Abstract
Adaptor proteins help proteases modulate substrate choice, ensuring that appropriate proteins are degraded at the proper time and place. SspB is an adaptor that delivers ssrA-tagged proteins to the AAA+ protease ClpXP for degradation. To identify new SspB-regulated substrates, we examined proteins captured by ClpXPtrap in sspB+ but not sspB- strains. RseA1-108, a fragment of a transmembrane protein that regulates the extracytoplasmic-stress response, fits this criterion. In response to stress, RseA is cleaved on each side of the membrane and is released as a cytoplasmic fragment that remains bound in an inhibitory complex with the σE transcription factor. Trapping experiments together with biochemical studies show that ClpXP functions in concert with SspB to efficiently recognize and degrade RseA1-108, and thereby releases σE. Genetic studies confirm that ClpX and SspB participate in induction of the σE regulon in vivo, acting at the final step of an activating proteolytic cascade. Surprisingly, the SspB-recognition sequence in RseA1-108 is unrelated to its binding sequence in the ssrA tag. Thus, these experiments elucidate the final steps in induction of the extracytoplasmic stress response and reveal that SspB delivers a broader spectrum of substrates to ClpXP than has been recognized.
Keywords: RseA, σE, ClpX, ClpP, RpoE
The AAA+ protease ClpXP performs a diverse array of cellular tasks, including degrading incomplete polypeptides, adjusting the activity of metabolic enzymes, and altering the levels of regulatory proteins in response to stress (Gottesman et al. 1998; Wang et al. 1999; Maurizi and Rasulova 2002; Flynn et al. 2003; Gottesman 2003). As a result, many substrates compete for degradation by a relatively small number of ClpXP protease molecules (Ortega et al. 2004). The priority of substrate recognition and degradation can also be controlled by adaptor proteins, which enhance or inhibit interactions between specific substrates and ClpXP or other AAA+ proteases (Dougan et al. 2002a). How widely adaptor proteins are used to control substrate choice is not currently understood.
In the ClpXP protease, ClpX—a hexameric-ring ATPase—binds native substrate proteins, denatures these molecules, and translocates the unfolded polypeptides into an internal degradation chamber of the ClpP peptidase (Maurizi et al. 1990, 1994; Wojtkowiak et al. 1993; Wang et al. 1997; Weber-Ban et al. 1999; Kim et al. 2000; Kim and Kim 2003). ClpX binds to short unstructured peptides called recognition signals or degradation tags, usually located near the amino or C terminus of substrates (Levchenko et al. 1997; Gottesman et al. 1998; Gonciarz-Swiatek et al. 1999; Flynn et al. 2003). The ssrA degradation tag is a well-characterized 11-residue peptide (AANDENYALAA), which is added cotranslationally to nascent polypeptides when ribosomes stall (Keiler et al. 1996). SsrA tagging frees these distressed ribosomes for new rounds of translation and targets the incomplete polypeptides for degradation by ClpXP and other proteases (Gottesman et al. 1998; Withey and Friedman 2003).
The SspB adaptor was originally identified by its ability to enhance ClpXP degradation of ssrA-tagged proteins (Levchenko et al. 2000) and is one of the best-characterized proteins that functions in substrate delivery (Wah et al. 2002, 2003; Dougan et al. 2003; Levchenko et al. 2003; Song and Eck 2003; Bolon et al. 2004). SspB enhances recognition of ssrA-tagged proteins by mediating the assembly of ternary complexes in which the substrate, adaptor, and protease are tethered by the following three sets of protein-peptide interactions: (1) the AAA+ domain of ClpX binds to the C-terminal LAA sequence of the ssrA tag; (2) the substrate-binding domain of SspB interacts with a sequence spanning the N-terminal seven residues of the ssrA tag; and (3) a short peptide sequence at the end of a flexible SspB tail binds directly to the N-terminal domain of ClpX (Levchenko et al. 2000, 2003; Flynn et al. 2001; Wah et al. 2003; Bolon et al. 2004). Whether SspB delivers any substrates without ssrA tags for ClpXP degradation has not been addressed.
Here, we show that SspB directs ClpXP recognition of Escherichia coli proteins, which are not ssrA tagged. One of these substrates, RseA, functions as a master regulator of the extracytoplasmic-stress response by inhibiting the transcription factor (σE) that activates expression of stress genes (De Las Penas et al. 1997b; Missiakas et al. 1997; Dartigalongue et al. 2001; Rezuchova et al. 2003). RseA is a transmembrane protein with an N-terminal cytoplasmic domain, which normally binds to and inhibits σE (De Las Penas et al. 1997b; Missiakas et al. 1997). In response to the stress-induced accumulation of unfolded or unassembled outer-membrane proteins in the periplasm, RseA is processed via multiple cleavage events in a sequential cascade. DegS protease initially cleaves RseA within its periplasmic domain, activating a second cleavage on the cytoplasmic side of the membrane by YaeL protease (Alba et al. 2001, 2002; Kanehara et al. 2002). These cleavage events release the cytoplasmic domain of RseA from the membrane, but this inhibitory domain remains bound to σE, and thus, additional steps are required before σE can activate gene expression (Missiakas et al. 1997; Campbell et al. 2003).
Our experiments demonstrate that ClpXP and SspB play a role in the final step of the proteolytic cascade that activates σE. Cleavage of RseA on the cytoplasmic side of the membrane generates a fragment ending in a ClpX recognition signal, similar to the LAA sequence at the end of the ssrA tag. By binding simultaneously to this RseA1-108 fragment and ClpX, SspB brings the σE · RseA1-108 complex and the ClpXP protease together. The RseA fragment is, however, the only component of this complex that is degraded. Surprisingly, the peptide sequences bound by SspB in RseA1-108 and the ssrA tag are not similar, suggesting the SspB has different modes of protein recognition. These results establish that the SspB adaptor recognizes and delivers different classes of cellular proteins for degradation by ClpXP.
Results
SspB influences recognition of a set of ClpXP substrates in vivo
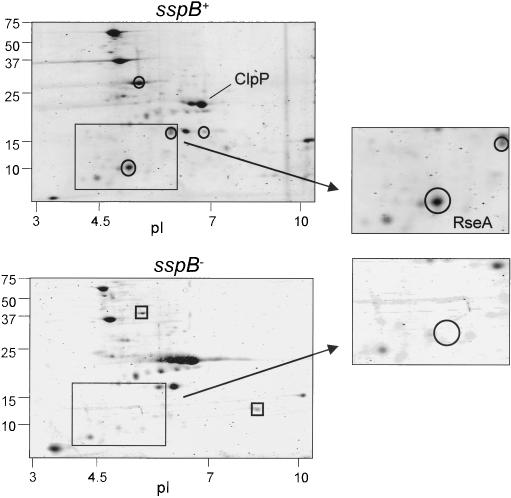
To investigate whether SspB controls ClpXP degradation of proteins without ssrA tags, we compared intracellular substrates captured in an inactive variant of ClpP (ClpPtrap) in the presence and absence of SspB (Flynn et al. 2003). Trapping strains were smpB-, which inactivates ssrA tagging (Karzai et al. 1999), and clpA-, which removes another ATPase capable of choosing substrates for ClpP. These mutations eliminate trapping of ssrA-tagged and ClpAP substrates. Following capture in sspB+ or sspB- strains, ClpXP substrates were visualized by staining following two-dimensional gel electrophoresis (Fig. 1). This experiment revealed that the majority of cellular substrates do not require SspB to interact with ClpXP. However, a handful of proteins were clearly more abundant in ClpPtrap when SspB was present. This differential trapping indicates that SspB influences the recognition of a subset of ClpXP substrates. Interestingly, a few proteins were more efficiently trapped when SspB was absent, suggesting that SspB may also inhibit ClpXP degradation of certain proteins.
