Abstract
Background
Warfarin is the most commonly used oral anticoagulant for the treatment and prevention of thromboembolic disorders. Pharmacogenomics studies have shown that variants in CYP2C9 and VKORC1 genes are strongly and consistently associated with warfarin dose variability. Although different populations from the Middle East and North Africa (MENA) region may share the same ancestry, it is still unclear how they compare in the genetic and non-genetic factors affecting their warfarin dosing.
Objective
To explore the prevalence of CYP2C9 and VKORC1 variants in MENA, and the effect of these variants along with other non-genetic factors in predicting warfarin dose.
Methods
In this systematic review, we included observational cross sectional and cohort studies that enrolled patients on stable warfarin dose and had the genetics and non-genetics factors associated with mean warfarin dose as the primary outcome. We searched PubMed, Medline, Scopus, PharmGKB, PHGKB, Google scholar and reference lists of relevant reviews.
Results
We identified 17 studies in eight different populations: Iranian, Israeli, Egyptian, Lebanese, Omani, Kuwaiti, Sudanese and Turkish. Most common genetic variant in all populations was the VKORC1 (-1639G>A), with a minor allele frequency ranging from 30% in Egyptians and up to 52% and 56% in Lebanese and Iranian, respectively. Variants in the CYP2C9 were less common, with the highest MAF for CYP2C9*2 among Iranians (27%). Variants in the VKORC1 and CYP2C9 were the most significant predictors of warfarin dose in all populations. Along with other genetic and non-genetic factors, they explained up to 63% of the dose variability in Omani and Israeli patients.
Conclusion
Variants of VKORC1 and CYP2C9 are the strongest predictors of warfarin dose variability among the different populations from MENA. Although many of those populations share the same ancestry and are similar in their warfarin dose predictors, a population specific dosing algorithm is needed for the prospective estimation of warfarin dose.
Introduction
Warfarin is the most widely used oral anticoagulant for the treatment and prevention of thromboembolic manifestations associated with atrial fibrillation, prosthetic heart valves, orthopedic surgery, and history of vascular thrombosis [1, 2]. For example, in the United States alone, around 30 million warfarin prescriptions are dispensed annually, and in the year of 2010, total direct expenditures on warfarin were estimated to be around $600 million [2, 3]. Warfarin is a vitamin K antagonist that mediates its anticoagulant effect through preventing the activation of several vitamin K dependent coagulation factors including:- factors II, VII, IX, and X [4, 5]. Because of its narrow therapeutic index and wide interpatient variability, warfarin therapy requires close monitoring and repeated dose adjustments to achieve and maintain therapeutic anticoagulation effect [6–8]. Studies have repeatedly shown that genetic and non-genetic factors are contributing to the warfarin dose variability [9–11]. The most important genes consistently affecting warfarin dose among different populations are the CYP2C9- a gene coding for cytochrome P450 2C9 enzyme which metabolizes the more potent S enantiomer of warfarin, and VKORC1- a gene coding for the vitamin K epoxide reductase which is an enzyme inhibited by warfarin [9, 10]. Mutations in the gene coding for the CYP4F2, a metabolizing enzyme for vitamin K, have also been shown to contribute to warfarin dose variability but to a lesser extent, and its effect was not consistent among all populations [12, 13]. The Clarification of Optimal Anticoagulation through Genetics (COAG) and the European Pharmacogenetics of Anticoagulant Therapy (EU-PACT) were the largest randomized controlled trials designed to assess the utility of genotype-guided dosing [14, 15]. Despite the negative results from the COAG trial indicating no benefit of genetic-guided dosing, compared to clinical dosing, it showed that the percent time in therapeutic range (PTTR) was significantly lower in blacks in the genetic-guided arm compared to the clinical dosing arm [14]. This is probably due to the fact that blacks may have other less-common genetic variants affecting their warfarin dose that were not well-represented in the genetic-algorithm used in the COAG trial [16]. The EU-PACT study on the other hand, compared pharmacogenetics-based doing versus fixed-dose strategy and was performed in a predominantly white population from Europe [15]. In this study, mean PTTR at 3 months was significantly higher in the genotype-guided group than in the control group [15].
Due to its strategic location and its resources, the Middle East and North Africa (MENA) region was always left at conflict, with different civilizations migrating in and out of its countries. That in turn have put populations of MENA through genetic admixture and racial mixing, creating heavily admixed populations with Asian, Caucasian, Arab and African ancestries. Numerous studies in different countries of MENA have been conducted to estimate the frequencies of the genetic mutations associated with warfarin dosing and its ability along with other non-genetic factors to predict warfarin dose. However, it is still not very clear to what extent the results of these studies are comparable and whether all or some of these results should be pooled together to come-up with a more accurate algorithm. As such, we felt that a summary of the evidence on warfarin pharmacogenetics in this region is needed in-order to identify the gaps and provide future directions. Our goal was to systematically review studies from the MENA region that estimate the impact of genetic and non-genetic factors on warfarin dose requirements.
Study objectives: to systematically review all observational studies that have explored the effect of both genetic and non-genetic factors on warfarin dose variability in the MENA region, estimate the prevalence of the studied genetic variations that are associated with warfarin dose variability, and compare the performance of different genetic-based algorithms in warfarin dose prediction.
Methods
Search Strategy
A systematic comprehensive search for observational studies was applied with no date or language restrictions. The search started on February 2016 and was completed by the end of March 2016.
Database search
PubMed, Scopus, MedLine, PharmGKB (Pharmacogenomics Knowledge Base), and PHGKB (Public Health Genomics Knowledge Base) were searched using different terms with the appropriate Booleans. Below is an example of the search terms and Boolean used to search PubMed. Similar terms were used to search other databases, for Scopus and MedLine results were refined by country.
Combination #1: “warfarin” AND “dose prediction” AND “genetic polymorphism” (In title and abstract).
Combination #2: “warfarin” AND “dosing” AND “genetic mutations” (In title and abstract).
Combination #3: “warfarin” AND “genotype guided” AND “algorithm” (ALL fields).
Combination #4: “warfarin” AND “polymorphism” AND “dose prediction”.
Combination #5: #1 OR #2 OR #3 OR #4.
Other resources
For grey literature, Google Scholar was searched without any language or date limits. Moreover, reference list of all kind of relevant review articles retrieved during our search, were hand searched to identify any potential study.
