Abstract
Background
Sphingosine-1-phosphate and lysophosphatidic acid (LPA) are ligands for two related families of G protein-coupled receptors, the S1P and LPA receptors, respectively. The lysophospholipid ligands of these receptors are structurally similar, however recognition of these lipids by these receptors is highly selective. A single residue present within the third transmembrane domain (TM) of S1P receptors is thought to determine ligand selectivity; replacement of the naturally occurring glutamic acid with glutamine (present at this position in the LPA receptors) has previously been shown to be sufficient to change the specificity of S1P1 from S1P to 18:1 LPA.
Results
We tested whether mutation of this "ligand selectivity" residue to glutamine could confer LPA-responsiveness to the related S1P receptor, S1P4. This mutation severely affected the response of S1P4 to S1P in a [35S]GTPγS binding assay, and imparted sensitivity to LPA species in the order 14:0 LPA > 16:0 LPA > 18:1 LPA. These results indicate a length restriction for activation of this receptor and demonstrate the utility of using LPA-responsive S1P receptor mutants to probe binding pocket length using readily available LPA species. Computational modelling of the interactions between these ligands and both wild type and mutant S1P4 receptors showed excellent agreement with experimental data, therefore confirming the fundamental role of this residue in ligand recognition by S1P receptors.
Conclusions
Glutamic acid in the third transmembrane domain of the S1P receptors is a general selectivity switch regulating response to S1P over the closely related phospholipids, LPA. Mutation of this residue to glutamine confers LPA responsiveness with preference for short-chain species. The preference for short-chain LPA species indicates a length restriction different from the closely related S1P1 receptor.
Background
Sphingosine-1-phosphate (S1P) and lysophosphatidic acid (LPA) are phospholipid growth factors which are present in normal serum and plasma. These lipids elicit diverse responses from a wide range of cell types, including enhanced cell survival, cell proliferation, induction of cytoskeletal changes and chemotaxis (reviewed in [1-4]. Some of these responses reflect activation of G protein-coupled receptors of the endothelial differentiation gene (Edg) family. The Edg receptor family is classified into two clusters based on ligand selectivity: S1P1/2/3/4/5 (formerly Edg1/5/3/6/8) specifically respond to S1P whilst LPA1/2/3 (formerly Edg2/4/7) respond to LPA [5]. Members of the S1P receptor family display higher sequence similarity to each other (approximately 40% identity) than to members of the LPA receptor family (approximately 30% identity). These homologies, coupled with observed differences in the structure of S1P and LPA receptor genes, suggest that these receptor families evolved from distinct ancestral genes. The S1P receptors contain a conserved glutamic acid residue present within the third TM that corresponds to glutamine in the LPA receptors. Interaction between distinct functional groups present on S1P and LPA with this residue was shown for the S1P1 and LPA1 receptors using computational modelling techniques [6,7] and was demonstrated as the basis for the ligand preference displayed by the receptors. Experimental characterisation confirmed that replacement of glutamic acid with glutamine in S1P1 changed ligand specificity from S1P to LPA, and the reciprocal mutation in LPA1 resulted in recognition of both LPA and S1P [7].
In the present study, the role of this residue in determining ligand selectivity for the S1P4 receptor was examined. Phylogenetic analysis of the Edg family of receptors indicates that S1P4 is more closely related to other S1P receptors than receptors which respond to LPA. However, S1P4 lies on the edge of the S1P family cluster and has been shown to bind S1P with lower affinity than other S1P receptors and hence it has been suggested that S1P is not the true endogenous agonist of this receptor [8]. We therefore decided to investigate whether replacement of this residue (E3.29(122)) with glutamine conferred LPA-responsiveness to the S1P4 receptor and hence determine the role of this residue in this lower-affinity S1P receptor. To achieve this, we expressed wild type and E3.29(122)Q mutant S1P4 receptors in CHO-K1 cells and studied responses to lysophospholipids using a [35S]GTPγS binding assay. Since CHO-K1 cells respond to LPA, we utilised fusion proteins constructed between the S1P4 receptor and a pertussis toxin-insensitive Gαi1(C351I) G protein. Expression of these proteins in CHO-K1 cells followed by treatment with pertussis toxin prior to harvest allowed elimination of any signal due to stimulation of endogenous LPA receptors. Within this study, we also examined how the length of the LPA acyl chain affected potency at the mutant S1P4 receptor, using a panel of naturally occurring LPA analogues. Computational models of complexes between the wild type or mutant S1P4 receptor and S1P and LPA species were used to provide a molecular interpretation of the experimental findings.
Results
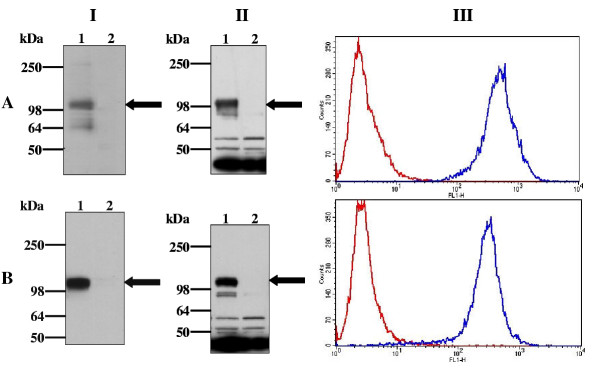
Human HA-S1P4 was mutated at position 122 to replace the naturally occurring glutamic acid with glutamine. The mutant and wild type receptors were stably expressed in CHO-K1 cells as in-frame GPCR-G protein fusions with pertussis toxin-insensitive Gαi1(C351I). Western blotting was used to detect expression of these fusion proteins. Membranes from HA-S1P4-Gαi1(C351I)- or HA-S1P4(E3.29(122)Q)-Gαi1(C351I)-transfected cells contained a polypeptide with an apparent molecular mass of approximately 110 kDa, which reacted with anti-HA and anti-SG1 antibodies (Figure 1A and 1B) and was consistent with expression of the GPCR-G protein fusion. Confirmation of comparable cell-surface expression of these proteins was obtained via FACS analysis using an anti-HA antibody directly conjugated with fluorescein (Figure 1 Panel III).
Figure 1.