Figure 1.
Proteins captured by ClpXPtrap with and without SspB. Two-dimensional gel analysis of proteins captured by ClpXPtrap in E. coli strains JF162 (sspB+clpA-; top) and JF259 (sspB- clpA-; bottom). Representative proteins trapped preferentially in the sspB+ strain are circled, whereas proteins trapped preferentially in the sspB- strain are marked by squares.
One SspB-dependent substrate is an N-terminal fragment of RseA
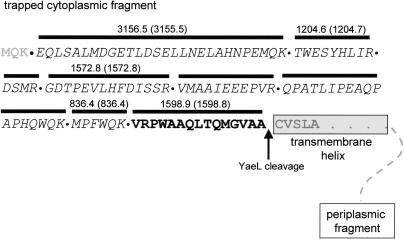
Tandem-mass spectrometry identified one of the most prominent SspB-dependent ClpXP trapped proteins as an N-terminal fragment of RseA. Tryptic digestion of the RseA spot followed by mass spectrometry identified peptides covering the N-terminal 108 amino acids of RseA (Fig. 2), including a peptide with a molecular weight corresponding to residues 94-108: VRPWAAQLTQMG VAA108. The fact that this peptide did not terminate with lysine or arginine (as expected for an internal tryptic fragment) indicated that alanine was the natural C terminus of the trapped protein. Thus, this analysis demonstrates that the trapped RseA fragment (RseA1-108) terminates with the sequence VAA-COOH (Fig. 2). This C-terminal sequence is a member of the well-characterized C-motif 1 class of ClpX recognition signals (Flynn et al. 2003), and thus, it makes sense that it would target the RseA fragment to ClpXP.
Figure 2.
Sequence analysis of the ClpXPtrap-captured RseA fragment. Tryptic fragments of the RseA fragment were analyzed by mass spectrometry. Identified peptides are marked with bold lines above the corresponding sequences; sequences identified by tandem mass spectrometry are italicized, and peptides identified by MALDI mass spectrometry have the experimental (expected) molecular weights listed. The peptide highlighted in bold was identified by MALDI mass spectrometry and is the C-terminal tryptic peptide of the trapped fragment.
Recognition of RseA by the cytoplasmic protease ClpXP must occur after YaeL cleavage releases the N-terminal fragment from the membrane (Alba et al. 2001; Kanehara et al. 2002). On the basis of the cleavage specificity of the homologous SP2 protease, Alba et al. (2002) proposed that YaeL might cleave RseA between A108 and C109, to generate the N-terminal fragment that we trapped and characterized.
RseA1-108 is a substrate for SspB and ClpXP in vitro
A fragment corresponding to RseA1-108 was cloned, overexpressed, and purified to investigate its susceptibility to ClpXP degradation in vitro. ClpXP degraded RseA1-108 in a reaction that required ATP (Fig. 3A; data not shown). A mutant variant in which the C-terminal sequence was VDD108 (RseA-DD1-108) was also purified and was found to be degraded 25-30 times more slowly than RseA1-108 (Fig. 3A, inset). Thus, we conclude that the C-terminal sequence of RseA1-108 is a critical signal that targets this protein for degradation by ClpXP.
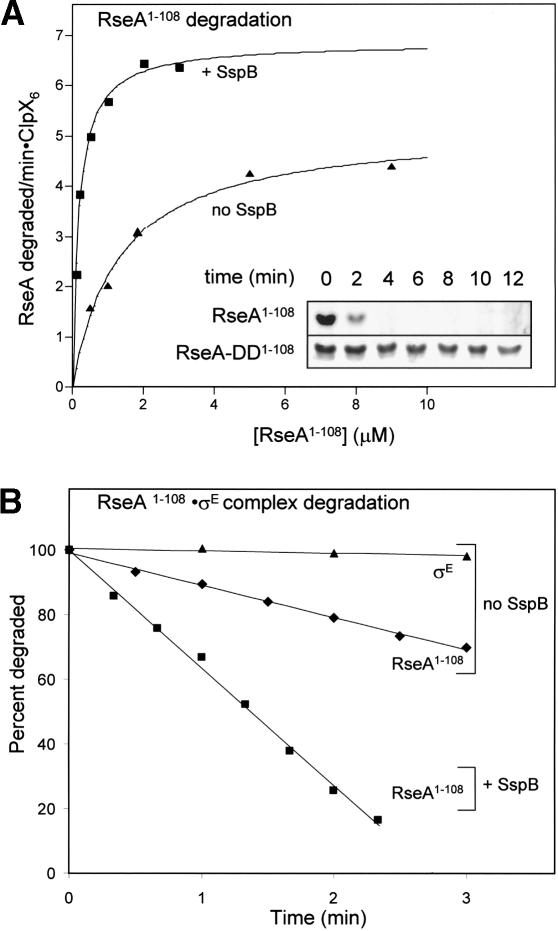
Figure 3.
ClpXP efficiently degrades purified RseA1-108 in an SspB-stimulated manner. (A) Rates of ClpXP-mediated degradation of 35S-labeled RseA1-108 by ClpX6 (50 nM) and ClpP14 (150 nM) were determined at different substrate concentrations in the presence or absence of SspB (200 nM). Degradation was assayed by changes in TCA-soluble radioactivity, and rates were plotted against the substrate concentration. The solid lines are fits to the Michaelis-Menten equation in the absence (Km = 1.3 μM, Vmax = 5.2 min-1) or presence (Km = 0.18 μM, Vmax = 6.8 min-1) of SspB. (Inset) Degradation of RseA-DD1-108 (2 μM) or RseA1-108 (2 μM) by ClpX6 (300 nM) and ClpP14 (800 nM) was assayed by SDS-PAGE. (B) ClpXP degradation of RseA1-108 complexed with σE. 35S-labeled RseA1-108 (500 nM) was incubated with unlabeled σE (500 nM) for 5 min at 30°C. Degradation by ClpX6 (50 nM) and ClpP14 (150 nM) was assayed by changes in TCA-soluble radioactivity in the presence (▪) or absence (♦) of SspB (200 nM). 35S-labeled σE was also incubated with unlabeled RseA1-108 and ClpXP degradation was monitored in the same manner (▴). No detectable σE degradation by ClpXP was observed in the presence of SspB.
As expected from the trapping results, SspB also stimulated degradation of RseA1-108 by ClpXP in vitro (Fig. 3). SspB reduced the Michaelis constant (Km) for ClpXP degradation of RseA1-108 approximately sevenfold from 1.3 to ∼0.2 μM and stimulated Vmax by ∼50% (Fig. 3A). Thus, SspB enhances productive interactions between RseA1-108 and ClpX, in a manner analogous to its role in delivering ssrA-tagged proteins for ClpXP degradation (Levchenko et al. 2000). A truncated SspB variant lacking the tails that bind ClpX did not enhance ClpXP degradation of RseA1-108, demonstrating that tethering interactions between SspB and ClpX are important for delivery of this substrate (data not shown).
