Abstract
Neurodegenerative diseases (NDs) often involve the formation of abnormal and toxic protein aggregates, which are thought to be the primary factor in ND occurrence and progression. Aged neurons exhibit marked increases in aggregated protein levels, which can lead to increased cell death in specific brain regions. As no specific drugs/therapies for treating the symptoms or/and progression of NDs are available, obtaining a complete understanding of the mechanism underlying the formation of protein aggregates is needed for designing a novel and efficient removal strategy. Intracellular proteolysis generally involves either the lysosomal or ubiquitin-proteasome system. In this review, we focus on the structure and assembly of the proteasome, proteasome-mediated protein degradation, and the multiple dynamic regulatory mechanisms governing proteasome activity. We also discuss the plausibility of the correlation between changes in proteasome activity and the occurrence of NDs. [BMB Reports 2016; 49(9): 459-473]
Keywords: Assembly, Gate opening, Neurodegenerative diseases, Post-translational modification, Proteasome, Regulators
INTRODUCTION
The 26S proteasome is a large multi-protein complex and plays a central role in ubiquitin-dependent proteolysis in eukaryotic cell. The 20S core proteasome (CP; ∼700 kDa) and the 19S regulatory proteasome (RP; ∼900 kDa) interact to form the 26S proteasome. Association of the 19S RP with either one or both ends of the 20S CP results in the formation of the 26S (RP1-CP) and 30S (RP2-CP), respectively. The term “26S proteasome” is commonly used to refer to both isoforms, as the RP2-CP, RP1-CP, and CP, as well as various assembly intermediates, all coexist in the cell (1). In rat cortex, RP2-CP comprises approximately 14% of the proteasome complement, whereas RP1-CP comprises approximately 43% (2). Instead of the 19S RP, the 26S proteasome can includes various ATP-independent regulatory molecules, such as PA28α/β, PA28γ, PA200, PI31, PC530, or the signalosome COP9 (1). These alternative regulatory molecules also associate with the ends of the 20S CP, potentially resulting in the formation of asymmetric 26S proteasome isoforms. Although they lack ATPase activity, the alternative regulatory molecules mediate ubiquitin-independent protein degradation. The subunits of the 26S proteasome are listed in Table 1.
Table 1. Subunits of the 26S proteasome.
| Subunit | S. cerevisiae | Mammals | Activity/Function | Size** (human) | ||
|---|---|---|---|---|---|---|
| 20S CP | α ring | α1 | Scl1/Prc2/Prs2/C7 | PSMA6/Pros27/Iota | 27.4 | |
| α2 | Pre8/Prs4/Y7 | PSMA2/C3/Lmpc3 | 25.8 | |||
| α3 | Pre9/Prs5/Y13 | PSMA4/C9 | Gate opening | 29.4 | ||
| α4 | Pre6 | PSMA7/C7/XAPC7 | 27.9 | |||
| α5 | Pup2/Doa5 | PSMA5/Zeta | 26.5 | |||
| α6 | Pre5 | PSMA1/C2/Pros30 | 29.6/30.1# | |||
| α7 | Pre10/Prc1/Prs1/C1 | PSMA3/C8 | 28.3 | |||
| β ring | β1 | Pre3 | PSMB6/Y/delta/LMPY/LMP19 | PGPH* activity | ||
| β1i | PSMB9/Lmp2/RING12/MC7 | Chymotrypsin-like activity | ||||
| β2 | Pup1 | PSMB7/Z/MC14/Lmp9 | Trypsin-like activity | 25.3 | ||
| β2i | PSMB10/MECL-1/Lmp10 | Trypsin-like activity | ||||
| β3 | Pup3 | PSMB3/C10 | 22.9 | |||
| β4 | Pre1/C11 | PSMB2/C7 | 22.8 | |||
| β5 | Pre2/Doa3/Prg1 | PSMB5/X/epsilon/MB1/Lmp17 | Chymotrypsin-like activity | 22.5 | ||
| β5i | PSMB8/Lmp7/RING10/Y2/C13 | Chymotrypsin-like activity | ||||
| β6 | Pre7/Prs3/Pts1/C5 | PSMB1/C5 | 23.5 | |||
| β7 | Pre4 | PSMB4/N3/beta/LMP3 | 24.4 | |||
| 19S RP | Base | Rpt1/S7 | Rpt1/Cim5/Yta3 | PSMC2/Mss1 | AAA ATPase | 48.6 |
| Rpt2/S4 | Rpt2/Yhs4/Yta5 | PSMC1 | AAA ATPase / Gate opening | 49.2 | ||
| Rpt3/S6b | Rpt3/Ynt1/Yta2 | PSMC4/Mip224/Tbp7 | AAA ATPase | 47.3 | ||
| Rpt4/S10b | Rpt4/Crl13/Pcs1/Sug2 | PSMC6/Sug2/p42 | AAA ATPase | 44.1 | ||
| Rpt5/S6a | Rpt5/Yta1 | PSMC3/Tbp1 | AAA ATPase / Ub receptor / Gate opening | 49.1 | ||
| Rpt6/S8 | Rpt6/Cim3/Crl3/Sug1/TbpY/Tby1 | PSMC5/p45/Sug1/Trip1 | AAA ATPase | 45.7 | ||
| Rpn1/S2 | Rpn1/Hrd2/Nas1/Rpd1 | PSMD2/p97/Trap2 | Ub receptor / Binding of substrate / Gate opening | 100.2 | ||
| Rpn2/S1 | Rpn2/Sen3 | PSMD1/p112 | Ub receptor / Binding of substrate / Gate opening | 105.9 | ||
| Rpn10/S5a | Rpn10/Sun1/Mcb1 | PSMD4/S5a/Mcb1 | Ub receptor / Interface of lid & base | 40.7 | ||
| S5b | - | PSMD5/KIAA0072 | ||||
| Rpn13 | Rpn13/Daq1 | ADRM1 | Ub receptor | 42.1 | ||
| Lid | Rpn3/S3 | Rpn3/Sun2 | PSMD3/p58 | 61.0 | ||
| Rpn4 | Rpn4/Son1/Ufd5 | - | ||||
| Rpn5/p55 | Rpn5/Nas5 | PSMD12/p55 | 52.9 | |||
| Rpn6/S9 | Rpn6/Nas4 | PSMD11/p44.5 | Binding of substrate (possible) | 47.4 | ||
| Rpn7/S10 | Rpn7 | PSMD6/p42a | 45.5 | |||
| Rpn8/S12 | Rpn8 | PSMD7/p40/Mov34 | 37.1 | |||
| Rpn9/Nas7 | Rpn9/Nas7 | PSMD13/p40.5 | 42.9 | |||
| Rpn11/S13 | Rpn11/Mpr1 | PSMD14/Pad1/Poh1 | Deubiquitinase | 34.6 | ||
| Rpn12/S14 | Rpn12/Nin1 | PSMD8/p31 | 30.0 | |||
| Rpn15/Sem1 | Rpn15/Sem1/Dsh1 | SHFM1/DSS1/SHFDG1 | 24.6 | |||
| p27 | Nas2 | PSMD9/p27 | ||||
| p28 | Nas6 | PSMD10/p28/Gankyrin | ||||
*PGPH: peptidylglutamyl-peptide hydrolyzing (also called caspase-like activity). **Unit of size: kDa. #Long and short isoform of the α6 subunit
20S CP
The eukaryotic 20S CP has a cylindrical structure consisting of four hetero-heptameric rings, two outer and two central. The two outer rings are composed of seven α-subunits (α1-α7) and the two central rings are composed of seven β-subunits (β1-β7) (3). Mammalian cells also contain immunoproteasomes that can be activated by treatment with interferon-γ (IFNγ), which stimulates the synthesis of three additional proteasome subunits: β1i, β2i, and β5i. The immune-20S CP (i-CP) is then formed by the substitution of β1, β2, and β5 with β1i, β2i, and β5i, respectively (4).
Catalytic activity of the 20S CP: Of the seven β-subunits, only three (β1, β2, and β5) exhibit threonine protease hydrolytic activity (3). These subunits cleave a different peptide sequence; for example, β1 hydrolyzes on the C-terminal side after acidic residues, β2 hydrolyzes on the C-terminal side after tryptic residues, and β5 hydrolyzes on the C-terminal side after hydrophobic residues. Subunits β1, β2, and β5 are referred to as exhibiting peptidylglutamyl-peptide hydrolyzing activity (PGPH; also called caspase-like activity), tryptic activity, and chymotryptic activity, respectively. The protease activity of subunits β1, β2, and β5 is self-activated via a two-step mechanism characterized by initial reduction of the length of the propeptide sequence and subsequent autocatalytic removal of the residual propeptide.
RNase activity: The 20S CP (particularly the α5 subunit) exhibits RNase activity dependent upon divalent cations such as Ca2+ and Mg2+, whereas the proteolytic activity does not require divalent cations (5). Interestingly, the α5-RNase activity is highly correlated with the extent of α6- and α7-subunit phosphorylation. Moreover, the α5-RNase activity increases markedly during erythroid differentiation and programmed cell death (5). Mittenberg et al. recently reported that most α-subunits (except α3) exhibit in vitro RNase activity against p53 mRNA (6). They also reported that the RNase activity of the α6- and α7-subunit is upregulated upon hemin-induced differentiation of proerythroleukemia cells, in parallel with changes in their phosphorylation and ubiquitination status (6).
19S RP
The 19S RP consists of base and lid complexes and functions as an ATP-dependent proteasome activator. The base complex comprises six AAA+-ATPase subunits (regulatory particle triple A-ATPase 1 [Rpt1]-Rpt6) and three non-ATPases (regulatory particle non-ATPase 1 [Rpn1], Rpn2, and Rpn10) (1). In mammals, the base complex also contains hRpn13 and S5b, whereas that of Saccharomyces cerevisiae contains Rpn13 (7). The six Rpt subunits form a heterohexameric ring in which the Rpn1 and Rpn2 are stacked (1). The lid complex comprises eight non-ATPases (Rpn3, 5-9, 11, and 12) (1). The lid is loosely (i.e., temporarily) associated with other subunits, such as p27, p28, and Rpn15, and in S. cerevisiae, the lid is also associated with Rpn4. Interaction between the base and the lid complex is stabilized by Rpn10 (1).
