Abstract
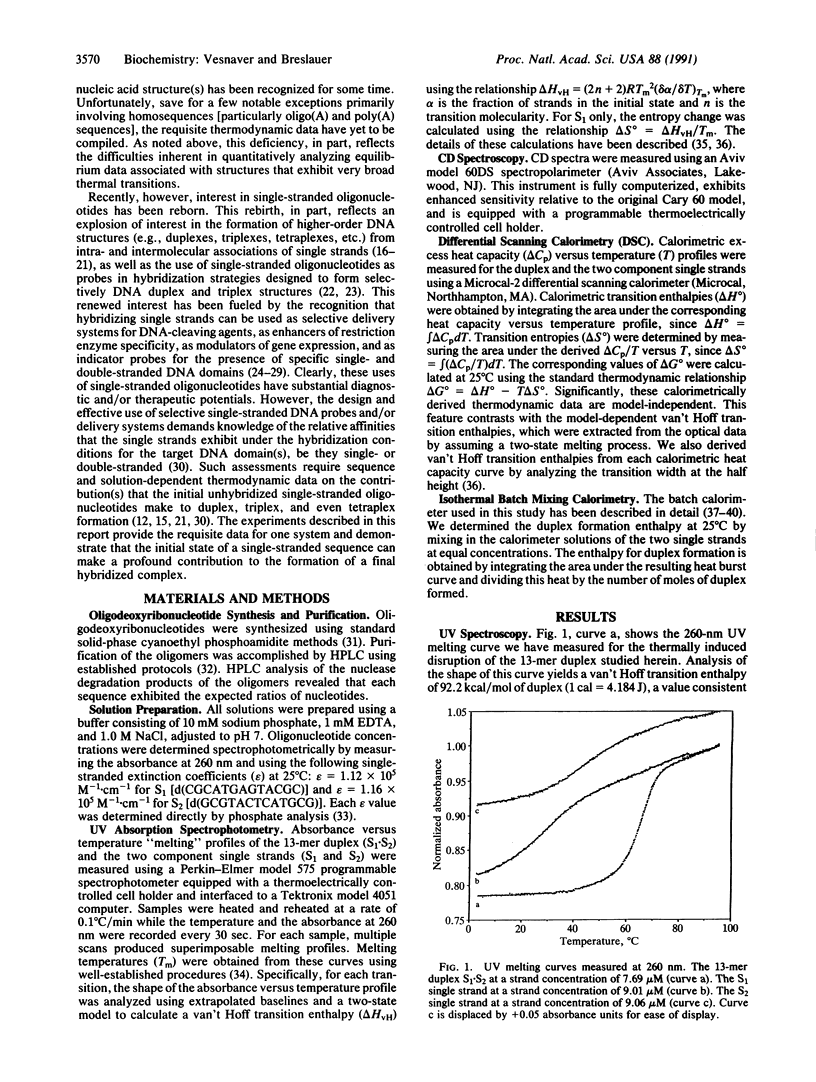
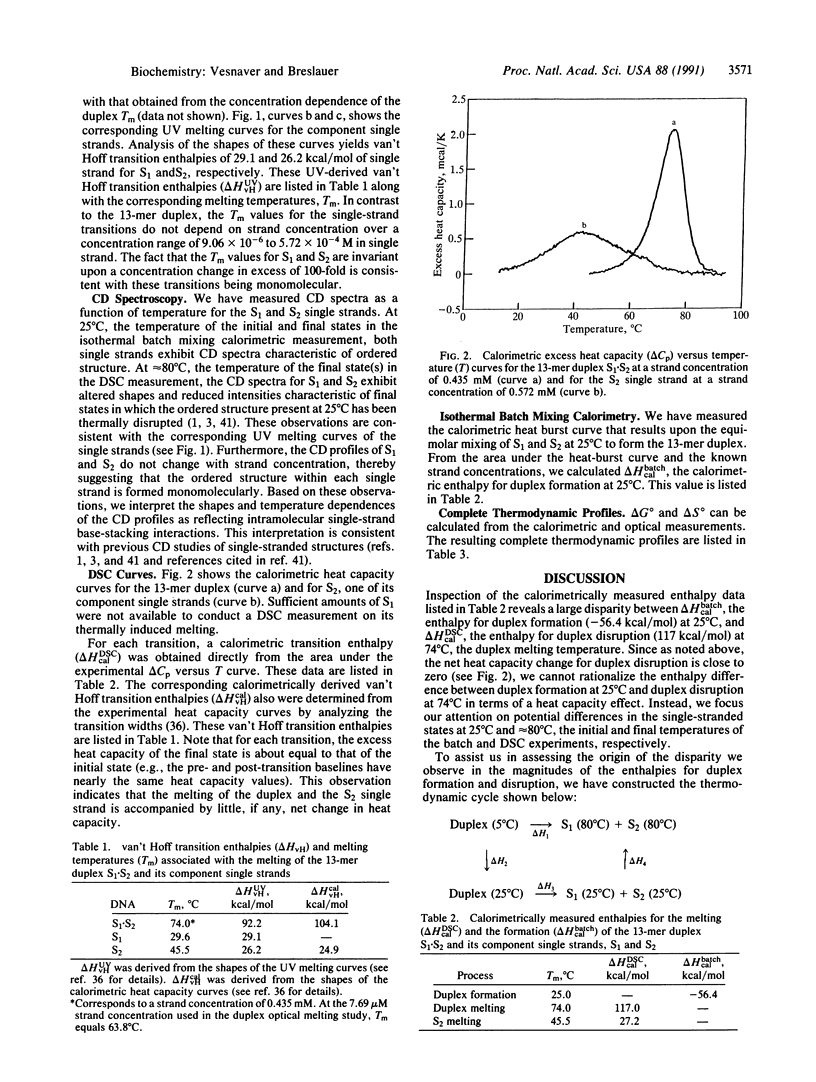
We report a direct determination of the thermodynamic contribution that DNA single-stranded order makes to DNA duplex formation. By using differential scanning calorimetry (DSC) and temperature-dependent UV absorbance spectroscopy, we have characterized thermodynamically the thermally induced disruption of the 13-mer duplex [d(CGCATGAGTACGC)].[d(GCGTACTCATGCG)] (henceforth called S1.S2) and its component single strands, [d(CGCATGAGTACGC)] (henceforth called S1) and [d(GCGTACTCATGCG)] (henceforth called S2). These spectroscopic and calorimetric measurements yield the following thermodynamic profiles at 25 degrees C: delta G degree = 20.0 kcal/mol, delta H degree = 117.0 kcal/mol, and delta S degree = 325.4 cal.degree-1.mol-1 for duplex melting of S1.S2; delta G degree = 0.45 kcal/mol, delta H degrees = 29.1 kcal/mol, and delta S degree = 96.1 cal.degree-1.mol-1 for single-strand melting of S1; delta G degree = 1.44 kcal/mol, delta H degree = 27.2 kcal/mol, and delta S degree = 86.4 cal.degree-1.mol-1 for single-strand melting of S2 (1 cal = 4.184 J). These data reveal that the two single-stranded structures S1 and S2 are only marginally stable at 25 degrees C, despite exhibiting rather substantial transition enthalpies. This behavior results from enthalpy and entropy contributions of similar magnitudes that compensate each other, thereby giving rise to relatively small free energies of stabilization for the single strands at 25 degrees C. By contrast, the S1.S2 duplex state is very stable at 25 degrees C since the favorable transition entropy associated with duplex disruption (325.4 cal.degree-1.mol-1) is more than compensated for by the extremely large duplex transition enthalpy (117.0 kcal/mol). We also measured directly an enthalpy change (delta H degree) of -56.4 kcal/mol for duplex formation at 25 degrees C using isothermal batch-mixing calorimetry. This duplex formation enthalpy of -56.4 kcal/mol at 25 degrees C is very different in magnitude from the duplex disruption enthalpy of 117.0 kcal/mol measured at 74 degrees C by DSC. Since the DSC measurement reveals the net transition heat capacity change to be close to zero, we interpret this large disparity between the enthalpies of duplex disruption and duplex formation as reflecting differences in the single-stranded structures at 25 degrees C (the initial states in the isothermal mixing