Expression of HA-S1P4-Gαi1 and HA-S1P4(E122Q)-Gαi1(C351I) in CHO-K1 cells. Membranes from untransfected CHO-K1 cells (lane 2) and CHO-K1 cells stably expressing HA-S1P4-Gαi1(C351I) (A) or HA-S1P4(E122Q)-Gαi1(C351I) (B) were analysed by Western blotting using anti-HA (panel I) or anti-Gαi1 (panel II) antibodies. Visualisation of immunoreactive proteins was achieved using chemiluminescence after incubation of the blot with appropriate HRP-conjugated secondary antibodies. The position of each HA-S1P4 fusion protein is indicated by an arrow. Cell-surface expressed HA-S1P4 receptor was detected by FACS analysis (panel III) using a Fluorescein conjugate of the anti-HA antibody (blue line). Cells were also stained with an isotype matched control antibody (red line). No staining of untransfected CHO-K1 cells was observed using the Fluorescein conjugate of the anti-HA antibody (not shown). Data are presented as overlay histograms and are representative of at least five independent experiments.
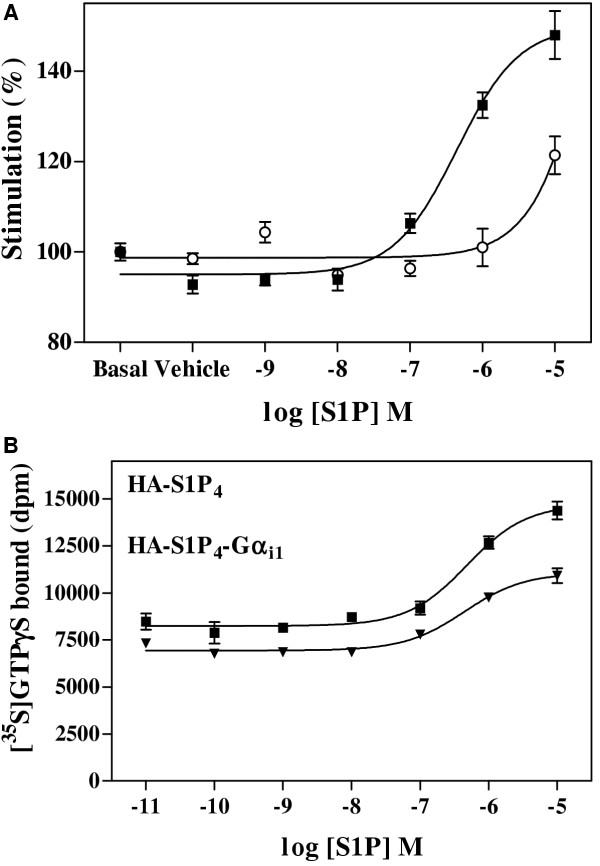
The response of HA-S1P4-Gαi1(C351I) and HA-S1P4(E3.29(122)Q)-Gαi1(C351I) to S1P was assessed using membranes from cells transfected to express these proteins and treated with pertussis toxin prior to harvest. S1P promoted dose-dependent increase in [35S]GTPγS binding to membranes containing HA-S1P4-Gαi1(C351I) with an EC50 of 355 nM ± 155 nM (n = 3); in contrast, membranes from HA-S1P4(E3.29(122)Q)-Gαi1(C351I)-expressing cells demonstrated severely impaired response to S1P (Figure 2A). The EC50 for S1P stimulation of the HA-S1P4-Gαi1 fusion protein (355 ± 155 nM) was not statistically significantly different from that obtained using the unfused HA-S1P4 receptor (439 ± 187 nM, Figure 2B) and compared favourably with published values for this receptor in HEK293T cells from two different research groups of 270 nM [9] and 790 nM [10].
Figure 2.
Sensitivity of HA-S1P4-Gαi1(C351I) and HA-S1P4(E3.29(122)Q)-Gαi1(C351I) to S1P in [35S]GTPγS binding assay. A. Membranes from CHO-K1 cells transfected with HA-S1P4-Gαi1(C351I) (filled squares) or HA-S1P4(E3.29(122)Q)-Gαi1(C351I) (open circles) which had been cultured in the presence of 100 ng/mL pertussis toxin for 24 hours prior to harvest were stimulated with varying concentrations of S1P for 30 minutes at 30°C in the [35S]GTPγS binding assay. Gαi G proteins were immunoprecipitated after solubilisation and preclearance with non-immune serum. Data are the mean of three determinations ± SEM from a single experiment, and are representative of three such experiments performed. B. Dose-dependent stimulation of wild-type HA-S1P4 (filled squares) and HA-S1P4-Gαi1 fusion protein (filled triangles) by S1P measured as described in panel A. These data are representative of three such experiments performed and analysis of mean EC50 values obtained for each protein showed them to be not statistically different.
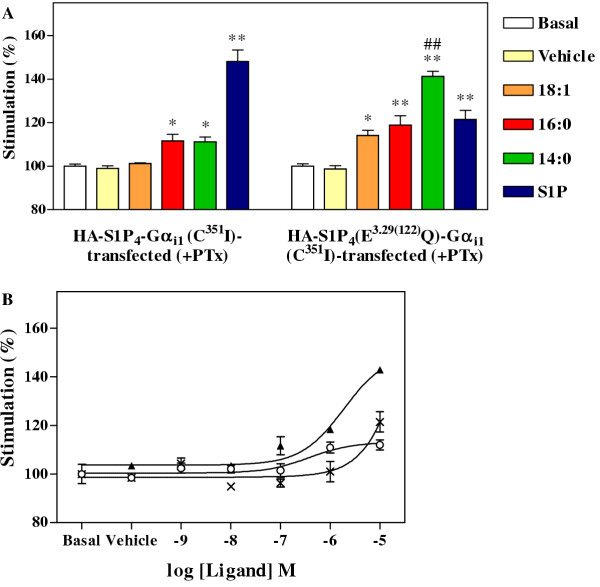
Structure activity relationships were determined by the [35S]GTPγS assay for S1P4 or its E3.29(122)Q mutant using S1P and LPA species with 14:0, 16:0 and 18:1 acyl chains at a single (10 μM) concentration. Of the lysophospholipids tested, only S1P induced a strong response over basal levels (approximately 48% ± 5%) in membranes containing HA-S1P4-Gαi1 (Figure 3A), whilst 18:1 LPA did not stimulate a statistically significant response; weak stimulation of [35S]GTPγS binding was observed with 14:0 and 16:0 LPA (approximately 11% ± 3% above basal in each case). In contrast, membranes containing HA-S1P4(E3.29(122)Q)-Gαi1(C351I) showed at least weak response to each ligand (Figure 3A). The weakest agonist at the E3.29(122)Q mutant S1P4 receptor was 18:1 LPA, which produced only 14% ± 2% stimulation over basal. S1P and 16:0 LPA gave approximately 21% ± 4% and 19% ± 4% stimulation over basal response, respectively. The best agonist for the E3.29(122)Q mutant was 14:0 LPA, which gave a 40% ± 2% enhancement over basal levels. The stimulation promoted by 14:0 LPA was statistically different from that produced by 18:1 LPA (p < 0.01). Sensitivity of the receptor to stimulation by each form of LPA, and particularly 14:0 LPA, was markedly increased after introduction of the E3.29(122)Q mutation and indicated that this position was important in influencing HA-S1P4 ligand preference.
