Abstract
Patients with advanced congestive heart failure (CHF) or chronic kidney disease often have increased angiotensin II (Ang II) levels and cachexia. We previously demonstrated that Ang II, via its type 1 receptor, causes muscle protein breakdown and apoptosis and inhibits satellite cell (SC) proliferation and muscle regeneration, likely contributing to cachexia in CHF and chronic kidney disease. In contrast, Ang II, via its type 2 receptor (AT2R) expression, is robustly induced during SC differentiation, and it potentiates muscle regeneration. To understand the mechanisms regulating AT2R expression and its potential role in muscle regeneration in chronic diseases, we used a mouse model of CHF and found that muscle regeneration was markedly reduced and that this was accompanied by blunted increase of AT2R expression. We performed AT2R promoter reporter analysis during satellite cell differentiation and found that the 70 bp upstream of the AT2R transcription start site contain a core promoter region, and regions upstream of 70 bp to 3 kbp are dispensable for AT2R induction. Instead, AT2R intron 2 acts as a transcriptional enhancer during SC differentiation. Further deletion/mutation analysis revealed that multiple transcription factor binding sites in the +286/+690 region within intron 2 coordinately regulate AT2R transcription. Importantly, +286/+690 enhancer activity was suppressed in CHF mouse skeletal muscle, suggesting that AT2R expression is suppressed in CHF via inhibition of AT2R intronic enhancer activity, leading to lowered muscle regeneration. Thus targeting intron 2 enhancer element could lead to the development of a novel intervention to increase AT2R expression in SCs and potentiate skeletal muscle regenerative capacity in chronic diseases.
Keywords: angiotensin II, heart failure, muscle regeneration, skeletal muscle, stem cells, transcription enhancer
Introduction
Overactivation of the renin-angiotensin system (RAS)2 contributes to the pathophysiology of many chronic disease conditions. Cachexia, or a wasting syndrome, is typically associated with chronic diseases such as cancer, chronic obstructive pulmonary disease, congestive heart failure (CHF), and chronic kidney disease (CKD), and both basic and clinical studies have suggested a link between the RAS and cachexia development. Penafuerte et al. (1) reported that plasma angiotensin II (Ang II) levels were significantly increased in pre-cachectic and cachectic cancer patients and proposed Ang II as a blood biomarker and the master upstream regulator of pre-cachexia. Also, patients with advanced CHF and CKD often have elevated circulating Ang II and cachexia, and treatment with an angiotensin-converting enzyme inhibitor can reduce weight loss (2). Cachexia is typically associated with skeletal muscle atrophy, and increasing evidence has suggested that elevated levels of Ang II lead to muscle atrophy in chronic disease conditions (3–9). Muscle damage, particularly to the muscle membrane, or sarcolemma, is a feature of muscle atrophy in these conditions. Although muscle damage normally activates skeletal muscle stem (satellite) cells (SCs) and triggers muscle regeneration, many of these conditions in which there is muscle atrophy are associated with reduced regeneration, such as disuse (10–14), denervation (15), chronic obstructive pulmonary disease (16–21), CKD (22, 23), burn injury (24, 25), diabetes (26–31), and cancer (32–36). We recently found that two types of Ang II receptors, type 1 (AT1R) and type 2 (AT2R), regulate different stages of SC differentiation. Quiescent and proliferating SCs express high levels of AT1R, and AT1R signaling inhibits SC proliferation, leading to depletion of the SC pool and reduced muscle regenerative capacity in a cardiotoxin (CTX) injury-induced model (37). In contrast, AT2R expression is robustly increased during SC differentiation both in vitro and in vivo. AT2R positively regulates SC differentiation; thus knockdown of AT2R significantly suppressed SC differentiation in vitro and muscle regeneration in vivo (38). These data strongly suggest that Ang II regulates different stages of the SC differentiation process via AT1R and AT2R and that balancing AT1R and AT2R signaling is critical to maintain muscle regenerative capacity mediated by SC function. Although our data showed that AT2R positively regulates SC differentiation, it is completely unknown how its expression is induced during SC differentiation. Furthermore, it has been shown that AT2R expression is robustly increased in many tissues/cells during embryogenesis, although its expression is below detectable levels in many adult tissues, suggesting that AT2R may regulate differentiation of multiple cell types, including SCs. Thus elucidating mechanisms of AT2R transcription have the potential to lead to the development of novel interventions to restore lowered skeletal muscle regenerative capacity in chronic diseases, and it may lead to identification of mechanisms that regulate differentiation of multiple cell lineages. In this study we aimed to identify mechanisms whereby AT2R expression is increased during SC differentiation.
Results
Skeletal Muscle Regeneration and Induction of AT2R Expression Are Reduced in CHF
We have shown that the RAS plays an important role in causing skeletal muscle wasting in CHF and possibly other chronic disease conditions (3–8, 39–41). Our recent findings indicate that different subtypes of Ang II receptors AT1R and AT2R regulate different stages of SC differentiation and skeletal muscle regeneration, potentially contributing to the development of muscle wasting in CHF (9, 37, 38). However, it is not clear whether skeletal muscle regeneration is impaired in the setting of CHF and whether Ang II signaling contributes to that process. To analyze skeletal muscle regenerative capacity in CHF, we used mice that had undergone left anterior descending artery (LAD) ligation, a myocardial infarction model of CHF. CHF mice had a significant increase in left ventricular end systolic diameter (1.70 ± 0.101 in sham versus 3.35 ± 0.216 in LAD, n = 6, p < 0.01) and left ventricular end diastolic diameter (3.58 ± 0.101 in sham versus 4.53 ± 0.231 in LAD, n = 6, p < 0.01) and a decrease in fractional shortening (51.9 ± 0.88 in sham versus 26.2 ± 1.58 in LAD, n = 6, p < 0.01). The hindlimb muscles of these mice were injured by CTX injection to induce skeletal muscle regeneration (Fig. 1A). Importantly, we found that skeletal muscle regenerative capacity was reduced 4 weeks after LAD ligation. Both the number and size of regenerating myofibers (myofibers with centralized nuclei) were reduced after 7 days of CTX injury (Fig. 1, B and C). Consistent with the reduced muscle regeneration, the expression of myogenin and eMyHC was reduced in CHF mouse hindlimb muscles (Fig. 1, D and E). As we have shown previously (37, 38), AT1R expression was slightly decreased, and AT2R expression was robustly increased during regeneration. In CHF mice, we found that the induction of AT2R expression after CTX injury was markedly blunted, whereas AT1R expression was not altered. We have previously shown that the AT2R positively regulates skeletal muscle regeneration and that knockdown of AT2R suppressed muscle regeneration (38). Taken together, these data strongly suggest that AT2R down-regulation in CHF mice plays a critical role in inhibiting skeletal muscle regenerative capacity.
FIGURE 1.
CHF suppresses skeletal muscle regenerative capacity and AT2R induction. A, experimental design. LAD ligation surgery was performed in C57BL/6 mice, and CTX was injected in gastrocnemius muscles 4 weeks after the surgery. The contralateral muscles were used as non-injury control. Muscles were harvested 3, 5, and 7 days after the CTX injection, and regenerating myofiber number (B) and size (C) and the expression of myogenin (D), eMyHC (E), AT1R (F), and AT2R (G) were analyzed by quantitative RT-PCR. n = 5–6; mean ± S.E.; *, p < 0.05; **, p < 0.01 between sham versus LAD.
