Abstract
Background
Tospovirus is a plant-infecting genus within the family Bunyaviridae, which also includes four animal-infecting genera: Hantavirus, Nairovirus, Phlebovirus and Orthobunyavirus. Compared to these members, the structures of Tospovirus proteins still are poorly understood. Despite multiple studies have attempted to identify candidate N protein regions involved in RNA binding and protein multimerization for tospovirus using yeast two-hybrid systems (Y2HS) and site-directed mutagenesis, the tospovirus ribonucleocapsids (RNPs) remains largely uncharacterized at the molecular level and the lack of structural information prevents detailed insight into these interactions.
Results
Here we used the nucleoprotein structure of LACV (La Crosse virus-Orthobunyavirus) and molecular dynamics simulations to access the structure and dynamics of the nucleoprotein from tospovirus GRSV (Groundnut ringspot virus). The resulting model is a monomer composed by a flexible N-terminal and C-terminal arms and a globular domain with a positively charged groove in which RNA is deeply encompassed. This model allowed identifying the candidate amino acids residues involved in RNA interaction and N-N multimerization. Moreover, most residues predicted to be involved in these interactions are highly conserved among tospoviruses.
Conclusions
Crucially, the interaction model proposed here for GRSV N is further corroborated by the all available mutational studies on TSWV (Tomato spotted wilt virus) N, so far. Our data will help designing further and more accurate mutational and functional studies of tospovirus N proteins. In addition, the proposed model may shed light on the mechanisms of RNP shaping and could allow the identification of essential amino acid residues as potential targets for tospovirus control strategies.
Electronic supplementary material
The online version of this article (doi:10.1186/s12859-016-1339-4) contains supplementary material, which is available to authorized users.
Keywords: Homology modeling, Molecular dynamics, Tospovirus, Nucleoprotein
Background
Tospovirus is a thrips-borne plant-infecting genus within the family Bunyaviridae, which also includes four animal-infecting genera: Hanta/Nairo/Phlebo- and Orthobunyavirus [1]. GRSV (Groundnut ringspot virus) is an emerging tospovirus, that has caused severe diseases in distinct vegetable crops in South America and is phylogenetically close to the tospovirus type-species TSWV (Tomato spotted wilt virus) [2]. Like all tospoviruses, GRSV contain a trisegmented negative single-stranded RNA (ssRNA) genome that encodes the viral RNA-dependent RNA polymerase (RdRp), two glycoproteins (Gn/Gc), the movement protein (NSm), the RNA silencing suppressor protein (NSs) and the nucleoprotein (N) [3]. N is a multifunctional protein involved in RNA protection, particle assembly, intracellular movement and might play a role in transcription/replication regulation [4–14]. Multiple copies of the N protein form oligomers that interact with the viral RNAs to build ribonucleoprotein complexes (RNPs) that are proposed to be transported via plasmodesmata and are functional templates for RNA replication and transcription [6, 15, 16].
Multiple studies have attempted to identify candidate N protein regions involved in RNA binding and protein multimerization for TSWV using yeast two-hybrid systems (Y2HS) and site-directed mutagenesis [4, 6, 17, 18], but the tospovirus RNPs remains largely uncharacterized at the molecular level and the lack of structural information prevents detailed insight into these interactions. The lack of a reverse genetics system, which is available for other bunyaviruses, has hampered tospovirus research. The N protein crystal structures of related RNA virus families (Arena/Orthomyxo/Bunyaviridae) have been elucidated [8, 19–26] and despite different size and distinct N-folding structures, there are common features and architectural principles by which these proteins form N-N multimers and N-RNA complexes [27]. Therefore, these available structures were used to predict a three-dimensional model for GRSV N (the most important and prevalent tospovirus in Brazil) using homology modeling.
