Abstract
Base excision repair (BER) is the most prominent DNA repair pathway in human mitochondria. BER also results in a temporary generation of AP-sites, single-strand breaks and nucleotide gaps. Thus, incomplete BER can result in the generation of DNA repair intermediates that can disrupt mitochondrial DNA replication and transcription and generate mutations. We carried out BER analysis in highly purified mitochondrial extracts from human cell lines U2OS and HeLa, and mouse brain using a circular DNA substrate containing a lesion at a specific position. We found that DNA ligation is significantly slower than the preceding mitochondrial BER steps. Overexpression of DNA ligase III in mitochondria improved the rate of overall BER, increased cell survival after menadione induced oxidative stress and reduced autophagy following the inhibition of the mitochondrial electron transport chain complex I by rotenone. Our results suggest that the amount of DNA ligase III in mitochondria may be critical for cell survival following prolonged oxidative stress, and demonstrate a functional link between mitochondrial DNA damage and repair, cell survival upon oxidative stress, and removal of dysfunctional mitochondria by autophagy.
Keywords: Mitochondrial DNA repair intermediates, Oxidative stress, Autophagy, Cell survival
1. Introduction
Mitochondrial DNA (mtDNA) is a ~16.5 kb circular genome that encodes for 22 transfer RNAs, two ribosomal RNAs, and 13 proteins that are all subunits of the oxidative phosphorylation system. Studies on human tissues and in animal models have unequivocally demonstrated the importance of mtDNA integrity for normal cell function and organ development [1–3]. MtDNA deletions were first reported in myopathies and Kearns–Sayer syndrome (KSS) in 1988 [4,5], and mtDNA point mutations have been found in Leber’s hereditary optic neuropathy (LHON) [6]. Since then, over 200 pathogenic mtDNA mutations have been identified [2]. Several human diseases are caused by defects in the pathways that control mtDNA maintenance [7,8]. Moreover, ample evidence suggests that accumulation of damaged and mutated mtDNA correlate with normal aging as well as with neurodegenerative diseases [9–11]. Cells contain several hundred to several thousand copies of mtDNA. The number of copies present in each mitochondrion varies between different cell types, depending on the energy demand within the tissue [11].
Several mechanisms generate cellular reactive oxygen species (ROS) including controlled ROS production by NADPH oxidase enzymes in phagocytic cells to defend against microorganism infection and in non-phagocytic cells such as colon epithelial and endothelial cells [12,13]. However, mitochondria are overall the primary site of cellular ROS production. ROS can react with DNA generating a variety of DNA lesions such as oxidized bases and DNA strand breaks. The close proximity of mtDNA to the oxidative phosphorylation system in the inner membrane is thought to make mtDNA particularly vulnerable to damage and mutations from ROS produced as a byproduct of the oxidative phosphorylation process. This may account for the higher mutation frequency observed in mtDNA relative to nuclear DNA [14].
Base excision repair (BER) is the major pathway for repair of ROS-induced DNA lesions that are frequent events in mtDNA [15,16]. Compared with the nucleus, mitochondria have limited DNA repair capacity. BER is the most prominent DNA repair pathway in mitochondria. BER is initiated by a DNA glycosylase that removes a damaged base leaving a baseless site or AP-site. The resulting AP-site is incised on the 5′-side by AP-endonuclease 1 (APE1) followed by 5′-dRP excision and gap filling performed by a DNA polymerase, and finally sealing of the repaired DNA by a DNA ligase. BER in mitochondria can proceed through single-nucleotide insertion (short-patch) or via incorporation of several nucleotides (long-patch) [17–19]. Long-patch BER may result in displacement of the 5′-dRP residue along with several nucleotides. The removal of the 5′-single stranded DNA or 5′-flap requires additional proteins [18–21]. Incomplete BER can give rise to unprocessed DNA repair intermediates like AP-sites and nicked DNA. These intermediates can be mutagenic, cause DNA strand breaks, and block DNA replication and transcription, which all affect mitochondrial function.
Nuclear and mitochondrial DNA ligase III are generated from a single gene by alternative translation initiation [22]. In the nucleus, DNA ligase III and XRCC1 are tightly associated and DNA ligase III level and activity has been shown to be dependent on XRCC1 expression [23,24]. However, DNA ligase III activity in mitochondria is independent of XRCC1 [25]. These observations led to the view that DNA ligase III is critical for the nuclear and mitochondrial single-strand DNA repair and BER. However, recently, two reports demonstrated that DNA ligase III is critical for mtDNA integrity and cell survival but not for nuclear DNA repair [3,26].
In this study, we show that in mitochondria, DNA ligation is slower than the preceding BER steps and that overexpression of DNA ligase III in mitochondria improves BER and reduces the cytotoxicity of menadione induced oxidative stress. Moreover, we show that the basal levels of autophagy are similar in cells overexpressing DNA ligase III in mitochondria and control cells; however, cells overexpressing DNA ligase III in mitochondria display lower rotenone-induced autophagy compared with control cells. Our results indicate that the ligation step in mtBER is important for cellular survival and suggest that mtDNA repair is a critical mitochondrial surveillance system.
2. Materials and methods
Synthetic oligonucleotides were from TAG Copenhagen. [γ-32P] ATP (3000 Ci/mmol) and [α-32P] dCTP (3000 Ci/mmol) were from Perkin Elmer. Restriction enzymes were from New England Biolabs and paramagnetic protein-A beads were from Dynal, Life Technologies. MitoTracker Red CMXRos was from Life Technologies.
2.1. Cell lines and preparation of mitochondrial extracts
HeLa and U2OS cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM-Glutamax, Gibco), or in DMEM containing galactose instead of glucose as indicated, with 10% fetal calf serum and 0.1 mg/ml gentamicin in 5% CO2. Mitochondrial extracts were prepared as described previously [17]. Mitochondria were routinely isolated from 35 dishes (150 mm) at ~95–100% confluence. Cells were detached by trypsination, collected by centrifugation at 400 × g, resuspended in hypotonic buffer (20 mM HEPES, pH 8.0, 5 mM KCl and 1 mM DTT) and incubated on ice until swollen. 2× MSH buffer (420 mM mannitol, 140 mM sucrose, 20 mM HEPES pH 7.4, 4 mM EDTA, 2 mM EGTA, and 5 mM DTT) was then added (1:1, v/v) and the cells were broken in a Dounce homogenizer with a B-type pestle. The homogenate was centrifuged at 1000 × g. This step was repeated until no nuclei were seen in the pellet (typically 2–3 times). The crude mitochondria were pelleted at 10,000 × g for 30 min, resuspended in 1× MSH/50% Percoll and loaded on top of a 1× MSH/50% Percoll mixture and centrifuged at 50,000 g for 75 min. Mitochondrial fraction was removed from the gradient and centrifuged in 1× MSH buffer at 3000 × g for 10 min to remove Percoll. The mitochondrial pellet was washed once in 1× MSH buffer without EDTA and EGTA and resuspended in 0.15 ml of the same buffer, and then divided into three or four tubes and treated with different concentrations of proteinase K at 37 °C for 20 min. Usually, treatment of isolated mitochondria with ~0.2 mg/ml proteinase K was sufficient to clear mitochondria of nuclear contamination. Phenyl-methylsulfonyl fluoride (PMSF) was added at final concentration of 5 mM to inactivate proteinase K. Mitochondria were pelleted at 10,000 × g for 10 min and washed twice with 0.2 ml of a protease inhibitor mix (Complete proteinase inhibitor (Roche), 2× MSH buffer (1:1, v/v) and 5 mM PMSF). The mitochondria were suspended in buffer I (10 mM HEPES, pH 8.0 and 200 mM KCl) and then an identical volume of buffer II (10 mM HEPES, pH 8.0, 2 mM DTT, 40% glycerol, 1% Nonidet P-40, 1% Triton X-100, Complete protease inhibitors, phosphatase inhibitor cocktails 1 and 2, 10 mM PMSF, and 200 mM KCl) and incubated at 4 °C for 90 min and briefly sonicated. Mitochondrial debris was removed by centrifugation at 16,200 × g for 10 min, and the supernatant was collected and stored at −80 °C. The protein concentration was measured using the Bradford method (BioRad). Mitochondria from mouse brain were isolated as described before [23]. Briefly, frozen whole brain was thawed in 5 ml of ice-cold 1× MSH, pH 7.4. The brain was cut into small pieces and homogenized in a Teflon Dounce homogenizer and centrifuged at 600 × g for 10 min. The supernatant was centrifuged at 7000 × g for 10 min. The supernatant was carefully removed and the pellet was suspended in 1× MSH/50% Percoll and loaded on top of a 1× MSH/50% Percoll mixture and centrifuged at 50,000 × g for 75 min. The mitochondrial fraction was removed from the gradient and mitochondrial extracts were prepared as described above.