Functions of the 19S RP
The 19S RP interacts with the 20S CP to trigger opening of the gate in the 20S CP through conformational rearrangement of the N-terminal region of the α-subunit (8). Base complex subunits Rpt2, Rpt5, Rpn1, and Rpn2 participate in the gate-opening process (9). Rpn10 then captures polyubiquitinated substrates via the C-terminal hydrophobic cluster. Whereas isolated Rpn10 protein interacts with three forms of polyubiquitin (Lys6, Lys11, or Lys 48) in equal ratios, the intact 26S proteasome interacts only with Lys48-linked polyubiquitin chains (10). Rpn1, Rpn2, Rpn13, and Rpt5 are also involved in capturing of polyubiquitinated substrates. Rpn1 and Rpn2 contain a leucine-rich repeat domain that is involved in non-specific protein-protein interactions and also plays an important role in substrate binding (11). The previously reported finding that ubiquitinated nuclear factor κB/p100 interacts with Rpn6 (12) suggests that Rpn6 is another proteasome substrate acceptor. Multiple Rpt subunits also interact with substrates. Before translocation into the 20S CP, substrates are fully unfolded by the 19S RP, utilizing energy derived from the hydrolysis of ATP (13). The lid subsequently releases the polyubiquitin chain from the substrates to facilitate their degradation, in which Rpn11 plays a role (14).
Alternative ATP-independent regulatory particles
PA28 (also called 11S regulator): The PA28 family consist of PA28α/β and PA28γ. PA28α/β has a heteroheptameric ringshaped structure and is composed of either an α4β3 or α3β4 complex (15). PA28α/β associates with the two ends of the 20S CP or the free end of asymmetric 19S-20S CPs to form 19S-20S-PA28. PA28 accelerates the degradation of small peptides, but it has no effect on the degradation of folded or ubiquitinated proteins (15). The PA28 α- and β-subunits promote proteasome activity and contain a highly conserved linker sequence between helices 2 and 3, which is referred as “the activation loop”, a structure that is important for the activity-stimulating effect of PA28 (15). Expression of PA28α/β is induced by IFNγ. Under conditions that induce an intensified immune response, PA28α/β associates tightly with the i-CP. In contrast to PA28α/β, IFNγ does not induce expression of the PA28γ subunit. In addition, the PA28γ subunit forms homoheptamers and associates with the 20S CP, activating primarily the tryptic activity (16).
PA200 (also called Blm10 or Blm3): PA200 is a monomeric protein with an asymmetric turban-like shape. PA200 attaches to one or both ends of the 20S CP to induce the hydrolysis of small peptides (but not folded proteins) in vitro, but the induction only of peptide hydrolysis is not seen with the yeast homologue, Blm10 (17). PA200 connects the 26S proteasome to the proteins that recognize regions of DNA molecules damaged by ionizing radiation (18). Once recruited to the region of damaged DNA, the 26S proteasome degrades the chromatin-associated proteins there to expose the damaged section of DNA for repair enzymes (18).
PI31: PI31 is a proline-rich protein that competes with other regulatory proteins for binding to the 20S CP. In vitro, PI31 forms an unstable complex with the 20S CP (1) and inhibits the chymotryptic and PGPH activity while stimulating the tryptic activity of the 20S CP. PI31 also plays a role in immunoproteasome biogenesis, instead of the usual ATP/ ubiquitin-dependent protein degradation (15).
PC530: PC530 is a multi-protein complex originally identified in starfish oocytes. PC530 associates with the 20S CP to regulate the removal of defective sperm in the epididymis and paternal mitochondria in fertilized eggs (19).
The COP9 Signalosome (CSN): The CSN consists of eight subunits (CSN1-8) that are highly homologous to the subunits of the 19S RP lid. The CSN replaces and assumes the function of the 19S RP lid in the 26S proteasome (1). The CSN also exhibits de-ubiquitination activity (20).
REGULATION AND UNDERLYING MOLECULAR MECHANISMS OF PROTEASOME ASSEMBLY AND PROTEOLYSIS
Regulation of 26S proteasome assembly and novel modulators
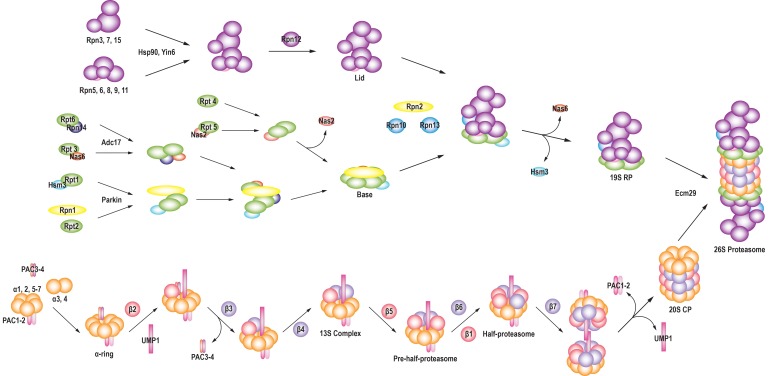
Considering the biological importance of the 26S proteasome, its assembly would be expected to proceed in a precise and tightly controlled manner. The process of 26S proteasome assembly is complex, however, as 66 proteins must be assembled into a functional complex in an exact order. In addition, some proteasome subunits must be processed during assembly. Previous studies revealed that several chaperone proteins actively mediate assembly of the highly complex proteasome in an extremely efficient and accurate process. A schematic illustration of the pathway for assembly of the entire 26S proteasome based on currently available data is shown in Fig. 1.
Fig. 1. Schematic diagram illustrating assembly of the 26S proteasome.
Assembly of the 20S CP
Archaeal CP: Each α- and β-subunit of the archaeal CP is encoded by a single-copy gene. Binding between subunits results in the formation of a homoheptameric ring (21). The α-subunit alone can form single or coupled heptameric rings via its N-terminal α-helix. In contrast, the β-subunit cannot self-assemble into multimeric structures and remains in the monomer state. Proteolytic cleavage of the β-subunit leads to the formation of inactive rings (21). Assembly of the archaeal CP is probably initiated by formation of the α-ring, which then attaches to the β-subunit to form a half-proteasome. Two half-proteasomes assemble into the pre-CP, which is finally converted into the mature CP after cleavage of the N-terminal propeptide of the β-subunit (21). Panfair et al. (22) proposed three plausible 20S CP assembly pathways: (a) α-subunits self-assemble in cis to form a single-ring (SR) that serves as a template for β-subunit entry; (b) interaction between multiple α-subunits and a β-subunit, independent of the SR, lead to the formation of the half-proteasome; and (c) α-subunits interacts with each other in trans, resulting in the formation of a double-ring structurally analogous to a half-proteasome.
Bacterial CP: The bacterial CP is composed of both α- and β-subunits (23). In contrast to the archaeal CP, bacterial α-subunits do not self-assemble to form a ring structure. In bacteria, one α- and one β-subunit form a heterodimer that assembles into a half-proteasome. Analyses of the crystal structure of a bacterial CP revealed that the β-subunit propeptide acts as an intramolecular chaperone, mediating CP assembly by inserting into the interface between two adjacent α-subunits and acting to hold them together (24).
Eukaryotic CP: Studying the assembly of the eukaryotic CP is difficult due to the heterogeneity and short half-lives of the ≥33 associated subunits. The involvement of several chaperone proteins adds to the complexity of the eukaryotic CP assembly process. Eukaryotic CP assembly usually involves four steps, as described below: Step 1: α-ring formation. Formation of the α-ring is promoted by several chaperone proteins, including proteasome assembling chaperone 1 (PAC1; also known as proteasome biogenesis-associated-1 [Pba1] in yeast)-PAC2 and PAC3-PAC4 heterodimers. The PAC1-PAC2 complex initiates formation of an α-ring intermediate composed of all α-subunits except α3 and α4 (21). The PAC1-PAC2 heterodimer remains attached to the α-ring until the assembly is completed to prevent aberrant dimerization of immature intermediate α-ring (25). Formation of the mature α-ring involves another chaperone, the PAC3-PAC4 heterodimer, which interacts with α5 at a site opposite the site at which PAC1-PAC2 interacts. It is important to promote the appropriate incorporation of α3 and α4 (25). Step 2: β-ring formation. Formation of the β-ring begins with the attachment of β2 to the α-ring, followed by the binding of β3 and β4. The β2 propeptide recruits β3, and then β2 wraps around β3 to initiate formation of the β-ring. Upon β3 incorporation, the PAC3-PAC4 heterodimer dissociates from the α-ring (25). The resulting intermediate, known as the 13S complex, consists of an α-ring, β2-β4, and the PAC1-PAC2 heterodimer. After PAC3-PAC4 dissociation, the β5 propeptide (which is required for β6 recruitment) interacts with underpinning maturation of proteasome 1 (UMP1), which then recruits β6 and β1 to the 13S complex. This process results in the formation of a pre-half-proteasome, also called the 15S precursor complex. Finally, β7 associates with the pre-half-proteasome and triggers the formation of the half-proteasome (also called the 15S complex) (25). Sahara et al. (26) recently identified two chaperones, transmembrane recognition complex 40 and Bag6. Knockdown of these chaperones leads to accumulation of CP assembly intermediates and inefficient β-subunit assembly on the α-ring. Step 3: dimerization of the half-proteasome. Dimerization of the half-proteasome occurs after incorporation of β7. The extended C-terminal tail of β7 is inserted between β1 and β2 on the opposing β-ring, forming a strong dimer composed of two half-proteasomes and stabilizing the active β1 conformation (27). The β5 propeptide and N-terminal extension of β6 also promote half-proteasome dimerization. In contrast, the β6 propeptide and UMP1 inhibit half-proteasome dimerization. UMP1 in particular functions to inhibit dimerization until all β-subunits are recruited onto the α-ring (21). Step 4: maturation. The dimerized half-proteasome is initially loosely packed, with a wider gate than the mature CP (28), but it is then re-arranged by sequential removal of β-propeptides. Finally, the dimerized proteasome degrades UMP1, the PAC1-PAC2 complex is released and recycled for a new round of proteasome assembly, and the gate is tightened (28, 29).
