Abstract
Drug resistance inevitably limits the efficacy of all targeted therapies including tyrosine kinase inhibitors (TKIs). Understanding the biological underpinnings of TKI resistance is key to the successful development of future therapeutic strategies. Traditionally, mechanisms of TKI resistance have been viewed under a dichotomous lens. Tumor cells are TKI-sensitive or TKI-refractory, exhibit intrinsic or acquired resistance, and accumulate alterations within or outside the target to promote their survival. Such classifications facilitate our comprehension of an otherwise complex biology, but are likely an oversimplification. Recent studies underscore the multifaceted, genetically heterogeneous nature of TKI resistance, which evolves dynamically with changes in therapy. In this Review, we provide a broad framework for understanding the diverse mechanisms of resistance at play in oncogene-driven lung cancers.
Keywords: Resistance, tyrosine kinase inhibitor (TKI), NSCLC
TKI resistance: a pervasive challenge
In 2002, I.B. Weinstein defined the phenomenon of ‘oncogene addiction,’ whereby cancer cells become excessively dependent on a particular ‘driver’ alteration for their survival [1]. Cancers with these dependencies exhibit exquisite vulnerability to drugs that inhibit the drivers, so-called ‘targeted therapies.’ The past decade has witnessed numerous successes in targeting specific molecular subsets of cancer across a spectrum of different human malignancies, most notably non-small-cell lung cancer (NSCLC). Somatic activating mutations in epidermal growth factor receptor (EGFR) were the first driver alterations characterized in NSCLC, and are found in 10–15% of non-Asian patients [2]. These mutations confer sensitivity to small molecule tyrosine kinase inhibitors (TKIs) of EGFR, resulting in high response rates and prolonged progression-free survival [3–8]. Rearrangements involving the anaplastic lymphoma kinase (ALK) gene are identified in 3–7% of NSCLC, and predict a comparable degree of clinical benefit from ALK-TKIs [9]. Genotype-directed therapy is now the standard of care in advanced NSCLC and has led to improvements in overall survival [10, 11].
The inevitable barrier that limits the effectiveness of TKI therapy and tempers enthusiasm is the issue of resistance—today’s pervasive challenge for long-term disease control. Cancer is at its core a microcosm of evolution. Its survival is driven by genetic diversity and longitudinal accumulation of mutations, influenced by the selective pressures of TKI therapy. These rudimentary yet intricate principles underlie the refractory nature of TKI resistance, which is traditionally categorized as primary (intrinsic) or secondary (acquired). In primary resistance, patients lack any treatment response to targeted therapy. In secondary resistance, patients initially achieve some clinical benefit, followed by disease progression. With the discovery of each oncogenic driver and targeted inhibitor, a growing number and diversity of resistance mechanisms are being defined. In this Review, we highlight the major concepts that have emerged from studying mechanisms of TKI resistance in NSCLC.
Overview of oncogenic drivers in NSCLC
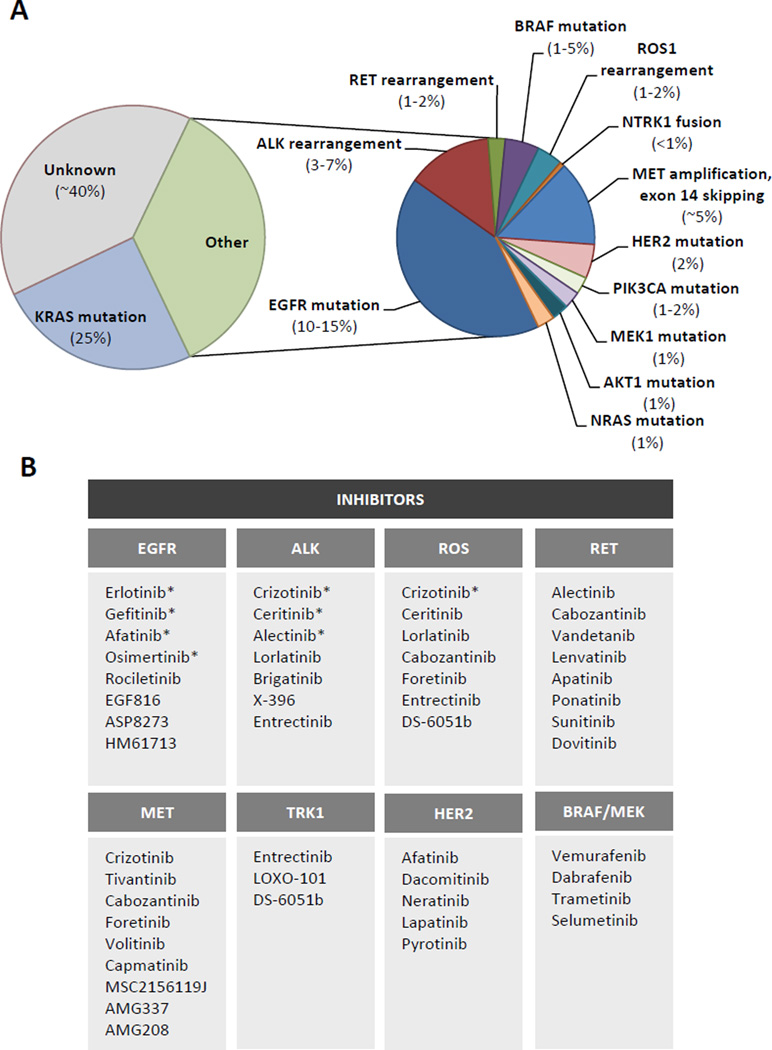
In addition to EGFR and ALK, several other oncogenic drivers have been discovered in NSCLC. In lung adenocarcinoma, the most common histologic subtype of NSCLC, actionable drivers are present in over 20–25% of patients [10, 11]. These alterations include ROS1 rearrangements, BRAF mutations, RET rearrangements, NTRK1 rearrangements, MET amplification and exon 14 skipping mutations, and HER2 mutations (Figure 1) [10–17]. An in-depth discussion of each driver and its signaling pathways is beyond the scope of this Review. Kinase inhibitors targeting each of these oncogenes are either the standard of care or undergoing active development in the clinic, providing a fertile ground for investigations of drug resistance.
Figure 1. Oncogenic Drivers in Lung Adenocarcinoma.
(A) The distribution of known oncogenic driver alterations in lung adenocarcinoma is shown, with estimated percentages for each driver. For an estimated 40% of lung adenocarcinomas, the underlying genetic alteration(s) remain unknown. Approximately 25% of lung adenocarcinomas carry an activating KRAS mutation, for which targeted therapies are not yet available. (B) There are many tyrosine kinase inhibitors (TKIs) currently in use in the clinic or undergoing active development, which target the validated oncogenic drivers in NSCLC. Examples are listed. Asterisks indicate TKIs which have been approved by the Food and Drug Administration for use in patients with NSCLC harboring the indicated genetic alterations.
Primary resistance
A review of the randomized trials using EGFR- or ALK-TKIs in the first-line setting for advanced EGFR- or ALK-positive NSCLC, respectively, suggests that primary resistance—when defined as “progressive disease as the best response”—occurs in 4–10% of newly diagnosed patients [3–9]. Mechanisms underlying intrinsic TKI resistance are not fully elucidated but include non-sensitizing alterations within the target. For example, in-frame exon 20 insertions account for 4–10% of EGFR mutations [18–20]. These activate EGFR signaling in vitro, but do not confer sensitivity to first-generation EGFR-TKIs [18–21]. EGFR T790M, the gatekeeper mutation, is a well-described mechanism of acquired resistance to first-generation EGFR-TKIs (discussed below). In treatment-naïve patients with classic activating EGFR mutations, pre-existing EGFR T790M-mutant clones may promote intrinsic resistance at a certain threshold of allelic frequency [22, 23]. The reported frequency of pre-existing T790M mutation has varied widely in the literature depending on the detection method (range, <10% to 65%) [22–24].