Analysis of AT2R Promoter Activity during Myoblast Differentiation
Although our data (Fig. 1) (38) suggest that restoration of AT2R expression could lead to a potential therapy for lowered muscle regeneration in chronic disease such as CHF, currently transcriptional mechanisms regulating AT2R expression are poorly understood. We have shown previously that AT2R mRNA and protein are robustly up-regulated in SCs and C2C12 myoblasts during differentiation in vitro (38). To identify transcription factor (TF)-binding site(s) responsible for AT2R transcriptional activation during myoblast differentiation, we performed a series of AT2R promoter luciferase reporter assays. First, we cloned the 3-kbp DNA fragment upstream of the AT2R transcription start site (TSS, +1) of mouse AT2R together with the sequence from exon 1 to the 5′-untranslated region (UTR) of exon 3 (+1/+1503) into pGL4.10(luc2) vector (AT2R-3000-luc vector, Fig. 2). We also generated luciferase vectors with 2 and 1 kbp and 500 bp of DNA fragments upstream of AT2R TSS (AT2R-2000-luc, AT2R-1000-luc, and AT2R-500-luc vectors, respectively). These vectors were transfected into primary cultured SCs. 2 days after transfection, these cells were induced to differentiate into myotubes, and the luciferase activity was measured after 1–3 days of differentiation. pGL4.74[hRluc/TK] vector, in which Renilla reniformis luciferase is inserted downstream of herpes simplex virus-thymidine kinase promoter, was co-transfected, and luciferase activity was calculated as a ratio of firefly to Renilla luciferase (Fluc/Rluc). As shown in Fig. 2, luciferase activity was increased during SC differentiation in all of the AT2R-luciferase vectors tested. AT2R-1000-luc and AT2R-500-luc vectors showed higher luciferase induction, suggesting that a cis element that suppressed AT2R existed within the −3000/−1000 region of AT2R. We also performed the same luciferase reporter assay in C2C12 myoblasts, and the data showed that AT2R-1000-luc and AT2R-500-luc vectors showed higher luciferase induction compared with AT2R-3000-luc and AT2R-2000-luc as in the case of SCs (Fig. 2). These changes in luciferase activity are consistent with AT2R protein increase in SC/C2C12 during differentiation (representative immunoblot images are shown in Fig. 2). These data suggest that AT2R transcription is regulated by a 500-bp region upstream of AT2R TSS both in SCs and C2C12 myoblasts.
FIGURE 2.
AT2R promoter activity during myoblast differentiation. Mouse AT2R genomic DNA sequence upstream of TSS (+1) together with exon 1, intron 1, exon 2, intron 2, and the 5′-UTR of exon 3 were subcloned into the pGL4.10[luc2] vector. Four vectors were generated with different sized 5′ sequence (3000, 2000, 1000, and 500 bp). These vectors were transfected into SCs and C2C12 myoblasts. After 2 days of transfection, cells were induced to differentiate and harvested on days 0–3, and luciferase activity was measured using Renilla luciferase as an internal control. Mean ± S.E., n = 4; *, p < 0.05; **, p < 0.01 compared with d0. Western blotting images show representative AT2R protein induction in primary cultured SCs and C2C12 myoblasts during differentiation.
AT2R Intron 2 Plays a Critical Role in Regulating AT2R Transcription during Myoblast Differentiation
By TRASFAC database search, we identified four putative TF-binding sites within the −500/+1 region of the AT2R putative promoter as follows: promyelocytic leukemia zinc finger protein at −479; heat shock factor protein-1 (HSF1) at −282; kidney, ischemia, and developmentally regulated gene-3 (KID3) at −240, and E-box at −170 (Fig. 3A). As reported previously for rat AT2R (42), mouse AT2R possesses a TATA box at −30. To identify TF-binding site(s) within this region, we further generated AT2R promoter reporter vectors with truncated 5′-putative promoter regions. Surprisingly, deletion of the −500/−70 region, which contains all of the TF-binding sites above, did not affect luciferase induction during myoblast differentiation. Deletion of the −70/−10 region, which contains the TATA box, significantly suppressed luciferase induction. These data suggest that the −70/−10 region acts as a core promoter of AT2R, whereas regions upstream of −70 are dispensable for AT2R transcriptional activation during myoblast differentiation. Although our data do not exclude the possibility that there is a TF-binding site that is not predicted by database search within the −70/+10 segment, we hypothesized that AT2R transcription is mediated by intronic sequences of AT2R. We generated luciferase reporter vectors with deletions of intron 1 and intron 2 sequences (Fig. 4). The deletion of intron 1 did not alter luciferase induction during myoblast differentiation, whereas intron 2 deletion almost completely abolished luciferase induction (Fig. 4). These data strongly suggest that the AT2R intron 2 sequence acts as a transcriptional enhancer during myoblast differentiation.
FIGURE 3.
AT2R 5′–promoter analysis during myoblast differentiation. A, DNA sequence of AT2R promoter (−500/+40). Four transcription factor-binding sites were predicted by TRANSFAC database search as follows: promyelocytic leukemia zinc finger protein (PLZF) at −479, heat shock factor protein-1 (HSF1) at −282, kidney, ischemia, and developmentally regulated gene-3 (KID3) at −240, and E-box at −170. TATA box and initiator element are located at −30/−23 and +3/+7, respectively. B, series of AT2R promoter reporter vectors were constructed and analyzed by luciferase reporter assay in C2C12 myoblasts/myotubes. All the AT2R luciferase reporter vectors contain exon 1, intron 1, exon 2, intron 2, and the 5′-UTR of exon 3. Numbers indicate the distance from the AT2R TSS (+1). Cells were induced to differentiate 2 days after plasmid transfection, and harvested before (day 0) and 2 days after differentiation (Diff). Luciferase activity was measured using Renilla luciferase as an internal control. Mean ± S.E., n = 4; **, p < 0.01.
FIGURE 4.

AT2R intron 2 regulates AT2R transcription during myoblast differentiation. A series of AT2R promoter reporter vectors were constructed and analyzed by luciferase reporter assay in C2C12 myoblasts/myotubes. AT2R-70-luc vector contains −70/+1503 DNA segment of AT2R upstream of firefly luciferase. Intron 1 and intron 2 were deleted from AT2R-70-luc vector to generate AT2R-70-ΔInt1-luc and AT2R-70-ΔInt2-luc vector. Both intron 1 and 2 were removed from AT2R-70-ΔInt1/2-luc vector. Cells were induced to differentiate 2 days after plasmid transfection and harvested before (day 0) and 2 days after differentiation (Diff). Luciferase activity was measured using Renilla luciferase as an internal control. Mean ± S.E., n = 4; **, p < 0.01 compared with d0 control; §§, p < 0.01 compared with AT2R-70-luc d2.
AT2R Intron 2 Acts as a Transcriptional Enhancer
To further analyze the potential enhancer activity of AT2R intron 2, we generated luciferase reporter vectors with minimal promoter sequence (pGL4.23[luc2/minP] vector, Promega). pGL4.23[luc2/minP] vector contains 31-bp minimal promoter sequence upstream of luciferase, allowing assessment of DNA fragments for their enhancer activities. AT2R intron 2 sequence was subcloned into pGL4.23[luc2/minP] vector, and luciferase activity was measured during myoblast differentiation (Fig. 5). Consistent with our hypothesis, the luciferase vector with the intron 2 showed an increase of luciferase activity during myoblast differentiation, whereas the vector with the AT2R 5′-region (−500/−1) did not show any change.
FIGURE 5.
Analysis of transcriptional enhancer activity in AT2R intron 2. A series of luciferase reporter vectors with minimal promoter and AT2R intron 2 sequence were constructed and analyzed by luciferase reporter assay in C2C12 myoblasts/myotubes. Different segments from the AT2R promoter/enhancer region were subcloned into pGL4.23[luc2/minP] vector, which contains 31-bp minimal promoter sequence upstream of firefly luciferase. The numbers indicate the location of AT2R promoter/enhancer. AT2R-500-luc and pGL4.23[luc2/minP] were used as positive and negative controls, respectively. Cells were induced to differentiate 2 days after plasmid transfection and harvested before (day 0) and 2 days after differentiation (Diff). Luciferase activity was measured using Renilla luciferase as an internal control. Mean ± S.E., n = 4; **, p < 0.01 compared with d0 control.
