Abstract
Molecular pathways involved in dauer formation, an alternate larval stage that allows Caenorhabditis elegans to survive adverse environmental conditions during development, also modulate longevity and metabolism. The decision to proceed with reproductive development or undergo diapause depends on food abundance, population density, and temperature. In recent years, the chemical identities of pheromone signals that modulate dauer entry have been characterized. However, signals derived from bacteria, the major source of nutrients for C. elegans, remain poorly characterized. To systematically identify bacterial components that influence dauer formation and aging in C. elegans, we utilized the individual gene deletion mutants in E. coli (K12). We identified 56 diverse E. coli deletion mutants that enhance dauer formation in an insulin-like receptor mutant (daf-2) background. We describe the mechanism of action of a bacterial mutant cyaA, that is defective in the production of cyclic AMP, which extends lifespan and enhances dauer formation through the modulation of TGF-β (daf-7) signaling in C. elegans. Our results demonstrate the importance of bacterial components in influencing developmental decisions and lifespan in C. elegans. Furthermore, we demonstrate that C. elegans is a useful model to study bacterial-host interactions.
The soil-dwelling nematode, C. elegans, assesses environmental cues including food availability, temperature, and population density1,2, to determine the choice between reproduction and survival2. Under favorable conditions, C. elegans attains reproductive maturity, but under adverse environments, worms enter dauer diapause, a long-lived stress resistant stage2. There are at least two types of chemical signals that dictate this developmental decision; a) dauer pheromones, a complex mixture of small molecules called ascarosides whose levels depend on worm population density3,4,5,6, and b) signals derived from bacteria that promote development3.
To detect microbial metabolites in the environment C. elegans has evolved multiple mechanisms, which allow discrimination between bacterial species that are appropriate food sources and those that are pathogenic or inferior in quality7,8. The chemosensory system senses bacterial metabolites in the local environment and modulates the organismal response to signaling through conserved molecular pathways1,2,9. These include the transforming growth factor beta (TGF-β), guanylyl cyclase, and insulin-like signaling (ILS) pathways, which ultimately converge to modulate the activity of key factors such as forkhead transcription factor (DAF-16/FOXO)2,3,4,10,11,12. The activity of DAF-16 is pivotal in determining the organism’s response, to either engage in reproductive growth or to enter dauer arrest state2,13. Although the genetic pathways that modulate dauer formation are relatively well understood, the identities of the signals that influence dauer formation remain poorly defined. C. elegans is known to respond to signals emanating from bacteria to detect food sources8, and a recent report identified bacterial fatty acids as a signal that modulates recovery from the dauer stage14. It is likely that the worm integrates a multitude of bacterial signals through the chemosensory system to make appropriate developmental decisions which remain undiscovered.
Dauer formation provides an easy to assess, binary readout of the type of food signals emanating from bacteria. Here we demonstrate how multiple individual gene knockouts in E. coli can influence development and survival in C. elegans. Through a systematic screen of ~4000 single gene knockout E. coli strains, we identified 56 E. coli mutants that enhanced dauer formation in a C. elegans insulin-like receptor mutant, daf-2, some of which also extended adult lifespan in control worms. One of the largest lifespan extensions was observed by feeding the bacterial mutant, cyaA (adenylate cyclase). We identify that cyaA influences TGF-β signaling to modulate dauer formation and lifespan. Our results demonstrate that the combination of bacterial and worm genetics can be a powerful tool to study bacterial-host interactions that influence nutrient signaling pathways and organismal physiology that may be relevant for host-microbiome interactions in other species.
Results
Bacterial mutants enhance dauer formation in daf-2 (e1370) mutants
To systematically study bacterial products that could act as food signals in C. elegans, we fed worms with bacterial strains from the E. coli Keio knockout library. The library consists of 3985 single-gene in-frame knockout mutants, with individual genes precisely excised and replaced with a Kanamycin resistance cassette in a K-12 BW25113 background15. To study the impact of bacteria on organismal physiology we screened for bacterial mutants that enhance dauer formation in a daf-2(e1370) mutant background. This worm strain carries a hypomorphic mutation in the insulin receptor ortholog gene that results in reduced flux through the ILS pathway and triggers constitutive dauer formation at high temperatures but undergoes normal development at low temperatures12,16,17,18,19. We carried out the screen at a semi-permissive temperature (21.5 °C), which allowed most worms on the control K-12 E. coli strain to develop to adulthood, with a small percentage entering the dauer stage. This assay created a sensitive readout to detect bacterial food signals in the diet, which influence organismal physiology detected by an increase in dauer formation in the worm.
In the primary screen, approximately 60 eggs were seeded onto plates with mutant E. coli strains and monitored for dauer formation after 4–5 days. The total number of dauers on the K-12 control strain was used as a reference and knockout strains that produced at least four more dauer animals than the control strain were marked as positive candidates and retested. This second, and more stringent, round of screening examined 150 knockout candidates in triplicate, and knockouts that at least doubled the total dauer percentage of worms developed on the K-12 control strain were scored as positive (Fig. 1a). Using this approach, we identified 56 mutant E. coli strains that robustly enhanced entry into the dauer stage (Table 1). PCR with gene-specific primers was performed to confirm gene deletion in the knockout mutants (Fig. S1A). As negative controls, ten knockout strains that were not identified in the primary screen were randomly chosen and thoroughly tested for dauer formation. These strains did not show any significant alteration in the rate of dauer formation when compared to controls (Fig. S1B).
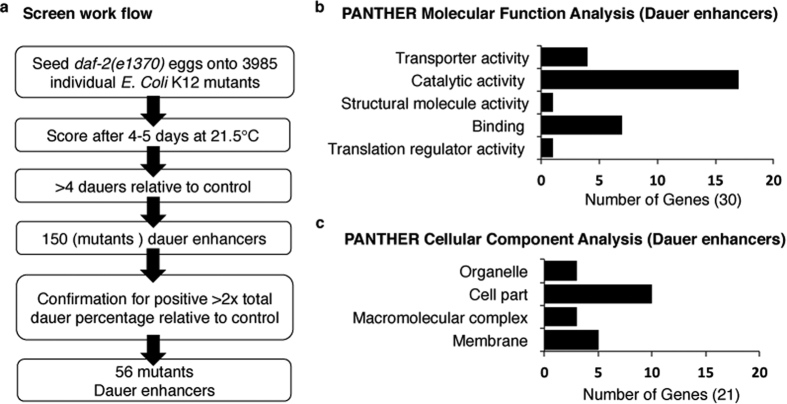
Figure 1. Schematic of the E. coli knockout screen for dauer formation in C. elegans and functional classifications of identified bacterial genes.
(a) Schematic of the workflow of E. coli knockout screen for dauer enhancement in C. elegans. The screen resulted in 56 mutant bacterial strains enhancing dauer formation in the worm. The knockout candidates and associated statistical information are available on Table 1. (b) Functional classification of candidate genes (dauer enhancers) from the screen. The molecular function analysis of the candidate genes was done using PANTHER pathway database. (c) The cellular component analysis of these genes was done using PANTHER pathway database. For details pertaining to biological function analysis and enrichment analysis see Fig. S1.
Table 1. List of bacterial mutants that enhance dauer formation.
