Abstract
To obtain insight into pH change-driven molecular dynamics, we studied the higher order structure changes of protein G´e at the molecular and sub-molecular levels in solution by using nanoESI- and IMS-mass spectrometry, CD spectroscopy, and protein chemical modification reactions. We found a dramatic change of the overall tertiary structure of protein G´e when the pH was changed from neutral to acidic, but at the same time secondary structure features remained nearly invariable. Limited proteolysis and surface-topology mapping of protein G´e by fast photochemical oxidation of proteins (FPOP) under neutral and acidic conditions reveal areas where higher order conformational changes occur on the amino-acid residue level. Under neutral solution conditions, lower oxidation occurs for residues of the first linker region, whereas greater oxidative modifications occur for amino-acid residues of the IgG-binding domains I and II. We propose a dynamic model of pH-induced structural changes in which protein G´e at neutral pH adopts an overall tight conformation with all four domains packed in a firm assembly, whereas at acidic pH, the three IgG-binding domains form an elongated alignment and the N-terminal, His-tag-carrying domain unfolds. At the same time the individual IgG-binding domains themselves seem to adopt a more compacted fold. As the secondary structure features are nearly unchanged at either pH, interchange between both conformations is highly reversible, explaining the high reconditioning power of protein G´e-based affinity chromatography columns.
Graphical Abstract
Introduction
The interactions and functionality of the protein G´-IgG affinity system have been well-characterized1–5 because it is an extremely important affinity chromatography tool that is used for purification and enrichment of biomolecules from many biological sources.6–11 This affinity system has numerous applications including in proteome research projects for fractionation of plasma proteins12,13 and in clinical apheresis therapies (e.g., with Rheumatoid Arthritis patients).14 The protein G´-IgG affinity chromatography/enrichment system provides both, strong and specific binding of target molecules for capture as well as fast and nearly complete release of ligands during elution. Switching from binding to release is commonly triggered by changing the pH of the media from neutral to acidic.15
Stabilities and folding dynamics of single domains of protein G´16 under different solvent and temperature conditions have been extensively studied.17–21 High resolution crystal structures of all three single IgG-binding domains of protein G´22,23 and of domain II in complex with the “crystallizable fragment (Fc)” from IgG are known,17,24 providing precise information on the binding surfaces between the two proteins. Yet, the commercially available, recombinantly engineered protein G´ is a four-domain, 26 kDa protein (termed protein G´e25) that strongly binds IgGs at neutral pH but permits nearly quantitative release of the antibodies upon acidification. Shifting back to neutral pH reconstitutes the IgG-binding capacity.15,26 Up to now, the assemblies of the three nearly identical IgG-binding domains plus that of the additional His-tag carrying N-terminal domain of protein G´e relative to each other at neutral pH and the alteration caused by shifting to acidic pH remain unknown.
In this study, we first investigated the pH-dependent conformational changes of protein G´e by using mass spectrometry-based methods including nanoESI-MS and IM-MS that are suitable to determine the higher order structure features of the protein.27–29 We then studied the dependence of the secondary structure of protein G´e on pH by using CD-spectroscopy.30,31 Furthermore, we estimated the boundary between the linker and domain I of protein G´e by using limited proteolysis with trypsin.32 Lastly, surface topology and sub-molecular structure changes upon pH shift were elucidated by FPOP (fast photochemical oxidation of proteins) in combination with mass spectrometric peptide mapping analysis.33
Protein footprinting by fast photochemical oxidation of proteins (FPOP) on the microsecond timescale coupled with mass spectrometry is of increasing interest for elucidation of dynamic structural changes in proteins.34 This method has been successfully employed for studying changes in higher order structures of protein therapeutics.35,36 The hydroxyl radical, which is the labeling reagent in the FPOP method, efficiently probes solvent-exposed reactive amino acid residues owing to its small size (comparable to water molecules) and its ability to form a stable, irreversible covalent modification on reactive amino-acid residues. In the hydroxyl radical labeling procedure, the key is to rapidly initiate and immediately quench the reaction afterwards, so that ideally only a single state of the protein is footprinted. The OH radical reactivity is intrinsic for each amino-acid side chain modulated by its solvent accessibility. FPOP, therefore, allows the interrogation of independent protein conformations that, in the case of protein G´e, are obtained by adjusting pH to either neutral or acidic.37,38
The results of our studies on all hierarchical protein structure levels afford a dynamic model of the pH-dependent conformational changes in protein G´e. The model is consistent with observations and practical experiences that others in laboratories around the world have made with fast binding and elution of the IgG-protein G´e affinity system.
Materials and methods
Materials
Protein G'e was purchased from Sigma (catalog no. P4689-5MG; lot no. SLBB8536V) as lyophilized powder. Formic acid, trifluoroacetic acid, L-glutamine, L-methionine, catalase, and HPLC-grade solvents were obtained from Sigma Aldrich (St. Louis, MO, USA). Trypsin and trypsin / LysC mix were purchased from Promega (Madison, WI, USA).
NanoESI MS acquisition conditions, Ion mobility – mass spectrometry, and CCS calculations
NanoESI MS analyses, nanoESI IMS measurements, and CCS calculations were performed as described in the Supporting information Methods file.32,39,40–42
CD spectroscopy acquisition conditions
Circular dichroism analysis of protein G´e were performed as described in the Supporting Information Methods file.43
Limited digestion of protein G´e with trypsin
Limited proteolysis of protein G´e with trypsin was performed as described in the Supporting Information Methods file.
FPOP labeling of protein G´e and LC-ESI MS analysis of intact proteins
FPOP labeling of protein G´e in 2% aqueous acetic acid (pH 2.5) and in 50 mM aqueous ammonium acetate (pH 7) and LC-ESI MS analyses were performed as described in the Supporting Information Methods file. 45
Enzymatic digestion of FPOP labeled protein G´e
Enzymatic digestion of FPOP labeled protein G´e was performed as described in the Supporting Information Methods file.
