Abstract
Background
A long-standing view of Indian biodiversity is that while rich in species, there are few endemics or in-situ radiations within the subcontinent. One exception is the Western Ghats biodiversity hotspot, an isolated mountain range with many endemic species. Understanding the origins of the montane-restricted species is crucial to illuminate both taxonomic and environmental history.
Results
With evidence from genetic, morphometric, song, and plumage data, we show that two songbird lineages endemic to the Western Ghats montane forest each have diversified into multiple distinct species. Historically labeled as single species of widespread Asian genera, these two lineages are highly divergent and do not group with the taxa in which they were previously classified but rather are distinct early divergences in larger Asian clades of flycatchers and babblers. Here we designated two new genera, the Western Ghats shortwings as Sholicola and the laughingthrushes as Montecincla, and evaluated species-limits to reflect distinct units by revising six previously named taxa and describing one novel species. Divergence dating showed that both these montane groups split from their Himalayan relatives during the Miocene, which is coincident with a shift towards arid conditions that fragmented the previously contiguous humid forest across peninsular India and isolated these lineages in the Western Ghats. Furthermore, these two genera showed congruent patterns of diversification across the Western Ghats Sky Islands, coincident with other climatic changes.
Conclusion
Our study reveals the existence of two independent endemic radiations in the high montane Western Ghats or Shola Sky Islands with coincident divergence times, highlighting the role of climate in the diversification of these ancient lineages. The endemic and highly divergent nature of these previously unrecognized species underscores the dearth of knowledge about the biogeography of the Asian tropics, even for comparatively well-known groups such as birds. The substantial increase in the diversity of this region underscores the need for more rigorous systematic analysis to inform biodiversity studies and conservation efforts.
Electronic supplementary material
The online version of this article (doi:10.1186/s12862-017-0882-6) contains supplementary material, which is available to authorized users.
Keywords: Phylogenetics, Birds, Shola, Passerine, Montane, Sky-islands, Taxonomy, Tropics
Background
The Western Ghats (WG), an isolated coastal mountain chain in the southwest of India, is a global biodiversity hotspot [1]. Despite a long history of human occupation, knowledge of this biodiversity remains poor [2, 3]. Modern systematic analyses to assess species distinctiveness and their responses to past climatic events are urgently needed to inform conservation efforts in such montane tropical systems as the Western Ghats, where diversity and threat levels are high [1, 4, 5].
In most current avian taxonomic treatments, WG lineages are a subset of the diverse avifaunal groups in the Himalayas and Southeast Asia. WG endemics are usually circumscribed as single species of larger Asian groups with limited differentiation across the WG mountain range [6–8]. Contrary to this traditional view, the hitherto first and only phylogenetic investigation of a WG endemic songbird challenged both these ideas – it revealed considerable genetic divergence between populations across the mountain range as well as from its congeners, rendering the traditional genus non-monophyletic [9]. The Indian subcontinent underwent dramatic climatic changes during the Cenozoic, which may have influenced species dispersal to and diversification within the WG [10, 11]. The long-favored ‘Satpura Hypothesis’ [12–16] suggested a specific colonization route for species to disperse from the Himalayas to the WG through the Satpura Hills, a narrow band of wet forest across central India. In the Miocene, peninsular India was much more humid with near continuous forest cover and since this period, changing climate conditions and local tectonic events led to gradual drying of northwestern and central India and the establishment of the distinct patches of wet-zone forested regions on the highlands [15, 17]. Until now, support of this hypothesis was mainly from observations of avian species distributions [6] due to the dearth of phylogenetic analyses of Indian birds [3].
The peaks of the WG range host a unique form of tropical montane cloud forest known as Shola, a natural matrix of forests and grasslands [18]. The Shola habitat is restricted to the highest elevation zone and is characterized by high rainfall, humidity and low temperatures relative to lower elevations [18, 19]. A variety of endemic taxa are exclusively found in this habitat [18, 19]. These species often have disjunct distributions across the WG mountain-tops, which have been likened to ‘islands’ of specific habitat and microclimatic conditions in an ‘ocean’ of unsuitable habitat [20]. While there are several classic examples of speciation and adaptive radiations across oceanic islands, only a few studies have examined similar patterns across montane or sky islands [2].