CP isoforms: Eukaryotic cells contain two additional CP isoforms: i-CP and thymo-CP (t-CP). The β-subunits of the i-CP and t-CP differ from those of the 20S CP. The i-CP contains β1i, β2i and β5i instead of β1, β2, and β5, respectively (4). The t-CP contains thymus-specific β5t instead of β5 or β5i (30). β1i is incorporated into the i-CP earlier than β1 and therefore can facilitate the incorporation of β2i. Thus, β1i and β2i are incorporated before the other β-subunits, with subsequent recruitment of β3, β4, β5i (or β5t), and β6. As with the 20S CP, the i-CP and t-CP half-proteasomes dimerize after the incorporated of β7 (29). The same chaperone proteins involved in the assembly of the 20S CP are also involved in the assembly of the i-CP and t-CP.
Assembly of the 19S RP
Base assembly: As with the CP, assembly of the RP base is tightly orchestrated and regulated by a number of chaperone proteins, including Nas2/p27, Nas6/gankyrin/p28, Rpn14/ PAAF1, Hsm3/S5b, parkin, Bag6, and Adc17, which bind to the C-terminus of specific Rpts to facilitate proper pairing. Two hypothetic models of the base assembly process have been proposed: CP-independent and CP-dependent assembly. CP-independent assembly. CP-independent assembly involves selective interaction of Hsm3 with Rpt1, Nas2 with Rpt5, Nas6 with Rpt3, and Rpn14 with Rpt6 (31). In addition, interaction between Rpt1 and Rpn1 is mediated by parkin (a protein linked to Parkinson disease) (32). Bag6 directly binds to Rpt5 and promotes the interaction between Rpt4 and the Nas2-Rpt5 complex (26). Adc17 interacts with the C-terminus of Rpt6 to facilitate formation of the Rpt6-Rpt3 complex (33). These interactions ultimately lead to the formation of three intermediates, Hsm3-Rpn1-Rpt1-Rpt2, Nas2-Rpt4-Rpt5, and Nas6- Rpt3-Rpt6-Rpn14, which then form a heterohexameric ring with Rpn2, Rpn10, and Rpn13 (31). CP-dependent assembly. In CP-dependent assembly, Rpt4, Rpt6, and possibly Rpt3 interact to form a complex directly on the CP. Rpn14 and Nas6 guide the complex to the appropriate position at which specific α-ring pockets of the CP can interact with the C-terminus of the Rpts. The Hsm3-Rpn1-Rpt1-Rpt2 complex is subsequently formed independent of the CP and recruited and positioned at the Hsm3-Rpn1-Rpt1-Rpt2 complex on the CP (34).
Lid assembly: Analyses of the intact lid using mass spectrometry (MS) have suggested two possible intermediate complexes; one comprised of Rpn5, Rpn6, Rpn8, Rpn9, and Rpn11, and another comprised of Rpn3, Rpn7, Rpn12, and Rpn15. Subunits Rpn3, Rpn7, Rpn12, and Rpn15 form a helical bundle that directs the ordered self-assembly of C-terminal helices and proteasome COP9 initiation factor 3 domains. These two intermediates are connected by the interaction between Rpn3 and Rpn5 (35). Finally, Rpn12 is incorporated into the lid complex to complete the assembly process (36). Prior to the attachment of Rpn12, this complex exists in a closed state with the N-terminal domain of Rpn6 folded inward, inhibiting binding to the base. During Rpn12 incorporation, the single α-helix of Rpn12 induces the repositioning of Rpn8 and Rpn11 and a conformational change in the N-terminus of Rpn6 that consequently promotes its binding to the base (36). Two chaperones are involved in lid assembly: Hsp90 and Yin6. Inactivation of the proteins or deletion of their gene results in lid disassembly (21). The lid complex is subsequently joined to the base via an interaction between Rpn10 and the helical N-terminal domain of Rpn9 (37). The observation that parkin overexpression markedly enhances the interaction between Rpn6 and Rpt5 suggests that parkin may help stabilize the base and lid complexes (32).
RP-CP association
Precise regulation of RP-CP association is critical for proper proteasome function because both CP maturation and gate opening require association with the RP. The RP and CP complex is assembled via interaction between the C-terminus of Rpts and α-subunit pockets (38). Tian et al. reported that the CP exhibits unexpected asymmetry, in that one side of the ring makes 1:1 contacts between Rpt2 and α4, Rpt6 and α3, and Rpt3 and α2, whereas on the opposite side, the tails of Rpt1, Rpt4 and Rpt5 cross-link with α5/α6, α7/α1, and α6/α7, respectively (38). The Rpt6-α3 interaction promotes both the Rpt3-α2 interaction through ATP-dependent binding with Nas6 and the release of Nas6 from the proteasome (39). Both Ecm29 and Hsp90 have been shown to regulate RP-CP assembly. Purified proteasomes from a bacterial strain lacking the Ecm29 gene exhibited CP-RP dissociation. Ecm29 stabilizes the 26S proteasome by enhancing the interaction between the RP and CP (21). Loss-of-function mutations in Hsp90 lead to dissociation of the 26S proteasome (21). However, the mechanism underlying the interaction between the RP and CP remains unclear. Interestingly, we recently found that the telomerase catalytic subunit hTERT exhibits a novel chaperone activity, selectively mediating the association between the RP and CP (Im and Chung, manuscript in preparation). Specifically, hTERT promotes the interactions between RP and CP subunits, thus facilitating proteasome assembly and subsequent proteasome-mediated proteolysis.
Gate opening within the 20S CP
Under resting conditions, 20S CP activity is auto-inhibited by the N-terminus of the α-subunit tail. Fully unfolded proteins are translocated by the RP into the chamber of the 20S CP through a gate formed by the CP tails (40). The first 12 amino acids of the α-subunit block the catalytic sites of the β-subunit (8). The tail of the α-subunit has a conserved ‘Tyr8-Asp9-Arg10’ (YDR) sequence. All α-subunits have Tyr8, whereas six of the seven α-subunits (except α2) have Asp9, and three of the seven α-subunits (α1, α4, and α5) have Arg10. In α3, the carboxylate group of Asp9 forms a salt bridge with the guanidinium group of Arg10 in α4. It also forms hydrogen bonds with the carboxylated group of α4-Asp9. Deletion of the 9-residue tail (GSRRYDSRT) of the α3 subunit leads to disordered formation of the other six α-subunits, resulting in opening of the 20S CP gate along the 7-fold pseudo-symmetric axis (40). Substitution of α3-Asp9 with Ala promotes normal gate opening, thus increasing peptide hydrolysis (40).
Gate opening can also occur upon binding of the CP to activator proteins, such as the 19S RP, PAN, PA26, and PA200. The ATPases in the 19S RP and PAN complexes contain a conserved C-terminal hydrophobic-tyrosine-X (HbYX) motif. This motif induces ATP-dependent gate opening and is important to the binding of PAN to the 20S CP (41). The C-terminal regions of Rpt2 and Rpt5 contain the HbYX motif and strongly stimulate gate opening, whereas Rpt3, which also contains the HbYX motif, does not appear to stimulate opening. Furthermore, Rpt1, which contains the YX sequence, induces only weak gate opening, whereas Rpt4 and Rpt6, which do not contain the HbYX motif, have no effect (41). The C-terminal domain of PAN inserts into the pockets between the α-subunits, where it interacts with the highly conserved residues Gly34 and Leu81 (41). Through these interactions, HbYX motif of PAN induces a rotation of the α-subunits of about 4° around Gly128, which is a key residue in the highly conserved α-subunit loop (Tyr126-Gly127-Gly128) (41).
The rotation of the α-subunit results in radial and lateral movement of Pro17 away from the pore and a reverse-turn interaction between four highly conserved residues (Tyr8, Asp9, Pro17, and Tyr26) that serves to stabilize the open-gate conformation (8, 41). Rosenzweig et al. also suggested that the Rpn1-Rpn2 interaction plays some role in regulating gate opening via binding of the Rpn1-Rpn2 complex with the 20S CP to facilitate the entry of small substrates (9). PA26, a PA28 homologue from Trypanosoma brucei, also induces gate opening via the same mechanism as PAN, although the C-terminal region of PA26 lacks the HbYX motif (42). Unlike PAN, PA26 function requires an activation loop (clustered between Arg141 and Gly149) that induces a reverse turn in the α-subunits that facilitates gate opening and stabilizes the open conformation (42). The C-terminal region of PA200 has a YYX motif (or less frequently, a YFX motif) that is analogous to the C-terminal HbYX motif of PAN and three ATPases. It is thought that this motif could trigger the degradation of proteasome substrates by inducing gate opening (43).
Regulation of 26S proteasome activity by post-translational modifications
Numerous reports have demonstrated that proteasomal subunits are subject to a diverse array of post-translational modifications (PTMs), including phosphorylation, N-acetylation, ubiquitination, SUMOylation, O-glycosylation, succinylation, poly-ADP ribosylation, oxidation, N-terminal processing, proteolytic cleavage by caspases, and N-myristoylation (Table 2). As most of these modifications were identified during large-scale proteomics analyses of cells or tissues, the specific roles of many of these PTMs and their targets have yet to be clearly elucidated. Nevertheless, the functional consequences or/and specific roles of PTMs of various enzymes and modulators have been examined.