LC-nanoESI MS/MS of tryptic peptides after FPOP
From each desalted digestion mixture 5 µL were analyzed on a LTQ Orbitrap XL (Thermo Fisher, San Jose, CA) operated in data-dependent acquisition mode. The sample was loaded via autosampler onto a nano-trap column (Acclaim PepMap100 C18, 5 µm, 100 Å, 100 µm (i.d.) × 2 cm (length) from Thermo Scientific, Waltham, MA USA) for desalting and then were separated on a 15 cm column with a PicoFrit tip (New Objective, Inc., Woburn, MA), custom-packed with C18 reversed-phase material (Magic, 0.075 mm × 150 mm, 5 µm, 100 Å; Michrom Bioresources Inc., Auburn, CA); the chromatograph was a Dionex UltiMate 3000 RSLCnano System. Peptides were eluted by a 95 min gradient with the flow of 260 nL/min coupled to the nanospray source. The gradient started with a linear increase from 2% of solvent B (CH3CN/0.1%FA) to 32% in 65 min, and then to 90% in 13 min, held at 90% B for 5 min, and re-equilibrated to solvent A (H2O/0.1%FA) for 12 min.45 Mass spectra were obtained at high mass resolving power (60,000 FWHM, for ions of m/z 400), and data-dependent scanning was set to “top-six” analysis with a dynamic exclusion window of 12 s to ensure good sampling of each elution peak.
To estimate the oxidation rate of protein G´e samples on a global level the remaining unoxidized fraction was calculated using the following equation:
Where I is the intensity of ion peak of the unmodified protein, Iox is the sum of intensities of ion peaks that correspond to oxidized protein.
Data analysis of oxidative modifications on the peptide level was performed using Mascot Distiller, version 2.5.1.0 (Matrix Science, London, UK) with in-built Thermo Xcalibur software. The chromatographic traces of all the samples were processed, and a list of MS signals corresponding to each peak was created. Only those ion signals that corresponded to peptides which eluted with retention times at which identifiable chromatographic peaks were obtained were considered. Next, tryptic peptides were searched using Mascot software, version 2.2.03 (Matrix Science, London, UK), against an custom-created database containing the sequence of protein G´e. Oxidations on Asp, Phe, His, Trp, Lys, Met, Asn, Pro, Arg, and Tyr were set as variable modifications. Product-ion spectra of all peptide ions were then processed to confirm the assignment and locate the modification sites of oxidized peptides.46
Results
Higher order structure characterization of protein G´e under neutral and acidic pH conditions
NanoESI-MS analysis of protein G´e under acidic conditions (pH 2.5) showed a series of multiply charged ions ranging from [M + 18H]18+ to [M + 27H]27+ with maximum abundance for the [M + 21H]21+ protein ion (Figure 1, Table 1, and Supplemental Figure 1A) in the low m/z range. This “charge structure” has an unfolded (denatured) protein conformation that remains unfolded and/or unfolds more in the gas phase owing to Coulombic repulsion between charged residues. Each of the multiply charged molecular ions from protein G´e was accompanied by satellite ions of the gluconoylated protein G´e. By contrast, we observed under neutral conditions broad protein-ion signals with low charge numbers from [M + 8H]8+ to [M + 11H]11+ with a maximum abundance for [M + 9H]9+ or [M + 10H]10+ in the relatively high m/z range. This observation suggests that the protein has a compact, globular (native) structure (Figure 1 and Supplemental Figure 1B) even in the gas phase. The pH-dependent overall charge structures are consistent with a change in a global tertiary structure of protein G´e in response to the pH shift that goes along with a change in antibody-binding properties of protein G´e. We determined a dissociation constant of 9.4 nM for the protein G´e-IgG interaction in a buffer containing 50 mM Tris at pH 7.6 (Table 1). At neutral pH, strong binding between antibodies and protein G´e occurs, whereas under acidic conditions, antibody binding to protein G´e is almost completely abolished.
Figure 1.
Ion mobility arrival times plotted as functions of charge states and nanoESI mass-spectra of protein G´e. A, B: Protein G´e was sprayed from 2% acetic acid:MeOH (95:5), pH 2.5. The drift time distribution for the 21+ ion (m/z 1239.09) is shown on the right. A series of narrow multiply charged ion signals with high charge numbers are recorded as doublets, indicating unfolded protein conformations. The satellite ion signal (on the right of the doublets) arises from gluconoylation. C, D: Protein G´e was sprayed from 50 mM ammonium acetate, pH 7. The drift time distribution for the 9+ ion (m/z 2909.82) is shown on the right. Broad ion signals with low charge numbers indicate globularly folded protein conformations.
Table 1.
Protein G´e structural analysis arranged according to molecular hierarchy a).
| method | functional / structural features | pH 7 | pH 2.5 | |
|---|---|---|---|---|
| A | thermophoresis b) | antibody - protein G: KD | 9.4 nM | >> |
| affinity chromatography |
antibody binding | yes | no | |
| B | nanoESI-MS | charges of intense molecular ion signals |
8+ to 11+ | 18+ to 25+ |
| ion mobility | collisional cross sections of charged molecular ions |
[Å2] | [Å2] | |
| 8+ | 2540 ± 19 c) | n.d. | ||
| 9+ | 2422 ± 14 c) | n.d. | ||
| 10+ | 2446 ± 13 c) | n.d. | ||
| 19+ | n.d. | 6143 ± 10 d) | ||
| 20+ | n.d. | 6167 ± 24 d) | ||
| 21+ | n.d. | 6165 ± 27d) | ||
| 22+ | n.d. | 6192 ± 26 d) | ||
| 23+ | n.d. | 6261 ± 8 d) | ||
| 24+ | n.d. | 6327 ± 17 d) | ||
| 25+ | n.d. | 6448 ± 12 d) | ||
| 26+ | n.d. | 6541 ± 16 d) | ||
| 27+ | n.d. | 6699 ± 15d) | ||
| C | Limited proteolysis | linker - domain border | cleavage sites | n.a. e) |
| accessible not accessible |
R16, R31, K40 K49, etc. |
|||
| CD spectroscopy f) | secondary structure content | [%] | [%] | |
| α-Helix | 28 ± 2 | 21 ± 2 | ||
| β-sheet | 47 ± 3 | 52 ± 2 | ||
| random/turn | 22 ± 2 | 27 ± 1 | ||
| D | FPOP g) | surface topology of domains | oxidized residues | oxidized residues |
| N-terminal domain | M20, M23, M 29 | M 20, M 23, M 29 | ||
| linker | n.d. | Y 48 | ||
| IgG-binding domain I | W 88, Y 90 | W 88 | ||
| spacer 1 | P 113 | P 113 | ||
| IgG-binding domain II | W 158, Y 160 | W 158 | ||
| spacer 2 | P 183 | P 183 | ||
| IgG-binding domain III | W 228 | W 228 | ||
A: quaternary / supra-molecular structure level; B: tertiary / molecular structure level; C: secondary / sub-molecular structure level; D: primary structure level / surface topology of amino acid residues
see ref. 25
average of 3 measurements at wave velocities 700, 800, and 900 m/s
average of 3 measurements at wave heights 30, 35, and 40 V
n.a.: not analyzed
average of 6 measurements
amino acid residues with differences in oxidation are printed in bold
We then determined the rotationally averaged collisional cross sections (CCS) for those ions whose abundances are sufficient to allow accurate recording of arrival times by gas-phase ion mobility measurements (Table 1). When protein G´e was sprayed from acidic solution, it exhibits an averaged CCS of 6353 ± 20 Å2 which is consistent with a dominant, nearly completely unfolded protein conformation that becomes slightly more extended with each additional charge (Figure 1 and Table 1). In contrast, ion mobility measurements of protein G´e sprayed from neutral solvents resulted in an average CCS of 2469 ± 15 Å2 according to ion-mobility arrival times of ions with charges 8+, 9+, and 10+, for which only one single conformation per charge was observed. This CCS value is approximately three times smaller than that obtained for the high charge states and, therefore, is assigned to a compact (i.e., globularly folded) conformation of protein G´e. There is no further collapse of the protein structure with additional decrease of charge. It should be noted, that for charge states 11+ up to 24+, multiple coexisting gas-phase conformations of protein G´e exist (Supplemental Figure 2).