| Gene Knockout | Brief Description | Func. Group | Mean(%)a | ±SEb | nc | p-Valued |
|---|---|---|---|---|---|---|
| K-12 | Control | / | 12.6 | 1.5 | 231 | / |
| moaD | molybdopterin synthase, small subunit | MET | 81.9 | 3.2 | 117 | <0.0001 |
| JW09292 | zapC: cell division factor, localises to the cytokinetics ring | OTR | 79.0 | 2.6 | 309 | <0.0001 |
| fdoG | α subunit of formate dehydrogenase O | MET | 77.7 | 0.9 | 63 | <0.0001 |
| JW1330 | abgT; predicted cryptic aminobenzoyl-glutamate transporter | MET | 72.1 | 3.5 | 222 | <0.0001 |
| Lar | Rac prophae: restriction alleviation protein | OTR | 70.2 | 2.0 | 145 | <0.0001 |
| panE | 2-dehydropantoate reductase, NADPH-specific | MET | 68.6 | 3.8 | 98 | <0.0001 |
| mdoH | glucan biosynthesis: glycosyl transferase | CEB | 67.5 | 7.4 | 96.0 | 0.00 |
| yjgB | predicted alcohol dehydrogenase, Zn-dependent and NAD(P)-binding | UNK | 67.5 | 3.3 | 180 | <0.0001 |
| elaA | predicted acyltransferase, Zn- dependent and NAD(P)-binding | UNK | 66.7 | 3.2 | 207 | <0.0001 |
| kdpB | potassium translocating ATPase, subunit B | ITM | 66.4 | 17.5 | 103 | 0.04 |
| dcuC | anaerobic C4-dicarboxylate transport | MET | 65.2 | 1.6 | 186 | <0.0001 |
| mutS | methyl-directed mismatch repair protein | DNA | 64.4 | 3.6 | 209 | <0.0001 |
| htrG | predicted signal transduction protein (SH3 domain) | UNK | 64.3 | 5.4 | 194 | <0.0001 |
| dhaH | fused predicted dihydroxyacetone-specific PTS enzymes: HPr component | MET | 63.4 | 11.4 | 63 | 0.01 |
| rimK | ribosomal protein S6 modification protein | TRN | 62.8 | 3.3 | 72 | <0.0001 |
| gloB | predicted hydroxyacylglutathione hydrolase | MET | 62.6 | 8.6 | 95.0 | 0.00 |
| clpX | ATPase and specificity subunit of ClpX-ClcP ATP-dependent serine protease | TRN | 62.4 | 2.3 | 175 | <0.0001 |
| tufa | protein chain elongation factor EF-Tu (duplicate of tuFB) | TRN | 60.7 | 3.0 | 69 | <0.0001 |
| ybeA | 23S rRNA mψ1915 methyltranferase | OTR | 60.2 | 7.6 | 250 | 0.00 |
| JW0632 | ybeB; predicted protein | UNK | 58.7 | 6.4 | 190 | 0.00 |
| rfaD | ADP-L-glycero-D-mannoheptose-6-epimerase | CEB | 58.3 | 5.0 | 146 | <0.0001 |
| yjiK | conserved protein | UNK | 56.4 | 5.4 | 151 | 0.00 |
| fdoH | formate dehydrogenase-O, Fe-S subunit | MET | 56.0 | 10.7 | 65 | 0.02 |
| yncA | predicted acyltransferase with acyl-CoA N-acyltransferase domain | CEB | 55.8 | 9.5 | 135 | 0.01 |
| fepB | iron-enterbactin transporter subunit | ITM | 53.2 | 10.3 | 309 | 0.01 |
| hsdS | specificity detereminant for hsdM and hsdR | DNA | 52.3 | 6.6 | 188 | 0.00 |
| ygiQ | conserved protein | UNK | 51.0 | 9.2 | 63 | 0.01 |
| Ung | iron-enterbactin transporter subunit | DNA | 50.6 | 11.6 | 70 | 0.03 |
| hdfR | DNA-binding transcriptional regulator | TRX | 50.4 | 16.3 | 200 | 0.08 |
| yjbD | conserved protein | UNK | 49.2 | 4.0 | 180 | 0.00 |
| fepC | iron-enterbactin transporter subunit | ITM | 47.8 | 4.9 | 142 | 0.00 |
| hypE | carbomoyl phosphate phosphatase, hydrogenase 3 maturation protein | TRN | 45.2 | 7.4 | 105 | 0.01 |
| JW5726 | phnl, phnH; carbon-phosphorus lyase complex subunit | ITM | 43.5 | 4.2 | 112 | 0.00 |
| ydhF | predicted oxidoreductase | UNK | 43.1 | 6.0 | 145 | 0.01 |
| yqeH | conserved protein with bipartite regulator domain | TRX | 42.1 | 4.4 | 117 | 0.00 |
| yjgN | conserved inner membrane protein | UNK | 41.4 | 4.8 | 97 | 0.00 |
| JW1321.5 | Unknown | UNK | 40.0 | 5.7 | 236 | 0.01 |
| cyaA | adenylate cyclase | OTR | 39.6 | 7.5 | 66.0 | 0.02 |
| moaE | molybdopterin synthase, small subunit | MET | 38.3 | 7.7 | 82 | 0.03 |
| sufA | Fe-S cluster assembly protein | MET | 37.6 | 5.1 | 229 | 0.01 |
| sufE | sulphur acceptor protein | MET | 36.6 | 5.1 | 211 | 0.01 |
| yggX | protein that protects iron-sulfur proteins against oxidative damage | TRN | 36.1 | 8.1 | 352 | 0.05 |
| yheM | predicted intracellular sulfur oxidation protein | ITM | 36.0 | 2.8 | 208 | 0.00 |
| yedR | predicted inner membrane protein | UNK | 33.5 | 8.1 | 294 | 0.07 |
| fepG | iron-enterbactin transporter subunit | ITM | 32.3 | 6.8 | 195 | 0.03 |
| yicM | purine ribonucleoside exporter | MET | 32.2 | 9.0 | 187 | 0.10 |
| dmsC | dimethyl sulfoxide reductase, anaerobic, subunit C | MET | 32.1 | 1.8 | 111 | 0.00 |
| yjgX | KpLE2 phage-like element; predicted protein (pseudogene) | UNK | 31.8 | 4.8 | 181 | 0.02 |
| yrfD | predicted pilus assembly protein | MET | 31.6 | 4.9 | 195 | 0.02 |
| srlA | glucitol/sorbitol -specific enzyme IIC component of PTS | MET | 30.6 | 4.0 | 323 | 0.01 |
| pepA | aminopeptidase A, a cyteinylglycinase | MET | 29.8 | 2.6 | 216 | 0.00 |
| yfaS | predicted protein (pseudogene) | UNK | 29.7 | 4.5 | 223 | <0.0001 |
| Yjhc | KpLE2 phage-like element; predicted oxidoreducatse | UNK | 29.7 | 1.6 | 247.0 | <0.001 |
| tolA | membrane anchored protein involved in colicin uptake | CEB | 29.2 | 3.8 | 186 | 0.02 |
| ygjQ | predicted thioredoxin-like | UNK | 28.6 | 1.7 | 252 | 0.00 |
| yqjH | predicted siderophore interacting protein | ITM | 28.3 | 1.0 | 367 | <0.0001 |
E. coli knockouts were identified as portrayed in Fig. 1a and functional groups were delineated as described in text and Fig. 1b. Brief descriptions were adapted from EcoCyc. Functional codes were abbreviated as follows: MET, metabolism; ITM, inorganic ion transport and metabolism; TRN, translation, posttranslational modification, protein turnover, chaperones; CEB, cell envelope biogenesis, outer membrane; DNA, DNA replication, recombination and repair; TRX, transcription; OTR, other; UNK, unknown.
aThe mean percentage of dauer staged animals.
bThe standard error of the mean in percent.
cThe total number of individuals scored (includes all replicates).
dThe p-value for a student’s two-tailed t-test comparing respective E. coli gene knockout strain to control strain.
To identify the biological pathways and cellular functions affected by the bacterial mutants that influenced dauer formation, the candidates were analyzed using PANTHER classification system20,21. Out of 56 genes, 9 genes are not characterized, and the remaining 47 genes were analyzed for molecular and biological functions (Fig. 1b and Fig. S1C). The following broad molecular functional categories were identified: translation regulator activity (GO:0045182) (4 genes). binding (GO:0005488) (7 genes), structural molecule activity (GO:0005198) (1 gene), catalytic activity (GO:0003824) (17 genes), transporter activity (GO:0005215) (4 genes) (Fig. 1b). Using the PANTHER classification system the cellular site of action of the product of these genes was identified:membrane (GO:0016020), macromolecular complex (GO:0032991), cell part (GO:0044464) and organelle (GO:0043226) (Fig. 1c). Furthermore, the candidate genes were analyzed and assigned functional groups utilizing Clusters of Orthologous Group (COG) terms22 and confirmed with MultiFun23 and Gene Ontology (GO) classifications24. Categorization of the genes revealed the enrichment in the following functional groups: metabolism (~39%); inorganic ion transport and metabolism (~17%); translation, post-translational modification, protein turnover (~12%), and chaperones; cell envelope biogenesis and outer membrane structures (~10%); DNA replication, recombination and repair (~7%); transcription (~5%); and rest other functions (~10%) (Table 1 and Fig. S1D). Furthermore, several of the genes from the screen fell within functional groups and transcriptional units, e.g. operons and regulons (Fig. S1E). Together these findings indicate that the bacterial genes with a broad range of cellular functions and processes can alter C. elegans physiology through changes in bacteria derived metabolites or nutrients.
Phenotypic analyses of bacterial mutants that enhance dauer formation for DAF-16 activation and lifespan extension
The phosphorylation cascade elicited by an activated ILS negatively regulates DAF-16 by sequestering it in the cytoplasm. Conversely, reduced ILS signaling leads to diminished phosphorylation of DAF-16, thereby allowing it to enter the nucleus and modify gene expression25,26. Given the effects of the bacterial mutants on dauer formation in the daf-2 mutant, we examined whether they would also impact DAF-16 in adult animals. A majority (∼63%) of the bacterial mutants promoted DAF-16 nuclear localization in a worm strain expressing a DAF-16::GFP (daf-16(mgDf47)) fusion construct (Fig. 2 and Fig. S2A). Once DAF-16 enters the nucleus it can modulate the transcription of its targets, such as the stress-responsive superoxide dismutase, SOD-327. Consistently, ~64% of the bacterial mutants also increased SOD-3p::GFP expression in an N2 control background, while ~80% of the bacterial knockouts increased sod-3 expression in the daf-2(e1370) background (Fig. 2 and Fig. S2A). No significant effects on sod-3 expression were noted in the 10 negative controls used previously, compared to animals fed control bacteria (data not shown). Lastly, of the 35 bacterial candidates that showed DAF-16 nuclear localization, 26 induced sod-3p::GFP in N2 background, while 32 induced GFP in the daf-2;sod-3p::GFP strain (Fig. 2). Taken together these data suggest that the majority of E. coli mutants from the screen stimulate DAF-16 translocation and/or subsequent regulation of its transcriptional targets in both the daf-2 mutant and control (N2) worms.
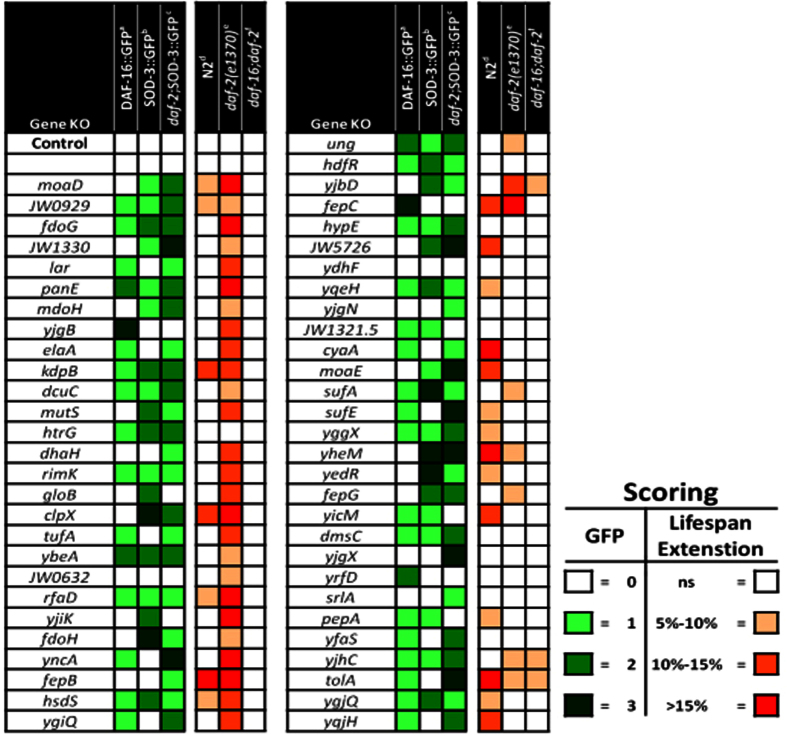
Figure 2. Analysis of DAF-16 activation and lifespan extension in bacterial knockouts that enhance C. elegans dauer formation.