Results and discussion
Three-dimensional model of GRSV N and oligomerization
The GRSV N and LACV N have similar protein fold with the predicted GRSV N monomer forming thirteen helical segments and two small beta-sheets (Figs. 1, 2a and e-f). The protein has a globular core domain (26–223 aa) containing a deep positively charged groove with the two chain terminals forming an N-terminus arm (1–25 aa) and a C-terminus arm (224–258 aa) (Fig. 2a-b and Fig. 3). The N- and C-arms extend outwards from the globular core domain and interacts with the globular core domain of neighboring monomers to mediate the multimerization, supporting the “head-to-tail” model proposed by [18]. Amino acids S2-V12 of the N-arm interact with the Q61-N82 of the core domain of one neighboring monomer (Fig. 2c and e) while K227-K249 of the C-arm interact with the K173–K198 of the core domain the other neighboring monomer (Fig. 2d and f). Specific residue-residue interactions have been listed in Table 1 for the two independent interfaces. According to PISA, the intermolecular interactions were mainly hydrogen bonds, but van der Waals and hydrophobic interactions also contribute to hold the monomers together (data not shown). This interaction model is further corroborated by the available mutational studies on TSWV N [4, 17, 18].
Fig. 1.
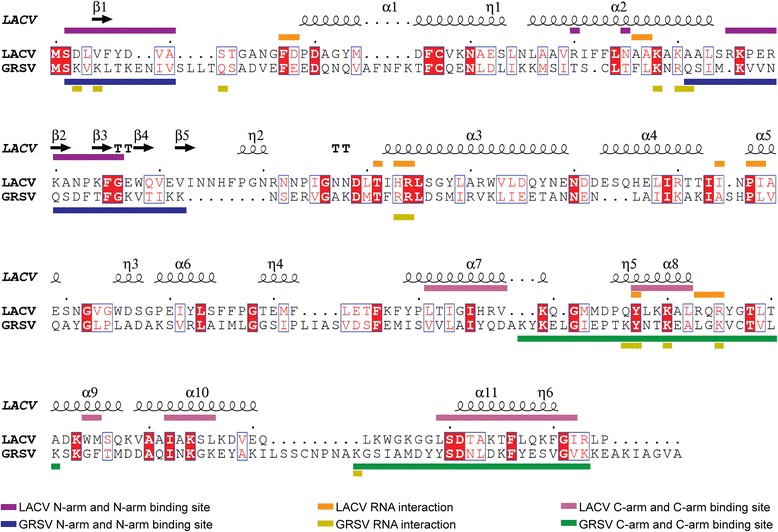
Groundnut ringspot virus (GRSV) and La crosse virus (LACV) Nucleoproteins sequence alignment. Key residues for GRSV N and LACV N oligomerization and for ssRNA binding are colored as indicated by the colored bars. The secondary structure of LACV N is shown above, and every 10 residues are indicated with a dot (.). Strictly conserved residues are highlighted in red with white letter and highly conserved residues are displayed by red letters. GRSV N GenBank accession number is AF251271 and LACV N UniProt accession code is P04873
Fig. 2.
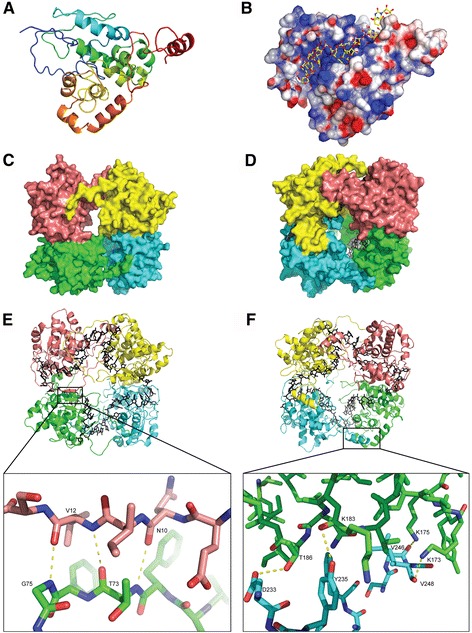
Monomeric and tetrameric structure of the Groundnut ringspot virus (GRSV) nucleoprotein (N). a Cartoon representation of monomeric GRSV N with rainbow coloring from N- (blue) to C-terminus (red). b Electrostatic surface of the GRSV N with a positively charged groove in complex with RNA shown as yellow (carbons) and red (oxygens) sticks. Positive and negative charges are blue and red, respectively. c N-terminus interaction surface representation of four GRSV N monomers A, B, C, D shown in color pink, yellow, cyan and green, respectively. d C-terminus interaction surface representation of the GRSV N tetrameric ring. The RNA is shown in black sticks deeply bound inside the tetrameric ring. e Cartoon representation with the N-arm oligomerization interface showing interacting residues. The N-terminal arm is in pink and the globular region is in green. The intermolecular hydrogen bonds are shown as yellow dotted lines. f 180° rotation of Fig. 2e, C-arm oligomerization interface showing interacting residues. The C-terminal arm is in cyan and the globular region is in green
Fig. 3.