Genetic alterations outside the target kinase can also contribute to decreased de novo sensitivity to TKIs. MET amplification has been reported in EGFR-mutant NSCLC prior to TKI exposure [25]. BIM, or BCL2L11, is a member of the BCL2 family that mediates apoptosis triggered by several TKIs. Recent studies showed that lower mRNA expression levels of BIM correlate with inferior efficacy of EGFR-TKIs [24, 26]. Additionally, in Asian patients, a BIM polymorphism resulting in isoforms that lack the pro-apoptotic BH3 domain was associated with poor response to EGFR-TKIs [27]. A similar role for BIM has not yet been described in other oncogene-driven lung cancers. Another potential modulator of intrinsic NSCLC sensitivity to EGFR-TKIs is NF-κB. In EGFR-mutant lung cancer cells, RNA interference-mediated knockdown of components of the NF-κB pathway enhanced erlotinib sensitivity, and similar effects were seen with pharmacologic inhibition of NF-κB [28]. In a cohort of 52 patients with EGFR-mutant lung cancer treated with erlotinib, higher expression of the NF-κB inhibitor IκB was associated with increased TKI response and survival [28], suggesting that an approach combining EGFR inhibition with NF-κB pathway inhibition could potentially improve responses to EGFR-TKIs in the clinic.
Alterations in genes leading to the induction of epithelial-to-mesenchymal transition (EMT) have also been reported as a cause of primary TKI resistance [29, 30]. For example, Park et al. demonstrated that overexpression of Cripto-1, an EGF-CFC family member, could confer erlotinib resistance in cell line and mouse xenograft models which was mediated by SRC activation [30]. In 85 NSCLC patients with EGFR-sensitizing mutations, higher Cripto-1 expression levels were associated with intrinsic resistance to EGFR-TKIs [30]. Further work is needed to validate these findings and establish whether Cripto-1 may be a major modulator of EGFR-TKI sensitivity in the clinic.
Finally, false-positive genotyping may account for at least a few cases of apparent primary resistance. In the case of ALK rearrangement, the gold standard diagnostic assay has been a break-apart fluorescence in-situ hybridization (FISH) assay. The most common ALK rearrangement, EML4-ALK, results from a small intrachromosomal inversion event, leading in some cases to a subtle splitting of the 5’ and 3’ signals [31]. This can lead to significant inter-observer variability and potential false-positives [31]. Secondary confirmation of ALK positivity using immunohistochemistry (IHC) or next-generation sequencing (NGS) can thus be helpful.
Secondary resistance
Extensive efforts in recent years led to the elucidation of multiple mechanisms of acquired TKI resistance. Broadly speaking, the general categories of resistance mechanisms include secondary alterations within the target, activation of an alternative (i.e. bypass) signaling pathway or downstream effector(s), and phenotypic transformation. Conceptually, all can be viewed as manifestations of evolution of cancer cells under the selective pressure of targeted therapies. Understanding each mechanism is paramount to developing therapeutic strategies to overcome, or even prevent, TKI resistance.
Mutations in the target
Secondary somatic mutations within the target kinase enable its persistent activation despite the presence of the inhibitor. In general, these alterations hinder the kinase’s ability to bind the drug or alter the kinase’s conformation when non-contact residues are involved. The classic example is the gatekeeper ABL T315I mutation in Philadelphia chromosome-positive chronic myelogenous leukemia (CML). This mutation was first described in CML patients treated with imatinib, and has since been found to confer resistance to all approved ABL-TKIs prior to ponatinib [32]. T315I affects a conserved amino acid within the catalytic cleft that determines the relative accessibility of inhibitors to a hydrophobic pocket, resulting in steric interference with the binding of ABL-TKIs, but preserved kinase activity [32].
In EGFR-mutant NSCLC, the T790M gatekeeper mutation in exon 20 of EGFR was one of the earliest TKI-resistance mechanisms reported. It represents the dominant cause of resistance to erlotinib or gefitinib, seen in 50–60% of cases (Table 1) [33–35]. Interestingly, the T790M substitution appears to render resistance primarily by enhancing the kinase affinity for ATP rather than by causing steric hindrance [36]. Other non-T790M resistance mutations within EGFR are rarely observed in the clinic. These include T854A, D761Y, and L747S [34–39]. In vitro, the non-T790M mutations confer less potent resistance, and the mechanisms by which they confer resistance are not as well understood.
Table 1.
Known Secondary Resistance Mutations in EGFR, ALK, ROS1, and NTRK1.†
| Mutation | EGFR | ALK | ROS1 | NTRK1 |
|---|---|---|---|---|
| Gatekeeper | T790M | L1196M | NR | NR |
| ATP-binding pocket | T854A | G1269A/S V1180L |
NR | G667C* |
| Solvent front | NR | G1202R D1203N S1206C/Y |
G2032R D2033N |
G595R* |
| Covalent drug binding residue | C797S | NR | NR | NR |
| Other known mutations | L747S D761Y |
G1123S I1151Tins L1152P/R C1156Y/T I1171T/N F1174C/L/V L1198F F1245C |
Only those mutations clinically identified in patients are shown.
Recently, studies have provided insights into resistance mutations against third-generation EGFR inhibitors that are designed to target T790M, including osimertinib, rociletinib, and WZ4002. These compounds bind EGFR covalently through the cysteine residue 797 (C797S). A novel EGFR C797S mutation, which alters the drug contact site, has now been observed in about 20% of patients who progress on osimertinib [40–43]. Notably, in vitro studies by Niederst et al. suggest that the allelic configuration of the C797S and T790M mutations may have an impact on the responsiveness of EGFR-mutant lung cancer cells to select groups of EGFR-TKIs [43]. Those cells with the C797S occurring in trans (on a different allele) with T790M were resistant to third-generation EGFR-TKIs, but retained sensitivity to combination therapy using a first-generation and third-generation TKI. In contrast, when the C797S occurred in cis (on the same allele) with T790M, the cells were resistant to all EGFR-TKIs [43]. Furthermore, if the cells acquired C797S in the absence of T790M (which has not been reported clinically but may be seen in the future if patients are treated with a third-generation EGFR inhibitor upfront), they were resistant to third-generation EGFR-TKIs but responded to first-generation inhibitors [43]. These findings may have future clinical implications in guiding the sequencing of EGFR-TKIs.
For ALK, numerous kinase domain mutations have been reported in patients with acquired TKI resistance (Table 1). The ALK-L1196M gatekeeper mutation confers resistance through steric interference, and it occurs at a lower prevalence compared to EGFR-T790M [44–47]. G1269A is the second most common ALK resistance mutation that causes resistance by interfering with TKI binding. Mutations I1151Tins, L1152R, and C1156Y are located near the αC-helix outside the drug-binding region, and may cause TKI resistance by increasing catalytic activity [45–47]. Solvent front mutations G1202R and S1206Y alter residues in the solvent-exposed region of ALK. These mutations lower the drug binding affinity [45–47]. The G1202R mutation in particular is refractory to most ALK-TKIs including crizotinib, ceritinib, alectinib, and brigatinib but is overcome by the third-generation TKI lorlatinib [48–50].
Most recently, the ALK L1198F resistance mutation was discovered in a patient who relapsed first on crizotinib due to an ALK C1156Y mutation and subsequently was treated with lorlatinib [51]. This mutation changes the very residue used to enhance the selectivity of lorlatinib for ALK over other tyrosine kinases, again demonstrating the inherent mutational capacity of cancer cells under selective pressures. Interestingly, with the addition of L1198F to C1156Y (and to other crizotinib-resistance mutations), lorlatinib-resistant cells could be re-sensitized to crizotinib [51]. This finding was demonstrated in cellular and biochemical assays, and also in the patient who re-responded to crizotinib after failing lorlatinib. This case highlights the clinical utility of multiple sequential ALK-TKIs and the importance of serial biopsies to guide their selection.
It is worth noting that there appears to be a much more diverse spectrum of resistance mutations identified in ALK compared to EGFR, where the EGFR T790M mutation is essentially the sole, clinically predominant resistance mechanism after failure of first- and second-generation EGFR-TKIs. This may be due in part to the presence of a pre-existing activating mutation in EGFR, which constrains the kinase’s ability to acquire additional mutations while preserving its function [47]. On the other hand, multiple different secondary mutations in ALK have been reported in resistant patient specimens. As a whole, these mutations account for only about a third of crizotinib-resistant cases. Interestingly, a narrower and distinct spectrum of resistance mutations is seen with each next-generation ALK-TKI (Gainor et al., submitted). However, in this setting, secondary resistance mutations are seen in 50–60% of patients, similar to the frequency of EGFR T790M in EGFR-TKI resistance cases (Table 2) [46, 47]. The lower prevalence of on-target alterations seen with crizotinib may reflect less potent target inhibition compared to next-generation ALK-TKIs, or EGFR-TKIs in EGFR-mutant NSCLC.