Study Types
Observational cross sectional or cohort studies were included. Randomized controlled studies, case control, case series, and qualitative studies were excluded from this review.
Participants
Adult patients from MENA region, who have been on warfarin for a sufficient amount of time to have a stable dose and their International Normalized Ratio (INR) within the therapeutic range.
Outcome Measure
Included studies were required to have mean warfarin dose as their primary outcome and to have estimated the variability in the dose explained by genetic and non-genetic factors.
Eligibility Criteria
Studies were considered eligible for this review if they have studied populations of the MENA region, have explored the effect of both genetic and non-genetic factors on warfarin dose, and have developed a dosing model.
Study Selection
Potentially relevant studies were screened first by title and abstract to exclude any irrelevant studies. Then studies were screened by text to exclude any study that does not apply the inclusion criteria. Both former steps were carried by two reviewers (HE and LB), independently. Whenever there was a disagreement, it was resolved by discussion.
Data Extraction and Management
Data were extracted with a pre-specified and piloted extraction tool that was prepared and reviewed by the authors. Abstracted data included: author and year of publication, population studied, sample size, genetic and non-genetic factors explored, mean warfarin dose, mean INR, and main findings.
Quality Assessment
Quality assessment tool for observational cohort and cross sectional studies, adopted from the NIH, National Heart, Lung, and Blood Institute [17] was used to assess the quality of all included studies.
Data Synthesis
All extracted data were summarized and presented descriptively.
Results
Study Selection
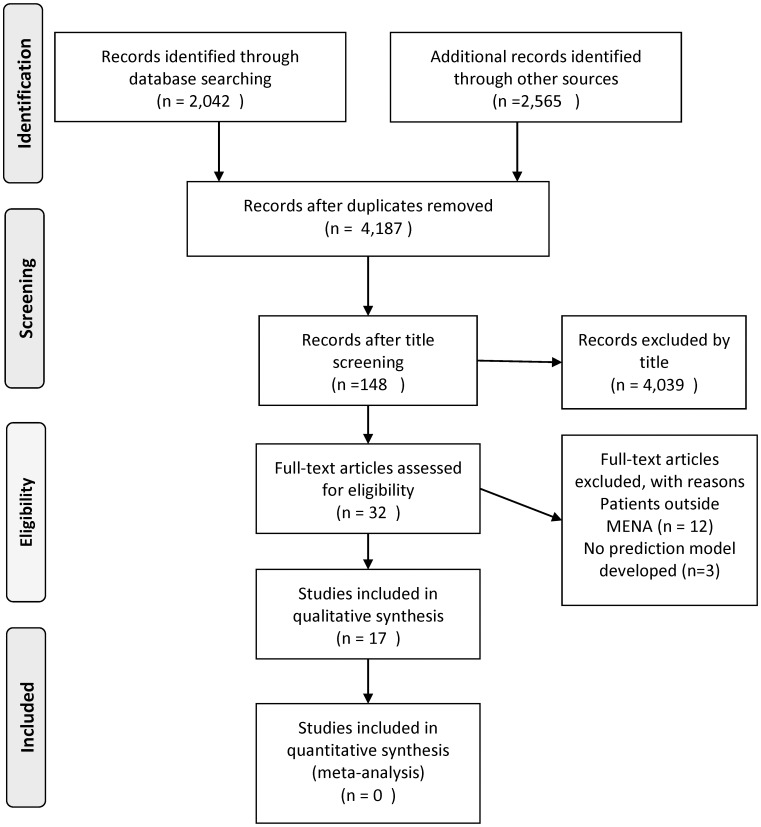
A total of 17 studies were included in this review, including 8 different populations. Fig 1 shows a flow chart of the included studies. Out of 32 studies screened for eligibility, 12 studies were excluded for not targeting patients from the MENA region [7, 9, 10, 18–26] and 3 studies were excluded [27–29] as they did not develop a dosing model. We identified studies from 8 different countries including: Egypt, Iran, Lebanon, Turkey, Sudan, Oman, Kuwait, and two studies in occupied Palestine on Israeli population. Characteristics of all studies are shown in Table 1.
Fig 1. PRISMA Flow Chart of Included Studies.
Table 1. study Characteristics.
| Study | Population | Sample size | Genetic factors explored | Non-genetic factors explored | Mean warfarin dose |
|---|---|---|---|---|---|
| Shahin et. al. 2011, 2013 [12, 30] | Egyptians | 207 | CYP2C9(*2, *3, *4, *5, *8),VKORC1 (3673G>A, Asp36Tyr), APOE, CYP4F2(Val433Met), CALU | Sex, age, BSA, use of aspirin, indication for warfarin, concomitant disease & smoking status | 36.8 ± 17.9 mg/wk |
| Bazan et. al. 2014 [31] | Egyptians | 63 | CYP2C9(*2, *3), VKORC1 | Age & smoking status | 7.3 ± 5.2 (1–30) mg/day |
| Ghozlan et. al. 2015 [32] | Egyptians with ACS | 80 | CYP2C9, VKORC1 | Age & height | 4.8 ± 1.96 (2–10) mg/day |
| Issac et. al. 2014 & Ekladious et. al. 2013 [33, 34] | Egyptian | 84 (50 for model, 34 for validation) | CYP2C9(1075A>C), VKORC1(1173C>T), R1, (C3435T), EPHX1(H139R), PZ(A-13G) | Age & gender | |
| Namazi et. al. 2010 [35] | Iranian | 100 total (100 CYP2C9, 99 CYP2C19, 81 VKORC1) 55 for the model | CYP2C9 (*2, *3), VKORC1 (-1639G>A), CYP2C19 (*2, *3) | Gender, age, BSA, weight & height | 7.3 ± 5.2 (1–30) mg/day |
| Loebstein et. al. 2005 [36, 37] | Israeli | 100 | GGCX, CALU, VKORC1, EPHX1, CYP2C9 (*2, *3) | Age, weight, concurrent medication & total vit.K plasma concentration | 5.7 ± 3.3 (1.1–20) mg/day |
| Alrashid et. al. 2016 [38] | Kuwaiti | 108 | CYP2C9, VKORC1 | Sex & BMI, age were adjusted for in the model but not included as predictors | 4.7 ± 2.7 mg/day |
| Esmerian et. al. 2011 [39] | Lebanese | 43 | CYP2C9 (*2, *3), VKORC1 (-1639G>A, 1173 C>T) | N/A | 31 ± 14 mg/wk |
| Pathare et. al. 2012 [40] | Omani | 212 (142 in the model derivation cohort, 70 for validation) | CYP2C9 (*2, *3), CYP4F2 *3, VKORC1 (3673G>A, 5808T>G, 6009C>T, 6484C>T, 9041G>A) | Simvastatin, amiodarone, hypertension, diabetes, atrial fibrillation, deep vein thrombosis, mechanical valve, age, weight & gender | 4.75 (3–5.5) mg/day |
| Shrif et. al. 2011 [41] | Sudanese | 203 patients 180 healthy volunteers | CYP2C9 (*2, *3, *5, *6, *8, *9, *11), VKORC1 (20 tagSNPs) | Body weight, concurrent medication, target INR, body surface area, height, age, indication for warfarin treatment & gender | 5.58 ± 2.48 (1.5–22.5) mg/day |