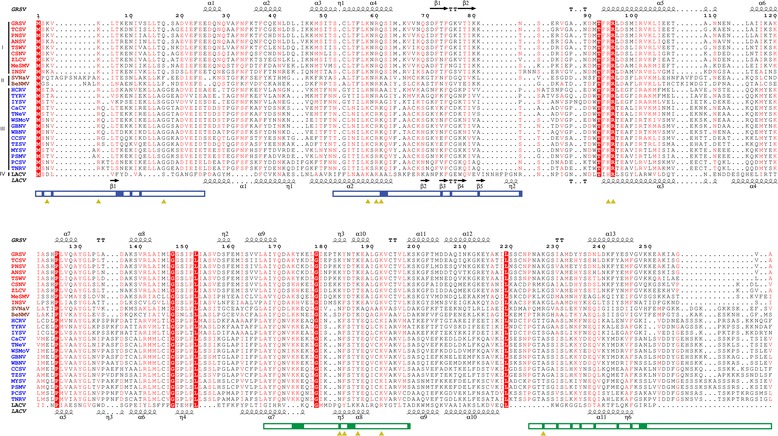
Sequence alignment of representative tospoviruses Nucleoproteins (N). The secondary structure of Groundnut ringspot virus (GRSV) is shown above and of La crosse virus (LACV) is shown at the bottom. Key residues for GRSV N ssRNA binding are marked with yellow triangles. GRSV N- and C-arms are marked with blue and green boxes respectively, with key residues for oligomerization highlighted. Strictly conserved residues are highlighted in red with white letter and highly conserved residues with red letter. I: Tospovirus American clade I; II: Tospovirus American clade II; III: Tospovirus Eurasian clade; IV: Orthobunyavirus. The sequence codes are supplied at the Additional file 2: Table S1
Table 1.
Pairs of interacting residues for GRSV N-N oligomerization
| N-arma | N-arm binding siteb | C-armc | C-arm binding siteb |
|---|---|---|---|
| S2d | S83 | A226 | K183 |
| V4 | N82 | S229 | K183 |
| T7 | Q61 | D233 | T186 |
| T7 | S62 | Y235 | K183 |
| K8 | S62 | N238 | K198 |
| N10 | T73 | Y243 | N185 |
| V12 | T73 | V246 | K175 |
| V12 | G75 | V248 | K173 |
| K249 | Y174 |
aN-arm amino acids residues of GRSV N
bInteracting amino acids residues of GRSV N globular core domain
cC-arm amino acids residues of GRSV N
dAmino acids residues position in the GRSV N sequence
Actually, the first assay to map functional domains of TSWV N, performing Y2HS and random serial deletions, showed that both the N- (1–39 aa) and C-terminals (233–248 aa) were important for N-N interaction [18], in clear agreement with the structural results presented here. Furthermore, [17] identified three crucial intermonomer binding regions: 42–56, 132–152 and 222–248 which have a clear correspondence with the predicted interaction residues of GRSV N located at N- and C-arms, or buried in the core of the model (Fig. 3). Moreover, amino acids residues located at the regions K103-A119 and L132-V135 are solvent accessible and therefore are able to interact with NSm, glycoproteins, viral polymerase or host proteins [6, 7]. Recently, studies have been performed attempting to identify N-NSm interactions [28, 29] which results are in perfect congruence with the GRSV N protein model. In both cases, the model proposed here represents an efficient tool to assist in planning experiments with mutations and deletion in the N protein.
In addition, the obtained model for N protein was submitted to molecular dynamics simulations in order to both refine the structure in aqueous solvent [30, 31] and access the protein conformational ensemble, further exploring its structural and functional roles. During the simulation time, the globular core domain did not reveal any loss of secondary structure, increase of radius of gyration or persistent increments on RMSD values, which supports the model quality. It is worthy to mention that RMSF calculations indicate the N-terminal arm (1–25 aa) as a very flexible region (Fig. 4c).
Fig. 4.
