Abstract
The basic helix-loop-helix (bHLH) factor Hand1 plays an essential role in cardiac morphogenesis, and yet its precise function remains unknown. Protein-protein interactions involving Hand1 provide a means of determining how Hand1-induced gene expression in the developing heart might be regulated. Hand1 is known to form either heterodimers with near-ubiquitous E-factors and other lineage-restricted class B bHLH proteins or homodimers with itself in vitro. To date, there have been no reported Hand1 protein interactions involving non-bHLH proteins. Heterodimer-versus-homodimer choice is mediated by the phosphorylation status of Hand1; however, little is known about the in vivo function of these dimers or, importantly, how they are regulated. In an effort to understand how Hand1 activity in the heart might be regulated postdimerization, we have investigated tertiary Hand1-protein interactions with non-bHLH factors. We describe a novel interaction of Hand1 with the LIM domain protein FHL2, a known transcriptional coactivator and corepressor expressed in the developing cardiovascular system. FHL2 interacts with Hand1 via the bHLH domain and is able to repress Hand1/E12 heterodimer-induced transcription but has no effect on Hand1/Hand1 homodimer activity. This effect of FHL2 is not mediated either at the level of dimerization or via an effect of Hand1/E12 DNA binding. In summary, our data describe a novel differential regulation of Hand1 heterodimers versus homodimers by association of the cofactor FHL2 and provide insight into the potential for a tertiary level of control of Hand1 activity in the developing heart.
The heart is the first organ to develop in the embryo proper, and its importance is underlined by the multitude of lethal phenotypes in animal models carrying null mutations in genes critical for cardiovascular development. Efforts to understand the molecular mechanisms underlying vertebrate heart development have recently shifted focus from the definition of cardiac-specific master regulatory factors to multiprotein complexes that coordinate programs of heart-specific gene expression.
Hand1 (eHAND, Hxt, Thing1) encodes a basic helix-loop-helix (bHLH) transcription factor that our investigators have previously shown plays an essential role in cardiac morphogenesis during murine embryonic development (44). As a member of the class B (lineage-restricted) bHLH transcription factors, Hand1 has been shown to heterodimerize with a class A (near-ubiquitous) E-factor, such as that encoded by E2A (E12/E47), and activate transcription of downstream target genes from “E-box” sequences (CANNTG [23]) or more specifically, in the case of Hand1, a degenerate “Thing1” box (CGTCTG [21]). Besides interacting with the more-ubiquitous E-factors, Hand1 has also been shown to heterodimerize with other lineage-restricted bHLH proteins. Hand1 is capable of forming efficient heterodimers with its most closely related bHLH factor, Hand2 (14), and interaction of both Hand1 and Hand2 has been demonstrated with the recently described Hey/Hrt family of bHLH proteins 1 to 3 (14). Furthermore, Hand1 may also form functional homodimers, in differentiated lineages, since Hand1/Hand1 dimers have been detected by immunoprecipitation and mammalian two-hybrid studies (46). That said, the DNA binding affinity of Hand1 homodimers is an order of magnitude lower than that of Hand1/E-factor heterodimers (46), suggesting that, for Hand1 homodimer formation to be functionally significant in vivo, such homodimers may need to be associated with other non-bHLH proteins in a higher-order complex to sufficiently alter their DNA binding affinity and transcriptional activity.
Having initially been classified as an activator of transcription (21), there is now substantial evidence that Hand1 can also function as a transcriptional repressor either via sequestration of class A bHLH factors from E-boxes (25) or through direct inhibition of transcriptional activity (14). Hand1 is able to antagonize the function of other bHLH proteins, such as Mash2, by competing for E-factor binding in a manner similar to MISTI and Id proteins (46) and can also negatively regulate MyoD-E47-tethered dimers (3), which are resistant to other proteins (such as the Ids) that act by binding and competing for E-proteins in the cell. Both Hand1 and Hand2 have been shown to operate independently of direct DNA binding (35, 52), adding another layer of complexity to their function. Most notably, misexpression of either Hand protein in the developing limb bud in transgenic mice induces ectopic digits, and this activity is dependent exclusively on the HLH region, excluding a previously identified transcriptional activation domain or the basic DNA binding region (35).
The broad dimerization profile of Hand1, along with a potential requirement for tertiary interaction with other proteins, reflects the complex mechanisms by which Hand1 regulates transcription. Recently, dimerization of the Hand proteins was shown to be modulated by site-specific phosphorylation (16) as an important step towards understanding how posttranslational modification may regulate Hand activity in a tissue-specific manner. However, little is known about the tertiary interaction of proteins following Hand dimerization, or how such interactions may further regulate the Hand proteins activity in vivo. The identity of Hand1 cofactors and the functional significance of any identified interaction are of interest, therefore, in accounting for novel mechanisms of action of Hand1 in the developing vertebrate heart.
In this study we screened for candidate cofactors of Hand1 homo- or heterodimers by testing for interaction with LIM domain-containing proteins, since LIM domains mediate protein dimerization, are present in oligomeric complexes in differentiated cell types (45), and are known to interact with other protein motifs, such as PDZ domains (8) and ankyrin repeats (53) as well as helix-loop-helix domains (26, 30). Precedent for a Hand1-LIM-containing protein interaction exists in the erythroid DNA binding complex containing the bHLH factor SCL/TAL, which shares significant amino acid homology with Hand1 (2) and interacts with the LIM-only protein LMO2 via its HLH domain (30, 54). Moreover, a considerable number of LIM domain-containing proteins are expressed in the developing heart (1, 18, 28) and of these, a legitimate candidate for interaction with Hand1 is FHL2, a member of the four-and-a-half LIM domain-containing family collectively expressed in skeletal muscle and the developing cardiovascular system (7). Fhl2 is restricted to the developing heart, where it is coexpressed with Hand1 between embryonic day 8.0 (E8.0) and E9.5 (7; this study). Functionally, FHL2 has been shown to play a role in modifying the response to hypertrophic stimuli in the heart (27), a putative function previously attributed to Hand1 (39, 52), and in the case of FHL2 this may be mediated by uncoupling of ERK1/2 from transcriptional pathways regulating cardiac differentiated growth (42). Moreover, like Hand1, FHL2 has been shown to act both as a transcriptional coactivator of the androgen receptor (38), AP-1 (37), WT1 (11), β-catenin (32, 56), and BRAC1 (58) and as a corepressor for the PLZF protein (36).
Here, we present evidence for a direct physical association between Hand1 and FHL2, as a bona fide, non-bHLH interaction with a member of the Hand family of class B bHLH proteins. We demonstrate a novel differential regulation of Hand1 homodimer- and Hand1/E12 heterodimer-induced transcriptional activity via the proposed interaction with FHL2 and determine that FHL2 can function as a repressor of Hand1 biological activity in the developing heart.
MATERIALS AND METHODS
Plasmids.
The full-length murine Hand1 plasmids, pcDNA3Hand1 and pFLAGHand1, have been described previously (46). The full-length human E-factor E12 plasmid pHBAPneo-E12 was obtained from Cornelius Murre, University of California San Diego. pcDNA3-E12 was constructed by PCR amplification of the human E12 cDNA from pHBAPneo-E12, subcloned into EcoRI-XhoI sites of pcDNA3. Tethered constructs pHand1∼Hand1 and pHand1∼ITF2 were obtained from Jay Cross, University of Calgary, and have been previously described (46).
A mock Hand1 target gene reporter plasmid was constructed by subcloning annealed sense and antisense oligonucleotides corresponding to a chick α-cardiac actin (α-CA) minimal promoter (−58 to +15) (6, 31) into the SacI-XhoI sites of pGL2-Basic, upstream of a luciferase reporter gene. Subsequently, annealed oligonucleotides corresponding to X6 concatemerized optimal Hand1 binding sites (Thing1 boxes) plus flanking nucleotides (21) were subcloned upstream of the α-CA minimal promoter into the KpnI-SacI sites of pGL2-Basic.
A full-length FLAG-tagged FHL2 cDNA (pFLAG-CMV-FHL2) was obtained from Ju Chen, UCSD (7). Full-length FHL2 was subcloned into pKS+, creating pKS-FHL2, for use as an in situ probe. Full-length FHL2 containing MYC N-terminal epitope tags, pMYC-FHL2, was constructed by PCR amplification of five MYC tags subcloned at the HindIII-BamHI sites of pcDNA3 to produce pcDNA3MYC. Subsequently, full-length FHL2 minus the ATG start codon was PCR amplified and subcloned at the EcoRI-XhoI sites of pcDNA3MYC downstream and in frame with the five MYC tags. To construct the FHL2 deletion mutants pFHL1/2-2, pFHL2-3, pFHL3-4, pFHL1/2-1, pFHL2, pFHL3, and pFHL4, the corresponding fragments (LIM1/2-2, amino acids 1 to 153; LIM2-3, amino acids 101 to 212; LIM3-4, amino acids 162 to 279; LIM1/2-1, amino acids 1 to 92; LIM2, amino acids 101 to 153; LIM3, amino acids 162 to 212; and LIM4, amino acids 221 to 279) were amplified by PCR and inserted at the BamHI-EcoRI sites of pcDNA3.
A GAL4 reporter plasmid was constructed as follows: five 17-mer GAL4 upstream activating sequence binding sites concatemerized were placed upstream of a minimal cytomegalovirus (CMV) promoter minus enhancer and subcloned into the SmaI-XhoI sites of pGL3-Basic to produce pGL3-CMVmin (CMVmin(Gal4)5-luciferase). To produce pGAL4DBDHand1, the GAL4 DNA binding domain from pGBDU was subcloned into the HindIII site of pcDNA3 containing an in-frame full-length Hand1 cDNA. The pMVP16 plasmid containing a fusion of the GAL4 DNA binding domain and VP16 activation domain is available commercially (Clontech).
Glutathione S-transferase (GST) fusion plasmids were constructed as follows: pGSTHand1 was constructed by taking an EcoRI-XhoI fragment containing the full-length Hand1 cDNA and cloning in frame at the EcoRI-XhoI sites of pGEX4T-1 (Promega). pGST-bHLH was constructed by PCR amplification of the bHLH region of Hand1 (amino acids 93 to 147) subcloned in frame at the EcoRI-XhoI sites of pGEX4T-1. pGST-FHL2 was constructed by PCR amplification of full-length FHL2 cDNA subcloned in frame at the EcoRI-XhoI sites of pGEX4T-1.