To identify a region within intron 2 that is responsible for AT2R induction during myoblast differentiation, we next generated a series of luciferase reporter vectors with the minimal promoter and truncated AT2R intron 2 (Fig. 5). Within intron 2, our data showed that the +691/+1080 and +1081/+1468 regions do not have enhancer activity, whereas +286/+690 showed significant increase in luciferase activity during myoblast differentiation. Deletion of +1081/+1468 region increased luciferase activity, suggesting that the +1081/+1467 region has a repressor activity. Consistent with the results in Figs. 3 and 4, the −500/−1 region did not have a transcriptional enhancer activity. Also, luciferase vectors that contain AT2R endogenous TSS (e.g. −500/+285 region) did not show luciferase activity. However, the +286/+1467 region showed significantly lower luciferase activity compared with AT2R-500-luc vector, suggesting that −500/−1 region and intron 2 may cooperatively regulate AT2R transcription. Nonetheless, our data showed that the +286/+690 region contains a critical transcriptional enhancer element that is responsible for AT2R induction during myoblast differentiation.
To further identify the precise enhancer element in intron 2, we generated a series of luciferase vectors with the minimal promoter and a truncated +286/+690 region (Fig. 6). The data showed that the deletion of +286/+375 markedly blunted the luciferase activity. However, +286/+375 region itself did not show luciferase induction. Deletion of independent parts within the +286/+690 region also blocked the luciferase induction (+475/+585 and +585/+690), whereas the deletion of +375/+475 segment did not significantly reduce luciferase activity. These data suggest that the AT2R intron 2 enhancer activity resides not in a single element but in multiple elements within the +286/+690 region.
FIGURE 6.
Analysis of transcriptional enhancer activity in +286/+690 sequence of AT2R intron 2. A series of luciferase reporter vectors with minimal promoter and AT2R intron 2 sequence from the +286/+690 region were constructed and analyzed by luciferase reporter assay in C2C12 myoblasts/myotubes. Different segments from AT2R promoter/enhancer region were subcloned into pGL4.23[luc2/minP] vector, which contains 31-bp minimal promoter sequence upstream of firefly luciferase. The numbers indicate the location of AT2R promoter/enhancer. AT2R (+286/+690)-minP-luc and pGL4.23[luc2/minP] were used as positive and negative control, respectively. Cells were induced to differentiate 2 days after plasmid transfection and harvested before (day 0) and 2 days after differentiation (Diff). Luciferase activity was measured using Renilla luciferase as an internal control. Mean ± S.E., n = 4; *, p < 0.05 compared with d0 control; §§, p < 0.01 compared with AT2R (+286/+690)-minP-luc d2.
To gain insight into the +286/+690 enhancer, we analyzed the evolutionary conservation of AT2R genomic DNA sequences from mouse (Mus musculus, UCSC Gene ID: uc009suq.3), human (Homo sapiens, ENST00000371906.4), pig (Sus scrofa, ENSSSCT00000034178.1), rabbit (Oryctolagus cuniculus, ENSOCUT00000012357.2), and dog (Canis lupus familiaris, ENSCAFT00000029009.3) (supplemental Fig. S1). Although AT2R intron 2 plays a critical role in regulating AT2R transcription during myoblast differentiation and the intron 1 sequence is dispensable, the nucleotide identity within the intron 1 (49.4%) is higher than that of intron 2 (27.7%, supplemental Fig. S1B). Similarly, although our data indicated that mouse AT2R +286/+690 acts as a critical transcriptional enhancer, the nucleotide identity within this region is lower than that of the +691/+1080 region (34.7%), which was dispensable for AT2R induction (Fig. 5). It has been shown that phylogenetic footprinting does not always work (43) due to the short (5–20 bp) length of the regulatory elements relative to the entire regulatory region searched in silico. Depending on the divergence of species compared, it is possible that the noise of the diverged nonfunctional background would overcome the short conserved signal. The supplemental Fig. S1C shows the nucleotide alignment of AT2R +286/+690 corresponding regions from mouse, human, pig, rabbit, and dog. These data show that, although the overall nucleotide identity is low in this region, there are several highly conserved short DNA elements, which may act as TF-binding sites. Potential TF-binding sites within +286/+690 enhancer were analyzed below.
Potential Transcription Factor-binding Sites within AT2R Intron 2
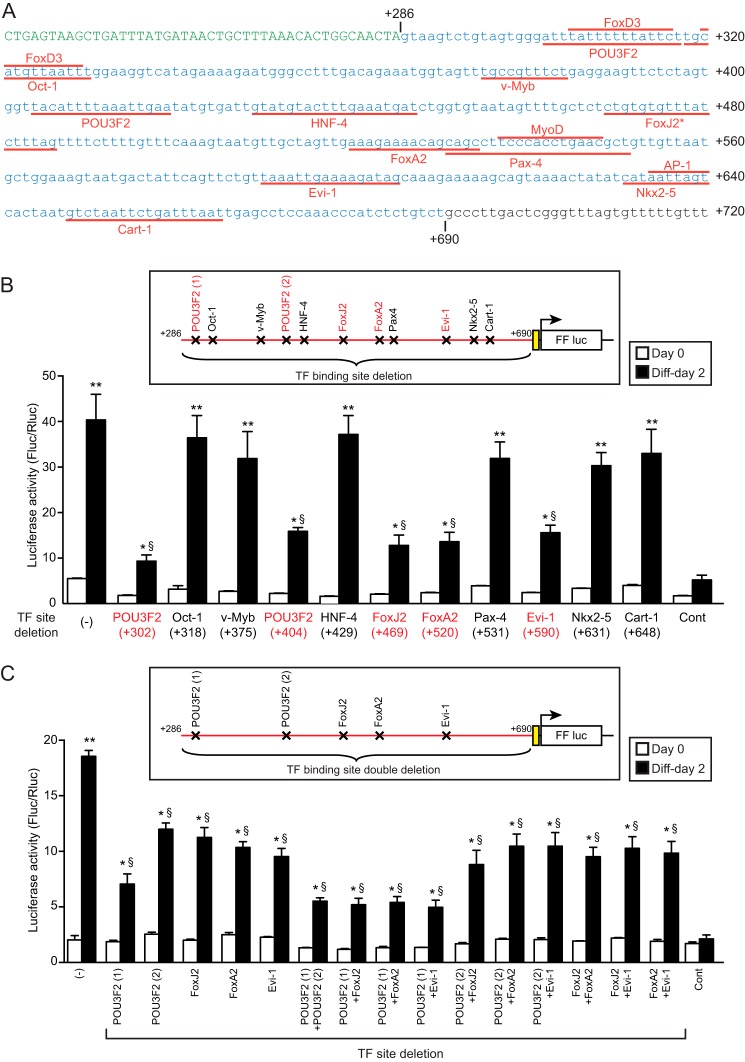
By TRANSFAC (44) database search, we identified 19 putative TF-binding sites within the +286/+690 region (Fig. 7A, it is of note that some of these sites are overlapping) as follows: POU domain, class 3, transcription factor 2 (POU3F2, +302 and +404); FOXD3 (+305 and +320); octamer transcription factor-1 (OCT-1, +318); v-MYB (+375); hepatocyte nuclear factor 4α (HNF-4α, +429); FoxJ2 (+469); FoxL1 (+470); FoxI1 (+471); FOXF1 (+471); GATA (+472); hepatocyte nuclear factor 3β (HNF-3β, +520); PAX-4 (+531); MYOD (+537); ecotropic virus integration site 1 protein homolog (EVI-1, +478 and +590); NK2 transcription factor related, locus 5 (NKX2–5, +631); activator protein-1 (AP-1), and cartilage paired-class homeoprotein 1 (CART-1, +648). To see whether these putative TF-binding sites are responsible for +286/+690 enhancer activity, we deleted each of the TF-binding sites from the +286/+690 enhancer reporter vector (Fig. 7B). It is of note that some of the TF-binding sites are overlapped with the other (e.g. MYOD site at +537/+548 is within the PAX-4 site at +531/+559). Thus, deletion of POU3F2 (+302/+317), OCT-1 (+318/+330), PAX-4 (+531/+559), and NKX2–5 (+631/+640) also delete FOXD3 (+305/+316), FOXD3 (+320/+329), MYOD (+537/+548), and AP-1 (+635/+640) sites, respectively. As shown in Fig. 7B, deletion of POU3F2 (+302 and +404)-, FOXJ2 (+469)-, HNF-3β (+520)-, and EVI-1 (+590)-binding sites partially suppressed the luciferase induction during myoblast differentiation, whereas the deletion of other sites did not alter luciferase activity. Becausethe deletions in Fig. 7B only partially blocked luciferase induction, we generated luciferase reporter vectors with double deletions of these putative TF-binding sites (Fig. 7C). Introduction of POU3F2 (+404), FOXJ2 (+469), HNF-3β (+520), and EVI-1 (+590) deletions into POU3F2 (+302) site-deleted vector did not further reduce the luciferase activity. Similarly, double deletions in any other combinations of putative TF-binding sites did not show additive effects on luciferase activity.