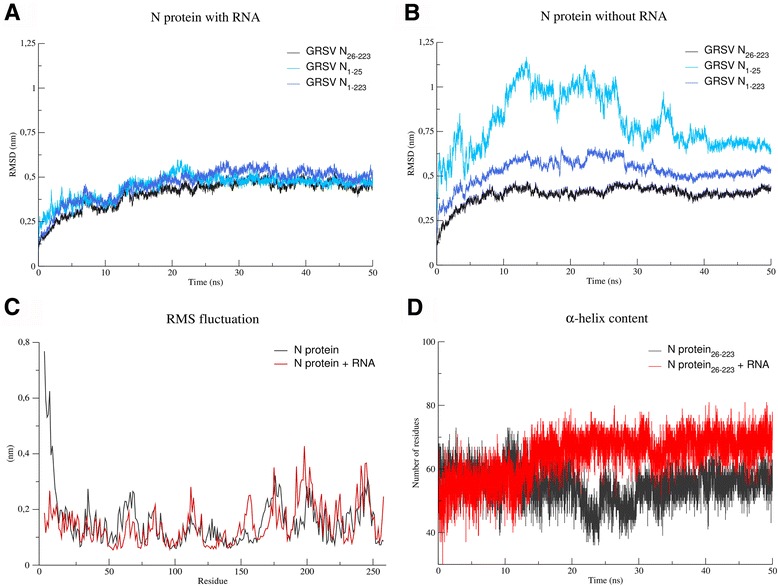
Molecular Dynamics of monomeric Nucleoprotein (N) of Groundnut ringspot virus (GRSV). Root Mean Square Deviation (RMSD) calculations for different set of atoms in both presence a and absence b of RNA. c Root Mean Square Fluctuations (RMSF) calculations for the entire N protein in both presence (red) and absence (black) of RNA. d Plot of α-helix content as function of time in both presence (red) and absence (black) of RNA
RNA interaction
According to the GRSV N protein model, the RNA is primarily bound at the central RNA-binding groove (Fig. 2b), and the key residues for this interaction (K3, K5, Q17, K58, R60, Q61, R94, R95, K183, Y184, K187, K192 and K227) are mainly located in this positively charged groove. This positively charged groove is only possible because residues F37, F56, F72, F74, I79, M91, F93 and L96 form a hydrophobic core, which is indispensable to stabilize the protein folding and to correctly orient the RNA interacting residues towards the groove. Importantly, these residues are highly conserved among all tospoviruses (Fig. 3). Note that the N-terminal arm is also involved in RNA binding and shielding RNA from the solvent (Fig. 2c-d). Residues F23, L54, F56, L57 and F93 were observed to modulate the RNA nucleobases dynamics during the performed simulation, while the N-terminal arm seems to play a stabilization role during MD simulations of GRSV N protein (Fig. 4a and b). In addition, the content of alpha-helices in GRSV N protein bound to RNA increased 25 % during the simulation in comparison to the free monomer (Fig. 4d), suggesting that, in the simulated timescale, the monomeric state does not present a lack of conformational stability in detriment of oligomeric states, as observed experimentally for other viruses [32, 33].
Recently, the residues R60, R94, and R95 were confirmed to interact with RNA [33], which also supports our results. RNA is strongly bent at each N-N interface and is largely solvent-inaccessible in the tetramer (Fig. 2d). The dimensions of the groove can accommodate ssRNA and PISA analysis showed that the majority of residue-nucleotide interactions occur with the ribose and the phosphate moieties, suggesting a non-sequence-specific RNA interaction. Indeed, Richmond et al. [4] carried out mutagenesis and gel shift assay studies to identify N regions important for ssRNA binding and demonstrated that the N-RNA complex is highly stable and non-sequence-specific, further supporting these results.
Conclusions
Taken together, these data will help designing further and more accurate mutational and functional studies of tospovirus N proteins. In addition, the proposed model may shed light on the mechanisms of RNP shaping and could allow the identification of essential amino acid residues as potential targets for tospovirus control strategies.
Methods
In silico homology modeling and model optimization
A template for modeling the GRSV N protein was searched in expasy SWISS-MODEL server [34] using the amino acid sequence of GRSV N as a reference. Template crystal structures of Orthobunyavirus genus were chosen due to their genetic relationship. The LACV (La Crosse virus-Orthobunyavirus) N tetrameric crystal structure in complex with ssRNA (PDB ID 4BHH) was selected as the template [20], aligned with GRSV N using T-Coffee server [35] and the resulting alignment was manually improved using BioEdit [36]. Aligned sequences were used with MODELLERv9.10 [37] to develop high quality tetrameric models along with or without RNA.