We then analyzed the secondary structure compositions of protein G´e in acidic and neutral solutions by CD spectroscopy. Surprisingly, the CD curves (Supplemental Figure 3) and the content of the secondary structure elements α-helix, β-sheet, and turns/random coils (Table 1) are almost the same under both pH conditions. We observed only a marginal decrease in α-helical content (from 27.7% to 21.2%) that was compensated by a slight increase in β-sheet and turn/random coil content of protein G´e upon shifting the pH from neutral to acidic. A plausible interpretation (see also next paragraph) is that the N-terminal α-helix in the His-tag-containing domain is lost under acidic conditions. From this result, we conclude that the secondary structure elements of the IgG-binding domains of protein G´e do not change dramatically upon pH-shift and ruled out loss of substantial secondary structure elements during pH drop.
Linker–domain border determination by limited proteolysis
Limited proteolysis of protein G´e with trypsin showed cleavages in the N-terminal region covering the His-tag-carrying N-terminal domain only (Figure 2). NanoESI MS analysis of this digest that was sprayed at neutral pH (Supplemental Figure 4) showed two major series of ions (“a” and “b”) with the dominant ions carrying few charges (maximum abundances were for the [M + 9H]9+ and [M + 10H]10+ ions). As an example, proteins with [M + 8H]8+ ion signals possess compactly folded structures (Supplemental Table 3) of the truncated protein G´e derivatives with CCS values of 2219 Å2 and 2317 Å2 for series “a” and “b”, respectively. These CCS values are slightly smaller than that of the compactly folded, intact protein G´e (Table 1) and are consistent with truncation of the protein. Interestingly, the shorter trypsin-derived fragment “b” presented a somewhat larger CCS value at each of the charge states in comparison with the longer fragment “a”. Both truncated protein G´e derivatives show steep increases in CCS values with increasing charge states. For instance, charge state CCS values of species “a” and “b” for both [M + 10H]10+ species are 3288 Å2 and 3845 Å2, respectively, as compared to the CCS values of 2470 ± 15 Å2 for the [M + 8H]8+, [M + 9H]9+, and [M + 10H]10+ ions of non-truncated, intact protein G´e. This increase of the CCS values for the smaller truncated protein G´e fragments can be explained by a new orientation of the truncated N-terminal domain with respect to the compactly folded assembly of the three IgG-binding domains. In the full-length protein G´e, the N-terminal region can form a compact structure together with the IgG-binding domains, whereas for truncated protein G´e derivatives, the N-terminal fragments may move away from the tightly folded assembly owing to fewer molecular interactions, yielding higher CCS values of these species. Noteworthy, when the pH of the solution containing trypsin-derived fragments was changed to acidic, a direct correlation of CCS values with the length of the molecule was observed. For instance, the [M + 20H]20+ charge state ions of full-length protein G´e, of fragment “a”, of fragment “b”, and of fragment “c” present decreasing cross sections of 6167 Å2, 5893 Å2, 5593 Å2, and 5422 Å2, respectively. Thus, limited proteolysis shows that the three IgG-binding domains remain stable under the applied digestion conditions, whereas the N-terminus is prone to facile enzymatic digestion. That the digestion occurred at R16, R31, and K40 proves that these residues exist in a rather unstructured and easily accessible region of protein G´e. By contrast, lack of cleavage at K49 and at tryptic cleavage sites thereafter (20 more in total) indicates that K49 and the subsequent lysine residues are shielded from protease attack, most likely because they are part of the compactly folded and firmly packed IgG-binding domains.
Figure 2.
Protein G´e amino acid sequence in single letter code. Oxidizable amino acid residues (M, Y, W, and P) are shown in bold. An N-terminal α-N-gluconoylation or α-N-6-phosphogluconoylation is indicated by “#”. Tn: numbers of tryptic peptides. Domains and higher order structure details are indicated with boxes and lines below the sequence. N: N-terminal region with His-tag; L: linker; I, II, III: IgG binding domains; S1, S2: spacer regions. Dashed boxes show regions that directly interact with the Fc parts of IgG. Fragments upon limited proteolysis with trypsin are dsignated “a”, “b” and “c” (cf. Supplemental Figure 4).
Sub-molecular structure/surface topology characterization of protein G´e under neutral and acidic pH conditions
FPOP allows insights into changes on different sub-structures of protein G´e, because oxidizable amino acid residues are distributed all over this protein. The N-terminal domain contains three M residues (Figure 2), and the three IgG-binding domains (I, II, and III) contain W and Y residues. Oxidizable amino-acid residues in the sequence of protein G´e are dispersed along the protein such that they also allow insights into the linker between the N-terminal domain and the first IgG-binding domain that contains a Y residue. Finally, the two spacer regions between the first and the second as well as between the second and the third IgG-binding domains contain oxidizable P residues.
