Abstract
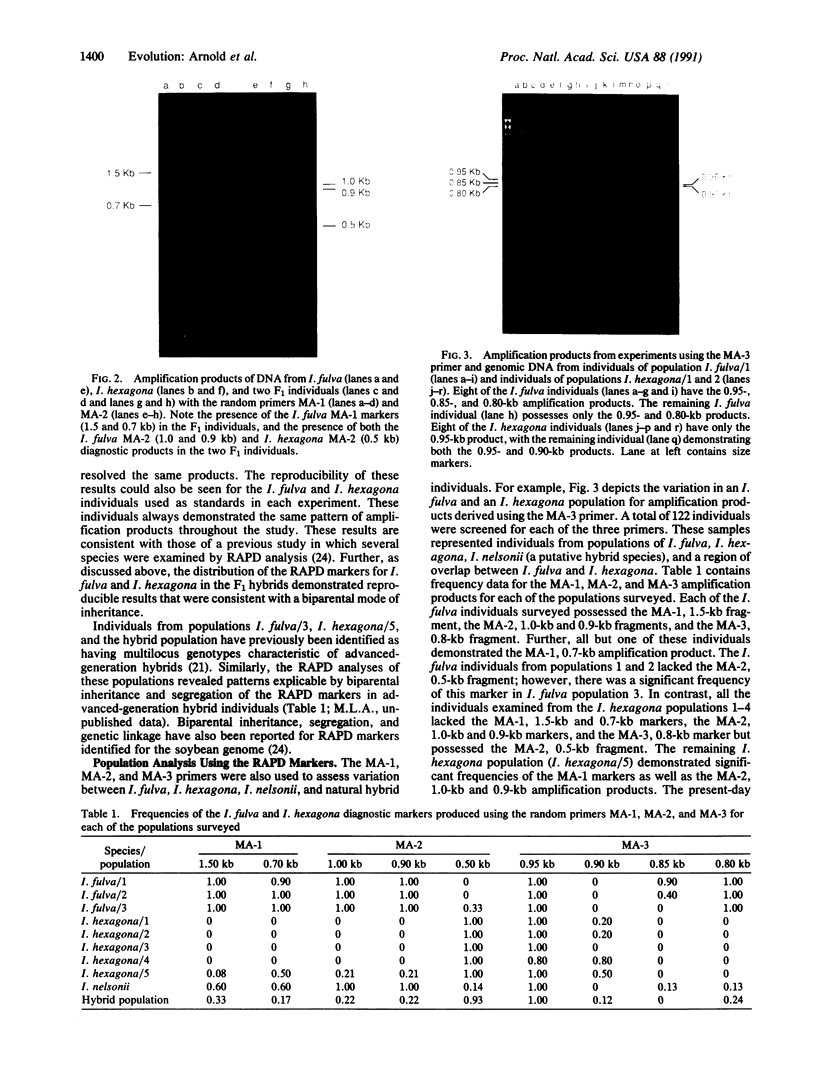
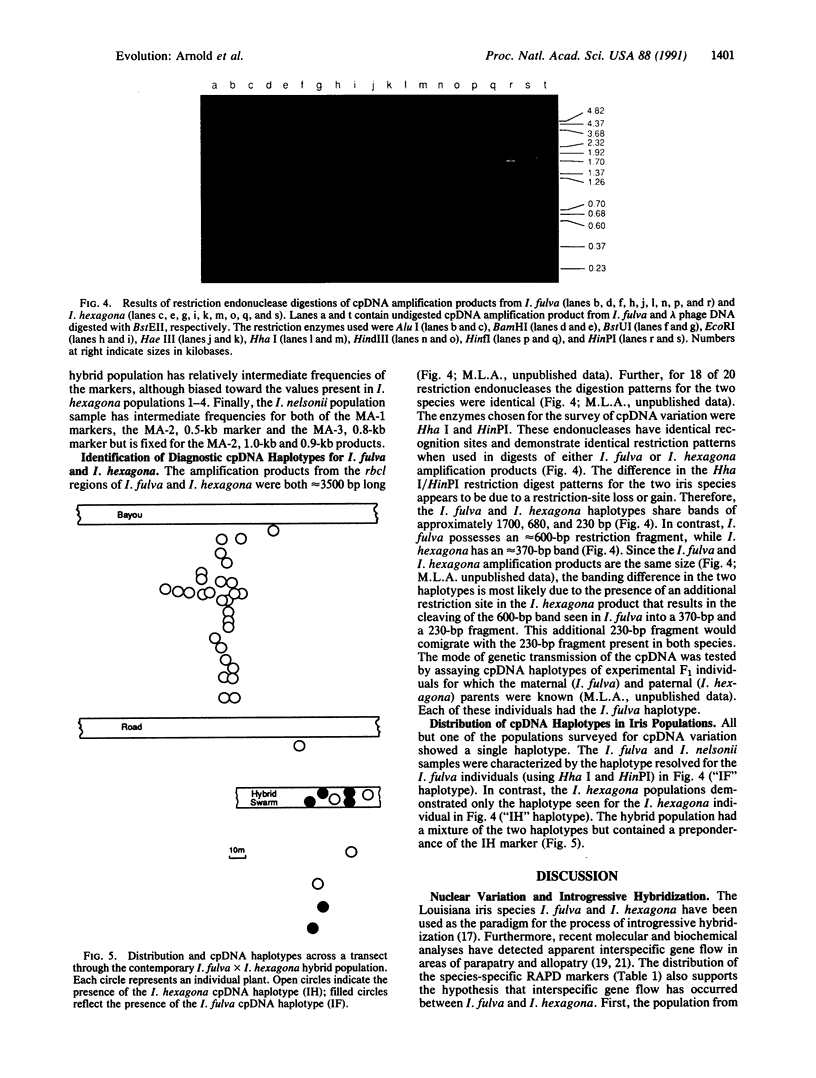
Populations of the "Louisiana iris" species Iris fulva, I. hexagona, and I. nelsonii were examined genetically to test for interspecific gene flow between I. fulva and I. hexagona, for pollen- versus seed-mediated introgression between these species, and for the presumed hybrid origin of I. nelsonii. Genetic markers were identified by using both a polymerase chain reaction-like method that allows the identification of random, nuclear markers and standard polymerase chain reaction experiments involving specific chloroplast DNA (cpDNA) oligonucleotides. Restriction endonuclease digestions of the cpDNA amplification products resolved diagnostic restriction site differences for I. fulva and I. hexagona. The distribution of the species-specific nuclear markers supports a hypothesis of bidirectional introgression between I. fulva and I. hexagona. Thus, individuals analyzed from a contemporary hybrid population demonstrate multilocus genotypes that are indicative of advanced-generation hybrid individuals. Furthermore, several markers from the alternate species were present in low frequency in one allopatric population each of I. fulva and I. hexagona. Data from the nuclear analysis also support the hypothesized hybrid origin of I. nelsonii from the interaction of I. fulva and I. hexagona. Finally, cpDNA data support the