Two endemic avian species groups in the Western Ghats sky islands, the Kerala or Black-chinned Laughingthrushes (formerly placed in Garrulax, currently Trochalopteron [8, 21] or Strophocincla [7, 22]) and the Western Ghats Shortwing (alternatively placed in Brachypteryx [8, 21], Myiomela [7, 23] or Callene [24]) have been surrounded by taxonomic confusion, complicating studies of their diversity. The laughingthrushes exhibit striking plumage variation across the different isolated sky island peaks and have been alternatively considered as one [6, 25], two [7, 8, 21, 22], or four [26] species. The shortwings have been described as one [6, 21, 25] or two [7, 8] species. Previous phylogenetic analysis for some populations of WG Shortwings showed deep divergences across the sky island complex [9].
Elucidating the origins of montane species is crucial to illuminate the evolutionary and environmental history of this landscape [2]. We reconstructed the phylogenies of the WG laughingthrush and shortwing complexes to determine their evolutionary history and test hypotheses of diversification in relation to past climatic events. Furthermore, we examined discrete variation across populations of these two lineages using multiple types of data – genetic, song, plumage and morphometric – to determine species-limits and assess differentiation within the WG range.
Methods
Sample collection
From January 2012 to May 2013 we conducted expeditions to survey and collect samples across the entire distributional range of both species complexes (Additional file 1: Figure S1). We followed Robin et al. [9] for field sampling techniques to capture birds with multiple 12m * 2 m mist-nets and collect blood samples from the brachial vein in Queen’s lysis buffer. For one location where field sampling proved difficult (Bababudan Hills) for the shortwings, we used two museum samples from the Natural History Museum (Tring, UK; NHMUK) for DNA analysis.
DNA sequencing
We extracted DNA (using the Qiagen Blood and Tissue Extraction Kit) and used standard procedures [9, 27] to sequence 26 and 31 individuals of the laughingthrush and the shortwing complexes, respectively, across their entire distributions (Additional file 1: Tables S1 and S2). We generated sequence data from five and eight loci, respectively, for both groups to match published phylogenies of their relatives [27–29]. For the laughingthrushes, we sequenced 5 loci: cytochrome b (CYTB), NADH dehydrogenase subunit-3 (ND3), the fifth intron of nuclear b-fibrinogen (FIB5), the third intron of the muscle-specific kinase Gene (MUSK), and the fifth intron of transforming growth factor β2 (TGF) using standard primers [30] and standard PCR procedures (see [9, 30, 31]). For shortwings, we sequenced 8 loci: NADH dehydrogenase subunit-2 (ND2), ND3, CYTB, cytochrome c oxidase 1 (CO1), intron 2 of myoglobin (MYO), introns 6 and 7 of ornithine decarboxylase (ODC), intron 11 of the glyceraldehyde-3-phosphodehydrogenase (GAPDH), and intron 3 of lactate dehydrogenase (LDH). DNA sequences were assembled, annotated, and aligned using Geneious 6.1.4.
Phylogenetic analyses
WG laughingthrushes
To reconstruct the evolutionary relationships of the WG Laughingthrushes, we re-analyzed a larger clade encompassing laughingthrushes in general (Leiothrichidae) as per recent studies on sylvioid songbirds [27, 32, 33]. We assembled a matrix by incorporating representatives of key members of other laughingthrush species with our data from all lineages found in the Western Ghats. As outgroups, we included one representative of each of the other major clades of babblers [27]. We assembled two matrices of five loci each: to examine divergences within the WG Laughingthrush complex, we included all 26 WG individuals and several other babblers as outgroups; and to examine the relationships of the WG Laughingthrushes with other babblers, we assembled a matrix comprising 73 taxa (Additional file 1: Table S1). PartitionFinder v1.01 [34] determined that the best partitioning scheme of gene regions divided by rates of evolution was four partitions: 1st position CYTB + 1st position ND3, 2nd position CYTB + 2nd position ND3, 3rd position CYTB + 3rd position ND3, FIB5 + MUSK + TGF.