We analyzed oxidation extents of protein G´e under neutral (pH 7) and acidic (pH 2.5) conditions by determining the molecular masses and abundances of the intact proteins by using nanoESI-MS directly after FPOP labeling (Supplemental Figure 5). We observed multiply charged ions of the unmodified protein G´e (ion series 1) and its oxidation products (ion series 2–4) as well as for the ions of the gluconoylated protein G´e (ion series 5) and its oxidation products (ion series 6 and 7). The FPOP-derived oxidation yields were higher under both pH conditions (ca. 20% remained unoxidized) as compared to the oxidation yields from chemical oxidation alone (no laser, ca. 60% remained unoxidized after incubation in H2O2). Interestingly, oxidation extents are similar when protein G´e is dissolved at neutral pH (81.1% oxidized) and at acidic pH (80.7% oxidized); substantiating the presence of secondary structure elements in solution also at low pH.
Next, we enzymatically digested oxidized protein G´e samples with trypsin/Lys-C and separated and analyzed the resulting peptides by LC/MS. The elution profiles of both, protein G´e oxidized under acidic or neutral conditions are similar: ca. 18 peaks, each (Figure 3). We integrated the mass spectra under each peak and found the clearest differences with respect to oxidized peptides with peak 11, which was present only in the elution profile of protein G´e and corresponds to oxidized material formed at neutral pH, and with peak 8, which represents more oxidized components when protein G´e was oxidized under acidic conditions (Supplemental Figure 6 and Supplemental tables 4–6).
Figure 3.
Chromatographic traces of nanoLC-separated tryptic peptides from protein G´e. A: After FPOP of protein G´e in 50 mM ammonium acetate, pH 7. B: After FPOP of protein G´e in 2% acetic acid, pH 2.5. Peak numbers with differential oxidation results between the two experiments are printed in bold.
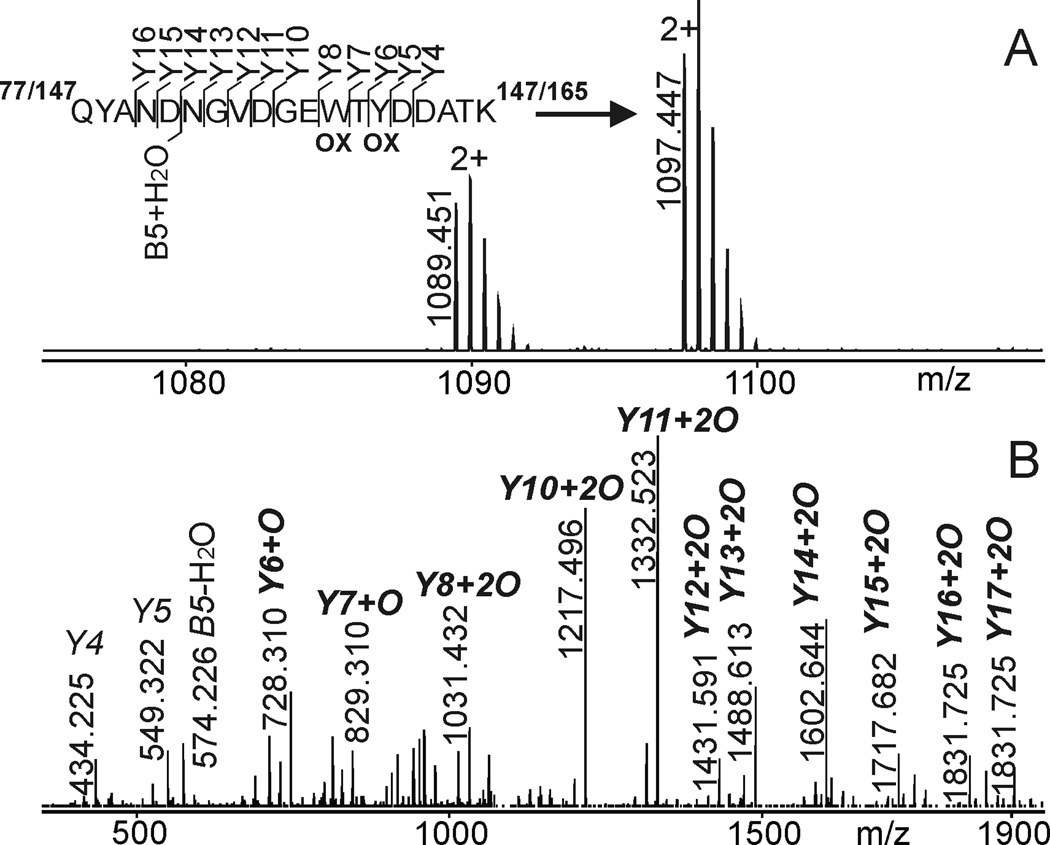
Peak 11 of the nanoLC-ESI-MS run represents the oxidized tryptic peptides T9 (from domain I) and/or T15 (from domain II) upon FPOP at neutral pH (Figure 4). As both peptides have the same amino acid sequence, they cannot be distinguished. The mass spectrum shows doubly charged peptide ions of m/z 1089.451 (in lesser abundance) for the singly oxidized and at m/z 1097.447 (higher abundance) for the doubly oxidized peptides. Product ions from the precursor ion at m/z 1097.447 are B-type and Y"-type fragment ions that pinpoint the oxidation sites on W88 and Y90 (from domain I) and/or W158 and Y160 (from domain II). In addition, peptides T9 and/or T15 in peaks 10 undergo just one oxidation upon FPOP of the W residues only. It should be mentioned that non-oxidized peptides T9 and/or T15 were found in chromatographic peaks 12 of both LC runs (Supplemental Figure 7). Chromatographic peak 13 represents a mixture of peptides with the “WTY” motif from the third IgG-binding domain of protein G´e consisting of non-oxidized T22 (doubly charged peptide ion signal at m/z 1066.464) as well as singly oxidized and deamidated T22 (triply charged peptide ion of m/z 716.988) with oxidation on W228. Whereas oxidation at W on all IgG-binding domains appears similar no matter the pH of the protein G´e solution, there is less oxidation at the Y residues under acidic conditions in the first and second domains. Both the W and the Y residues are located in the middle of their respective IgG-binding domains. This result indicates that the IgG-binding domains adopt a more compact conformation under acidic conditions as compared to their conformation at neutral pH.
Figure 4.