Fluorescent fusion proteins were constructed for fluorescence resonance energy transfer (FRET) microscopy as follows. The Hand1 and FHL2 coding regions (including Kozak sequence) minus stop codons were independently amplified by PCR and, in the case of Hand1, subcloned upstream of enhanced yellow fluorescent protein as N-terminal fusion at the EcoRI-KpnI sites in pEYFP-N1 and, in the case of FHL2, subcloned upstream of enhanced cyan fluorescent protein at the EcoR1-BamHI sites of pECFP-N1 (Clontech), to produce pHand1-EYFP and pFHL2-ECFP, respectively.
All plasmids were verified by sequencing, and their relative expression in cells, prior to the mammalian two-hybrid and reporter assays, were determined by Western blot analyses.
Whole-mount in situ hybridization.
Whole-mount in situ hybrdization was performed as previously described (48, 57), with minor modifications. A 0.5% 3-[(3-choladmiopropyl)-dimethylammonio]-1-propanesulfonate detergent was added to the hybridization buffer, and hybridization was carried out at 68°C using a probe for FHL2 (full-length FHL2; pKS-FHL2). Blocking buffer additionally contained 1% bovine serum albumin (BSA) and 10% sheep serum. Antisense RNA probes were labeled with digoxigenin. Nitro Blue Tetrazolium and 5-bromo-4-chloro-3-indolyl-phosphate (NBT/BCIP tablets; Amersham Pharmacia) were used as color reagents for probe detection. To stop the NBT/BCIP reaction, embryos were washed in phosphate-buffered saline (PBS) with 0.1% Triton X-100 (PBT) and fixed overnight (4% paraformaldehyde in PBS at 4°C). To reduce background, embryos were cleared in 100% methanol for 10 min on a Nutator and rehydrated through a methanol-PBS gradient. Finally, embryos were stored at 4°C in PBS containing 1% azide prior to photography or histological sectioning.
Whole-mount embryo sectioning and staining.
Following in situ hybridization, embryos were dehydrated in a series of 30-min ethanol-saline washes and equilibrated with Histoclear and then with molten paraffin wax as previously described (48). Embryos were orientated in molds and allowed to set, and sagittal sections of 10-μm thickness were cut on a microtome. Sections were dewaxed in Histoclear (two 5-min incubations) and washed in serial ethanol dilutions, as previously described (48). After two 5-min washes in distilled water, sections were counterstained with 0.5% eosin prior to mounting and visualization on an Olympus SZ4045TR microscope.
Cell lines, transfections, and derivation of cardiomyocytes.
NIH 3T3 and H9c2 cells were cultured in Dulbecco's modified Eagle's medium supplemented with 10% bovine calf serum (for NIH 3T3) or 10% fetal bovine serum (for H9c2 cells). P19 cells were cultured in Dulbecco's modified Eagle's medium supplemented with 7.5% bovine calf serum and 2.5% fetal bovine serum plus 0.1 M β-mercaptoethanol. Transient-transfection assays were carried out on NIH 3T3 and H9c2 cells in six-well plates. Cells were transfected at between 40 to 60% confluence using the Effectene method as per the manufacturer's instructions (QIAGEN). The total amount of transfected DNA per well was kept constant (between 1 and 2 μg) by adding the corresponding amounts of empty expression plasmid or pcDNA3. A total of 200 ng of reporter plasmid, 200 ng of each expression plasmid, or 125 ng of each Gal-fusion plasmid was transfected into each well. A 50-ng aliquot of a β-actin-β-galactodsidase-expressing plasmid was cotransfected to normalize luciferase activity for transfection efficiency. Luciferase activity was assayed 48 h posttransfection using a Promega luciferase assay system according to the manufacturer's instructions (Promega). β-Galactosidease activity was assessed following incubation of transfected cell lysate with assay buffer (120 mM Na2HPO4, 80 mM NaH2PO4, 2 mM MgCl2, 0.1 M β-mercaptoenthanol, 1.5 mg of o-nitrophenyl-β-d-galactopyranoside/ml) at 37°C for between 30 min and 3 h. The reaction was stopped with the addition of 1 M Na2CO3, and the degree of β-galactosidase precipitate was assessed by reading the A420. Wells were transfected in duplicate, and each experiment was repeated at least five times.
To derive cardiomyocytes for endogenous immunoprecipitation experiments, P19 cells were aggregated in the presence of dimethyl sulfoxide (DMSO) and induced to differentiate over a period of 8 days as previously described (33, 34). Briefly, pluripotent P19 cells were grown to confluence and resuspended in bacteriological-grade dishes at 105 cells/ml of medium supplemented with 0.8% (vol/vol) DMSO. After 2 days of incubation medium was replenished with fresh DMSO-containing medium, and after 4 days in suspension aggregates were plated onto tissue-grade surfaces in medium lacking DMSO. By 5 to 6 days after the initiation of aggregate formation, rhythmically contracting cardiac muscle cells were observed in the cultures which express the appropriate transcription factors and exhibit equivalent biochemical and physiological characteristics to that of embryological cardiomyocytes (47).
GST pull-down assay.
GST and GST-fusion proteins were expressed and purified according to the manufacturer's protocol (Amersham Pharmacia), with the induction of protein expression performed at 25°C overnight. The expression vectors for Hand1, FHL2, and derivatives were used for in vitro transcription and translation using a TNT kit (Promega) in the presence of [35S]methionine. The 35S-labeled proteins were incubated with either 0.5 or 2 μg of GST derivatives bound to glutathione-Sepharose beads in 150 ml of binding buffer (20 mM Tris-HCl [pH 8.0], 100 mM NaCl, 1 mM EDTA, 0.5% NP-40, and protease inhibitors from Roche). The binding reaction was performed at 4°C overnight, and the beads were subsequently washed four times with washing buffer (same as the binding buffer). The beads were eluted in 20 μl of 2× Lamelli protein loading buffer and boiled for 3 to 5 min, and protein interactions were analyzed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) followed by autoradiography.
Antibodies, coimmunoprecipitation, and Western blot analyses.
A polyclonal anti-Hand1 antibody (α-Hand1; C-terminal epitope α-eHAND) was purchased from Santa Cruz. Polyclonal and monoclonal α-FHL2 antibodies were a kind gift from R. Schule, Freiburg, Germany (38). α-Epitope tag antibodies α-FLAG and α-MYC were either purchased from Sigma (anti-FLAG M2 affinity gel) or were a kind gift from S. Parkinson, Imperial Cancer Research Fund, London, England (α-MYC).
For the overexpression coimmunoprecipitation experiments, NIH 3T3 cells were transfected in 10-cm dishes as described above. For CO-IP of endogenous Hand1 and FHL2, lysates from P19-derived cardiomyocytes at day 8 of differentiation were prepared as described above. For in vitro detection of the interaction of Hand1 with FHL2, NIH 3T3 cells were transfected with pFLAGHand1 and pMYCFHL2; for determining whether FHL2 can disrupt either Hand1 homodimerization or Hand1 E-factor heterodimerization, NIH 3T3 cells were transfected with either pcDNA3Hand1 alone or pcDNA3Hand1 in combination with pcDNA3-E12 in the presence or absence of FHL2. Forty-eight hours posttransfection, cells were washed twice with PBS. Transfected cells and P19-derived cardiomyocytes were lysed in 0.8 ml of lysis buffer (50 mM Tris-HCl [pH 7.4], 150 mM NaCl, 1 mM EDTA, 1% Triton X-100, 1 mM dithiothreitol [DTT], 0.5 mM phenylmethylsulfonyl fluoride [PMSF], and protease inhibitors from Roche). After brief shearing with a 21-gauge needle, the lysate was centrifuged at 12,000 × g for 5 min. Protein A-G-Sepharose beads (Amersham Pharmacia) were preblocked with PBS containing 1% BSA for 30 min at room temperature. One hundred microliters of preblocked beads and either 4 μg of α-FLAG M2 antibody (overexpression) or 3 μg of FHL2 monoclonal antibody (endogenous) was used for each immunoprecipitation. Immunoprecipitation was performed overnight at 4°C. The beads were centrifuged at 4,000 rpm for 1 min and washed four times in ice-cold PBS. The precipitates were resuspended in 20 μl of 2× SDS-PAGE loading buffer and loaded on an SDS-12% PAGE gel, followed by Western blotting according to standard procedures. Five percent of the input crude extract was used for determining protein expression levels. The MYC-tagged FHL2 was detected using a 1:500 dilution of the α-MYC polyclonal antibody, and endogenous Hand1 was detected using a 1:300 dilution of the α-Hand1 polyclonal antibody described above.
Immunofluorescence and FRET analysis.
For endogenous Hand1 and FHL2 staining, H9c2 cells were cultured on 0.1% gelatine-coated coverslips until approximately 60% confluent. Cells were washed twice with PBS, fixed with either 100% methanol, 2% paraformaldehyde-PBS, or 4% paraformaldehyde-PBS for 10 min on ice, permeabilized with 0.5% Triton X-100-PBS for 5 min, and blocked in 1% BSA-PBS for 30 min. Staining with the polyclonal α-Hand1 (diluted 1:100 in 1% BSA-PBS) and the polyclonal α-FHL2 antibody (diluted 1:500 in 1% BSA-PBS) was carried out overnight at 4°C. Cells were then washed three times with PBS. Secondary antibodies fluorescein isothiocyanate-labeled α-goat and tetramethyl rhodamine isocyanate-labeled α-rabbit immunoglobulin G (Dako) were used at dilutions of 1:20 and 1:30, respectively, in 1% BSA-PBS. Nuclei were stained with 5 μg of bis-benzamide (Hoechst 33342; Sigma)/ml in PBS for 10 min at room temperature. Cells were mounted with coverslips secured in 50% glycerol-PBS, and fluorescence was visualized using a Zeiss Axioskop 2 microscope with 20× and 40× Zeiss Plan-Neofluar objectives.