Optimization of the models was achieved using energy minimization protocols available at Yasara [38] and Chiron [39] servers. Quality of the 3D models were evaluated with ERRAT (version 2.0) [40] and MOL probity [41]. Ramachandran plots for the models were assessed and Ramachandran outlier residues were fixed with COOT [42] and energy minimization. The highest quality model with 90.1 % residues in favored region and 8.4 % in allowed region while 1.5 % outlier at Ramachandran plot was selected after visual inspection (see Additional file 1: Figure S1). The model was subjected to the PISA program [43] for interface analysis at EBI-EMBL server and the retrieved PISA data was analyzed for binding patterns using PyMOL [44].
Molecular dynamics
Molecular dynamics techniques were applied using GROMACS suite [45] in order to evaluate the stability and consistency of the obtained N protein monomeric model and investigate GRSV N protein-RNA interactions over time. Therefore, N protein model was simulated in the presence and absence of the modeled RNA, in two analytical systems. Amber99SB-ILDN force field [46] was used to generate proper topologies. The models were placed at the center of a dodecahedral box and solvated with TIP3P water model [47]. Counterions were used to neutralize the net charge of the system, and 0.15 M of NaCl was added to the box in order to simulate cellular ionic environment.
After a minimization protocol using steepest descent and conjugate gradient to eliminate possible clashes and bad contacts, NVT ensemble with restraint forces of 1000 kJ/mol was carried for 4 ns at 300 K. Moreover, five subsequent equilibration steps in NPT ensemble were carried out at 1 bar with restraint forces of 800 kJ/mol on heavy atoms, 600 Kcal/(mol x nm) and 400 kJ/mol on mainchain, 200 kJ/mol on backbone and 100 kJ/mol on alpha-carbons, totalizing 13 ns. Finally, production runs with no restraints were carried for 50 ns using an integration step of 2 fs and LINCS algorithm [48]. Also, Particle Mesh Ewald method [49] was applied for Coulombic and Lennard-Jones interactions longer than 1 nm.
Acknowledgments
Declarations
This article has been published as part of BMC Bioinformatics Volume 17 Supplement 18, 2016. Proceedings of X-meeting 2015: 11th International Conference of the AB3C + Brazilian Symposium on Bioinformatics: bioinformatics. The full contents of the supplement are available online https://bmcbioinformatics.biomedcentral.com/articles/supplements/volume-17-supplement-18.
Funding
Publication of this paper has been funded by CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico), CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior), and FAPDF (Fundação de Apoio à Pesquisa do Distrito Federal), Brazil.
Availability of data and materials
The datasets supporting the conclusions of this article are included within the article and its supplementary files.
Authors’ contributions
Conceived and designed the experiments: JARGB MF RNL. Performed the homology modeling: MF RNL. Performed the Molecular Dynamics: HV MDP. Analyzed the data: FLM HV JARGB MDP MF RNL ROR. Wrote the paper: FLM HV JARGB MDP MF RNL ROR. All authors read and approved the final manuscript.
Competing interests
The authors declare that they have no competing interests.
Consent for publication
Not applicable.
Ethics approval and consent to participate
Not applicable.