Peak 11 of the nanoLC-ESI-MS run of protein G´e-derived tryptic peptides after FPOP in 50 mM ammonium acetate (cf. Figure 3A). A: The mass spectrum shows ions of m/z 1089.451 and 1097.447, which are assigned to peptides T9 and/or T15 with one and two oxidations, respectively. For ion signal assignments see supplemental table 5. B: MS/MS fragment ions from the precursor ion at m/z 1097.447. The corresponding amino acid sequence is shown in the insert and oxidation sites on W88 and/or W158 and Y90 and/or Y160 are depicted together with the fragment ion type assignments. B-type and Y"-type fragment ion signals are labeled; Y"n are labeled as Yn for simplicity.
Under acidic conditions, FPOP-oxidized peptide T4, which represents the linker between the N-terminal domain and the first IgG-binding domain, elutes at a retention time of 1613 s represented by chromatographic peak 8 and gives rise to an ion of m/z 498.273 (Supplemental Figure 8). MS/MS fragmentation of this ion yields B and Y" fragment ions, from which the position of the oxidation can be narrowed down to “YK”, and, because trypsin does not cleave at oxidized K residues, oxidation was assigned to Y48. It should be noted that Y48 is not oxidized when protein G´e is at neutral pH (Supplemental Figure 9), indicating that at least partial unfolding of the border region between the linker and IgG-binding domain I occurs when this residue becomes more exposed under acidic conditions. With most of the secondary structural features of the IgG-binding domains remaining intact at acidic conditions, an increase of oxidation at an amino acid residue may occur by a conformational rearrangement of still mostly globularly folded protein G´e domains with respect to each other and exposure of a previously buried residue of the linker border.
Analyzing potential pH-induced conformational changes on the two spacer regions S1 and S2, which connect the IgG-binding domains, is possible by monitoring oxidation on peptides T10 (between IgG-binding domain I and II) and/or T16 (between IgG-binding domain II and III); the sequences of the spacer peptides are identical. Chromatographic peaks 14 of both chromatographic runs represent the oxidized peptides (triply-charged peptide ion signal at m/z 886.126; Supplemental Figure 10). Fragment B- and Y"-type ions point to oxidation of these peptides on P113 and P183, respectively. Peak 15 represents the non-oxidized spacer peptides (Supplemental Figure 11). As the oxidation results are the same in both FPOP experiments, we conclude that the spacer sequences in between the globularly folded IgG-binding domains are accessible in a comparable manner irrespective of whether the protein is dissolved in neutral or acidic solutions.
The mass spectra of the ions from chromatographic peaks 1–4 showed a quadruply charged peptide ion that corresponds to peptide T2–T3 with double oxidation at neutral pH (Supplemental Figure 12). Fragmentation of the peptide of m/z 625.018 revealed a mixture of oxidations at two methionine amino acid residues (M20, M23 and / or M29). The mass spectrum of the component corresponding to chromatographic peak 6 shows the quadruply charged non-oxidized peptide T2–T3 ion signal (Supplemental Figure 13). It should be noted that oxidation of M residues was already quite prominent with exposure of protein G´e to H2O2 (i.e., without laser-induced FPOP (Supplemental Table 4)). Nevertheless, the amount of M oxidation increased when FPOP was used and remained nearly unchanged no matter whether protein G´e was dissolved in either acetic acid (acidic pH) or in ammonium acetate (neutral pH). Oxidation of M20, M23, and M29 residues (Supplemental tables 4–6) may be favored by their intrinsic reactivities. Further, because these residues are located on the N-terminus of the flanking region of protein G´e, which carries the His-tag, they should extrude from the protein surface and remain accessible to oxidation under all tested conditions.
Model of protein G´e conformational changes
Taking all our experimental data together, we propose that protein G´e adopts under neutral conditions an overall “compactly folded” conformation in which all three independently globularly folded IgG-binding domains plus the somewhat less tightly folded N-terminal His-tag containing domain are assembled more or less at the corner positions of a tetrahedron (Figure 5). Linkers and spacers connect the domains that are positioned such that the IgG-binding sequences are exposed on the domain surfaces and face towards the solvent. In such a domain assembly, some amino-acid residues from the domains remain exposed on the surface of not only the domains but also of the whole molecule, whereas other residues, although on the surface of their respective domains, participate in forming a “buried core” in the center of the molecule assembled as a tetrahedron.
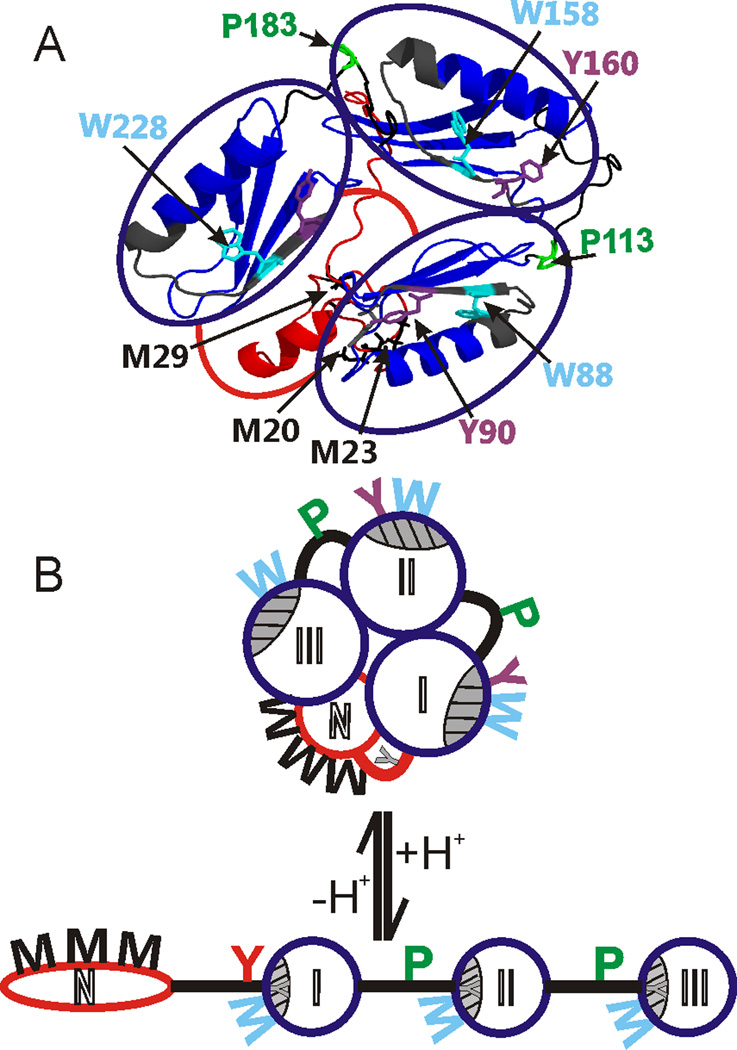
Figure 5.