WG shortwings
Preliminary analyses found that shortwings belong within the flycatcher/chat complex. To examine variation within the WG Shortwings, we assembled a phylogenetic matrix of 34 taxa for four loci: ND2, ND3, CYTB, CO1 (Additional file 1: Table S2). To examine their placement in a broader phylogenetic context, we compiled data for species across a larger clade based on other published studies [28, 29, 35] and included all distinct WG Shortwing lineages for a total of 96 species for six loci -- ND2, CYTB, MYO, ODC, GAPDH, and LDH (Additional file 1: Table S2). The best partitioning scheme for this dataset according to PartitionFinder was 8 partitions: 1st position CYTB, 2nd position CYTB, 3rd position CYTB, 1st position ND2, 2nd position ND2, 3rd position ND2, MYO, and GAPDH + LDH + ODC.
For both groups, we conducted maximum likelihood (ML) phylogenetic analyses using RAxML 8 [36] by partitioning genes according to similar evolutionary rates determined with PartitionFinder. We conducted 1000 random bootstrap replicates that were subsequently used to search for the best ML tree. We compared relationships from this analysis to those using different optimality criteria such as maximum parsimony, using PAUP* [37], and Bayesian inference, using MrBayes 3.2 [38]. In PAUP*, we treated all characters with equal weights and ran heuristic searches of 1000 random addition replicates. In MrBayes, we used the same partition scheme as ML analyses and ran two Monte Carlo Markov Chain runs of four chains each for 20 million generations, sampling every 500th generation. We used default priors and unlinked parameters across partitions except for branch length calculations. We assessed convergence and stationarity of runs using Tracer v1.6 [39] and AWTY [40], discarding the first 5000 generations of each run as burn-in.
Divergence dating
We used BEAST v1.75 [41] to estimate the timing of lineage divergence. For both groups, we used one representative of each distinct lineage for divergence dating analysis. In each analysis, we unlinked substitution and clock rates, and linked tree models for each locus. To calibrate the laughingthrush/babbler phylogeny, we used a secondary calibration for the divergence time between the families Timaliidae, Pellorneidae + Leiotrichidae as 20.92 Ma (standard deviation of 2.11 Ma), taken from [29], a recent time-calibrated phylogeny of Asian passerines using 13 corroborated clade ages based on fossil and biogeographic calibrations (see Additional file 1 in [29]). To calibrate the shortwing/flycatcher tree, we used the same source to date the split between the families Muscicapidae and Turdidae as 21 Ma (standard deviation of 2 Ma) [29]. We understand that dates for the bird tree of life can be controversial but we chose this source [29] because it was the most densely-sampled, recent phylogeny for taxa relevant to this study. For each analysis, we used an uncorrelated lognormal relaxed clock model with a birth-death speciation tree prior. We conducted two runs of 20 million generations, sampling every 1000th and discarding the first 5000 as burn-in. We used Tracer v1.6 [39] to ensure stationarity of chains for all parameters (ESS values >200).
Ancestral area reconstruction
We used Lagrange v. 20130526 [42] to reconstruct ancestral areas for both groups. For laughingthrushes and relatives, we defined eight areas – Africa, Peninsular India, Himalayas, Southeast Asia, China, Sundaland, Philippines, and Assam Hills. For shortwings and relatives, we defined nine areas based on the geographic extent of these species: Africa + Southwest Asia, Peninsular India, Himalayas, Southeast Asia, China, Sundaland, Philippines, Eurasia, and Australasia/New World. In Lagrange, we assigned the ranges of each species based on their current geographical distributions. We allowed the ancestral area reconstructions to include any combination of areas. We used the time-calibrated tree from our BEAST analysis to reconstruct the likelihood of ancestral changes in distribution.