2.2. Western blot analysis (WB)
20–50 μg of the protein extracts were separated in 10% Tris–glycine SDS-PAGE (Invitrogen). For LC3B, 4–20% Tris–glycine SDS-gel was used. The following primary antibodies were used: PCNA (sc-56, Santa Cruz), lamin (sc-6215, Santa Cruz), COX IV (AMB-7487, BioSite), VDAC-1 (sc-58649, Santa Cruz), Ku86 (sc-1485 Santa Cruz), DNA ligase III (611877, BD Transduction Laboratories), LC3B (NB600-1384, Novus Biologicals), cleaved PARP-1 antibody (1074-1, Epitomics), Polγ (ab128899, Abcam), TFAM (B01P, Abnova). TOPOIIα antibody was a home-made rabbit polyclonal antibody. The secondary antibodies, polyclonal rabbit-mouse IgG/HRP or peroxidase-labeled polyclonal swine-rabbit IgG were from Dako Cytomation.
For autophagy, the experiments were carried out as described previously [27]. In brief, 1 × 106 cells were seeded on 100 mm dishes. 24 h later, cells were treated with 5 μM rotenone. Cells were incubated for 24 h, harvested by trypsination and lysed in 0.15 ml lysis buffer (50 mM Tris–HCl, pH 7.4, 150 mM NaCl, 1 mM EDTA, 1% Triton X-100, 1 mM DTT and protease inhibitor cocktail, Roche) for 30 min on ice. The lysate was briefly sonicated three times at 5 W for 5 s on ice. The extracts were centrifuged at 16,000 × g for 10 min and the supernatant collected.
2.3. BER assay
The BER assay was performed as described previously [28,29]. Briefly, an uracil-containing oligonucleotide (5′-GATCCTCTAGAGTUGACCTGCA-3′) was annealed to ssDNA derived from pGEM-3Zf(+) plasmid containing guanine opposite uracil (supplementary Fig. S1). DNA synthesis was carried out with T4 gene 32 ssDNA-binding protein, T4 DNA polymerase, T4 DNA ligase, and dNTPs at 37 °C for 120 min. The covalently closed double-strand circular DNA was purified from agarose gel. C:G DNA substrate was prepared using oligo containing C instead of U and used as control.
BER was measured using 30 μg mitochondrial extract in, 40 mM HEPES–KOH pH 7.8, 1 mM DTT, 5 mM MgCl2, 100 mM KCl, 2 mM ATP, 0.36 mg/ml BSA, 20 μM dATP, 20 μM dTTP, 20 μM dGTP, 10 μM dCTP, 4.5 mM phosphocreatine, 50 ng/μl creatine kinase, 250 ng DNA substrate, and 80 nCi/μl [α-32P]dCTP at 37 °C for the indicated times in a volume of 25 μl. The reaction was stopped by adding EDTA and further incubated with SDS and proteinase K at 37 °C for 30 min. DNA was purified with phenol/chloroform extraction and salt precipitation. DNA was resuspended in 10 mM Tris–HCl pH 8.5 and divided into two aliquots; one was incubated with 1 μl T4 DNA ligase (Roche) at 37 °C for 60 min and one was left untreated. T4 DNA ligase was inactivated at 70 °C for 10 min and DNA was digested with XbaI and HindIII restriction enzymes and separated in 12% denaturing polyacrylamide gel at 300 V for 1 h.
2.4. Cloning of YFP fusion constructs and confocal microscopy
C-terminal YFP-tagged DNA ligase III lacking the stop codon was prepared by PCR amplification using a cDNA that contains the entire coding region of DNA ligase III as template (SC109386, Origene) and forward primer (F1) 5′-AGATCTGCTAGCATGTCTTTGGCTTTCAAGATC-3′ and reverse primer (R1) 5′-GGATCCACCGGTCGGCAGGGAGCTACCAGTCTC-3′. The PCR products were digested with Nhe I and Age I and cloned into YFP-N1 vector. Then, the second start codon (M2) in ligase III was changed to ATC using site-directed mutagenesis and oligonucleotides: 5′-CCCTGTGAGATCGCTGAGCAACGG-3′ and 5′-CCGTTGCTCAGCGATCTCACAGGG-3′. Next, the start codon for YFP was changed to ATC using oligonucleotides 5′-CCGGTCGCCACCATCGTGAGCAAGGGCG-3′ and 5′-CGCCCTTGCTCACGATGGTGGCGACCGG-3. Finally, the mitochondrial localization signal from SOD2 (5′-ATGTTGAGCCGGGCAGTGTGCGGCACCAGCAGG CAGCTGG CTCCGGCTT TGGGGTATCTGGGCTCCAGGCAG-3′) was inserted into the NheI restriction site in frame with DNA lig III.
Fluorescence images of live cells stably expressing MLS-lig III-YFP and MLS-YFP control cells preincubated with MitoTracker were produced using a Zeiss LSM 700 microscope equipped with WN-Achroplan 40×/0.75 M27 objective. Images were taken using the following settings: YFP excitation at λ = 488 nm, detection at λ = 405/488, MitoTracker, excitation at λ = 555 nm, detection at λ = 555/634 nm.