Additional files
Ramachandran plot analysis of predicted structure of Groundnut ringspot virus (GRSV) N protein. The regions covered by light blue lines show most favored regions, while the regions covered by dark blue lines show allowed regions. Other regions of the plot show the disallowed region. The pink dots show the outliers (PNG 87 kb)
The Genbank acession numbers of the viruses used at this work (TABLEDOCX 19 kb)
References
- 1.Walter CT, Barr JN. Recent advances in the molecular and cellular biology of bunyaviruses. J Gen Virol. 2011;92(Pt 11):2467–2484. doi: 10.1099/vir.0.035105-0. [DOI] [PubMed] [Google Scholar]
- 2.de Avila AC, de Haan P, Kormelink R, Resende Rde O, Goldbach RW, Peters D. Classification of tospoviruses based on phylogeny of nucleoprotein gene sequences. J Gen Virol. 1993;74(Pt 2):153–159. doi: 10.1099/0022-1317-74-2-153. [DOI] [PubMed] [Google Scholar]
- 3.Pappu HR, Jones RA, Jain RK. Global status of tospovirus epidemics in diverse cropping systems: successes achieved and challenges ahead. Virus Res. 2009;141(2):219–236. doi: 10.1016/j.virusres.2009.01.009. [DOI] [PubMed] [Google Scholar]
- 4.Richmond KE, Chenault K, Sherwood JL, German TL. Characterization of the nucleic acid binding properties of tomato spotted wilt virus nucleocapsid protein. Virology. 1998;248(1):6–11. doi: 10.1006/viro.1998.9223. [DOI] [PubMed] [Google Scholar]
- 5.Ribeiro D, Borst JW, Goldbach R, Kormelink R. Tomato spotted wilt virus nucleocapsid protein interacts with both viral glycoproteins Gn and Gc in planta. Virology. 2009;383(1):121–130. doi: 10.1016/j.virol.2008.09.028. [DOI] [PubMed] [Google Scholar]
- 6.Soellick T, Uhrig JF, Bucher GL, Kellmann JW, Schreier PH. The movement protein NSm of tomato spotted wilt tospovirus (TSWV): RNA binding, interaction with the TSWV N protein, and identification of interacting plant proteins. Proc Natl Acad Sci U S A. 2000;97(5):2373–2378. doi: 10.1073/pnas.030548397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Feng Z, Chen X, Bao Y, Dong J, Zhang Z, Tao X: Nucleocapsid of Tomato spotted wilt tospovirus forms mobile particles that traffic on an actin/endoplasmic reticulum network driven by myosin XI-K. New Phytol. 2013;200(4):1212-24. [DOI] [PubMed]
- 8.Ariza A, Tanner SJ, Walter CT, Dent KC, Shepherd DA, Wu W, et al. Nucleocapsid protein structures from orthobunyaviruses reveal insight into ribonucleoprotein architecture and RNA polymerization. Nucleic Acids Res. 2013;41(11):5912–5926. doi: 10.1093/nar/gkt268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Guu TS, Zheng W, Tao YJ. Bunyavirus: structure and replication. Adv Exp Med Biol. 2012;726:245–266. doi: 10.1007/978-1-4614-0980-9_11. [DOI] [PubMed] [Google Scholar]
- 10.de Oliveira AS, Melo FL, Inoue-Nagata AK, Nagata T, Kitajima EW, Resende RO. Characterization of bean necrotic mosaic virus: a member of a novel evolutionary lineage within the genus tospovirus. PLoS One. 2012;7(6):e38634. doi: 10.1371/journal.pone.0038634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Snippe M, Willem Borst J, Goldbach R, Kormelink R. Tomato spotted wilt virus Gc and N proteins interact in vivo. Virology. 2007;357(2):115–123. doi: 10.1016/j.virol.2006.06.037. [DOI] [PubMed] [Google Scholar]
- 12.Mir MA, Panganiban AT. The bunyavirus nucleocapsid protein is an RNA chaperone: possible roles in viral RNA panhandle formation and genome replication. RNA (New York, NY) 2006;12(2):272–282. doi: 10.1261/rna.2101906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Mir MA, Panganiban AT. The hantavirus nucleocapsid protein recognizes specific features of the viral RNA panhandle and is altered in conformation upon RNA binding. J Virol. 2005;79(3):1824–1835. doi: 10.1128/JVI.79.3.1824-1835.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Brennan B, Welch SR, Elliott RM. The consequences of reconfiguring the ambisense S genome segment of rift valley fever virus on viral replication in mammalian and mosquito cells and for genome packaging. PLoS Pathog. 2014;10(2):e1003922. doi: 10.1371/journal.ppat.1003922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Li W, Lewandowski DJ, Hilf ME, Adkins S. Identification of domains of the tomato spotted wilt virus NSm protein involved in tubule formation, movement and symptomatology. Virology. 2009;390(1):110–121. doi: 10.1016/j.virol.2009.04.027. [DOI] [PubMed] [Google Scholar]