Heat map showing various phenotypes observed with bacterial mutants (left to right). (a) Semi-quantitative analysis of DAF-16 nuclear localization is shown following feeding on listed E. coli knockouts. Increased green shading correlates with increasing nuclear localization of DAF-16. (Fig. S2) (b) sod-3p::GFP expression is shown following feeding on listed E. coli knockouts. Increased green shading correlates with increasing sod-3p::GFP expression. (c) daf-2(e1370); sod-3p::GFP expression is shown following feeding on listed E. coli knockouts. Increased green shading correlates with increasing daf-2(e1370);sod-3p::GFP expression. In each visual marker (a–c) knockout strains are given a score relative to its K-12 control, which was consistently set at a score of 0 (white shading). (d–f) Lifespan analysis of respective worm strains following feeding on bacterial knockout strains throughout life. Deeper red shading indicates increased average lifespan compared to K-12 control strain. Representative lifespan curves and lifespan data analysis are shown in Fig. S2 and Table 2 respectively.
In addition to regulating dauer entry during development, DAF-16 exhibits transcriptional control over genes determining longevity in the adult worm27,28. Thus, we assessed the effect of each bacterial strain identified in the screen on the survival of N2, daf-2, and daf-16; daf-2 strains. Feeding several of the E. coli knockout strains resulted in increased lifespan in both the daf-2 and control (N2) background. However, ~61% of the bacterial mutants significantly extended the average lifespan of daf-2 mutants compared to ~38% in the control N2 strain. (Fig. 2, Table 2 and Fig. S2B). These observations were similar to the results obtained with the sod-3::gfp expression (Fig. 2), which can potentially be explained as the screen was carried out in a sensitized daf-2 mutant background. Although most of the knockout bacteria appear to modulate lifespan in a daf-16-dependent manner (Fig. 2 and Table 2), some bacterial mutants (e.g. tolA, and yjbD) extended lifespan in daf-16;daf-2 mutant worms, indicating that a genetically modified diet may also affect longevity by daf-16-independent mechanisms.
Table 2. Lifespans of C. elegans strains fed on different bacterial mutants.
| Gene Knockout | N2 |
daf-2(e1370) |
daf-16(mu86); daf-2(e1370) |
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Meana | Changeb | nc | p-Valued | Meana | changeb | nc | p-Valued | Meana | Changeb | nc | p-Valued | |
| K-12 | 13.9 ± 0.4 | / | 248 | — | 26.4 ± 0.9 | / | 278 | — | 12.7 ± 0.3 | / | 264 | |
| moaD | 15.1 | 8.7% | 11 | 0.0076 | 31.3 | 18.5% | 106 | <0.0001 | 13.2 | / | 96 | |
| JW09292 | 14.6 | 5.2% | 131 | <0.0001 | 28.6 | 8.2% | 116 | <0.0001 | 13.1 | / | 116 | |
| fdoG | 12.9 | / | 93 | 30.4 | 15.1% | 107 | <0.0001 | 13 | / | 91 | ||
| JW1330 | 14.5 | / | 124 | 28.5 | 7.9% | 109 | 0.0315 | 12.6 | / | 73 | ||
| Lar | 13.5 | / | 99 | 29.4 | 11.5% | 109 | <0.0001 | 12.6 | / | 125 | ||
| panE | 14.3 | / | 96 | 30.6 | 15.9% | 105 | <0.0001 | 10.7 | / | 140 | ||
| mdoH | 14.1 | / | 109 | 28.2 | 6.7% | 113 | 0.029 | 11.8 | / | 146 | ||
| yjgB | 13.1 | / | 131 | 28.2 | 11.3% | 109 | <0.0001 | 12.2 | / | 137 | ||
| elaA | 12.7 | / | 83 | 30.1 | 14% | 105 | <0.0001 | 13.2 | / | 100 | ||
| kdpB | 15.6 | 11.9% | 140 | 0.0375 | 29.4 | 11.4% | 106 | <0.0001 | 12.6 | / | 99 | |
| dcuC | 13.6 | / | 94 | 28.7 | 8.8% | 118 | 0.0013 | 12.7 | / | 112 | ||
| mutS | 11.3 | / | 69 | 29.5 | 11.7% | 104 | <0.0001 | 12.2 | / | 94 | ||
| htrG | 13 | / | 112 | 27.6 | / | 108 | 12.4 | / | 140 | |||
| dhaH | 11.8 | / | 92 | 29.4 | 11.3% | 111 | <0.0001 | 12.5 | / | 99 | ||
| rimK | 12.7 | / | 117 | 30.2 | 14.4% | 108 | <0.0001 | 13.2 | / | 127 | ||
| gloB | 13.2 | / | 112 | 29.2 | 10.8% | 109 | <0.001 | 12.4 | / | 94 | ||
| clpX | 15.9 | 14.4% | 96 | <0.0001 | 32.5 | 23% | 109 | <0.0001 | 12.6 | / | 79 | |
| tufA | 12.7 | / | 123 | 29.7 | 12.4% | 107 | <0.0001 | 12.6 | / | 87 | ||
| ybeA | 13.4 | / | 126 | 28.1 | 6.6% | 110 | 0.02 | 13.2 | / | 82 | ||
| JW0632 | 13.4 | / | 119 | 28.3 | 7.3% | 108 | 0.018 | 13.9 | / | 129 | ||
| rfaD | 14.8 | 6.2% | 128 | 0.45 | 30.9 | 16.9% | 106 | <0.0001 | 13.2 | / | 127 | |
| yjiK | 13.1 | / | 123 | 30.8 | 16.8% | 107 | <0.0001 | 12.3 | / | 105 | ||
| fdoH | 12.7 | / | 84 | 28.6 | 8.3% | 110 | <0.0001 | 12.1 | / | 103 | ||
| yncA | 12.6 | / | 83 | 31.2 | 18.2% | 106 | <0.0001 | 12.4 | / | 89 | ||
| fepB | 16.7 | 19.8% | 89 | <0.0001 | 32 | 21.3% | 90 | <0.0001 | 12.3 | / | 146 | |
| hsdS | 14.9 | 7.3% | 112 | 0.0172 | 29.3 | 11% | 108 | <0.0001 | 13.4 | / | 128 | |
| ygiQ | 13.8 | / | 108 | 29.2 | 10.7% | 108 | <0.0001 | 12.8 | / | 56 | ||
| ung | 12.2 | / | 77 | 29 | 9.7% | 110 | <0.0001 | 12 | / | 92 | ||
| hdfR | 14.7 | / | 141 | 27.6 | / | 112 | 12.8 | / | 135 | |||
| yjbD | 13 | / | 99 | 29.4 | 11.3% | 112 | <0.0001 | 13.8 | 8.20% | 124 | <0.0001 | |
| fepC | 15.6 | 12.4% | 150 | <0.0001 | 31.5 | 19.3% | 106 | <0.0001 | 12.9 | / | 126 | |
| hypE | 14.4 | / | 135 | 26.8 | / | 98 | 12.7 | / | 145 | |||
| JW5726 | 15.3 | 10.4% | 115 | 0.0125 | 26.9 | / | 103 | 12.3 | / | 102 | ||
| ydhF | 14.3 | / | 205 | 26.8 | / | 107 | 12.2 | / | 113 | |||
| yqeH | 15.1 | 8.3% | 141 | 0.0266 | 25.6 | / | 98 | 12.4 | / | 139 | ||
| yjgN | 14.3 | / | 101 | 26.4 | / | 113 | 12.2 | / | 150 | |||
| JW1321.5 | 13.4 | / | 135 | 25.7 | / | 106 | 12.3 | / | 142 | |||
| cyaA | 18.9 | 36.3% | 135 | <0.001 | 23.4 | / | 53 | 13 | / | 144 | ||
| moaE | 15.5 | 12.2% | 116 | 0.0015 | 26 | / | 53 | 12.2 | / | 90 | ||
| sufA | 14.7 | / | 129 | 28.1 | 6.5% | 103 | <0.0001 | 12.5 | / | 105 | ||
| sufE | 15.2 | 9.4% | 133 | 0.02 | 27.4 | / | 105 | 12.2 | / | 128 | ||
| yggX | 15.2 | 9.3% | 145 | 0.0137 | 26.3 | / | 107 | 12.6 | / | 146 | ||
| yheM | 16.6 | 19.3% | 141 | <0.0001 | 28.3 | 7.3% | 104 | <0.0001 | 11.8 | / | 105 | |
| yedR | 15.2 | 9.5% | 143 | 0.021 | 26.2 | / | 107 | 12.3 | / | 84 | ||
| fepG | 14.4 | / | 83 | 28.8 | 9% | 103 | <0.0001 | 11.8 | / | 139 | ||
| yicM | 15.9 | 14.2% | 146 | <0.0001 | 27.1 | / | 101 | 12.6 | / | 139 | ||
| dmsC | 15.8 | 19.2% | 129 | <0.0001 | 23.3 | 8.8% | 103 | <0.0001 | 11.89 | / | 102 | |
| yjgX | 14.8 | / | 179 | 25.6 | / | 111 | 12.3 | / | 144 | |||
| yrfD | 14.8 | / | 172 | 27.4 | / | 110 | 12.4 | / | 144 | |||
| srlA | 14.9 | / | 131 | 22.2 | / | 72 | 11.6 | / | 121 | |||
| pepA | 15.2 | 9.7% | 137 | 0.0119 | 25.7 | / | 110 | 12.1 | / | 113 | ||
| yfaS | 13.2 | / | 97 | 26.8 | / | 106 | 12.5 | / | 128 | |||
| yjhc | 14.4 | / | 98 | 28.2 | 6.9% | 108 | 0.0258 | 13.5 | 6.40% | 127 | 0.0039 | |
| tolA | 16.8 | 21.1% | 162 | <0.0001 | 27.7 | 5% | 107 | <0.0001 | 13.5 | 6.30% | 122 | <0.0001 |
| ygjQ | 15.1 | 8.7% | 143 | 0.035 | 26.3 | / | 107 | 12.7 | / | 127 | ||
| yqjH | 15.3 | 10% | 113 | 0.0074 | 26.6 | / | 107 | 12.5 | / | 124 | ||
Lifespan of C. elegans when feeding on bacterial knockouts: All experimental lifespan assays were performed in triplicate and survival data was grouped. Control lifespan assays performed at 2–3 different time points were done in triplicate and the survival curves were averaged and used to compare to experimental lifespan sets.
