tail
- LRR
- leucine-rich repeat
- NEBD
- nuclear envelope breakdown
- AD
- the GAL4 activation domain
- BD
- the GAL4 DNA-binding domain
- CBB
- Coomassie Brilliant Blue
- PMP
- protein metallophosphatases
- PBD
- Polo-binding domain
- CFP
- cyan fluorescent protein
- CENP
- centromere-associated protein
- KD
- kinase-deficient mutant.
References
- 1. Holland A. J., and Cleveland D. W. (2009) Boveri revisited: chromosomal instability, aneuploidy and tumorigenesis. Nat. Rev. Mol. Cell Biol. 10, 478–487 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Lampson M. A., Renduchitala K., Khodjakov A., and Kapoor T. M. (2004) Correcting improper chromosome-spindle attachments during cell division. Nat. Cell Biol. 6, 232–237 [DOI] [PubMed] [Google Scholar]
- 3. Welburn J. P., Vleugel M., Liu D., Yates J. R. 3rd, Lampson M. A., Fukagawa T., and Cheeseman I. M. (2010) Aurora B phosphorylates spatially distinct targets to differentially regulate the kinetochore-microtubule interface. Mol. Cell 38, 383–392 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Zannini L., Delia D., and Buscemi G. (2014) CHK2 kinase in the DNA damage response and beyond. J. Mol. Cell Biol. 6, 442–457 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Pinsky B. A., Kotwaliwale C. V., Tatsutani S. Y., Breed C. A., and Biggins S. (2006) Glc7/protein phosphatase 1 regulatory subunits can oppose the Ipl1/Aurora protein kinase by redistributing Glc7. Mol. Cell Biol. 26, 2648–2660 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Kim Y., Holland A. J., Lan W., and Cleveland D. W. (2010) Aurora kinases and protein phosphatase 1 mediate chromosome congression through regulation of CENP-E. Cell 142, 444–455 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Liu D., Vleugel M., Backer C. B., Hori T., Fukagawa T., Cheeseman I. M., and Lampson M. A. (2010) Regulated targeting of protein phosphatase 1 to the outer kinetochore by KNL1 opposes Aurora B kinase. J. Cell Biol. 188, 809–820 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Ruchaud S., Carmena M., and Earnshaw W. C. (2007) Chromosomal passengers: conducting cell division. Nat. Rev. Mol. Cell Biol. 8, 798–812 [DOI] [PubMed] [Google Scholar]
- 9. Tanaka T. U., Rachidi N., Janke C., Pereira G., Galova M., Schiebel E., Stark M. J., and Nasmyth K. (2002) Evidence that the Ipl1-Sli15 (Aurora kinase-INCENP) complex promotes chromosome bi-orientation by altering kinetochore-spindle pole connections. Cell 108, 317–329 [DOI] [PubMed] [Google Scholar]
- 10. DeLuca K. F., Lens S. M., and DeLuca J. G. (2011) Temporal changes in Hec1 phosphorylation control kinetochore-microtubule attachment stability during mitosis. J. Cell Sci. 124, 622–634 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Musacchio A., and Salmon E. D. (2007) The spindle-assembly checkpoint in space and time. Nat. Rev. Mol. Cell Biol. 8, 379–393 [DOI] [PubMed] [Google Scholar]
- 12. Mo F., Zhuang X., Liu X., Yao P. Y., Qin B., Su Z., Zang J., Wang Z., Zhang J., Dou Z., Tian C., Teng M., Niu L., Hill D. L., Fang G., Ding X., Fu C., and Yao X. (2016) Acetylation of Aurora B by TIP60 ensures accurate chromosomal segregation. Nat. Chem. Biol. 12, 226–232 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Li H., Liu X. S., Yang X., Wang Y., Wang Y., Turner J. R., and Liu X. (2010) Phosphorylation of CLIP-170 by Plk1 and CK2 promotes timely formation of kinetochore-microtubule attachments. EMBO J. 29, 2953–2965 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Zitouni S., Nabais C., Jana S. C., Guerrero A., and Bettencourt-Dias M. (2014) Polo-like kinases: structural variations lead to multiple functions. Nat. Rev. Mol. Cell Biol. 15, 433–452 [DOI] [PubMed] [Google Scholar]
- 15. Strebhardt K., and Ullrich A. (2006) Targeting polo-like kinase 1 for cancer therapy. Nat. Rev. Cancer 6, 321–330 [DOI] [PubMed] [Google Scholar]