Species limits
We conducted focused mist-net based sampling over 3 years (2011 – 2014) and several years (2000 – 2010) of mixed sampling effort (point counts, transects, ad libitum observations, and mist-net sampling). During this period, we captured and examined 359 WG laughingthrushes and 430 WG shortwings. We collected a variety of vocal, plumage, morphometric and genetic data, along with extensive natural history observations of these species.
To assess species limits, we compared all population clades to determine statistically discrete units of nucleotide substitutions as well as distinct differences in plumage characteristics, morphometric variation, and song features. We chose this method because it allows using all available evidence to infer absence of gene flow and to delineate evolutionarily distinct units as species. Our species delineations are consistent with multiple species concepts - phylogenetic species concept, biological species concept, general lineage concept, and integrative taxonomy.
Coalescent-based test of species delimitation
We utilized a coalescent-based approach to statistically test species delimitation using the program BPP v3.1 [43]. This method employs the multispecies coalescent to compare different models of species delimitation and phylogeny in a Bayesian framework while accounting for incomplete lineage sorting due to ancestral polymorphism and gene tree-species tree conflicts [43]. We used the joint species delimitation and species tree analysis in BPP to test if the number of distinct units/species as delineated by our character-based analysis (see below) was statistically significant. For each lineage, we used the genetic dataset assembled to examine within-group variation and designated individuals by population/clade. For population size parameters, we assigned the gamma prior G (2, 1000), with mean 2/2000 = 0.001. We ran each analysis at least twice to confirm consistency between runs. Each run was for 100,000 samples, sampling frequency was 5, and the burnin was set to 20,000.
Morphometrics
All captured birds were banded and measurements of the length of right tarsus, flattened chord right wing, bill (from base of skull) and tail were taken following SAFRING manual [44] and USDA [45] guidelines. Each measurement of a feature was repeated thrice to examine measurement error [46]. Tarsus and bill measurements were taken with Mitutoyo ABS Digimatic Caliper (Mitutoyo Corp Japan) with accuracy of 0.02mm. Wing and tail measurements were taken with a wing rule (WING15ECON Avinet Inc.) that had a flush stop and calibration from both directions.
We analyzed the four morphometric variables (bill length, tarsus length, wing length and tail length; Additional file 1: Tables S3, S4) using multivariate statistics. For each species complex, we considered each distinct population clade separately and built a linear discriminant function with a common covariance matrix for all groups (JMP version 8). This was visualized with a canonical plot including a 95% confidence ellipse of the mean of each group and biplot rays indicating the direction of variables in canonical space. The groups with non-overlapping ellipses are considered different in morphology with the direction of biplot rays indicating the variables contributing to the observed differences. Wing and tail lengths for WG Shortwings were found to be collinear, but no significant collinearity among morphometric variables was detected in WG Laughingthrushes. However, a repetition of the discriminant analyses with reduced data set of three principal components (comprising 89.76% of the variation for WG Shortwings, 85% for WG Laughingthrushes) did not alter the results, hence we retained the analyses with the original variables for easier interpretation.
Plumage
For plumage comparisons, we examined and noted features of each population by studying live birds (sampled in the field), photographs of sampled birds and museum specimens (Bombay Natural History Society; BNHS, India) to make side-by-side comparisons, and descriptions in the literature. We noted differences in coloration across discrete feather patches to assess whether populations had unique plumage patterns.
Song
Field recording of 38 individual singing males was carried out between 0700 to 1100 h following Robin et al. [47]. Each recording consisted of only one continuous song bout from a single individual recorded using a Sennheisser shotgun microphone (ME66-K6) and Marantz Digital Audio Recorder (PMD660). Recordings were converted into spectrograms in Raven Pro 1.3 [48] at a sampling rate of 48 kHz. For each song, we collected data on frequency and note length parameters for each note, with multiple songs per individual and multiple individuals per population. A detailed statistical analysis of songs and syllables across multiple populations is presented in Purushotham and Robin [49].
