Abstract
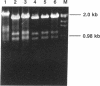
Nuclear extracts from several tissue culture cell lines (human, primate, and murine) contain an endonuclease that specifically cleaves sequences at the herpes simplex virus 1 (HSV-1) segment inversion site. Mapping studies identified the preferential site of cleavage as a set of tandemly repeated dodecamers, the DR2 repeats. Endonuclease levels vary according to the proliferative state of the cell; little or no activity is detectable in extracts from quiescent cells, whereas high levels are expressed in dividing cells. Also, infection of density-arrested BSC-1 cells with HSV-1 induces a substantial increase (at least 35-fold) in endonucleolytic activity, which is first detectable at about 1 hr after infection at 32 degrees C. The elevated levels of enzyme activity then persist throughout the viral life cycle. In addition to the HSV-1 DR2 repeats, certain other G+C-rich sequences with an asymmetric distribution of purines and pyrimidines on the DNA strands and with appropriate sequences and lengths are substrates for the nuclease. These data indicate that target site recognition by the enzyme is conformation specific rather than sequence specific.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Beasty A. M., Behe M. J. An oligopurine sequence bias occurs in eukaryotic viruses. Nucleic Acids Res. 1988 Feb 25;16(4):1517–1528. doi: 10.1093/nar/16.4.1517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Behe M. J. The DNA sequence of the human beta-globin region is strongly biased in favor of long strings of contiguous purine or pyrimidine residues. Biochemistry. 1987 Dec 1;26(24):7870–7875. doi: 10.1021/bi00398a050. [DOI] [PubMed] [Google Scholar]
- Chou J., Roizman B. Characterization of DNA sequence-common and sequence-specific proteins binding to cis-acting sites for cleavage of the terminal a sequence of the herpes simplex virus 1 genome. J Virol. 1989 Mar;63(3):1059–1068. doi: 10.1128/jvi.63.3.1059-1068.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chou J., Roizman B. Isomerization of herpes simplex virus 1 genome: identification of the cis-acting and recombination sites within the domain of the a sequence. Cell. 1985 Jul;41(3):803–811. doi: 10.1016/s0092-8674(85)80061-1. [DOI] [PubMed] [Google Scholar]
- Côté J., Renaud J., Ruiz-Carrillo A. Recognition of (dG)n.(dC)n sequences by endonuclease G. Characterization of the calf thymus nuclease. J Biol Chem. 1989 Feb 25;264(6):3301–3310. [PubMed] [Google Scholar]
- Dalziel R. G., Marsden H. S. Identification of two herpes simplex virus type 1-induced proteins (21K and 22K) which interact specifically with the a sequence of herpes simplex virus DNA. J Gen Virol. 1984 Sep;65(Pt 9):1467–1475. doi: 10.1099/0022-1317-65-9-1467. [DOI] [PubMed] [Google Scholar]
- Davison A. J., Wilkie N. M. Nucleotide sequences of the joint between the L and S segments of herpes simplex virus types 1 and 2. J Gen Virol. 1981 Aug;55(Pt 2):315–331. doi: 10.1099/0022-1317-55-2-315. [DOI] [PubMed] [Google Scholar]
- Gottlieb J., Muzyczka N. In vitro excision of adeno-associated virus DNA from recombinant plasmids: isolation of an enzyme fraction from HeLa cells that cleaves DNA at poly(G) sequences. Mol Cell Biol. 1988 Jun;8(6):2513–2522. doi: 10.1128/mcb.8.6.2513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gottlieb J., Muzyczka N. Purification and characterization of HeLa endonuclease R. A G-specific mammalian endonuclease. J Biol Chem. 1990 Jul 5;265(19):10836–10841. [PubMed] [Google Scholar]
- Gottlieb J., Muzyczka N. Substrate specificity of HeLa endonuclease R. A G-specific mammalian endonuclease. J Biol Chem. 1990 Jul 5;265(19):10842–10850. [PubMed] [Google Scholar]
- Hanvey J. C., Klysik J., Wells R. D. Influence of DNA sequence on the formation of non-B right-handed helices in oligopurine.oligopyrimidine inserts in plasmids. J Biol Chem. 1988 May 25;263(15):7386–7396. [PubMed] [Google Scholar]
- Hanvey J. C., Shimizu M., Wells R. D. Intramolecular DNA triplexes in supercoiled plasmids. II. Effect of base composition and noncentral interruptions on formation and stability. J Biol Chem. 1989 Apr 5;264(10):5950–5956. [PubMed] [Google Scholar]