Figure 3.
Ligand preference of HA-S1P4(E122Q)-Gαi1(C351I) in [35S]GTPγS binding assay. Membranes were stimulated for 30 minutes at 30°C with lysophospholipid ligands in the [35S]GTPγS binding assay and Gαi G proteins immunoprecipitated after solubilisation and preclearance with non-immune serum. (A) Membranes from CHO-K1 cells transfected to express HA-S1P4-Gαi1(C351I) or the mutant HA-S1P4(E3.29(122)Q)-Gαi1(C351I) and incubated with 100 ng/mL pertussis toxin for 24 hours prior to harvest, were left untreated (basal), or treated with vehicle or 10 μM concentrations of 18:1 LPA, 16:0 LPA, 14:0 LPA or S1P. Data are the mean of three determinations ± SEM from a single experiment and are representative of three such experiments performed. Statistical significance from the basal responses of each set of membranes tested is denoted by * (P < 0.05) or ** (P < 0.01); ## denotes statistical significance from the response to 18:1 LPA (P < 0.01) for the HA-S1P4(E3.29(122)Q)-Gαi1(C351I)-transfected membranes. (B) Membranes from CHO-K1 cells transfected to express HA-S1P4(E122Q)-Gαi1(C351I) and incubated with 100 ng/mL pertussis toxin for 24 hours prior to harvest, were stimulated with various concentrations of S1P (crosses), 18:1 LPA (open circles) or 14:0 LPA (filled triangles). Data are the mean of three determinations ± SEM and are representative of three such determinations performed.
These results indicate that introduction of the E3.29(122)Q mutation in the S1P4 receptor confers LPA-responsiveness, and that a short form of LPA was a more effective agonist than the intermediate and longer forms, when tested at this single concentration. Dose response curves were constructed for ligand-induced activation of the E3.29(122)Q S1P4 mutant by the 14:0 and 18:1 forms of LPA as well as S1P (Figure 3B). An EC50 could only be determined for the 14:0 form of LPA as S1P and 18:1 LPA caused minimal stimulation at only the highest concentration tested. The EC50 value for activation of HA-S1P4(E3.29(122)Q)-Gαi1(C351I) was calculated to be 3.8 ± 1.4 μM. However, since a plateau of maximal stimulation was not achieved, interpretation of this EC50 value needs caution. This result clearly showed that 14:0 LPA was a weak agonist of HA-S1P4(E3.29(122)Q)-Gαi1(C351I) and hence confirmed the involvement of residue 122 in S1P4 ligand preference. Similar results were obtained using a second CHO-K1 clone expressing this fusion protein (not shown).
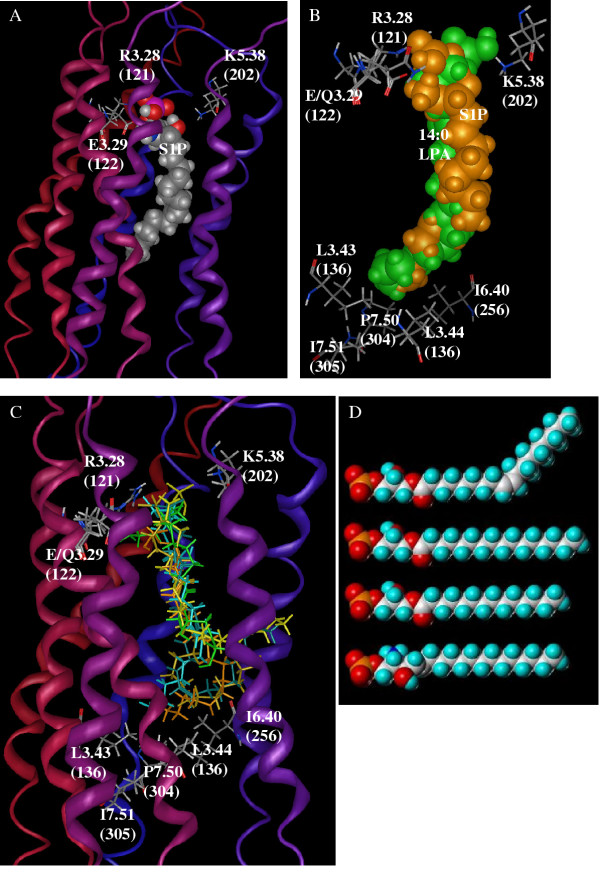
Computational modelling of the S1P complex with the wild type S1P4 receptor identifies the best S1P binding site within the TM with the phosphate group at the extracellular end (Figure 4A). Ion pairs appear between the phosphate group of S1P and two cationic amino acids, R3.28(121) and K5.38(202). An additional ion pair occurs between the cationic ammonium of S1P and E3.29(122). Hydrophobic residues from TM2, TM3, TM5 and TM6 line the binding pocket and surround the alkyl chain of S1P.
Figure 4.
Computational models of wild type S1P4 and its E3.29(122)Q mutant with S1P and LPA species. Computational models of the complexes between the wild type S1P4 or its E3.29(122)Q mutant with S1P or various LPA species generated by Autodock 3.0 and minimised using the MMFF94 forcefield in the MOE program. Complexes in each panel are shown from the same viewpoint with the extracellular end of the receptors oriented to the top of the figure. Standard element color codes are used with grey, white red, blue and magenta representing carbon, hydrogen, oxygen, nitrogen and phosphorous. Ribbons are shaded from red at the amino-terminus to blue at the carboxy-terminus. (A) Model of the complex between S1P (spacefilling) and the wild type S1P4 receptor. Residues in the receptor involved in ion pairs with S1P are shown as stick models and labelled. (B) Superimposition of the wild type S1P4 complex with S1P (orange) and the E3.29(122)Q S1P4 mutant complex with 14:0 LPA (green). For clarity, the only position at which the modelled amino acid position is shown for both receptor models is 3.29(122). Other residues had very similar optimised positions in the two model structures. (C) Superimposition of wild type S1P4 complexes with 18:1 LPA (cyan), 16:0 LPA (yellow) and 14:0 LPA (green) on E3.29(122)Q mutant complexes with 18:1 LPA (blue-green), 16:0 LPA (gold) and S1P (orange). For clarity, the only position at which modelled amino acid position is shown for both the wild type and mutant receptor models is 3.29(122). Other residues had very similar optimised positions in all model structures. (D) Space-filling models which represent the minimised extended conformation of each structure were constructed using SYBYL 6.9 software (Tripos Inc., St. Louis, MO., U.S.A.). The distance between phosphorus and terminal carbon atoms was predicted for each structure listed from top to bottom: 18:1 LPA, 27.0 Å; 16:0 LPA, 26.7 Å; 14:0 LPA, 24.2 Å; S1P, 24.0 Å.