A: Ribbon structure model of protein G´e. Domain assemblies at neutral pH. IgG-binding domains are circled in blue. Dark grey: IgG-binding surfaces. Red: N-terminal domain. Oxidized amino acid residues in the N-terminal domain, the three IgG binding domains, and linker and spacer peptides connecting the domains are shown in colored wireframes (black: M; purple: Y; cyan: W; green: P). B: Suggested models of protein G´e domain assemblies at neutral pH (top) and at acidic pH (bottom). Open circles represent the N-terminal domain (N; red), that is shown to be largely unstructured under acidic pH conditions, and the three IgG binding domains (I, II, III; blue), that maintain their secondary structure features under neutral and acidic conditions but become tighter at low pH. Thick lines represent linker (red) and spacer (black) peptides connecting the domains. Grey shaded and hatched areas depict IgG-binding surfaces. Amino acid residues (M, Y, W, P) are depicted to extrude from the structure cartoons when oxidized and buried in the interior when not oxidized (not accessible).
Partial unfolding of protein G´e under acidic conditions (e.g. to afford a nearly linear line-up of domains) merely requires rearrangement of the still compactly folded domains (e.g. by “stretching” linkers and spacers). This process increases the exposure of oxidizable amino-acid residues that are located in these linkers and spacers and were previously buried (here Y48, which is considered to be located at the border of the linker and IgG-binding domain I). The domain surfaces that previously participated in forming the “buried core”, however, are now prone to rearrange within their domains to prevent complete unfolding of the latter. As a result, the individual domain structures may become condensed, causing two effects: (i) previously exposed amino-acid residues of the domains diminish in accessibility (here Y90 and/or Y160) and (ii) the IgG-binding surfaces on these domains shrink or disappear by conformational rearrangements. Ultimately, previously bound IgG molecules at neutral pH will be released when the solution becomes acidic because conformational rearrangements occur on protein G´e.
The results from all our structure-related analyses of protein G´e indicate that together with the global structural changes, the surface topology of distinct molecule areas become substantially altered whereas most of the secondary structure features are nearly unchanged under acidic conditions as compared to neutral conditions. Because the protein has these structural properties, reconditioning/regeneration can be easily achieved.
Discussion
Although X-ray crystallography provides structural resolution at the atomic level, it presents a snapshot of proteins and protein complexes in their “folded” solid states.47 Most protein interactions, however, involve dynamic rearrangements of their conformations48 in solution as does protein G´e when it switches from the IgG-bound to the unbound state. Deepening our understanding of the underlying principles of the “affinity switch” between protein G´ and antibodies will be helpful for further development and designing of milder and more efficient elution conditions that might prevent (irreversible) denaturation of the interacting partners and allow longer regeneration of immuno-affinity columns.49
Given that FPOP seems to modify amino acid residues with nearly equal reactivity independent of pH38, we were able to differentiate oxidizable amino acid residues into two categories with respect to their accessibility by applying FPOP to protein G´e at neutral or acidic pH. Category 1 encompasses those amino acid residues whose oxidation by FPOP does not change despite shifting pH, and category 2 contains those whose oxidation by FPOP does change when shifting pH. In category 1, we found methionine residue side chains. In case of protein G´e, M residues, which are part of the rather loosely structured N-terminal domain, are readily chemically modified even prior to FPOP, indicating they are easily accessible independent of pH. Similarly, that the proline residues of protein G´e become oxidized at both pH states indicates good solvent accessibility of the spacer regions to which these residues belong, independent of the pH chosen here. Tryptophan residues are known to be easily oxidizable even with relatively low solvent exposure.50,51 This (photo)chemical behavior explains their rather facile oxidation in protein G´e, independent of pH and despite their involvement in formation of the hydrophobic cores of their respective single domains even though they should be relatively well shielded from the solvent.
Category 2 amino acid side chains are those whose oxidation yields are different when comparing the two pH states in which protein G´e was oxidized. They are comprised of tyrosine residues from different regions/domains of the protein. As expected, oxidizability of Y48 increased at acidic vs. neutral pH because protein G´e is denatured at pH 2.5. The fact that Y48 is thought to belong to the IgG-binding domain I is also supported by our limited proteolysis results. Most interestingly, the oxidation behavior of the Y90 and Y160 residues is totally different. Both tyrosine residues are part of their IgG-binding domains and involved in forming the respective domain structures. Interestingly, the side-chain hydroxyl groups of Y90, Y160, and Y230 are involved in hydrogen bonds with D92, D162, and D232, respectively, and these hydrogen bonds remain “native-like” even when 90% of the protein is unfolded.18 We suggest that steric effects explain the diminished oxidation propensity of Y90 and Y160 under acidic conditions, assuming that the individual domains condense somewhat under these conditions owing to the assumed domain line-up assembly in which the hydrophobic cores need to be “hidden”, in response to opening up the overall globular conformation. In addition, inductive electronic effects that should occur in the aromatic ring of the tyrosine residues, owing to formation of a hydrogen bond between the tyrosines´ hydroxyl groups and the side chains of aspartic acid residues, may also influence the oxidation propensity of these tyrosine residues.
At neutral pH, the carboxyl groups of the aspartic acid side chains should be mostly deprotonated and function as good hydrogen bonders with the tyrosine hydroxyl groups. As a consequence, the negative inductive effect of the aromatic ring becomes stronger resulting in higher oxidation at the ortho position of the aromatic tyrosine ring. On the contrary, under acidic conditions, the carboxyl groups of the aspartic acid residues should be mostly protonated and then they may function as donors of hydrogen bonds between Y and D residues. As a consequence, the negative inductive effect in the aromatic ring is weakened, and less oxidation occurs at the ortho positions.
Of note, the third domain, which possesses a slightly different amino acid sequence from two other domains, possesses a glutamic acid (E) residue in position 209 (instead of an A in domains I and II) whose side-chain carboxyl group is located in close spatial proximity to the hydroxyl group of Y230. Thus, in domain III, under neutral solvent conditions, the carboxyl group of E209 may serve as an additional proton donor to Y230, reducing the negative inductive effect in the aromatic ring of the Y230 residue and diminishing the oxidation of the Y230 residue.