2.5. DNA ligase activity assay
For DNA ligase substrate, oligonucleotide: 5′-GCTTGGCACTGGCCGTCGTTTTACATTTGAGACGTC CCAGCTTTTTTGCTGGGACGTCTC-3′ was radio-labeled at the 5′ end using [γ-32P]ATP and T4 polynucleotide kinase (New England Biolabs) and annealed to oligonucleotide: 5′-AAATGTAAAACGACGGCCAGTG-3′, by heating at 85 °C for 3 min and gradual cooling to room temperature. For immunoprecipitation, paramagnetic protein-A beads were covalently coupled to anti-GFP antibody (ab290, Abcam) according to the producer’s protocol. Beads were incubated with 0.2 mg mitochondrial extract at 4 °C for 3 h under constant rotation. The beads were washed in 20 mM HEPES, pH 7.4, 1 mM DTT, 1 M NaCl, and Proteinase inhibitor cocktail (Roche), and suspended in 20 μl reaction containing ligase buffer (Roche) and 5 nM DNA substrate and incubated at 37 °C for 1 h. Loading buffer was added, heated at 85 °C, and the sample was separated in 8% denaturing polyacrylamide gel. Reactions containing immunoprecipitates from MLS-YFP cells mitochondrial extract and T4 DNA ligase were used as negative and positive controls, respectively.
2.6. Cell survival and colony forming assay
The cell survival assay was performed essentially as described previously [30]. 15,000 cells were plated in each well in 96-well plates with complete growth medium. Twenty-four hours later, the cells were washed in PBS and exposed to the indicated concentrations of menadione (in growth medium without FBS) for 3 h. Then, cells were washed with PBS and incubated in complete medium for 19 h. Viability was determined by the ability of cells to reduce the WST-1 dye (Roche). Ten microliters of WST-1 was added to each well and incubated at 37 °C for 3 h and the absorbance was read (480–595 nm). For colony forming assay, 2000 cells were seeded on 100 mm growth dishes. After 18 h, the cells were treated with 10 μM menadione for 3 h washed with PBS and incubated in complete medium for 6 days. Cells were fixed in 3:1 solution of methanol:acetic acid for 1 h and stained in crystal violet staining solution (1 mg/ml crystal violet in water) overnight.
2.7. Determination of mitochondrial ROS
Cells were incubated in Hank’s balanced salt solution medium containing 5 μM MitoSOX Red (Molecular Probes, Invitrogen) at 37 °C for 30 min, trypsinized and washed in PBS. Determination of mitochondrial ROS was carried out using a FACScalibur (BD Bioscience). MitoSOX Red was excited at 488 nm and data collected at FSC, SSC and 585/42 nm (FL2) channels. A minimum of 50,000 events was collected. Appropriate color compensation was applied to distinguish signals from YFP and MitoSOX Red.
2.8. Statistical analysis
Data are plotted as mean ± SEM. Statistical significance of the changes was determined by one-way ANOVA followed by Bonferroni post hoc test. Statistical analysis was performed using Excel software.
3. Results
3.1. DNA ligation is slower than the preceding mtBER steps
Nuclear genes can encode both nuclear-targeted and mitochondrial-targeted forms of BER proteins either by alternative splicing or by use of alternative sites of translation initiation [9,31]. Thus, preparation of mitochondrial extracts free of nuclear BER proteins is critical for mitochondrial BER (mtBER) analysis. We have demonstrated previously that a mild treatment of isolated mitochondria with proteinase K provides a rapid and simple method to make mitochondrial extracts free of nuclear protein contaminants [17]. Fig. 1 shows results of a typical Western blot analysis of mitochondria after treatment with 0.2 mg/ml proteinase K at 37 °C for 20 min. Total cell extract was used as a positive control for the antibodies. The apparent absence of abundant nuclear proteins TOPO IIα, Ku 86, lamin and PCNA strongly suggests that, under our experimental conditions, we were able to prepare mitochondrial extracts free of nuclear contaminants. Proteinase K treatment slightly degraded voltage-dependent anion channel 1 (VDAC-1) protein (which is located at the outer mitochondrial membrane), TFAM and Polγ, but had little or no detectable effect on the inner membrane protein COX IV. Altogether, these results suggest that proteinase K treatment did not cause significant degradation of mitochondrial proteins.
Fig. 1.

Representative image of Western blot analysis of mitochondrial extracts prepared from proteinase K treated, purified mitochondria. 20 μg total cell extracts and 50 μg mitochondrial extracts were used in each experiment. Antibodies against TOPO IIα, Ku 86, lamin and PCNA were used to assess contamination of the mitochondrial extracts with nuclear proteins. VDAC-1 and COX IV were used as the marker of the outer- and the inner-mitochondrial membrane proteins, respectively, and Polγ and TFAM as the core mitochondrial nucleoid proteins.
Mitochondria BER analyses were carried out using a DNA substrate containing a lesion (uracil) at a specific position (Supplementary Fig. S1). The DNA was incubated in the mitochondrial extracts, and then purified from the extracts by phenol/chloroform purification and divided into two aliquots, one was incubated with T4 DNA ligase (to determine the level of unligated but ligatable BER intermediates after repair) and one was left untreated. T4 DNA ligase treatment of the DNA resulted in ~2-fold increase in the intensity of the fully repaired 24 nucleotide band at the 40 and 60 min incubation times, indicating the presence of substantial amounts of unligated, but ligatable, BER intermediates in the samples (Fig. 2A and B). These results suggest that the DNA ligation reaction is less efficient than the preceding mtBER steps. Next, we carried out BER analysis in mitochondrial extracts from mouse brain. Western blot analysis showed that these mitochondrial extracts were not contaminated with the nuclear proteins Ku 86, lamin and PCNA (Supplementary Fig. S2). Similar to HeLa and U2OS cells, incubation of the repaired DNA substrate with T4 DNA ligase resulted in ~1.6 and 1.85 fold more ligated products after 40 and 60 min incubation, respectively (Fig. 2C), indicating the presence of a substantial amount of unligated, but ligatable, BER intermediates in the sample. Altogether, these results suggest that DNA ligation is the kinetic “bottleneck” in BER in mitochondrial extracts from HeLa, U2OS cells and mouse brain.
Fig. 2.
BER analysis of mitochondrial extracts. BER analysis was carried out in duplicate using two independently prepared mitochondrial extracts from HeLa (A) and U2OS (B) cell lines. The increased level of the upper band after T4 DNA ligase treatment relative to the untreated samples indicates the presence of unligated DNA repair intermediates (rep. int.) in the sample. (C) BER analysis of mitochondrial extracts from mouse brain as per panels A and B. The bar charts show the relative intensity of the upper band (fully repaired 24 nucleotides) to the lower bands (repair intermediates) in each lane. The error bars show standard deviation of the mean. Absence of detectable repair products in G:C control DNA substrate indicates site specific repair of uracil in U:G DNA substrate.
3.2. Overexpression of DNA ligase III in mitochondria
To test whether the low DNA ligase activity observed could be compensated for by increasing the concentration of DNA ligase III in mitochondria, we transfected U2OS cells with a construct encoding DNA ligase III, C-terminally fused with yellow fluorescent protein (YFP). The human DNA ligase III gene encodes both a nuclear and a mitochondrial form of the protein by alternative translation initiation [22]. We had difficulty obtaining cells that stably expressed the longer form of DNA ligase III (which contains a mitochondrial localization signal (MLS) at its N-terminus), in mitochondria. Over time the protein was localized to both the nucleus and mitochondria (not shown). However, by utilizing an additional MLS from superoxide dismutase 2 (SOD2) to DNA ligase III, we were able to prepare cells stably overexpressing DNA ligase III in mitochondria (henceforward denoted as MLS-lig III-YFP). The mitochondrial localization of the fusion protein was confirmed using MitoTracker, a fluorescence dye that specifically labels mitochondria in live cells (Fig. 3I and II). Cells stably expressing MLS-YFP without ligase III were also prepared (Fig. 3III and IV) and were used as a control in the following experiments.