The best complex of 14:0 LPA in the E3.29(122)Q S1P4 mutant receptor model has striking similarity to the best complex of S1P in the wild type S1P4 receptor model (Figure 4B). Both models demonstrate ion pairing between the phosphate group and two cationic amino acids, R3.28(121) and K5.38(202). Each ligand interacts with the amino acid at position 3.29(122), S1P by an ion pair with the carboxylate of the wild type glutamate and 14:0 LPA by a hydrogen bond with the mutated glutamine. Multiple hydrophobic residues surround the nonpolar tails of the lipid ligands. The superimposition of the two complexes (Figure 4B) also demonstrates that the ligands occupy almost identical volumes. Common interactions and overlap volumes are qualitatively consistent with the experimental findings that these ligands give similar 48% and 40% maximal stimulation over basal for S1P at the wild type and 14:0 LPA at the mutant receptor, respectively.
In contrast to the complexes of 14:0 LPA with E3.29(122)Q S1P4 and S1P with wild type S1P4, the remaining complexes show much less common volume (Figure 4C). Most complexes exhibit the phosphate interactions described for 14:0 LPA with E3.29(122)Q S1P4 and S1P with wild type S1P4. Of particular interest is the observation that the best complexes generated by Autodock for the 18:1 LPA species with wild type S1P4 has a very high positive van der Waals interaction energy, > 3000 kcal/mol, compared to values well under 200 kcal/mol for every other complex studied. In the best complexes found for 16:0 LPA and 18:1 LPA in both constructs, the terminal six to eight carbons of the hydrophobic tails fold into L-shaped conformations quite different from the extended conformations observed in the S1P complex with wild type S1P4 or the 14:0 LPA complex with the E3.29(122)Q S1P4 mutant. The terminal carbons in several complexes curl out of the receptor between TM5 and TM6 (Figure 4C) due to the restricted length of the binding pocket. These results suggest that the complete lack of S1P4 activation in response to 18:1 LPA is likely due to failure to form a complex. The strongest complexes formed, S1P with wild type S1P4 and 14:0 LPA with the E3.29(122)Q S1P4 mutant, have complementary interactions with the residue at position 3.29(122). These strong complexes give the most robust activation. Weak complexes are formed for other combinations due to mismatched interactions with position 3.29(122) or excessive length of the hydrophobic tail. The presence of hydrophobic tails of 16:0 or 18:1 LPA between transmembrane domains may additionally impair the conformational change necessary for full agonist responses.
Discussion
Parental CHO-K1 cells respond to LPA in functional assays, reflecting expression of endogenous LPA1 (G. Holdsworth, et al., manuscript in preparation). For this reason, fusion proteins between wild type or mutant HA-S1P4 and the pertussis toxin-insensitive Gαi1(C351I) G protein were used in these studies. Expression of these proteins in CHO-K1 cells followed by treatment with pertussis toxin prior to harvest allowed elimination of any signal due to stimulation of endogenous LPA receptors. McAllister et al. [11] (.(adopted a similar approach for studies of the LPA1 receptor.
We examined the role of residue E3.29(122) in controlling S1P4 ligand selectivity using functional and computational methods. This residue, which is conserved throughout the S1P receptors, has been shown to control ligand specificity for the related S1P1 receptor [7]. Introduction of the E3.29(122)Q mutation severely affected the response of S1P4 to S1P: in dose-response experiments S1P caused minimal stimulation at only the highest concentration of ligand used. This is in agreement with published observations for activation of the equivalent S1P1 mutant [7]. 14:0 LPA was able to induce dose-dependent stimulation of S1P4(E3.29(122)Q) with an EC50 of approximately 3.8 μM but only promoted minimal stimulation of the wild type S1P4 receptor. The modelled complexes of 14:0 LPA with E3.29(122)Q S1P4 and S1P with wild type S1P4 demonstrate nearly identical volumes occupied by the two ligands and very similar interactions between these ligands and their respective receptors. Of particular importance are amino acid residues at positions 3.28(121), 3.29(122) and 5.38(202), which either ion pair with the phosphate or interact with the 2-amino or 2-hydroxyl group in S1P and 14:0 LPA, respectively. The importance of interactions with amino acids at positions 3.28 and 3.29 has been previously noted for the S1P1 [6,7] and LPA1,2,3 [7,12] receptors.
The S1P4 receptor exhibits marked constitutive (agonist-independent) activity (Figure 5) which was unaffected by the introduction of the E3.29(122)Q mutation (data not shown). This indicates that the mutation perturbs S1P recognition without affecting the ability of the receptor to spontaneously adopt an active conformation. Similar observations have been reported for the β2AR, where a mutation in the sixth transmembrane domain abolished agonist activation but not constitutive activity [13].
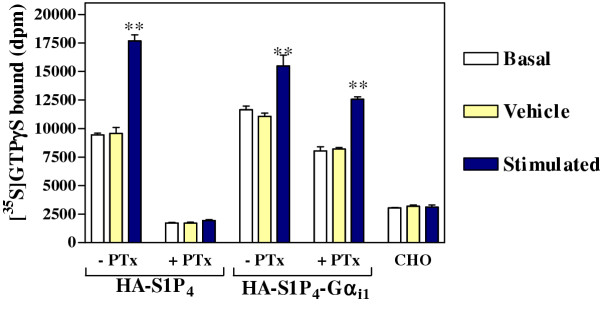
Figure 5.
Constitutive activity of HA-S1P4 and HA-S1P4-Gαi1(C351I). Comparison of basal, vehicle and stimulated (10 μM S1P) GTPγS binding for membranes prepared from CHO cells transfected to express HA-S1P4 or HA-S1P4 fused to PTx insensitive Gαi1. Where indicated, cells were treated with 100ng/ml PTx for 24 hours prior to harvesting. PTx treatment of HA-S1P4-transfected membranes prevented activation by S1P and also caused a dramatic reduction in basal signalling, indicative of constitutive activity. In contrast, when the HA-S1P4-Gαi1 fusion-expressing membranes were treated with PTx, there was only a slight reduction in basal signalling and the receptor still responded to exogenous S1P, indicating that the receptor signalled via the tethered, PTx-insensitive G protein. Basal signalling in PTx-treated HA-S1P4-Gαi1-expressing membranes exceeded that seen with membranes from untransfected CHO cells.