- 16.Singh P, Indi SS, Savithri HS. Groundnut bud necrosis virus encoded NSm associates with membranes via its C-terminal domain. PLoS One. 2014;9(6):e99370. doi: 10.1371/journal.pone.0099370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kainz M, Hilson P, Sweeney L, Derose E, German TL. Interaction between tomato spotted wilt virus N protein monomers involves nonelectrostatic forces governed by multiple distinct regions in the primary structure. Phytopathology. 2004;94(7):759–765. doi: 10.1094/PHYTO.2004.94.7.759. [DOI] [PubMed] [Google Scholar]
- 18.Uhrig JF, Soellick TR, Minke CJ, Philipp C, Kellmann JW, Schreier PH. Homotypic interaction and multimerization of nucleocapsid protein of tomato spotted wilt tospovirus: identification and characterization of two interacting domains. Proc Natl Acad Sci U S A. 1999;96(1):55–60. doi: 10.1073/pnas.96.1.55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zheng W, Olson J, Vakharia V, Tao YJ. The crystal structure and RNA-binding of an orthomyxovirus nucleoprotein. PLoS Pathog. 2013;9(9):e1003624. doi: 10.1371/journal.ppat.1003624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Reguera J, Malet H, Weber F, Cusack S. Structural basis for encapsidation of genomic RNA by La Crosse orthobunyavirus nucleoprotein. Proc Natl Acad Sci U S A. 2013;110(18):7246–7251. doi: 10.1073/pnas.1302298110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Dong H, Li P, Elliott RM, Dong C. Structure of schmallenberg orthobunyavirus nucleoprotein suggests a novel mechanism of genome encapsidation. J Virol. 2013;87(10):5593–5601. doi: 10.1128/JVI.00223-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Niu F, Shaw N, Wang YE, Jiao L, Ding W, Li X, et al. Structure of the leanyer orthobunyavirus nucleoprotein-RNA complex reveals unique architecture for RNA encapsidation. Proc Natl Acad Sci U S A. 2013;110(22):9054–9059. doi: 10.1073/pnas.1300035110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Raymond DD, Piper ME, Gerrard SR, Skiniotis G, Smith JL. Phleboviruses encapsidate their genomes by sequestering RNA bases. Proc Natl Acad Sci U S A. 2012;109(47):19208–19213. doi: 10.1073/pnas.1213553109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Carter SD, Surtees R, Walter CT, Ariza A, Bergeron E, Nichol ST, et al. Structure, function, and evolution of the Crimean-Congo hemorrhagic fever virus nucleocapsid protein. J Virol. 2012;86(20):10914–10923. doi: 10.1128/JVI.01555-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ferron F, Li Z, Danek EI, Luo D, Wong Y, Coutard B, et al. The hexamer structure of rift valley fever virus nucleoprotein suggests a mechanism for its assembly into ribonucleoprotein complexes. PLoS Pathog. 2011;7(5):e1002030. doi: 10.1371/journal.ppat.1002030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Brunotte L, Kerber R, Shang W, Hauer F, Hass M, Gabriel M, et al. Structure of the Lassa virus nucleoprotein revealed by X-ray crystallography, small-angle X-ray scattering, and electron microscopy. J Biol Chem. 2011;286(44):38748–38756. doi: 10.1074/jbc.M111.278838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Reguera J, Cusack S, Kolakofsky D. Segmented negative strand RNA virus nucleoprotein structure. Curr Opin Virol. 2014;5:7–15. doi: 10.1016/j.coviro.2014.01.003. [DOI] [PubMed] [Google Scholar]
- 28.Tripathi D, Raikhy G, Pappu HR. Movement and nucleocapsid proteins coded by two tospovirus species interact through multiple binding regions in mixed infections. Virology. 2015;478:137-47. [DOI] [PubMed]
- 29.Leastro MO, Pallas V, Resende RO, Sanchez-Navarro JA. The movement proteins (NSm) of distinct tospoviruses peripherally associate with cellular membranes and interact with homologous and heterologous NSm and nucleocapsid proteins. Virology. 2015;478c:39–49. doi: 10.1016/j.virol.2015.01.031. [DOI] [PubMed] [Google Scholar]