Table 2. Modifications of 26S proteasome subunits.
| Subunit | N-terminal processing | Cleavage by caspase | N-myristoylation | Phosphorylation | Ubiquitination | SUMOylation | N-acetylation | O-Glycosylation | Succinylation | Poly-ADP ribosylation | Carbonylation | HNE modification | S-glutathionylation | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Cas3 | Cas7 | SUMO1 | SUMO2 | SUMO3 | SUMO4 | ||||||||||||
| α1 | O (m) | K59 (H) | O (y) | S5 (m) | O (r) | ||||||||||||
| α2 | O (H) | Y120 (m) | O (H) | O (y) | O (m) | O (r) | |||||||||||
| α3 | S74 (H) | O (y) | O (H) | O (y) | O (m) | O (y) | O (r) | ||||||||||
| α4 | Y106 | K115 (H) | O (y) | S130 (m) | O (y) | O (r) | |||||||||||
| Y153 (H) | |||||||||||||||||
| α5 | O (H) | O (y) | S198 | O (r) | C76 | ||||||||||||
| (H, m) | C11 | ||||||||||||||||
| C221 (y) | |||||||||||||||||
| α6 | O (H) | K115 | O (H) | O (y) | S110 (m) | O (r) | C66 (y) | ||||||||||
| K208 (H) | |||||||||||||||||
| α7 | S243 | O (y) | O (y) | O (H) | C42 | ||||||||||||
| S250 (M) | C76 (y) | ||||||||||||||||
| α-sub-unit | T84 | ||||||||||||||||
| T178 | |||||||||||||||||
| T202 (b) | |||||||||||||||||
| β1 | O (y) | O (y) | |||||||||||||||
| β1i | O (r) | ||||||||||||||||
| β2 | O (y) | O (m) | O (y) | O (m) | |||||||||||||
| β3 | O (m) | O (y) | O (y) | O (r) | |||||||||||||
| β4 | O (H) | O (y) | O (r) | ||||||||||||||
| β5 | O (y) | O (y) | O (m) | ||||||||||||||
| β6 | O (y) | S57 | O (y) | O (r) | |||||||||||||
| S208 (m) | |||||||||||||||||
| β7 | O (y) | O (m) | |||||||||||||||
| 20S CP | O (H) | ||||||||||||||||
| Rpt1 | O (H) | O (H) | O (H) | O (H) | |||||||||||||
| Rpt2 | D288 (m) | O (H, m, y) | K237 (H) | O (H) | |||||||||||||
| Rpt3 | T25 (m) | O (y) | O (y) | O (H) | |||||||||||||
| Rpt4 | O (y) | ||||||||||||||||
| Rpt5 | D27 (H) | O (H, m) | O (y) | O (H) | |||||||||||||
| Rpt6 | D248 (m) | S120 (r) | O (H) | O (y) | O (y) | ||||||||||||
| Rpn1 | O (y) | O (H) | O (H) | ||||||||||||||
| Rpn2 | D85 (H) | O (H) | O (y) | O (y) | O (H) | ||||||||||||
| Rpn10 | D11 D258 (H) | K71# | O (H) | O (H) | O (H) | ||||||||||||
| K84# | |||||||||||||||||
| K99# | |||||||||||||||||
| K112 | |||||||||||||||||
| K126 | |||||||||||||||||
| K135 | |||||||||||||||||
| K262 | |||||||||||||||||
| K268# | |||||||||||||||||
| K365 (H) | |||||||||||||||||
| Rpn13 | O (y) | O (y) | |||||||||||||||
| Rpn3 | O (y) | O (y) | |||||||||||||||
| Rpn5 | O (y) | ||||||||||||||||
| Rpn6 | S14 (H, m) | O (H) | O (y) | ||||||||||||||
| Rpn7 | O (y) | ||||||||||||||||
| Rpn8 | O (y) | ||||||||||||||||
| Rpn9 | O (y) | ||||||||||||||||
| Rpn11 | O (y) | ||||||||||||||||
| Rpn12 | O (y) | O (H) | O (y) | ||||||||||||||
| PA28 | O (H) | ||||||||||||||||
| PI31 | O (d) | ||||||||||||||||
*Abbreviation, H: human, M: monkey, m: mouse, r: rat, d: drosophila, y: yeast, b: bacteria. #Same site with human and yeast.
Irreversible PTMs
N-terminal processing: Five β-subunits (except β3 and β4) of the 20S CP contain N-terminal propeptides that prevent inactivation of the Thr1-containing active site required for proper proteasome assembly via acetylation (44). During assembly of the 20S CP, a conserved Gly-Thr dipeptide within the β1, β2, and β5-subunits is cleaved between the Gly and Thr residues in an autocatalytic fashion to facilitate generation of the active subunit (1). An (evolutionarily conserved) Lys residue located in close proximity to the Gly-Thr cleavage site is also necessary for correct processing of the β subunits (1). Finally, the β6 and β7-subunits are further processed by the neighboring active subunits, whereas β3 and β4 remain unprocessed (3).
Cleavage by caspase: Several caspases cleave proteasome subunits. For example, during apoptosis, Rpt5, Rpn2, and Rpn10 are cleaved by caspase-3 (45), whereas in MCF-7 cells stimulated with the 5-lipoxygenease inhibitor, nordihydroguaiaretic acid, α2, α6, and Rpt1 are cleaved by caspase-7 (46). Cleavage of the above-mentioned subunits by caspse-3 or -7 impairs proteasome activity, resulting in the accumulation of ubiquitinated proteins. However, caspase-3 cleavage of Rpt2 and Rpt6 was shown to increase proteasome activity in staurosporine- or etoposide-treated C2C12 myotubes, although the underlying mechanism has yet to be elucidated (47).
N-myristoylation: N-myristoylation is a modification in which a myristoyl lipid group is covalently attached to the N-terminal Gly residue of the target protein. MS analyses indicated that Rpt2 contains a highly conserved N-myristoylation motif of at the N-terminus (48). Although the functional consequence of N-myristoylation of Rpt2 is unclear, as myristoylation promotes protein-protein and protein-membrane interactions, the modification is thought to play a role in mediating interactions between the 26S proteasome and membranes or membrane-bound regulatory proteins (49).
Reversible PTMs
Phosphorylation: As is the case with many biological processes, proteasome activity is most commonly regulated via phosphorylation of component proteins. Alterations in proteasome subunit conformation resulting from phosphorylation control the stability, assembly, and proteolytic activity of the proteasome. For example, several subunits of the 20S CP are known to be phosphorylated. Phosphorylation of the α-subunit in the Mycobacterium tuberculosis proteasome impedes assembly of the proteasome complex and consequently enhances mycobacterial resistance to H2O2 (50). PKA phosphorylates several subunits of the mammalian 20S CP, including α1-α3, β2, β3 and β7, thus increasing the proteasomal chymotryptic and PGPH activities (51). Phosphorylation of α3 in particular affects the binding of the 19S RP to the 20S CP because the phosphorylated residue is located close to the end of cylinder-shaped 20S CP (49). Under conditions of oxidative stress or ionizing irradiation, the α4-subunit is also phosphorylated on Tyr153 by c-Abl and Arg tyrosine kinase (52), ultimately inhibiting proteasomal activity. The c-Abl kinase also phosphorylates the α4-subunit on Tyr 106, in turn promoting the degradation of the α4-subunit via BRCA1-mediated ubiquitination (52). Phosphorylation of the α7-subunit by CKII does not affect proteasomal activity but does stabilize the association between the 19S RP and 20S CP. Bose et al. reported that phosphorylation of PA28 is markedly upregulated by INFγ, stimulating the peptidase activity of the proteasome via conformational changes in PA28 (53). Conversely, dephosphorylation of PA28 leads to its dissociation from the 20S CP (53).
The ATPase activity of the 19S RP is also regulated by phosphorylation (54). Phosphorylation of the 19S RP also likely affects substrate recognition and gate opening of the 20S CP as well as the assembly of 19S RP subunits due to phosphorylation-induced conformational changes (55). Examples of specific phosphorylation of 19S RP subunits, including targeted residues and the functional consequences and underlying regulatory mechanisms of phosphorylation, have been reported by many investigators, including ourselves. For instance, we demonstrated that under a variety of stressful conditions, the ATPase activity of Rpt5 is inhibited as a result of direct phosphorylation by ASK1 (56). During cell cycle progression, the Rpt3 subunit of the 19S RP is phosphorylated on Thr25 by DYRK2, enhancing substrate translocation to the proteasome complex and subsequent degradation (57). Eventually, it increases tumorigenesis by proteasome-addicted human breast cancer cells in mice (57). Activated PKA phosphorylates Rpn6 on Ser14, whereas both PKA and CaMKII phosphorylate Rpt6 (58). Phosphorylation of Rpt6 by PKA and CaMKII initiates proteasome assembly by enhancing the direct association between Rpt6 and α2 (59, 60). In contrast, inhibition of CaMKII prevents the retrieval-induced increase in proteasome activity as well as the phosphorylation of Rpt6 on Ser120 in the amygdala (60).
Ubiquitination and SUMOylation: Several subunits of 26S proteasome have been identified as targets for conjugation with ubiquitin and small ubiquitin-like modifier (SUMO). Besche et al. reported that RING figure E3 ligase Rnf181 attaches mono-ubiquitin to Rpt1 (61). In addition, Rpn10 can be ubiquitinated by three different ubiquitin ligases: RING finger E3 ligase (MuRF1, Siah2, Parkin, SCFβTRCP1, and APC), U-box-containing E3 (CHIP), and HECT domain E3 (E6AP and Nedd4) (62). Moreover, the ubiquitin-protein ligase Rsp5 (mammalian homologue Nedd4) catalyzes the Lys63-linked polyubiquitination of Rpn10, whereas Ubp2, a deubiquitinating enzyme, removes the polyubiquitin chain (63). These two enzymes mediate the monoubiquitination of Rpn10 at four Lys residues. Lys84-mono-ubiquitination decreases the ability of Rpn10 to bind polyubiquitinated substrates, consequently inhibiting proteasome activity (63). Mono-ubiquitinated Rpn10 is further modified for Lys48-linked polyubiquitination by the HECT domain E3 enzyme, Hul5. Lys48-linked polyubiquitinated Rpn10 with a Lys84-mono-ubiquitination is then degraded by the 26S proteasome (64). Another ubiquitin receptor, Rpn13, is Lys29- and Lys48-linked polyubiquitinated by 19S RP-associated ubiquitin ligase Hul5 under conditions of proteotoxic stress and reduced proteasomal capacity (61). Ubiquitinated Rpn13 then inhibits the degradation of ubiquitinated proteins (but not peptide substrates) by blocking the binding of the 20S CP to ubiquitin conjugates (61). Furthermore, ubiquitination of the α2-subunit of the 20S CP leads to its interaction with ALAD, a proteasome-associated protein that specifically inhibits endogenous proteasome chymotryptic activity (65).