| Özer et. al. 2013 [42] | Turkish | 107 | CYP2C9 (*1,*2), VKORC1(-1639G>A, 1173C>T), CYP4F2, EPHX1 | Age, height, weight, No. of cigarettes & daily consumed tea and green vegetables | 5.16 ± 1.95 (1.43–10) mg/day |
| Ozgon et. al. 2008 [43] | Turkish | 205 | CYP2C9 (*2,*3,*4,*5), VKORC1 (-1639G>A) | Age & non-indication of VT | 34.2 ± 16.78 (6.25–125) mg/wk |
| Yildirim et. al. 2014 [44] | Turkish | 101 | CYP2C9 (*2,*3), VKORC1(-1639G>A), factor VII (-401G>T) | Age, BMI & INR | 4.07 ± 1.6(1.13–7.86) mg/day |
| Ozer et. al. 2010 [45] | Turkish | 100 | CYP2C9(*2,*3), VKORC1(-1639G>A) | Age & BSA | 4.11 (1.16–9.33) mg/day |
After assessing for quality, most studies were found to be ranging from fair to good quality. However, the study by Alrashid et al [38] was considered of poor quality as there was no reporting of the r2 value for the dosing model. The sample-size used in some of the studies was less than 100, which is relatively low and may have affected the power of their conclusions [31, 32, 34, 39]. Testing for the Hardy-Weinberg equilibrium was reported in all studies except those by Yildirim et al [44] and Ghozlan et al [32].
Targeted populations were required to be on stable warfarin dose as part of the inclusion criteria. Due to the variability in the definition of stable warfarin dose among studies, we reported the different definitions used by authors of each included study (Table 2). Overall, having an INR within therapeutic range for three consecutive visits while on the same warfarin dose was considered “stable warfarin dose” description by almost all studies.
Table 2. Stable Warfarin Dose Definition Variation Among Studies.
| Studies | Definition |
|---|---|
| Shrif et. al. 2013 [41] | Mean of the daily warfarin doses at which INR measurements were within target therapeutic levels for three consecutive clinic visits over more than 3 months |
| Namazi et. al. 2010 [35] | Warfarin dose that was constant for ≥ 3 consecutive visits over a minimum period of 3 months with INR value variation ≤ 15% |
| Esmerian et. al. 2011 [39] | Therapy for at least 2 months, same weekly dose of warfarin over the past 3 INR examinations, patients considered adequately coagulated if the INR value at recruitment fell within a range of 1.7–4 |
| Pathare et. al. 2012 [40] | A patient was considered to have a stable INR when his/her INR was between 2 and 3 on at least 3 consecutive assessments, 3 months after initiating the therapy |
| Alrashid et. al. 2016 [38] | Patients were defined as those whose warfarin dose requirement has remained constant for at least three consecutive clinic visits, at which point the INR was within the therapeutic range |
| Bazan et. al. 2014 [31] | Patients having a stable warfarin dose requirement for at least 3 consecutive times with dose titration to an INR target range of 2–3.5 |
| Shahin et. al. 2011, 2013 [12] | A dose that did not vary by more than 10% between clinic visit, for three consecutive visits, occurring over a minimum time-period of 2 months |
| Ghozlan et. al. 2015 [32] | Steady-state dose that leads to stable anticoagulation levels in three consecutive clinic visits for which the INR measurements are within the range of 2–3 |
| Yildirim et. el. 2014 [44] | Not clearly defined |
| Özer et. al. 2013 [42] | Patients were included if they had been on therapy for > 4 months and their last three INR measurements were within therapeutic range for the same mean daily dose |
| Ozgon et. al. 2008 [43] | Patients were included if they had been on therapy >2 months and their last three INR measurements were considered stable by their doctors, whether or not they correspond to their target INR |
| Loebstein et. al. 2005 [37] | Having therapeutic INR over 4 consecutive visit and receiving the same daily dose of warfarin before sample collection on index visit |
| Issac et. al. 2014 & Ekladious et. al. 2013 [33, 34] | Having at least three consecutive INRs in the therapeutic (2–3) range for the same daily maintenance dose after at least 3 months of therapy |
| Ozer et. al. 2010 [45] | Three consecutive clinic visits for which INR measurements were within therapeutic range for the same daily dose |
Prevalence of the studied genetic variants
In-order to estimate the prevalence of different genetic variants we reported the minor allele frequency (MAF) of these variants. For the most common genetic variants in VKORC1 and CYP2C9, MAF’s are presented in Table 3. Variants in the VKORC1 gene had the highest frequencies among all populations followed by the *2 variant of CYP2C9 gene and *3, with *2 being relatively higher in most of the populations.