YaeL cleavage releases the N-terminal fragment of RseA from the membrane, but does not disrupt its binding to σE (Alba et al. 2002; Kanehara et al. 2002). We asked, therefore, whether SspB could deliver the σE · RseA1-108 complex to ClpXP for disassembly and degradation. As shown in Figure 3B, ClpXP degraded RseA1-108 bound to σE, and SspB stimulated this degradation. At the concentrations tested, the rate of degradation of free RseA1-108 was similar to that of complexed RseA1-108. This result indicates that binding of σE to RseA1-108 does not inhibit degradation or provide any critical contacts that enhance recognition of RseA1-108 by ClpX. Importantly, σE in the σE · RseA1-108 complex was not degraded. In addition, as expected, SspB remained undegraded throughout the reaction (data not shown).
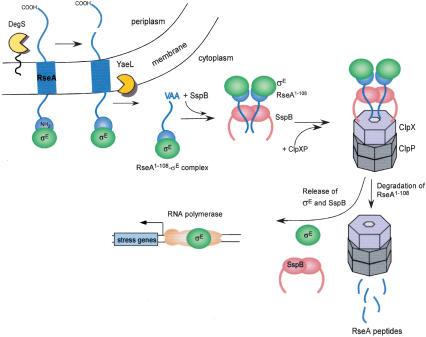
On the basis of this analysis, we conclude that SspB can deliver the σE · RseA1-108 complex to ClpXP, leading to the targeted degradation of RseA1-108. These results are integrated into a model for σE activation shown in Figure 4. Following DegS and YaeL cleavage of RseA, SspB delivers the σE · RseA1-108 complex to ClpX, which selectively denatures RseA1-108 and translocates it into ClpP for degradation. This processing of the RseA fragment by ClpXP releases SspB and σE from the enzyme complex. As a consequence, σE is liberated to bind to core RNA polymerase and activate transcription.
Figure 4.
Activation of σE mediated by a cascade of RseA proteolysis.
SspB and ClpX enhance activation of the σE regulon in vivo
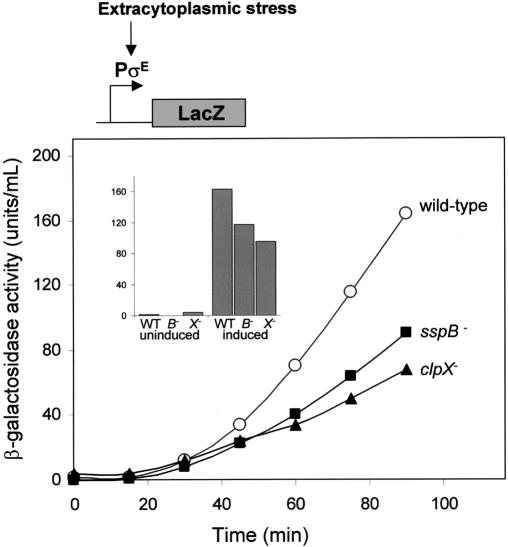
Taken together, the results presented so far suggest that degradation mediated by ClpXP and SspB controls the intracellular levels of RseA1-108 and should therefore influence σE activity. To test for roles for ClpX and SspB in the extracytoplasmic stress response, we monitored induction of a σE-controlled lacZ reporter gene following induction of the stress response in sspB- or clpX- cells. Extracytoplasmic stress was induced using a plasmid-encoded fusion protein, ending with a YYF sequence, which is targeted to the periplasm and activates DegS degradation of RseA (Walsh et al. 2003). Following induction, σE-dependent β-galactosidase synthesis was delayed in both the clpX- and sspB- cells (Fig. 5). These data show that ClpX and SspB participate in activation of σE during the stress response. The clpX- cells had a larger defect than the sspB- cells, in accordance with the observation that SspB is not essential for ClpXP degradation of RseA1-108 in vitro. Although clearly reduced, the σE-reporter gene was still induced in the absence of ClpX, suggesting that proteases in addition to ClpXP also participate in the activation of σE by degrading RseA1-108 (see Discussion).
Figure 5.
Induction of the σE regulon is attenuated in sspB- and clpX- strains. The extracytoplasmic-stress response was induced in wild-type (CAG43583), sspB::kan (CAG43583) and clpX::kan (CAG43583) strains with L-arabinose at time 0. Samples were analyzed for β-galactosidase activity at the times indicated. The clpX::kan and sspB::kan strains grow slightly slower than wild type. When the cultures were at a similar O.D.600, however, the clpX::kan and sspB::kan strains still exhibited reduced levels of β-galactosidase (see inset). The uninduced samples were measured at time 0 when the cultures were at an O.D.600 of 0.15. The induced samples were measured when the cultures reached an O.D.600 of 0.45.
SspB forms stable delivery complexes with RseA1-108 and with σE · RseA1-108
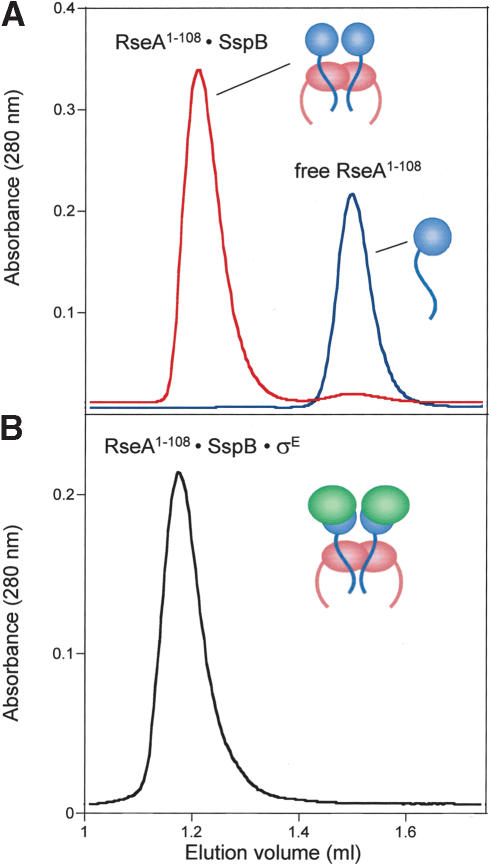
Mutagenic and crystallographic studies have identified detailed interactions between the ssrA tag and SspB, and peptide-binding studies have established a strong consensus sequence for SspB recognition of the tag (Levchenko et al. 2000, 2003; Flynn et al. 2001; Song and Eck 2003). Inspection of the RseA1-108 sequence, however, failed to identify any sequences with significant homology to the SspB-recognition sequence in the ssrA tag. Thus, we sought to determine whether SspB forms a specific complex with RseA1-108 as it does with the ssrA tag using gel filtration as a binding assay. SspB and RseA1-108 coeluted on a Superose 12 column at a position distinct from free RseA1-108 (Fig. 6A). Moreover, a larger ternary complex was formed when SspB, σE, and RseA1-108 were mixed (Fig. 6B). The presence of SspB, σE, and RseA1-108 in this complex was confirmed by SDS-PAGE (data not shown). Stable formation of this ternary complex provides further support for the model that SspB binds the σE · RseA1-108 complex and delivers this complex to ClpXP.