Table 2.
Mechanisms of Acquired Resistance in EGFR- and ALK-Positive NSCLC Treated with Tyrosine Kinase Inhibitors (TKIs).†
| Category | Alteration | Estimated frequency (%) | Refs |
|---|---|---|---|
| Resistance to EGFR-TKI | EGFR target alteration | ~60 | |
| T790M | 50–60 (for 1st-generation EGFR-TKI) | [33–35] | |
| D761Y, T854A, L747S | 1–2 | [34–39] | |
| C797S | ~20 (for 3rd-generation EGFR-TKI) | [40–43] | |
| EGFR amplification | 8–10 | [33,39] | |
| Bypass signaling tracks | ~20 | ||
| MET amplification | 5–22 | [33,34,59] | |
| HGF overexpression | 1 of 2 cases reported | [60] | |
| HER2 amplification | 12 | [33,34] | |
| FGFR3 activation | 1 case reported | [63] | |
| BRAF mutations | 1 | [64] | |
| CRKL amplification | 1 of 11 cases reported | [65] | |
| NF1 reduced expression | 4 of 10 cases reported | [66] | |
| Phenotypic changes | 3–10 | ||
| Transformation to SCLC | 3–10 | [33–35] | |
| Unknown mechanism | 10–20 | ||
| Resistance to ALK-TKI | ALK target alteration | ~28–46 | |
| Secondary mutations in ALK | 22–36 | [44–47] | |
| ALK amplification | 7–18 | [46,47] | |
| Bypass signaling tracks | ~40–50 | ||
| EGFR activation | Up to 44 | [45–47] | |
| c-KIT amplification and SCF overexpression |
15 | [46] | |
| IGF-1R activation | 4 of 5 cases | [62] | |
| MEK1 mutations | 1 case reported | [63] | |
| PIK3CA mutations | 1 case reported | [63] | |
| MET amplification | 1 case reported (for alectinib) | [61] | |
| SRC activation | Unknown % | [63] | |
| Phenotypic changes | <5 | ||
| Transformation to SCLC | < 5 (case reports) | [72–74] | |
| Unknown mechanism | ~15–30 |
Only those mechanisms clinically identified in patients are shown.
Abbreviations: EGFR, epidermal growth factor receptor; HGF, hepatocyte growth factor; HER2, human epidermal growth factor receptor; FGFR3, fibroblast growth factor receptor 3; CRKL, V-Crk avian sarcoma virus CT10 oncogene homolog-like; NF1, neurofibromin 1; SCLC, small cell lung cancer; ALK, anaplastic lymphoma kinase; SCF, stem cell factor; IGF-1R, insulin-like growth factor 1 receptor; MEK1, mitogen-activated protein kinase kinase; PIK3CA, phosphatidylinositol-4,5-bisphosphonate 3-kinase, catalytic subunit alpha.
Secondary mutations in the target are now being reported in other oncogene-driven lung cancers as well. Solvent front mutations in ROS1, G2032R (analogous to ALK G1202R) and D2033N cause crizotinib resistance in ROS1-rearranged NSCLC (Table 1) [52, 53]. The ROS1 gatekeeper mutation L2026M confers resistance to crizotinib in vitro [54] but has not yet been observed clinically. For NTRK1, secondary resistance mutations against entrectinib were recently described in colorectal cancer, including the G595R solvent front mutation (analogous to ALK G1202R), and the G667C mutation (analogous to ALK G1269A and EGFR T854A) [55].
Target amplification
Increased gene dosage through target amplification is a well-known cause of acquired TKI resistance. EGFR amplification has been identified in tumors resistant to EGFR-TKIs [33, 39]. In one series of 37 patients with TKI-resistant EGFR-mutant NSCLC, three patients (8%) acquired EGFR amplification post-treatment [33]. All three patients, however, also acquired a T790M mutation with selective amplification of the T790M allele in two of these patients—suggesting that EGFR amplification may serve to enhance the resistance phenotype of the gatekeeper mutation [33]. Intriguingly, loss of the activating EGFR has also been reported in cases of EGFR-TKI resistance [56, 57]. However, in these cases, there was concomitant reactivation of a downstream survival pathway [56] or histologic transformation [57], which can abrogate dependence on the target oncogene for survival. Therefore, it is more likely that the loss of EGFR is a reflection of this state of target independence than a direct cause of acquired resistance.
Crizotinib resistance due to ALK amplification has similarly been observed in 7–18% of cases, and can occur alone or together with an ALK resistance mutation [46, 47]. In one in vitro model, partial crizotinib resistance acquired through wild-type EML4-ALK amplification was then followed by a higher degree of resistance conferred by the emergence of ALK L1196M, suggesting a stepwise evolution of acquired resistance [58]. In all of these examples, tumor cells remain dependent on the target kinase.
Bypass signaling pathway activation
Tumor cells orchestrate a network of signaling pathways that sustain survival. When a dominant pathway is inhibited by a TKI, diversion to parallel signaling pathways can allow cells to reactivate critical downstream effectors, allowing continued tumor survival and growth. As an example, MET amplification was one of the first described mechanisms of EGFR-TKI resistance in 2007 [59], and has since been identified in 5–10% of resistant patients [33, 34]. MET amplification leads to phosphorylation of HER3 and reactivation of PI3K/AKT signaling, bypassing EGFR. Overexpression of the MET ligand, hepatocyte growth factor HGF, similarly results in EGFR-TKI resistance [60].
Oncogenic drivers in NSCLC are essentially mutually exclusive, but can operate in a complementary manner in the resistant setting. MET amplification mediating EGFR resistance is one such example. In ALK-rearranged NSCLC, crizotinib resistance is mediated by EGFR activation in up to 44% of cases, although caused by upregulation of the receptor and/or overexpression of EGFR ligands rather than by genetic alterations of EGFR [45–47]. MET amplification has also been reported as a mechanism of resistance to the next-generation ALK inhibitor alectinib in ALK-rearranged NSCLC [61].
Other examples of bypass signaling which have been validated clinically in patient tumor samples include (Table 2): c-KIT amplification and ligand SCF overexpression, IGF-1R activation, and SRC signaling upregulation in ALK-TKI resistance [46, 62, 63]; HER2 amplification, FGFR3 activation, BRAF mutations, CRKL amplification, and NF1 downregulation in EGFR-TKI resistance [33, 34, 63–66]; and EGFR activation in crizotinib-resistant ROS1-positive NSCLC [67]. A number of additional resistance mechanisms have been proposed based upon preclinical studies but a description of these is beyond the scope of this Review. In all of these cases, the bypass track reactivates a key downstream pathway in NSCLC, namely the PI3K/AKT/mTOR or RAF/MEK/ERK pathway [14].
Downstream effector activation
Tumor cells can also acquire alterations in downstream signaling effectors leading to drug resistance. A salient example occurs in BRAF-mutant melanomas, which develop resistance to BRAF inhibitor monotherapy through reactivation of the MAPK pathway [68]. MAPK reactivation has also been identified as a resistance mechanism to dabrafenib in a patient with BRAF-mutant NSCLC [69]. These preclinical findings provided the rationale for the development of combinatorial regimens of BRAF and MEK inhibitors in both melanoma and NSCLC harboring activating BRAF mutations.
MAPK activation has similarly been identified as a predominant resistance mechanism in both EGFR- and ALK-positive NSCLC after treatment with EGFR- and ALK-TKIs, respectively. This is of particular interest, as multiple combination regimens of EGFR-TKI or ALK-TKI with a MEK inhibitor have entered early phase testing based on preclinical findings. For example, in EGFR-mutant lung cancer, resistance to EGFR inhibitors can be acquired through reactivation of ERK1/2. This resistance is overcome by treatment with MEK inhibitors such as trametinib [70]. Trickler et al. additionally showed that acquired resistance to EGFR-TKIs could be prevented by upfront combination therapy using an EGFR inhibitor plus MEK inhibitor in EGFR-mutant lung cancer cell line and mouse models [70].