- Hanvey J. C., Shimizu M., Wells R. D. Intramolecular DNA triplexes in supercoiled plasmids. Proc Natl Acad Sci U S A. 1988 Sep;85(17):6292–6296. doi: 10.1073/pnas.85.17.6292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayward G. S., Jacob R. J., Wadsworth S. C., Roizman B. Anatomy of herpes simplex virus DNA: evidence for four populations of molecules that differ in the relative orientations of their long and short components. Proc Natl Acad Sci U S A. 1975 Nov;72(11):4243–4247. doi: 10.1073/pnas.72.11.4243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hogan M., LeGrange J., Austin B. Dependence of DNA helix flexibility on base composition. Nature. 1983 Aug 25;304(5928):752–754. doi: 10.1038/304752a0. [DOI] [PubMed] [Google Scholar]
- Jaworski A., Blaho J. A., Larson J. E., Shimizu M., Wells R. D. Tetracycline promoter mutations decrease non-B DNA structural transitions, negative linking differences and deletions in recombinant plasmids in Escherichia coli. J Mol Biol. 1989 Jun 5;207(3):513–526. doi: 10.1016/0022-2836(89)90461-0. [DOI] [PubMed] [Google Scholar]
- Jeffreys A. J., Wilson V., Thein S. L. Hypervariable 'minisatellite' regions in human DNA. Nature. 1985 Mar 7;314(6006):67–73. doi: 10.1038/314067a0. [DOI] [PubMed] [Google Scholar]
- Jenkins F. J., Roizman B. Herpes simplex virus 1 recombinants with noninverting genomes frozen in different isomeric arrangements are capable of independent replication. J Virol. 1986 Aug;59(2):494–499. doi: 10.1128/jvi.59.2.494-499.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klein R. D., Wells R. D. Effects of neighboring DNA homopolymers on the biochemical and physical properties of the Escherichia coli lactose promoter. I. Cloning and characterization studies. J Biol Chem. 1982 Nov 10;257(21):12954–12961. [PubMed] [Google Scholar]
- Knopf C. W. Recombinational hotspots within the herpes simplex virus DNA polymerase gene? Nucleic Acids Res. 1987 Sep 25;15(18):7647–7648. doi: 10.1093/nar/15.18.7647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohwi-Shigematsu T., Kohwi Y. Poly(dG)-poly(dC) sequences, under torsional stress, induce an altered DNA conformation upon neighboring DNA sequences. Cell. 1985 Nov;43(1):199–206. doi: 10.1016/0092-8674(85)90024-8. [DOI] [PubMed] [Google Scholar]
- Kohwi Y., Kohwi-Shigematsu T. Magnesium ion-dependent triple-helix structure formed by homopurine-homopyrimidine sequences in supercoiled plasmid DNA. Proc Natl Acad Sci U S A. 1988 Jun;85(11):3781–3785. doi: 10.1073/pnas.85.11.3781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu L. F., Wang J. C. Supercoiling of the DNA template during transcription. Proc Natl Acad Sci U S A. 1987 Oct;84(20):7024–7027. doi: 10.1073/pnas.84.20.7024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lyamichev V. I., Mirkin S. M., Frank-Kamenetskii M. D. A pH-dependent structural transition in the homopurine-homopyrimidine tract in superhelical DNA. J Biomol Struct Dyn. 1985 Oct;3(2):327–338. doi: 10.1080/07391102.1985.10508420. [DOI] [PubMed] [Google Scholar]
- McCall M., Brown T., Kennard O. The crystal structure of d(G-G-G-G-C-C-C-C). A model for poly(dG).poly(dC). J Mol Biol. 1985 Jun 5;183(3):385–396. doi: 10.1016/0022-2836(85)90009-9. [DOI] [PubMed] [Google Scholar]
- McLean M. J., Blaho J. A., Kilpatrick M. W., Wells R. D. Consecutive A X T pairs can adopt a left-handed DNA structure. Proc Natl Acad Sci U S A. 1986 Aug;83(16):5884–5888. doi: 10.1073/pnas.83.16.5884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mirkin S. M., Lyamichev V. I., Drushlyak K. N., Dobrynin V. N., Filippov S. A., Frank-Kamenetskii M. D. DNA H form requires a homopurine-homopyrimidine mirror repeat. Nature. 1987 Dec 3;330(6147):495–497. doi: 10.1038/330495a0. [DOI] [PubMed] [Google Scholar]