aThe mean lifespan in days.
bThe percent change of mean lifespan relative to K-12 control lifespan.
cThe total number of individuals scored.
dThe p-value of log-rank test comparing survival curves of gene knockout fed to control strain fed worm.
Several of the identified bacterial genes appear to have highly directed roles in micronutrient or metabolite production or transport (Table 1 and Fig. S1E). For example, fepB, C, and G have roles in the iron import system. dhaH has an enzymatic role in the production of dihydroxyacetone phosphate, while srlA, yjhc, and mdoH are involved in the production of sorbitol, N-acetylneuraminic acid (sialic acid), and periplasmic glucans, respectively. One of the largest lifespan extension was observed with cyaA, which encodes adenylate cyclase that catalyzes the conversion of ATP to Adenosine 3′,5′-cyclic monophosphate (cAMP) and pyrophosphate15. In bacteria, intracellular cAMP signaling modulates the expression of catabolic proteins with biosynthetic and ribosomal proteins according to cellular metabolic needs during exponential growth29. We further characterized cyaA mutant bacteria to determine the role of bacterial cAMP in modulating insulin signaling, dauer formation and lifespan in C. elegans.
cAMP modulates lifespan and dauer formation in C.elegans
We examined whether the effect of cyaA mutant bacteria on dauer formation was a consequence of reduced levels of cAMP. We confirmed that cyaA gene is disrupted (Fig. S1) and cAMP levels were indeed reduced in cyaA mutant bacteria (Fig. 3a). Next, we found that supplementation with 2 mM exogenous cAMP could restore levels back to those observed in K12 bacteria (Fig. 3a). Also, we did not see any significant difference in the bacterial growth curves between cyaA mutant bacteria as compared K-12 bacteria under minimal media conditions (Fig. S3A). The uptake of cAMP has been shown to be variable in bacteria and exhibits saturation kinetics based on the culture conditions30. Also, worms fed on cyaA mutant bacteria did not display any apparent adverse effects on worm physiology, as we did not observe any significant change in physiological parameters such as body size, lawn leaving behavior or pharyngeal pumping in control animals (N2) fed on cyaA mutant bacteria (Fig. S3B–G). Furthermore, bacterial choice assays suggested that the N2 worms preferred K-12 control bacteria to cyaA and other mutant bacteria that enhance dauer formation (Fig. S3H).
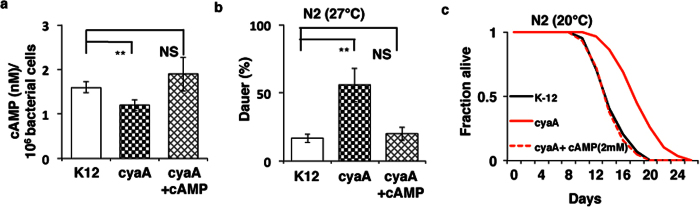
Figure 3. Feeding bacterial cyaA mutants modulates lifespan and dauer formation in C. elegans.
(a) The amount of cAMP in K-12 (control), mutant cyaA bacteria, cyaA mutant bacteria supplemented with cAMP (2 mM), (b) Bacterial knockout of cyaA enhances the dauer formation in N2 strain. Control (N2) animals fed on K-12 (control) bacteria, mutant cyaA bacteria and cyaA mutant bacteria supplemented with cAMP (2 mM) on minimal media agar plate. Data is represented as mean percent ± SD of greater than 3 biological replicates, n > 200. (c) Feeding E. coli cyaA mutants extends the lifespan of N2 strain in C. elegans. ***P < 0.0001.
Higher temperature has also been shown to enhance dauer formation31. We observed that at 27 °C, ~56% of N2 animals grown on cyaA bacteria formed dauers, compared with only ~20% dauers when grown on K12 bacteria (Fig. 3b, Table S1 and Fig. S4A). However, the dauer enhancement was rescued when cyaA bacteria were supplemented with 2 mM cAMP. Next, we performed a dauer assay with daf-2(m41) mutant worms that make almost 100% dauer at semi-permissive temperature. We observed a ~20% inhibition of dauer formation on the addition of cAMP (2 mM) in daf-2(m41) mutant worms (Fig. S4B). In lifespan experiments, N2 worms fed on cyaA mutant bacteria displayed a ~35% increase in lifespan. The addition of 2 mM cAMP to cyaA bacteria significantly suppressed the lifespan extension observed in the cyaA mutant bacteria alone (Fig. 3c, Table S2 and Fig. S4C). Thus both the dauer and lifespan effects of cyaA mutant bacteria could be rescued by exogenous addition of cAMP.
Bacterial cyaA mutant modulates dauer formation and lifespan through TGF-β signaling and DAF-16 activation
To investigate the signaling mechanism by which cyaA mutant bacteria modulates host C. elegans physiology to enhance dauer formation and extend lifespan, we examined the gene expression changes modulated by cyaA bacteria in control animals, using microarray profiling in adult animals fed cyaA bacteria and K12 bacteria (Fig. 4a and Table S3). Using GoMiner high-throughput analysis, we discovered multiple enriched biological processes within our data set (Fig. S5A). Of particular interest to us was the significant parent-child GO-term set of aging, multicellular organismal aging, and determination of adult lifespan. Comparison of changes in gene expression in the daf-2 background from previously published32 and our own data sets with transcriptionally altered genes in cyaA, showed considerable overlap (Fig. S5B,C). Collectively, these data suggest that feeding cyaA mutant E. coli activates DAF-16 to enhance dauer formation and extend lifespan.
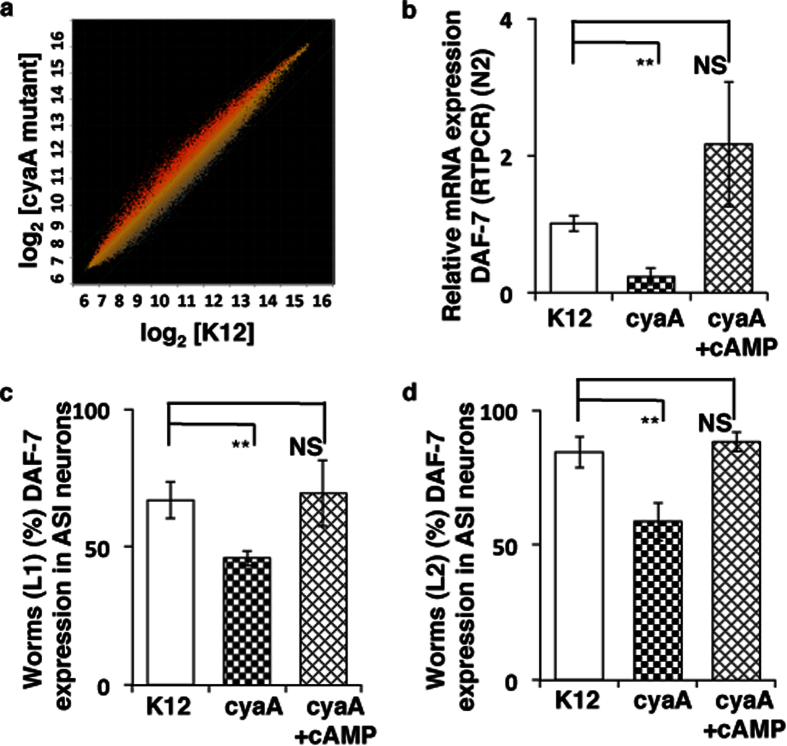
Figure 4. Feeding bacterial cyaA mutants alters DAF-7 expression in C. elegans.
(a) A scatter plot of log2 transcript probe intensities for daf-2(e1370) versus N2, and K-12 (control) versus cyaA fed control worm comparisons. Genes which show only p-values of <0.05 and fold changes >1.3 and <0.8 were shortlisted. (b) Relative expression of daf-7 mRNA in N2 animals on feeding K-12(control), cyaA mutant bacteria, and cyaA mutant bacteria supplemented with cAMP(2 mM). (c,d) cAMP induces the expression of DAF-7 in ASI neuron. The fraction of animals show DAF-7 expression at L1 and L2 stage on feeding K-12 (control), cyaA mutant bacteria, and cyaA mutant bacteria supplemented with cAMP (2 mM). The data is represented as percent mean ± S.D. of greater than 3 biological replicates, n > 200, **P < 0.0001.