experiment) and the single-stranded structures at approximately 80 degrees C (the final states in the DSC experiment). In fact, the enthalpy for duplex formation at 25 degree C (-56.4 kcal/mol) can be combined with the sum of the integral enthalpies requires to melt each single strand from 25 to 80 degree C (23.6 kcal/mol for S1 and 27.2 kcal/mol for S2) to calculate a delta H degree of -107.2 kcal/mol for the hypothetical process of duplex formation from "random-coil" "unstacked" single strands at 25 degree C. The magnitude of this predicted delta H degree value for duplex formation is in good agreement with the corresponding parameter we measure directly by DSC for duplex disruption (117.0 kcal/mol), thereby lending credence to our interpretation and analysis of the data. Thus, our results demonstrate that despite being only marginally stable at 25 degree C, single strands can exhibit intramolecular interactions that enthalpically poise them for duplex formation. For the duplex studied herein, prior to association at 25 degree C, the two complementary single strands already possess > 40% of the total enthalpy (50.8/117) that ultimately stabilizes the final duplex state. This feature of single-stranded structure near room temperature can reduce significantly the enthalpic driving force one might predict for duplex formation from nearest-neighbor data, since such data generally are derived from measurements in which the single strands are in their random-coil states. Consequently, potential contributions from single-stranded structure must be recognized and accounted for when designing hybridization experiments and when using isothermal titration and/or batch mixing techniques to study the formation of duplexes and higher-order DNA structures (e.g., triplexes, tetraplexes, etc.) from their component single strands.
Full text
PDF


Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Appleby D. W., Kallenbach N. R. Theory of oligonucleotide stabilization. I. The effect of single-strand stacking. Biopolymers. 1973;12(9):2093–2120. doi: 10.1002/bip.1973.360120915. [DOI] [PubMed] [Google Scholar]
- Applequist J., Damie V. Thermodynamics of the one-stranded helix-coil equilibrium in polyadenylic acid. J Am Chem Soc. 1966 Sep 5;88(17):3895–3900. doi: 10.1021/ja00969a001. [DOI] [PubMed] [Google Scholar]
- Brahms J., Michelson A. M., Van Holde K. E. Adenylate oligomers in single- and double-strand conformation. J Mol Biol. 1966 Feb;15(2):467–488. doi: 10.1016/s0022-2836(66)80122-5. [DOI] [PubMed] [Google Scholar]