Fig. 3.
Overexpression of DNA ligase III in mitochondria. U2OS cells stably expressing DNA ligase III fused with YFP and the mitochondrial localization signal of SOD2 (MLS-lig III-YFP, panels I and II) or MLS-YFP alone as control (III and IV), were established. Representative images of live MLS-lig III-YFP and MLS-YFP cells pre-incubated with MitoTracker Red showing mitochondrial localization of the fusion proteins (see merged images). Panels II and IV are high magnification, z-stacked image of the mitochondrial network in a MLS-lig III-YFP and a MLS-YFP cell, respectively.
Overexpression of DNA ligase III in mitochondria did not cause detectable levels of apoptotic cell death as determined by Western blot analysis of total cell extracts for PARP-1 (indicator of apoptosis) (Supplementary Fig. S3), nor any increase in mitochondrial ROS production (Supplementary Fig. S4). Altogether, these results indicate that overexpression of DNA ligase III-YFP in mitochondria does not induce apoptosis or increase mitochondrial ROS production.
3.3. Overexpression of DNA ligase III in mitochondria enhances BER in mitochondrial extracts
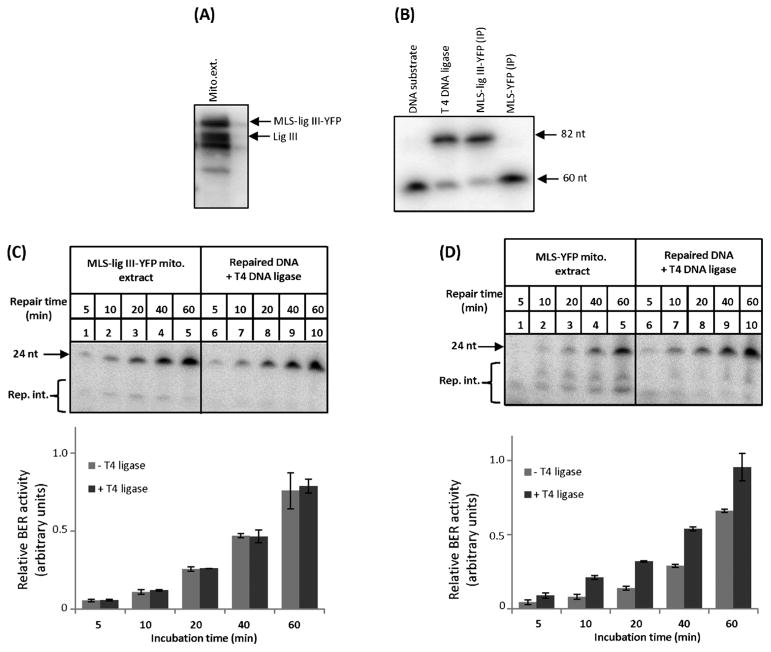
Western blot analysis of mitochondrial extract from MLS-lig III-YFP cells detected an additional DNA ligase III band that corresponded to the predicated size of DNA ligase III and YFP combined (Fig. 4A). Proteinase K treatment of mitochondria from MLS-lig III-YFP cells did not cause detectable levels of degradation of endogenous DNA ligase III or MLS-lig III-YFP III (Supplementary Fig. S5). The amount of DNA ligase III in the mitochondria in MLS-lig III-YFP cells was ~1.5–2 fold higher than the endogenous ligase III. First, we tested whether the fusion protein was catalytically active. We immunoprecipitated (IP) MLS-ligase III-YFP from mitochondrial extracts using anti-GFP (also recognizes YFP) coupled beads and tested it for DNA ligase activity. The result showed that the fusion protein had DNA ligase activity (Fig. 4B). Next, we carried out BER analysis in mitochondrial extracts from MLS-lig III-YFP cells under the same conditions as shown in Fig. 2, and found that over-expression of DNA ligase III in mitochondria improved BER activity in mitochondrial extracts (Fig. 4C). Incubation of repaired DNA with T4 DNA ligase only slightly increased the intensity of the upper bands (Fig. 4C and the diagram), indicating that overexpression of DNA ligase III in mitochondria enhanced overall mtBER. BER analysis of mitochondrial extracts from MLS-YFP cells was included as a control (Fig. 4D).
Fig. 4.
Mitochondrial extracts from MLS-lig III-YFP cells show increased rate of BER compared with control cells. (A) Western blot analysis of mitochondria isolated from MLS-lig III-YFP cells using antibody against DNA ligase III. (B) To test whether MLS-lig III-YFP fusion protein is catalytically active, MLS-ligase III-YFP was immunoprecipitated (IP) from mitochondrial extracts using an antibody against GFP (also recognizes YFP). The IP was assayed for DNA ligase activity for 1 h, and DNA was separated in denaturing polyacrylamide gel. T4 DNA ligase and the IP from mitochondrial extracts of MLS-YFP cells were used as positive and negative controls, respectively. Upper band corresponds to the fully ligated 82 nucleotides substrate (82 nt), and the lower band are unligated 60 nucleotide end-labeled substrate. (C) BER analysis of mitochondrial extracts from MLS-lig III-YFP, and (D) MLS-YFP cells.
3.4. Overexpression of DNA ligase III in mitochondria enhances viability of the cells after menadione treatment
Menadione treatment leads to the accumulation of ROS in the mitochondria that attack mtDNA and proteins and cause mitochondrial dysfunction and cell death [32]. To investigate whether MLS-lig III-YFP cells were more resistant to ROS-induced cell death, we exposed the cells to different concentrations of menadione. MLS-lig III-YFP cells showed a significantly higher rate of survival compared with U2OS and MLS-YFP control cells (Fig. 5A). To substantiate these findings, we used colony forming ability as a cytotoxic assay and assessed the sensitivity of the cell lines to menadione treatment. Both U2OS and MLS-YFP control cells showed similar sensitivity to menadione treatment, while MLS-lig III-YFP cells were significantly more resistant to this agent (Fig. 5B). Replacing media glucose with galactose forces cells to use OXPHOS for ATP production [33]. We cultured U2OS cells in media with high glucose (25 mM), low glucose (5 mM), or 10 mM galactose. Cells grew with comparable viability and growth rate in all three media indicating U2OS cells have functional mitochondria and use OXPHOS for ATP production. Next, we examined sensitivity of the cells grown in the galactose containing medium to menadione treatment. Also under these conditions, the cell lines showed a rate of cell survival comparable to that shown in Fig. 5A (Supplementary Fig. S6). These results indicate that U2OS cells rely on mitochondrial OXOPHOS for ATP production and that overexpression of DNA ligase III in mitochondria protects cells against menadione induced oxidative stress.
Fig. 5.
Overexpression of DNA ligase III in mitochondria enhances cell survival following menadione-induced oxidative stress. (A) Sensitivity of the cells to menadione was measured using a WST-1 assay after treatment for 3 h. Cells were allowed to recover for 19 h before WST-1 analysis. (B) Colony forming ability assay. Cells were treated with 10 μM menadione for 3 h. After six days, cells were fixed and stained with crystal violet and the colonies were counted. Colony forming ability of the MLS-lig III-YFP cells following menadione treatment was significantly higher than the control U2OS and MLS-YFP cells. The experiment was performed three times, and each experiment was carried out in triplicate. Data are presented as mean ±SEM. *p < 0.05 one-way analysis of variance (ANOVA) followed by Bonferroni’s post hoc test.