Large-scale proteomics analyses have revealed that similar to ubiquitination, many proteasome subunits undergo SUMOylation, although the functional role of SUMOylation of specific targets remains to be determined. Whereas only four subunits are modified by SUMO-1 in yeast (i.e., Rpn1, Rpn7, Rpn12, and α3), many more proteasome subunits are SUMOylated in human (66). Manza et al. identified that several components of the 19S RP and 20S CP are SUMOylated following exposure to oxidative stress. Whereas Rpn10 is modified by SUMO-1, SUMO-3 covalently attaches to the α6-subunit and Rpn10 in the cytosol and to Rpt1 in the nucleus (67). Golebiowski et al. reported that eight proteasome subunits (α3, α5, β4, Rpn1, Rpn2, Rpn6, Rpn10, and Rpn12) are conjugated to SUMO-2. Under heat shock treatment, SUMO-2 is conjugated to α5 and β4, and their levels are considerably increased (66). In addition, under conditions of serum starvation, human Rpt1 and Rpt6 are modified by SUMO-4 (66).
N-acetylation: N-Acetylation of proteasome components occurs via a reversible mechanism mediated by N-acetyltransferase (NAT) and various histone deacetylases (HDACs). For N-terminal acetylation to proceed, the N-terminal methionine must be cleaved before NAT-catalyzed replacement with an acetyl group from acetyl-CoA. There are at least three types of NATs capable of modifying proteasome subunits: NatA (or NAT1), NatB (or NAT3), and NatC (or MAK3). NatA acetylates α1-α4, α7, β3, Rpt4-Rpt6, Rpn2, Rpn3, Rpn5, Rpn6, and Rpn8 (68, 69), whereas NatB acetylates β4, Rpt3, and Rpn11, and NatC acetylates α5 and α6 (68, 69). Acetylation of α-subunits by NatA appears to contribute to gate opening. Non-N-acetylated α-subunits carry a positive charge that induces a conformational change in the 20S CP and gate opening (68). N-Acetylation of the β1-, β2-, and β5-subunits, which form the active site for 20S CP-mediated proteolysis, impedes assembly of the 20S CP (44). Liquid chromatography-tandem mass spectrometry (LC-MS/MS) analyses revealed that human and mouse cardiac proteasome subunits are also acetylated and that acetylation enhances proteasomal capacity under both healthy and diseased conditions (70). In addition, polyubiquitinated α2-subunits are also acetylated in cells treated with an HDAC inhibitor (65).
O-glycosylation: A number of nuclear and cytoplasmic proteins, including proteasome subunits, are modified with O-linked N-acetylglucosamine, which is attached to the hydroxyl groups on Ser or Thr residue of target proteins, a process known as O-glycosylation. Glycosylation of Rpt2 reduces its native ATPase activity and a consequent reduction in proteasome activity (71). Zong et al. identified six glycosylated subunits in murine cardiac 20S proteasomes: α1-α3, β2, β5, and β6 (72). Overath et al. reported that glycosylated subunits of the 20S proteasome in murine spleen and brain tissues include α1, α4-α6, and β6 (73). To date, glycosylation of only one subunit, α5, has been reported in human erythrocytes (74).
Succinylation: Lys residues in target proteins can be modified with a succinyl group. LC-MS/MS analyses have identified 13 proteasome subunits that are succinylated in yeast: α3, α4, α7, β3, β4, β6, Rpt3, Rpt6, Rpn2, Rpn3, Rpn9, Rpn12, and Rpn13 (75). The functional consequences of succinylation of any of these subunits remain unknown, however. Interestingly, Weinert et al. reported that 10 of the 13 succinylation sites in the yeast subunits are also targets of acetylation, suggesting the possibility of crosstalk between succinylation and acetylation in the proteasome complex (75).
Poly-ADP ribosylation: Poly-ADP ribosylation involves the addition of one or more ADP-ribose moieties from NAD+ to Glu, Asp, or Lys residues of a substrate, catalyzed by poly (ADP-ribose) polymerase 1 (PARP1). ADP-ribosylation of the nuclear 20S CP by PARP1 increases proteasomal chymotryptic activity and enhances the degradation of damaged histones and oxidized nuclear proteins (76). PI31 is also modified and regulated by ADP-ribosylation, which disrupts the interaction between PI31 and the α-subunits of the 20S CP, upregulating assembly of the 26S proteasome (77).
Oxidation: Proteasome subunits are sensitive to oxidation by carbonylation, 4-hydroxy-2-nonenal (HNE) modification, and S-glutathionylation. Carbonylation introduces a carbonyl group onto the native amino acid side chains of the target proteins. Rpt3 is carbonylated in response to electrophile-induced oxidative stress, resulting in impaired ATPase activity and subsequently reduced proteasomal activity (78). Divald et al. showed that carbonylation of human cardiac Rpt5 subunit increases following ischemia and reperfusion injury (79).
Lipid peroxidation by free radicals produced during oxidative stress can lead to HNE modification. The HNE moiety is highly reactive and readily forms covalent linkages to the side chains of Cys, His, and Lys residues in substrate protein via nucleophilic attack. Bulteau et al. demonstrated that the tryptic activity of purified myocardial 20S CPs isolated following ischemia and reperfusion injury is dramatically suppressed (80). In addition, coronary occlusion and reperfusion injury results in selective HNE modification of α1-, α2-, and α4-subunits (81). Purified 20S CPs from HNE-treated heart tissue exhibit HNE modification of the α5-, α6-, and β6-subunits (81). In the liver, only α6 is modified as a result of HNE treatment, resulting in inhibition of chymotryptic activity. Based on analyses of 20S CPs purified from HME-treated liver, Farout et al. reported that seven proteasome subunits (α2-α5, β3, β4, and β1i) are HNE modified (81). In addition, based on phage display studies using recombinant antibodies, Just et al. recently identified an unstable HNE modification on the human α7-subunit (82).
S-glutathionylation involves the addition of either glutathione (GSH) to cysteine residues or oxidized glutathione (GSSG) to reduced thiol residues, with the resulting formation of mixed disulfides in proteins. The mammalian 20S CP exhibits a biphasic response to S-glutathionylation. In the presence of micromolar concentrations of GSH or GSSG, the chymotryptic activity of purified proteasomes increases, whereas in the presence of millimolar GSH or GSSG concentrations, chymotryptic activity declines. An oxidative stress-associated decrease in activity was also reported for S-glutathionylated purified yeast 20S CPs (83). S-glutathionylation seems to affect 20S CP gate opening. For example, Silva et al. identified two S-glutathionylated Cys residues in α5-subunits purified from yeast cells (84). Proteasomes in which α5-subunits were S-glutathionylated at Cys76 and Cys221 degraded both oxidized and partially unfolded proteins more effectively than proteasomes in which these residues were in the reduced form. Furthermore, Silva et al. identified four additional modified residues in α-subunits: α5-Cys117, α6-Cys66, α7-Cys42, and α7-Cys76 (84). S-glutathionylation of α5 on Cys76 and Cys221 was shown to produce conformational changes that lead to gate opening in the 20S CP (84, 85). In α5-C76S mutant yeast, the gate remains closed, whereas in α5-C221A mutants, the gate was shown to remain open in over 90% of 20S CPs. The 20S CP gate opens to its maximum diameter when α5 Cys76 is S-glutathionylated (85). GSH and H2O2 treatment has also been shown to result in S-glutathionylation of 19S RP subunits, including Rpn1 and Rpn2 from neutrophils, HEK293 cells, and purified human 26S proteasomes (85). Analyses of extracts of mouse lung in which catalase has been inactivated also revealed that Rpn1 and Rpn2 are S-glutathionylated (86).
CORRELATION BETWEEN 26S PROTEASOME DYSFUNCTION AND NEURODEGENERATIVE DISEASES
The formation of toxic protein aggregates plays an important role in the pathogenesis of many NDs, including Alzheimer’s disease (AD), Parkinson’s disease (PD), Huntington disease (HD), amyotrophic lateral sclerosis (ALS), and prion disease. The cytotoxicity of these abnormal protein aggregates leads to cell death and concomitant tissue degeneration. Severe impairment of proteasome function in specific areas of brain is observed in many NDs.
AD
AD is characterized by progressive memory loss, personality change, and degeneration of language skills. Two characteristic types of protein aggregates are found in the brain of AD patients: senile plaque composed mainly by amyloid beta (Aβ), and neurofibrillary tangle composed of hyperphosphorylated tau protein. Postmortem analyses of brain tissues from AD patients revealed a reduction in proteasome activity in the hippocampus, related limbic structures, and the inferior parietal lobe (87). Aβ inhibits proteasome activity in a dose-dependent manner. Among several processed forms of Aβ identified to date, Aβ40 was shown to inhibit proteasome chymotryptic activity via selective localization inside the active site of the 20S CP; aggregated forms of Aβ42 inhibit proteasome activity by competing with substrates for the active site (88). Hyperphosphorylated tau also inhibits proteasome activity after cross-linking with HNE as well as the ubiquitinated form (89). Tau-associated inhibition of proteasome activity can be attenuated by treatment with PKA activators, which probably act through the phosphorylation of proteasome subunit(s) (90).
PD
PD patients exhibit both the selective loss of dopaminergic neurons in the substantia nigra pars compacta (SNpc) and the accumulation of ubiquitinated proteins. Mutations in several genes, including α-synuclein, parkin, PINK1, and LRRK2, have been identified in familiar PD patients. As in AD patients, proteasome activity is greatly reduced in the SNpc region of the brain in PD patients. PD patients also exhibit a decline in the levels of 20S CP α-subunits in the brain, PA700, and PA28 (91). Both monomeric and aggregated α-synuclein were shown to inhibit proteasome activity through interaction with the 20S CP (92). Direct binding of parkin to Rpn10 may facilitate transfer of the conjugates to the proteasome (93). Interestingly, parkin also enhances proteasome assembly by enhancing the interactions between non-ATPase and ATPase 19S subunits (32). Various PD-linked mutations in parkin have been shown to abolish the enhancement of proteasome assembly. Decreased activity of both the β5 and β1 proteasome subunits was observed in a mouse model of PD induced by injection of neurotoxin 1-methyl-4-phenylpyridinium into the SNpc area of the midbrain (94).
HD
HD progression is associated with severe neuronal losses in the thalamus, hippocampus, and spinal cord. In addition, large amounts of aggregated polyubiquitinated huntingtin protein and inclusion bodies are observed in the above-mentioned brain regions. The fibrillary form of huntingtin was shown to inhibit proteasome activity (95). The polyQ sequence interacts with the α-ring of the 20S CP, impeding substrate recognition and binding to the 19S RP (96). Proteasome activity is generally inhibited in brain of HD patients. Jeon et al. showed that transfer of the proteasome activator PA28γ gene into YAC128 HD mice enhances proteasome activity resulting in improved motor behavior (97).