All in all, the dynamic model that we propose assumes an overall change of the assembly of the IgG-binding domains and unfolding of the N-terminal domain of protein G´e depending on pH. On the basis of our results, both “global” and at the residue level, we propose that protein G´e changes its overall globular conformation at neutral pH to adopt an elongated conformation when the pH decreases. Although the secondary structure architecture of its domains does not change dramatically, it still condenses or becomes “tighter”. As a result of the molecular reorientation, the IgG-binding surfaces on each IgG-binding domain undergo steric reorganization. In addition, residues that can be protonated on the IgG-binding surfaces may change their interactions when the pH changes from neutral to acid. Both effects taken together explain, at least in part, the antibody release from protein G´e under acidic conditions and fast and almost completely reversible reconditioning of protein G´e on affinity columns when changing back to neutral pH.
Supplementary Material
Acknowledgments
We express our thanks to Dr. Wolfgang Baumann and Ms. Anja Koenig for their help with CD spectroscopy measurements and to Mr. Kreutzer for his invaluable support with data analysis. We acknowledge the European Union IRSES grant “MS-LIFE” for researcher exchange (PIRSES269256), German Academic Exchange Service (DAAD) for providing scholarships for Y. Yefremova and K. Opuni. Research at Washington University in St. Louis was supported by the US NIH, NIGMS (Grant P41GM103422).
Abbreviations
- Protein G´
protein G prime
- Protein G´e
protein G prime e (extended)
- IgG
immunoglobulin G
- Fc
crystallizable fragment
- CD
circular dichroism
- DC
direct current
- RF
radio frequency
- ESI
electrospray ionization
- EDC
enhanced duty cycle
- CCS
collisional cross section
- UV
ultra violet
- FPOP
fast photochemical oxidation of proteins
- HPLC
high performance liquid chromatography
- ToF
time of flight
- TW-IMS
traveling-wave ion mobility spectroscopy
- Q
quadrupole
- FA
fluoroacetic acid
- ACN
acetonitrile
- MeOH
methanol
- Kd
dissociation constant
References
- 1.Akerström B, Björck L. J. Biol. Chem. 1986;261:240–247. [PubMed] [Google Scholar]
- 2.Akerström B, Nielsen E, Björck L. J. Biol. Chem. 1987;262:13388–13391. [PubMed] [Google Scholar]
- 3.Alexander P, Fahnestock S, Lee T, Orban J, Bryan P. Biochemistry. 1992;31:3597–3603. doi: 10.1021/bi00129a007. [DOI] [PubMed] [Google Scholar]
- 4.Guss B, Eliasson M, Olsson A, Uhlén M, Frej AK, Jörnvall H, Flock JI, Lindberg M. EMBO J. 1986;5:1567–1575. doi: 10.1002/j.1460-2075.1986.tb04398.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kronvall G. J. Immunol. 1973;111:1401–1406. [PubMed] [Google Scholar]
- 6.Akerstrom B, Brodin T, Reis K, Bjorck L. J. Immunol. 1985;135:2589–2592. [PubMed] [Google Scholar]
- 7.Björck L, Blomberg J. Eur. J. Clin. Microbiol. 1987;6:428–429. doi: 10.1007/BF02013104. [DOI] [PubMed] [Google Scholar]
- 8.Boström TNJ, Hober S. In: Protein Purification. Ahmad R, editor. InTech; 2012. p. 224. [Google Scholar]
- 9.Crowley SC, Walters RR. J. Chromatogr. 1983;266:157–162. doi: 10.1016/s0021-9673(01)90888-x. [DOI] [PubMed] [Google Scholar]
- 10.Dancette OP, Taboureau JL, Tournier E, Charcosset C, Blond P. J. Chromatogr. B. 1999;723:61–68. doi: 10.1016/s0378-4347(98)00470-8. [DOI] [PubMed] [Google Scholar]
- 11.Faulkner S, Elia G, Hillard M, O'Boyle P, Dunn M, Morris D. Proteomics. 2011;11:2329–2335. doi: 10.1002/pmic.201000364. [DOI] [PubMed] [Google Scholar]
- 12.Echan LA, Tang HY, Ali-Khan N, Lee K, Speicher DW. Proteomics. 2005;5:3292–3303. doi: 10.1002/pmic.200401228. [DOI] [PubMed] [Google Scholar]
- 13.Omenn GS, States DJ, Adamski M, Blackwell TW, Menon R, Hermjakob H, Apweiler R, Haab BB, Simpson RJ, Eddes JS, Kapp EA, Moritz RL, Chan DW, Rai AJ, Admon A, Aebersold R, Eng J, Hancock WS, Hefta SA, Meyer H, Paik YK, Yoo JS, Ping P, Pounds J, Adkins J, Qian X, Wang R, Wasinger V, Wu CY, Zhao X, Zeng R, Archakov A, Tsugita A, Beer I, Pandey A, Pisano M, Andrews P, Tammen H, Speicher DW, Hanash SM. Proteomics. 2005;5:3226–3245. doi: 10.1002/pmic.200500358. [DOI] [PubMed] [Google Scholar]
- 14.Kienbaum M, Koy C, Montgomery HV, Drynda S, Lorenz P, Illges H, Tanaka K, Kekow J, Guthke R, Thiesen HJ, Glocker MO. Proteom. Clin. Appl. 2009;3:797–809. doi: 10.1002/prca.200800232. [DOI] [PubMed] [Google Scholar]
- 15.Zachariou M. Methods Mol. Biol. 2008;421:vii–viii. doi: 10.1007/978-1-59745-582-4. [DOI] [PubMed] [Google Scholar]
- 16.Goward CR, Murphy JP, Atkinson T, Barstow DA. Biochem. J. 1990;267:171–177. doi: 10.1042/bj2670171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Gronenborn AM, Clore GM. J. Mol. Biol. 1993;233:331–335. doi: 10.1006/jmbi.1993.1514. [DOI] [PubMed] [Google Scholar]
- 18.Khare D, Alexander P, Antosiewicz J, Bryan P, Gilson M, Orban J. Biochemistry. 1997;36:3580–3589. doi: 10.1021/bi9630927. [DOI] [PubMed] [Google Scholar]