- 16. Zhou L., Tian X., Zhu C., Wang F., and Higgins J. M. (2014) Polo-like kinase-1 triggers histone phosphorylation by Haspin in mitosis. Embo. Rep. 15, 273–281 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Ghenoiu C., Wheelock M. S., and Funabiki H. (2013) Autoinhibition and Polo-dependent multisite phosphorylation restrict activity of the histone H3 kinase Haspin to mitosis. Mol. Cell 52, 734–745 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Wang F., Dai J., Daum J. R., Niedzialkowska E., Banerjee B., Stukenberg P. T., Gorbsky G. J., and Higgins J. M. (2010) Histone H3 Thr-3 phosphorylation by haspin positions Aurora B at centromeres in mitosis. Science 330, 231–235 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Kelly A. E., Ghenoiu C., Xue J. Z., Zierhut C., Kimura H., and Funabiki H. (2010) Survivin reads phosphorylated histone H3 threonine 3 to activate the mitotic kinase Aurora B. Science 330, 235–239 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Chu Y., Yao P. Y., Wang W., Wang D., Wang Z., Zhang L., Huang Y., Ke Y., Ding X., and Yao X. (2011) Aurora B kinase activation requires survivin priming phosphorylation by PLK1. J Mol. Cell. Biol. 3, 260–267 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Heroes E., Lesage B., Görnemann J., Beullens M., Van Meervelt L., and Bollen M. (2013) The PP1 binding code: A molecular-lego strategy that governs specificity. FEBS J. 280, 584–595 [DOI] [PubMed] [Google Scholar]
- 22. Trinkle-Mulcahy L., Andrews P. D., Wickramasinghe S., Sleeman J., Prescott A., Lam Y. W., Lyon C., Swedlow J. R., and Lamond A. I. (2003) Time-lapse imaging reveals dynamic relocalization of PP1γ throughout the mammalian cell cycle. Mol. Biol. Cell 14, 107–117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Posch M., Khoudoli G. A., Swift S., King E. M., Deluca J. G., and Swedlow J. R. (2010) Sds22 regulates Aurora B activity and microtubule-kinetochore interactions at mitosis. J. Cell Biol. 191, 61–74 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Meadows J. C., Shepperd L. A., Vanoosthuyse V., Lancaster T. C., Sochaj A. M., Buttrick G. J., Hardwick K. G., and Millar J. B. (2011) Spindle checkpoint silencing requires association of PP1 to both Spc7 and kinesin-8 motors. Dev. Cell 20, 739–750 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Rosenberg J. S., Cross F. R., and Funabiki H. (2011) KNL1/Spc105 recruits PP1 to silence the spindle assembly checkpoint. Curr. Biol. 21, 942–947 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Ohkura H., and Yanagida M. (1991) S. pombe gene sds22+ essential for a midmitotic transition encodes a leucine-rich repeat protein that positively modulates protein phosphatase-1. Cell 64, 149–157 [DOI] [PubMed] [Google Scholar]
- 27. Dinischiotu A., Beullens M., Stalmans W., and Bollen M. (1997) Identification of sds22 as an inhibitory subunit of protein phosphatase-1 in rat liver nuclei. FEBS Lett. 402, 141–144 [DOI] [PubMed] [Google Scholar]
- 28. Renouf S., Beullens M., Wera S., Van Eynde A., Sikela J., Stalmans W., and Bollen M. (1995) Molecular cloning of a human polypeptide related to yeast sds22, a regulator of protein phosphatase-1. FEBS Lett. 375, 75–78 [DOI] [PubMed] [Google Scholar]
- 29. Ceulemans H., Vulsteke V., De Maeyer M., Tatchell K., Stalmans W., and Bollen M. (2002) Binding of the concave surface of the Sds22 superhelix to the α4/α5/α6-triangle of protein phosphatase-1. J. Biol. Chem. 277, 47331–47337 [DOI] [PubMed] [Google Scholar]
- 30. Wurzenberger C., Held M., Lampson M. A., Poser I., Hyman A. A., and Gerlich D. W. (2012) Sds22 and repo-man stabilize chromosome segregation by counteracting Aurora B on anaphase kinetochores. J. Cell Biol. 198, 173–183 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Rodrigues N. T., Lekomtsev S., Jananji S., Kriston-Vizi J., Hickson G. R., and Baum B. (2015) Kinetochore-localized PP1-Sds22 couples chromosome segregation to polar relaxation. Nature 524, 489–492 [DOI] [PubMed] [Google Scholar]
- 32. Eiteneuer A., Seiler J., Weith M., Beullens M., Lesage B., Krenn V., Musacchio A., Bollen M., and Meyer H. (2014) Inhibitor-3 ensures bipolar mitotic spindle attachment by limiting association of SDS22 with kinetochore-bound protein phosphatase-1. EMBO J. 33, 2704–2720 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Xue Y., Ren J., Gao X., Jin C., Wen L., and Yao X. (2008) GPS 2.0, a tool to predict kinase-specific phosphorylation sites in hierarchy. Mol. Cell. Proteomics 7, 1598–1608 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Han G., Ye M., Zhou H., Jiang X., Feng S., Jiang X., Tian R., Wan D., Zou H., and Gu J. (2008) Large-scale phosphoproteome analysis of human liver tissue by enrichment and fractionation of phosphopeptides with strong anion exchange chromatography. Proteomics 8, 1346–1361 [DOI] [PubMed] [Google Scholar]
- 35. Olsen J. V., Vermeulen M., Santamaria A., Kumar C., Miller M. L., Jensen L. J., Gnad F., Cox J., Jensen T. S., Nigg E. A., Brunak S., and Mann M. (2010) Quantitative phosphoproteomics reveals widespread full phosphorylation site occupancy during mitosis. Sci. Signal. 3, ra3. [DOI] [PubMed] [Google Scholar]