We then examined the mechanism of activation of DAF-16 by measuring the expression of genes upstream of DAF-16 known to be involved in dauer formation. We observed that daf-7, a gene encoding a TGF-β-like ligand, was down-regulated in control worms grown on cyaA bacteria. To confirm this, we examined the expression of mRNA of daf-7 by RT-PCR in one-day adult control animals on cyaA mutant bacteria, compared with animals fed with K-12 or cyaA supplemented with cAMP (2 mM). DAF-7 expression was significantly reduced (~50%) on feeding animals with cyaA mutant bacteria and was rescued by addition of cAMP (2 mM) to the bacterial lawns (Fig. 4b). We also examined DAF-7::GFP expression in the ASI chemosensory neuron in L1 and L2 when fed with cyaA mutant bacteria. We observed that cyaA mutant bacteria inhibited the expression DAF-7::GFP in L1 and L2 stage in comparison to K-12 bacteria (Fig. 4c,d). Conversely, supplementation with 2 mM exogenous cAMP to cyaA mutant bacteria increased the expression of DAF-7 in a fraction of L1 and L2 animals (Fig. 4c,d). These results suggest that cAMP levels in the bacteria can modulate the expression of DAF-7 in worms, which influences dauer formation and insulin signaling33.
To further examine how cyaA mutant bacteria and cAMP modulate dauer formation we carried out epistasis analysis on C. elegans mutants using mutants in the three major pathways involved in dauer formation16,17,34. As shown in Fig. 3 dauer entry in daf-2(1370) mutants, was significantly increased when fed on cyaA mutant bacteria but was reversed by addition of exogenous cAMP. However, neither the daf-7(e1372) mutant, which is defective in TGF-β signaling nor the daf-11(m47) mutant, which has defective cyclic GMP signaling and is involved in DAF-7 secretion, showed any enhancement of dauer formation when fed on cyaA mutant bacteria (Fig. 5a–c and Table S1). This data supports the hypothesis that reduced bacterial cAMP enhances dauer formation by a similar mechanism as inhibition cGMP and TGF-β signaling pathways in worms.
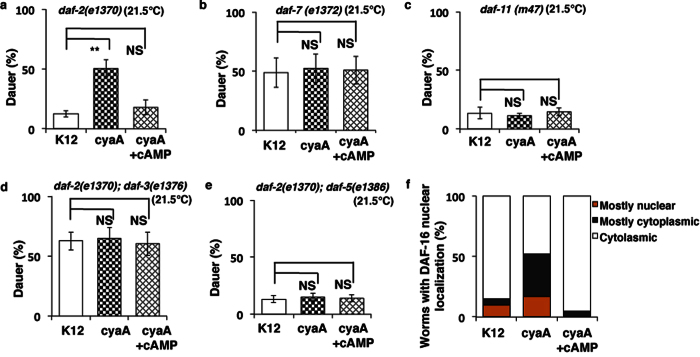
Figure 5. Bacterial cyaA mutants affect dauer formation through TGF-β pathway and DAF-16.
(a–e) Dauer formation of daf-2(e1370), daf-7(e1372), daf-11(m47) and daf-2(e1370); daf-3(e1376), daf-2(e1370); daf-5(e1386) animals on feeding K-12 (control), cyaA mutant bacteria, and cyaA mutant bacteria supplemented with cAMP (2 mM). (f) DAF-16::GFP worms in the N2 background on feeding K-12 (control), cyaA mutant bacteria, and cyaA mutant bacteria supplemented with cAMP(2 mM). Quantification of DAF-16::GFP localization on feeding K-12 (control), cyaA mutant bacteria, and cyaA mutant bacteria supplemented with cAMP(2 mM). In each case, the data is represented as mean percent ± S.D of three replicates, **P < 0.0001, n > 200.
The constitutive dauer formation phenotype of daf-7(e1372), but not daf-2(e1370), is suppressed by mutations in daf-7 downstream factors daf-3 and daf-52. The introduction of either daf-3(e1376) or daf-5(e1386) into the daf-2(e1370) background prevented cyaA mutant bacteria from enhancing dauer formation (Fig. 5d–e and Table S1). This suggests that cyaA mutant bacteria acts through TGF-β signaling to activate DAF-16, and enhancing dauer formation. We confirmed our finding from the screen that worms fed on cyaA mutant bacteria show increased nuclear localization of DAF-16 (Fig. 5f). Furthermore, not only was the nuclear localization of DAF-16::GFP enhanced on cyaA mutant bacteria compared with the K12 control, but it was also reversed upon supplementation of bacterial lawns with exogenous cAMP (Fig. 5f).
Conserved molecular pathways that regulate dauer formation like Insulin/IGF-1 signaling (IIS) and TGF-β signaling pathways also regulate longevity. This also appears to be the mechanism by which cyaA bacteria extend lifespan, since cyaA bacteria did not extend lifespan in either the daf-7(e1372) or the daf-11(m47) mutant backgrounds, nor did it extend lifespan in daf-2(e1370); daf-3(e1376) or daf-2(e1370); daf-5(e1386) double mutant backgrounds (Fig. 6a–f and Table S2). Together, these data indicates that reduced bacterial cAMP acts via the TGF-β signaling pathway to enhance DAF-16 resulting in increased dauer formation and lifespan.
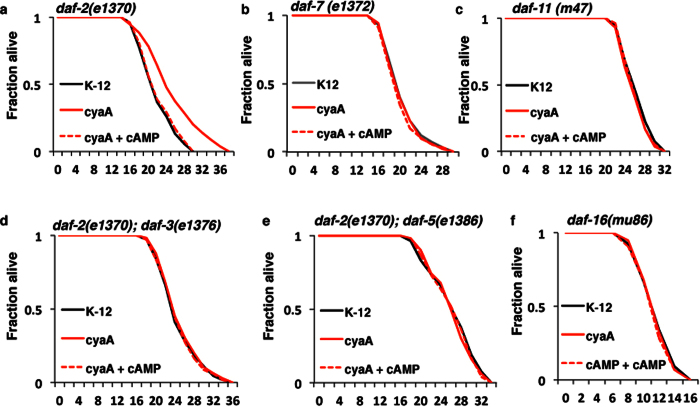
Figure 6. Bacterial cyaA mutants affect lifespan through TGF-β pathway and DAF-16.
(a–f) Kaplan–Meier survival curves of synchronously aging hermaphrodite worms: daf-2(e1370), daf-7(e1372), daf-11(m47), daf-2(e1370); daf-3(e1376), daf-2(e1370); daf-5(e1386) and daf-16(mu86) animals fed on K-12 (control), cyaA mutant bacteria, and cyaA mutant bacteria supplemented with cAMP (2 mM). n = 100, **p < 0.01; average ± std. dev (n = 3).
Discussion
The type of bacterial food source modulates C. elegans development and lifespan35,36,37,38,39,40,41,42,43,44. In this study, we have screened for bacterial mutants that regulate dauer formation in C. elegans to understand the role of bacterial genes in modulating nutrient sensing pathways. We used the Keio mutant E. coli library15 to identify individual gene knockout mutants that affect bacterial signals that enhance dauer formation in C. elegans. Genes mutated among the bacterial mutants identified in the dauer screen have varied functions (Fig. 1b) including, metabolism, translation, biogenesis of membrane and DNA replication (Fig. 1b and Table 1). These results suggest that bacteria act not only as a food source but also provide signals that influence nutrient signaling pathways in C. elegans. A better understanding of E. coli genes that influence these signaling pathways could give us a better understanding of complex host-bacterial relationships in other species.
In the absence of bacterial food, C. elegans show reduced rates of growth and progeny production and lifespan extension45. These observations can be explained by the idea that worms subjected to suboptimal nutrition or the absence of toxic or pathogenic components of bacteria, extend lifespan. Previously, different bacterial diets10, and bacterial metabolites like folate36 and fatty acids14 have been proposed to modulate lifespan. Our results identify several bacterial genes that are involved in the production or transport of various downstream metabolites to regulate dauer formation and lifespan in worms. Furthermore, we observed that the worms fed on dead bacteria show increase in dauer formation (Fig. S5D), which suggests that viable secondary metabolites can have a profound effect on bacterial physiology that can modulate dauer formation in worms. To further examine if the cAMP-mediated dauer rescue was due to the direct uptake of cAMP by the worm, or an indirect effect on bacterial metabolism, we grew worms on UV-killed bacteria supplemented with cAMP. In these experiments, we found that there was an increase in basal dauer formation in both the K12 and cyaA controls, but that the addition of cAMP to UV-killed cyaA bacteria did not significantly reduce the dauer entry phenotype (Fig. S5D). These experiments suggest that action of cAMP requires live bacteria to mediate its effects on dauer formation.
There are several pathways involved in dauer formation, including guanylyl cyclase, ILS, and TGF-β2. A critical downstream factor of some of these pathways is DAF-16 which has been shown to integrate inputs from various upstream dauer formation pathways2,46. Most of the bacterial mutants that enhance dauer formation from our screen do not extend lifespan in daf-16 null mutant animals and enhance nuclear localization of DAF-16 to a variable extent (Fig. 2). These bacterial mutants also enhance the expression of SOD-3, a direct target of DAF-1647,48. These data suggest that several but not all of these bacterial mutants require DAF-16/FOXO for increased longevity and improved health span, implicating bacterial metabolites in modulating nutrient sensing signaling pathways49.
We also identified that bacteria deficient in cyaA inhibit TGF-β signaling to enhance DAF-16/FOXO to enhance dauer formation and lifespan. Dauer assays using daf-7 and daf-11 null mutants suggests that the effects of feeding on the cyaA mutant bacteria on dauer formation require TGF-β and the guanylyl cyclase (DAF-11) expression2,27,46. DAF-11 is localized primarily in the ciliary endings of some of the amphid neurons and integrates environmental signals from G-protein coupled receptors (GPCR)50. As mentioned before, ILPs and DAF-7 are expressed and released primarily by neurons and ASI neurosensory cells, respectively16. Thus we speculate that cyaA mutant bacteria differentially impact these sensory neurons to regulate changes in dauer formation and lifespan. We speculate that this regulation may be carried out by reducing cAMP levels produced by the E. coli, resulting in changes in secondary metabolites or cAMP itself29. Based on our observation it appears that feeding cyaA mutant bacteria modulates DAF-11 signaling, decreasing DAF-7 neuronal expression (Fig. 7). Further studies are required to uncover the cross talk between cyaA dependent bacterial products on C. elegans signaling pathways.