Figure 6.
SspB forms stable complexes with RseA1-108 and RseA1-108 · σE. Gel-filtration on a Superose 12 column (4°C) of RseA1-108 · SspB complex (A, top trace), free RseA (A, bottom trace), and RseA1-108 · σE · SspB complex (B).
Truncation experiments established that a sequence near the C terminus of RseA1-108 was required for stable complex formation with SspB. A truncated variant ending at residue 89 (RseA1-89) failed to coelute with SspB during gel filtration, whereas a slightly longer variant, RseA1-99, retained the ability to bind SspB stably (Fig. 7A). To determine which portion of RseA1-108 bound SspB, we looked for sites protected from tryptic cleavage in the complex. Incubation of RseA1-99 with trypsin resulted in two major stable fragments; the larger fragment resulted from digestion after K93, whereas the smaller fragment was generated by trypsin digestion after both R58 and K93 (Fig. 7B). In the presence of SspB, two larger fragments were also observed as a result of partial suppression of the cleavage following K93. These data, like the truncation experiments, implicate the sequence surrounding residue 93 in SspB · RseA complex formation.
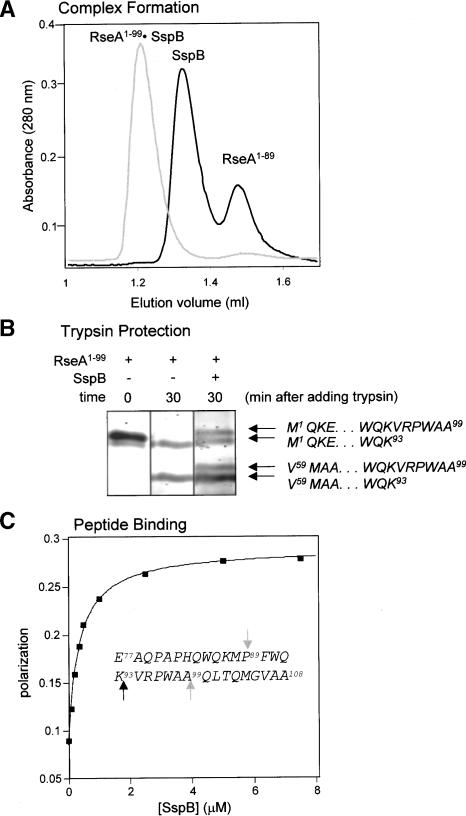
Figure 7.
SspB interacts with residues 77-99 of RseA. (A) RseA1-99 forms a stable complex with SspB (gray trace), whereas RseA1-89 does not form this complex (black trace). The RseA variants and SspB were incubated at 30°C for 5 min and then chromatographed on a Superose 12 gel-filtration column (4°C). (B) Protection of RseA1-99 by SspB from tryptic cleavage. RseA1-99 (5 μM) was incubated with trypsin in the absence or presence of SspB (15 μM). Electrospray mass spectrometry and N-terminal sequencing determined the identity of the resulting fragments. (C) The RseA77-108 peptide binds to SspB. Binding of fluorescently labeled RseA77-108 peptide to SspB at 30°C was measured by an increase in polarization. The solid line is a fit for a Kd of 0.35 μM. Unlabeled RseA77-108 peptide was able to compete for binding to the fluoresceinated peptide. The sequence of the RseA77-108 peptide is given in the inset. Gray arrows correspond to the C-terminal residues of the fragments tested for complex formation in A. The black arrow corresponds to the protected trypsin site in B.
Peptide-binding studies confirm that the C-terminal region of RseA1-108 mediates its interaction with SspB. A synthetic fluorescein-labeled peptide containing RseA residues 77-108 bound SspB with a Kd of 0.35 μM as determined by changes in fluorescence polarization (Fig. 7C). This binding was competed both by excess RseA1-108 and by an ssrA peptide (data not shown). Furthermore, a mutation in the peptide-binding cleft of SspB (Bolon et al. 2004) prevented binding of both molecules. These experiments suggest that the C-terminal region of RseA1-108 and the ssrA peptide bind to at least some common sites within the peptide-binding cleft on SspB, despite the lack of significant sequence homology.
Discussion
ClpXP and SspB regulate σE activity via RseA destruction
The activity of σE, the transcription factor for the extracytoplasmic-stress response, is tightly controlled by its binding to and inhibition by the transmembrane regulator, RseA (Fig. 4; De Las Penas et al. 1997b; Missiakas et al. 1997). Stress induces sequential cleavages of RseA on each side of the membrane by the DegS and YaeL proteases, respectively, releasing the σE · RseA1-108 complex into the cytoplasm (Ades 2004). Our results show that ClpXP, with the assistance of SspB, recognizes the inhibited σE · RseA1-108 complex and catalyzes release of active σE through selective proteolytic destruction of RseA1-108. Thus, SspB and ClpXP participate in the final stage of a proteolytic cascade, which begins in the periplasm and, ultimately, releases an active transcription factor in the cytoplasm.
ClpXP is especially well suited to recognize and degrade proteins with C-terminal signals generated by prior proteolytic cleavage. Cleavage of RseA from the membrane generates a fragment that terminates with VAACOOH, a sequence that belongs to the C-motif 1 class of ClpXP recognition signals (Flynn et al. 2003). For this class of peptide sequences, which includes the ssrA tag, the nonpolar side chains and the free α-carboxyl group are both important for ClpX recognition (Kim et al. 2000; Flynn et al. 2001). Thus, a VAA or LAA sequence is recognized poorly, if at all, at an internal position in a protein. Degradation of the SOS response repressor, LexA, also illustrates this type of regulation (Neher et al. 2003a). Full-length LexA is not a ClpXP substrate, but damage-induced auto-cleavage creates an N-terminal LexA fragment, ending with VAA-COOH, which is degraded efficiently by ClpXP (Neher et al. 2003a). Thus, certain internal peptide sequences function as cryptic degradation signals, which remain hidden until revealed by protein cleavage. Cryptic signals permit coordinated protein destruction, allowing a single protein processing event—such as cleavage in response to an environmental cue—to trigger recognition by ClpXP.
Structural and biochemical studies demonstrate that complexes of σE with RseA are very stable and incompatible with transcriptional activation. The cocrystal structure of RseA1-90 bound to σE reveals extensive contacts in which the first 66 amino acids of RseA are sandwiched between the two domains of σE in a manner that would directly block σE-RNA polymerase interaction (Campbell et al. 2003). We found that the σE · RseA1-108 complex copurified over several columns without detectable dissociation during a period of days (J. Flynn and I. Levchenko, unpubl.), and direct experiments estimate the half-life of the complex in vitro to be well in excess of 2 h (I. Grigorova and C. Gross, pers. comm.). Response to extracytoplasmic stress, in contrast, occurs in minutes, a time-scale similar to the rate of ClpXP degradation of RseA1-108 in a σE · RseA1-108 complex. Therefore, ClpX must actively pull the two proteins in the σE · RseA1-108 complex apart to release σE and allow degradation of RseA1-108. The proteolytic activity of ClpP in the ClpXP complex may assist in activation of σE by destroying RseA1-108 to prevent reformation of the RseA1-108 · σE complex. Thus, a key feature of σE activation is the mechanical disassembly of the σE · RseA1-108 complex by ClpXP.