Results
Phylogenetic analyses
The larger, family-level phylogenetic analyses indicated that both WG lineages diverged early within their respective clades (Fig. 1). Each of these two lineages is a distinct clade not closely related to the genus historically or currently grouped within. Additionally, both were found to be discrete clades deeply divergent from their closest relatives. The WG Laughingthrushes did not group with any of the other traditional laughingthrush clades (including ones designated as Trochalopteron or Strophocincla), but instead were sister to a clade composed of Heterophasia, Minla, Actinodura, Leiothrix, Liocichla and Crocias (Fig. 1a). Similarly, we reconstructed the WG Shortwings as sister to a newly uncovered clade comprised of several genera of mainly Asian blue flycatchers, the Niltavinae [28, 35], and not closely related to traditional shortwings in the genera Myiomela or Brachypteryx (Fig. 1b).
Fig. 1.
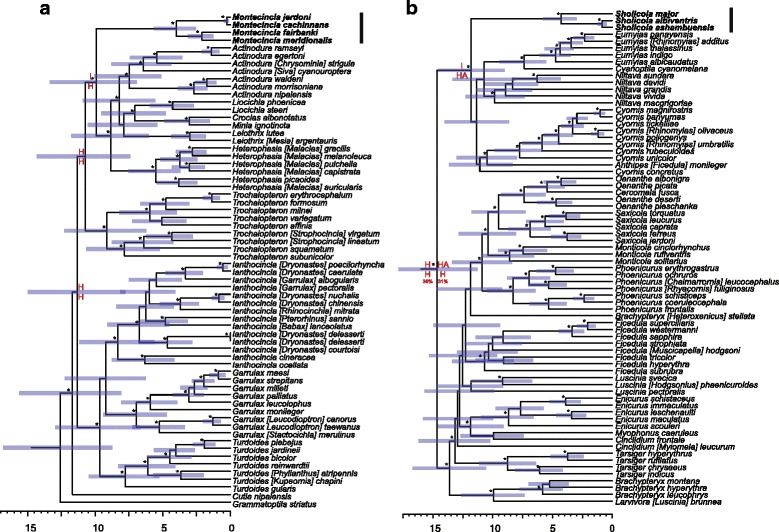
Time-calibrated phylogenies of the (a) laughingthrush/babbler and (b) shortwing/flycatcher clades. The WG Laughingthrushes, Montecincla, did not group with any of the other laughingthrush clades and are sister to a clade of birds traditionally not included in laughingthrushes. The WG Shortwings, Sholicola, are not part of shortwings (Brachypteryx or Myiomela) but are sister to a clade of Asian blue flycatchers, the Niltavinae. Generic designations follow Clements Checklist [8]; Handbook of the Birds of the World [22] names are shown in brackets if different. Node bars show 95% HPD estimates of divergence dates and stars indicate ML bootstrap of 70% or higher and Bayesian posterior probability of 95% or higher. Letters (in red) show ancestral area reconstruction at relevant nodes (I = Peninsular India; H = Himalayas; A = Southeast Asia)
Both WG lineages are not only divergent from their closest relatives, they are also not sister to any single genus as defined under current taxonomic treatments. Thus, they cannot easily be lumped or subsumed under an existing clade without an extensive reorganization of multiple genera. As a better alternative, we designated new genus names: Western Ghats Laughingthrushes as Montecincla and Western Ghats Shortwings as Sholicola (see below for descriptions), to recognize their distinctiveness and to avoid confusion with older names associated with other taxa that are not part of these new groups.
The ancestral area reconstruction showed dispersal into the WG for Montecincla as most likely from the Himalayas and for Sholicola from the Himalayas + Southeast Asia (Additional file 1: Figures. 1, S2, S3). The two WG lineages have a similar estimated timing of separation from their closest relatives (Table 1). These dates also coincide with climatic events that led to the drying of peninsular India (Table 1), perhaps leading to discontinuing gene flow and a differentiation between peninsular forms from Himalayan / Southeast Asian forms. Furthermore, the divergence of populations within both lineages across the sky islands exhibits similar timing of diversification in response to the same biogeographic barriers (see below).