Figure 7. Model of how bacterial cAMP signaling regulates dauer formation and lifespan in worms.
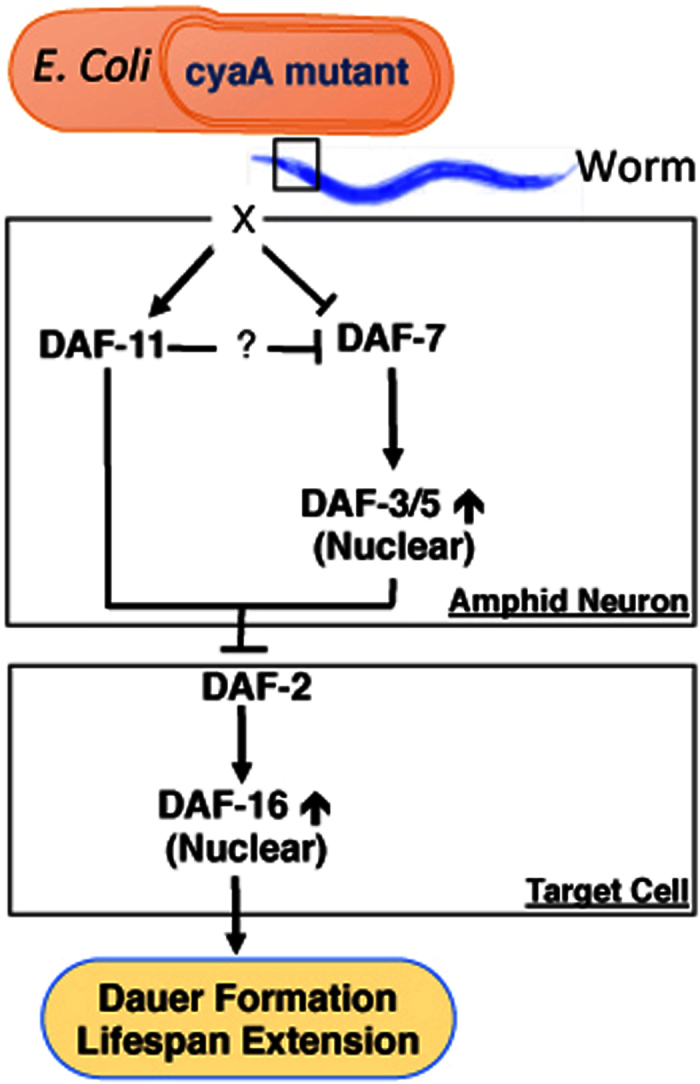
Lack of bacterial cAMP inhibits the expression of DAF-7, resulting in the inhibition of DAF-3/5. This leads to the suppression of IGF-1 pathway in animals, and enhances the dauer formation and lifespan extension in DAF-16/FOXO dependent manner.
Bacterial products as part of the gut microbiome affect many biological processes such as metabolism, longevity, and health span in mammals16,35. The function of some of these bacteria has been well described in several human diseases including obesity, liver diseases, metabolic syndrome, autoimmune disorders, and diabetes51,52,53. Some of these microbiome related products modulate diverse host metabolic activities resulting in pathologies related to insulin sensitivity54,55. However, the mechanisms by which individual bacterial gene products interact with the host to modulate insulin signaling pathways are not known49. Gut microbiome’s influence on biological processes has been observed across different species from insects to mammals. In this study, we describe how mono-association of bacterial mutants with C. elegans can help decipher how specific bacterial signals modulate host physiology. Thus interaction between bacteria and C. elegans can be used to understand conserved signals that are likely to play a role in host-microbiome interactions in humans which may influence diseases like Type II diabetes and obesity.
Methods
C. elegans strains and growth conditions
All strains were obtained from the Caenorhabditis Genetics Center (CGC). The strains obtained from the CGC were outcrossed 6 times to N2 (control) worms. Strains used in this manuscript from CGC are: CB1370 daf-2(e1370) III, CB1372 daf-7(e1372) III, DR47 daf-11(m47) V, CF1038 daf-16(mu86) I, HT1608 daf-2(e1370) III; daf-3(e1376) X,CF1553 muIs84 [(pAD76) sod-3p::GFP + rol-6(su1006)], DR1624 daf-5(e1386) II; daf-2(e1370) III, CF1139 muIs61 [(pKL78) daf16::GFP + rol-6(su1006)], DR1564 daf-2(m41) III, FK181 ksIs2 [daf-7p::GFP + rol-6(su1006)], TM151 sod-2(sj173) I; daf-2(e1370) III; sod-3(sj134) X and PR678 tax-4(p678) III; daf-11(m47) V56. Strains were maintained on nematode growth media plates at 20 °C seeded with E. coli (K12) bacteria grown in Luria broth. For egg collection, gravid adults were soaked in hypochlorite solution as described previously57.
E. coli strains
E. coli mutant strains, including the parent K12 BW25113 strain, were obtained from the Keio mutant collection15. The strain library was shipped and stored as a frozen stock in 96-well plates. Stock plates were pin-replicated to produce a working copy plate containing Luria Broth (LB) and 25 μg/ml kanamycin. For assays used in this study, E. coli strains were stab cultured and grown overnight with shaking at 37 °C in minimal media as described below. Overnight cultures were seeded on minimal media plates and dried at 37 °C for 12 hours prior to transfer of C. elegans.
Media
Luria broth (LB) and nematode growth media (NGM) were used to maintain stock cultures of E. coli and C. elegans strains, respectively. Minimal media was used for assays in this study as described previously58. Liquid media for bacterial growth contained: 0.05 M NaCl, 0.04 M NH4Cl, 0.01 M CaCl2, 0.025 M phosphate buffer, 0.4% glucose, 1 μg /mL thiamine, and 0.001 M MgSO4. Minimal media plates for subsequent C. elegans experiments contained: 2% agar, 0.05 M NaCl, 0.04 M NH4Cl, 0.001 M CaCl2, 1 μg/mL cholesterol, 0.025 M phosphate buffer, 0.4% glucose, 1 μg/mL thiamine, and 0.001 M MgSO4. All compounds supplemented to the agar plates for rescue experiments were purchased from Sigma-Aldrich.
PCR confirmation
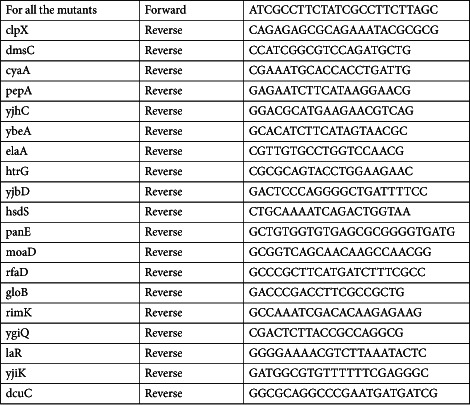
Bacterial strains for kanamycin cassette confirmation were streaked onto LB plates containing 25 μg/ml kanamycin. Strains were colony purified, and genomic PCR was performed. The primer sets were designed for individual knockout strains around the outside of the target gene. The forward primer was designed to be within 200 bp upstream of the 5′ end, and the reverse primer was within 200 bp downstream of the 3′ end of the target gene. The thermal cycler parameters were as follows: initial 5 min at 94 °C, then 25 cycles of 30 s at 94 °C, 10 s at 56 °C, 1 min at 72 °C, and final 5 min at 72 °C. Products were run on an agarose gel and amplified products were examined for appropriate length. Representative list of primers (5′ to 3′).
 |
RT-PCR validation
Quantitative real-time PCR was carried out using SYBR Green I (Sigma Aldrich) assay reagents to verify gene expression profiles. 200 ηg of total RNA were reverse transcribed to cDNA and the real-time PCR reactions were performed on a 7500 Fast System Real-Time PCR cycler (Applied Biosystems, Foster City, CA), according to the manufacturer’s instructions. The gene expression fold changes between different treatment groups were calculated using the delta Ct method59.
 |
Dauer assay
Dauer assay using bacterial mutants
Bacterial strains were grown to stationary phase in minimal medium prior to being seeded on minimal media agar. Mutant and control E. coli plates were left to grow overnight at 37 °C. Eggs were placed on plates containing the E. coli mutants. Plates were incubated at 21.5 °C for 4–5 days prior to assessing dauer or non-dauer state worms. Worm larval states were analyzed using a dissecting microscope and manually counted.
Measurement of cAMP
cAMP levels from bacterial strains and worm strains were measured using the cAMP- Glo TM assay kit purchased from Promega (Madison, WI). For bacteria, 5 ml cultures of K-12, cyaA were grown with or without the presence of cAMP (2 mM) in minimal media at 37 °C for overnight. Cells were counted in 50 μl bacterial culture and were sonicated for complete lysis, and 5 μl of the lysate was used to measure cAMP. The number of bacteria was counted with standard bacterial colony count protocol and the amount of cAMP in different bacteria was expressed per 106 bacterial cells. PBS for complete lysis. 5 μl sample of the lysate was used for cAMP measurement, and the amount of cAMP was expressed per 200 animals.
Fluorescence Microscopy
Age-synchronized L1-stage animals carrying DAF-16::GFP integrated transgenes (daf-16(mgDf47) I) were placed on plates seeded with overnight cultures of bacterial strains as described above. Animals were grown at 15 °C for 2–3 days and then moved to 21.5 °C for one day unless otherwise noted. The fluorescent images were captured using reflected light fluorescence microscopy (Olympus IX3) and the images were then processed for densitometry analysis using ImageJ.