Unlike S1P, which exists as a single species in vivo, the term LPA actually refers to a family of molecules that take the general form 1-o-acyl-2-hydroxy-sn-glyceryl-3-phosphate. Naturally occurring forms of LPA contain acyl chains of differing lengths, with differing degrees of saturation. Investigations into the effect of the length and degree of saturation of the acyl chain of LPA have been undertaken for the LPA receptors [14,15], but limited SAR information is available for S1P receptors (22). The LPA-responsive E3.29(122)Q S1P4 mutant facilitates structure activity relationship (SAR) studies due to the greater availability of LPA analogs relative to S1P analogs. Comparison of space-filling models of the structures of S1P and three analogues of LPA (Figure 4D) revealed that 14:0 LPA most closely resembled S1P in terms of apparent length. [35S]GTPγS binding assays demonstrated greater agonist activity of 14:0 LPA at the mutant receptor relative to 18:1 or 16:0 LPA. This SAR indicates a length restriction for the S1P4 agonist binding site. Model complexes of 16:0 and 18:1 LPA contained alkyl chains that fold at the bottom of the binding pocket, defined by a cluster of hydrophobic amino acids. Three of these differ either in position of sidechain branching or size relative to LPA receptors and the other S1P receptors. Position 2.46, I88 in S1P4, is leucine in LPA1–3 and other S1P receptors. Residue I6.40(256) is larger than the valine found in the other four S1P receptors, LPA1 and LPA3. Finally, I7.51(305) corresponds to the smaller valine in S1P2 and S1P3 and the much smaller alanine in LPA2. These findings provide a molecular explanation for a similar SAR observed using para-alkyl amide analogs of S1P [9]. SAR obtained with the S1P4 mutant are in contrast to that shown by LPA receptors, which exhibit the general trend of 18:1 ≥ 16:0 > 14:0 for potency and maximal stimulation [15].
Since mutation of residue 122 in the S1P4 receptor from the naturally occurring glutamic acid to glutamine conferred responsiveness to 14:0 LPA and severely affected responses to S1P, our observations support the hypothesis that this conserved residue in the third transmembrane domain of the S1P receptors is involved in ligand recognition. This is in contrast to a recent paper describing models of several GPCRs, including S1P4, which had been generated using novel first principle methods [16]. In this model of S1P4, interactions between S1P and residues T7.34(127) and W7.37(291) and E7.30(284) were observed. Interaction of E7.30(284) with the ammonium group of S1P appeared to control ligand selectivity since the other residues appeared to interact with the phosphate group, which is present on both LPA and S1P. It is therefore surprising that none of these residues are conserved throughout the S1P or LPA receptor families. The data presented here support the assertion that glutamic acid residue 3.29 present in the third transmembrane domain of the S1P receptors controls ligand selectivity and suggest that the S1P4 model described by Vaidehi et al. [16] is inaccurate.
The current study provides new information for the development of more selective S1P receptor agonists. In particular, an S1P analog with its hydrophobic chain extended by either 2 or 4 carbons would be a very poor agonist of the S1P4 receptor. On the other hand, the activation of the S1P1-E3.29(121)Q mutant by 18:1 LPA [7] indicates that a chain-extended S1P analog should retain agonist activity at the S1P1 receptor. S1P receptor agonists with differing selectivity profiles will be useful tools to more completely map the physiological and pathophysiological roles of these receptors.
Conclusions
These studies confirm that glutamic acid residue 3.29, present in the third transmembrane domain of the S1P receptors is important for the selective recognition of S1P, versus the closely related lipid, LPA. Mutation of E3.29 to glutamine diminished response to S1P and allowed structure activity studies using the diverse available LPA species. The mutant S1P4 receptor is stimulated most strongly by LPA 14:0 and is not activated by the longer LPA 18:1, in contrast with a previous report on the analogous S1P1 receptor mutant that responded to LPA 18:1. Thus the S1P4 receptor ligand binding pocket is shorter in length than the S1P1 ligand binding pocket.
Methods
Residue nomenclature
Amino acids within the TM of S1P4 can be assigned index positions to facilitate comparison between receptors with different numbers of amino acids, as described by Weinstein and coworkers [17]. An index position is in the format x.xx. The first number denotes the TM in which the residue appears. The second number indicates the position of that residue relative to the most highly conserved residue in that TM which is arbitrarily assigned position 50. E3.29, then, indicates the relative position of glutamate 122 in TM3 relative to the highly conserved arginine 143 in the E(D)RY motif which is assigned index position 3.50 [17].
Materials
Materials for tissue culture were supplied by Invitrogen Ltd. (Paisley, Scotland, U.K.). Foetal bovine serum was obtained from Helena Biosciences Ltd., (Sunderland, U.K.) or PAA Labs GmbH., (Linz, Austria). Pertussis toxin was purchased from CN Biosciences Ltd., (Nottingham, U.K.). Lysophosphatidic acid (18:1, 16:0 and 14:0) and S1P were from Avanti Polar Lipids Inc., (Alabaster, AL., U.S.A.). The SG1 antiserum was produced previously [18]. All other chemicals were from Sigma Aldrich Company Ltd., (Gillingham, Dorset, U.K.) or BDH Ltd., (Poole, Dorset, U.K.) unless stated otherwise.
Construction of receptor expression plasmids
The S1P4 coding sequence was cloned from a human PBMC cDNA library using the sense primer 5'-GAGAGAGCGGCCGCCACCATGTATCCATATGATGTTCCAGATTATGCTAACGCCACGGGGACCCCGGTG-3', which contains a NotI restriction site (bold) and the haemagglutinin HA epitope tag (YPYDVPVYA, underlined) immediately after the initiator methionine, and the antisense primer 5'-GAGAGAGAATTCGGCGATGCTCCGCACGCTGGAGATG-3', which contains an EcoRI restriction site (bold) and changes the S1P4 stop codon to alanine (underlined). A C351I mutant of the Gαi1 G protein (previously produced, [19]) was amplified using PCR with the sense cloning primer 5'-GAGAGAGAATTCGCCA CCATGGGCTGCACACTGAGCG-3', which contains the EcoRI restriction site (bold), and the antisense cloning primer 5'-GAGAGAGGATCCTTAGAAGAGACCGATGTCTTTTA G-3', which contains a BamHI restriction site (bold). After digestion of each PCR product with the appropriate restriction enzymes, fragments were ligated into the pIRESpuro mammalian expression vector (Invitrogen Ltd.) to generate an in-frame fusion between HA-S1P4 and Gαi1(C351I).