- 30.Kairys V, Gilson MK, Fernandes MX. Using protein homology models for structure-based studies: approaches to model refinement. TheScientificWorldJOURNAL. 2006;6:1542–1554. doi: 10.1100/tsw.2006.250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sellers BD, Nilmeier JP, Jacobson MP. Antibodies as a model system for comparative model refinement. Proteins. 2010;78(11):2490–2505. doi: 10.1002/prot.22757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Dong H, Li P, Bottcher B, Elliott RM, Dong C. Crystal structure of schmallenberg orthobunyavirus nucleoprotein-RNA complex reveals a novel RNA sequestration mechanism. RNA (New York, NY) 2013;19(8):1129–1136. doi: 10.1261/rna.039057.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Li J, Feng Z, Wu J, Huang Y, Lu G, Zhu M, et al. Structure and function analysis of nucleocapsid protein of tomato spotted wilt virus interacting with RNA using homology modeling. J Biol Chem. 2015;290(7):3950–3961. doi: 10.1074/jbc.M114.604678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Biasini M, Bienert S, Waterhouse A, Arnold K, Studer G, Schmidt T, et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 2014;42(Web Server issue):W252–W258. doi: 10.1093/nar/gku340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Notredame C, Higgins DG, Heringa J. T-coffee: a novel method for fast and accurate multiple sequence alignment. J Mol Biol. 2000;302(1):205–217. doi: 10.1006/jmbi.2000.4042. [DOI] [PubMed] [Google Scholar]
- 36.Hall T. BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp Ser. 1999;41:95–98. [Google Scholar]
- 37.Sali A, Blundell TL. Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol. 1993;234(3):779–815. doi: 10.1006/jmbi.1993.1626. [DOI] [PubMed] [Google Scholar]
- 38.Krieger E, Joo K, Lee J, Lee J, Raman S, Thompson J, et al. Improving physical realism, stereochemistry, and side-chain accuracy in homology modeling: four approaches that performed well in CASP8. Proteins. 2009;77(Suppl 9):114–122. doi: 10.1002/prot.22570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Ramachandran S, Kota P, Ding F, Dokholyan NV. Automated minimization of steric clashes in protein structures. Proteins. 2011;79(1):261–270. doi: 10.1002/prot.22879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Colovos C, Yeates TO. Verification of protein structures: patterns of nonbonded atomic interactions. Protein sci pub protein soc. 1993;2(9):1511–1519. doi: 10.1002/pro.5560020916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Chen VB, Arendall WB, 3rd, Headd JJ, Keedy DA, Immormino RM, Kapral GJ, et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr D Biol Crystallogr. 2010;66(Pt 1):12–21. doi: 10.1107/S0907444909042073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Emsley P, Lohkamp B, Scott WG, Cowtan K. Features and development of coot. Acta Crystallogr D Biol Crystallogr. 2010;66(Pt 4):486–501. doi: 10.1107/S0907444910007493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Krissinel E, Henrick K. Inference of macromolecular assemblies from crystalline state. J Mol Biol. 2007;372(3):774–797. doi: 10.1016/j.jmb.2007.05.022. [DOI] [PubMed] [Google Scholar]
- 44.Delano W. The PyMOL molecular graphics system. San Carlos: DeLano Scientific; 2002. [Google Scholar]
- 45.Abraham MJ, Murtola T, Schulz R, Páll S, Smith JC, Hess B, et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX. 2015;1–2:19–25. doi: 10.1016/j.softx.2015.06.001. [DOI] [Google Scholar]
- 46.Lindorff-Larsen K, Piana S, Palmo K, Maragakis P, Klepeis JL, Dror RO, et al. Improved side-chain torsion potentials for the amber ff99SB protein force field. Proteins. 2010;78(8):1950–1958. doi: 10.1002/prot.22711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Jorgensen WL, Madura JD. Quantum and statistical mechanical studies of liquids. 25. Solvation and conformation of methanol in water. J Am Chem Soc. 1983;105(6):1407–1413. doi: 10.1021/ja00344a001. [DOI] [Google Scholar]
- 48.Hess B, Bekker H, Berendsen HJC, Fraaije JGEM. LINCS: a linear constraint solver for molecular simulations. J Comput Chem. 1997;18(12):1463–1472. doi: 10.1002/(SICI)1096-987X(199709)18:12<1463::AID-JCC4>3.0.CO;2-H. [DOI] [Google Scholar]
- 49.Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG. A smooth particle mesh ewald method. J Chem Phys. 1995;103(19):8577–8593. doi: 10.1063/1.470117. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets supporting the conclusions of this article are included within the article and its supplementary files.