Lifespan assay
Late L4 larvae growing at 15 °C were transferred to fresh minimal media plates with FUdR (5 μg/ml) added to the bacterial lawn as indicated in the results. The first day of adulthood is day 1 in survival curves. Animals were scored as alive, dead or lost every other day. Animals that failed to display touch-provoked movement were scored as dead. Animals that died from causes other than aging, such as sticking to the plate walls, internal hatching or bursting in the vulval region, were scored as lost. Animals were transferred to fresh plates and fresh E. coli every 2 days with cAMP added to have final concentration 2 mM. Plates were dried for 15 minutes at 20 °C before the worms were transferred. All lifespan experiments were performed at 21.5 °C. Survival curves were plotted, and statistical analyses (log-rank tests) were performed using Prism 4 software (Graphpad Software, Inc., San Diego, CA, USA). Statistical significance between lifespan curves was determined by p-value of <0.05 and a mean lifespan of >5% difference compared to the control mean as described before32,60.
Behavioral assays: Pharyngeal pumping and body bends
Worms were observed using stereomicroscope at 40x magnification. Pumps per minute were counted as described previously61. Body bends per minute were counted as described previously62.
Bacterial choice assay
Age-synchronized day 1 adult worms were used. Minimal media plates were used for this assay: 2% agar, 0.05 M NaCl, 0.04 M NH4Cl, 0.001 M CaCl2, 5 μg/mL cholesterol, 0.025 M phosphate buffer, 0.4% glucose, 1 μg/mL thiamine, and 0.001 M MgSO4. Overnight bacterial culture with OD 600 nm of 2.0 was spotted (30 μl) equidistant from the center of the plate and dried for 15 minutes at 37 °C. Both K12 and mutant bacteria had approximately the same cellular density: K12 bacteria at 0.8 × 107 ± 1.2 × 107 colony-forming units (cfu) per ml and dmsC mutant bacteria at 0.7 × 107 ± 1.3 × 107 colony-forming units (cfu) per ml. Worms were washed and prepared as previously described63. The worms were placed in the center of the plate, and positions of the worms were observed using stereomicroscope and images taken at 15 minutes interval. Percentage of worms at each lawn at (1 hr) were plotted as a histogram.
C. elegans feeding behaviour assays
Feeding behavior was assessed based on bacteria depletion as described before64. Minimal media plates were used for this assay: 2% agar, 0.05 M NaCl, 0.04 M NH4Cl, 0.001 M CaCl2, 5 μg/mL cholesterol, 0.025 M phosphate buffer, 0.4% glucose, 1 μg/mL thiamine, and 0.001 M MgSO4. Overnight bacterial culture with OD 600 nm of 0.8 was spotted (60 μl) in center of the plate with streotomycin (300 ng/ml), carbenicillin & Kanamycin (50 μg/ml), and FUdR (5 μg/ml) and the plates were dried for 15 minutes at 37 °C. Age synchronized late L4 worms (n = 60) were washed and prepared as previously described63,64 and introduced on the plates. The worms were left on the plates overnight at 20 °C. Control plates with bacteria spotted as described above with no worms were also incubated at 20 °C overnight. After 24 hrs the bacteria was washed off the plates and checked for OD at 600 nm.
Lawn leaving assay
Age-synchronized day 1 adult animals (n = 20 per plate) were transferred to bacterial lawns cultured under conditions described above. Assay was performed for 5 minutes and lawn leaving events were manually counted by observing under the microscope and averaged to lawn leaving events per minute as previously described65.
Statistical Analysis
Microarray data analysis was performed using SPSS Professional Edition software (IBM). Descriptive statistics including mean and s.e.m. along with one-way ANOVAs followed by multiple comparison tests and two-tailed T-test were used to determine significant differences. P < 0.05 was considered significant as described before32,66. For lifespan assays, log-rank tests were performed by the Prism 4 software as described before32,60. Pearson product-moment correlation coefficient was calculated to measure the correlation between data sets.
Additional Information
How to cite this article: Khanna, A. et al. A genome-wide screen of bacterial mutants that enhance dauer formation in C. elegans. Sci. Rep. 6, 38764; doi: 10.1038/srep38764 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Material
Acknowledgments
We would like to thank Gordon Lithgow, Jennifer Garrison, Mark Watson, Mark Lucanic and Matthew McGee for their comments and contribution to the manuscript. We would like to thank members of the Kapahi and Lithgow labs for discussions and suggestions. We would like to thank Cori Bargmann for tyra-3 rescue strains. This work was funded by grants from American Federation for Aging Research and the NIH (R01AG038688 and RL1 AAG032113).
Footnotes
The authors declare no competing financial interests.
Author Contributions M.V., J.K., A.K., T.M. and P.K. designed the study. M.V., T.M., D.K. and J.K. established the dauer and lifespan assays conditions. N.N., A.K., L.B., J.K. and M.V. performed the dauer and lifespan assays. S.K. helped with cAMP estimation. A.K., M.V., A.S. and P.L. performed DAF-16 localization experiments. A.K., J.K., R.B., D.K., M.G., S.D.M. and C.N. analyzed expression data. A.K., M.V. and P.K. wrote the manuscript.
References
- Hashmi S. et al. A C. elegans model to study human metabolic regulation. Nutr Metab (Lond) 10, 31, doi: 10.1186/1743-7075-10-31 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu P. J. Dauer. WormBook: the online review of C. elegans biology 1–19, doi: 10.1895/wormbook.1.144.1 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Golden J. W. & Riddle D. L. ACaenorhabditis elegans dauer-inducing pheromone and an antagonistic component of the food supply. J Chem Ecol 10, 1265–1280, doi: 10.1007/BF00988553 (1984). [DOI] [PubMed] [Google Scholar]
- Golden J. W. & Riddle D. L. A pheromone influences larval development in the nematode Caenorhabditis elegans. Science 218, 578–580 (1982). [DOI] [PubMed] [Google Scholar]
- Butcher R. A., Ragains J. R., Kim E. & Clardy J. A potent dauer pheromone component in Caenorhabditis elegans that acts synergistically with other components. Proceedings of the National Academy of Sciences of the United States of America 105, 14288–14292, doi: 10.1073/pnas.0806676105 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ludewig A. H. & Schroeder F. C. Ascaroside signaling in C. elegans. WormBook: the online review of C. elegans biology, 1–22, doi: 10.1895/wormbook.1.155.1 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Balla K. M. & Troemel E. R. Caenorhabditis elegans as a model for intracellular pathogen infection. Cellular microbiology 15, 1313–1322, doi: 10.1111/cmi.12152 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beale E., Li G., Tan M. W. & Rumbaugh K. P. Caenorhabditis elegans senses bacterial autoinducers. Appl Environ Microbiol 72, 5135–5137, doi: 10.1128/AEM.00611-06 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ben Arous J., Laffont S. & Chatenay D. Molecular and sensory basis of a food related two-state behavior in C. elegans. PloS one 4, e7584, doi: 10.1371/journal.pone.0007584 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Avery L. & You Y. J. C. elegans feeding. WormBook 1–23, doi: 10.1895/wormbook.1.150.1 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larsen P. L., Albert P. S. & Riddle D. L. Genes that regulate both development and longevity in Caenorhabditis elegans. Genetics 139, 1567–1583 (1995). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riddle D. L., Swanson M. M. & Albert P. S. Interacting genes in nematode dauer larva formation. Nature 290, 668–671 (1981). [DOI] [PubMed] [Google Scholar]
- Ogg S. et al. The Fork head transcription factor DAF-16 transduces insulin-like metabolic and longevity signals in C. elegans. Nature 389, 994–999, doi: 10.1038/40194 (1997). [DOI] [PubMed] [Google Scholar]
- Kaul T. K., Reis Rodrigues P., Ogungbe I. V., Kapahi P. & Gill M. S. Bacterial fatty acids enhance recovery from the dauer larva in Caenorhabditis elegans. PloS one 9, e86979, doi: 10.1371/journal.pone.0086979 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baba T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol 2, 2006 0008, doi: 10.1038/msb4100050 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barna J. et al. Heat shock factor-1 intertwines insulin/IGF-1, TGF-beta and cGMP signaling to control development and aging. BMC developmental biology 12, 32, doi: 10.1186/1471-213X-12-32 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cassada R. C. & Russell R. L. The dauerlarva, a post-embryonic developmental variant of the nematode Caenorhabditis elegans. Dev Biol 46, 326–342 (1975). [DOI] [PubMed] [Google Scholar]
- Vowels J. J. & Thomas J. H. Genetic analysis of chemosensory control of dauer formation in Caenorhabditis elegans. Genetics 130, 105–123 (1992). [DOI] [PMC free article] [PubMed] [Google Scholar]
- al-Waiz M., Mikov M., Mitchell S. C. & Smith R. L. The exogenous origin of trimethylamine in the mouse. Metabolism 41, 135–136 (1992). [DOI] [PubMed] [Google Scholar]
- Kanehisa M. et al. From genomics to chemical genomics: new developments in KEGG. Nucleic acids research 34, D354–357, doi: 10.1093/nar/gkj102 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mi H. et al. The PANTHER database of protein families, subfamilies, functions and pathways. Nucleic acids research 33, D284–288, doi: 10.1093/nar/gki078 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tatusov R. L., Koonin E. V. & Lipman D. J. A genomic perspective on protein families. Science 278, 631–637 (1997). [DOI] [PubMed] [Google Scholar]
- Serres M. H. & Riley M. MultiFun, a multifunctional classification scheme for Escherichia coli K-12 gene products. Microbial & comparative genomics 5, 205–222 (2000). [DOI] [PubMed] [Google Scholar]
- Burnell A. M., Houthoofd K., O’Hanlon K. & Vanfleteren J. R. Alternate metabolism during the dauer stage of the nematode Caenorhabditis elegans. Exp Gerontol 40, 850–856, doi: 10.1016/j.exger.2005.09.006 (2005). [DOI] [PubMed] [Google Scholar]