The E3.29(122)Q mutation was introduced into the S1P4 sequence in parallel PCR reactions. Complementary oligonucleotides were designed across the residue which was to be mutated such that each primer contained the necessary base change to mutate residue 122 to glutamine (underlined in each primer): sense mutational primer: 5'-CAGTGGTTCCTACGGCAGGGCCTGCTCTTCAC-3'; antisense mutational primer: 5'-GTGAAGAGCAGGCCCTGCCGTAGGAACCACTG-3'. Mutational sense or antisense primers were used in parallel PCR reactions with the appropriate antisense or sense cloning primer, with HA-S1P4 plasmid DNA as template. Equimolar amounts of each purified PCR product were mixed and amplified in a further reaction, using the cloning primers described above. The resultant product was digested with the appropriate restriction enzymes and ligated with the Gαi1 sequence in the pIRESpuro expression vector to generate an in-frame fusion between HA-S1P4(E3.29(122)Q) and Gαi1(C351I).
Cell culture and transfection
CHO-K1 cells were maintained at 37°C with 5% CO2 in Dulbecco's modified Eagle's medium (DMEM), supplemented with 10% foetal bovine serum (FBS), 2 mM L-glutamine and non-essential amino acids. Sub-confluent cell monolayers were stably transfected to express either HA-S1P4-Gαi1(C351I) or HA-S1P4(E3.29(122)Q)-Gαi1(C351I) fusion proteins using Lipofectamine reagent (Invitrogen). 72 hours post-transfection, cells were seeded in media supplemented with 7.5 μg/mL puromycin and the resultant clones examined for expression of cell surface receptor using FACS analysis. Clonal cell lines were expanded in complete DMEM containing 7.5 μg/mL puromycin and were transferred to serum free DMEM approximately 24 hours prior to harvesting. Where indicated, 100 ng/mL pertussis toxin was included in the serum free medium.
It should be noted that we initially expressed S1P4 in RH7777 cells, which are unresponsive to S1P and LPA and have been commonly used for studies of Edg family receptors [20]. Unfortunately, our attempts to detect activation of S1P4 expressed in these cells using a variety of functional assays were unsuccessful. Therefore, we used CHO-K1 cells as an alternative host in these studies; expression of functional S1P4 in CHO-K1 cells has also been reported by Mandala et al. [21].
FACS analysis
The amino-terminal HA-epitope tag was detected using a fluorescein conjugate of the anti-HA antibody, clone 3F10 (Roche Molecular Biochemicals Ltd., Lewes, U.K.). Cells were harvested non-enzymatically and washed with FACS buffer (PBS containing 3% FBS and 0.1% NaN3) then stained with the 3F10 antibody (or an isotype matched control) for 40 minutes at 4°C in the dark. After washing with FACS buffer, cells were analysed using a FACScalibur flow cytometer (BD Biosciences, Oxon., U.K.).
Preparation of cell membranes
Cells were harvested non-enzymatically, washed with PBS and resuspended in "assay buffer" (20 mM Hepes, pH 7.4, 3 mM MgCl2, 100 mM NaCl), supplemented with "complete" protease inhibitors (Roche Molecular Biochemicals Ltd.). Cells were homogenised in a nitrogen cavitation chamber (500 psi for 15 minutes). Unbroken cells and nuclei were pelleted by centrifugation (500 × g, 10 minutes, 4°C) and the supernatant fraction was centrifuged at 45,000 × g for 45 minutes at 4°C. Membrane pellets were resuspended in assay buffer, titurated through a fine gauge needle and stored at -80°C until required.
Immunoblot analysis
Samples were resolved by SDS-Page on 4–20% Tris-Glycine gels (Invitrogen) and were transferred to Immobilon-P membrane (Millipore Ltd., Herts., U.K.). The membrane was blocked using 2.5% Marvel in PBS before incubating with primary antibodies which had been diluted in PBS/0.1% Tween-20 containing 1% Marvel. The high affinity rat anti-HA antibody was diluted 1 in 500; the anti-Gαi1 antibody (Autogen Bioclear Ltd., Wilts., U.K.) was diluted 1 in 1000. Immunoreactivity was detected using an appropriate horseradish peroxidase-conjugated secondary antibody, diluted 1 in 10,000 in PBS/0.1% Tween-20 containing 1% Marvel, followed by detection using SuperSignal reagents (Perbio Science Ltd., Cheshire, U.K.).
[35S]GTPγS binding assay
[35S]GTPγS binding experiments were performed essentially as described previously [22]. Briefly, membranes were incubated with or without the indicated ligand for 30 minutes at 30°C in assay buffer containing [35S]GTPγS (100 nCi/point), saponin (20 μg/point) and 0.1 μM GDP. 18:1 LPA was prepared as a 2 mM DMSO stock whilst 16:0 and 14:0 LPA were prepared as 2 mM stock solutions in 1:1 ethanol:water per supplier recommendation due to their poor solubility in DMSO. S1P had previously been dispensed as thin film aliquots (dissolved in MeOH and the solvent evaporated under nitrogen) in brown glass vials and stored at -70°C prior to use. Lipids (S1P or LPA forms) were diluted in assay buffer containing 1% fatty acid free BSA, such that the final concentration of BSA in the assay was 0.1%. Following incubation, membrane protein was solubilised with 1.25% NP-40 and 0.4% SDS and after pre-clearance using non-immune serum, Gαi1/2 subunits were immunoprecipitated with SG1 antiserum, used at a dilution of 1 in 200. Non-specific binding was determined by the addition of 100 μM GTPγS. Bound radioactivity was measured using liquid scintillation counting.
Experimental data analysis
Numerical data are expressed as means ± standard error, shown as error bars in the appropriate figures. Statistical comparisons were made using one-way ANOVA with Dunnett's multiple comparison post test.