- Breslauer K. J., Frank R., Blöcker H., Marky L. A. Predicting DNA duplex stability from the base sequence. Proc Natl Acad Sci U S A. 1986 Jun;83(11):3746–3750. doi: 10.1073/pnas.83.11.3746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Breslauer K. J., Sturtevant J. M. A calorimetric investigation of single stranded base stacking in the ribo-oligonucleotide A7. Biophys Chem. 1977 Nov;7(3):205–209. doi: 10.1016/0301-4622(77)87023-3. [DOI] [PubMed] [Google Scholar]
- Breslauer K. J., Sturtevant J. M., Tinoco I., Jr Calorimetric and spectroscopic investigation of the helix-to-coil transition of a ribo-oligonucleotide: rA7U7. J Mol Biol. 1975 Dec 25;99(4):549–565. doi: 10.1016/s0022-2836(75)80171-9. [DOI] [PubMed] [Google Scholar]
- Davis R. C., Tinoco I., Jr Temperature-dependent properties of dinucleoside phosphates. Biopolymers. 1968;6(2):223–242. doi: 10.1002/bip.1968.360060206. [DOI] [PubMed] [Google Scholar]
- Erie D. A., Jones R. A., Olson W. K., Sinha N. K., Breslauer K. J. Melting behavior of a covalently closed, single-stranded, circular DNA. Biochemistry. 1989 Jan 10;28(1):268–273. doi: 10.1021/bi00427a037. [DOI] [PubMed] [Google Scholar]
- Erie D., Sinha N., Olson W., Jones R., Breslauer K. A dumbbell-shaped, double-hairpin structure of DNA: a thermodynamic investigation. Biochemistry. 1987 Nov 3;26(22):7150–7159. doi: 10.1021/bi00396a042. [DOI] [PubMed] [Google Scholar]
- Filimonov V. V., Privalov P. L. Thermodynamics of base interaction in (A)n and (A.U)n. J Mol Biol. 1978 Jul 15;122(4):465–470. doi: 10.1016/0022-2836(78)90422-9. [DOI] [PubMed] [Google Scholar]
- François J. C., Saison-Behmoaras T., Barbier C., Chassignol M., Thuong N. T., Hélène C. Sequence-specific recognition and cleavage of duplex DNA via triple-helix formation by oligonucleotides covalently linked to a phenanthroline-copper chelate. Proc Natl Acad Sci U S A. 1989 Dec;86(24):9702–9706. doi: 10.1073/pnas.86.24.9702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Freier S. M., Hill K. O., Dewey T. G., Marky L. A., Breslauer K. J., Turner D. H. Solvent effects on the kinetics and thermodynamics of stacking in poly(cytidylic acid). Biochemistry. 1981 Mar 17;20(6):1419–1426. doi: 10.1021/bi00509a003. [DOI] [PubMed] [Google Scholar]
- Gaffney B. L., Jones R. A. Thermodynamic comparison of the base pairs formed by the carcinogenic lesion O6-methylguanine with reference both to Watson-Crick pairs and to mismatched pairs. Biochemistry. 1989 Jul 11;28(14):5881–5889. doi: 10.1021/bi00440a026. [DOI] [PubMed] [Google Scholar]