3.5. MLS-lig III-YFP cells show lower rotenone-induced autophagy
Autophagy is a process for clearance of damaged cytosolic components including mitochondria, and may in that regard act as a backup repair mechanism when mtBER fails. Next, we examined whether overexpression of DNA ligase III in mitochondria could affect the extent of autophagy. Inhibition of the mitochondrial electron transport chain complex I with rotenone results in the release of superoxide to the matrix side of the inner membrane (in close proximity to mtDNA), and induces autophagy [34]. We monitored autophagy by Western blotting and by measuring the ratio of conversion of the cytosolic LC3B (LC3B-I, 18 kDa) to the autophagic membrane-bound form of LC3 (LC3B-II, 16 kDa) as described before [27,35,36]. We found that the basal autophagy was similar in MLS-lig III-YFP and MLS-YFP control cells, but rotenone-induced autophagy was significantly reduced in MLS-lig III-YFP cells (Fig. 6). Altogether, these results show that improving mtBER by overexpression of DNA ligase III reduces autophagic signaling during excessive ROS production (which also takes place in damaged mitochondria) and thus links mtDNA damage and repair to removal of dysfunctional mitochondria by autophagy.
Fig. 6.
Rotenone-induced autophagy is lower in MLS-lig III-YFP cells. Western blots of MLS-YFP and MLS-lig III-YFP cells for the autophagosome marker LC3B. Cells were treated with 5 μM rotenone (rot. autophagy inducer) or mock treated with DMSO (<0.1%). The experiment was carried out in triplicate. The ratio of the lower band (LC3B II) to the upper band (LC3B I) was used to calculate the rotenone-induced autophagy activity. Data are presented as mean ± SEM. *p < 0.05 one-way ANOVA with Bonferroni’s post hoc test. Autophagy activity in MLS-lig III-YFP cells treated with rotenone was significantly lower than the control cells exposed to rotenone.
4. Discussion
Mitochondria play a critical role in cellular bioenergetics, and the integrity of mtDNA is essential for proper mitochondrial function. In the past two decades, a number of inherited mitochondrial diseases have been identified that are caused by mutations of mtDNA or of nuclear genes that encode proteins for mtDNA maintenance [8,37]. Acquired, somatic mtDNA mutations have also been implicated in the process of aging and the incidence of age-related diseases [11,38–42]. Most somatic mtDNA mutations are thought to be generated by DNA replication and repair errors followed by clonal expansion of the mutated mtDNA [11,43–45]. BER is the most prominent DNA repair pathway in mitochondria. There is evidence that mtBER activity changes with age [10,46–48], which in part may be related to a decline in the capacity of mitochondria to import BER proteins [48]. Altogether, these studies suggest that alterations in mtBER capacity over time might be a potential source of somatic mtDNA mutations. The oxidative environment of mitochondria makes mtDNA more susceptible to damage than nuclear DNA. Moreover, BER in mitochondria lacks the diversity and complexity of nuclear BER [31,49], which involves many proteins that are not strictly required in BER, but enhance its efficiency and fidelity [50,51]. It appears that hydrogen peroxide-induced DNA lesions persist longer in mtDNA than in nuclear DNA [52], suggesting that mtBER is less efficient than nuclear BER in repairing oxidative DNA lesions and/or mtDNA is more vulnerable to damage than nuclear DNA. However, this might also depend on the type of DNA damage [53], because removal of 8-oxoG (a major product of oxidative base damage) from nuclear DNA was reported to be slower than from mtDNA [54]. Previous studies have investigated the effect of overexpression of DNA glycosylases in mitochondria on mitochondrial function and cell survival with some apparent contradictory results. Thus, whereas in some studies overexpression of OGG1 improved cell survival upon stress [55,56], others found that it resulted in mtDNA deletions, increased intracellular levels of ROS and cell death [57,58]. BER is a multistep process and incomplete BER results in the generation of DNA repair intermediates that can lead to mutations, DNA strand breaks, and disruption of DNA replication and transcription, which can all potentially lead to mitochondrial dysfunction. MtDNA deletions are highly prevalent in neurons of the substantia nigra, the primary site of neurodegeneration in Parkinson disease (PD), compared with other brain areas [59–61]. This is probably because of high ROS generation through the metabolism of dopamine [62]. Excision of adenine opposite 8-oxoG in mtDNA by DNA glycosylase MUTYH in the absence of OGG1 resulted in the accumulation of the SSB BER intermediate, which triggered cell death and neurodegeneration [63,64]. Thus, imbalanced concentration of mtBER proteins can be lethal, especially under chronic and acute stress conditions [64].
Here, we investigated the efficiency of mtBER in mitochondrial extracts from human cell lines U2OS and HeLa and mouse brain, using a covalently closed circular double-strand DNA substrate containing a uracil opposite a guanine at a defined position. We identified DNA ligation to be kinetically less efficient than the preceding mtBER steps. Overexpression of DNA ligase III in U2OS cell mitochondria enhanced mtBER, suggesting that the concentration of DNA ligase III in mitochondria is the rate-determining factor in these cells. Interestingly, the increased concentration of DNA ligase III in mitochondria also improved cell survival after a short exposure time of the cells to menadione, suggesting a functional relationship between mtDNA repair and cell survival upon acute oxidative stress.
Selective removal of abnormal mitochondria by autophagy is an important quality control mechanism that maintains a functional population of mitochondria [65]. Overexpression of the mitochondrial antioxidant manganese-superoxide dismutase (SOD2) reduced rotenone-induced autophagy and conversion of LC3-I to LC3-II [34], indicating that this approach can be used to directly assess the effect of different factors on mitochondrial damage and their removal by autophagy. Our observation that overexpression of DNA ligase III in mitochondria enhances mtBER capacity and reduces autophagy activity in cells treated with autophagy-inducer rotenone suggest that mtDNA damage and repair may play a role in a mitochondrial surveillance system.
In a previous report, antisense-mediated reduction of DNA ligase III expression was shown to result in reduced cellular mtDNA content and increased mtDNA single-strand breaks [66]. In a very recent report, however, cells severely depleted of DNA ligase III maintained normal mtDNA copy number [67], but showed increased mtDNA degradation following H2O2 exposure [67], suggesting a critical role for DNA ligase III in mtBER. Accordingly, our results show that DNA ligase III plays an important role in mtBER.
The baseline autophagy activity of MLS-Lig III-YFP was comparable to that of control cells (Fig. 6. DMSO treated control cells). Moreover, the survival rate of MLS-lig III-YFP and the control cells was comparable at lower menadione concentrations, and was only significantly different at 40 μM and higher concentrations (Fig. 5A). This suggests that the endogenous amount of DNA ligase III in mitochondria was sufficient to repair mtDNA damage generated by menadione at those concentrations. Notably, we found that the exposure of cells to 10 μM menadione for one hour resulted in ~15–20% increase in cellular ROS production (not shown), suggesting that mtBER is somewhat able to repair DNA damage that is generated following acute ROS production.