FIGURE 7.
Analysis of putative transcription factor-binding sites in AT2R intron 2. A, AT2R +241/+720 DNA sequence is shown. Green characters indicate part of the exon 2 sequence, and +286/+690 sequence is shown in blue. Putative transcription factor-binding sites predicted by TRANSFAC are shown in red as follows: POU domain, class 3, transcription factor 2 (POU3F2, +302 and +404); FOXD3 (+305 and +320); octamer transcription factor-1 (OCT-1, +318); v-MYB (+375); hepatocyte nuclear factor 4α (HNF-4α, +429), FOXJ2 (+469); FOXL1 (+470); FOXI1 (+471); FOXF1 (+471); GATA (+472); hepatocyte nuclear factor 3β (HNF-3β, +520); PAX-4 (+531); MYOD (+537); ecotropic virus integration site 1 protein homolog (EVI-1, +478 and +590); NK2 transcription factor related, locus 5 (NKX2–5, +631); activator protein-1 (AP-1) and cartilage paired-class homeoprotein 1 (CART-1, +648). Five transcription factor binding sites overlaps at the FOXJ2 site (asterisk): FOXl1 (+470/+486); FOXI1 (+471/+483); FOXF1 (+471/+483); GATA (+472/+482); EVI-1 (+478/+492). B, predicted transcription factor binding sites shown in A were deleted from the AT2R (+286/+690)-minP-luc vector, and luciferase activity was analyzed in C2C12 myoblasts/myotubes. The vector map indicates the location of the deletion. In each vector, the nucleotides indicated by red lines in A were deleted. It is of note that the deletion of POU3F2 (+302/+317), OCT-1 (+318/+330), PAX-4 (+531/+559), and NKX2–5 (+631/+640) also delete FOXD3 (+305/+316), FOXD3 (+320/+329), MYOD (+537/+548), and AP-1 (+635/+640) sites, respectively. AT2R (+286/+690)-minP-luc and pGL4.23[luc2/minP] were used as positive and negative controls, respectively. Cells were induced to differentiate 2 days after plasmid transfection and harvested before (day 0) and 2 days after differentiation. Luciferase activity was measured using Renilla luciferase as an internal control. C, AT2R (+286/+690)-minP-luc vectors with the indicated double deletions at the predicted transcription factor-binding sites were generated, and luciferase activity was analyzed in C2C12 myoblasts/myotubes as in B. Mean ± S.E., n = 4; *, p < 0.05, and **, p < 0.01, compared with d0 control; §, p < 0.05 compared with AT2R (+286/+690)-minP-luc d2.
CHF Suppresses AT2R Intron 2 Transcriptional Enhancer Activity
We aimed to identify mechanisms whereby AT2R transcription is activated during myoblast differentiation and suppressed in the CHF model. To test whether the identified AT2R +286/+690 sequence acts as an enhancer in vivo, we electroporated AT2R (+286/+690)-minP-luc vector into the gastrocnemius muscles of C57BL/6 mice. We previously reported that an electric pulse causes damage to skeletal muscle and induces skeletal muscle regeneration (5, 38) without CTX injection. In this experimental condition, there was a robust increase of luciferase activity in AT2R (+286/+690)-minP-luc vector-electroporated muscles but no increase in control (no electric pulse applied or control pGL4.23[luc2/minP] vector electroporated) muscles (Fig. 8A). The luciferase activity reached the maximal level on day 5, which is the same time point at which the AT2R expression is induced after injury (Fig. 1D) (38). To analyze whether the activity of the +286/+690 enhancer element is altered in the setting of CHF, we electroporated AT2R (+286/+690)-minP-luc vector into gastrocnemius muscles of C57BL/6 mice after 4 weeks of LAD ligation surgery (Fig. 8B). Importantly, the luciferase induction of AT2R (+286/+690)-minP-luc vector was significantly suppressed in the muscles of CHF mice. These data strongly suggest that induction of AT2R expression after muscle injury is regulated by the +286/+690 enhancer, and the enhancer activity is suppressed in the setting of CHF. However, it is also possible that the reduction of luciferase activity is due to increased cell death caused by LAD ligation. Cell death is also caused by electroporation in this experimental setting, and it peaked on the 3rd and 5th days after electroporation (Fig. 8D). We did not observe significant difference in cell death between sham and LAD groups with or without electroporation (Fig. 8E). Also, the Renilla luciferase activity under the control of herpes simplex virus-thymidine kinase promoter from pGL4.74[Rluc/TK] vector, which was used as an internal control, was not different between sham and LAD ligation groups (Fig. 8F). These data suggest that the reduction in +286/+690 enhancer activity is suppressed in the CHF condition independently from muscle cell death, potentially leading to reduced induction of AT2R in this chronic disease state.
FIGURE 8.
AT2R intron 2 enhancer activity during muscle regeneration in CHF. A, AT2R intron 2 (+286/+690) enhancer activity was measured during skeletal muscle regeneration in vivo. AT2R (+286/+690)-minP-luc (A) and control pGL4.23[luc2/minP] (C) vectors were electroporated (EP) into C57BL/6 mouse gastrocnemius muscles and harvested on d3, d5, and d7. Muscles injected with the vectors without electric pulse were used as non-injury control. Luciferase activity was measured using Renilla luciferase as an internal control. B, experimental design of luciferase activity measurement in CHF mice. LAD ligation surgery was performed in C57BL/6 mice, and AT2R (+286/+690)-minP-luc and control pGL4.23[luc2/minP] vectors were electroporated into gastrocnemius muscles 4 weeks after the surgery. Muscles were harvested 5 days after the electroporation, and luciferase activity was measured. C, luciferase activity in gastrocnemius muscles harvested from B. Sham surgery was used as a control, and luciferase activity was measured using Renilla luciferase as an internal control. D, electroporation-mediated cell death was measured by Cell Death Detection ELISA after 3, 5, and 7 days of electroporation. Measurement was normalized to protein amount. E, cell death was measured after LAD ligation surgery and electroporation in the gastrocnemius muscles harvested in B. Non-electroporation muscles were used as control. F, Renilla luciferase activity was shown in muscles from C and normalized to protein amount. Mean ± S.E., n = 6; *, p < 0.05; **, p < 0.01; ns, not significant; ND, not detected.
Discussion
Ang II levels have been shown to increase with aging and in patients with chronic diseases, and accumulating evidence suggests that increased Ang II has a causal role in the development of cachexia and sarcopenia development. We have shown that AT1R signaling inhibits SC proliferation and suppresses skeletal muscle regeneration, potentially contributing to muscle wasting. In contrast, we found that AT2R signaling potentiates SC differentiation and increases skeletal muscle regeneration. These data suggest that the balance between AT1R and AT2R signaling in SCs plays a critical role in regulating SC function and that this balance may be altered in chronic diseases, leading to lowered muscle regenerative capacity. Indeed, we now report that skeletal muscle regenerative capacity is decreased in a mouse myocardial infarction model of CHF (LAD ligation, Fig. 1). Importantly, the decline in muscle regeneration was associated with a blunted induction of AT2R expression during muscle regeneration (Fig. 1E), suggesting that suppression of AT2R expression plays a critical role in decreasing skeletal muscle regeneration in the CHF condition. Because the AT2R positively regulates SC differentiation, activation of AT2R in SCs has the potential to mitigate the decrease in muscle regeneration present in the elderly and in patients with chronic diseases. However, for the development of effective therapeutic interventions, it is critical to understand the precise molecular mechanisms whereby AT2R is regulated during SC differentiation.