ALS
ALS is characterized by the progressive degeneration of motor neurons, leading to muscle weakness. Abnormal protein aggregates are formed due to mutation in Cu, Zn-superoxide dismutase (SOD1). Studies of SOD1G93A transgenic mouse revealed that proteasome activity is inhibited in ALS primarily due to PTM of β5 and decreases in α-subunit levels (98).
Prion disease
The hallmarks of prion diseases are the accumulation of misfolded prion protein (PrPSc) and loss of normal isoform (PrPc). Progressive inhibition of proteasome activity results from direct binding of accumulating PrPSc to the 20S CP, which stabilizes the closed conformation of the gate and prevents substrate entry into the proteasome (99). The onset of proteasome dysfunction is closely correlated with PrPSc deposition, preceding the earliest behavioral deficits and neuronal loss (100).
CONCLUSION AND FUTURE PERSPECTIVE
In this review, we discussed recent findings regarding the mechanisms that regulate proteasome formation and function, including proteasome assembly, gate opening, and the effects of various PTMs. Over the past 30 years, intensive investigations of the proteasome machinery of organisms ranging from archaea to eukaryotes have greatly enhanced our understanding of the structure and function of these large multi-protein complexes. The complex nature of the structure of proteasomes and the mechanisms regulating their function has been particularly well established in yeast. However, details regarding the specific roles of most mammalian proteasome subunits remain unclear, as does the specific mechanism of substrate unfolding and translocation. We also reviewed current understanding of the putative link between proteasome dysfunction and the occurrence of NDs. Many studies have suggested that changes in proteasome activity are closely associated with a number of NDs. However, whether changes in proteasome activity lead to pathologic increases in protein accumulation or alternatively, whether final aggregate formation leads to proteasome inhibition, remains unclear. In order to fully understand and exploit the potential of the proteasome as a possible therapeutic target, whether (and if so, how) the proteasome coordinates the detoxification and clearance of aggregated proteins must be explored further.
In conclusion, obtaining a complete understanding of the specific mechanisms of proteasome assembly, proteolysis, regulation, and the pathophysiologic correlation between proteasome dysfunction and the occurrence of NDs is critical. Obtaining such an understanding could facilitate the development of therapeutics for treating NDs.
Acknowledgments
This research was supported by grants from the National Research Foundation of Korea (NRF) (2015R1A2A2A01003 080 and 2014M3C7A1064545 to KCC) funded by the Ministry of Science, ICT & Future Planning (MSIP), Republic of Korea, and by grant from the Korea Healthcare Technology R&D Project (HI14C0093 to KCC) through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea. This work was also supported in part by the Yonsei University Research Fund of 2014 (2014-12-0134 to EI), and by the Yonsei University Future-leading Research Initiative of 2015 (2015- 22-0055 to KCC).
References
- 1.Sorokin AV, Kim ER, Ovchinnikov LP. Proteasome system of protein degradation and processing. Biochemistry (Mosc) (2009);74:1411–1442. doi: 10.1134/S000629790913001X. [DOI] [PubMed] [Google Scholar]
- 2.Asano S, Fukuda Y, Beck F, et al. Proteasome. A molecular census of 26S proteasomes in intact neurons. Science. (2015);347:439–442. doi: 10.1126/science.1261197. [DOI] [PubMed] [Google Scholar]
- 3.Groll M, Ditzel L, Löwe J, et al. Structure of the 20S proteasome from yeast at 2.4 Å resolution. Nature. (1997);386:463–471. doi: 10.1038/386463a0. [DOI] [PubMed] [Google Scholar]
- 4.Griffin TA, Nandi D, Cruz M, et al. Immunoproteasome assembly: cooperative incorporation of interferon gamma (IFN-gamma)-inducible subunits. J Exp Med. (1998);187:97–104. doi: 10.1084/jem.187.1.97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kulichkova VA, Tsimokha AS, Fedorova OA, et al. 26S proteasome exhibits endoribonuclease activity controlled by extra-cellular stimuli. Cell Cycle. (2010);9:840–849. doi: 10.4161/cc.9.4.10829. [DOI] [PubMed] [Google Scholar]
- 6.Mittenberg AG, Moiseeva TN, Kuzyk VO, Barlev NA. Regulation of Endoribonuclease Activity of Alpha-Type Proteasome Subunits in Proerythroleukemia K562 Upon Hemin-Induced Differentiation. Protein J. (2016);35:17–23. doi: 10.1007/s10930-015-9642-x. [DOI] [PubMed] [Google Scholar]
- 7.Qiu XB, Ouyang SY, Li CJ, Miao S, Wang L, Goldberg AL. hRpn13/ADRM1/GP110 is a novel proteasome subunit that binds the deubiquitinating enzyme, UCH37. EMBO J. (2006);25:5742–5753. doi: 10.1038/sj.emboj.7601450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Latham MP, Sekhar A, Kay LE. Understanding the mechanism of proteasome 20S core particle gating. Proc Natl Acad Sci U S A. (2014);111:5532–5537. doi: 10.1073/pnas.1322079111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Rosenzweig R, Osmulski PA, Gaczynska M, Glickman MH. The central unit within the 29S regulatory particle of the proteasome. Nat Struct Mol Biol. (2008);15:573–580. doi: 10.1038/nsmb.1427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Thrower JS, Hoffman L, Rechsteiner M, Pickart CM. Recognition of the polyubiquitin proteolytic signal. EMBO J. (2000);19:94–102. doi: 10.1093/emboj/19.1.94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Sakata E, Bohn S, Mihalache O, et al. Localization of the proteasomal ubiquitin receptors Rpn10 and Rpn13 by electron cryomicroscopy. Proc Natl Acad Sci U S A. (2012);109:1479–1484. doi: 10.1073/pnas.1119394109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Fong A, Zhang M, Neely J, Sun SC. S9, a 19 S proteasome subunit interacting with ubiquitinated NF-kappaB2/p100. J Biol Chem. (2002);277:40697–40702. doi: 10.1074/jbc.M205330200. [DOI] [PubMed] [Google Scholar]
- 13.Glickman MH, Ciechanover A. The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol Rev. (2002);82:373–428. doi: 10.1152/physrev.00027.2001. [DOI] [PubMed] [Google Scholar]
- 14.Verma R, Aravind L, Oania R, et al. Role of Rpn11 metalloprotease in deubiquitination and degradation by the 26S proteasome. Science. (2002);298:611–615. doi: 10.1126/science.1075898. [DOI] [PubMed] [Google Scholar]
- 15.Dahlmann B. Proteasome. Essays Biochem. (2005);41:31–48. doi: 10.1042/bse0410031. [DOI] [PubMed] [Google Scholar]
- 16.Gao X, Li J, Pratt G, Wilk S, Rechsteiner M. Purification procedures determine the proteasome activation properties of REGγ (PA28γ). Arch Biochem Biophys. (2004);425:158–164. doi: 10.1016/j.abb.2004.03.021. [DOI] [PubMed] [Google Scholar]
- 17.Sadre-Bazzaz K, Whitby FG, Robinson H, Formosa T, Hill CP. Structure of a Blm10 complex reveals common mechanisms for proteasome binding and gate opening. Mol Cell. (2010);37:728–735. doi: 10.1016/j.molcel.2010.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ustrell V, Hoffman L, Pratt G, Rechsteiner M. PA200, a nuclear proteasome activator involved in DNA repair. EMBO J. (2002);21:3516–3525. doi: 10.1093/emboj/cdf333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Sakai N, Sawada MT, Sawada H. Non-traditional roles of ubiquitin-proteasome system in fertilization and gametogenesis. Int J Biochem Cell Biol. (2004);36:776–784. doi: 10.1016/S1357-2725(03)00263-2. [DOI] [PubMed] [Google Scholar]
- 20.Zhou C, Wee S, Rhee E, Naumann M, Dubiel W, Wolf DA. Fission yeast COP9/signalosome suppresses cullin activity through recruitment of the deubiquitylating enzyme Ubp12p. Mol Cell. (2003);11:927–938. doi: 10.1016/S1097-2765(03)00136-9. [DOI] [PubMed] [Google Scholar]
- 21.Murata S, Yashiroda H, Tanaka K. Molecular mechanisms of proteasome assembly. Nat Rev Mol Cell Biol. (2009);10:104–115. doi: 10.1038/nrm2630. [DOI] [PubMed] [Google Scholar]
- 22.Panfair D, Ramamurthy A, Kusmierczyk AR. Alpha-ring independent assembly of the 20S proteasome. Sci Rep. (2015);5:13130. doi: 10.1038/srep13130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Tamura T, Nagy I, Lupas A, et al. The first characterization of a eubacterial proteasome: the 20S complex of Rhodococcus. Curr Biol. (1995);5:766–774. doi: 10.1016/S0960-9822(95)00153-9. [DOI] [PubMed] [Google Scholar]
- 24.Kwon YD, Nagy I, Adams PD, Baumeister W, Jap BK. Crystal structures of the Rhodococcus proteasome with and without its pro-peptides: implications for the role of the pro-peptide in proteasome assembly. J Mol Biol. (2004);335:233–245. doi: 10.1016/j.jmb.2003.08.029. [DOI] [PubMed] [Google Scholar]
- 25.Hirano Y, Kaneko T, Okamoto K, et al. Dissecting β-ring assembly pathway of the mammalian 20S proteasome. EMBO J. (2008);27:2204–2213. doi: 10.1038/emboj.2008.148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Sahara K, Kogleck L, Yashiroda H, Murata S. The mechanism for molecular assembly of the proteasome. Adv Biol Regul. (2014);54:51–58. doi: 10.1016/j.jbior.2013.09.010. [DOI] [PubMed] [Google Scholar]