Table 3. Minor Allele Frequency (MAF) of Most Common Genetic Variants.
| Population | Gene | VKORC1 | CYP2C9 | |
|---|---|---|---|---|
| Variant | -1639 G>A (rs9923231)* | *2(rs179853) | *3(rs1057910) | |
| Egyptians [12, 30] | 46.2% | 11.7 | 9.2% | |
| Egyptians [34] | 72.05% | N/A | 10.7% | |
| Egyptians [31] | 51% | 7% | 9.6% | |
| Egyptians [32] | 30% | 8% | 4.3% | |
| Iranian [35] | 56% | 27% | 9% | |
| Israeli [36] | N/A | 12.5% | 11% | |
| Kuwaiti [38] | 40% | 14% | 5% | |
| Lebanese [39] | 52% | 15% | 7% | |
| Omani [40] | 35% | 6% | 6% | |
| Sudanese [41] | 37% | 5% | 0% | |
| Turkish [43] | 50% | 13% | 10% | |
| Turkish [42] | 49% | N/A | N/A | |
| Turkish [45] | 40% | 13% | 15% | |
| Turkish [44] | 51% | 17% | 27% | |
* This rs ID refers to VKORC1 (-1639G>A) and VKORC1(3673G>A)
Other less common genetic variants have been explored in Egyptians, Israeli, Omani, and Turkish populations. These include variants in the following genes: CYP4F2, EPHX1, GGCX1, MDR1, factor VII, APOE, CALU and PZ. The MAF for the CYP4F2 (rs2108622) was 40% among Turkish [42], 30% in Omani [40] and 42% in Egyptians [12]. The EPHX1 encodes for a putative subunit of the vitamin K epoxide reductase complex called: microsomal epoxide hydrolase 1 [10]. The MAF of EPHX1 (rs2292566) was 16% and 26.19% in Turkish and Egyptians, respectively [33, 42], while another variant the (rs1051740) was 25% in Israeli [36, 37]. The multidrug resistance gene (MDR1 C3435T) and Protein Z A-13G were only studied in Egyptians and their MAF were 42.86% and 0%, respectively [33]. Other gene variants that are associated with warfarin dose variability are the APOE and CALU [10]. The rs429358 and rs7412 variants of the APOE gene had a MAF of 6.7% and 7.4%, in Egyptians, respectively [12]. On the other hand, the rs339097 variant of the CALU gene was less common in Egyptians than the APOE variants with a MAF of 2.3% [12]. Other rare genetic variants that have been explored are the Factor VII (-401G>A) which had a MAF of 35% in Turkish [42], and the GGCX (rs699664) which had a MAF of 29.5% in Israeli [36].
Significant Predictors of Warfarin Dose
Using univariate and multivariate linear regression, authors of all studies were able to identify the most significant predictors of warfarin dose and the most significant dosing model which explained the highest percent of the dose variability. Table 4 shows the significant predictors in the different populations and to what extent were they able to predict warfarin dose.
Table 4. Most Significant Predictors and % Variability Explained.
| Population | Significant Genetic Predictors | Significant Non-Genetic Predictors | Variability explained by the model |
|---|---|---|---|
| Egyptians [12, 30] |
|
|
36.5% |
| Egyptians [32] |
|
|
30.6% |
| Egyptians [31] |
|
|
43.4% |
| Egyptians [33, 34] |
|
|
20.9% |
| Iranian [35] |
|
|
41.3% |
| Israeli [37] |
|
|
63% |
| Kuwaiti [38] |
|
|
N/A |
| Lebanese [39] |
|
N/A | 33.9% |
| Omani [40] |
|
|
63% |
| Sudanese [41] |
|
|
36.75 |
| Turkish [43] |
|
|
43% |
| Turkish [45] |
|
|
60.4% |
| Turkish [42] |
|
|
39.3% |
| Turkish [44] |
|
18.2% |
Discussion
Summary of Evidence
The term MENA refers to countries from Morocco to Iran and down to Sudan. Studies of maternally inherited mitochondrial DNA have suggested that all modern populations, including those of MENA [46–48], have originated through a single wave from Sub Saharan Africa and populated other parts of the world [49]. Later-on, several historical events took place and facilitated genetic migration like the early Islamic conquests, the Ottoman Empire expansion and the continuous migrations in and out of the Arabian Peninsula and Europe. Other examples include the silk and spice road, which used to connect China with Europe through the Middle East [50]. Countries like Egypt and Oman, their strategic crossroad position between Africa and Eurasia have left them with high intrapopulation diversity [46]. On the other hand, other countries like Qatar and Yemen have the highest rates of consanguinity, 44.7% and 54%, respectively, leaving them with less genetic variability [51]. All of the aforementioned reasons have made these countries of particular interest in genetic studies.
Pharmacogenetics studies have shown that variants of VKORC1 and CYP2C9 are strongly associated with warfarin dose. VKORC1(-1639G>A) alone can explain up to 40% of the dose variation in some populations [52]. Along with these genetic predictors, other genetic factors and non-genetic factors such as age, weight, gender, race, ethnicity, concurrent medication, and comorbidities can boost the preemptive estimation of warfarin dosing to more than 50% [10].
The fact that VKORC1 variants are the most significant predictors of warfarin dose followed by those of CYP2C9, was consistent across all studies included in this systematic review. However, the frequency of these genetic variations and the extent of their effect on dose variability varied from population to another. Similarities were seen between Sudanese and African Americans in that CYP2C9*5, *6, and *11 were better predictors of warfarin dose than the *2 and *3 variants [53]. Moreover, the variability explained by variants of VKORC1 (rs9934438, rs9923231, rs8050894) were comparable to those of African Americans [1, 53]. CYP2C9*8 (rs9332094) is another rare variant that affects around 12% of African Americans and has been associated with reduced S-warfarin clearance [54]. This SNP was only investigated in two populations of MENA; Egyptians and Sudanese [12, 41]. Despite some of the similarities found between these two populations and African Americans, the CYP2C9*8 did not shown any significant association with warfarin dose requirements in Egyptians and Sudanese as it did in African Americans. This might be in part due the fact that this variant was not very common in these populations, it showed a MAF of 0.008 and 0.01 in Egyptians and Sudanese, respectively [12, 41]. On the other hand, the frequencies of CYP2C9 *2 and *3 and VKORC1 (3673G>A) reported in Turkish [43] were closer to Caucasians than to Asians and African Americans [1].
Variants of the VKORC1 gene were the most prevalent among all populations. Great discrepancy was seen in the frequency of the VKORC1 in Egyptians, it ranged from 30% in the study be Ghozlan et. al and up to 72% in the study by Ekladious et. al. [32]. Such discrepancy could be in part due to the small sample sizes used in the different studies concerning the Egyptian population, in particular the studies by Ghozlan, Ekladious, and Bazan and their colleagues [31, 32, 34]. Moreover, the Egyptian population is a heavily admixed population with different ethnic origins [12] which may have led to different frequencies depending on the ethnicity of the studied population. However, none of these studies have reported the different ethnic origins of their studied population and whether it was a contributing factor or not. Carriers of the -1639G>A variant required lower warfarin dose compared to the wild type in all studies. Shrif and colleagues were able to identify four different SNPs downstream from the VKORC1 gene (rs7199949, rs11865038, rs11864839, and rs750952). These SNPs were found to be located in newly identified gene which is only 2 kb downstream from the VKORC1 gene in the same reverse strand named: POL3S (polyserase 3S) [41]. Amino acid sequences of the POL3S are highly conserved among humans, chimpanzees, dogs, cows, and mice [55]. While function of the encoded protein is still unknown in-vivo, it has shown to degrade the alpha chain of fibrinogen as well as pro-urokinase type plasminogen in vitro, indicating its potential role in the coagulation process [55]. Out of the identified SNPs in the POL3S, rs7199949 (p.Pro406Ala) proved to be a significant predictor of warfarin dose and its addition to the regression model explained 7% of the variability [41].