To visualize an interaction between Hand1 and FHL2 in living cells, NIH 3T3 cells were cotransfected with pHand1-EYFP (donor) and pFHL2-ECFP (acceptor) on coverslips as described above. Nuclei were counterstained with bis-benzamide as previously described. The fluorescence imaging workstation was a Leica multiphoton CLS microscope.
Regions of colocalization of the donor pHand1-EYFP and acceptor pFHL2-ECFP in individual cotransfected cells were obtained using the yellow fluorescent protein (YFP) (excitation, 500/20 nm; emission, 535/30 nm) and cyan fluorescent protein (CFP) (excitation, 436/10 nm; emission, 470/30 nm) filter sets, respectively. FRET was carried out by acceptor bleaching: control images were acquired of the acceptor by using the CFP filter set, and the acceptor was then bleached. A time course of fluorescence measurements was taken using a FRET filter set (excitation, 500/20 nm; emission, 470/30 nm) in selected regions of colocalization and regions outside where localization of donor and acceptor was mutually exclusive. Measurements were taken from a minimum of 10 individual cotransfected cells. FRET was recorded if in a region of colocalization within the cell excitation of YFP resulted in emission from CFP immediately following bleaching of the CFP-fused acceptor. Efficient energy transfer implies that the YFP and CFP must lie within a 10- by 10- by 10-nm volume of one another (i.e., less than 50 Å apart or typical Forster distances of 2 to 7 nm), which requires direct interaction of the proteins of interest fused to these fluorescent moieties.
EMSAs.
Electrophoretic mobility shift assays (EMSAs) were carried out with either in vitro-translated (IVT) proteins or transfected cell protein extracts. IVT proteins were synthesized using the TNT kit (Promega). NIH 3T3 cells were transfected as described above. Whole-cell extracts were prepared by harvesting cotransfected cells in lysis buffer (20 mM HEPES [pH 7.9], 450 mM NaCl, 25% glycerol, 0.5 mM DTT, 0.25 mM EDTA, and protease inhibitors from Roche) and carrying out three freeze-thaw cycles: snap freezing in liquid nitrogen followed by rapid thawing at 30°C. Lysates were centrifuged at 13,000 rpm, and the supernatant was stored at −80°C for up to three reactions.
IVT proteins were mixed and preincubated for 30 min at 37°C to facilitate interaction. Typically 3 to 5 μl of IVT protein or lysate was used per binding reaction mixture, ensuring the protein concentration was at least 10-fold higher than the DNA concentration. Annealed 26-bp oligonucleotides 5′-CAACCACAATGGCGTCGTCTGGCATTTTT-3′ and 5′-TCGAAAAAATGCCAGACGCCATTGTG-3′, containing an optimal Hand1/E12 binding site (shown in bold) (Thing1 box, CGTCTG) (21), and flanking nucleotides were 5′-end labeled. In a typical binding reaction the protein extract was mixed with 40,000 cpm (approximately 40 fmol) of labeled oligo in 20 μl of binding buffer (50 mM HEPES [pH 7.9], 200 mM NaCl, 30% glycerol, 4 mM MgCl2, 1 mM EGTA, 1 mM DTT, 100 μg of BSA/ml) with 1 mg of poly(dI-dC)/ml. The binding reaction mixture was incubated at room temperature for 20 min in the absence of labeled oligo to ensure subunit exchange, and the complete reaction mixture including labeled oligo with or without 100-fold cold oligo was then incubated for a further 20 min at room temperature. Two microliters of loading buffer (50 mM Tris-HCl [pH 7.6], 20% glycerol, 0.25% bromophenol blue) was added, and each sample was run through a 6% polyacrylamide gel. The gel was then dried and subjected to autoradiography.
RESULTS
Hand1 is coexpressed with FHL2 in the developing heart.
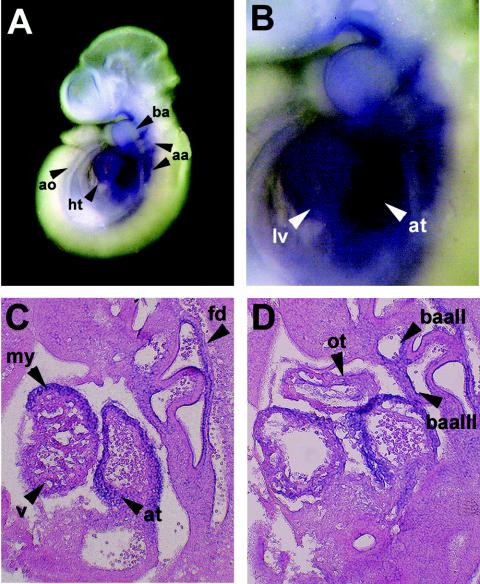
Based on its reported expression pattern (5, 7) and putative ability to act as a dominant repressor in the myocardium (42), FHL2 was deemed a potential candidate LIM protein interactor with Hand1 in the developing murine heart. We therefore set out to confirm that FHL2 was coexpressed with Hand1 in the heart by whole-mount in situ hybridization followed by sectioning of wild-type embryos at E9.5. As shown in Fig. 1, FHL2 was expressed throughout the heart at E9.5 (Fig. 1A), and expression was strong in both the presumptive atrial and ventricular chambers (Fig. 1B to D) with weaker expression evident in the developing outflow tract (Fig. 1D). We and others have previously shown that Hand1 is similarly expressed at high levels in the presumptive ventricle, most notably in the outer curvature of the ventricular myocardium as well as in the proximal outflow tract (44, 48). Both FHL2 (Fig. 1C and D) and Hand1 (48) are specifically localized to cardiomyocytes which make up the compact layer and trabeculae of the ventricular myocardium. FHL2 is also expressed in the first branchial arch (Fig. 1B), as is Hand1 (48), and in mutually exclusive regions such as the dorsal aorta (Fig. 1A) second and third branchial arch arteries (Fig. 1C) and in cells lining the pharyngeal region of the foregut diverticulum (Fig. 1D).
FIG. 1.
FHL2 is coexpressed with Hand1 in the developing heart. (A and B) FHL2 mRNA expression in E9.5 wild-type embryos as detected by whole-mount in situ hybridization. (C and D) Sagittal sections through E9.5 wild-type hearts following whole-mount in situ hybridization for FHL2 and counterstaining with eosin. Abbreviations: aa, arch arteries; ao, dorsal aorta; at, atrium; ba, branchial arch; baII, second branchial arch artery; baIII, third branchial arch artery; fd, foregut diverticulum; lv, left ventricle; my, myocardium; ot, outflow tract; v, ventricle.
Hand1 interacts specifically with FHL2 via the bHLH domain.
Since Hand1 appears to play an essential role in cardiac morphogenesis, we set out to test for interaction of Hand1 with FHL2 as a first step towards understanding how Hand1 protein interactions might regulate cardiac muscle gene expression.
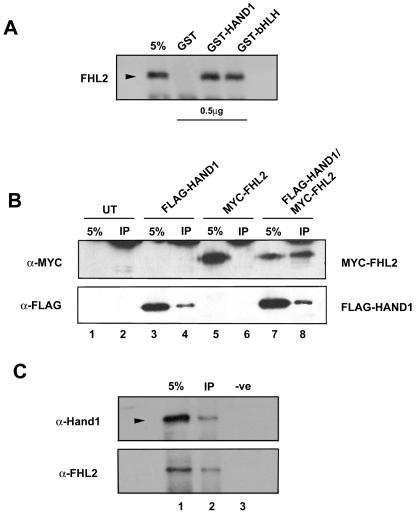
IVT FHL2 was incubated with equal amounts of either GST-Hand1, GST-bHLH (containing the isolated bHLH domain of Hand1), or GST alone. GST-Hand1 and GST-bHLH, but not GST alone, interacted with FHL2 (Fig. 2A), suggesting Hand1 and FHL2 can physically associate in vitro and that the interaction involves the bHLH domain of Hand1. To determine whether this interaction occurs in vivo, we took a cotransfection approach, given the lack of cell lines expressing both Hand1 and FHL2 at appropriate levels. NIH 3T3 cells were transiently transfected either with the FLAG-Hand1 vector alone (Fig. 2B, lane 3), a vector for MYC epitope-tagged FHL2 (MYC-FHL2) alone (Fig. 2B, lane 5), or a combination of vectors for FLAG-Hand1 and MYC-FHL2 (lane 7). Extracts from the transfected cells, along with untransfected controls, were first immunoprecipitated with an α-FLAG M2 antibody (lanes 2, 4, 6, and 8). The associated proteins were then subjected to Western blot analysis with an α-MYC antibody (9E10.2). An intense band corresponding to MYC-FHL2 was readily detected only when FLAG-Hand1 was expressed and immunoprecipitated by the FLAG antibody (lane 8), indicating that Hand1 specifically interacted with FHL2 in vivo. To further determine whether endogenous Hand1 and FHL2 associate in cardiomyocytes, we carried out CO-IP experiments on lysates prepared from P19-derived cardiomyocytes using a α-FHL2 monoclonal antibody followed by Western analysis with an α-Hand1 antibody (Fig. 2C). A distinct band corresponding to Hand1 was readily detected when FHL2 was expressed and immunoprecipitated by the α-FHL2 antibody (lane 2), indicating that the two proteins physically interact in cardiomyocytes. Since P19 cells can differentiate into spontaneously beating cardiomyocytes which exhibit the biochemical and physiological properties of their embryological equivalents (47), this finding can be extrapolated to an interaction between endogenous Hand1 and FHL2 in the developing heart.
FIG. 2.