Receptor model development
A model of human S1P4 (GenBank™ accession number AAP84350) was developed by homology to the experimentally-validated model of S1P1 [23]. Alignment of the S1P receptor sequences was performed using the MOE software package (version 2003. 01 ed. Chemical Computing Group, Montreal, Canada). The alignment was optimised by the manual removal of gaps within the TM, and alignment in the region of TM5 was shifted one position to correctly orient K5.38(202) toward the interior of the helical bundle (Pham, et al., unpublished data). A preliminary model was generated by homology modelling using default parameters and subsequently manually refined to optimise interhelical hydrogen bonding. Cis-amide bonds present in the loop regions were converted to the trans conformation by manual rotation followed by the minimisation of two residues on either side of the amide linkage to a root mean square (RMS) gradient of 0.1 kcal/mol·Å using the MMFF94 forcefield [24]. After these manual refinements, the receptor model was optimised using the MMFF94 forcefield to an RMS gradient of 0.1 kcal/mol·Å.
A model of S1P4 with the E3.29(122)Q mutation was developed by performing the appropriate mutation in MOE, and saturating the residue with hydrogen atoms. To allow the sidechains of the other residues in the binding pocket to adapt to the presence of the new moiety, the backbone atoms of the receptor were fixed and the receptor was optimised to an RMS gradient of 0.1 kcal/mol·Å using the MMFF94 forcefield [24].
Ligand model development
Computational models of the naturally-occurring stereoisomers of 14:0 LPA, 16:0 LPA, 18:1 LPA, and S1P were built using the MOE software package. The -1 ionization state for the phosphate functionality was chosen for all ligands, and the +1 ionization state was chosen for the amine moiety of S1P. Previous docking studies using the -2 ionization state for phosphate in related systems yield essentially identical geometries as studies using the -1 ionization state. These ligands were geometry optimised using the MMFF94 force field [24].
Docking
Using the AUTODOCK 3.0 software package [25], 14:0 LPA, 16:0 LPA, 18:1 LPA, and S1P were docked into the S1P4 wild type and S1P4 E3.29(122)Q mutant receptor models. Each docking box was centered near F3.33(126) with dimensions of 30.75 × 23.25 × 23.25 or 32.25 × 23.25 × 23.25 Å for shorter (S1P and 14:0 LPA) or longer (16:0 and 18:1 LPA) ligands, respectively. At least 20 putative complexes were generated for each receptor:ligand pair using docking parameters at default values with the exception of the number of energy evaluations (2.5 × 108), generations (10000) and maximum iterations (3000). Resultant complexes were evaluated based on final docked energy, Van der Waals interaction energies from the MMFF94 forcefield as well as visual analysis. The complexes with the lowest final docked energies and others of interest were geometry optimised using the MMFF94 force field [24], and the lowest energy complex after minimisation was chosen as the final complex structure.
Abbreviations
CHO, Chinese hamster ovary; Edg, endothelial differentiation gene; ERK, extracellularly regulated kinase; FACS, fluorescence activated cell sorter; G protein, guanine nucleotide-binding protein; GPCR, G protein-coupled receptor; HA, haemagglutinin; LPA, lysophosphatidic acid; MAP, mitogen-activated protein kinase; PBMC, peripheral blood mononuclear cell; PTx, pertussis toxin; SIP, sphingosine-1-phosphate; TM, transmembrane domain
Authors' contributions
G Holdsworth performed and interpreted all studies with experimental S1P4 fusion proteins and drafted the manuscript. D Osborne performed and interpreted docking studies to generate all mutant complexes with all LPA species and S1P and all wild type complexes with LPA species. TC Pham generated the homology model of the human S1P4 receptor. J Fells performed docking studies of S1P with the wild type S1P4 receptor. G Hutchinson and G Milligan participated in the design and coordination of the experimental studies with S1P4 fusion proteins. A Parrill participated in the design and coordination of the modelling studies and edited the manuscript.
Acknowledgments
Acknowledgments
We gratefully acknowledge the assistance of Jim Turner (Celltech R&D Ltd.) for production of lipid space-filling models and Gabor Tigyi (University of Tennessee Health Sciences Center) for critically reading the manuscript. This work was supported in part by grants from NIH (1 RO1 CA92160-01) and the American Heart Association (awards 0050006N and 0355199B). The Chemical Computing Group generously donated the MOE program.
Contributor Information
Gill Holdsworth, Email: gill.holdsworth@celltechgroup.com.
Daniel A Osborne, Email: daosborn@memphis.edu.
TrucChi Thi Pham, Email: tcpham@memphis.edu.
James I Fells, Email: jfells@memphis.edu.
Gillian Hutchinson, Email: gillian.hutchinson@celltechgroup.com.
Graeme Milligan, Email: G.Milligan@bio.gla.ac.uk.
Abby L Parrill, Email: aparrill@memphis.edu.
References
- Pyne S, Pyne NJ. Sphingosine 1-phosphate Signalling and Termination at Lipid Phosphate Receptors. Biochim Biophys Acta. 2002;1582:121–131. doi: 10.1016/S1388-1981(02)00146-4. [DOI] [PubMed] [Google Scholar]
- Fukushima N, Ishii I, Contos JJ, Weiner JA, Chun J. Lysophospholipid Receptors. Annu Rev Pharmacol Toxicol. 2001;41:507–534. doi: 10.1146/annurev.pharmtox.41.1.507. [DOI] [PubMed] [Google Scholar]
- Spiegel S, Milstien S. Sphingosine-1-phosphate: An Enigmatic Signalling Lipid. Nat Rev Mol Cell Biol. 2003;4:397–407. doi: 10.1038/nrm1103. [DOI] [PubMed] [Google Scholar]
- Hla T. Signaling and Biological Actions of Sphingosine 1-phosphate. Pharmacol Res. 2003;47:401–407. doi: 10.1016/S1043-6618(03)00046-X. [DOI] [PubMed] [Google Scholar]