Variants in the CYP2C9 were less common than those of the VKORC1 in all populations with the *2 variant being slightly more common than the *3 variant in majority of cases. The *2 variant was highest among Iranian (27%) [35], while the *3 variant was highest among Turkish (27%) and it was not detected in Sudanese [41, 44]. MAF of *2 and *3 were closely similar among Egyptians, Israeli, and Turkish [30, 36, 43]. Both *2 and *3 variants were associated with lower warfarin dose requirements in all populations.
The VKORC1 (-1639G>A) variant was the most significant predictor of warfarin dose in all population. This variant alone was able to explain 45% of the dose variability in Omani [40]. The VKORC1 Asp36Tyr was significantly associated with higher warfarin doses in Egyptians and when added to Shahin et al. dosing model it further explained the variability by only 2% [30].
CYP4F2 is the gene coding for the cytochrome P450 4F2 enzyme, which is a vitamin K oxidase. The V433M allele is associated with reduced metabolism of vitamin K1 and higher warfarin doses [56]. Caldwell et al. has shown that CYP4F2 V433M (rs2108622) variant has an impact on warfarin dose and that carriers of at least one minor allele required 4–12% higher doses than those with homozygous wild type genotype [57]. Only 3 studies in three different populations of MENA -Egyptian, Omani, and Turkish- have investigated the association between warfarin dose and that gene variant. These studies have shown no significant association between CYP4F2 gene variant and warfarin dose in Egyptians neither in Omani, despite its high frequency [12, 40]. A significant association between CYP4F2 gene variant and warfarin dose was among Turkish by Özer et. al [42]. Furthermore, Turkish carriers of the variant allele required higher warfarin doses than those with the wildtype. The APOE gene encodes for an apolipoprotein that is involved in the hepatic clearance of vitamin K [10]. In the Egyptian cohort studied by Shahin. et. al., carriers of the APOE ε2 gene variant required lower warfarin dose compared to the wild type [12]. Moreover, that variant explained 2.53% of the dose variability. No association was found between MDR1 C3435T, EPHX1 H139R, GGCX and protein Z A-13G gene variations and warfarin dose in populations of MENA. However, when combined, these genetic variants were able to explain 3% of the dose variability in Egyptians [33]. Calumenin (CALU) is a regulator of vitamin K epoxide reductase (VKOR) as identified in studies of warfarin resistance rat and it have been linked to warfarin sensitivity [58]. Shahin et al. was able to confirm the association of CALU rs339097 gene polymorphism with higher doses of warfarin, however, when added to the regression model it failed to show any significance [12]. The authors reported that failing to show significance may have been due to lack of power, since only nine variant carriers were found in their cohort. In African Americans, it was shown, that for every CALU rs339097 variant allele warfarin dose increases by almost 11% [58]. Factor VII is a vitamin K dependent factor, factor VII -401G>T variant was able to predict only 2.2% of warfarin dose variability among Turkish, even though it had an MAF of 35% [44]. Carriers of the minor allele required lower warfarin dose as opposed to the carriers of the major allele [44].
After developing their dosing algorithm, Pathare et al. compared the performance of their algorithm with that of the International Warfarin Pharmacogenetics Consortium (IWPC) in predicting warfarin dose in their Omani cohort [40]. The IWPC algorithm was able to explain only 33.6% of the variability, while Pathare et al. algorithm explained 63% of the variability [40]. Moreover, warfarin dose predicted by the IWPC algorithm was 13% higher than the patients’ actual dose [40]. Even though the IWPC algorithm has been developed and validated using data from different racial groups including Asians, Blacks, and Whites across four different countries, and even though the Omani population comes from Asian, African and Caucasian ancestries, the IWPC algorithm did not perform well in the Omani population [40]. Such variation in the algorithm performance could be owed to the fluctuation in racial definitions, clinical variations, social/lifestyle contexts, population migration and historical perspectives [59]. Such observation highlights the need for dosing algorithm that are more population-tailored.
Beside the genetic variants other clinical and demographic variables were shown to be significantly associated with warfarin dose. Age, weight, gender, body mass index and body surface area were the most significant predictors of warfarin dose and added up to the variability explained.
Limitations
Some of the limitations of this review were having only two reviewers carrying the whole process independently which may have led us to overlook some important findings or observations. Another limitation is that we only considered mean warfarin dose as our primary outcome, while other outcomes like Time to Therapeutic Range (TTR) and risk of bleeding might be predicted by genetic factors. Furthermore, SNPs like the GGCX, Factor VII, CALU, APOE, MDR1, EPHX1, and PZ were investigated in limited number of studies, which made it unfeasible for us to compare their effect on warfarin dose between different populations of MENA.
Conclusions and Future Directions
VKORC1 & CYP2C9 were the most useful variants in predicting warfarin dose. Despite the shared significant predictors in all dosing models, their performance still varied largely between populations. As such, micro-geographically defined population specific dosing algorithms are needed. Warfarin pharmacogenetics have been explored in limited number of populations in MENA region. Thus, more pharmacogenetics studies on other populations are needed to further explain the impact of genetic factors on warfarin dose in the region. Future clinical utility and cost effectiveness studies will confirm if warfarin genetic dosing strategy should replace clinical dosing.
Data Availability
All relevant data are freely available and are within the paper.