Hand1 and FHL2 interact in vitro and in vivo in embryonic cardiomyocytes. (A) In vitro pull-down assay showing Hand1 interaction with FHL2. SDS-PAGE of 35S-labeled FHL2 were incubated with 0.5 μg of GST-Hand1 or GST-bHLH, respectively. FHL2 was pulled down by GST-Hand1 and GST-bHLH but not by GST alone. (B) Immunoprecipitation of MYC-FHL2 by full-length FLAG-Hand1 demonstrating that Hand1 and FHL2 can interact in vivo. Transient transfections of full-length MYC-FHL2 together with full-length FLAG-Hand1 constructs (lanes 7 and 8) were performed in NIH 3T3 cells as described in Materials and Methods. Immunoprecipitation (IP) was performed with α-FLAG M2 antibody and immunoblotting with α-MYC 9E10.2. Lanes 1 and 2 represent untransfected and lanes 3 to 6 represent single transfection controls (lanes 3 and 4, FLAG-Hand1; lanes 5 and 6, MYC-FHL2). Note the single band in lane 8, which represents MYC-FHL2 following immunoprecipitation by α-FLAG compared to the blank control lanes 2, 4, and 6. (C) Immunoprecipitation of endogenous FHL2 and Hand1 in P19-derived cardiomyocytes, demonstrating that the two proteins interact in the heart. Cardiomyocytes were derived from aggregated P19 cells differentiated in 0.8% DMSO as described in Materials and Methods. IP was performed on lysates with a monoclonal α-FHL2 antibody and immunoblotting with α-Hand1. Lane 1 represents 5% of the lysate input. Lane 2, CO-IP lane; lane 3, a no-primary-antibody negative control. Note the single band in lane 2, representing Hand1 following IP by α-FHL2 compared to the blank control lane 3. Expression of FHL2 in the starting lysate and CO-IP lanes was confirmed by immunoblotting with a polyclonal α-FHL2 antibody.
This interaction of FHL2 with Hand1 appeared to be relatively specific, since MyoD and E12/E47, both of which play essential roles in the activation of skeletal muscle genes, did not appear to interact with FHL2 according to in vitro protein binding assays (data not shown).
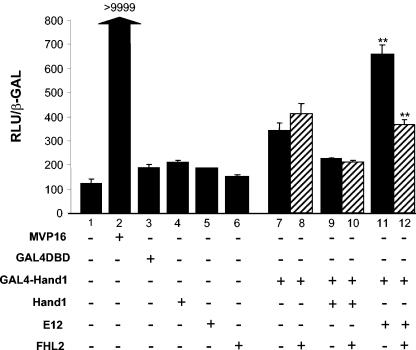
The association between Hand1 and FHL2 was corroborated by a mammalian two-hybrid system. For these studies, expression plasmids encoding a Gal4-Hand1 fusion plus Hand1, the E-protein E12, or FHL2 were cotransfected into NIH 3T3 cells along with the reporter gene CMVmin(Gal4)5-luciferase (Fig. 3). The chimeric protein Gal4-Hand1 containing the entire Hand1 coding region in frame with the DNA binding domain of Gal4 only weakly activated the (Gal4)5-luc reporter by approximately threefold in NIH 3T3 cells, consistent with that previously reported (21). Transfection of either E12 alone or FHL2 alone had no apparent effect on reporter gene activation (lanes 4 to 6). Coexpression of FHL2 had no effect on levels of activation of reporter arising from Gal4-Hand1 with or without Hand1 (lanes 6, 9, and 10). By contrast, coexpression of Gal4-Hand1 with E12 generated a significant activation of the (Gal4)5-luc reporter gene (lane 11), presumably as a result of the heterodimer formation between Hand1 and E12. Furthermore, FHL2 significantly inhibited the Gal4-Hand1-E12 transactivation of the reporter gene (lane 12), indicating that FHL2 is capable of interacting with a Hand1-E-protein heterodimer. This was confirmed in a reciprocal experiment whereby Gal4-E12 synergistic transactivation of reporter with Hand1 was inhibited by FHL2 (data not shown).
FIG. 3.
Mammalian two-hybrid analyses of Hand1 and FHL2 interactions in NIH 3T3 cells. NIH 3T3 cells were cotransfected with GAL4-DBD, GAL4-Hand1, and the CMVmin(Gal)5-luc reporter gene along with MVP16 as a positive control, Hand1, E12, or FHL2 or Hand1 plus FHL2 or E12 plus FHL2. MVP16 dramatically activated luciferase expression to confirm that the system is working (lane 2). Hand1, E12, or FHL2 in isolation had no effect on luciferase expression (lanes 4 to 6). GAL4-Hand1 activated luciferase expression by approximately threefold (lane 7) compared to reporter alone (lane 1). Coexpression of FHL2 had no significant effect on this level of activation (lane 8). Hand1 in combination with GAL4-Hand1 (lane 9) inhibited the activation observed with GAL4-Hand1 alone (lane 7). FHL2 had no effect on the Hand1-induced inhibition (lane 10). Coexpression of Gal4-Hand1 with E12 significantly activated luciferase expression (approximately sixfold; lane 11). FHL2 significantly inhibited GAL4-Hand1/E12-induced activation (lane 12), suggesting that FHL2 can functionally interact with Hand1-E protein heterodimers in vitro. Error bars reflect the standard errors of the means from a minimum of three independent transfections. **, P < 0.05.
Hand1 and FHL2 colocalize in the nuclei of H9c2 cells.
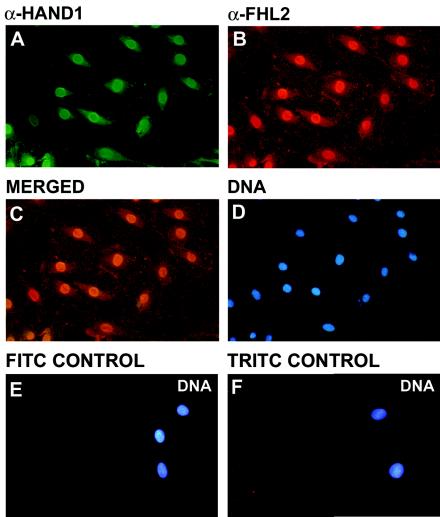
To investigate how the interaction between Hand1 and FHL2 translates in living cells, we first examined the cellular localization of the endogenous proteins in H9c2 cells. H9c2 is a rat cardioblast cell line derived from E18 ventricle in which both Hand1 and FHL2 are expressed, as determined by reverse transcription-PCR (data not shown). Immunofluorescence using polyclonal anti-mouse α-Hand1 and α-FHL2 antibodies (which cross-react with rat orthologues) revealed the two proteins to have predominantly nuclear staining (Fig. 4), suggesting that their interaction occurs in the nucleus, consistent with the previously reported transcriptional roles for both Hand1 and FHL2.
FIG. 4.
Subcellular colocalization of endogenous Hand1 and FHL2 in H9c2 cells. H9c2 cells were cultured on coverslips, fixed with 4% paraformaldehyde, and immunostained with polyclonal α-Hand1 (A) and α-FHL2 (B) antibodies followed by fluorescein isothiocyanate-conjugated anti-goat (A) and tetramethyl rhodamine isocyanate-conjugated anti-rabbit (B) antibodies, respectively. (C to E) Nuclei were visualized following staining with bis-benzamide (Hoechst 33342 [D]). Merged images of immunostained H9c2 cells demonstrated nuclear colocalization of Hand1 and FHL2 (C). Secondary antibody-alone negative controls are shown in panels D and E.
Hand1 and FHL2 can interact in living cells, as visualized by fluorescent protein two-fusion FRET.
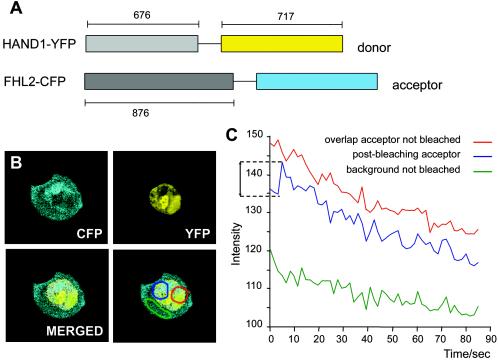
To further analyze the spatio-regulation of the Hand1-FHL2 interaction in an intact mammalian cell line, we performed “acceptor bleach” FRET analysis. We first generated chimeric fusion proteins of Hand1 and EYFP and FHL2 and ECFP to act as the donor and acceptor proteins, respectively (Fig. 5A). NIH 3T3 cells were cotransfected with the Hand1-EYFP and FHL2-ECFP constructs, and areas within cotransfected nuclei of overlapping and mutually exclusive expression were selected prior to performing FRET analysis (Fig. 5B). The CFP acceptor channel was used to bleach a region of overlap, which was subsequently excited using the YFP donor channel. As shown in Fig. 5C, emission from the acceptor, postbleaching, occurred following donor excitation, indicating FRET compared to the nonbleached region of overlap and the mutually exclusive region controls. Efficient FRET implies that the Hand1-EYFP and FHL2-ECFP fusion proteins colocalize in a subnuclear compartment at a distance of less than 50 Å apart and are therefore assumed to be physically associated.
FIG. 5.
Hand1 and FHL2 are spatially associated in intact living cells. FRET analysis of Hand1-YFP and FHL2-CFP fusion proteins in cotransfected NIH 3T3 cells. (A) N-terminal fluorescent fusion proteins were constructed as shown (numbers refer to size of each cassette, in base pairs) with Hand1-YFP as the donor and FHL2-CFP as the acceptor. (B) Images of the nuclear localization of the FHL2-CFP acceptor and Hand1-YFP donor were merged to confirm areas of overlap, and then three regions were selected prior to FRET analysis. The red region represents overlap of donor and acceptor in which the acceptor was not bleached. Blue represents a region of overlap where the acceptor was bleached. Green represents a mutually exclusive region not bleached by way of a background control. (C) Acceptor bleach FRET analysis was carried out whereby the three selected regions (red, blue, and green) were excited by the donor YFP channel (excitation, 500/20 nm), following in the case of the blue region appropriate prebleaching of the acceptor. Emission was recorded using the CFP channel (emission, 470/30 nm) as a measure of fluorescence intensity over time. A distinct increase in CFP intensity was observed for the blue overlap region, postbleaching of the acceptor and following donor YFP excitation, indicating efficient FRET and a direct physical association of the two fusion proteins. The data presented in panel C represent the averages of 10 independent FRET measurements.
FHL2 represses Hand1-E12 heterodimer-induced transcription but has no effect on Hand1-Hand1 homodimer activity.
Next we wanted to examine the possible functional significance of the observed interaction. As Hand1 can function as either a transcriptional activator or repressor (3, 21, 25, 46) depending on cell type or developmental context, we wanted to test whether the association with FHL2 might regulate the transcriptional activity of Hand1 and, therefore, impact on its biological activity in the developing heart.