Recent work using patient-derived cell lines identified the MEK1 activating mutation, MAP2K1 K57N, as a driver of resistance to ALK inhibition in a case of ALK-rearranged NSCLC [63]. In this model, dual blockade of MEK and ALK was effective in overcoming resistance. Primary dependency of ALK on MEK as the dominant effector oncoprotein in ALK-rearranged lung adenocarcinomas may account for this finding, as proposed by Hrustanovic et al. [71]. In their in vitro assays, inhibition of MEK—but not PI3K/AKT or JAK—could recapitulate the growth suppression phenotype conferred by ALK inhibition. Conversely, constitutive activation of MEK could rescue the cells from ALK inhibition [71]. Notably, treatment of the ALK-positive NSCLC cells with both an ALK inhibitor (ceritinib or crizotinib) and MEK inhibitor (trametinib) suppressed the emergence of resistance in cell line and mouse xenograft models [71], providing a compelling rationale for the clinical evaluation of this combination approach.
Phenotypic and histologic transformation
Cancers may acquire TKI resistance through phenotypic transformation. Change in histology from adenocarcinoma to small-cell lung cancer (SCLC) is observed in 3–10% of cases of resistance to EGFR inhibitors, including third-generation EGFR-TKIs (Table 2) [33–35]. Sequencing of EGFR from repeat biopsies has revealed that the activating mutation in EGFR seen in the original adenocarcinoma is retained in SCLC, suggesting a phenotypic evolution of tumor cells rather than a de novo SCLC [33–35]. Several case reports have now also been published of ALK-rearranged NSCLC patients with SCLC transformation after progression on crizotinib or alectinib [72–74].
How histologic transformation mediates TKI resistance remains to be elucidated. In one study by Niederst et al., RB loss was detected in 100% of the 10 EGFR-mutant NSCLC patient tumor samples that had transformed to SCLC at the time of drug resistance (versus 11% of the 9 cases that retained NSCLC histology) [75]. Furthermore, RNA expression profiling and hierarchical clustering analysis showed the resistant, SCLC-transformed EGFR-mutant lung cancer cells to be more closely related to classical SCLC than other, TKI-resistant NSCLC cell lines [75]. Hence, this subset of TKI-resistant lung cancers may adopt the genetic (and perhaps epigenetic) features of classical SCLC, which is generally insensitive to EGFR-TKI therapy.
Another mediator of TKI resistance is phenotypic change to NSCLC via epithelial-to-mesenchymal transition (EMT), an evolutionarily conserved program of transdifferentiation and a driver of tumor invasiveness [76]. EMT was observed clinically in 5% of patients with EGFR-TKI resistance [33], and reported in cell line models of ALK-TKI resistance [77]. This change may involve the activation of AXL through augmented expression of AXL or its ligand GAS6, observed in 20% and 25% of EGFR-TKI-resistant tumors, respectively [78]. IGF1R and SRC/FAK signaling have also been implicated in EMT in the context of TKI resistance [79, 80].
Drug-tolerant persister cells
Intrinsic versus acquired resistance has traditionally been perceived as dichotomous processes, but in actuality, the demarcation may be less distinct (Figure 2). In 2010, Sharma et al. proposed the notion of ‘drug-tolerant persister cells’ [81]. In NSCLC and other cancers, small subpopulations of cells (range 0.3–5%) can survive the initial exposure to drugs by adopting a reversible, quiescent drug-tolerant state in response to the lethal stress. The emergence of drug tolerance is dynamic. Persister cells can emerge de novo from single cell-derived, drug-sensitive populations [81]. These persister cells subsequently propagate until a permanent, genetically-based resistance mechanism is acquired [81]. Clinically, the majority of cancer patients with an objective tumor response to TKIs actually have incomplete (or partial) responses. While this residual disease burden may be secondary to multiple mechanisms (including pharmacokinetic issues that limit complete target inhibition), it could also be accounted for by the presence of persister cells, which ultimately lead to clinical relapse [82]. Therefore, understanding the mechanisms of persister cell survival may be critical in order to transform partial into complete responses and reduce the chance of resistance emerging.
Figure 2. An Evolutionary Model of Drug Resistance in Cancer Cells.
This figure depicts different models of the evolution of acquired resistance to targeted therapies in cancer cells. In the ‘pre-existing clones model,’ a few tumor cells harbor a pre-existing resistance mechanism prior to the exposure to a tyrosine kinase inhibitor (TKI). These tumor cells with pre-established resistance mechanisms can survive and rapidly proliferate under the selective pressure of a TKI, becoming the predominant cell population. The tumor population as a whole will continue to be heterogeneous. Alternatively, in the ‘persister evolution model,’ a small subpopulation of cells (represented in light blue) may adopt a relatively quiescent, reversible drug-tolerant persister state induced by the initial insult of the TKI exposure. This allows the survival of the persister cells and their subsequent propagation. The persister cells present a window of opportunity for the acquisition of resistance mechanisms, leading to the establishment of fully drug-resistant cells. In the ‘mixed model,’ there are pre-existing drug-resistant clones as well as drug-tolerant persister cells in the TKI-naïve cell population, which evolve under the selective pressure of the TKI until fully drug-resistant tumor clones are established. The ratio of various drug-resistant clones will differ in each tumor and will dynamically change over time in concert with changes in therapy.
Studies have demonstrated the importance of epigenetic programs in the emergence and survival of persister cells. Sharma et al. found that in some EGFR-mutant NSCLC cell lines, the development of EGFR-TKI tolerance requires activation of IGF-1R signaling. Increased IGF-1R phosphorylation was observed in persister cells, and treatment of the parental EGFR-mutant lung cancer cells with an EGFR inhibitor together with an IGF-1R inhibitor (AEW541) could suppress the emergence of persisters [81]. Interestingly, IGF-1R-mediated drug tolerance required the histone-demethylating activity of KDM5A, whose expression was upregulated in the persister cells [81]. KDM5A was previously shown to associate with histone deacetylases (HDACs), and its yeast ortholog Msc1 reduces histone H3K9/K14 acetylation. Indeed, the drug-tolerant EGFR-mutant lung cancer cells exhibited significantly reduced acetylation of histone H3K14, and treatment of these cells with an HDAC inhibitor (trichostatin A) resulted in cell death [81]. Altogether, these findings suggest a potential therapeutic role for HDAC inhibitors and/or IGF-1R inhibitors in suppressing the emergence of drug tolerance.
More recent work has identified additional mechanisms by which the persister cell state can be induced. For example, Blakely et al. reported that the treatment of EGFR-mutant lung cancer cells with an EGFR inhibitor could stimulate adaptive hyperactivation of NF-κB via ubiquitination events [83]. This NF-κB activation resulted in the upregulated expression of interleukin (IL)-6, activation of downstream JAK/STAT signaling pathway, and persister cell survival. In EGFR-mutant lung cancer cell line and murine xenograft models, pharmacologic inhibition of NF-κB together with EGFR inhibition led to enhanced responsiveness to EGFR-TKI treatment, suppression of residual disease, and suppression of the emergence of acquired resistance, providing the rationale for clinical testing of this approach [83]. Additional work has shown that therapy-induced factors secreted into the tumor microenvironment (either by tumor or stromal cells) may also regulate the survival of persister cells [84]. Therefore, tumor cells likely utilize a variety of different mechanisms to survive the initial TKI exposure.
Once persisters survive the initial exposure to TKI, they may then go on to acquire a range of full-blown resistance mechanisms such as T790M in EGFR (Figure 2) [85, 86]. Hata et al. studied the evolution of persister cells [86], and detected two distinct populations of EGFR-TKI-resistant tumor cells in EGFR-mutant NSCLC cell lines: ‘early-resistant’, pre-existing EGFR-T790M-mutant cells; and ‘late-resistant’, de novo EGFR-T790M-mutant cells. These late-resistant cells shared the molecular hallmarks of drug-tolerant persister cells [86], suggesting that persister cells likely served as the reservoir for subsequent T790M mutant resistant clones. Interestingly, compared to the pre-existing EGFR-T790M-mutant cells, the late-resistant cells exhibited diminished apoptotic response to treatment with a third-generation EGFR-TKI, WZ4002 [86]. The combination of WZ4002 with a BCL2-inhibitor, navitoclax, was able to induce enhanced apoptosis and tumor regression [86]. Based on these results, a clinical trial is underway examining the efficacy of osimertinib combined with navitoclax in patients with EGFR-mutant NSCLC after disease progression on first-line EGFR-TKIs.
Heterogeneity and polyclonal resistance
A binary view of resistance mechanisms fails in more ways than one. To this day, a resistant tumor is often simply labeled as either harboring one particular resistance mechanism or not. However, genetic heterogeneity is a defining feature of cancer and impacts TKI resistance, imbuing yet another layer of complexity.