Funding Statement
This research was funded by the student grant provided by College of Pharmacy, Qatar University (QUST-CPH-SPR-15/16-8, http://www.qu.edu.qa/offices/research/academic/internal-fund.php), to HE. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Anderson JL, Horne BD, Stevens SM, Grove AS, Barton S, Nicholas ZP, et al. Randomized trial of genotype-guided versus standard warfarin dosing in patients initiating oral anticoagulation. Circulation. 2007. November 27;116(22):2563–70. 10.1161/CIRCULATIONAHA.107.737312 [DOI] [PubMed] [Google Scholar]
- 2.Kirley K, Qato DM, Kornfield R, Stafford RS, Alexander GC. National trends in oral anticoagulant use in the United States, 2007 to 2011. Circ Cardiovasc Qual Outcomes. 2012. September 1;5(5):615–21. 10.1161/CIRCOUTCOMES.112.967299 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Wysowski DK, Nourjah P, Swartz L. Bleeding complications with warfarin use: a prevalent adverse effect resulting in regulatory action. Archives of internal medicine. 2007. July 9;167(13):1414–9. 10.1001/archinte.167.13.1414 [DOI] [PubMed] [Google Scholar]
- 4.Thomas DD. In: Thomas DD, editor. Hemostasis and Thrombosis: Springer International Publishing Switzerland; 2015; 2014. p. 1–7. [Google Scholar]
- 5.In: Offermanns S, Rosenthal W, editors. Encyclopedia of Molecular Pharmacology. 2nd ed Berlin, Heidelberg: Springer; 2008. p. 948–50. [Google Scholar]
- 6.Landefeld CS, Beyth RJ. Anticoagulant-related bleeding: clinical epidemiology, prediction, and prevention. Am J Med. 1993. September;95(3):315–28. [DOI] [PubMed] [Google Scholar]
- 7.Klein TE, Altman RB, Eriksson N, Gage BF, Kimmel SE, Lee MT, et al. Estimation of the warfarin dose with clinical and pharmacogenetic data. N Engl J Med. 2009. February 19;360(8):753–64. 10.1056/NEJMoa0809329 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Johnson JA. Warfarin: an old drug but still interesting. Pharmacotherapy. 2008. September;28(9):1081–3. 10.1592/phco.28.9.1081 [DOI] [PubMed] [Google Scholar]
- 9.Kamali F, Khan TI, King BP, Frearson R, Kesteven P, Wood P, et al. Contribution of age, body size, and CYP2C9 genotype to anticoagulant response to warfarin. Clin Pharmacol Ther. 2004. March;75(3):204–12. 10.1016/j.clpt.2003.10.001 [DOI] [PubMed] [Google Scholar]
- 10.Wadelius M, Chen LY, Eriksson N, Bumpstead S, Ghori J, Wadelius C, et al. Association of warfarin dose with genes involved in its action and metabolism. Hum Genet. 2007. March;121(1):23–34. 10.1007/s00439-006-0260-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Yuan HY, Chen JJ, Lee MT, Wung JC, Chen YF, Charng MJ, et al. A novel functional VKORC1 promoter polymorphism is associated with inter-individual and inter-ethnic differences in warfarin sensitivity. Hum Mol Genet. 2005. July 1;14(13):1745–51. 10.1093/hmg/ddi180 [DOI] [PubMed] [Google Scholar]
- 12.Shahin MH, Khalifa SI, Gong Y, Hammad LN, Sallam MT, El Shafey M, et al. Genetic and nongenetic factors associated with warfarin dose requirements in Egyptian patients. Pharmacogenet Genomics. 2011. March;21(3):130–5. 10.1097/FPC.0b013e3283436b86 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Takeuchi F, McGinnis R, Bourgeois S, Barnes C, Eriksson N, Soranzo N, et al. A genome-wide association study confirms VKORC1, CYP2C9, and CYP4F2 as principal genetic determinants of warfarin dose. PLoS Genet. 2009. March;5(3):e1000433 10.1371/journal.pgen.1000433 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kimmel SE, French B, Kasner SE, Johnson JA, Anderson JL, Gage BF, et al. A pharmacogenetic versus a clinical algorithm for warfarin dosing. N Engl J Med. 2013. December 12;369(24):2283–93. 10.1056/NEJMoa1310669 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Pirmohamed M, Burnside G, Eriksson N, Jorgensen AL, Toh CH, Nicholson T, et al. A randomized trial of genotype-guided dosing of warfarin. N Engl J Med. 2013. December 12;369(24):2294–303. 10.1056/NEJMoa1311386 [DOI] [PubMed] [Google Scholar]
- 16.Johnson JA, Cavallari LH. Warfarin pharmacogenetics. Trends Cardiovasc Med. 2015. January;25(1):33–41. 10.1016/j.tcm.2014.09.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Health NIo. Quality Assessment Tool for Observatioanl Cohort and Cross-Sectional Studies. In: Natioanl Heart L, and Blood Institute, editor.2014.
- 18.Carlquist JF, Horne BD, Mower C, Park J, Huntinghouse J, McKinney JT, et al. An evaluation of nine genetic variants related to metabolism and mechanism of action of warfarin as applied to stable dose prediction. J Thromb Thrombolysis. 2010. October;30(3):358–64. 10.1007/s11239-010-0467-3 [DOI] [PubMed] [Google Scholar]
- 19.Caldwell MD, Berg RL, Zhang KQ, Glurich I, Schmelzer JR, Yale SH, et al. Evaluation of genetic factors for warfarin dose prediction. Clin Med Res. 2007. March;5(1):8–16. 10.3121/cmr.2007.724 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Carlquist JF, Horne BD, Muhlestein JB, Lappe DL, Whiting BM, Kolek MJ, et al. Genotypes of the cytochrome p450 isoform, CYP2C9, and the vitamin K epoxide reductase complex subunit 1 conjointly determine stable warfarin dose: a prospective study. J Thromb Thrombolysis. 2006. December;22(3):191–7. 10.1007/s11239-006-9030-7 [DOI] [PubMed] [Google Scholar]
- 21.Gage BF, Eby C, Milligan PE, Banet GA, Duncan JR, McLeod HL. Use of pharmacogenetics and clinical factors to predict the maintenance dose of warfarin. Thromb Haemost. 2004. January;91(1):87–94. 10.1160/TH03-06-0379 [DOI] [PubMed] [Google Scholar]