- 19.Lindman S, Xue WF, Szczepankiewicz O, Bauer MC, Nilsson H, Linse S. Biophys. J. 2006;90:2911–2921. doi: 10.1529/biophysj.105.071050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kmiecik S, Kolinski A. Biophys. J. 2008;94:726–736. doi: 10.1529/biophysj.107.116095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Park SH, Shastry MCR, Roder H. Nat. Struct. Biol. 1999;6:943–947. doi: 10.1038/13311. [DOI] [PubMed] [Google Scholar]
- 22.Derrick JP, Wigley DB. J. Mol. Biol. 1994;243:906–918. doi: 10.1006/jmbi.1994.1691. [DOI] [PubMed] [Google Scholar]
- 23.Schmidt HLF, Sperling LJ, Gao YG, Wylie BJ, Boettcher JM, Wilson SR, Rienstra CA. J. Phys. Chem. B. 2007;111:14362–14369. doi: 10.1021/jp075531p. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Sauer-Eriksson AE, Kleywegt GJ, Uhlén M, Jones TA. Structure. 1995;3:265–278. doi: 10.1016/s0969-2126(01)00157-5. [DOI] [PubMed] [Google Scholar]
- 25.Yefremova Y, Al-Majdoub M, Opuni KF, Koy C, Cui W, Yan Y, Gross ML, Glocker MO. J. Am. Soc. Mass Spectrom. 2015;26:482–492. doi: 10.1007/s13361-014-1053-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Al-Majdoub M, Opuni KFM, Yefremova Y, Koy C, Lorenz P, El-Kased RF, Thiesen HJ, Glocker MO. J. Mol. Recognit. 2014;27:566–574. doi: 10.1002/jmr.2375. [DOI] [PubMed] [Google Scholar]
- 27.Konijnenberg A, Yilmaz D, Ingolfsson HI, Dimitrova A, Marrink SJ, Li Z, Venien-Bryan C, Sobott F, Kocer A. Proc. Natl. Acad. Sci. U. S. A. 2014;111:17170–17175. doi: 10.1073/pnas.1413118111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Loo JA, Berhane B, Kaddis CS, Wooding KM, Xie Y, Kaufman SL, Chernushevich IV. J. Am. Soc. Mass Spectrom. 2005;16:998–1008. doi: 10.1016/j.jasms.2005.02.017. [DOI] [PubMed] [Google Scholar]
- 29.Przybylski M, Glocker MO. Angew. Chem. Int. Ed. Engl. 1996;35:807–826. [Google Scholar]
- 30.Greenfield NJ. Nat. Protoc. 2006;1:2876–2890. doi: 10.1038/nprot.2006.202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kelly SM, Price NC. Curr. Protein Pept. Sci. 2000;1:349–384. doi: 10.2174/1389203003381315. [DOI] [PubMed] [Google Scholar]
- 32.Bantscheff M, Glocker MO. Eur. Mass Spectrom. 1998;4:279–285. [Google Scholar]
- 33.Zhang H, Wen J, Huang RY, Blankenship RE, Gross ML. Int. J. Mass Spectrom. 2012;312:78–86. doi: 10.1016/j.ijms.2011.07.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gau BC, Sharp JS, Rempel DL, Gross ML. Anal. Chem. 2009;81:6563–6571. doi: 10.1021/ac901054w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Jones LM, Zhang H, Cui W, Kumar S, Sperry JB, Carroll JA, Gross ML. J. Am. Soc. Mass Spectrom. 2013;24:835–845. doi: 10.1007/s13361-013-0582-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zhang Y, Rempel DL, Zhang H, Gross ML. J. Am. Soc. Mass Spectrom. 2015;26:526–529. doi: 10.1007/s13361-014-1055-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Hambly DM, Gross ML. In: Comprehensive Analytical Chemistry. Whitelegge JP, editor. Elsevier BV; 2009. pp. 151–177. [Google Scholar]
- 38.Vahidi S, Stocks BB, Liaghati-Mobarhan Y, Konermann L. Anal. Chem. 2012;84:9124–9130. doi: 10.1021/ac302393g. [DOI] [PubMed] [Google Scholar]
- 39.Happersberger HP, Przybylski M, Glocker MO. Anal. Biochem. 1998;264:237–250. doi: 10.1006/abio.1998.2836. [DOI] [PubMed] [Google Scholar]
- 40.Bush MF, Hall Z, Giles K, Hoyes J, Robinson CV, Ruotolo BT. Anal. Chem. 2010;82:9557–9565. doi: 10.1021/ac1022953. [DOI] [PubMed] [Google Scholar]
- 41.Hofmann J, Struwe WB, Scarff CA, Scrivens JH, Harvey DJ, Pagel K. Anal. Chem. 2014;86:10789–10795. doi: 10.1021/ac5028353. [DOI] [PubMed] [Google Scholar]
- 42.Ruotolo BT, Benesch JL, Sandercock AM, Hyung SJ, Robinson CV. Nat. Protoc. 2008;3:1139–1152. doi: 10.1038/nprot.2008.78. [DOI] [PubMed] [Google Scholar]
- 43.Al-Majdoub M, Opuni KFM, Koy C, Glocker MO. Anal. Chem. 2013;85:10479–10487. doi: 10.1021/ac402559m. [DOI] [PubMed] [Google Scholar]
- 44.Yan YT, Chen GD, Wei H, Huang RYC, Mo JJ, Rempel DL, Tymiak AA, Gross ML. J. Am. Soc. Mass Spectrom. 2014;25:2084–2092. doi: 10.1007/s13361-014-0993-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Zhang H, Gau BC, Jones LM, Vidavsky I, Gross ML. Anal. Chem. 2011;83:311–318. doi: 10.1021/ac102426d. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Gau BC, Chen J, Gross ML. Biochim. Biophys. Acta. 2013;1834:1230–1238. doi: 10.1016/j.bbapap.2013.02.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Henzler-Wildman K, Kern D. Nature. 2007;450:964–972. doi: 10.1038/nature06522. [DOI] [PubMed] [Google Scholar]
- 48.Hensen U, Meyer T, Haas J, Rex R, Vriend G, Grubmuller H. PLoS One. 2012;7:e33931. doi: 10.1371/journal.pone.0033931. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Moser AC, Hage DS. Bioanalysis. 2010;2:769–790. doi: 10.4155/bio.10.31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Hambly DM, Gross ML. J. Am. Soc. Mass Spectrom. 2005;16:2057–2063. doi: 10.1016/j.jasms.2005.09.008. [DOI] [PubMed] [Google Scholar]
- 51.Lins L, Thomas A, Brasseur R. Protein Sci. 2003;12:1406–1417. doi: 10.1110/ps.0304803. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.