Studies have demonstrated that in a given TKI-resistant tumor biopsy specimen, or in separate tumor sites within an individual patient, two or more discrete resistance mechanisms can be identified. Examples of polyclonal resistance in EGFR-mutant NSCLC include the co-occurrence of T790M with MET amplification [33–35, 59], EGFR amplification [33], HER2 amplification [34], or SCLC transformation [33, 34]; and SCLC transformation with MET amplification [34] or PIK3CA mutation [33]. In ALK-rearranged lung cancer, concurrent ALK G1202R mutation and c-KIT amplification [46], G1269A and ALK amplification [47], and MEK mutation and PIK3CA mutation [63] have been reported. Clinically, though, durable disease control is often achieved by targeting the predominant mechanism of resistance despite this heterogeneity. One example can be found in T790M-positive, EGFR-mutant NSCLC, where despite the known heterogeneity (also see below for further discussion), third-generation EGFR-TKIs appear to be typically uniformly effective [87, 88].
In the clinic, heterogeneity in TKI resistance is also manifested temporally as patients undergo changes in therapy. ‘Disease flares’ occur in up to 25% of lung cancer patients who develop TKI resistance, usually within days of stopping the drug [89, 90]. In parallel, ‘re-treatment responses’ are seen: tumors presumed to be TKI-resistant re-acquire sensitivity after a ‘drug holiday’ [89, 90]. These phenomena represent clinical corollaries of the longitudinal evolution of heterogeneous tumors in concert with changes in therapy. Tumors with acquired resistance contain a mixed population of cells within which a small subset remains TKI-sensitive and proliferates rapidly upon the removal of the TKI, explaining the phenomenon of disease flare. During a drug holiday, re-population of the cancer by TKI-sensitive cells leads to potentiation of responses upon TKI re-challenge [91].
More recently, Piotrowska et al. reported that T790-wild-type EGFR clones emerge after the T790M-positive EGFR-mutant lung cancers acquire resistance to rociletinib [88]. Single cell cloning of a T790M-‘positive’ pre-rociletinib specimen revealed both T790M-positive and -wild-type cells, suggesting that the apparent ‘loss’ of T790M at the time of resistance was actually due to the selection of these pre-existing T790-wild-type cells [88]. Similarly, Thress et al. found that some of the T790M-positive EGFR-mutant lung cancer cells treated with osimertinib could acquire resistance by ‘losing’ the T790M mutation [40]. This dynamic, polyclonal nature of resistance highlights the need for serial repeat biopsies to delineate key steps in the longitudinal evolution of resistance. As biopsies typically involve a single site and may not always be feasible, the development of noninvasive methods such as circulating tumor cell or cell free DNA analyses will be critical for evaluating the heterogeneity of resistance mechanisms.
Strategies targeting TKI resistance
Ongoing studies of TKI resistance mechanisms have shed light on the profound complexities of tumor biology and have guided the design of new therapeutic strategies to combat drug resistance. A detailed discussion of these strategies is beyond the scope of this Review; further information is available elsewhere [2, 92]. Here, we close with a brief overview of the use of next-generation TKIs and TKI-based combinatorial regimens.
Secondary resistance alterations within the kinase have spurred the development of newer-generation TKIs. In EGFR-mutant NSCLC, several third-generation TKIs have been developed to target T790M, including rociletinib, osimertinib, EGF816, ASP8273, and HM61713. Osimertinib is associated with a response rate of 61% and a median progression-free survival (PFS) of 9.6 months among patients with T790M-positive NSCLC who progressed on prior EGFR-TKIs [87]. The estimated duration of response is 6 months or longer in 88% of these patients [87]. Osimertinib is now FDA-approved for use in this patient population. Studies are underway to evaluate the efficacy of osimertinib in the front-line setting.
In ALK-rearranged NSCLC, the second-generation ALK inhibitors ceritinib and alectinib are currently FDA-approved for patients previously treated with crizotinib. In the ASCEND-1 trial, ceritinib showed a response rate of 56%, median duration of response of 8.3 months, and median PFS of 6.9 months in ALK inhibitor-pretreated, advanced ALK-rearranged NSCLC [93]. In ALK inhibitor-naïve patients, the response rate was 72%, with a median duration of response of 17 months and median PFS of 18.4 months [93]. In two phase II studies, alectinib demonstrated a ~50% response rate, median duration of response of 11.2–13.5 months, and estimated median PFS of 8.1–8.9 months in patients with advanced ALK-rearranged NSCLC who had progressed after crizotinib [94, 95]. Clinical trials are ongoing to evaluate the efficacy of both ceritinib and alectinib in the first-line setting. Under active investigation are also a number of other next-generation ALK-TKIs, including lorlatinib, brigatinib, ensartinib, and entrectinib. Lorlatinib in particular has demonstrated clinical activity in patients who have failed multiple ALK-TKIs. Notably, each ALK-TKI is structurally distinct with varying degrees of activity against the different ALK resistance mutations in vitro and in the clinic [96]. These unique activity profiles may form the basis for selecting and sequencing next-generation ALK-TKIs.
The importance of bypass tracks and downstream effectors in mediating TKI resistance suggests that combinatorial approaches may be required. Indeed, numerous clinical trials are underway to evaluate the efficacy and tolerability of different drug combinations to overcome resistance. These include the inhibition of EGFR together with MET, HGF, mTOR, PI3K, MEK, BCL2, AXL/FLT3, SRC, IGF, JAK, or HSP90. In ALK-TKI resistant disease, co-inhibition of ALK with mTOR, CDK4/6, VEGF, MEK, or HSP90 is being studied. Several early-phase studies are also assessing the feasibility of coupling TKIs with immunotherapies, although this is based on limited preclinical data. The number of possible combination regimens is vast. The bottleneck in many cases will be the side effects that emerge when drugs are given in combination [2, 92]. Use of alternative dosing schedules such as pulsed or intermittent dosing may help in certain cases. In parallel, greater rigor is warranted in the design of early studies to ensure the integration of patients’ genotypes and resistance mechanisms in the rational design of combination strategies.
Concluding remarks
Fourteen years have now passed since I.B. Weinstein’s initial analogy of oncogene addictions to Achilles’ heel [1]. Each year, novel tumor vulnerabilities are being discovered, and new therapeutic strategies are moving from bench to bedside. Amidst these advances, understanding TKI resistance mechanisms has become an urgent imperative. Many examples provided in this Review focus on our collective experience with EGFR- or ALK-positive NSCLC. However, TKI resistance is pervasive, and themes presented herein are likely universal. On the whole, the evolution of drug resistance is complex and dynamic—in many ways reminiscent of the multi-headed Hydra in Greek mythology [92].
Many questions on TKI resistance remain unanswered (see Outstanding Questions). Over the next decade and beyond, our ability to combat resistance will require collaborative efforts among bench, translational, and clinical researchers. More studies on the evolutionary dynamics of resistance are critical. Efforts will need to focus on the development of noninvasive diagnostic tools to capture the full breadth of the evolving resistance mechanisms in patients. Thus far, our therapeutic strategies have focused on overcoming TKI resistance that is already established. It is likely that greater impact will be derived from novel therapeutic strategies that prevent the emergence of resistance upfront, target the drug-tolerant persister cells, and/or integrate targeted therapies with other treatment modalities such as radiation or immunotherapy. These approaches hold the promise of truly transforming cancer treatment and prolonging patients’ lives.
Outstanding Questions.
What molecular mechanisms underlie primary or intrinsic resistance to TKI therapy? On the opposite end of the spectrum, a small percentage of patients demonstrate prolonged responses to TKIs lasting many years. What are the molecular underpinnings of exceptional responses?
In some cases of resistance, cancers develop recurrent on-target gene alterations and remain oncogene-addicted. In other cases of resistance, cancers activate alternative signaling pathways and quickly lose their oncogene addiction. What is the biological basis for cancers evolving on-target versus off-target resistance mechanisms?
How do histologic (i.e., SCLC transformation) and phenotypic (i.e., EMT) changes drive TKI resistance, and will these changes predominate as we develop increasingly effective targeted therapy approaches?
What are the key pathways enabling survival of drug-tolerant persister cells? What is the balance between rare pre-existing resistant cells and drug-tolerant persister cells, and how can therapeutic strategies be developed to target both populations?