- 27.Marques AJ, Glanemann C, Ramos PC, Dohmen RJ. The C-terminal extension of the beta7 subunit and activator complexes stabilize nascent 20 S proteasomes and promote their maturation. J Biol Chem. (2007);282:34869–34876. doi: 10.1074/jbc.M705836200. [DOI] [PubMed] [Google Scholar]
- 28.Kock M, Nunes MM, Hemann M, et al. Proteasome assembly from 15S precursors involves major conformational changes and recycling of the Pba1-Pba2 chaperone. Nat Commun. (2015);6:6123. doi: 10.1038/ncomms7123. [DOI] [PubMed] [Google Scholar]
- 29.Bai M, Zhao X, Sahara K, et al. Assembly mechanisms of specialized core particles of the proteasome. Biomolecules. (2014);4:662–677. doi: 10.3390/biom4030662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Murata S, Sasaki K, Kishimoto T, et al. Regulation of CD8+ T cell development by thymus-specific proteasomes. Science. (2007);316:1349–1353. doi: 10.1126/science.1141915. [DOI] [PubMed] [Google Scholar]
- 31.Park S, Roelofs J, Kim W, et al. Hexameric assembly of the proteasomal ATPases is templated through their C termini. Nature. (2009);459:866–870. doi: 10.1038/nature08065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Um JW, Im E, Lee HJ, et al. Parkin directly modulates 26S proteasome activity. J Neurosci. (2010);30:11805–11814. doi: 10.1523/JNEUROSCI.2862-09.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Hanssum A, Zhong Z, Rousseau A, Krzyzosiak A, Sigurdardottir A, Bertolotti A. An inducible chaperone adapts proteasome assembly to stress. Mol Cell. (2014);55:566–577. doi: 10.1016/j.molcel.2014.06.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Besche HC, Peth A, Goldberg AL. Getting to first base in proteasome assembly. Cell. (2009);138:25–28. doi: 10.1016/j.cell.2009.06.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Estrin E, Lopez-Blanco JR, Chacon P, Martin A. Formation of an intricate helical bundle dictates the assembly of the 26S proteasome lid. Structure. (2013);21:1624–1635. doi: 10.1016/j.str.2013.06.023. [DOI] [PubMed] [Google Scholar]
- 36.Tomko RJ, Jr, Taylor DW, Chen ZA, Wang HW, Rappsilber J, Hochstrasser M. A Single α Helix Drives Extensive Remodeling of the Proteasome Lid and Completion of Regulatory Particle Assembly. Cell. (2015);163:432–444. doi: 10.1016/j.cell.2015.09.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Hu Y, Wu Y, Li Q, Zhang W, Jin C. Solution structure of yeast Rpn9: insights into proteasome lid assembly. J Biol Chem. (2015);290:6878–6889. doi: 10.1074/jbc.M114.626762. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Tian G, Park S, Lee MJ, et al. An asymmetric interface between the regulatory and core particles of the proteasome. Nat Struct Mol Biol. (2011);18:1259–1267. doi: 10.1038/nsmb.2147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Sokolova V, Li F, Polovin G, Park S. Proteasome Activation is Mediated via a Functional Switch of the Rpt6 C-terminal Tail Following Chaperone-dependent Assembly. Sci Rep. (2015);5:14909. doi: 10.1038/srep14909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Groll M, Bajorek M, Köhler A, et al. A gated channel into the proteasome core particle. Nature Struct Biol. (2000);7:1062–1067. doi: 10.1038/80992. [DOI] [PubMed] [Google Scholar]
- 41.Rabl J, Smith DM, Yu Y, Chang SC, Goldberg AL, Cheng Y. Mechanism of gate opening in the 20S proteasome by the proteasomal ATPases. Mol Cell. (2008);30:360–368. doi: 10.1016/j.molcel.2008.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Forster A, Masters EI, Whitby FG, Robinson H, Hill CP. The 1.9 A° structure of a proteasome-11S activator complex and implications for proteasome-PAN/PA700 interactions. Mol Cell. (2005);18:589–599. doi: 10.1016/j.molcel.2005.04.016. [DOI] [PubMed] [Google Scholar]
- 43.Dange T, Smith D, Noy T, et al. Blm10 protein promotes proteasomal substrate turnover by an active gating mechanism. J Biol Chem. (2011);286:42830–42839. doi: 10.1074/jbc.M111.300178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Arendt CS, Hochstrasser M. Eukaryotic 20S proteasome catalytic subunit propeptides prevent active site inactivation by N-terminal acetylation and promote particle assembly. EMBO J. (1999);18:3575–3585. doi: 10.1093/emboj/18.13.3575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Sun XM, Butterworth M, MacFarlane M, Dubiel W, Ciechanover A, Cohen GM. Caspase activation inhibits proteasome function during apoptosis. Mol Cell. (2004);14:81–93. doi: 10.1016/S1097-2765(04)00156-X. [DOI] [PubMed] [Google Scholar]
- 46.Jang M, Park BC, Lee AY, et al. Caspase-7 mediated cleavage of proteasome subunits during apoptosis. Biochem Biophys Res Commun. (2007);363:388–394. doi: 10.1016/j.bbrc.2007.08.183. [DOI] [PubMed] [Google Scholar]
- 47.Wang XH, Zhang L, Mitch WE, Ledoux JM, Hu J, Du J. Caspase-3 cleaves specific 19S proteasome subunits in skeletal muscle stimulating proteasome activity. J Biol Chem. (2010);285:21249–21257. doi: 10.1074/jbc.M109.041707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Gomes AV, Zong C, Edmondson RD, et al. Mapping the murine cardiac 26S proteasome complexes. Circ Res. (2006);99:362–371. doi: 10.1161/01.RES.0000237386.98506.f7. [DOI] [PubMed] [Google Scholar]
- 49.Wang X, Chen CF, Baker PR, Chen PL, Kaiser P, Huang L. Mass spectrometric characterization of the affinity-purified human 26S proteasome complex. Biochemistry. (2007);46:3553–3565. doi: 10.1021/bi061994u. [DOI] [PubMed] [Google Scholar]
- 50.Anandan T, Han J, Baun H, et al. Phosphorylation regulates mycobacterial proteasome. J Microbiol. (2014);52:743–754. doi: 10.1007/s12275-014-4416-2. [DOI] [PubMed] [Google Scholar]
- 51.Zong C, Gomes AV, Drews O, et al. Regulation of murine cardiac 20S proteasomes: role of associating partners. Circ Res. (2006);99:372–380. doi: 10.1161/01.RES.0000237389.40000.02. [DOI] [PubMed] [Google Scholar]
- 52.Li D, Dong Q, Tao Q, et al. c-Abl regulates proteasome abundance by controlling the ubiquitinproteasomal degradation of PSMA7 subunit. Cell Rep. (2015);10:484–496. doi: 10.1016/j.celrep.2014.12.044. [DOI] [PubMed] [Google Scholar]
- 53.Bose S, Stratford FL, Broadfoot KI, Mason GG, Rivett AJ. (2004) Phosphorylation of 20s proteasome alpha subunit C8 (Alpha7) stabilizes the 26s proteasome and plays a role in the regulation of proteasome complexes by gamma-interferon. Biochem J. (2004);378:177–184. doi: 10.1042/bj20031122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Kikuchi J, Iwafune Y, Akiyama T, et al. Co- and post-translational modifications of the 26S proteasome in yeast. Proteomics. (2010);10:2769–2779. doi: 10.1002/pmic.200900283. [DOI] [PubMed] [Google Scholar]
- 55.Konstantinova IM, Tsimokha AS, Mittenberg AG. Role of proteasomes in cellular regulation. Int Rev Cell Mol Biol. (2008);267:59–124. doi: 10.1016/S1937-6448(08)00602-3. [DOI] [PubMed] [Google Scholar]
- 56.Um JW, Im E, Park J, et al. ASK1 negatively regulates the 26 S proteasome. J Biol Chem. (2010);285:36434–36446. doi: 10.1074/jbc.M110.133777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Guo X, Wang X, Wang Z. Site-specific proteasome phosphorylation controls cell proliferation and tumorigenesis. Nat Cell Biol. (2016);18:202–212. doi: 10.1038/ncb3289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Lokireddy S, Kukushkin NV, Goldberg AL. cAMP-induced phosphorylation of 26S proteasomes on Rpn6/PSMD11 enhances their activity and the degradation of misfolded proteins. Proc Natl Acad Sci U S A. (2015);112:E7176–7185. doi: 10.1073/pnas.1522332112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Satoh K, Sasajima H, Nyoumura KI, Yokosawa H, Sawada H. Assembly of the 26S proteasome is regulated by phosphorylation of the P45/Rpt6 ATPase subunit. Biochemistry. (2001);40:314–319. doi: 10.1021/bi001815n. [DOI] [PubMed] [Google Scholar]
- 60.Jarome TJ, Ferrara NC, Kwapis JL, Helmstetter FJ. CaMKII regulates proteasome phosphorylation and activity and promotes memory destabilization following retrieval. Neurobiol Learn Mem. (2016);128:103–109. doi: 10.1016/j.nlm.2016.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Besche HC, Sha Z, Kukushkin NV, et al. Autoubiquitination of the 26S proteasome on Rpn13 regulates breakdown of ubiquitin conjugates. EMBO J. (2014);33:1159–1176. doi: 10.1002/embj.201386906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Uchiki T, Kim HT, Zhai B, et al. The ubiquitininteracting motif protein, S5a, is ubiquitinated by all types of ubiquitin ligases by a mechanism different from typical substrate recognition. J Biol Chem. (2009);284:12622–12632. doi: 10.1074/jbc.M900556200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Isasa M, Katz EJ, Kim W, et al. Monoubiquitination of RPN10 regulates substrate recruitment to the proteasome. Mol Cell. (2010);38:733–745. doi: 10.1016/j.molcel.2010.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Crosas B, Hanna J, Kirkpatrick DS, et al. Ubiquitin chains are remodeled at the proteasome by opposing ubiquitin ligase and deubiquitinating activities. Cell. (2006);127:1401–1413. doi: 10.1016/j.cell.2006.09.051. [DOI] [PubMed] [Google Scholar]