In the cocrystal structure of σE · RseA1-90, the first 66 residues of RseA form a stable domain that binds σE, whereas the last 24 residues are not visible and are presumably unstructured (Campbell et al. 2003). Although previously there was no known function for this unstructured extension of N-RseA, our data indicates that this region functions to interact with both SspB and ClpX during the final step of activation of σE. SspB, σE and N-RseA form a stable delivery complex, in which σE interacts with the first 66 residues of RseA, and SspB binds to the C-terminal unstructured tail. Why is RseA1-108 the only member of this stable complex degraded? Both SspB and σE probably lack degradation signals that would allow ClpX to engage these proteins to initiate protein degradation. Alternatively, the geometry of the complex might place RseA1-108 but not the other proteins in a position that allows engagement by the enzyme.
σE function is essential in E. coli (De Las Penas et al. 1997a) but ClpX, ClpP, and SspB are nonessential proteins, suggesting that other proteases also degrade RseA1-108 and release active σE. clpX- and sspB- cells show reduced induction of a σE-regulated promoter, rather than no induction. In fact, recent experiments demonstrate that several different proteases participate in degradation of RseA1-108, although ClpXP plays the single largest role (R. Chaba and C. Gross, pers. comm.). Hence, RseA1-108 must contain targeting signals for several proteases, emphasizing the critical nature of its destruction.
Adaptors like SspB expand and regulate the substrate repertoire of proteases
Prior to this study, ssrA-tagged proteins were the only known substrate partners for SspB (Levchenko et al. 2000). Identification of RseA1-108 as a new SspB partner provides the opportunity to compare mechanisms of substrate delivery. There are many similarities. Both RseA1-108 and ssrA-tagged proteins contain a C-motif 1 degradation tag at the extreme C terminus, and SspB binds to a nearby region within 10-30 residues. For both classes of substrates, SspB enhances ClpXP degradation principally by decreasing Km, and therefore serves to stabilize enzyme-substrate interactions. Finally, RseA1-108 and the ssrA tag appear to occupy overlapping binding sites in the peptide-binding cleft on SspB.
Despite these similarities, the sequences within RseA1-108 and the ssrA tag that bind SspB are not similar. Experiments presented here reveal that the SspB-binding site in RseA1-108 lies between residues 77 and 99 (see Fig. 7). This region, as well as the rest of RseA1-108, is devoid of sequences resembling the ssrA tag consensus for SspB binding ([AGPSV]1-[ASV]2-[NH]3-[DCE]4-X5-X6-[FWY]7; Flynn et al. 2001). Studies are currently in progress to define more clearly how RseA1-108 binds to SspB and how the peptide-binding cleft of SspB can interact strongly and specifically with two, seemingly unrelated, sequences.
The studies reported here revealed several different proteins that were trapped in sspB+ but not sspB- strains. In addition to RseA1-108, trapping of both AceA (isocitrate lyase) and Cdd (deoxycytidine deaminase) was also stimulated by the presence of SspB (data not shown). Delivery of ssrA-tagged substrates or RseA1-108 for ClpXP degradation is clearly a direct consequence of SspB function, and we suspect that additional proteins will also be directly delivered by SspB. However, adaptors also can have indirect effects on substrate selection by AAA+ proteases. For example, by mediating efficient degradation of specific substrates, an adaptor may serve to free the protease to degrade other substrates more efficiently. In addition, targeted degradation of transcription factors, translation regulators, chaperones, and proteases has the potential to cause large changes in protein levels, leading to indirect changes in the repertoire of substrates available for degradation.
Although SspB is a positive regulator of RseA1-108 recognition, it also has the potential to act as an inhibitor. In our experiments, ClpXP trapped a few substrates more efficiently when SspB was absent (see Fig. 1). SspB binding could prevent ClpXP degradation of certain proteins by masking their degradation tags. In fact, both SspB and the ClpS adaptor protein inhibit ClpAP recognition of ssrA-tagged proteins (Flynn et al. 2001; Dougan et al. 2002b). Alternatively, absence of competition could lead to improved degradation of substrates or substrate-adaptor complexes that compete with SspB for tethering to ClpX.
It is becoming increasingly clear that many proteins are targeted for disassembly and destruction by AAA+ ATPases both by intrinsic recognition tags and by extrinsic tethering mediated by adaptor proteins. How many adaptors exist for each enzyme, and their overall impact on recognition is not yet known. In addition to SspB, E. coli ClpXP uses the RssB adaptor that delivers the stationary σ factor σS to ClpXP for degradation during non-starvation conditions (Muffler et al. 1996; Zhou and Gottesman 1998). Furthermore, the UmuD subunit of the UmuD UmuD′ heterodimer functions as an SspB-like adaptor for UmuD′ degradation by ClpXP during recovery from DNA damage (Neher et al. 2003b).
Why do certain substrates use adaptors? One answer is that adaptor proteins can increase the efficiency of recognition at low substrate concentrations. For example, SspB improves ClpXP recognition of RseA1-108 in vivo, as shown both by trapping and σE-induction experiments, even though RseA1-108 is a good ClpXP substrate in the absence of SspB in vitro. Furthermore, the use of adaptors can lead to the degradation of a group of proteins, allowing coregulation. The results of our trapping experiments indicate that up-regulation or down-regulation of SspB would be likely to change the efficiency of degradation of a group of substrate proteins in a coordinated manner. In fact, we have observed that overproduction of SspB improves activation of σE during stress (data not shown). We suspect that additional adaptors remain to be discovered. These proteins, like SspB, will probably also bind a spectrum of substrates, thereby controlling the breadth and efficiency of recognition by their partner AAA+ enzymes.
Materials and methods
Strains and plasmids
Genes encoding RseA1-108 and RseA1-99 were amplified by PCR from E. coli genomic DNA using primers encoding NdeI and BamHI restriction sites. The amplified DNA was cleaved with both restriction enzymes and cloned between the NdeI and BamHI sites of pET3a to generate pET3a-rseA1-108 and pET3arseA1-99. A plasmid expressing RseA-DD1-108 was constructed by site-directed mutagenesis of the rseA1-108 gene. The gene encoding σE (rpoE) was PCR amplified from E. coli chromosomal DNA and cloned into the NdeI and BclI sites of the pT7LysS plasmid (I. Levchenko, unpubl.) to generate pT7LysS-rpoE.
The chromosomally encoded sspB gene was replaced by a FRT-flanked kanamycin resistance cassette following the method of Datsenko and Wanner (2000). The sspB::kan cassette was then transferred into W3110 clpP::cat ΔsmpB-1 cells by P1 transduction. KmR mutants were transformed with pCP20 encoding the Flipase enzyme, and resulting transformants were tested for loss of the kanamycin resistance as described in Datsenko and Wanner (2000). The deletion was confirmed by PCR analysis. A clpA · kan cassette was then introduced by P1 transduction and finally pJF105 (Flynn et al. 2003) encoding the ClpPtrap was transformed into the strain (JF259). CAG43583 (Walsh et al. 2003) was a gift from Carol Gross (University of California at San Francisco, San Francisco, CA). The sspB · kan and clpX · kan cassettes were introduced into the strain by P1 transduction.