Tumor heterogeneity has been widely described in resistant cancers, yet in many cases, one resistance mechanism predominates (e.g., EGFR-T790M). What are the best methods to assess tumor heterogeneity and to evaluate its clinical relevance?
What are the best upfront combination regimens to use to prevent acquired resistance?
Trends.
Tyrosine kinase inhibitors (TKIs) represent a highly efficacious class of cancer therapeutics. However, drug resistance invariably limits the clinical activity of most TKIs.
Universal, well-established mechanisms of TKI resistance include secondary resistance mutations within the target kinase, activation of bypass signaling tracks, dysregulation of downstream effector proteins, and phenotypic transformation.
TKI resistance evolves dynamically under the selective pressure exerted by each targeted therapy. In some (but not all) cases, the evolution of TKI resistance can lead to significant tumor heterogeneity within a given cancer.
Drug-tolerant or ‘persister’ cells can survive initial TKI therapy and serve as a reservoir for drug-resistant clones.
Rational therapeutic strategies targeting mechanisms of TKI resistance are critical to improving clinical outcomes.
Acknowledgments
We apologize to the numerous colleagues whose important contributions could not be cited in this Review due to space constraints. A.T.S. is supported by grants from the National Cancer Institute (5R01CA164273) and the National Foundation for Cancer Research.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Weinstein IB. Addiction to oncogenes –the Achilles heal of cancer. Science. 2002;297:63–64. doi: 10.1126/science.1073096. [DOI] [PubMed] [Google Scholar]
- 2.Camidge DR, et al. Acquired resistance to TKIs in solid tumours: learning from lung cancer. Nat. Rev. Clin. Oncol. 2014;11:473–481. doi: 10.1038/nrclinonc.2014.104. [DOI] [PubMed] [Google Scholar]
- 3.Maemondo M, et al. Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. N. Engl. J. Med. 2010;362:2380–2388. doi: 10.1056/NEJMoa0909530. [DOI] [PubMed] [Google Scholar]
- 4.Rosell R, et al. Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): a multicentre, open-label, randomised phase 3 trial. Lancet Oncol. 2012;13:239–246. doi: 10.1016/S1470-2045(11)70393-X. [DOI] [PubMed] [Google Scholar]
- 5.Zhou C, et al. Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): a multicentre, open-label, randomised, phase 3 study. Lancet Oncol. 2011;12:735–742. doi: 10.1016/S1470-2045(11)70184-X. [DOI] [PubMed] [Google Scholar]
- 6.Mitsudomi T, et al. Gefitinib versus cisplatin plus docetaxel in patients with non-small-cell lung cancer harbouring mutations of the epidermal growth factor receptor (WJTOG3405): an open label, randomised phase 3 trial. Lancet Oncol. 2010;11:121–128. doi: 10.1016/S1470-2045(09)70364-X. [DOI] [PubMed] [Google Scholar]
- 7.Sequist LV, et al. Phase III study of afatinib or cisplatin plus pemetrexed in patients with metastatic lung adenocarcinoma with EGFR mutations. J. Clin. Oncol. 2013;31:3327–3334. doi: 10.1200/JCO.2012.44.2806. [DOI] [PubMed] [Google Scholar]
- 8.Wu YL, et al. Afatinib versus cisplatin plus gemcitabine for first-line treatment of Asian patients with advanced non-small-cell lung cancer harbouring EGFR mutations (LUX-Lung 6): an open-label, randomised phase 3 trial. Lancet Oncol. 2014;15:213–222. doi: 10.1016/S1470-2045(13)70604-1. [DOI] [PubMed] [Google Scholar]
- 9.Solomon BJ, et al. First-line crizotinib versus chemotherapy in ALK-positive lung cancer. N. Engl. J. Med. 2014;371:2167–2177. doi: 10.1056/NEJMoa1408440. [DOI] [PubMed] [Google Scholar]
- 10.Kris MG, et al. Using multiplexed assays of oncogenic drivers in lung cancers to select targeted drugs. JAMA. 2014;311:1998–2006. doi: 10.1001/jama.2014.3741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Barlesi F, et al. Routine molecular profiling of patients with advanced non-small-cell lung cancer: results of a 1-year nationwide programme of the French Cooperative Thoracic Intergroup (IFCT) Lancet. 2016 doi: 10.1016/S0140-6736(16)00004-0. [DOI] [PubMed] [Google Scholar]
- 12.Swanton C, Govindan R. Clinical implications of genomic discoveries in lung cancer. N. Engl. J. Med. 2016;374:1864–1873. doi: 10.1056/NEJMra1504688. [DOI] [PubMed] [Google Scholar]
- 13.Takeuchi K, et al. RET, ROS1, and ALK fusions in lung cancer. Nat. Med. 2012;18:378–381. doi: 10.1038/nm.2658. [DOI] [PubMed] [Google Scholar]
- 14.Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature. 2014;511:543–550. doi: 10.1038/nature13385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Bergethon K, et al. ROS1 rearrangements define a unique molecular class of lung cancers. J. Clin. Oncol. 2012;30:863–870. doi: 10.1200/JCO.2011.35.6345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Shaw AT, et al. Crizotinib in ROS1-rearranged non-small-cell lung cancer. N. Engl. J. Med. 2014;371:1963–1971. doi: 10.1056/NEJMoa1406766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Awad MM, et al. MET exon 14 mutations in non-small-cell lung cancer are associated with advanced age and stage-dependent MET genomic amplification and c-Met overexpression. J. Clin. Oncol. 2016;34:721–730. doi: 10.1200/JCO.2015.63.4600. [DOI] [PubMed] [Google Scholar]
- 18.Oxnard GR, et al. Natural history and molecular characteristics of lung cancers harboring EGFR exon 20 insertions. J. Thorac. Oncol. 2013;8:179–184. doi: 10.1097/JTO.0b013e3182779d18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Arcila ME, et al. EGFR exon 20 insertion mutations in lung adenocarcinomas: prevalence, molecular heterogeneity, and clinicopathologic characteristics. Mol. Cancer Ther. 2013;12:220–229. doi: 10.1158/1535-7163.MCT-12-0620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Yang JC, et al. Clinical activity of afatinib in patients with advanced non-small-cell lung cancer harbouring uncommon EGFR mutations: a combined post-hoc analysis of LUX-Lung 2, LUX-Lung 3, and LUX-Lung 6. Lancet Oncol. 2015;16:830–838. doi: 10.1016/S1470-2045(15)00026-1. [DOI] [PubMed] [Google Scholar]
- 21.Yasuda H, et al. Structural, biochemical, and clinical characterization of epidermal growth factor receptor (EGFR) exon 20 insertion mutations in lung cancer. Sci. Transl. Med. 2013 doi: 10.1126/scitranslmed.3007205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Yu HA, et al. Poor response to erlotinib in patients with tumors containing baseline EGFR T790M mutations found by routine clinical molecular testing. Ann. Oncol. 2014;25:423–428. doi: 10.1093/annonc/mdt573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Su KY, et al. Pretreatment epidermal growth factor receptor (EGFR) T790M mutation predicts shorter EGFR tyrosine kinase inhibitor response duration in patients with non-small-cell lung cancer. J. Clin. Oncol. 2012;30:433–440. doi: 10.1200/JCO.2011.38.3224. [DOI] [PubMed] [Google Scholar]
- 24.Costa C, et al. The impact of EGFR T790M mutations and BIM mRNA expression on outcome in patients with EGFR-mutant NSCLC treated with erlotinib or chemotherapy in the randomized phase III EURTAC trial. Clin. Cancer Res. 2014;20:2001–2010. doi: 10.1158/1078-0432.CCR-13-2233. [DOI] [PubMed] [Google Scholar]