Altogether, our results suggest that under normal conditions, mtBER is able to maintain genome integrity; however, under conditions that cause abnormal and persistent levels of mitochondrial ROS (e.g. in dysfunctional mitochondria), the level of mtDNA damage may exceed the capacity of mtBER, resulting in the accumulation of BER intermediates and mtDNA instability.
In conclusion, we have shown that DNA ligation is kinetically the “bottleneck” step in mtBER and that overexpression of DNA ligase III in mitochondria improves mtBER and enhances cell survival after oxidative stress. Our results demonstrate a functional link between mtBER, cell survival and autophagy, and provide supporting evidence that an accumulation of mtBER intermediates may lead to the generation of sporadic mitochondrial genome instability in somatic cells. Given the importance of mtDNA maintenance in normal cell and tissue function, our findings may have important implications for understanding the mechanisms underlying human aging and disease.
Supplementary Material
Acknowledgments
We would like to thank Dr. Peter Sykora, Dr. Deborah L. Croteau, Dr. Tadokoro Takashi and Dr. Huiming Lu for critically reading of the manuscript. We thank Dr. Agla Jael Rubner Fridriksdottir for help with FACS cell sorting. Confocal and electron microscopy was carried out at the Core Facility of Integrated Microscopy, University of Copenhagen. This work was funded by the NORDEA Foundation, Denmark.
Appendix A. Supplementary data
Supplementary data associated with this article can be found, in the online version, at http://dx.doi.org/10.1016/j.dnarep.2014.01.015.
Footnotes
Conflict of interest statement
The authors declare no conflict of interest.
References
- 1.Lauritzen KH, Moldestad O, Eide L, Carlsen H, Nesse G, Storm JF, Mansuy IM, Bergersen LH, Klungland A. Mitochondrial DNA toxicity in forebrain neurons causes apoptosis, neurodegeneration, and impaired behavior. Mol Cell Biol. 2010;30:1357–1367. doi: 10.1128/MCB.01149-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.McFarland R, Taylor RW, Turnbull DM. Mitochondrial disease – its impact, etiology, and pathology. Curr Top Dev Biol. 2007;77:113–155. doi: 10.1016/S0070-2153(06)77005-3. [DOI] [PubMed] [Google Scholar]
- 3.Simsek D, Furda A, Gao Y, Artus J, Brunet E, Hadjantonakis AK, Van Houten B, Shuman S, McKinnon PJ, Jasin M. Crucial role for DNA ligase III in mitochondria but not in Xrcc1-dependent repair. Nature. 2011;471:245–248. doi: 10.1038/nature09794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Holt IJ, Harding AE, Morgan-Hughes JA. Deletions of muscle mitochondrial DNA in patients with mitochondrial myopathies. Nature. 1988;331:717–719. doi: 10.1038/331717a0. [DOI] [PubMed] [Google Scholar]
- 5.Zeviani M, Moraes CT, DiMauro S, Nakase H, Bonilla E, Schon EA, Rowland LP. Deletions of mitochondrial DNA in Kearns–Sayre syndrome. Neurology. 1988;38:1339–1346. doi: 10.1212/wnl.38.9.1339. [DOI] [PubMed] [Google Scholar]
- 6.Wallace DC, Singh G, Lott MT, Hodge JA, Schurr TG, Lezza AM, Elsas LJ, EK, Nikoskelainen Mitochondrial DNA mutation associated with Leber’s hereditary optic neuropathy. Science. 1988;242:1427–1430. doi: 10.1126/science.3201231. [DOI] [PubMed] [Google Scholar]
- 7.Copeland WC. Inherited mitochondrial diseases of DNA replication. Annu Rev Med. 2008;59:131–146. doi: 10.1146/annurev.med.59.053006.104646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Stumpf JD, Copeland WC. Mitochondrial DNA replication and disease: insights from DNA polymerase gamma mutations. Cell Mol Life Sci. 2011;68:219–233. doi: 10.1007/s00018-010-0530-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.de Souza-Pinto NC, Wilson DM, 3rd, Stevnsner TV, Bohr VA. Mitochondrial DNA, base excision repair and neurodegeneration. DNA Rep (Amst) 2008;7:1098–1109. doi: 10.1016/j.dnarep.2008.03.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Gredilla R, Bohr VA, Stevnsner T. Mitochondrial DNA repair and association with aging – an update. Exp Gerontol. 2010;45:478–488. doi: 10.1016/j.exger.2010.01.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Taylor RW, Turnbull DM. Mitochondrial DNA mutations in human disease. Nat Rev Genet. 2005;6:389–402. doi: 10.1038/nrg1606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Novo E, Parola M. Redox mechanisms in hepatic chronic wound healing and fibrogenesis. Fibrogen Tissue Rep. 2008;1:5. doi: 10.1186/1755-1536-1-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Orient A, Donko A, Szabo A, Leto TL, Geiszt M. Novel sources of reactive oxygen species in the human body. Nephrol Dial Transplant. 2007;22:1281–1288. doi: 10.1093/ndt/gfm077. [DOI] [PubMed] [Google Scholar]
- 14.Brown WM, George M, Jr, Wilson AC. Rapid evolution of animal mitochondrial DNA. Proc Natl Acad Sci U S A. 1979;76:1967–1971. doi: 10.1073/pnas.76.4.1967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Akbari M, Krokan HE. Cytotoxicity and mutagenicity of endogenous DNA base lesions as potential cause of human aging. Mech Ageing Dev. 2008;129:353–365. doi: 10.1016/j.mad.2008.01.007. [DOI] [PubMed] [Google Scholar]
- 16.Bjelland S, Seeberg E. Mutagenicity, toxicity and repair of DNA base damage induced by oxidation. Mutat Res. 2003;531:37–80. doi: 10.1016/j.mrfmmm.2003.07.002. [DOI] [PubMed] [Google Scholar]
- 17.Akbari M, Visnes T, Krokan HE, Otterlei M. Mitochondrial base excision repair of uracil and AP sites takes place by single-nucleotide insertion and long-patch DNA synthesis. DNA Rep (Amst) 2008;7:605–616. doi: 10.1016/j.dnarep.2008.01.002. [DOI] [PubMed] [Google Scholar]