There are very few reports describing AT2R promoter analysis; therefore, the AT2R transcriptional machinery is poorly understood. Ichiki and Inagami (45) determined that the −1497/+56 region of mouse AT2R accounted for 70% of the basal promoter activity in a mouse fibroblast cell line R3T3. In contrast, the −1497/−874 segment suppressed AT2R promoter activity in a rat pheochromocytoma-derived cell line PC12W (46). Xue et al. (42) found eight glucocorticoid-response elements within the −1853/+13 region of the rat AT2R promoter, and deletion of each glucocorticoid-response element increased luciferase reporter activity in a cardiac myoblast cell line H9c2. However, these previous studies analyzed only the AT2R basal promoter activity, which is likely to be different from the activity in differentiation processes, during which AT2R expression is robustly increased in many tissues/cells (47).
In this study, we aimed to identify a novel AT2R transcriptional machinery in myoblasts. Analysis of AT2R promoter activity during myoblast differentiation in vitro revealed that the AT2R 5′-sequence distant from TSS (−3000/−1000) contains repressive elements for AT2R induction during SC differentiation. Existence of a repressor element is consistent with the data from Ichiki and Inagami (46) on PC12W cells but is inconsistent with the data on R3T3 cells by the same authors (45). It is possible that different transcriptional mechanisms exist depending on the cellular context. Our data indicate that 70 bp upstream of AT2R TSS contains a core promoter region, including a TATA box that is necessary for initiation of AT2R transcription (Fig. 3B). Importantly, however, we found that AT2R intron 2, but not the 5′-upstream sequence of TSS, plays a critical role in inducing AT2R transcription during myoblast differentiation (Figs. 4 and 5).
Recent advances in ChIP-seq-based studies have revealed a highly complex picture of transcriptional regulatory networks. The classical model in which TFs bind to the promoter sequence adjacent to TSS was observed in fact only in limited cases. Studies have shown that critical tissue-specific TFs bind at large distances from the gene, either in intergenic regions or within introns. For instance, genome-wide binding profile analysis of the essential hematopoietic factors GATA1, TAL1, LDB1, and RUNX1 showed that these factors occupy distant elements (intronic/intergenic) in up to 90% of the cases (48). Our data indicate that the +286/+690 segment in intron 2 acts as a critical intronic transcriptional enhancer (Fig. 5). Further truncation of this segment showed that multiple DNA elements within the +286/+690 sequence coordinately regulate AT2R transcription during myoblast differentiation, and no one segment within this region is sufficient to activate AT2R expression (Fig. 6). The +286/+690 segment is predicted to have 19 TF-binding sites (Fig. 7A), and deletion of TF-binding sites for POU3F2 (both +302 and +404), FoxJ2, FoxA2, and Evi-1 significantly, but not completely, blocked the luciferase induction (Fig. 7B).
In vivo TF binding studies using chromatin immunoprecipitation (ChIP) revealed that a TF binds to only a small subset of motifs that exist in the genome, and the binding sites differ depending on cellular contexts. For example, TGF-β signaling, mediated through Smad2/3, directs different responses in different cell types. Cell type-specific TFs direct Smad3 to different locations of the genome to induce cell type-specific responses to TGF-β (e.g. Oct4 in embryonic stem cells, Myod1 in myotubes, and PU.1 in pro-B cells) (49, 50). These data indicate that a TF motif alone is not sufficient to direct TFs, but additional factors are required. Our data indicate that independent segments within the +286/+690 region are required for luciferase induction (Figs. 6 and 7), suggesting multiple TFs cooperatively regulate AT2R transcription. Among the above identified putative TF-binding sites (POU3F2, FoxJ2, FoxA2, and Evi-1), deletion of POU3F2/FoxD3 site at +302 showed the strongest reduction in luciferase activity. Although both POU3F2 (51) and FoxD3 (52) have been suggested to be involved in myogenesis, we found that there was no detectable expression level of these TFs during myoblast differentiation in vitro (quantitative RT-PCR, data not shown), suggesting that this site is bound and regulated by a TF not predicted by the database search.
Despite the recent development of TF-binding motif databases and algorithms to predict TF-binding sites within the genome (53–55), accurate in silico prediction of TFs that bind to the promoter/enhancer region remains a challenge. In addition to searching the TRANSFAC database using the MATCH program (Fig. 7), we ran multiple TF-binding site prediction programs for the +286/+690 enhancer (supplemental Fig. 1 and supplemental Table 2 to 5) as follows: AliBaba2.1 (alignments for analysis of binding sites) predicts TF-binding sites using the TRANSFAC database (56); the JASPAR database holds collections of position frequency matrix nucleotide profiles based on published experiments from diverse sources (57); and ConSite combines TF-binding site prediction with cross-species comparison (phylogenic footprinting), which allows identification of TF-binding sites conserved between different species (58). Our supplemental Fig. S2 shows the predicted TF-binding sites within the +286/+690 region, and the predicted sites and TFs for each site are vastly different between prediction programs. The only TF sites commonly predicted in multiple predictions are FoxD3 (+305/+317 in MATCH and +301/+312 in ConSite), GATA (+472/+482 in MATCH and +473/+482 in AliBaba), and FoxL1 (+470/+485 in MATCH and +474/+480 in JASPAR). ConSite search results showed four candidate TF-binding sites (supplemental Fig. S2D), and interestingly, the c-Fos-binding site at +287/+294 is conserved among the five species analyzed (supplemental Figs. S1C and S2D).
We also sought to identify potential binding factors to the +286/+690 enhancer and analyzed available ChIP-seq data on mouse ENCODE database: MyoD (wgEncodeEM002127 to 002130), myogenin (wgEncodeEM002131 to 002133 and 002136), acetyl-histone H3 (wgEncodeEM002148 and 002149), trimethyl histone H3 (Lys-4, wgEncodeEM002141 and 002142), and trimethyl histone H3 (Lys-27, wgEncodeEM002138 and 002143) in C2C12 myoblast/myocyte during differentiation. However, none of these data indicated a significant binding of these factors within +286/+690 region (data not shown).
In silico analysis above indicate that the +286/+690 enhancer element would not be able to be identified by analyzing available dataset or TF-binding site prediction. Further analyses, such as EMSA and ChIP-seq, are required to verify whether predicted TFs regulate the +286/+690 enhancer activity or identify TFs that are not predicted by database search. Nonetheless, our data strongly suggest that multiple TFs coordinately regulate AT2R induction (and myoblast differentiation) through binding to the +286/+690 enhancer.
We hypothesized that the AT2R transcriptional machinery is suppressed in chronic diseases such as CHF and therefore that the enhancer activity of the +286/+690 sequence is repressed in the condition of CHF, leading to blunted AT2R expression and lowered myoblast differentiation and skeletal muscle regeneration. By hindlimb muscle electroporation, we found that the +286/+690 element acts as an AT2R transcriptional enhancer in vivo (Fig. 7A). Importantly, this luciferase activity was significantly suppressed in the gastrocnemius muscles of CHF mice (Fig. 8, B and C). These data strongly suggest that the +286/+690 enhancer activity is inhibited in CHF, potentially leading to suppression of AT2R expression and skeletal muscle regeneration. The important question is whether AT2R suppression occurs in other chronic disease states, leading to lowered muscle regenerative capacity. Our hypothesis is that AT2R induction is suppressed in different chronic diseases via inhibition of the +286/+690 enhancer, and further experiments in other disease models will be required to determine whether these regulatory mechanisms are relevant to other diseases.
Another important future study is to understand the transcriptional regulatory mechanisms of AT1R in SCs. In contrast to the robust increase in AT2R during SC differentiation, we have shown that AT1R expression is decreased in parallel to AT2R increase (37). Our hypothesis is that AT1R and AT2R coordinately regulate SC proliferation and differentiation processes (38), and it would be important to understand the regulatory machinery of AT1R in SCs to fully understand the mechanisms of RAS-mediated SC function and skeletal muscle regeneration.