- 22.Huang SW, Xiang DK, Wu HL, Chen BL, An BQ, Li GF. [Impact of five genetic polymorphisms on inter-individual variation in warfarin maintenance dose]. Zhonghua Yi Xue Yi Chuan Xue Za Zhi. 2011. December;28(6):661–5. 10.3760/cma.j.issn.1003-9406.2011.06.014 [DOI] [PubMed] [Google Scholar]
- 23.Zhu Y, Shennan M, Reynolds KK, Johnson NA, Herrnberger MR, Valdes R Jr., et al. Estimation of warfarin maintenance dose based on VKORC1 (-1639 G>A) and CYP2C9 genotypes. Clin Chem. 2007. July;53(7):1199–205. 10.1373/clinchem.2006.078139 [DOI] [PubMed] [Google Scholar]
- 24.Wadelius M, Chen LY, Downes K, Ghori J, Hunt S, Eriksson N, et al. Common VKORC1 and GGCX polymorphisms associated with warfarin dose. Pharmacogenomics J. 2005;5(4):262–70. 10.1038/sj.tpj.6500313 [DOI] [PubMed] [Google Scholar]
- 25.Santos PC, Dinardo CL, Schettert IT, Soares RA, Kawabata-Yoshihara L, Bensenor IM, et al. CYP2C9 and VKORC1 polymorphisms influence warfarin dose variability in patients on long-term anticoagulation. Eur J Clin Pharmacol. 2012. April;69(4):789–97. 10.1007/s00228-012-1404-5 [DOI] [PubMed] [Google Scholar]
- 26.Aquilante CL, Langaee TY, Lopez LM, Yarandi HN, Tromberg JS, Mohuczy D, et al. Influence of coagulation factor, vitamin K epoxide reductase complex subunit 1, and cytochrome P450 2C9 gene polymorphisms on warfarin dose requirements. Clin Pharmacol Ther. 2006. April;79(4):291–302. 10.1016/j.clpt.2005.11.011 [DOI] [PubMed] [Google Scholar]
- 27.Pathar A, Alkindi S, Al Zadjali S, Misquith R, Wasim F, Berbar T, et al. Combined effect of CYP2C9 and VKORC1 polymorphisms on warfarin maintance dose in Omani patients. Open Journal of Genetics; 2012. [Google Scholar]
- 28.Yildirim R, Gundogdu M, Kurnaz F, Yildirim A, Aksoy H, Erdem F, et al. CYP2C9 gene polymorphisms and warfarin dose requirement: a single-center experience in Turkey. Turkish Journal of Medical Sciences. 2012. December;42(6):981–6. [Google Scholar]
- 29.Yildirim R, Yildirim A, Laloglu E, Cadirci K, Erdem F, Kiki I, et al. Effect of VKORC1-1639 G > A polymorphism on warfarin response in east of Turkey. Turkish Journal of Biochemistry-Turk Biyokimya Dergisi. 2015 2015;40(4):271–4. [Google Scholar]
- 30.Shahin MH, Cavallari LH, Perera MA, Khalifa SI, Misher A, Langaee T, et al. VKORC1 Asp36Tyr geographic distribution and its impact on warfarin dose requirements in Egyptians. Thromb Haemost. 2013. June;109(6):1045–50. 10.1160/TH12-10-0789 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Bazan N, Sabry N, Rizk A, Mokhtar S, Badary O. Factors affecting warfarin dose requirements and quality of anticoagulation in adult Egyptian patients: role of gene polymorphism. Irish Journal of Medical Science. 2014;183(2):161–72. 10.1007/s11845-013-0978-y [DOI] [PubMed] [Google Scholar]
- 32.Ghozlan MF, Foad DA, Darwish YW, Saad AA. Impact of CYP2C9 and VKORC1 genetic polymorphisms upon warfarin dose requirements in Egyptian patients with acute coronary syndrome. Blood Coagulation and Fibrinolysis. [Article]. 2015;26(5):499–504. 10.1097/MBC.0000000000000272 [DOI] [PubMed] [Google Scholar]
- 33.Issac MS, El-Nahid MS, Wissa MY. Is there a role for MDR1, EPHX1 and protein Z gene variants in modulation of warfarin dosage? a study on a cohort of the Egyptian population. Mol Diagn Ther. 2014. February;18(1):73–83. 10.1007/s40291-013-0055-2 [DOI] [PubMed] [Google Scholar]
- 34.Ekladious SM, Issac MS, El-Atty Sharaf SA, Abou-Youssef HS. Validation of a proposed warfarin dosing algorithm based on the genetic make-up of Egyptian patients. Mol Diagn Ther. 2013. December;17(6):381–90. 10.1007/s40291-013-0046-3 [DOI] [PubMed] [Google Scholar]
- 35.Namazi S, Azarpira N, Hendijani F, Khorshid MB, Vessal G, Mehdipour AR. The impact of genetic polymorphisms and patient characteristics on warfarin dose requirements: A cross-sectional study in Iran. Clinical Therapeutics. [Article]. 2010;32(6):1050–60. 10.1016/j.clinthera.2010.06.010 [DOI] [PubMed] [Google Scholar]
- 36.Vecsler M, Loebstein R, Almog S, Kurnik D, Goldman B, Halkin H, et al. Combined genetic profiles of components and regulators of the vitamin K-dependent γ-carboxylation system affect individual sensitivity to warfarin. Thrombosis and Haemostasis. [Article]. 2006;95(2):205–11. 10.1160/TH05-06-0446 [DOI] [PubMed] [Google Scholar]
- 37.Loebstein R, Vecsler M, Kurnik D, Austerweil N, Gak E, Halkin H, et al. Common genetic variants of microsomal epoxide hydrolase affect warfarin dose requirements beyond the effect of cytochrome P450 2C9. Clinical Pharmacology and Therapeutics. [Article]. 2005;77(5):365–72. 10.1016/j.clpt.2005.01.010 [DOI] [PubMed] [Google Scholar]
- 38.Alrashid MH, Al-Serri A, Alshemmari SH, Koshi P, Al-Bustan SA. Association of Genetic Polymorphisms in the VKORC1 and CYP2C9 Genes with Warfarin Dosage in a Group of Kuwaiti Individuals. Mol Diagn Ther. 2016. April;20(2):183–90. 10.1007/s40291-016-0190-7 [DOI] [PubMed] [Google Scholar]
- 39.Esmerian MO, Mitri Z, Habbal MZ, Geryess E, Zaatari G, Alam S, et al. Influence of CYP2C9 and VKORC1 polymorphisms on warfarin and acenocoumarol in a sample of lebanese people. Journal of Clinical Pharmacology. [Article]. 2011;51(10):1418–28. 10.1177/0091270010382910 [DOI] [PubMed] [Google Scholar]
- 40.Pathare A, Al Khabori M, Alkindi S, Al Zadjali S, Misquith R, Khan H, et al. Warfarin pharmacogenetics: development of a dosing algorithm for Omani patients. Journal of Human Genetics. 2012. October;57(10):665–9. 10.1038/jhg.2012.94 [DOI] [PubMed] [Google Scholar]