- 65.Schmitt SM, Neslund-Dudas C, Shen M, Cui C, Mitra B, Dou QP. Involvement of ALAD-20S proteasome complexes in ubiquitination and acetylation of proteasomal α2 subunits. J Cell Biochem. (2016);117:144–151. doi: 10.1002/jcb.25259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Cui Z, Scruggs SB, Gilda JE, Ping P, Gomes AV. Regulation of cardiac proteasomes by ubiquitination, SUMOylation, and beyond. J Mol Cell Cardiol. (2014);71:32–42. doi: 10.1016/j.yjmcc.2013.10.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Manza LL, Codreanu SG, Stamer SL, et al. Global shifts in protein sumoylation in response to electrophile and oxidative stress. Chem Res Toxicol. (2004);17:1706–1715. doi: 10.1021/tx049767l. [DOI] [PubMed] [Google Scholar]
- 68.Kimura Y, Takaoka M, Tanaka S, et al. Nα-acetylation and proteolytic activity of the yeast 20 S proteasome. J Biol Chem. (2000);275:4635–4639. doi: 10.1074/jbc.275.7.4635. [DOI] [PubMed] [Google Scholar]
- 69.Kimura Y, Saeki Y, Yokosawa H, Polevoda B, Sherman F, Hirano H. N-Terminal modifications of the 19S regulatory particle subunits of the yeast proteasome. Arch Biochem Biophys. (2003);409:341–348. doi: 10.1016/S0003-9861(02)00639-2. [DOI] [PubMed] [Google Scholar]
- 70.Wang D, Fang C, Zong NC, et al. Regulation of acetylation restores proteolytic function of diseased myocardium in mouse and human. Mol Cell Proteomics. (2013);12:3793–3802. doi: 10.1074/mcp.M113.028332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Zhang F, Su K, Yang X, Bowe DB, Paterson AJ, Kudlow JE. O-GlcNAc modification is an endogenous inhibitor of the proteasome. Cell. (2003);115:715–725. doi: 10.1016/S0092-8674(03)00974-7. [DOI] [PubMed] [Google Scholar]
- 72.Zong C, Young GW, Wang Y, et al. Two-dimensional electrophoresis-based characterization of post-translational modifications of mammalian 20S proteasome complexes. Proteomics. (2008);8:5025–5037. doi: 10.1002/pmic.200800387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Overath T, Kuckelkorn U, Henklein P, et al. Mapping of O-GlcNAc sites of 20 S proteasome subunits and Hsp90 by a novel biotin-cystamine tag. Mol Cell Proteomics. (2012);11:467–477. doi: 10.1074/mcp.M111.015966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Wang Z, Park K, Comer F, Hsieh-Wilson LC, Saudek CD, Hart GW. Site-specific GlcNAcylation of human erythrocyte proteins: potential biomarker(s) for diabetes. Diabetes. (2009);58:309–317. doi: 10.2337/db08-0994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Weinert BT, Schölz C, Wagner SA, et al. Lysine succinylation is a frequently occurring modification in prokaryotes and eukaryotes and extensively overlaps with acetylation. Cell Rep. (2013);4:842–851. doi: 10.1016/j.celrep.2013.07.024. [DOI] [PubMed] [Google Scholar]
- 76.Catalgol B, Wendt B, Grimm S, Breusing N, Ozer NK, Grune T. Chromatin repair after oxidative stress: role of parp-mediated proteasome activation. Free Radic Biol Med. (2010);48:673–680. doi: 10.1016/j.freeradbiomed.2009.12.010. [DOI] [PubMed] [Google Scholar]
- 77.Cho-Park PF, Steller H. Proteasome regulation by ADP-ribosylation. Cell. (2013);153:614–627. doi: 10.1016/j.cell.2013.03.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Ishii T, Sakurai T, Usami H, Uchida K. Oxidative modification of proteasome: identification of an oxidation-sensitive subunit in 26S proteasome. Biochemistry. (2005);44:13893–13901. doi: 10.1021/bi051336u. [DOI] [PubMed] [Google Scholar]
- 79.Divald A, Kivity S, Wang P. Myocardial ischemic preconditioning preserves postischemic function of the 26S proteasome through diminished oxidative damage to 19S regulatory particle subunits. Circ Res. (2010);106:1829–1838. doi: 10.1161/CIRCRESAHA.110.219485. [DOI] [PubMed] [Google Scholar]
- 80.Bulteau AL, Lundberg KC, Humphries KM, et al. Oxidative modification and inactivation of the proteasome during coronary occlusion/reperfusion. J Biol Chem. (2001);276:30057–30063. doi: 10.1074/jbc.M100142200. [DOI] [PubMed] [Google Scholar]
- 81.Farout L, Mary J, Vinh J, Szweda LI, Friguet B. Inactivation of the proteasome by 4-hydroxy-2-nonenal is site specific and dependant on 20S proteasome subtypes. Arch Biochem Biophys. (2006);453:135–142. doi: 10.1016/j.abb.2006.02.003. [DOI] [PubMed] [Google Scholar]
- 82.Just J, Jung T, Friis NA, et al. Identification of an unstable 4-hydroxynoneal modification on the 20S proteasome subunit α7 by recombinant antibody technology. Free Radic Biol Med. (2015);89:786–792. doi: 10.1016/j.freeradbiomed.2015.10.405. [DOI] [PubMed] [Google Scholar]
- 83.Demasi M, Silva GM, Netto LE. 20 S proteasome from Saccharomyces cerevisiae is responsive to redox modifications and is S-glutathionylated. J Biol Chem. (2003);278:679–685. doi: 10.1074/jbc.M209282200. [DOI] [PubMed] [Google Scholar]
- 84.Silva GM, Netto LE, Simões V, et al. Redox control of 20S proteasome gating. Antioxid Redox Signal. (2012);16:1183–1194. doi: 10.1089/ars.2011.4210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Demasi M, Netto LE, Silva GM, et al. Redox regulation of the proteasome via S-glutathionylation. Redox Biol. (2014);2:44–51. doi: 10.1016/j.redox.2013.12.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Zmijewski JW, Banerjee S, Abraham E. S-glutathionylation of the Rpn2 regulatory subunit inhibits 26 S proteasomal function. J Biol Chem. (2009);284:22216–22221. doi: 10.1074/jbc.M109.028902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Keller JN, Hanni KB, Markesbery WR. Impaired proteasome function in Alzheimer’s disease. J Neurochem. (2000);75:436–439. doi: 10.1046/j.1471-4159.2000.0750436.x. [DOI] [PubMed] [Google Scholar]
- 88.Crouch PJ, Harding SM, White AR, Camakaris J, Bush AI, Masters CL. Mechanisms of Abeta mediated neurodegeneration in Alzheimer’s disease. Int J Biochem Cell Biol. (2008);40:181–198. doi: 10.1016/j.biocel.2007.07.013. [DOI] [PubMed] [Google Scholar]
- 89.Cripps D, Thomas SN, Jeng Y, Yang F, Davies P, Yang AJ. Alzheimer disease-specific conformation of hyperphosphorylated paired helical filament-tau is polyubiquitinated through lys-48, lys-11, and lys-6 ubiquitin conjugation. J Biol Chem. (2006);281:10825–10838. doi: 10.1074/jbc.M512786200. [DOI] [PubMed] [Google Scholar]
- 90.Myeku N, Clelland CL, Emrani S. Tau-driven 26S proteasome impairment and cognitive dysfunction can be prevented early in disease by activating cAMP-PKA signaling. Nat Med. (2016);22:46–53. doi: 10.1038/nm.4011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.McNaught KS, Belizaire R, Isacson O, Jenner P, Olanow CW. Altered proteasomal function in sporadic Parkinson’s disease. Exp Neurol. (2003);179:38–46. doi: 10.1006/exnr.2002.8050. [DOI] [PubMed] [Google Scholar]
- 92.Snyder H, Mensah K, Theisler C, Lee J, Matouschek A, Wolozin B. Aggregated and monomeric alpha- synuclein bind to the S6’ proteasomal protein and inhibit proteasomal function. J Biol Chem. (2003);278:11753–11759. doi: 10.1074/jbc.M208641200. [DOI] [PubMed] [Google Scholar]
- 93.Sakata E, Yamaguchi Y, Kurimoto E, et al. Parkin binds the Rpn10 subunit of 26S proteasomes through its ubiquitin-like domain. EMBO Rep. (2003);4:301–306. doi: 10.1038/sj.embor.embor764. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Caneda-Ferron B, De Girolamo LA, Costa T, Beck KE, Layfield R, Billett EE. Assessment of the direct and indirect effects of MPP+ and dopamine on the human proteasome: implications for Parkinson’s disease aetiology. J Neurochem. (2008);105:225–238. doi: 10.1111/j.1471-4159.2007.05130.x. [DOI] [PubMed] [Google Scholar]
- 95.Goswami A, Dikshit P, Mishra A, Mulherkar S, Nukina N, Jana NR. Oxidative stress promotes mutant huntingtin aggregation and mutant huntingtin-dependent cell death by mimicking proteasomal malfunction. Biochem Biophys Res Commun. (2006);342:184–190. doi: 10.1016/j.bbrc.2006.01.136. [DOI] [PubMed] [Google Scholar]
- 96.Ortega Z, Diaz-Hernandez M, Lucas JJ. Is the ubiquitin-proteasome system impaired in Huntington’s disease. Cell Mol Life Sci. (2007);64:2245–2257. doi: 10.1007/s00018-007-7222-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Jeon J, Kim W, Jang J, Isacson O, Seo H. Gene therapy by proteasome activator, PA28γ, improves motor coordination and proteasome function in Huntington's disease YAC128 mice. Neuroscience. (2016);324:20–28. doi: 10.1016/j.neuroscience.2016.02.054. [DOI] [PubMed] [Google Scholar]
- 98.Kabashi E, Agar JN, Taylor DM, Minotti S, Durham HD. Focal dysfunction of the proteasome: a pathogenic factor in a mouse model of amyotrophic lateral sclerosis. J Neurochem. (2004);89:1325–1335. doi: 10.1111/j.1471-4159.2004.02453.x. [DOI] [PubMed] [Google Scholar]
- 99.Andre R, Tabrizi SJ. Misfolded PrP and a novel mechanism of proteasome inhibition. Prion. (2012);6:32–36. doi: 10.4161/pri.6.1.18272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.McKinnon C, Goold R, Andre R, et al. Prionmediated neurodegeneration is associated with early impairment of the ubiquitin-proteasome system. Acta Neuropathol. (2016);131:411–425. doi: 10.1007/s00401-015-1508-y. [DOI] [PMC free article] [PubMed] [Google Scholar]