Although further analysis is required to identify TFs responsible for AT2R transcription during myoblast differentiation, our data are the first to identify intron 2 as an AT2R transcriptional enhancer. Because the AT2R is highly expressed in many embryonic tissues and thought to be involved in cellular differentiation processes such as neuronal cell differentiation and nerve regeneration (47), it will be of interest to determine whether the AT2R intronic enhancer also regulates AT2R expression and controls differentiation/regeneration of other cells/tissues. In summary, our data indicate that the intron 2 enhancer plays a critical role in regulating AT2R expression in chronic disease conditions, suggesting that targeting intron 2 could lead to the development of a novel intervention to increase AT2R expression in SCs and potentiate skeletal muscle regenerative capacity.
Experimental Procedures
Animals
Animal protocols were approved by the University of Missouri Animal Care and Use Committee (ACUC). 8–10-Week-old male C57BL/6 mice (Charles River) were used in this study. CTX (10 μm) was injected into eight different locations of gastrocnemius muscles (total 40 μl). LAD artery ligation was performed as described previously (59). Sham-operated animals underwent the same procedure without ligation of the coronary artery. All mice underwent echocardiography and were euthanized after 6 weeks.
Quantitative RT-PCR
Quantitative RT-PCR was performed as described previously (5), and PCR primers used in this study were obtained commercially (SABiosciences, PPM04482A for myogenin, PPM41610F for eMyHC, PPM05162A for AT1aR, PPM04811A for AT2R, and PPM03559F for Hprt).
Vector Construction
AT2R genomic DNA sequence was cloned from C57BL/6 mouse tail DNA using PfuUltra II Fusion HS DNA polymerase (Agilent Technologies) following the manufacturer's instruction. The primer combinations to construct luciferase vectors are shown in supplemental Table 1, and DNA fragments were subcloned into the pGL4.10[luc2] vector (Promega) at KpnI and XhoI sites. To avoid artificial promoter/enhancer activities derived from the vector, pGL4.10[luc2] multiple cloning site and linker sequences (between XhoI site and start ATG) were removed by PCR-based site-directed deletion (see below) using the following primers: 5′-GTT GCT GCA GTT CAA TAT GGA AGA TGC CAA AAA CAT-3′ and 5′-TTT TGG CAT CTT CCA TAT TGA ACT GCA GCA ACT CCA-3′. Construction of all the plasmids was confirmed by DNA sequencing. Site-directed deletion/mutation of plasmid vector was performed by PCR amplification with mutated primers followed by DpnI digestion and nick ligation in E. coli (60). AT2R enhancer elements were subcloned into pGL4.23[luc2/minP] vector (Promega). The primer combinations used to construct each vector are shown in supplemental Table 1.
Cell Culture
SCs were collected as described previously (61, 62). Briefly, hindlimb muscles (gastrocnemius, soleus, tibialis anterior, and extensor digitorum longus) were incubated with 0.2% (w/v) collagenase type II in DMEM and 10% horse serum for 2 h at 37 °C. Digested muscles were dissociated by triturating with a Pasteur pipette, washed, and further digested with 1 unit/ml dispase for 1 h at 37 °C. Cells were filtered, washed, and incubated in proliferation medium (DMEM supplemented with 20% FBS (Mediatech), 10% horse serum, 2% chicken embryo extract (US Biological), 1 mm sodium pyruvate, 2 mm l-glutamine, and penicillin/streptomycin) for 40 min to allow the fibroblasts to attach to the plate, and the unattached cells were plated onto Matrigel-coated plates in the proliferation medium at a density of 1 × 104 cells/cm2. C2C12 myoblasts (ATCC) were cultured in C2C12 growth medium (DMEM supplemented with 20% FBS, 1 mm sodium pyruvate, and 2 mm l-glutamine). After satellite cells and C2C12 cells reached confluence, cells were switched to the differentiation medium (DMEM supplemented with 2% horse serum, 1 mm sodium pyruvate, and 2 mm l-glutamine) to induce differentiation.
Luciferase Reporter Assay in Cultured Cells
In vitro luciferase reporter assays were performed in 24-well cell culture plates. Cells were plated at 1.67 × 105 cells/well, and plasmids (Fluc 500 ng and Rluc 25 ng per well) were transfected using Lipofectamine 2000 (Thermo Fisher Scientific) following the manufacturer's instructions. 2 days after transfection, cells were induced for differentiation by replacing medium with the differentiation medium. Cells were harvested before and 2 days after differentiation, and luciferase activity was measured by Dual-Luciferase® reporter assay system (Promega) following the manufacturer's instructions.
Electroporation in Vivo and Luciferase Assay
Mice were anesthetized with a mixture of 75 mg/kg ketamine, 15 mg/kg xylazine hydrochloride administered by i.p. injection. Mice hindlimbs were shaved, and plasmids (1 μg/μl in saline, 20 μg of AT2R (+286/+690)-minP-Fluc, and 1 μg of pGL4.74[hRluc/TK]) were injected into five different sites in each gastrocnemius muscle with a 22-gauge needle syringe (Hamilton model 705). Transcutaneous pulses were applied by two stainless steel plate electrodes (Caliper Electrode model 384, BTX) with a distance between the two plates of 0.5 cm, and conductive gel was used to ensure the electrical contact. Electric pulses with a standard square wave were delivered by an electroporator (ECM830 Electro Square Porator, BTX). Eight pulses of 50 V/cm were administered to the muscle with a delivery rate of 1 pulse/s with each pulse 20 ms in duration. Gastrocnemius muscles were collected at indicated time points. Skeletal muscle lysates were prepared, and luciferase activity was measured by Dual-Luciferase® reporter assay system (Promega) following the manufacturer's instruction.
Measurement of Cell Death
Cell death was measured in gastrocnemius muscles after electroporation using Cell Death Detection ELISA PLUS kit (Roche Applied Science) according to the manufacturer's instruction. Gastrocnemius muscles were homogenized in lysis buffer, and necrotic and apoptotic cell death was measured in this condition.
Prediction of Transcription Factor-binding Sites
Putative TF-binding sites were searched against TRANSFAC database (44) using Matrix Search for Transcription Factor Binding Sites (MATCH) program with the setting to minimize the sum of false-positive/false-negative error rates. TF-binding site search were also performed using programs Alibaba2.1 (56), JASPAR (57), and ConSite (58) with the settings indicated in supplemental Tables S2–S5.
Statistical Analysis
All data represent mean ± S.E., and results were analyzed using Student's t test when data from two experimental groups were compared or analysis of variance followed by Bonferroni's multiple comparison test when data from three or more groups were studied.
Author Contributions
T. Y. designed the experimental strategy, conducted the experiments, analyzed the results, interpreted the data, and wrote the manuscript. P. D. designed the experimental strategy, interpreted the data, and wrote the manuscript.
Supplementary Material
This work was supported by National Institutes of Health Grants R01-HL070241 and R01-HL080682 from NHLBI and P20-GM103629, P30-GM103337, and U54-GM104940 from NIGMS, American Heart Association Grant 15SDG25240022, and University of Missouri Department of Medicine Research Council. The authors declare that they have no conflicts of interest with the contents of this article. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.

This article contains supplemental Tables S1–S5 and Figs. S1 and S2.
- RAS
- renin-angiotensin system
- CHF
- congestive heart failure
- CKD
- chronic kidney disease
- Ang II
- angiotensin II
- AT1R
- angiotensin II type 1 receptor
- AT2R
- Ang II type 2 receptor
- SC
- satellite cell
- TSS
- transcription start site
- CTX
- cardiotoxin
- LAD
- left anterior descending artery
- TF
- transcription factor.