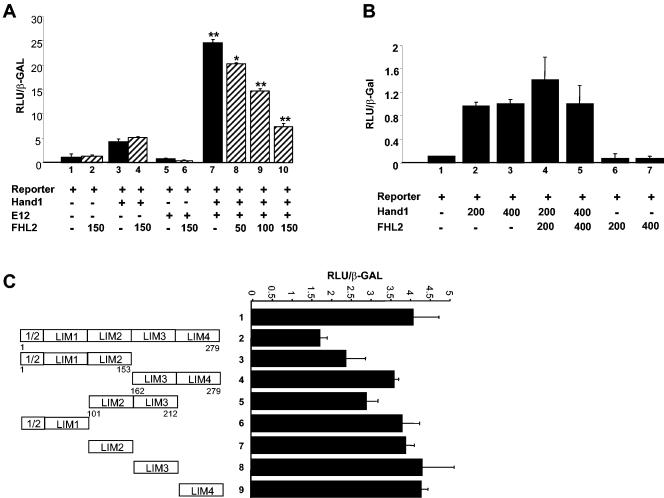
In the absence of a direct target gene for Hand1, we devised a reporter system comprising six high-affinity Thing1 box sequences (21) upstream of a minimal α-CA promoter (6, 31) and a luciferase cassette. Transfection of NIH 3T3 cells with reporter alone or reporter plus FHL2 revealed background luciferase activity (Fig. 6A, lanes 1 and 2). Cotransfection of reporter plus Hand1 activated the system approximately two- to threefold above reporter alone (Fig. 6A, lanes 1 and 3, and B, lanes 1 to 3). This relatively weak activation of the reporter by Hand1 is presumably brought about by low-affinity binding of the Thing1 sequences by Hand1 homodimers and is consistent with that previously reported (21, 46). FHL2 had no significant effect on Hand1 homodimer activation of the reporter (Fig. 6A, lanes 3 and 4) even when the dose of FHL2 was increased (Fig. 6B, lanes 2 to 4). Cotransfection of Hand1 with an E-protein, E12, a known interactor with Hand1, brought about significant coactivation of the mock target gene with luciferase activity approximately sevenfold above that of reporter alone (Fig. 6A, lane 7). This is consistent with the known ability of Hand1/E12 heterodimers to bind the Thing1 box with high affinity and activate transcription (21, 46). Surprisingly, given the lack of effect observed on Hand1 homodimer activation, FHL2 significantly inhibited Hand1/E12 coactivation of the target gene reporter. Furthermore, titration experiments with an increasing amount of FHL2 (50 to 150 ng) revealed a dose-dependent effect of FHL2 on the level of repression (Fig. 6A, lanes 8 to 10). The same results were obtained in a different cellular background with H9c2 cells, suggesting that the interaction observed in this assay is independent of other tissue-specific cofactors (data not shown). These data suggest that FHL2 can potentially regulate Hand1 biological activity by inhibiting Hand1-E protein heterodimer-induced activation of downstream target genes. Moreover, since FHL2 has apparently no effect on Hand1 homodimer transcriptional activity, the association of FHL2 with Hand1 represents a novel mechanism for tertiary differential regulation of Hand1 activity following dimerization choice.
FIG. 6.
FHL2 inhibits Hand1/E-protein heterodimer-induced transcriptional activation but has no effect on Hand1 homodimer activity. (A) Mock Hand1 target gene reporter, consisting of six concatemerized Thing1 boxes upstream of an α-CA minimal promoter driving luciferase, was cotransfected into NIH 3T3 cells with either Hand1, E12, or Hand1 plus E12 in the presence or absence of FHL2. Hand1 activated the system approximately two- threefold above reporter alone (lanes 1 and 3). FHL2 (150 ng) had no significant effect on Hand1 homodimer activation of the reporter (lanes 3 and 4). Cotransfection with E12 alone had no effect on reporter activation, and this remained unaltered in the presence of FHL2 (lanes 5 and 6). Cotransfection of Hand1 with E12 resulted in a sevenfold activation above that of reporter alone (lane 7). FHL2 significantly repressed Hand1/E12 coactivation of the mock target gene. Titration experiments with an increasing amount of FHL2 (50 to 150 ng) revealed a dose-dependent effect of FHL2 on the level of repression (lanes 8 to 10). (B) Increasing the dose of FHL2 (200 and 400 ng) had no effect on Hand1-induced activation of the mock target reporter (lanes 2 to 5). These findings suggest that the interaction of FHL2 with Hand1 impairs Hand1-E12 heterodimer function but has no effect on Hand1 homodimer activity. (C) Characterization of the FHL2 LIM domains involved in inhibition of Hand1-E12 transcriptional activity. Full-length FHL2 is required for efficient inhibition of Hand1-E12 activity (lanes 1 and 2). The effects of different FHL2 deletion constructs were analyzed on the Hand1-E12-activated mock target reporter. The N-terminal FHL2 (1/2-2) and the 2-3 constructs retained some inhibitory activity (lanes 3 and 5), suggesting that LIM domain 2 may play a prominent role. Single LIM domains in isolation had no effect on Hand1-E12 activity (lanes 6 to 9). Error bars reflect the standard errors of the means from a minimum of three independent transfections. **, P < 0.05; *, P < 0.1.
To investigate domains of FHL2 required to mediate the observed transcriptional inhibition, deletion constructs were coexpressed with Hand1 and E12 in the reporter assay. This revealed that the full-length FHL2 was required for optimal inhibition of Hand1/E12-induced transcriptional activation (Fig. 6C, lanes 1 and 2). No repression was observed for any of the LIM domains in isolation (Fig. 6C, lanes 6 to 9), and it appeared that the C-terminal LIM domains (3 and 4) were dispensable for FHL2 function in this assay because deletion of them resulted in a protein that retained its repressor-like activity (Fig. 6C, lane 4). However, the two constructs LIM (1/2-2) and LIM (2-3) did weakly repress the Hand1/E12 activation (Fig. 6C, lanes 3 and 5), suggesting that the second LIM domain may play a prominent role but that it requires the presence of additional neighboring LIM domains for full effect.
FHL2 repression of Hand1-E12 heterodimer transcriptional activity is not mediated via an effect on dimerization.
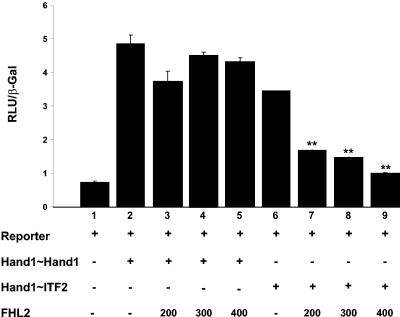
Next, we set out to determine the mechanism by which FHL2 is able to repress Hand1-E protein heterodimer-induced transcription. We first hypothesized that FHL2, through direct interaction with Hand1, may specifically impair Hand1/E-protein heterodimer formation (but not Hand1 homodimer formation), thus preventing the Hand1/E12 heterodimer binding to the Thing1 box target DNA sequences in our reporter gene assay. In order to investigate this, we made use of tethered Hand1-E-protein (Hand1∼ITF2) constructs in which complete monomers of Hand1 and ITF2 (murine functional analogue of E12 [49]) are joined to each other by a flexible polypeptide tether. Such enforced dimerization produces a dominant-positive heterodimer, which is insulated from negative regulation by other proteins (40). Cotransfection of NIH 3T3 cells with our mock target gene and a Hand1∼Hand1-tethered homodimer resulted in a sixfold activation of the reporter gene, which was not affected by increasing amounts of FHL2 (Fig. 7, lanes 1 to 5). Coexpression of reporter with a Hand1∼ITF2-forced heterodimer induced reporter gene activation by approximately sixfold (Fig. 7, lane 6). FHL2 was still able to significantly repress this heterodimer-induced activation in a dose-dependent manner (Fig. 7, lanes 7 to 9).
FIG. 7.
FHL2 inhibition of Hand1-E12-induced transcription is not mediated by an effect on heterodimerization per se. Cotransfection of NIH 3T3 cells with the mock target gene and a Hand1∼Hand1-tethered homodimer resulted in a sixfold activation of the reporter gene which was not effected by increasing amounts (200 to 400 ng) of FHL2 (lanes 1 to 5). Coexpression of reporter with a Hand1∼ITF2-forced heterodimer induced reporter gene activation by approximately sixfold (lane 6). FHL2 was still able to significantly repress Hand1∼ITF2-induced activation in a dose-dependent manner (lanes 7 to 9), suggesting a mechanism of FHL2 function distinct from disruption of Hand1-E protein heterodimerization. Error bars reflect the standard errors of the means from a minimum of three independent transfections. **, P < 0.05.
This finding supports the differential regulation of FHL2 on Hand1 E-protein heterodimers versus Handl homodimers and demonstrates that the mechanism of FHL2-induced inhibition of heterodimer transcriptional activity is not mediated via a negative effect on dimerization. As further confirmation, we also determined that FHL2 was unable to prevent CO-IP of MYC-tagged E12 with Hand1 following overexpression of the two proteins in NIH 3T3 cells (data not shown).
FHL2 does not appear to affect Hand1-E12 DNA binding.
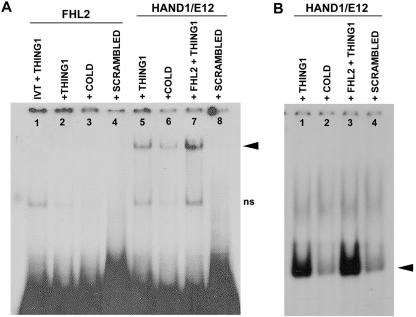
A second putative mechanism underlying FHL2 repression of Hand1-E protein-induced transcription might be at the level of DNA binding. FHL2 could directly inhibit the DNA binding of the Hand1-E factor protein complex, since it is feasible that FHL2 interaction with Hand1 results in a change in complex confirmation, which prevents the E-factor from entering the DNA binding pocket. Moreover, since localized structural differences among bHLH domains account for distinct dimerization and DNA binding properties (12), interaction of FHL2 with Hand1 need not necessarily affect Hand1 homodimer DNA binding. We investigated the possibility that FHL2 influences the DNA binding properties associated with the heterodimer complex by using EMSAs. IVT Hand1, E12, and FHL2 and proteins from cotransfected NIH 3T3 cell lysates were incubated with 32P-labeled Thing1 box and scrambled control oligonucleotide probes, and the generated protein-DNA complexes were resolved by native PAGE and autoradiography. As shown in Fig. 8A, IVT FHL2 alone does not bind to the Thing1 box sequence (lane 2). Hand1 and E12 in combination, as expected, bound the optimal Thing1 box such that a shifted band representing a Hand1-E12 heterodimer complex bound to DNA was observed (21, 46), and this could be competitively blocked by an excess of cold Thing1 box probe (Fig. 8A, lanes 5 and 6). Preincubation of Hand1/E12 with IVT FHL2 actually resulted in a slight enhancement in the bound Hand1/E12 complex (Fig. 8A, lane 7). This modest increase in Hand1/E12 DNA binding brought about by the addition of FHL2 was confirmed using cell lysates from NIH 3T3 cells cotransfected with all three proteins (Fig. 8B, lanes 1 and 4).