- 25.Turke AB, et al. Preexistence and clonal selection of MET amplification in EGFR mutant NSCLC. Cancer Cell. 2010;17:77–88. doi: 10.1016/j.ccr.2009.11.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Karachaliou N, et al. BIM and mTOR expression levels predict outcome to erlotinib in EGFR-mutant non-small-cell lung cancer. Sci. Rep. 2015 doi: 10.1038/srep17499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Nie W, et al. The BIM deletion polymorphism is a prognostic biomarker of EGFR-TKIs response in NSCLC: A systematic review and meta-analysis. Oncotarget. 2015;6:25696–25700. doi: 10.18632/oncotarget.4678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Bivona TG, et al. FAS and NF-κB signalling modulate dependence of lung cancers on mutant EGFR. Nature. 2011;471:523–526. doi: 10.1038/nature09870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Byers LA, et al. An epithelial-mesenchymal transition gene signature predicts resistance to EGFR and PI3K inhibitors and identifies Axl as a therapeutic target for overcoming EGFR inhibitor resistance. Clin. Cancer Res. 2013;19:279–290. doi: 10.1158/1078-0432.CCR-12-1558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Park KS, et al. CRIPTO1 expression in EGFR-mutant NSCLC elicits intrinsic EGFR-inhibitor resistance. J. Clin. Invest. 2014;124:3003–3015. doi: 10.1172/JCI73048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sholl LM, et al. Combined use of ALK immunohistochemistry and FISH for optimal detection of ALK-rearranged lung adenocarcinomas. J. Thorac. Oncol. 2013;8:322–328. doi: 10.1097/JTO.0b013e31827db604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Jabbour EJ, et al. Resistance to tyrosine kinase inhibition therapy for chronic myelogenous leukemia: a clinical perspective and emerging treatment options. Clin. Lymphoma Myeloma Leuk. 2013;13:515–529. doi: 10.1016/j.clml.2013.03.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Sequist LV, et al. Genotypic and histological evolution of lung cancers acquiring resistance to EGFR inhibitors. Sci. Transl. Med. 2011 doi: 10.1126/scitranslmed.3002003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Yu HA, et al. Analysis of tumor specimens at the time of acquired resistance to EGFR-TKI therapy in 155 patients with EGFR-mutant lung cancers. Clin. Cancer Res. 2013;19:2240–2247. doi: 10.1158/1078-0432.CCR-12-2246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Arcila ME, et al. Rebiopsy of lung cancer patients with acquired resistance to EGFR inhibitors and enhanced detection of the T790M mutation using a locked nucleic acid-based assay. Clin. Cancer Res. 2011;17:1169–1180. doi: 10.1158/1078-0432.CCR-10-2277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yun CH, et al. The T790M mutation in EGFR kinase causes drug resistance by increasing the affinity for ATP. Proc. Natl. Acad. Sci. U.S.A. 2008;105:2070–2075. doi: 10.1073/pnas.0709662105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Bean J, et al. Acquired resistance to epidermal growth factor receptor kinase inhibitors associated with a novel T854A mutation in a patient with EGFR-mutant lung adenocarcinoma. Clin. Cancer Res. 2008;14:7519–7525. doi: 10.1158/1078-0432.CCR-08-0151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Balak MN, et al. Novel D761Y and common secondary T790M mutations in epidermal growth factor receptor-mutant lung adenocarcinomas with acquired resistance to kinase inhibitors. Clin. Cancer Res. 2006;12:6494–6501. doi: 10.1158/1078-0432.CCR-06-1570. [DOI] [PubMed] [Google Scholar]
- 39.Costa DB, et al. Differential responses to erlotinib in epidermal growth factor receptor (EGFR)-mutated lung cancers with acquired resistance to gefitinib carrying the L747S or T790M secondary mutations. J. Clin. Oncol. 2008;26:1182–1184. doi: 10.1200/JCO.2007.14.9039. [DOI] [PubMed] [Google Scholar]
- 40.Thress KS, et al. Acquired EGFR C797S mutation mediates resistance to AZD9291 in non-small cell lung cancer harboring EGFR T790M. Nat. Med. 2015;21:560–562. doi: 10.1038/nm.3854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ercan D, et al. EGFR mutations and resistance to irreversible pyrimidine-based EGFR inhibitors. Clin. Cancer Res. 2015;21:3913–3923. doi: 10.1158/1078-0432.CCR-14-2789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Yu HA, et al. Acquired resistance of EGFR-mutant lung cancer to a T790M-specific EGFR inhibitor: emergence of a third mutation (C797S) in the EGFR tyrosine kinase domain. JAMA Oncol. 2015;1:982–984. doi: 10.1001/jamaoncol.2015.1066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Niederst MJ, et al. The allelic context of the C797S mutation acquired upon treatment with third-generation EGFR inhibitors impacts sensitivity to subsequent treatment strategies. Clin. Cancer Res. 2015;21:3924–3933. doi: 10.1158/1078-0432.CCR-15-0560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Choi YL, et al. EML4-ALK mutations in lung cancer that confer resistance to ALK inhibitors. N. Engl. J. Med. 2010;363:1734–1739. doi: 10.1056/NEJMoa1007478. [DOI] [PubMed] [Google Scholar]
- 45.Sasaki T, et al. A novel ALK secondary mutation and EGFR signaling cause resistance to ALK kinase inhibitors. Cancer Res. 2011;71:6051–6060. doi: 10.1158/0008-5472.CAN-11-1340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Katayama R, et al. Mechanisms of acquired crizotinib resistance in ALK-rearranged lung cancers. Sci. Transl. Med. 2012 doi: 10.1126/scitranslmed.3003316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Doebele RC, et al. Mechanisms of resistance to crizotinib in patients with ALK gene rearranged non-small cell lung cancer. Clin. Cancer Res. 2012;18:1472–1482. doi: 10.1158/1078-0432.CCR-11-2906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Ignatius Ou SH, et al. Next-generation sequencing reveals a novel NSCLC ALK F1174V mutation and confirms ALK G1202R mutation confers high-level resistance to alectinib (CH5424802/RO5424802) in ALK-rearranged NSCLC patients who progressed on crizotinib. J. Thorac. Oncol. 2014;9:549–553. doi: 10.1097/JTO.0000000000000094. [DOI] [PubMed] [Google Scholar]
- 49.Fontana D, et al. Activity of second-generation ALK inhibitors against crizotinib-resistant mutants in an NPM-ALK model compared to EML4-ALK. Cancer Med. 2015;4:953–965. doi: 10.1002/cam4.413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Zou HY, et al. PF-06463922, an ALK/ROS1 inhibitor, overcomes resistance to first and second generation ALK inhibitors in preclinical models. Cancer Cell. 2015;28:70–81. doi: 10.1016/j.ccell.2015.05.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Shaw AT, et al. Resensitization of crizotinib by the lorlatinib ALK resistance mutation L1198F. N. Engl. J. Med. 2015;374:54–61. doi: 10.1056/NEJMoa1508887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Awad MM, et al. Acquired resistance to crizotinib from a mutation in CD74-ROS1. N. Engl. J. Med. 2013;368:2395–2401. doi: 10.1056/NEJMoa1215530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Drilon A, et al. A novel crizotinib-resistant solvent-front mutation responsive to cabozantinib therapy in a patient with ROS1-rearranged lung cancer. Clin. Cancer Res. 2015 doi: 10.1158/1078-0432.CCR-15-2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Song A, et al. Molecular changes associated with acquired resistance to crizotinib in ROS1-rearranged non-small cell lung cancer. Clin. Cancer Res. 2015;21:2379–2387. doi: 10.1158/1078-0432.CCR-14-1350. [DOI] [PubMed] [Google Scholar]
- 55.Russo M, et al. Acquired resistance to the TRK inhibitor entrectinib in colorectal cancer. Cancer Discov. 2016;6:36–44. doi: 10.1158/2159-8290.CD-15-0940. [DOI] [PubMed] [Google Scholar]
- 56.Tabara K, et al. Loss of activating EGFR mutant gene contributes to acquired resistance to EGFR tyrosine kinase inhibitors in lung cancer cells. PLoS One. 2012 doi: 10.1371/journal.pone.0041017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Niederst MJ, et al. RB loss in resistant EGFR mutant lung adenocarcinomas that transform to small-cell lung cancer. Nat. Commun. 2015 doi: 10.1038/ncomms7377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Katayama R, et al. Therapeutic strategies to overcome crizotinib resistance in non-small cell lung cancers harboring the fusion oncogene EML4-ALK. Proc. Natl, Acad. Sci, U. S. A. 2011;108:7535–7540. doi: 10.1073/pnas.1019559108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Engelman JA, et al. MET amplification leads to gefitinib resistance in lung cancer by activating ERBB3 signaling. Science. 2007;316:1039–1043. doi: 10.1126/science.1141478. [DOI] [PubMed] [Google Scholar]
- 60.Yano S, et al. Hepatocyte growth factor induces gefitinib resistance of lung adenocarcinoma with epidermal growth factor receptor-activating mutations. Cancer Res. 2008;68:9479–9487. doi: 10.1158/0008-5472.CAN-08-1643. [DOI] [PubMed] [Google Scholar]
- 61.Gouji T, et al. Crizotinib can overcome acquired resistance to CH5424802: is amplification of the MET gene a key factor? J. Thorac. Oncol. 2014 doi: 10.1097/JTO.0000000000000113. [DOI] [PubMed] [Google Scholar]