References
- 1. Penafuerte C. A., Gagnon B., Sirois J., Murphy J., MacDonald N., and Tremblay M. L. (2016) Identification of neutrophil-derived proteases and angiotensin II as biomarkers of cancer cachexia. Br. J. Cancer 114, 680–687 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Anker S. D., Negassa A., Coats A. J., Afzal R., Poole-Wilson P. A., Cohn J. N., and Yusuf S. (2003) Prognostic importance of weight loss in chronic heart failure and the effect of treatment with angiotensin-converting-enzyme inhibitors: an observational study. Lancet 361, 1077–1083 [DOI] [PubMed] [Google Scholar]
- 3. Brink M., Wellen J., and Delafontaine P. (1996) Angiotensin II causes weight loss and decreases circulating insulin-like growth factor I in rats through a pressor-independent mechanism. J. Clin. Invest. 97, 2509–2516 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Song Y.-H., Li Y., Du J., Mitch W. E., Rosenthal N., and Delafontaine P. (2005) Muscle-specific expression of IGF-1 blocks angiotensin II-induced skeletal muscle wasting. J. Clin. Invest. 115, 451–458 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Yoshida T., Semprun-Prieto L., Sukhanov S., and Delafontaine P. (2010) IGF-1 prevents ANG II-induced skeletal muscle atrophy via Akt- and Foxo-dependent inhibition of the ubiquitin ligase atrogin-1 expression. Am. J. Physiol. Heart Circ Physiol. 298, H1565–H1570 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Rezk B. M., Yoshida T., Semprun-Prieto L., Higashi Y., Sukhanov S., and Delafontaine P. (2012) Angiotensin II infusion induces marked diaphragmatic skeletal muscle atrophy. PLoS ONE 7, e30276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Tabony A. M., Yoshida T., Galvez S., Higashi Y., Sukhanov S., Chandrasekar B., Mitch W. E., and Delafontaine P. (2011) Angiotensin II upregulates protein phosphatase 2Cα and inhibits AMP-activated protein kinase signaling and energy balance leading to skeletal muscle wasting. Hypertension 58, 643–649 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Tabony A. M., Yoshida T., Sukhanov S., and Delafontaine P. (2014) Protein phosphatase 2C-α knockdown reduces angiotensin II-mediated skeletal muscle wasting via restoration of mitochondrial recycling and function. Skeletal Muscle 4, 20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Yoshida T., and Delafontaine P. (2015) Mechanisms of cachexia in chronic disease states. Am. J. Med. Sci. 350, 250–256 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Mitchell P. O., and Pavlath G. K. (2004) Skeletal muscle atrophy leads to loss and dysfunction of muscle precursor cells. Am. J. Physiol. Cell Physiol. 287, C1753–C1762 [DOI] [PubMed] [Google Scholar]
- 11. Mozdziak P. E., Pulvermacher P. M., and Schultz E. (2001) Muscle regeneration during hindlimb unloading results in a reduction in muscle size after reloading. J. Appl. Physiol. 91, 183–190 [DOI] [PubMed] [Google Scholar]
- 12. Wanek L. J., and Snow M. H. (2000) Activity-induced fiber regeneration in rat soleus muscle. Anat. Rec. 258, 176–185 [DOI] [PubMed] [Google Scholar]
- 13. Guo B.-S., Cheung K.-K., Yeung S. S., Zhang B.-T., and Yeung E. W. (2012) Electrical stimulation influences satellite cell proliferation and apoptosis in unloading-induced muscle atrophy in mice. PLoS ONE 7, e30348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Matsuba Y., Goto K., Morioka S., Naito T., Akema T., Hashimoto N., Sugiura T., Ohira Y., Beppu M., and Yoshioka T. (2009) Gravitational unloading inhibits the regenerative potential of atrophied soleus muscle in mice. Acta Physiol. 196, 329–339 [DOI] [PubMed] [Google Scholar]
- 15. Jejurikar S. S., and Kuzon W. M. (2003) Satellite cell depletion in degenerative skeletal muscle. Apoptosis 8, 573–578 [DOI] [PubMed] [Google Scholar]
- 16. Langen R. C., Schols A. M., Kelders M. C., van der Velden J. L., Wouters E. F., and Janssen-Heininger Y. M. (2006) Muscle wasting and impaired muscle regeneration in a murine model of chronic pulmonary inflammation. Am. J. Respir. Cell Mol. Biol. 35, 689–696 [DOI] [PubMed] [Google Scholar]
- 17. Thériault M.-E., Paré M.-È., Lemire B. B., Maltais F., and Debigaré R. (2014) Regenerative defect in vastus lateralis muscle of patients with chronic obstructive pulmonary disease. Respir. Res. 15, 35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Nguyen T., Shrager J., Kaiser L., Mei L., Daood M., Watchko J., Rubinstein N., and Levine S. (2000) Developmental myosin heavy chains in the adult human diaphragm: coexpression patterns and effect of COPD. J. Appl. Physiol. 88, 1446–1456 [DOI] [PubMed] [Google Scholar]
- 19. Vogiatzis I., Simoes D. C., Stratakos G., Kourepini E., Terzis G., Manta P., Athanasopoulos D., Roussos C., Wagner P. D., and Zakynthinos S. (2010) Effect of pulmonary rehabilitation on muscle remodelling in cachectic patients with COPD. Eur. Respir. J. 36, 301–310 [DOI] [PubMed] [Google Scholar]
- 20. Fermoselle C., Rabinovich R., Ausín P., Puig-Vilanova E., Coronell C., Sanchez F., Roca J., Gea J., and Barreiro E. (2012) Does oxidative stress modulate limb muscle atrophy in severe COPD patients? Eur. Respir. J. 40, 851–862 [DOI] [PubMed] [Google Scholar]
- 21. Crul T., Spruit M. A., Gayan-Ramirez G., Quarck R., Gosselink R., Troosters T., Pitta F., and Decramer M. (2007) Markers of inflammation and disuse in vastus lateralis of chronic obstructive pulmonary disease patients. Eur. J. Clin. Invest. 37, 897–904 [DOI] [PubMed] [Google Scholar]
- 22. Cheung W. W., and Mak R. H. (2012) Melanocortin antagonism ameliorates muscle wasting and inflammation in chronic kidney disease. Am. J. Physiol. Renal Physiol. 303, F1315–F1324 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Zhang L., Wang X. H., Wang H., Du J., and Mitch W. E. (2010) Satellite cell dysfunction and impaired IGF-1 signaling cause CKD-induced muscle atrophy. J. Am. Soc. Nephrol. 21, 419–427 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Corrick K. L., Stec M. J., Merritt E. K., Windham S. T., Thomas S. J., Cross J. M., and Bamman M. M. (2015) Serum from human burn victims impairs myogenesis and protein synthesis in primary myoblasts. Front. Physiol. 6, 184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Wu X., Walters T. J., and Rathbone C. R. (2013) Skeletal muscle satellite cell activation following cutaneous burn in rats. Burns 39, 736–744 [DOI] [PubMed] [Google Scholar]
- 26. Krause M. P., Moradi J., Nissar A. A., Riddell M. C., and Hawke T. J. (2011) Inhibition of plasminogen activator inhibitor-1 restores skeletal muscle regeneration in untreated type 1 diabetic mice. Diabetes 60, 1964–1972 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Wieteska-Skrzeczynska W., Grzelkowska-Kowalczyk K., Jank M., and Maciejewski H. (2009) Transcriptional dysregulation of skeletal muscle protein metabolism in streptozotocin-diabetic mice. J. Physiol. Pharmacol. 60, 29–36 [PubMed] [Google Scholar]
- 28. Aragno M., Mastrocola R., Catalano M. G., Brignardello E., Danni O., and Boccuzzi G. (2004) Oxidative stress impairs skeletal muscle repair in diabetic rats. Diabetes 53, 1082–1088 [DOI] [PubMed] [Google Scholar]
- 29. Su Z., Robinson A., Hu L., Klein J. D., Hassounah F., Li M., Wang H., Cai H., and Wang X. H. (2015) Acupuncture plus low-frequency electrical stimulation (Acu-LFES) attenuates diabetic myopathy by enhancing muscle regeneration. PLoS ONE 10, e0134511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Gulati A. K., and Swamy M. S. (1991) Regeneration of skeletal muscle in streptozotocin-induced diabetic rats. Anat. Rec. 229, 298–304 [DOI] [PubMed] [Google Scholar]