- 18.Szczesny B, Tann AW, Longley MJ, Copeland WC, Mitra S. Long patch base excision repair in mammalian mitochondrial genomes. J Biol Chem. 2008;283:26349–26356. doi: 10.1074/jbc.M803491200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zheng L, Zhou M, Guo Z, Lu H, Qian L, Dai H, Qiu J, Yakubovskaya E, Bogenhagen DF, Demple B, Shen B. Human DNA2 is a mitochondrial nuclease/helicase for efficient processing of DNA replication and repair intermediates. Mol Cell. 2008;32:325–336. doi: 10.1016/j.molcel.2008.09.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Liu P, Qian L, Sung JS, de Souza-Pinto NC, Zheng L, Bogenhagen DF, Bohr VA, Wilson DM, 3rd, Shen B, Demple B. Removal of oxidative DNA damage via FEN1-dependent long-patch base excision repair in human cell mitochondria. Mol Cell Biol. 2008;28:4975–4987. doi: 10.1128/MCB.00457-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Tann AW, Boldogh I, Meiss G, Qian W, Van Houten B, Mitra S, Szczesny B. Apoptosis induced by persistent single-strand breaks in mitochondrial genome: critical role of EXOG (5′-EXO/endonuclease) in their repair. J Biol Chem. 2011;286:31975–31983. doi: 10.1074/jbc.M110.215715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lakshmipathy U, Campbell C. The human DNA ligase III gene encodes nuclear and mitochondrial proteins. Mol Cell Biol. 1999;19:3869–3876. doi: 10.1128/mcb.19.5.3869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Caldecott KW, McKeown CK, Tucker JD, Ljungquist S, Thompson LH. An interaction between the mammalian DNA repair protein XRCC1 and DNA ligase III. Mol Cell Biol. 1994;14:68–76. doi: 10.1128/mcb.14.1.68. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Caldecott KW, Tucker JD, Stanker LH, Thompson LH. Characterization of the XRCC1-DNA ligase III complex in vitro and its absence from mutant hamster cells. Nucleic Acids Res. 1995;23:4836–4843. doi: 10.1093/nar/23.23.4836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lakshmipathy U, Campbell C. Mitochondrial DNA ligase III function is independent of Xrcc1. Nucleic Acids Res. 2000;28:3880–3886. doi: 10.1093/nar/28.20.3880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gao Y, Katyal S, Lee Y, Zhao J, Rehg JE, Russell HR, McKinnon PJ. DNA ligase III is critical for mtDNA integrity but not Xrcc1-mediated nuclear DNA repair. Nature. 2011;471:240–244. doi: 10.1038/nature09773. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Scheibye-Knudsen M, Ramamoorthy M, Sykora P, Maynard S, Lin PC, Minor RK, Wilson DM, 3rd, Cooper M, Spencer R, de Cabo R, Croteau DL, Bohr VA. Cockayne syndrome group B protein prevents the accumulation of damaged mitochondria by promoting mitochondrial autophagy. J Exp Med. 2012;209:855–869. doi: 10.1084/jem.20111721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Akbari M, Otterlei M, Pena-Diaz J, Aas PA, Kavli B, Liabakk NB, Hagen L, Imai K, Durandy A, Slupphaug G, Krokan HE. Repair of U/G and U/A in DNA by UNG2-associated repair complexes takes place predominantly by short-patch repair both in proliferating and growth-arrested cells. Nucleic Acids Res. 2004;32:5486–5498. doi: 10.1093/nar/gkh872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Frosina G, Cappelli E, Fortini P, Dogliotti E. In vitro base excision repair assay using mammalian cell extracts. Methods Mol Biol. 1999;113:301–315. doi: 10.1385/1-59259-675-4:301. [DOI] [PubMed] [Google Scholar]
- 30.Canugovi C, Maynard S, Bayne AC, Sykora P, Tian J, de Souza-Pinto NC, Croteau DL, Bohr VA. The mitochondrial transcription factor A functions in mitochondrial base excision repair. DNA Rep (Amst) 2010;9:1080–1089. doi: 10.1016/j.dnarep.2010.07.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sykora P, Wilson DM, 3rd, Bohr VA. Repair of persistent strand breaks in the mitochondrial genome. Mech Ageing Dev. 2012;133:169–175. doi: 10.1016/j.mad.2011.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Fukui M, Choi HJ, Zhu BT. Rapid generation of mitochondrial superoxide induces mitochondrion-dependent but caspase-independent cell death in hippocampal neuronal cells that morphologically resembles necroptosis. Toxicol Appl Pharmacol. 2012;262:156–166. doi: 10.1016/j.taap.2012.04.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Marroquin LD, Hynes J, Dykens JA, Jamieson JD, Will Y. Circumventing the Crabtree effect: replacing media glucose with galactose increases susceptibility of HepG2 cells to mitochondrial toxicants. Toxicol Sci. 2007;97:539–547. doi: 10.1093/toxsci/kfm052. [DOI] [PubMed] [Google Scholar]
- 34.Chen Y, McMillan-Ward E, Kong J, Israels SJ, Gibson SB. Mitochondrial electron-transport-chain inhibitors of complexes I and II induce autophagic cell death mediated by reactive oxygen species. J Cell Sci. 2007;120:4155–4166. doi: 10.1242/jcs.011163. [DOI] [PubMed] [Google Scholar]
- 35.Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T, Kominami E, Ohsumi Y, Yoshimori T. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J. 2000;19:5720–5728. doi: 10.1093/emboj/19.21.5720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Levine B, Kroemer G. Autophagy in the pathogenesis of disease. Cell. 2008;132:27–42. doi: 10.1016/j.cell.2007.12.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Schapira AH. Mitochondrial diseases. Lancet. 2012;379:1825–1834. doi: 10.1016/S0140-6736(11)61305-6. [DOI] [PubMed] [Google Scholar]
- 38.Greaves LC, Barron MJ, Campbell-Shiel G, Kirkwood TB, Turnbull DM. Differences in the accumulation of mitochondrial defects with age in mice and humans. Mech Ageing Dev. 2011;132:588–591. doi: 10.1016/j.mad.2011.10.004. [DOI] [PubMed] [Google Scholar]
- 39.Michikawa Y, Mazzucchelli F, Bresolin N, Scarlato G, Attardi G. Aging-dependent large accumulation of point mutations in the human mtDNA control region for replication. Science. 1999;286:774–779. doi: 10.1126/science.286.5440.774. [DOI] [PubMed] [Google Scholar]
- 40.Mootha VK, Lindgren CM, Eriksson KF, Subramanian A, Sihag S, Lehar J, Puigserver P, Carlsson E, Ridderstrale M, Laurila E, Houstis N, Daly MJ, Patterson N, Mesirov JP, Golub TR, Tamayo P, Spiegelman B, Lander ES, Hirschhorn JN, Altshuler D, Groop LC. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003;34:267–273. doi: 10.1038/ng1180. [DOI] [PubMed] [Google Scholar]
- 41.Petersen KF, Befroy D, Dufour S, Dziura J, Ariyan C, Rothman DL, DiPietro L, Cline GW, Shulman GI. Mitochondrial dysfunction in the elderly: possible role in insulin resistance. Science. 2003;300:1140–1142. doi: 10.1126/science.1082889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Polyak K, Li Y, Zhu H, Lengauer C, Willson JK, Markowitz SD, Trush MA, Kinzler KW, Vogelstein B. Somatic mutations of the mitochondrial genome in human colorectal tumours. Nat Genet. 1998;20:291–293. doi: 10.1038/3108. [DOI] [PubMed] [Google Scholar]