- 41.Shrif NE, Won HH, Lee ST, Park JH, Kim KK, Kim MJ, et al. Evaluation of the effects of VKORC1 polymorphisms and haplotypes, CYP2C9 genotypes, and clinical factors on warfarin response in Sudanese patients. Eur J Clin Pharmacol. 2011. November;67(11):1119–30. 10.1007/s00228-011-1060-1 [DOI] [PubMed] [Google Scholar]
- 42.Özer M, Demirci Y, Hizel C, Sarikaya S, Karalti İ, Kaspar Ç, et al. Impact of genetic factors (CYP2C9, VKORC1 and CYP4F2) on warfarin dose requirement in the Turkish population. Basic Clin Pharmacol Toxicol. 2013. March;112(3):209–14. 10.1111/bcpt.12024 [DOI] [PubMed] [Google Scholar]
- 43.Oner Ozgon G, Langaee TY, Feng H, Buyru N, Ulutin T, Hatemi AC, et al. VKORC1 and CYP2C9 polymorphisms are associated with warfarin dose requirements in Turkish patients. Eur J Clin Pharmacol. 2008. September;64(9):889–94. 10.1007/s00228-008-0507-5 [DOI] [PubMed] [Google Scholar]
- 44.Yildirim E, Erol K, Birdane A. Warfarin dose requirement in Turkish patients: The influences of patient characteristics and polymorphisms in CYP2C9, VKORC1 and factor VII. Hippokratia. [Article]. 2014;18(4):319–27. [PMC free article] [PubMed] [Google Scholar]
- 45.Ozer N, Cam N, Tangurek B, Ozer S, Uyarel H, Oz D, et al. The impact of CYP2C9 and VKORC1 genetic polymorphism and patient characteristics upon warfarin dose requirements in an adult Turkish population. Heart and Vessels. [Article]. 2010;25(2):155–62. 10.1007/s00380-009-1177-7 [DOI] [PubMed] [Google Scholar]
- 46.Luis JR, Rowold DJ, Regueiro M, Caeiro B, Cinnioglu C, Roseman C, et al. The Levant versus the Horn of Africa: evidence for bidirectional corridors of human migrations. Am J Hum Genet. 2004. March;74(3):532–44. 10.1086/382286 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Abu-Amero KK, Gonzalez AM, Larruga JM, Bosley TM, Cabrera VM. Eurasian and African mitochondrial DNA influences in the Saudi Arabian population. BMC Evol Biol. 2007;7:32 10.1186/1471-2148-7-32 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Zalloua PA, Xue Y, Khalife J, Makhoul N, Debiane L, Platt DE, et al. Y-chromosomal diversity in Lebanon is structured by recent historical events. Am J Hum Genet. 2008. April;82(4):873–82. 10.1016/j.ajhg.2008.01.020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Behar DM, Villems R, Soodyall H, Blue-Smith J, Pereira L, Metspalu E, et al. The dawn of human matrilineal diversity. Am J Hum Genet. 2008. May;82(5):1130–40. 10.1016/j.ajhg.2008.04.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Zalloua PA, Platt DE, El Sibai M, Khalife J, Makhoul N, Haber M, et al. Identifying genetic traces of historical expansions: Phoenician footprints in the Mediterranean. Am J Hum Genet. 2008. November;83(5):633–42. 10.1016/j.ajhg.2008.10.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Cadenas AM, Zhivotovsky LA, Cavalli-Sforza LL, Underhill PA, Herrera RJ. Y-chromosome diversity characterizes the Gulf of Oman. Eur J Hum Genet. 2008. March;16(3):374–86. 10.1038/sj.ejhg.5201934 [DOI] [PubMed] [Google Scholar]
- 52.Wu AH, Wang P, Smith A, Haller C, Drake K, Linder M, et al. Dosing algorithm for warfarin using CYP2C9 and VKORC1 genotyping from a multi-ethnic population: comparison with other equations. Pharmacogenomics. 2008. February;9(2):169–78. 10.2217/14622416.9.2.169 [DOI] [PubMed] [Google Scholar]
- 53.Cavallari LH, Langaee TY, Momary KM, Shapiro NL, Nutescu EA, Coty WA, et al. Genetic and clinical predictors of warfarin dose requirements in African Americans. Clin Pharmacol Ther. 2010. April;87(4):459–64. 10.1038/clpt.2009.223 [DOI] [PubMed] [Google Scholar]
- 54.Liu Y, Jeong H, Takahashi H, Drozda K, Patel SR, Shapiro NL, et al. Decreased warfarin clearance associated with the CYP2C9 R150H (*8) polymorphism. Clin Pharmacol Ther. 2012. April;91(4):660–5. 10.1038/clpt.2011.269 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Cal S, Peinado JR, Llamazares M, Quesada V, Moncada-Pazos A, Garabaya C, et al. Identification and characterization of human polyserase-3, a novel protein with tandem serine-protease domains in the same polypeptide chain. BMC Biochem. 2006;7:9 10.1186/1471-2091-7-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.McDonald MG, Rieder MJ, Nakano M, Hsia CK, Rettie AE. CYP4F2 is a vitamin K1 oxidase: An explanation for altered warfarin dose in carriers of the V433M variant. Mol Pharmacol. 2009. June;75(6):1337–46. 10.1124/mol.109.054833 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Caldwell MD, Awad T, Johnson JA, Gage BF, Falkowski M, Gardina P, et al. CYP4F2 genetic variant alters required warfarin dose. Blood. 2008. April 15;111(8):4106–12. 10.1182/blood-2007-11-122010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Voora D, Koboldt DC, King CR, Lenzini PA, Eby CS, Porche-Sorbet R, et al. A polymorphism in the VKORC1 regulator calumenin predicts higher warfarin dose requirements in African Americans. Clin Pharmacol Ther. 2010. April;87(4):445–51. 10.1038/clpt.2009.291 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Caulfield T, Fullerton SM, Ali-Khan SE, Arbour L, Burchard EG, Cooper RS, et al. Race and ancestry in biomedical research: exploring the challenges. Genome Med. 2009;1(1):8 10.1186/gm8 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All relevant data are freely available and are within the paper.