- 62.Lovly CM, et al. Rationale for co-targeting IGF-1R and ALK in ALK fusion-positive lung cancer. Nat. Med. 2014;20:1027–1034. doi: 10.1038/nm.3667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Crystal AS, et al. Patient-derived models of acquired resistance can identify effective drug combinations for cancer. Science. 2014;346:1480–1486. doi: 10.1126/science.1254721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Ohashi K, et al. Lung cancers with acquired resistance to EGFR inhibitors occasionally harbor BRAF gene mutations but lack mutations in KRAS, NRAS, or MEK1. Proc. Natl. Acad. Sci. U.S.A. 2012;109:E2127–E2133. doi: 10.1073/pnas.1203530109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Cheung HW, et al. Amplification of CRKL induces transformation and epidermal growth factor receptor inhibitor resistance in human non-small cell lung cancers. Cancer Discov. 2011;1:608–625. doi: 10.1158/2159-8290.CD-11-0046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.De Bruin EC, et al. Reduced NF1 expression confers resistance to EGFR inhibition in lung cancer. Cancer Discov. 2014;4:606–619. doi: 10.1158/2159-8290.CD-13-0741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Davies KD, et al. Resistance to ROS1 inhibition mediated by EGFR pathway activation in non-small cell lung cancer. PLoS One. 2013 doi: 10.1371/journal.pone.0082236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Queirolo P, et al. Combined BRAF and MEK inhibition for the treatment of BRAF-mutated metastatic melanoma. Cancer Treat. Rev. 2015;41:519–526. doi: 10.1016/j.ctrv.2015.04.010. [DOI] [PubMed] [Google Scholar]
- 69.Lin L, et al. Mapping the molecular determinants of BRAF oncogene dependence in human lung cancer. Proc. Natl. Acad. Sci. U.S.A. 2014;111:E748–E757. doi: 10.1073/pnas.1320956111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Trickler EM, et al. Combined EGFR/MEK inhibition prevents the emergence of resistance in EGFR-mutant lung cancer. Cancer Discov. 2015;5:960–971. doi: 10.1158/2159-8290.CD-15-0063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Hrustanovic G, et al. RAS-MAPK dependence underlies a rational polytherapy strategy in EML4-ALK-positive lung cancer. Nat. Med. 2015;21:1038–1047. doi: 10.1038/nm.3930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Miyamoto S, et al. Transformation to small-cell lung cancer as a mechanism of acquired resistance to crizotinib and alectinib. Jap. J. Clin. Oncol. 2015;46:170–173. doi: 10.1093/jjco/hyv173. [DOI] [PubMed] [Google Scholar]
- 73.Fujita S, et al. Transformation to SCLC after treatment with the ALK inhibitor alectinib. J. Thorac. Oncol. 2016 doi: 10.1016/j.jtho.2015.12.105. [DOI] [PubMed] [Google Scholar]
- 74.Takegawa N, et al. Transformation of ALK rearrangement-positive adenocarcinoma to small cell lung cancer in association with acquired resistance to alectinib. Ann. Oncol. 2016 doi: 10.1093/annonc/mdw032. [DOI] [PubMed] [Google Scholar]
- 75.Niederst MJ, et al. RB loss in resistant EGFR mutant lung adenocarcinomas that transform to small-cell lung cancer. Nat. Commun. 2015 doi: 10.1038/ncomms7377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Singh A, Settleman J. EMT, cancer stem cells and drug resistance: an emerging axis of evil in the war on cancer. Oncogene. 2010;29:4741–4751. doi: 10.1038/onc.2010.215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Kim HR, et al. Epithelial-mesenchymal transition leads to crizotinib resistance in H2228 lung cancer cells with EML4-ALK translocation. Mol. Oncol. 2013;7:1093–1102. doi: 10.1016/j.molonc.2013.08.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Zhang Z, et al. Activation of the AXL kinase causes resistance to EGFR-targeted therapy in lung cancer. Nat. Genet. 2012;44:852–860. doi: 10.1038/ng.2330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Zhou J, et al. Implication of epithelial-mesenchymal transition in IGF1R-induced resistance to EGFR-TKIs in advanced non-small cell lung cancer. Oncotarget. 2015 doi: 10.18632/oncotarget.6293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Wilson C, et al. Overcoming EMT-associated resistance to anti-cancer drugs via Src/FAK pathway inhibition. Oncotarget. 2014;5:7328–7341. doi: 10.18632/oncotarget.2397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Sharma SV, et al. A chromatin-mediated reversible drug-tolerant state in cancer cell subpopulations. Cell. 2010;141:69–80. doi: 10.1016/j.cell.2010.02.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Bivona TG, Doebele RC. A framework for understanding and targeting residual disease in oncogene-driven solid cancers. Nat. Med. 2016;22:472–478. doi: 10.1038/nm.4091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Blakely CM, et al. NF-κB activating complex engaged in response to EGFR oncogene inhibition drives tumor cell survival and residual disease in lung cancer. Cell Rep. 2015;11:98–110. doi: 10.1016/j.celrep.2015.03.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Obenauf AC, et al. Therapy-induced tumour secretomes promote resistance and tumour progression. Nature. 2015;520:368–372. doi: 10.1038/nature14336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Ramirez M, et al. Diverse drug-resistance mechanisms can emerge from drug-tolerant cancer persister cells. Nat. Commun. 2016 doi: 10.1038/ncomms10690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Hata AN, et al. Tumor cells can follow distinct evolutionary paths to become resistant to epidermal growth factor receptor inhibition. Nat. Med. 2016 doi: 10.1038/nm.4040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Jänne PA, et al. AZD9291 in EGFR inhibitor-resistant non-small-cell lung cancer. N. Engl. J. Med. 2015;372:1689–1699. doi: 10.1056/NEJMoa1411817. [DOI] [PubMed] [Google Scholar]
- 88.Piotrowska Z, et al. Heterogeneity underlies the emergence of EGFRT790 wild-type clones following treatment of T790M-positive cancers with a third-generation EGFR inhibitor. Cancer Discov. 2015;5:713–722. doi: 10.1158/2159-8290.CD-15-0399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Chaft JE, et al. Disease flare after tyrosine kinase inhibitor discontinuation in patients with EGFR-mutant lung cancer and acquired resistance to erlotinib or gefitinib: implications for clinical trial design. Clin. Cancer Res. 2011;17:6298–6303. doi: 10.1158/1078-0432.CCR-11-1468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Browning ET, et al. Response to crizotinib rechallenge after initial progression and intervening chemotherapy in ALK lung cancer. J. Thorac. Oncol. 2013 doi: 10.1097/JTO.0b013e31827a892c. [DOI] [PubMed] [Google Scholar]
- 91.Chmielecki J, et al. Optimization of dosing for EGFR-mutant non-small cell lung cancer with evolutionary cancer modeling. Sci. Transl. Med. 2011 doi: 10.1126/scitranslmed.3002356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Chong CR, Jänne PA. The quest to overcome resistance to EGFR-targeted therapies in cancer. Nat. Med. 2013;19:1389–1400. doi: 10.1038/nm.3388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Kim DW, et al. Activity and safety of ceritinib in patients with ALK-rearranged non-small-cell lung cancer (ASCEND-1): updated results from the multicentre, open-label, phase 1 trial. Lancet Oncol. 2016 doi: 10.1016/S1470-2045(15)00614-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Shaw AT, et al. Alectinib in ALK-positive, crizotinib-resistant, non-small-cell lung cancer: a single-group, multicentre, phase 2 trial. Lancet Oncol. 2016;17:234–242. doi: 10.1016/S1470-2045(15)00488-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Ou SH, et al. Alectinib in crizotinib-refractory ALK-rearranged non-small-cell lung cancer: a phase II global study. J. Clin. Oncol. 2016;34:661–668. doi: 10.1200/jco.2015.63.9443. [DOI] [PubMed] [Google Scholar]
- 96.Awad MM, Shaw AT. ALK inhibitors in non-small cell lung cancer: crizotinib and beyond. Clin. Adv. Hematol. Oncol. 2014;12:429–439. [PMC free article] [PubMed] [Google Scholar]