FIG. 8.
FHL2 does not appear to affect Hand1-E12 DNA binding. EMSAs with IVT Hand1, E12, and FHL2 and proteins from cotransfected NIH 3T3 cell lysates were incubated with 32P-labeled Thing1 box and scrambled control oligonucleotide probes, and the generated protein-DNA complexes were resolved by native PAGE and autoradiography. (A) IVT FHL2 alone does not bind to the Thing1 box sequence (lane 2). Hand1 and E12 in combination bind the optimal Thing1 box such that a shifted band representing a Hand1-E12 heterodimer complex bound to DNA is observed (21, 46), and this can be competitively blocked by an excess of cold Thing1 probe (lanes 5 and 6). Preincubation of Hand1/E12 with IVT FHL2 actually resulted in a slight enhancement in the bound Hand1-E12 complex (lane 7). (B) The enhanced Hand1-E12 DNA binding brought about by the addition of FHL2 was confirmed using cell lysates from NIH 3T3 cells cotransfected with all three proteins (lanes 1 and 3). Arrowheads indicate specific DNA binding complexes. ns, nonspecific bands.
Despite the described physical interaction of FHL2 with Hand1 and Hand1-E-protein heterodimers, we never detected FHL2 in the Hand1-E12 DNA complex in these EMSA studies. Thus, although there appears to be slightly elevated Hand1-E12 binding to the optimal Thing1 box in the presence of FHL2, the latter likely dissociates from these proteins once the bHLH-DNA complex enters the gel. This is an interesting observation and is consistent with that observed for muscle LIM protein in promoting DNA interaction of a MyoD-E47 heterodimer complex during muscle development (26). Moreover, it is clear that since we never observed a negative effect of FHL2 on Hand1-E factor DNA binding the repression of Hand1-E12 transcriptional activity described in this study is almost certainly not mediated at the level of binding of the bHLH heterodimer to its target sequence.
DISCUSSION
The Hand subfamily of class B bHLH factors consists of two members, Hand1 and Hand2, both of which are dynamically expressed in a number of embryologically distinct lineages during development (for review, see reference 15 and references therein). Both Hand proteins have been shown to play essential roles in the developing murine heart from gene targeting studies. Hand1 is thought to be essential for cardiac looping morphogenesis (44), whereas Hand2 is required for specification of the right ventricle and formation of the aortic arches (50). To date, though, the precise mode of function for either Hand protein remains unknown, although many studies collectively suggest these factors play roles in multiple gene programs and that the mechanism of their regulation is relatively complex.
In vitro studies have demonstrated for Hand1 the potential to both activate and repress transcription depending on the target sequence (consensus E-box or degenerate Thing1/D-box) and dimerization partner (21, 25, 46). Efforts to better understand the functional role of Hand proteins have focused on their dimerization partners, and in this respect they are unlike the majority of class B bHLH factors, since they exhibit a promiscuous ability to dimerize with themselves and other class B proteins such as members of the Hey/Hrt bHLH family (14). More recently, posttranslational modification of Hand proteins has been implicated in regulating choice of dimerization partner during the transition from proliferating to differentiating cell lineages. Notably, during differentiation of Rcho1 cells, Hand1 phosphorylation increases coincident with downregulation of the phosphatase PP2A active subunit, B56δ, and a reduced affinity of Hand1 for E-proteins (16). The fact that the phosphorylation status of Hand1 affects its dimerization affinities with E-proteins provides a mechanism for changing the dynamics of bHLH equilibrium within the cell and significantly advances our understanding as to just how the formation of specific Hand dimers may activate tissue-specific gene programs. However, once the dimer pool has reached intracellular equilibrium, there is clearly a requirement for a further level of regulation to broaden transcriptional control. Such regulation may manifest as “secondary” posttranslational modifications, such as phosphorylation of residues in the basic domain, to affect DNA binding or via protein-protein interaction(s) of Hand dimers (homo- or hetero-) with transcriptional cofactors.
Here we set out to investigate whether Hand1 could interact with non-bHLH cofactors and, more specifically, with known LIM domain-containing proteins coexpressed with Hand1 in the developing heart. We have demonstrated that Hand1 can interact with FHL2, a member of the four-and-a-half LIM domain protein family, and that this interaction impacts on transcriptional regulation by Hand1. This interaction supports the notion that Hand1 functions as a member of a cardiac-specific transcriptional complex and that its biological activity is regulated not only by dimerization choice but also at the level of tertiary protein interaction.
LIM domain proteins comprise a superfamily of cofactors characterized by the presence of one or more signature LIM domains consisting of a cysteine-rich motif with the consensus sequence CX2CX16-23HX2-CX2CX2CX16-21CX(C,H,D) that coordinately binds two zinc atoms. LIM domains interact specifically with other LIM domains and with many different protein domains and are thought to function as protein interaction modules mediating specific contacts between functional complexes and modulating the activity of the constituent proteins (for review, see reference 10). Many LIM proteins exert an influence over developmental events by affecting gene expression (17, 24, 55). LIM proteins that lack a DNA binding homeodomain have been postulated to function in regulating cellular differentiation, and of these a number have been reported to physically interact with bHLH factors in a tissue-specific manner: muscle LIM protein interacts with the myogenic bHLH factor MyoD to promote skeletal muscle myogenesis (26). LMO1 (RBTN1) and LMO2 (RBTN2), LIM domain proto-oncogenes activated in T-cell leukemia, interact with the bHLH factor SCL/TAL1 during erythroid development (54) and similarly, in pancreatic cells, Imx-1 and Pan-1 (E47) interact via their respective LIM and bHLH domains to synergistically activate expression of the insulin gene (19).
Thus, the interactions between tissue-specific bHLH factors and LIM proteins may represent a common mechanism that is utilized by many different developmental systems to enhance transcriptional activity.
FHL2 has been characterized as a transcriptional coactivator and corepressor, a dual role in which FHL2 is thought to stabilize the transcriptional complex as a type of bridging factor (36). In this study FHL2, while able to repress Hand1/E-protein transcriptional coactivation of a mock Hand1 target gene, had no effect on Hand1/Hand1 homodimer-induced transcription. To our knowledge this is the first reported instance of a differential regulation of bHLH heterodimers versus homodimers by an associated cofactor. Interaction with lineage-restricted non-bHLH proteins in complex to activate or repress transcription represents a tertiary level of regulation of Hand bHLH biological activity and a further mechanism, aside from dimerization choice, by which the more widely expressed Hand proteins may regulate different subsets of downstream target genes in distinct tissues. Clearly, this has functional implications for the activation of transcriptional pathways in the developing heart, since cardiomyocyte differentiation in ventricular myocardium may be accompanied not only by a switch in Hand1 dimerization affinity but also by the downregulation of Hand1-E-protein activity following interaction with FHL2.
FHL2 appears to influence the transcriptional activity of Hand1-E-protein heterodimers through a direct physical interaction. This interaction is specific, as FHL2 does not complex with the skeletal muscle bHLH MyoD or with E-proteins alone. A possible consequence of this interaction could be the disruption of the Hand1/E-protein heterodimer. To test this directly, we made use of tethered bHLH dimers, in which a single polyprotein encoded both Hand1 and the E-protein ITF2 separated by a flexible linker. The tethered Hand1∼ITF2 is resistant to disruption by other proteins present in the cell; consequently, any effect on the enforced heterodimer activity occurs independently of dimerization. In this study, FHL2 was still able to induce inhibition of the tethered heterodimer transcriptional activity in a manner equivalent to that observed for the separate Hand1 and E-factor proteins, thus precluding any effect of FHL2 on heterodimer formation per se.
The ability of FHL2 to interact with Hand1/E-protein heterodimers suggests that FHL2 and E-proteins interact with Hand1 at different positions. E-factor and Hand1 association depends on the HLH region (46); therefore, FHL2 may impact on the basic region of either Hand1 or its dimerization partner in a structural conformation which adversely effects DNA binding. That said, the function of FHL2 in this study could not be explained by a reduction in DNA binding of the Hand1/E12 complex, since we failed to observe a significant change in DNA binding with the addition of FHL2 in EMSAs. Moreover, our inability to detect a significant shift in the size of the Hand1-E12 complex binding to the Thing1 target sequences suggests that FHL2 is either absent from the DNA binding complex or that it may be part of the complex but is relatively unstable and thus dissociates during the EMSA. Alternatively, the interaction with FHL2 within the transcriptional complex may actually target Hand1/E-protein heterodimers away from the promoters of the genes it regulates. Consequently, any transcriptional effect is entirely disassociated from direct DNA binding and, as such, this represents a unique form of regulation which has hitherto not been described for other bHLH family members.