Solutions
Buffer A is 50 mM Tris-HCl (pH 7.0), 50 mM NaCl, 0.5 mM DTT, 2 mM EDTA, and 5% glycerol. GF buffer is 50 mM Tris-HCl (pH 7.0), 150 mM KCl, 1 mM DTT, and 5% glycerol. PD buffer is as described (Kim et al. 2000).
Proteins
ClpX (Levchenko et al. 1997) and ClpP (Kim et al. 2000) were purified as described; SspB was a gift from David Wah (Massachusetts Institute of Technology, Cambridge, MA).
RseA1-108 was purified from E. coli ER2556 pLysS/pET3a-RseA1-108 cells grown in LB broth with 100 μg/mL ampicillin and 30 μg/mL chloramphenicol. Cells were grown at 37°C to an O.D.600 of 0.6 and protein expression was induced with IPTG for 2 h. All purification steps were performed at 4°C. Cell pellets were resuspended at a concentration of 3 mL/g of cells in buffer A plus 6 M of guanidine and Protease Inhibitor Cocktail III (Calbiochem). Following lysis for 1 h, the lysate was centrifuged for 30 min at 25,000 × g and the supernatant was dialyzed overnight against 4 L of buffer A with one buffer change. Insoluble proteins were removed by centrifugation, and ammonium sulfate was added to the supernatant to a final concentration of 30%. After mixing for 20 min, the precipitate containing RseA1-108 was collected by centrifugation, resuspended in buffer A, and desalted into the same buffer using a PD-10 desalting column (Amersham Biosciences). This sample was loaded onto a MonoQ HR 5/5 column (Amersham Biosciences) equilibrated in buffer A. The column was washed with 10-column volumes of buffer A, and the bound protein was eluted with a gradient to 1 M NaCl. The peak, including RseA1-108, was collected, and TFA was added to a final concentration of 0.06%. The sample was applied to a C4 HPLC column equilibrated in 0.06% TFA to separate full-length RseA1-108 from degradation products, and eluted with a gradient to 80% acetonitrile. RseA1-108 was lyophilized, resuspended in buffer A, and dialyzed against the same buffer overnight. RseA1-108 concentration was determined by UV absorbance (ε280 = 24040 M-1 cm-1).
RseA1-99 was purified from E. coli BL-21/pET3a-rseA1-99 cells using a similar protocol, except lysis in 50 mM Tris (pH 8.0), 0.l5 M NaCl, 1 mM EDTA, and 5% glycerol was performed by French press, and a Superdex 75 column (Amersham Biosciences) was used in place of HPLC as the final purification step. The resulting protein was >95% pure as determined by Commassie staining on a SDS-polyacrylamide gel.
The σE · RseA1-108 and σE · RseA1-108DD complexes were purified from ER2566 E. coli cells coexpressing either pET3arseA1-108 or pET3a-rseA1-108DD and pT7LysS-rpoE plasmids. The binary complexes were purified on a Sephacryl S-200 gel filtration column (Amersham Biosciences) followed by chromatography on Source15Q (Amersham Biosciences). The RseA fragment purified from the RseA1-108 · σE complex contained a number of C-terminal degradation products. The smallest degradation product (RseA1-89) was isolated by a C4 HPLC column as described above, and had a molecular weight of 10,251 Da by electrospray mass spectrometry.
35S-labeled RseA1-108 and σE were purified as a complex from ER2566 E. coli cells coexpressing pET3a-rseA1-108 and pT7LysS-rpoE. 35S-labeling was performed as described (Kim et al. 2000). Cells were lysed in guanidine, the lysate was dialyzed against buffer A, and a 30% ammonium sulfate cut was performed. The precipitate was resuspended in buffer A, and applied to a protein C4 HPLC column equilibrated in 0.06% TFA. RseA1-108 and σE, were separated by a gradient to 80% acetonitrile, lyophilized, resuspended in buffer A, and dialyzed against the same buffer overnight. σE concentration was determined by UV absorbance (ε280 = 14650 M-1 cm-1).
Synthetic fluorescein-labeled peptides containing residues 77-108 of RseA (EAQPAPHQWQKMPFWQKVRPWAAQLTQ MGVAA) and of an SsrA tag sequence (NKKGRHGAANDE NYALAA) were synthesized by the Massachusetts Institute of Technology Biopolymers Laboratory and purified by reverse-phase chromatography on a C4 HPLC column (Vydac).
Protein trapping
Trapped proteins were isolated from an sspB+ strain (JF162; W3110 clpP::cat clpA::kan ΔsmpB-1/pJF105) or an sspB- strain (JF259; see above) and analyzed by two-dimensional gels as described (Flynn et al. 2003). Protein spots from the gel were excised, digested with trypsin, and analyzed by microcapillary reverse-phase HPLC nano-electrospray tandem mass spectrometry using a Finnigan LCQ DECA quadropole ion trap mass spectrometer (Harvard Microchemistry Facility). The two-dimensional spot corresponding to RseA1-108 was subjected to in-gel tryptic digestion as described (Rosenfeld et al. 1992; Hellman et al. 1995) and peptides were analyzed by MALDI mass spectrometry at the MIT Biopolymers Facility.
Degradation assays
ClpX6, ClpP14, ATP (4 mM), and an ATP regeneration system (50 μg/mL creatine kinase and 2.5 mM creatine phosphate) were mixed in PD buffer and incubated for 2 min at 30°C. For gel analysis, RseA1-108 or RseA-DD1-108 (2 μM) was added, and samples were removed at different times and analyzed by SDS-PAGE. Bands were visualized using Sypro Orange protein stain (Molecular Probes) on a Fluorimager 595 (Molecular Dynamics). Degradation of 35S-labeled proteins were assayed by changes in TCA-soluble radioactivity as described in Burton et al. (2001). When present, the SspB concentration was 0.2 μM (monomer equivalents).
Gel filtration of protein complexes
Gel filtration was performed on a SMART system (Amersham Biosciences) using a Superose 12 column equilibrated in GF buffer at 4°C. RseA1-108, RseA1-99, RseA1-89, or the RseA1-108 · σE complex (8 μM) was incubated with or without SspB (8 μM monomer equivalents) in GF buffer for 5 min at 30°C prior to chromatography.
Limited trypsin proteolysis
A total of 5 μM RseA1-99 was incubated with or without 15 μM SspB in 100 mM Tris-HCl (pH 8.9) for 5 min at 30°C. Trypsin and RseA1-108 were mixed in a 1:93 ratio and samples were taken at different times and analyzed by 18% Tris-Tricine SDS-PAGE. To identify the resulting RseA fragments, a portion of each time point was analyzed by electrospray mass spectrometry and another portion was separated by SDS-PAGE, transferred onto PVDF membrane (Millipore), stained by Ponceau red stain, and subjected to N-terminal sequencing at the Massachusetts Institute of Technology Biopolymers Facility.