- Chun J, Goetzl EJ, Hla T, Igarashi Y, Lynch KR, Moolenaar W, Pyne S, Tigyi G. International Union of Pharmacology. XXXIV. Lysophospholipid Receptor Nomenclature. Pharmacol Rev. 2002;54:265–269. doi: 10.1124/pr.54.2.265. [DOI] [PubMed] [Google Scholar]
- Parrill AL, Baker DL, Wang D, Fischer DJ, Bautista DL, van Brocklyn J, Spiegel S, Tigyi G. Structural Features of EDG1 Receptor-Ligand Complexes Revealed by Computational Modeling and Mutagenesis. In: Goetzl E J and Lynch K R, editor. Lysophospholipids and Eicosanoids in Biology and Pathophysiology. Vol. 905. New York, New York Academy of Sciences; 2000. pp. 330–339. [DOI] [PubMed] [Google Scholar]
- Wang D, Lorincz Z, Bautista DL, Liliom K, Tigyi G, Parrill AL. A Single Amino Acid Determines Ligand Specificity of the S1P1 (EDG1) and LPA1 (EDG2) Phospholipid Growth Factor Receptors. J Biol Chem. 2001;276:49213–49220. doi: 10.1074/jbc.M107301200. [DOI] [PubMed] [Google Scholar]
- Lynch KR, S. Im. D. Life on the Edg. Trends Pharmacol Sci. 1999;20:473–475. doi: 10.1016/S0165-6147(99)01401-7. [DOI] [PubMed] [Google Scholar]
- Clemens JJ, Davis MD, Lynch KR, Macdonald TL. Synthesis of Para-Alkyl Aryl Amide Analogues of Sphingosine-1-phosphate: Discovery of Potent S1P Receptor Agonists. Bioorg & Med Chem Letters. 2003;13:3401–3404. doi: 10.1016/S0960-894X(03)00812-6. [DOI] [PubMed] [Google Scholar]
- Brinkmann V, Davis MD, Heise CE, Albert R, Cottens S, Hof R, Bruns C, Prieschl E, Baumruker T, Hiestand P, Foster CA, Zollinger M, Lynch KR. The Immune modulator, FTY720, Targets Sphingosine 1-Phosphate Receptors. J Biol Chem. 2002;277:21453–21457. doi: 10.1074/jbc.C200176200. [DOI] [PubMed] [Google Scholar]
- McAllister G, Stanton JA, Salim K, Handford EJ, Beer MS. Edg2 Receptor Functionality: Gialpha1 Coexpression and Fusion Protein Studies. Mol Pharmacol. 2000;58:407–412. doi: 10.1124/mol.58.2.407. [DOI] [PubMed] [Google Scholar]
- Sardar VM, Bautista DL, Fischer DJ, Yokoyama K, Nusser N, Virag T, Wang D, Baker DL, Tigyi G, Parrill AL. Molecular Basis for Lysophosphatidic Acid Receptor Antagonist Selectivity. Biochim Biophys Acta. 2002;1582:309–317. doi: 10.1016/S1388-1981(02)00185-3. [DOI] [PubMed] [Google Scholar]
- Hannawacker A, Krasel C, Lohse MJ. Mutation of Asn293 to Asp in Transmembrane Helix VI Abolishes Agonist-induced but not Constitutive Activity of the beta(2)-adrenergic Receptor. Mol Pharmacol. 2002;62:1431–1437. doi: 10.1124/mol.62.6.1431. [DOI] [PubMed] [Google Scholar]
- Bandoh K, Aoki J, Hosono H, Kobayashi S, Kobayashi T, Murakami-Murofushi K, Tsujimoto M, Arai H, Inoue K. Molecular Cloning and Characterization of a Novel Human G-protein-coupled Receptor, EDG7, for Lysophosphatidic Acid. J Biol Chem. 1999;274:27776–27785. doi: 10.1074/jbc.274.39.27776. [DOI] [PubMed] [Google Scholar]
- Im DS, Heise CE, Harding MA, George SR, O'Dowd BF, Theodorescu D, Lynch KR. Molecular Cloning and Characterization of a Lysophosphatidic Acid Receptor, Edg-7, Expressed in Prostate. Mol Pharmacol. 2000;57:753–759. [PubMed] [Google Scholar]
- Vaidehi N, Floriano WB, Trabanino R, Hall SE, Freddolino P, Choi EJ, Zamanakos G, Goddard WAIII. Prediction of Structure and Function of G Protein-Coupled Receptors. Proc Nat Acad Sci, USA. 2002;99:12622–12627. doi: 10.1073/pnas.122357199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ballesteros JA, Weinstein H. Chapter 19. In: Conn P M and Sealfon S C, editor. Methods in Neurosciences. Vol. 25. San Diego, Academic Press; 1995. pp. 366–428. [Google Scholar]
- Grant KR, Harnett W, Milligan G, Harnett MM. Differential G-protein Expression During B- and T-cell Development. Immunology. 1997;90:564–571. doi: 10.1046/j.1365-2567.1997.00196.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bahia DS, Wise A, Fanelli F, Lee M, Rees S, Milligan G. Hydrophobicity of Residue351 of the G Protein Gi1 alpha Determines the Extent of Activation by the Alpha 2A-Adrenoceptor. Biochemistry. 1998;37:11555–11562. doi: 10.1021/bi980284o. [DOI] [PubMed] [Google Scholar]
- Zhang G, Contos JJ, Weiner JA, Fukushima N, Chun J. Comparative Analysis of Three Murine G-protein Coupled Receptors Activated by Sphingosine-1-phosphate. Gene. 1999;227:89–99. doi: 10.1016/S0378-1119(98)00589-7. [DOI] [PubMed] [Google Scholar]
- Mandala S, Hajdu R, Bergstrom J, Quackenbush E, Xie J, Milligan J, Thornton R, Shei G, Card D, Keohane C, Rosenbach M, Hale J, Lynch CL, Rupprecht K, Parsons W, Rosen H. Alteration of Lymphocyte Trafficking by Sphingosine 1-Phosphate Receptor Agonists. Science. 2002;296:346–349. doi: 10.1126/science.1070238. [DOI] [PubMed] [Google Scholar]
- Stevens PA, Pediani J, Carrille JJ, Milligan G. Coordinated Agonist Regulation of Receptor and G Protein Palmitoylation and Functional Rescue of Palmitoylation-Deficient Mutants of the G Protein G11alpha Following Fusion to the Alpha1b-adrenoreceptor: Palmitoylation of G11alpha is not Required for Interaction with Beta*gamma Complex. J Biol Chem. 2001;276:35883–35890. doi: 10.1074/jbc.M103816200. [DOI] [PubMed] [Google Scholar]
- Parrill AL, Wang D-A, Bautista DL, Van Brocklyn JR, Lorincz Z, Fischer DJ, Baker DL, Liliom K, Spiegel S, Tigyi G. Identification of Edg1 Receptor Residues that Recognize Sphingosine 1- Phosphate. J Biol Chem. 2000;275:39379–39384. doi: 10.1074/jbc.M007680200. [DOI] [PubMed] [Google Scholar]
- Halgren TA. Merck Molecular Force Field. I. Basis, Form, Scope, Parameterization, and Performance of MMFF94*. J Comp Chem. 1996;17:490–519. [Google Scholar]
- Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, Olson AJ. Automated Docking Using a Lamarckian Genetic Algorithm and an Empirical Binding Free Energy Function. J Comput Chem. 1998;19:1639–1662. doi: 10.1002/(SICI)1096-987X(19981115)19:14<1639::AID-JCC10>3.0.CO;2-B. [DOI] [Google Scholar]