- Hanvey J. C., Shimizu M., Wells R. D. Site-specific inhibition of EcoRI restriction/modification enzymes by a DNA triple helix. Nucleic Acids Res. 1990 Jan 11;18(1):157–161. doi: 10.1093/nar/18.1.157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Häner R., Dervan P. B. Single-strand DNA triple-helix formation. Biochemistry. 1990 Oct 23;29(42):9761–9765. doi: 10.1021/bi00494a001. [DOI] [PubMed] [Google Scholar]
- Jin R. Z., Breslauer K. J., Jones R. A., Gaffney B. L. Tetraplex formation of a guanine-containing nonameric DNA fragment. Science. 1990 Oct 26;250(4980):543–546. doi: 10.1126/science.2237404. [DOI] [PubMed] [Google Scholar]
- Leng M., Felsenfeld G. A study of polyadenylic acid at neutral pH. J Mol Biol. 1966 Feb;15(2):455–466. doi: 10.1016/s0022-2836(66)80121-3. [DOI] [PubMed] [Google Scholar]
- Maher L. J., 3rd, Wold B., Dervan P. B. Inhibition of DNA binding proteins by oligonucleotide-directed triple helix formation. Science. 1989 Aug 18;245(4919):725–730. doi: 10.1126/science.2549631. [DOI] [PubMed] [Google Scholar]
- Marky L. A., Breslauer K. J. Calculating thermodynamic data for transitions of any molecularity from equilibrium melting curves. Biopolymers. 1987 Sep;26(9):1601–1620. doi: 10.1002/bip.360260911. [DOI] [PubMed] [Google Scholar]
- Marky L. A., Curry J., Breslauer K. J. Netropsin binding to polyd(AT) X polyd(AT) and to polydA X polydT: a comparative thermodynamic study. Prog Clin Biol Res. 1985;172B:155–173. [PubMed] [Google Scholar]
- Marky L. A., Snyder J. G., Remeta D. P., Breslauer K. J. Thermodynamics of drug-DNA interactions. J Biomol Struct Dyn. 1983 Oct;1(2):487–507. doi: 10.1080/07391102.1983.10507457. [DOI] [PubMed] [Google Scholar]
- Moser H. E., Dervan P. B. Sequence-specific cleavage of double helical DNA by triple helix formation. Science. 1987 Oct 30;238(4827):645–650. doi: 10.1126/science.3118463. [DOI] [PubMed] [Google Scholar]
- Mudd C., Berger R. L., Hopkins H. P., Friauf W. S., Gibson C. An optimized differential heat conduction solution microcalorimeter for thermal kinetic measurements. J Biochem Biophys Methods. 1982 Aug;6(3):179–203. doi: 10.1016/0165-022x(82)90042-2. [DOI] [PubMed] [Google Scholar]