Peptide-binding assays
Binding of SspB to the fluorescein-labeled RseA75-108 peptide (0.1 μM) was assayed by fluorescence polarization (excitation 467 nm; emission 511 nm) at 30°C in PD buffer lacking NP-40 using a Fluoromax-2 instrument (ISA, Jobin-Yvon). Binding curves were fit using Kaleidagraph (Synergy Software).
β-Galactosidase assays
Overnight cultures were diluted 1:100 to an O.D.600 of ∼0.025 and grown at 30°C in LB broth with appropriate antibiotics. The cultures were then grown at 30°C to an O.D.600 of 0.15 and overexpression of the OmpC fusion protein was induced by 0.2% L-(+)-arabinose. β-Galactosidase activities were measured as described (Miller 1972; Mecsas et al. 1993; Ades et al. 1999).
Acknowledgments
We thank Carol Gross for strains and advice, and Rachna Chaba, Irena Grigorova, and Carol Gross for sharing unpublished results. We thank the Harvard Microchemistry Facility for mass spectrometry and members of the Baker and Sauer labs for help, advice, and comments on the manuscript. Supported by NIH grant AI-16892 and HHMI. T.A.B. is an employee of HHMI.
Article and publication are at http://www.genesdev.org/cgi/doi/10.1101/gad.1240104.
References
- Ades S.E. 2004. Control of the alternative σ factor σ(E) in Escherichia coli. Curr. Opin. Microbiol. 7: 157-162. [DOI] [PubMed] [Google Scholar]
- Ades S.E., Connolly, L.E., Alba, B.M., and Gross, C.A. 1999. The Escherichia coli σ(E)-dependent extracytoplasmic stress response is controlled by the regulated proteolysis of an anti-σ factor. Genes & Dev. 13: 2449-2461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alba B.M., Zhong, H.J., Pelayo, J.C., and Gross, C.A. 2001. degS (hhoB) is an essential Escherichia coli gene whose indispensable function is to provide σ (E) activity. Mol. Microbiol. 40: 1323-1333. [DOI] [PubMed] [Google Scholar]
- Alba B.M., Leeds, J.A., Onufryk, C., Lu, C.Z., and Gross, C.A. 2002. DegS and YaeL participate sequentially in the cleavage of RseA to activate the σ(E)-dependent extracytoplasmic stress response. Genes & Dev. 16: 2156-2168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolon D.N., Wah, D.A., Hersch, G.L., Baker, T.A., and Sauer, R.T. 2004. Bivalent tethering of SspB to ClpXP is required for efficient substrate delivery: A protein-design study. Mol. Cell 13: 443-449. [DOI] [PubMed] [Google Scholar]
- Burton R.E., Siddiqui, S.M., Kim, Y.I., Baker, T.A., and Sauer, R.T. 2001. Effects of protein stability and structure on substrate processing by the ClpXP unfolding and degradation machine. EMBO J. 20: 3092-3100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Campbell E.A., Tupy, J.L., Gruber, T.M., Wang, S., Sharp, M.M., Gross, C.A., and Darst, S.A. 2003. Crystal structure of Escherichia coli σE with the cytoplasmic domain of its anti-sigma RseA. Mol. Cell 11: 1067-1078. [DOI] [PubMed] [Google Scholar]
- Dartigalongue C., Missiakas, D., and Raina, S. 2001. Characterization of the Escherichia coli σ E regulon. J. Biol. Chem. 276: 20866-20875. [DOI] [PubMed] [Google Scholar]
- Datsenko K.A. and Wanner, B.L. 2000. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. 97: 6640-6645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Las Penas A., Connolly, L., and Gross, C.A. 1997a. σE is an essential σ factor in Escherichia coli. J. Bacteriol. 179: 6862-6864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ____. 1997b. The σE-mediated response to extracytoplasmic stress in Escherichia coli is transduced by RseA and RseB, two negative regulators of σE. Mol. Microbiol. 24: 373-385. [DOI] [PubMed] [Google Scholar]
- Dougan D.A., Mogk, A., Zeth, K., Turgay, K., and Bukau, B. 2002a. AAA+ proteins and substrate recognition, it all depends on their partner in crime. FEBS Lett. 529: 6-10. [DOI] [PubMed] [Google Scholar]
- Dougan D.A., Reid, B.G., Horwich, A.L., and Bukau, B. 2002b. ClpS, a substrate modulator of the ClpAP machine. Mol. Cell 9: 673-683. [DOI] [PubMed] [Google Scholar]
- Dougan D.A., Weber-Ban, E., and Bukau, B. 2003. Targeted delivery of an ssrA-tagged substrate by the adaptor protein SspB to its cognate AAA+ protein ClpX. Mol. Cell 12: 373-380. [DOI] [PubMed] [Google Scholar]
- Flynn J.M., Levchenko, I., Seidel, M., Wickner, S.H., Sauer, R.T., and Baker, T.A. 2001. Overlapping recognition determinants within the ssrA degradation tag allow modulation of proteolysis. Proc. Natl. Acad. Sci. 98: 10584-10589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flynn J.M., Neher, S.B., Kim, Y.I., Sauer, R.T., and Baker, T.A. 2003. Proteomic discovery of cellular substrates of the ClpXP protease reveals five classes of ClpX-recognition signals. Mol. Cell 11: 671-683. [DOI] [PubMed] [Google Scholar]
- Gonciarz-Swiatek M., Wawrzynow, A., Um, S.J., Learn, B.A., McMacken, R., Kelley, W.L., Georgopoulos, C., Sliekers, O., and Zylicz, M. 1999. Recognition, targeting, and hydrolysis of the λ O replication protein by the ClpP/ClpX protease. J. Biol. Chem. 274: 13999-14005. [DOI] [PubMed] [Google Scholar]
- Gottesman S. 2003. Proteolysis in bacterial regulatory circuits. Annu. Rev. Cell. Dev. Biol. 19: 565-587. [DOI] [PubMed] [Google Scholar]
- Gottesman S., Roche, E., Zhou, Y., and Sauer, R.T. 1998. The ClpXP and ClpAP proteases degrade proteins with carboxy-terminal peptide tails added by the SsrA-tagging system. Genes & Dev. 12: 1338-1347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hellman U., Wernstedt, C., Gonez, J., and Heldin, C.H. 1995. Improvement of an `In-Gel' digestion procedure for the micropreparation of internal protein fragments for amino acid sequencing. Anal. Biochem. 224: 451-455. [DOI] [PubMed] [Google Scholar]
- Kanehara K., Ito, K., and Akiyama, Y. 2002. YaeL (EcfE) activates the σE pathway of stress response through a site-2 cleavage of anti-E, RseA. Genes & Dev. 16: 2147-2155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karzai A.W., Susskind, M.M., and Sauer, R.T. 1999. SmpB, a unique RNA-binding protein essential for the peptide-tagging activity of SsrA (tmRNA). EMBO J. 18: 3793-3799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keiler K.C., Waller, P.R., and Sauer, R.T. 1996. Role of a peptide tagging system in degradation of proteins synthesized from damaged messenger RNA. Science 271: 990-993. [DOI] [PubMed] [Google Scholar]