hypothesis that the localized and the dispersed introgression are largely due to pollen transfer. In addition to the biological implications, this study demonstrates the power of the polymerase chain reaction methodology for the rapid identification of random and specific genetic markers for testing evolutionary genetic hypotheses.
Full text
PDF


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Hill W. G. DNA fingerprints applied to animal and bird populations. Nature. 1987 May 14;327(6118):98–99. doi: 10.1038/327098a0. [DOI] [PubMed] [Google Scholar]
- Hiratsuka J., Shimada H., Whittier R., Ishibashi T., Sakamoto M., Mori M., Kondo C., Honji Y., Sun C. R., Meng B. Y. The complete sequence of the rice (Oryza sativa) chloroplast genome: intermolecular recombination between distinct tRNA genes accounts for a major plastid DNA inversion during the evolution of the cereals. Mol Gen Genet. 1989 Jun;217(2-3):185–194. doi: 10.1007/BF02464880. [DOI] [PubMed] [Google Scholar]
- Sears B. B. Elimination of plastids during spermatogenesis and fertilization in the plant kingdom. Plasmid. 1980 Nov;4(3):233–255. doi: 10.1016/0147-619x(80)90063-3. [DOI] [PubMed] [Google Scholar]
- Williams J. G., Kubelik A. R., Livak K. J., Rafalski J. A., Tingey S. V. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res. 1990 Nov 25;18(22):6531–6535. doi: 10.1093/nar/18.22.6531. [DOI] [PMC free article] [PubMed] [Google Scholar]