In general, LIM domains are thought to function as molecular adapters mediating the assembly of multiprotein complexes. Therefore, FHL2 may simply act as a scaffold for other cofactors (in this case repressors) to link the Hand1/E-factor complex to RNA polymerase II machinery or chromatin-remodeling complexes. Chromatin remodeling itself is an important prerequisite for transcriptional activity and provides a critical level of control independent of protein-protein interaction or direct DNA binding. The bHLH transcription factor Twist inhibits chromatin remodeling by blocking the activity of histone acetyltransferases (HATs) such as p300 (CBP, CREB binding protein) and PCAF (p300/CBP-associated protein), and this is thought to be the mechanism underlying its known inhibitory effects on transcription and differentiation of multiple cell lineages (20). The Hand factors are highly related to twist-like bHLH genes, and more recently Hand2 has been shown to physically interact with the CH3 and HAT domains of p300, in complex with GATA4, to synergistically activate the ANF promoter (9). In the context of this study, it is plausible that Hand1 also interacts with p300 and that FHL2 may disrupt this interaction exclusively in Hand1/E-protein complexes to prevent HAT activity and transcriptional derepression. This provides a mechanism of FHL2 repression which does not require FHL2 to act at the level of either Hand1-E protein heterodimerization or Hand1/E-protein binding of target DNA sequences but would be manifested in the transcriptional reporter assay experiments, since transiently transfected plasmids are rapidly chromatinized (41, 43, 51).
Unlike Hand1 (13, 44), FHL2 is not required for normal cardiovascular development (27); however, this does not detract from the functional implication of a Hand1-FHL2 interaction in the developing heart. The lack of a cardiac phenotype in FHL2-null embryos could be due to functional redundancy with other FHL family members, but it may also reflect a more subtle role for FHL2 during cardiac morphogenesis, such as mediating appropriate ventricular maturation or maintenance of cardiac cyto-architecture. Sustained β-adrenergic stimulation in adult FHL2 knockout mice provokes an exaggerated hypertrophic response in the heart compared to the response in wild-type littermates (27) and, more recently, FHL2 has been shown to interact with the mitogen-activated protein kinase ERK2 and partially antagonize a cardiac hypertrophic response in transgenic mice with constitutively active MEK1/2-ERK1/2 (42). FHL2 has also been shown to interact with the cardiac sarcomere protein Titin and recruit metabolic enzymes to sites of high energy consumption, thus contributing to maintenance of sarcomere integrity and ensuring that energy levels are sufficient to meet the demands of muscle contraction (29). These studies indicate that FHL2 may modulate ventricular function via structural remodeling and/or alterations in energy consumption in response to hypertrophic stimuli. Such a role has previously been attributed to developmental genes, required for patterning ventricular myocardium. In a recent study on CLP-1 (cardiac lineage protein 1)-null mice, it was suggested that an observed downregulation of Hand1 in the hearts of CLP-1 embryos directly promoted a fetal form of cardiac hypertrophy (22). In mice deficient for the homeobox transcription factor Irx4, adult-onset cardiomyopathy is preceded by significantly reduced expression of Hand1 in the left ventricle at midgestation stages (4). Hand1 has also been shown to be downregulated in human ischemic and dilated cardiomyopathies (39) and following induction of cardiac hypertrophy in adult rodent hearts (52). It would appear, therefore, that both Hand1 and FHL2 are capable of modifying cardiac responses in the myocardium to genetic insult or environmental stresses and that collectively they may play an important role during hypertrophic signaling.
Further functional studies should assist in clarifying the precise mechanism of action of FHL2 on Hand1 biological activity and the impact this may have on pathways in both the developing heart and those underlying adult onset cardiac disease.
Acknowledgments
We thank C. Murre and J. C. Cross for providing plasmids, R. Schule for the polyclonal α-FHL2 antibody, and R. B. Dhand and P. J. Scambler for discussion and comments.
This work was supported by grants from the British Heart Foundation and the Medical Research Council to P.R.R.
REFERENCES
- 1.Arber, S., J. J. Hunter, J. Ross, Jr., M. Hongo, G. Sansig, J. Borg, J. C. Perriard, K. R. Chien, and P. Caroni. 1997. MLP-deficient mice exhibit a disruption of cardiac cytoarchitectural organization, dilated cardiomyopathy, and heart failure. Cell 88:393-403. [DOI] [PubMed] [Google Scholar]
- 2.Begley, C. G., J. Visvader, A. R. Green, P. D. Aplan, D. Metcalf, I. R. Kirsch, and N. M. Gough. 1991. Molecular cloning and chromosomal localization of the murine homolog of the human helix-loop-helix gene SCL. Proc. Natl. Acad. Sci. USA 88:869-873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bounpheng, M. A., T. A. Morrish, S. G. Dodds, and B. A. Christy. 2000. Negative regulation of selected bHLH proteins by eHAND. Exp. Cell. Res. 257:320-331. [DOI] [PubMed] [Google Scholar]
- 4.Bruneau, B. G., Z. Z. Bao, D. Fatkin, J. Xavier-Neto, D. Georgakopoulos, C. T. Maguire, C. I. Berul, D. A. Kass, M. L. Kuroski-de Bold, A. J. de Bold, D. A. Conner, N. Rosenthal, C. L. Cepko, C. E. Seidman, and J. G. Seidman. 2001. Cardiomyopathy in Irx4-deficient mice is preceded by abnormal ventricular gene expression. Mol. Cell. Biol. 21:1730-1736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chan, K. K., S. K. Tsui, S. M. Lee, S. C. Luk, C. C. Liew, K. P. Fung, M. M. Waye, and C. Y. Lee. 1998. Molecular cloning and characterization of FHL2, a novel LIM domain protein preferentially expressed in human heart. Gene 210:345-350. [DOI] [PubMed] [Google Scholar]
- 6.Chang, K. S., K. N. Rothblum, and R. J. Schwartz. 1985. The complete sequence of the chicken alpha-cardiac actin gene: a highly conserved vertebrate gene. Nucleic Acids Res. 13:1223-1237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chu, P. H., P. Ruiz-Lozano, Q. Zhou, C. Cai, and J. Chen. 2000. Expression patterns of FHL/SLIM family members suggest important functional roles in skeletal muscle and cardiovascular system. Mech. Dev. 95:259-265. [DOI] [PubMed] [Google Scholar]
- 8.Cuppen, E., H. Gerrits, B. Pepers, B. Wieringa, and W. Hendriks. 1998. PDZ motifs in PTP-BL and RIL bind to internal protein segments in the LIM domain protein RIL. Mol. Biol. Cell 9:671-683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Dai, Y. S., P. Cserjesi, B. E. Markham, and J. D. Molkentin. 2002. The transcription factors GATA4 and dHAND physically interact to synergistically activate cardiac gene expression through a p300-dependent mechanism. J. Biol. Chem. 277:24390-24398. [DOI] [PubMed] [Google Scholar]
- 10.Dawid, I. B., J. J. Breen, and R. Toyama. 1998. LIM domains: multiple roles as adapters and functional modifiers in protein interactions. Trends Genet. 14:156-162. [DOI] [PubMed] [Google Scholar]
- 11.Du, X., P. Hublitz. T. Gunther, D. Wilhelm, C. Englert, and R. Schule. 2002. The LIM-only coactivator FHL2 modulates WT1 transcriptional activity during gonadal differentiation. Biochim. Biophys. Acta 1577:93-101. [DOI] [PubMed] [Google Scholar]
- 12.Ellenberger, T., D. Fass, M. Arnaud, and S. C. Harrison. 1994. Crystal structure of transcription factor E47: E-box recognition by a basic region helix-loop-helix dimer. Genes Dev. 8:970-980. [DOI] [PubMed] [Google Scholar]
- 13.Firulli, A. B., D. G. McFadden, Q. Lin, D. Srivastava, and E. N. Olson. 1998. Heart and extra-embryonic mesodermal defects in mouse embryos lacking the bHLH transcription factor Hand1. Nat. Genet. 18:266-270. [DOI] [PubMed] [Google Scholar]
- 14.Firulli, B. A., D. B. Hadzic, J. R. McDaid, and A. B. Firulli. 2000. The basic helix-loop-helix transcription factors dHAND and eHAND exhibit dimerization characteristics that suggest complex regulation of function. J. Biol. Chem. 275:33567-33573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Firulli, A. B. 2003. A HANDful of questions: the molecular biology of the heart and neural crest derivatives (HAND)-subclass of basic helix-loop-helix transcription factors. Gene 312:27-40. [DOI] [PubMed] [Google Scholar]
- 16.Firulli, B. A., M. J. Howard, J. R. McDaid, L. McIlreavey, K. M. Dionne, V. E. Centonze, P. Cserjesi, D. M. Virshup, and A. B. Firulli. 2003. PKA, PKC, and the protein phosphatase 2A influence HAND factor function: a mechanism for tissue-specific transcriptional regulation. Mol. Cell 12:1225-1237. [DOI] [PubMed] [Google Scholar]
- 17.Freyd, G., S. K. Kim, and H. R. Horvitz. 1990. Novel cysteine-rich motif and homeodomain in the product of the Caenorhabditis elegans cell lineage gene lin-11. Nature 344:876-879. [DOI] [PubMed] [Google Scholar]
- 18.Fung, Y. W., R. Wang, and C. C. Liew. 1996. Characterization of a human cardiac gene which encodes for a LIM domain protein and is developmentally expressed in myocardial development. J. Mol. Cell. Cardiol. 28:1203-1210. [DOI] [PubMed] [Google Scholar]
- 19.German, M. S., J. Wang, R. B. Chadwick, and W. J. Rutter. 1992. Synergistic activation of the insulin gene by a LIM-homeodomain protein and a basic helix-loop-helix protein: building a functional insulin minienhancer complex. Genes Dev. 6:2165-2176. [DOI] [PubMed] [Google Scholar]