- Kim D.Y. and Kim, K.K. 2003. Crystal structure of ClpX molecular chaperone from Helicobacter pylori. J. Biol. Chem. 278: 50664-50670. [DOI] [PubMed] [Google Scholar]
- Kim Y.I., Burton, R.E., Burton, B.M., Sauer, R.T., and Baker, T.A. 2000. Dynamics of substrate denaturation and translocation by the ClpXP degradation machine. Mol. Cell 5: 639-648. [DOI] [PubMed] [Google Scholar]
- Levchenko I., Smith, C.K., Walsh, N.P., Sauer, R.T., and Baker, T.A. 1997. PDZ-like domains mediate binding specificity in the Clp/Hsp100 family of chaperones and protease regulatory subunits. Cell 91: 939-947. [DOI] [PubMed] [Google Scholar]
- Levchenko I., Seidel, M., Sauer, R.T., and Baker, T.A. 2000. A specificity-enhancing factor for the ClpXP degradation machine. Science 289: 2354-2356. [DOI] [PubMed] [Google Scholar]
- Levchenko I., Grant, R.A., Wah, D.A., Sauer, R.T., and Baker, T.A. 2003. Structure of a delivery protein for an AAA+ protease in complex with a peptide degradation tag. Mol. Cell 12: 365-372. [DOI] [PubMed] [Google Scholar]
- Maurizi M.R. and Rasulova, F. 2002. Degradation of L-glutamate dehydrogenase from Escherichia coli: Allosteric regulation of enzyme stability. Arch. Biochem. Biophys. 397: 206-216. [DOI] [PubMed] [Google Scholar]
- Maurizi M.R., Clark, W.P., Katayama, Y., Rudikoff, S., Pumphrey, J., Bowers, B., and Gottesman, S. 1990. Sequence and structure of Clp P, the proteolytic component of the ATP-dependent Clp protease of Escherichia coli. J. Biol. Chem. 265: 12536-12545. [PubMed] [Google Scholar]
- Maurizi M.R., Thompson, M.W., Singh, S.K., and Kim, S.H. 1994. Endopeptidase Clp: ATP-dependent Clp protease from Escherichia coli. Methods Enzymol. 244: 314-331. [DOI] [PubMed] [Google Scholar]
- Mecsas J., Rouviere, P.E., Erickson, J.W., Donohue, T.J., and Gross, C.A. 1993. The activity of σ E, an Escherichia coli heat-inducible σ-factor, is modulated by expression of outer membrane proteins. Genes & Dev. 7: 2618-2628. [DOI] [PubMed] [Google Scholar]
- Miller J.H. 1972. Experiments in molecular genetics. Cold Spring Harbor Press, Cold Spring Harbor, NY.
- Missiakas D., Mayer, M.P., Lemaire, M., Georgopoulos, C., and Raina, S. 1997. Modulation of the Escherichia coli σE (RpoE) heat-shock transcription-factor activity by the RseA, RseB and RseC proteins. M. Microbiol. 24: 355-371. [DOI] [PubMed] [Google Scholar]
- Muffler A., Fischer, D., Altuvia, S., Storz, G., and Hengge-Aronis, R. 1996. The response regulator RssB controls stability of the σ(S) subunit of RNA polymerase in Escherichia coli. EMBO J. 15: 1333-1339. [PMC free article] [PubMed] [Google Scholar]
- Neher S.B., Flynn, J.M., Sauer, R.T., and Baker, T.A. 2003a. Latent ClpX-recognition signals ensure LexA destruction after DNA damage. Genes & Dev. 17: 1084-1089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neher S.B., Sauer, R.T., and Baker, T.A. 2003b. Distinct peptide signals in the UmuD and UmuD′ subunits of UmuD/D′ mediate tethering and substrate processing by the ClpXP protease. Proc. Natl. Acad. Sci. 100: 13219-13224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ortega J., Lee, H.S., Maurizi, M.R., and Steven, A.C. 2004. ClpA and ClpX ATPases bind simultaneously to opposite ends of ClpP peptidase to form active hybrid complexes. J. Struct. Biol. 146: 217-226. [DOI] [PubMed] [Google Scholar]
- Rezuchova B., Miticka, H., Homerova, D., Roberts, M., and Kormanec, J. 2003. New members of the Escherichia coli σE regulon identified by a two-plasmid system. FEMS Microbiol. Lett. 225: 1-7. [DOI] [PubMed] [Google Scholar]
- Rosenfeld J., Capdevielle, J., Guillemot, J.C., and Ferrara, P. 1992. In-gel digestion of proteins for internal sequence analysis after one- or two-dimensional gel electrophoresis. Anal. Biochem. 203: 173-179. [DOI] [PubMed] [Google Scholar]
- Song H.K. and Eck, M.J. 2003. Structural basis of degradation signal recognition by SspB, a specificity-enhancing factor for the ClpXP proteolytic machine. Mol. Cell 12: 75-86. [DOI] [PubMed] [Google Scholar]
- Wah D.A., Levchenko, I., Baker, T.A., and Sauer, R.T. 2002. Characterization of a specificity factor for an AAA+ ATPase: Assembly of SspB dimers with ssrA-tagged proteins and the ClpX hexamer. Chem. Biol. 9: 1237-1245. [DOI] [PubMed] [Google Scholar]
- Wah D.A., Levchenko, I., Rieckhof, G.E., Bolon, D.N., Baker, T.A., and Sauer, R.T. 2003. Flexible linkers leash the substrate binding domain of SspB to a peptide module that stabilizes delivery complexes with the AAA+ ClpXP protease. Mol. Cell 12: 355-363. [DOI] [PubMed] [Google Scholar]
- Walsh N.P., Alba, B.M., Bose, B., Gross, C.A., and Sauer, R.T. 2003. OMP peptide signals initiate the envelope-stress response by activating DegS protease via relief of inhibition mediated by its PDZ domain. Cell 113: 61-71. [DOI] [PubMed] [Google Scholar]
- Wang J., Hartling, J.A., and Flanagan, J.M. 1997. The structure of ClpP at 2.3 A resolution suggests a model for ATP-dependent proteolysis. Cell 91: 447-456. [DOI] [PubMed] [Google Scholar]
- Wang L., Elliott, M., and Elliott, T. 1999. Conditional stability of the HemA protein (glutamyl-tRNA reductase) regulates heme biosynthesis in Salmonella typhimurium. J. Bacteriol. 181: 1211-1219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weber-Ban E.U., Reid, B.G., Miranker, A.D., and Horwich, A.L. 1999. Global unfolding of a substrate protein by the Hsp100 chaperone ClpA. Nature 401: 90-93. [DOI] [PubMed] [Google Scholar]
- Withey J.H. and Friedman, D.I. 2003. A salvage pathway for protein structures: tmRNA and trans-translation. Annu. Rev. Microbiol. 57: 101-123. [DOI] [PubMed] [Google Scholar]
- Wojtkowiak D., Georgopoulos, C., and Zylicz, M. 1993. Isolation and characterization of ClpX, a new ATP-dependent specificity component of the Clp protease of Escherichia coli. J. Biol. Chem. 268: 22609-22617. [PubMed] [Google Scholar]
- Zhou Y. and Gottesman, S. 1998. Regulation of proteolysis of the stationary-phase σ factor RpoS. J. Bacteriol. 180: 1154-1158. [DOI] [PMC free article] [PubMed] [Google Scholar]