- 36. Rigbolt K. T., Prokhorova T. A., Akimov V., Henningsen J., Johansen P. T., Kratchmarova I., Kassem M., Mann M., Olsen J. V., and Blagoev B. (2011) System-wide temporal characterization of the proteome and phosphoproteome of human embryonic stem cell differentiation. Sci. Signal. 4, rs3. [DOI] [PubMed] [Google Scholar]
- 37. Bian Y., Song C., Cheng K., Dong M., Wang F., Huang J., Sun D., Wang L., Ye M., and Zou H. (2014) An enzyme assisted RP-RPLC approach for in-depth analysis of human liver phosphoproteome. J. Proteomics 96, 253–262 [DOI] [PubMed] [Google Scholar]
- 38. Liu D., Vader G., Vromans M. J., Lampson M. A., and Lens S. M. (2009) Sensing chromosome bi-orientation by spatial separation of Aurora B kinase from kinetochore substrates. Science 323, 1350–1353 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Xia P., Zhou J., Song X., Wu B., Liu X., Li D., Zhang S., Wang Z., Yu H., Ward T., Zhang J., Li Y., Wang X., Chen Y., Guo Z., and Yao X. (2014) Aurora A orchestrates entosis by regulating a dynamic MCAK-TIP150 interaction. J Mol. Cell Biol. 6, 240–254 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Liu D., Davydenko O., and Lampson M. A. (2012) Polo-like kinase-1 regulates kinetochore-microtubule dynamics and spindle checkpoint silencing. J. Cell Biol. 198, 491–499 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Dai J., Sultan S., Taylor S. S., and Higgins J. M. (2005) The kinase haspin is required for mitotic histone H3 Thr 3 phosphorylation and normal metaphase chromosome alignment. Genes Dev. 19, 472–488 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Dou Z., Liu X., Wang W., Zhu T., Wang X., Xu L., Abrieu A., Fu C., Hill D. L., and Yao X. (2015) Dynamic localization of Mps1 kinase to kinetochores is essential for accurate spindle microtubule attachment. Proc. Natl. Acad. Sci. U.S.A. 112, E4546–E4555 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Santaguida S., Tighe A., D'Alise A. M., Taylor S. S., and Musacchio A. (2010) Dissecting the role of MPS1 in chromosome biorientation and the spindle checkpoint through the small molecule inhibitor reversine. J. Cell Biol. 190, 73–87 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Meiselbach H., Sticht H., and Enz R. (2006) Structural analysis of the protein phosphatase 1 docking motif: molecular description of binding specificities identifies interacting proteins. Chem. Biol. 13, 49–59 [DOI] [PubMed] [Google Scholar]
- 45. Wang X., Zhuang X., Cao D., Chu Y., Yao P., Liu W., Liu L., Adams G., Fang G., Dou Z., Ding X., Huang Y., Wang D., and Yao X. (2012) Mitotic regulator SKAP forms a link between kinetochore core complex KMN and dynamic spindle microtubules. J. Biol. Chem. 287, 39380–39390 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Yuan K., Li N., Jiang K., Zhu T., Huo Y., Wang C., Lu J., Shaw A., Thomas K., Zhang J., Mann D., Liao J., Jin C., and Yao X. (2009) PinX1 is a novel microtubule-binding protein essential for accurate chromosome segregation. J. Biol. Chem. 284, 23072–23082 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Shao H., Huang Y., Zhang L., Yuan K., Chu Y., Dou Z., Jin C., Garcia-Barrio M., Liu X., and Yao X. (2015) Spatiotemporal dynamics of Aurora B-PLK1-MCAK signaling axis orchestrates kinetochore bi-orientation and faithful chromosome segregation. Sci. Rep. 5, 12204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Xia P., Wang Z., Liu X., Wu B., Wang J., Ward T., Zhang L., Ding X., Gibbons G., Shi Y., and Yao X. (2012) EB1 acetylation by P300/CBP-associated factor (PCAF) ensures accurate kinetochore-microtubule interactions in mitosis. Proc. Natl. Acad. Sci. U.S.A. 109, 16564–16569 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Zhu L., Wang Z., Wang W., Wang C., Hua S., Su Z., Brako L., Garcia-Barrio M., Ye M., Wei X., Zou H., Ding X., Liu L., Liu X., and Yao X. (2015) Mitotic protein CSPP1 interacts with CENP-H protein to coordinate accurate chromosome oscillation in mitosis. J. Biol. Chem. 290, 27053–27066 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Jiang H., Wang W., Zhang Y., Yao W. W., Jiang J., Qin B., Yao W. Y., Liu F., Wu H., Ward T. L., Chen C. W., Liu L., Ding X., Liu X., and Yao X. (2015) Cell polarity kinase MST4 cooperates with cAMP-dependent kinase to orchestrate histamine-stimulated acid secretion in gastric parietal cells. J. Biol. Chem. 290, 28272–28285 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.