- 20.Hamamori, Y., V. Sartorelli, V. Ogryzko, P. L. Puri, H. Y. Wu, J. Y. Wang, Y. Nakatani, and L. Kedes. 1999. Regulation of histone acetyltransferases p300 and PCAF by the bHLH protein twist and adenoviral oncoprotein E1A. Cell 96:405-413. [DOI] [PubMed] [Google Scholar]
- 21.Hollenberg, S. M., R. Sternglanz, P. F. Cheng, and H. Weintraub. 1995. Identification of a new family of tissue-specific basic helix-loop-helix proteins with a two-hybrid system. Mol. Cell. Biol. 15:3813-3822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Huang, F., M. Wagner, and M. A. Siddiqui. 2004. Ablation of the CLP-1 gene leads to down-regulation of the HAND1 gene and abnormality of the left ventricle of the heart and fetal death. Mech. Dev. 121:559-572. [DOI] [PubMed] [Google Scholar]
- 23.Johnson, J. E., S. J. Birren, T. Saito, and D. J. Anderson. 1992. DNA binding and transcriptional regulatory activity of mammalian achaete-scute homologous (MASH) proteins revealed by interaction with a muscle-specific enhancer. Proc. Natl. Acad. Sci. USA 89:3596-3600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Karlsson, O., S. Thor, T. Norberg, H. Ohlsson, and T. Edlund. 1990. Insulin gene enhancer binding protein Isl-1 is a member of a novel class of proteins containing both a homeo- and a Cys-His domain. Nature 344:879-882. [DOI] [PubMed] [Google Scholar]
- 25.Knofler, M., G. Meinhardt, S. Bauer, T. Loregger, R. Vasicek, D. J. Bloor, S. J. Kimber, and P. Husslein. 2002. Human Hand1 basic helix-loop-helix (bHLH) protein: extra-embryonic expression pattern, interaction partners and identification of its transcriptional repressor domains. Biochem. J. 36:641-651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kong, Y., M. J. Flick, A. J. Kudla, and S. F. Konieczny. 1997. Muscle LIM protein promotes myogenesis by enhancing the activity of MyoD. Mol. Cell. Biol. 17:4750-4760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kong, Y., J. M. Shelton, B. Rothermel, X. Li, J. A. Richardson, R. Bassel-Duby, and R. S. Williams. 2001. Cardiac-specific LIM protein FHL2 modifies the hypertrophic response to beta-adrenergic stimulation. Circulation 103:2731-2738. [DOI] [PubMed] [Google Scholar]
- 28.Kotaka, M., Y. M. Lau, K. K. Cheung, S. M. Lee, H. Y. Li, W. Y. Chan, K. P. Fung, C. Y. Lee, M. M. Waye, and S. K. Tsui. 2001. Elfin is expressed during early heart development. J. Cell Biochem. 83:463-472. [DOI] [PubMed] [Google Scholar]
- 29.Lange, S., D. Auerbach, P. McLoughlin, E. Perriard, B. W. Schafer, J. C. Perriard, and E. Ehler. 2002. Subcellular targeting of metabolic enzymes to titin in heart muscle may be mediated by DRAL/FHL-2. J. Cell Sci. 115:4925-4936. [DOI] [PubMed] [Google Scholar]
- 30.Larson, R. C., I. Lavenir, T. A. Larson, R. Baer, A. J. Warren, I. Wadman, K. Nottage, and T. H. Rabbitts. 1996. Protein dimerization between Lmo2 (Rbtn2) and Tal1 alters thymocyte development and potentiates T cell tumorigenesis in transgenic mice. EMBO J. 15:1021-1027. [PMC free article] [PubMed] [Google Scholar]
- 31.Lee, T. C., Y. Zhang, and R. J. Schwartz. 1994. Bifunctional transcriptional properties of YY1 in regulating muscle actin and c-myc gene expression during myogenesis. Oncogene 9:1047-1052. [PubMed] [Google Scholar]
- 32.Martin, B., R. Schneider, S. Janetzky, Z. Waibler, P. Pandur, M. Kuhl, J. Behrens, K. von der Mark, A. Starzinski-Powitz, and V. Wixler. 2002. The LIM-only protein FHL2 interacts with beta-catenin and promotes differentiation of mouse myoblasts. J. Cell Biol. 159:113-122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.McBurney, M. W., E. M. Jones-Villeneuve, M. K. Edwards, and P. J. Anderson. 1982. Control of muscle and neuronal differentiation in a cultured embryonal carcinoma cell line. Nature 299:165-167. [DOI] [PubMed] [Google Scholar]
- 34.McBurney, M. W. 1993. P19 embryonal carcinoma cells. Int. J. Dev. Biol. 37:135-140. [PubMed] [Google Scholar]
- 35.McFadden, D. G., J. McAnally, J. A. Richardson, J. Charitem, and E. N. Olson. 2002. Misexpression of dHAND induces ectopic digits in the developing limb bud in the absence of direct DNA binding. Development 129:3077-3088. [DOI] [PubMed] [Google Scholar]
- 36.McLoughlin, P., E. Ehler, G. Carlile, J. D. Licht, and B. W. Schafer. 2002. The LIM-only protein DRAL/FHL2 interacts with and is a corepressor for the promyelocytic leukemia zinc finger protein. J. Biol. Chem. 277:37045-37053. [DOI] [PubMed] [Google Scholar]
- 37.Morlon, A., and P. Sassone-Corsi. 2003. The LIM-only protein FHL2 is a serum-inducible transcriptional coactivator of AP-1. Proc. Natl. Acad. Sci. USA 100:3977-3982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Muller, J. M., U. Isele, E. Metzger, A. Rempel, M. Moser, A. Pscherer, T. Breyer, C. Holubarsch, R. Buettner, and R. Schule. 2000. FHL2, a novel tissue-specific coactivator of the androgen receptor. EMBO J. 19:359-369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Natarajan, A., H. Yamagishi, F. Ahmad, D. Li, R. Roberts, R. Matsuoka, S. Hill, and D. Srivastava. 2001. Human eHAND, but not dHAND, is down-regulated in cardiomyopathies. J. Mol. Cell. Cardiol. 33:1607-1614. [DOI] [PubMed] [Google Scholar]
- 40.Neuhold, L. A., and B. Wold. 1993. HLH forced dimers: tethering MyoD to E47 generates a dominant positive myogenic factor insulated from negative regulation by Id. Cell 74:1033-1042. [DOI] [PubMed] [Google Scholar]
- 41.Pullner, A., J. Mautner, T. Albert, and D. Eick. 1996. Nucleosomal structure of active and inactive c-myc genes. J. Biol. Chem. 271:31452-31457. [DOI] [PubMed] [Google Scholar]
- 42.Purcell, N. H., D. Darwis, O. F. Bueno, J. M. Muller, R. Schule, and J. D. Molkentin. 2004. Extracellular signal-regulated kinase 2 interacts with and is negatively regulated by the LIM-only protein FHL2 in cardiomyocytes. Mol. Cell. Biol. 24:1081-1095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Reeves, R., C. M. Gorman, and B. Howard. 1985. Minichromosome assembly of non-integrated plasmid DNA transfected into mammalian cells. Nucleic Acids Res. 13:3599-3615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Riley, P., L. Anson-Cartwright, and J. C. Cross. 1998. The Hand1 bHLH transcription factor is essential for placentation and cardiac morphogenesis. Nat. Genet. 18:271-275. [DOI] [PubMed] [Google Scholar]
- 45.Schmeichel, K. L., and M. C. Beckerle. 1994. The LIM domain is a modular protein-binding interface. Cell 79:211-219. [DOI] [PubMed] [Google Scholar]
- 46.Scott, I. C., L. Anson-Cartwright, P. Riley, D. Reda, and J. C. Cross. 2000. The Hand1 basic helix-loop-helix transcription factor regulates trophoblast differentiation via multiple mechanisms. Mol. Cell. Biol. 20:530-541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Skerjanc, I. S. 1999. Cardiac and skeletal muscle development in P19 embryonal carcinoma cells. Trends Cardiovasc. Med. 9:139-143. [DOI] [PubMed] [Google Scholar]
- 48.Smart, N., A. A. Hill, J. C. Cross, and P. R. Riley. 2002. A differential screen for putative targets of the bHLH transcription factor Hand1 in cardiac morphogenesis. Mech. Dev. 119(Suppl. 1):S65-S71. [DOI] [PubMed] [Google Scholar]
- 49.Soosaar, A., A. Chiaramello, M. X. Zuber, and T. Neuman. 1994. Expression of basic-helix-loop-helix transcription factor ME2 during brain development and in the regions of neuronal plasticity in the adult brain. Brain Res. Mol. Brain Res. 25:176-180. [DOI] [PubMed] [Google Scholar]
- 50.Srivastava, D., T. Thomas, Q. Lin, M. L. Kirby, D. Brown, and E. N. Olson. 1997. Regulation of cardiac mesodermal and neural crest development by the bHLH transcription factor, dHAND. Nat. Genet. 16:154-160. [DOI] [PubMed] [Google Scholar]
- 51.Stanfield-Oakley, S. A., and J. D. Griffith. 1996. Nucleosomal arrangement of HIV-1 DNA: maps generated from an integrated genome and an EBV-based episomal model. J. Mol. Biol. 256:503-516. [DOI] [PubMed] [Google Scholar]
- 52.Thattaliyath, B. D., C. B. Livi, M. E. Steinhelper, G. M. Toney, and A. B. Firulli. 2002. HAND1 and HAND2 are expressed in the adult-rodent heart and are modulated during cardiac hypertrophy. Biochem. Biophys. Res. Commun. 297:870-875. [DOI] [PubMed] [Google Scholar]
- 53.Tu, Y., F. Li, S. Goicoechea, and C. Wu. 1999. The LIM-only protein PINCH directly interacts with integrin-linked kinase and is recruited to integrin-rich sites in spreading cells. Mol. Cell. Biol. 19:2425-2434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Valge-Archer, V. E., H. Osada, A. J. Warren, A. Forster, J. Li, R. Baer, and T. H. Rabbitts. 1994. The LIM protein RBTN2 and the basic helix-loop-helix protein TAL1 are present in a complex in erythroid cells. Proc. Natl. Acad. Sci. USA 91:8617-8621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Way, J. C., and M. Chalfie. 1988. mec-3, a homeobox-containing gene that specifies differentiation of the touch receptor neurons in C. elegans. Cell 54:5-16. [DOI] [PubMed] [Google Scholar]
- 56.Wei, Y., C. A. Renard, C. Labalette, Y. Wu, L. Levy, C. Neuveut, X. Prieur, M. Flajolet, S. Prigent, and M. A. Buendia. 2003. Identification of the LIM protein FHL2 as a coactivator of beta-catenin. J. Biol. Chem. 278:5188-5194. [DOI] [PubMed] [Google Scholar]
- 57.Wilkinson, D. G. 1992. In situ hybridisation: a practical approach, p. 75. IRL Press, Oxford, England.
- 58.Yan, J., J. Zhu, H. Zhong, Q. Lu, C. Huang, and Q. Ye. 2003. BRCA1 interacts with FHL2 and enhances FHL2 transactivation function. FEBS Lett. 553:183-189. [DOI] [PubMed] [Google Scholar]