- 43.Crespo-Biel N, Theunis C, Van Leuven F. Protein tau: prime cause of synaptic and neuronal degeneration in Alzheimer’s disease. Int J Alzheimers Dis. 2012;2012:251426. doi: 10.1155/2012/251426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Park CB, Larsson NG. Mitochondrial DNA mutations in disease and aging. J Cell Biol. 2011;193:809–818. doi: 10.1083/jcb.201010024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Taylor RW, Barron MJ, Borthwick GM, Gospel A, Chinnery PF, Samuels DC, Taylor GA, Plusa SM, Needham SJ, Greaves LC, Kirkwood TB, Turnbull DM. Mitochondrial DNA mutations in human colonic crypt stem cells. J Clin Invest. 2003;112:1351–1360. doi: 10.1172/JCI19435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Chen D, Cao G, Hastings T, Feng Y, Pei W, O’Horo C, Chen J. Age-dependent decline of DNA repair activity for oxidative lesions in rat brain mitochondria. J Neurochem. 2002;81:1273–1284. doi: 10.1046/j.1471-4159.2002.00916.x. [DOI] [PubMed] [Google Scholar]
- 47.Imam SZ, Karahalil B, Hogue BA, Souza-Pinto NC, Bohr VA. Mitochondrial and nuclear DNA-repair capacity of various brain regions in mouse is altered in an age-dependent manner. Neurobiol Aging. 2006;27:1129–1136. doi: 10.1016/j.neurobiolaging.2005.06.002. [DOI] [PubMed] [Google Scholar]
- 48.Szczesny B, Hazra TK, Papaconstantinou J, Mitra S, Boldogh I. Age-dependent deficiency in import of mitochondrial DNA glycosylases required for repair of oxidatively damaged bases. Proc Natl Acad Sci U S A. 2003;100:10670–10675. doi: 10.1073/pnas.1932854100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Hegde ML, Mantha AK, Hazra TK, Bhakat KK, Mitra S, Szczesny B. Oxidative genome damage and its repair: implications in aging and neurodegenerative diseases. Mech Ageing Dev. 2012;133:157–168. doi: 10.1016/j.mad.2012.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Harrigan JA, Wilson DM, 3rd, Prasad R, Opresko PL, Beck G, May A, Wilson SH, Bohr VA. The Werner syndrome protein operates in base excision repair and cooperates with DNA polymerase beta. Nucleic Acids Res. 2006;34:745–754. doi: 10.1093/nar/gkj475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Tadokoro T, Ramamoorthy M, Popuri V, May A, Tian J, Sykora P, Rybanska I, Wilson DM, 3rd, Croteau DL, Bohr VA. Human RECQL5 participates in the removal of endogenous DNA damage. Mol Biol Cell. 2012;23:4273–4285. doi: 10.1091/mbc.E12-02-0110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Yakes FM, Van Houten B. Mitochondrial DNA damage is more extensive and persists longer than nuclear DNA damage in human cells following oxidative stress. Proc Natl Acad Sci U S A. 1997;94:514–519. doi: 10.1073/pnas.94.2.514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Higuchi Y, Linn S. Purification of all forms of HeLa cell mitochondrial DNA and assessment of damage to it caused by hydrogen peroxide treatment of mitochondria or cells. J Biol Chem. 1995;270:7950–7956. doi: 10.1074/jbc.270.14.7950. [DOI] [PubMed] [Google Scholar]
- 54.Thorslund T, Sunesen M, Bohr VA, Stevnsner T. Repair of 8-oxoG is slower in endogenous nuclear genes than in mitochondrial DNA and is without strand bias. DNA Rep (Amst) 2002;1:261–273. doi: 10.1016/s1568-7864(02)00003-4. [DOI] [PubMed] [Google Scholar]
- 55.Druzhyna NM, Hollensworth SB, Kelley MR, Wilson GL, Ledoux SP. Targeting human 8-oxoguanine glycosylase to mitochondria of oligodendrocytes protects against menadione-induced oxidative stress. Glia. 2003;42:370–378. doi: 10.1002/glia.10230. [DOI] [PubMed] [Google Scholar]
- 56.Wang J, Wang Q, Watson LJ, Jones SP, Epstein PN. Cardiac overexpression of 8-oxoguanine DNA glycosylase 1 protects mitochondrial DNA and reduces cardiac fibrosis following transaortic constriction. Am J Physiol Heart Circ Physiol. 2011;301:H2073–H2080. doi: 10.1152/ajpheart.00157.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Zhang H, Mizumachi T, Carcel-Trullols J, Li L, Naito A, Spencer HJ, Spring PM, Smoller BR, Watson AJ, Margison GP, Higuchi M, Fan CY. Targeting human 8-oxoguanine DNA glycosylase (hOGG1) to mitochondria enhances cisplatin cytotoxicity in hepatoma cells. Carcinogenesis. 2007;28:1629–1637. doi: 10.1093/carcin/bgm072. [DOI] [PubMed] [Google Scholar]
- 58.Zhang H, Xie C, Spencer HJ, Zuo C, Higuchi M, Ranganathan G, Kern PA, Chou MW, Huang Q, Szczesny B, Mitra S, Watson AJ, Margison GP, Fan CY. Obesity and hepatosteatosis in mice with enhanced oxidative DNA damage processing in mitochondria. Am J Pathol. 2011;178:1715–1727. doi: 10.1016/j.ajpath.2010.12.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Bender A, Krishnan KJ, Morris CM, Taylor GA, Reeve AK, Perry RH, Jaros E, Hersheson JS, Betts J, Klopstock T, Taylor RW, Turnbull DM. High levels of mitochondrial DNA deletions in substantia nigra neurons in aging and Parkinson disease. Nat Genet. 2006;38:515–517. doi: 10.1038/ng1769. [DOI] [PubMed] [Google Scholar]
- 60.Kraytsberg Y, Kudryavtseva E, McKee AC, Geula C, Kowall NW, Khrapko K. Mitochondrial DNA deletions are abundant and cause functional impairment in aged human substantia nigra neurons. Nat Genet. 2006;38:518–520. doi: 10.1038/ng1778. [DOI] [PubMed] [Google Scholar]
- 61.Soong NW, Hinton DR, Cortopassi G, Arnheim N. Mosaicism for a specific somatic mitochondrial DNA mutation in adult human brain. Nat Genet. 1992;2:318–323. doi: 10.1038/ng1292-318. [DOI] [PubMed] [Google Scholar]
- 62.Halliwell B. Reactive oxygen species and the central nervous system. J Neurochem. 1992;59:1609–1623. doi: 10.1111/j.1471-4159.1992.tb10990.x. [DOI] [PubMed] [Google Scholar]
- 63.Oka S, Ohno M, Tsuchimoto D, Sakumi K, Furuichi M, Nakabeppu Y. Two distinct pathways of cell death triggered by oxidative damage to nuclear and mitochondrial DNAs. EMBO J. 2008;27:421–432. doi: 10.1038/sj.emboj.7601975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Sheng Z, Oka S, Tsuchimoto D, Abolhassani N, Nomaru H, Sakumi K, Yamada H, Nakabeppu Y. 8-Oxoguanine causes neurodegeneration during MUTYH-mediated DNA base excision repair. J Clin Invest. 2012;122:4344–4361. doi: 10.1172/JCI65053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Wang K, Klionsky DJ. Mitochondria removal by autophagy. Autophagy. 2011;7:297–300. doi: 10.4161/auto.7.3.14502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Lakshmipathy U, Campbell C. Antisense-mediated decrease in DNA ligase III expression results in reduced mitochondrial DNA integrity. Nucleic Acids Res. 2001;29:668–676. doi: 10.1093/nar/29.3.668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Shokolenko IN, Fayzulin RZ, Katyal S, McKinnon PJ, Wilson GL, Alexeyev MF. Mitochondrial DNA ligase is dispensable for the viability of cultured cells but essential for mtDNA maintenance. J Biol Chem. 2013;288:26594–26605. doi: 10.1074/jbc.M113.472977. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.