- Plum G. E., Park Y. W., Singleton S. F., Dervan P. B., Breslauer K. J. Thermodynamic characterization of the stability and the melting behavior of a DNA triplex: a spectroscopic and calorimetric study. Proc Natl Acad Sci U S A. 1990 Dec;87(23):9436–9440. doi: 10.1073/pnas.87.23.9436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poland D., Vournakis J. N., Scheraga H. A. Cooperative interactions in single-strand oligomers of adenylic acid. Biopolymers. 1966;4(2):223–235. doi: 10.1002/bip.1966.360040209. [DOI] [PubMed] [Google Scholar]
- Powell J. T., Richards E. G., Gratzer W. B. The nature of stacking equilibria in polynucleotides. Biopolymers. 1972 Jan;11(1):235–250. doi: 10.1002/bip.1972.360110118. [DOI] [PubMed] [Google Scholar]
- Praseuth D., Perrouault L., Le Doan T., Chassignol M., Thuong N., Hélène C. Sequence-specific binding and photocrosslinking of alpha and beta oligodeoxynucleotides to the major groove of DNA via triple-helix formation. Proc Natl Acad Sci U S A. 1988 Mar;85(5):1349–1353. doi: 10.1073/pnas.85.5.1349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Privalov P. L., Potekhin S. A. Scanning microcalorimetry in studying temperature-induced changes in proteins. Methods Enzymol. 1986;131:4–51. doi: 10.1016/0076-6879(86)31033-4. [DOI] [PubMed] [Google Scholar]
- Puglisi J. D., Wyatt J. R., Tinoco I., Jr A pseudoknotted RNA oligonucleotide. Nature. 1988 Jan 21;331(6153):283–286. doi: 10.1038/331283a0. [DOI] [PubMed] [Google Scholar]
- Sklenár V., Feigon J. Formation of a stable triplex from a single DNA strand. Nature. 1990 Jun 28;345(6278):836–838. doi: 10.1038/345836a0. [DOI] [PubMed] [Google Scholar]
- Stannard B. S., Felsenfeld G. The conformation of polyriboadenylic acid at low temperature and neutral pH. A single-stranded rodlike structure. Biopolymers. 1975 Feb;14(2):299–307. doi: 10.1002/bip.1975.360140205. [DOI] [PubMed] [Google Scholar]
- Sun J. S., François J. C., Montenay-Garestier T., Saison-Behmoaras T., Roig V., Thuong N. T., Hélène C. Sequence-specific intercalating agents: intercalation at specific sequences on duplex DNA via major groove recognition by oligonucleotide-intercalator conjugates. Proc Natl Acad Sci U S A. 1989 Dec;86(23):9198–9202. doi: 10.1073/pnas.86.23.9198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suurkuusk J., Alvarez J., Freire E., Biltonen R. Calorimetric determination of the heat capacity changes associated with the conformational transitions of polyriboadenylic acid and polyribouridylic acid. Biopolymers. 1977 Dec;16(12):2641–2652. doi: 10.1002/bip.1977.360161206. [DOI] [PubMed] [Google Scholar]