- 31. Brannon M. A., Dodson M. V., Wheeler B. A., Mathison B. D., and Mathison B. A. (1989) Satellite cells derived from streptozotocin-diabetic rats display altered fusion parameters in vitro. Metab. Clin. Exp. 38, 348–352 [DOI] [PubMed] [Google Scholar]
- 32. Ramamoorthy S., Donohue M., and Buck M. (2009) Decreased Jun-D and myogenin expression in muscle wasting of human cachexia. Am. J. Physiol. Endocrinol Metab. 297, E392–E401 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. He W. A., Berardi E., Cardillo V. M., Acharyya S., Aulino P., Thomas-Ahner J., Wang J., Bloomston M., Muscarella P., Nau P., Shah N., Butchbach M. E., Ladner K., Adamo S., Rudnicki M. A., et al. (2013) NF-κB-mediated Pax7 dysregulation in the muscle microenvironment promotes cancer cachexia. J. Clin. Invest. 123, 4821–4835 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Mastrocola R., Reffo P., Penna F., Tomasinelli C. E., Boccuzzi G., Baccino F. M., Aragno M., and Costelli P. (2008) Muscle wasting in diabetic and in tumor-bearing rats: role of oxidative stress. Free Radic. Biol. Med. 44, 584–593 [DOI] [PubMed] [Google Scholar]
- 35. Penna F., Costamagna D., Fanzani A., Bonelli G., Baccino F. M., and Costelli P. (2010) Muscle wasting and impaired myogenesis in tumor bearing mice are prevented by ERK inhibition. PLoS ONE 5, e13604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Mehl K. A., Davis J. M., Berger F. G., and Carson J. A. (2005) Myofiber degeneration/regeneration is induced in the cachectic ApcMin/+ mouse. J. Appl. Physiol. 99, 2379–2387 [DOI] [PubMed] [Google Scholar]
- 37. Yoshida T., Galvez S., Tiwari S., Rezk B. M., Semprun-Prieto L., Higashi Y., Sukhanov S., Yablonka-Reuveni Z., and Delafontaine P. (2013) Angiotensin II inhibits satellite cell proliferation and prevents skeletal muscle regeneration. J. Biol. Chem. 288, 23823–23832 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Yoshida T., Huq T. S., and Delafontaine P. (2014) Angiotensin type 2 receptor signaling in satellite cells potentiates skeletal muscle regeneration. J. Biol. Chem. 289, 26239–26248 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Yoshida T., Semprun-Prieto L., Wainford R. D., Sukhanov S., Kapusta D. R., and Delafontaine P. (2012) Angiotensin II reduces food intake by altering orexigenic neuropeptide expression in the mouse hypothalamus. Endocrinology 153, 1411–1420 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Semprun-Prieto L. C., Sukhanov S., Yoshida T., Rezk B. M., Gonzalez-Villalobos R. A., Vaughn C., Michael Tabony A., and Delafontaine P. (2011) Angiotensin II induced catabolic effect and muscle atrophy are redox dependent. Biochem. Biophys. Res. Commun. 409, 217–221 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Yoshida T., Tabony A. M., Galvez S., Mitch W. E., Higashi Y., Sukhanov S., and Delafontaine P. (2013) Molecular mechanisms and signaling pathways of angiotensin II-induced muscle wasting: potential therapeutic targets for cardiac cachexia. Int. J. Biochem. Cell Biol. 45, 2322–2332 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Xue Q., Dasgupta C., Chen M., and Zhang L. (2011) Foetal hypoxia increases cardiac AT(2)R expression and subsequent vulnerability to adult ischaemic injury. Cardiovasc. Res. 89, 300–308 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Blanchette M., and Tompa M. (2002) Discovery of regulatory elements by a computational method for phylogenetic footprinting. Genome Res. 12, 739–748 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Matys V., Kel-Margoulis O. V., Fricke E., Liebich I., Land S., Barre-Dirrie A., Reuter I., Chekmenev D., Krull M., Hornischer K., Voss N., Stegmaier P., Lewicki-Potapov B., Saxel H., Kel A. E., and Wingender E. (2006) TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34, D108–D110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Ichiki T., and Inagami T. (1995) Expression, genomic organization, and transcription of the mouse angiotensin II type 2 receptor gene. Circ. Res. 76, 693–700 [DOI] [PubMed] [Google Scholar]
- 46. Ichiki T., and Inagami T. (1995) Transcriptional regulation of the mouse angiotensin II type 2 receptor gene. Hypertension 25, 720–725 [DOI] [PubMed] [Google Scholar]
- 47. de Gasparo M., Catt K. J., Inagami T., Wright J. W., and Unger T. (2000) International union of pharmacology. XXIII. The angiotensin II receptors. Pharmacol. Rev. 52, 415–472 [PubMed] [Google Scholar]
- 48. Stadhouders R., van den Heuvel A., Kolovos P., Jorna R., Leslie K., Grosveld F., and Soler E. (2012) Transcription regulation by distal enhancers: who's in the loop? Transcription 3, 181–186 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Yáñez-Cuna J. O., Dinh H. Q., Kvon E. Z., Shlyueva D., and Stark A. (2012) Uncovering cis-regulatory sequence requirements for context-specific transcription factor binding. Genome Res. 22, 2018–2030 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Mullen A. C., Orlando D. A., Newman J. J., Lovén J., Kumar R. M., Bilodeau S., Reddy J., Guenther M. G., DeKoter R. P., and Young R. A. (2011) Master transcription factors determine cell-type-specific responses to TGF-β signaling. Cell 147, 565–576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Liu W., He R., and Xue Z. (2001) Developmental expression of the POU transcription factor qBrn-2 during somitic myogenesis in quail. Int. J. Dev. Biol. 45, 415–420 [PubMed] [Google Scholar]
- 52. Plank J. L., Suflita M. T., Galindo C. L., and Labosky P. A. (2014) Transcriptional targets of Foxd3 in murine ES cells. Stem Cell Res. 12, 233–240 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Wasserman W. W., and Sandelin A. (2004) Applied bioinformatics for the identification of regulatory elements. Nat. Rev. Genet. 5, 276–287 [DOI] [PubMed] [Google Scholar]
- 54. Yáñez-Cuna J. O., Kvon E. Z., and Stark A. (2013) Deciphering the transcriptional cis-regulatory code. Trends Genet. 29, 11–22 [DOI] [PubMed] [Google Scholar]
- 55. Deplancke B., Alpern D., and Gardeux V. (2016) The genetics of transcription factor DNA binding variation. Cell 166, 538–554 [DOI] [PubMed] [Google Scholar]
- 56. Grabe N. (2002) AliBaba2: context specific identification of transcription factor binding sites. In Silico Biol. 2, S1–S15 [PubMed] [Google Scholar]
- 57. Mathelier A., Zhao X., Zhang A. W., Parcy F., Worsley-Hunt R., Arenillas D. J., Buchman S., Chen C.-Y., Chou A., Ienasescu H., Lim J., Shyr C., Tan G., Zhou M., Lenhard B., et al. (2014) JASPAR 2014: an extensively expanded and updated open-access database of transcription factor binding profiles. Nucleic Acids Res. 42, D142–D147 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Sandelin A., Wasserman W. W., and Lenhard B. (2004) ConSite: web-based prediction of regulatory elements using cross-species comparison. Nucleic Acids Res. 32, W249–W252 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Michael L. H., Entman M. L., Hartley C. J., Youker K. A., Zhu J., Hall S. R., Hawkins H. K., Berens K., and Ballantyne C. M. (1995) Myocardial ischemia and reperfusion: a murine model. Am. J. Physiol. 269, H2147–H2154 [DOI] [PubMed] [Google Scholar]
- 60. Liu H., and Naismith J. H. (2008) An efficient one-step site-directed deletion, insertion, single and multiple-site plasmid mutagenesis protocol. BMC Biotechnol. 8, 91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Conboy I. M., Conboy M. J., Smythe G. M., and Rando T. A. (2003) Notch-mediated restoration of regenerative potential to aged muscle. Science 302, 1575–1577 [DOI] [PubMed] [Google Scholar]
- 62. Conboy I. M., and Rando T. A. (2002) The regulation of Notch signaling controls satellite cell activation and cell fate determination in postnatal myogenesis. Dev. Cell 3, 397–409 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.