- Jensen V. L., Simonsen K. T., Lee Y.-H., Park D. & Riddle D. L. RNAi Screen of DAF-16/FOXO Target Genes in C. elegans Links Pathogenesis and Dauer Formation. PloS one 5, e15902, doi: 10.1371/journal.pone.0015902 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henderson S. T. & Johnson T. E. daf-16 integrates developmental and environmental inputs to mediate aging in the nematode Caenorhabditis elegans. Current biology: CB 11, 1975–1980 (2001). [DOI] [PubMed] [Google Scholar]
- Murphy C. T. et al. Genes that act downstream of DAF-16 to influence the lifespan of Caenorhabditis elegans. Nature 424, 277–283, doi: 10.1038/nature01789 (2003). [DOI] [PubMed] [Google Scholar]
- Lee S. S., Kennedy S., Tolonen A. C. & Ruvkun G. DAF-16 target genes that control C. elegans life-span and metabolism. Science 300, 644–647, doi: 10.1126/science.1083614 (2003). [DOI] [PubMed] [Google Scholar]
- You C. et al. Coordination of bacterial proteome with metabolism by cyclic AMP signalling. Nature 500, 301–306, doi: 10.1038/nature12446 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldenbaum P. E. & Hall G. A. Transport of cyclic adenosine 3′,5′-monophosphate across Escherichia coli vesicle membranes. Journal of bacteriology 140, 459–467 (1979). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee S. J., Murphy C. T. & Kenyon C. Glucose shortens the life span of C. elegans by downregulating DAF-16/FOXO activity and aquaporin gene expression. Cell metabolism 10, 379–391, doi: 10.1016/j.cmet.2009.10.003 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen D. et al. Germline signaling mediates the synergistically prolonged longevity produced by double mutations in daf-2 and rsks-1 in C. elegans. Cell reports 5, 1600–1610, doi: 10.1016/j.celrep.2013.11.018 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pierce S. B. et al. Regulation of DAF-2 receptor signaling by human insulin and ins-1, a member of the unusually large and diverse C. elegans insulin gene family. Genes Dev 15, 672–686, doi: 10.1101/gad.867301 (2001). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fielenbach N. & Antebi A. C. elegans dauer formation and the molecular basis of plasticity. Genes & development 22, 2149–2165, doi: 10.1101/gad.1701508 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ottaviani E. et al. Gut microbiota as a candidate for lifespan extension: an ecological/evolutionary perspective targeted on living organisms as metaorganisms. Biogerontology 12, 599–609, doi: 10.1007/s10522-011-9352-5 (2011). [DOI] [PubMed] [Google Scholar]
- Virk B. et al. Excessive folate synthesis limits lifespan in the C. elegans: E. coli aging model. BMC biology 10, 67, doi: 10.1186/1741-7007-10-67 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brooks K. K., Liang B. & Watts J. L. The influence of bacterial diet on fat storage in C. elegans. PloS one 4, e7545, doi: 10.1371/journal.pone.0007545 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiao R. et al. RNAi Interrogation of Dietary Modulation of Development, Metabolism, Behavior, and Aging in C. elegans. Cell reports 11, 1123–1133, doi: 10.1016/j.celrep.2015.04.024 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samuel B. S., Rowedder H., Braendle C., Felix M. A. & Ruvkun G. Caenorhabditis elegans responses to bacteria from its natural habitats. Proceedings of the National Academy of Sciences of the United States of America 113, E3941–3949, doi: 10.1073/pnas.1607183113 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shtonda B. B. & Avery L. Dietary choice behavior in Caenorhabditis elegans. The Journal of experimental biology 209, 89–102, doi: 10.1242/jeb.01955 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lynn D. A. et al. Omega-3 and -6 fatty acids allocate somatic and germline lipids to ensure fitness during nutrient and oxidative stress in Caenorhabditis elegans. Proceedings of the National Academy of Sciences of the United States of America 112, 15378–15383, doi: 10.1073/pnas.1514012112 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pang S. & Curran S. P. Adaptive capacity to bacterial diet modulates aging in C. elegans. Cell metabolism 19, 221–231, doi: 10.1016/j.cmet.2013.12.005 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pang S., Lynn D. A., Lo J. Y., Paek J. & Curran S. P. SKN-1 and Nrf2 couples proline catabolism with lipid metabolism during nutrient deprivation. Nature communications 5, 5048, doi: 10.1038/ncomms6048 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steinbaugh M. J. et al. Lipid-mediated regulation of SKN-1/Nrf in response to germ cell absence. eLife 4, doi: 10.7554/eLife.07836 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kang C. & Avery L. Systemic regulation of starvation response in Caenorhabditis elegans. Genes & development 23, 12–17, doi: 10.1101/gad.1723409 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murphy C. T. The search for DAF-16/FOXO transcriptional targets: approaches and discoveries. Exp Gerontol 41, 910–921, doi: 10.1016/j.exger.2006.06.040 (2006). [DOI] [PubMed] [Google Scholar]
- Honda Y. & Honda S. The daf-2 gene network for longevity regulates oxidative stress resistance and Mn-superoxide dismutase gene expression in Caenorhabditis elegans. FASEB journal: official publication of the Federation of American Societies for Experimental Biology 13, 1385–1393 (1999). [PubMed] [Google Scholar]
- Oh S. W. et al. Identification of direct DAF-16 targets controlling longevity, metabolism and diapause by chromatin immunoprecipitation. Nature genetics 38, 251–257, doi: 10.1038/ng1723 (2006). [DOI] [PubMed] [Google Scholar]
- Gipson G. T. et al. Multi-platform investigation of the metabolome in a leptin receptor defective murine model of type 2 diabetes. Mol Biosyst 4, 1015–1023, doi: 10.1039/b807332e (2008). [DOI] [PubMed] [Google Scholar]
- Birnby D. A. et al. A transmembrane guanylyl cyclase (DAF-11) and Hsp90 (DAF-21) regulate a common set of chemosensory behaviors in caenorhabditis elegans. Genetics 155, 85–104 (2000). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicholson J. K., Connelly J., Lindon J. C. & Holmes E. Metabonomics: a platform for studying drug toxicity and gene function. Nat Rev Drug Discov 1, 153–161, doi: 10.1038/nrd728 (2002). [DOI] [PubMed] [Google Scholar]
- Nicholson J. K., Holmes E., Lindon J. C. & Wilson I. D. The challenges of modeling mammalian biocomplexity. Nat Biotechnol 22, 1268–1274, doi: 10.1038/nbt1015 (2004). [DOI] [PubMed] [Google Scholar]
- Nicholson J. K. & Wilson I. D. Opinion: understanding ‘global’ systems biology: metabonomics and the continuum of metabolism. Nat Rev Drug Discov 2, 668–676, doi: 10.1038/nrd1157 (2003). [DOI] [PubMed] [Google Scholar]
- Backhed F., Ley R. E., Sonnenburg J. L., Peterson D. A. & Gordon J. I. Host-bacterial mutualism in the human intestine. Science 307, 1915–1920, doi: 10.1126/science.1104816 (2005). [DOI] [PubMed] [Google Scholar]
- Chen Z. et al. Incorporation of therapeutically modified bacteria into gut microbiota inhibits obesity. The Journal of clinical investigation 124, 3391–3406, doi: 10.1172/JCI72517 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coburn C. M., Mori I., Ohshima Y. & Bargmann C. I. A cyclic nucleotide-gated channel inhibits sensory axon outgrowth in larval and adult Caenorhabditis elegans: a distinct pathway for maintenance of sensory axon structure. Development 125, 249–258 (1998). [DOI] [PubMed] [Google Scholar]
- Rogers A. N. et al. Life span extension via eIF4G inhibition is mediated by posttranscriptional remodeling of stress response gene expression in C. elegans. Cell Metab 14, 55–66, doi: 10.1016/j.cmet.2011.05.010 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim H. W., Matin A. & Rhee M. S. Microgravity Alters the Physiological Characteristics of Escherichia coli O157:H7 ATCC 35150, 43889, and 43895 Under Different Nutrient Conditions. Applied and environmental microbiology, doi: 10.1128/AEM.04037-13 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hellemans J., Mortier G., De Paepe A., Speleman F. & Vandesompele J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome biology 8, R19, doi: 10.1186/gb-2007-8-2-r19 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen D., Thomas E. L. & Kapahi P. HIF-1 modulates dietary restriction-mediated lifespan extension via IRE-1 in Caenorhabditis elegans. PLoS genetics 5, e1000486, doi: 10.1371/journal.pgen.1000486 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raizen D., Song B. M., Trojanowski N. & You Y. J. Methods for measuring pharyngeal behaviors. WormBook: the online review of C. elegans biology 1–13, doi: 10.1895/wormbook.1.154.1 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawin E. R., Ranganathan R. & Horvitz H. R. C. elegans locomotory rate is modulated by the environment through a dopaminergic pathway and by experience through a serotonergic pathway. Neuron 26, 619–631 (2000). [DOI] [PubMed] [Google Scholar]
- Glater E. E., Rockman M. V. & Bargmann C. I. Multigenic natural variation underlies Caenorhabditis elegans olfactory preference for the bacterial pathogen Serratia marcescens. G3 4, 265–276, doi: 10.1534/g3.113.008649 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gomez-Amaro R. L. et al. Measuring Food Intake and Nutrient Absorption in Caenorhabditis elegans. Genetics 200, 443–454, doi: 10.1534/genetics.115.175851 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bendesky A., Tsunozaki M., Rockman M. V., Kruglyak L. & Bargmann C. I. Catecholamine receptor polymorphisms affect decision-making in C. elegans. Nature 472, 313–318, doi: 10.1038/nature09821 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Portman D. S. & Emmons S. W. Identification of C. elegans sensory ray genes using whole-genome expression profiling. Developmental biology 270, 499–512, doi: 10.1016/j.ydbio.2004.02.020 (2004). [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.