- Mocarski E. S., Deiss L. P., Frenkel N. Nucleotide sequence and structural features of a novel US-a junction present in a defective herpes simplex virus genome. J Virol. 1985 Jul;55(1):140–146. doi: 10.1128/jvi.55.1.140-146.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mocarski E. S., Post L. E., Roizman B. Molecular engineering of the herpes simplex virus genome: insertion of a second L-S junction into the genome causes additional genome inversions. Cell. 1980 Nov;22(1 Pt 1):243–255. doi: 10.1016/0092-8674(80)90172-5. [DOI] [PubMed] [Google Scholar]
- Mocarski E. S., Roizman B. Herpesvirus-dependent amplification and inversion of cell-associated viral thymidine kinase gene flanked by viral a sequences and linked to an origin of viral DNA replication. Proc Natl Acad Sci U S A. 1982 Sep;79(18):5626–5630. doi: 10.1073/pnas.79.18.5626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mocarski E. S., Roizman B. Site-specific inversion sequence of the herpes simplex virus genome: domain and structural features. Proc Natl Acad Sci U S A. 1981 Nov;78(11):7047–7051. doi: 10.1073/pnas.78.11.7047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mocarski E. S., Roizman B. Structure and role of the herpes simplex virus DNA termini in inversion, circularization and generation of virion DNA. Cell. 1982 Nov;31(1):89–97. doi: 10.1016/0092-8674(82)90408-1. [DOI] [PubMed] [Google Scholar]
- Poffenberger K. L., Roizman B. A noninverting genome of a viable herpes simplex virus 1: presence of head-to-tail linkages in packaged genomes and requirements for circularization after infection. J Virol. 1985 Feb;53(2):587–595. doi: 10.1128/jvi.53.2.587-595.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poffenberger K. L., Tabares E., Roizman B. Characterization of a viable, noninverting herpes simplex virus 1 genome derived by insertion and deletion of sequences at the junction of components L and S. Proc Natl Acad Sci U S A. 1983 May;80(9):2690–2694. doi: 10.1073/pnas.80.9.2690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rahmouni A. R., Wells R. D. Stabilization of Z DNA in vivo by localized supercoiling. Science. 1989 Oct 20;246(4928):358–363. doi: 10.1126/science.2678475. [DOI] [PubMed] [Google Scholar]
- Ruiz-Carrillo A., Renaud J. Endonuclease G: a (dG)n X (dC)n-specific DNase from higher eukaryotes. EMBO J. 1987 Feb;6(2):401–407. doi: 10.1002/j.1460-2075.1987.tb04769.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smiley J. R., Fong B. S., Leung W. C. Construction of a double-jointed herpes simplex viral DNA molecule: inverted repeats are required for segment inversion, and direct repeats promote deletions. Virology. 1981 Aug;113(1):345–362. doi: 10.1016/0042-6822(81)90161-6. [DOI] [PubMed] [Google Scholar]
- Umene K. Conversion of a fraction of the unique sequence to part of the inverted repeats in the S component of the herpes simplex virus type 1 genome. J Gen Virol. 1986 Jun;67(Pt 6):1035–1048. doi: 10.1099/0022-1317-67-6-1035. [DOI] [PubMed] [Google Scholar]
- Umene K., Enquist L. W. Isolation of novel herpes simplex virus type 1 derivatives with tandem duplications of DNA sequences encoding immediate-early mRNA-5 and an origin of replication. J Virol. 1985 Feb;53(2):607–615. doi: 10.1128/jvi.53.2.607-615.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weber P. C., Challberg M. D., Nelson N. J., Levine M., Glorioso J. C. Inversion events in the HSV-1 genome are directly mediated by the viral DNA replication machinery and lack sequence specificity. Cell. 1988 Jul 29;54(3):369–381. doi: 10.1016/0092-8674(88)90200-0. [DOI] [PubMed] [Google Scholar]
- Wells R. D., Collier D. A., Hanvey J. C., Shimizu M., Wohlrab F. The chemistry and biology of unusual DNA structures adopted by oligopurine.oligopyrimidine sequences. FASEB J. 1988 Nov;2(14):2939–2949. [PubMed] [Google Scholar]
- Wohlrab F., McLean M. J., Wells R. D. The segment inversion site of herpes simplex virus type 1 adopts a novel DNA structure. J Biol Chem. 1987 May 5;262(13):6407–6416. [PubMed] [Google Scholar]
- Wohlrab F., Wells R. D. Slight changes in conditions influence the family of non-B-DNA conformations of the herpes simplex virus type 1 DR2 repeats. J Biol Chem. 1989 May 15;264(14):8207–8213. [PubMed] [Google Scholar]