Table 1.
Estimates of divergences dates and geological events
| Event | Timing estimates (Ma) |
|---|---|
| Montecincla vs. sister group - split WG/Himalayas | 11.57 (14.65 - 8.70) |
| Sholicola vs. sister group - split WG/Himalayas | 11.83 (14.59 - 9.00) |
| within Montecincla - split across Palghat Gap [AB,CD] | 4.7 (6.34 - 3.19) |
| within Sholicola - split across Palghat Gap [AB,CD] | 4.33 (5.79 - 2.98) |
| within Montecincla - split across Shencottah Gap [C,D] | 2.51 (3.60 - 1.60) |
| within Sholicola - split across Shencottah Gap [C,D] | 0.86 (1.24 - 0.49) |
| within Montecincla - split across Chaliyar Valley [A,B] | 0.33 (0.55 - 0.15) |
| Himalayas - peak constructional phase [55] | 15 - 10.5 |
| Tibetan Plateau - significantly uplift [55, 66] | 10 - 8 |
| Climate - enhanced aridity of Asia; onset of Indian monsoons [56–58, 66] | 9 - 8 |
| Vegetation - C4 plants start to replace C3 [56, 57] | 6 |
| Climate - major global shift; monsoons weakened [58] | 2.6 |
Species limits
Phylogenetic analysis of both groups showed clades corresponding to four major biogeographic regions in the WG, north to south: A- Bababudan & Banasura hills; B- Nilgiri hills; C- Anamalai, Palani, and Meghamalai hills; D- Ashambu hills (Fig. 2). Montecincla was comprised of four reciprocally monophyletic clades (Fig. 2b). Bayesian species delimitation using BPP also showed that these four clades were distinct species. Elevating former subspecies names, these are: Montecincla jerdoni (range A), M. cachinnans (B), M. fairbanki (C), M. meridionalis (D). Similarly, analysis of Sholicola revealed three reciprocally monophyletic clades (Fig. 2c) that were distinct species according to BPP. While two existing subspecies names that match these units can be elevated, namely Sholicola major (range A, B) and S. albiventris (C), the third (D) is a new species we designate below. Analyses of fixed characters of plumage, morphometrics and song data (below) match the distinct units uncovered in the phylogenetic and species delimitation analyses. Therefore multiple lines of data support four species in the Montecincla complex and three species in the Sholicola complex.
Fig. 2.
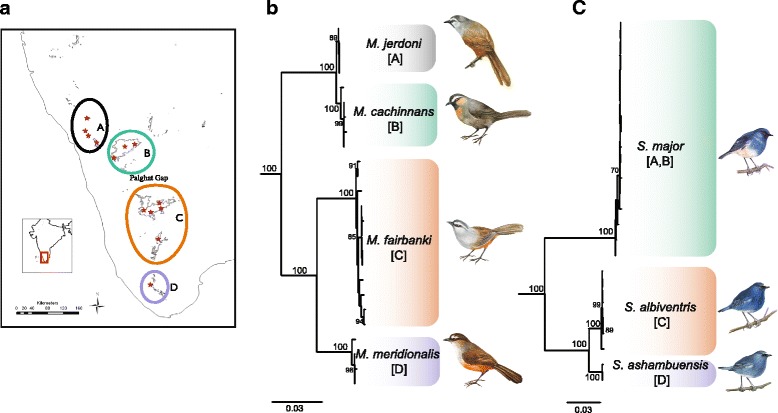
Ranges (a) and phylogenetic relationships of Montecincla (b) and Sholicola (c) species. a Inset map of the Western Ghats shows sampling localities (stars), and divisions of the sky islands based on differentiated taxa. b & c ML bootstrap values are shown at nodes. Bird illustrations by Maya Ramaswamy
Plumage
Our analysis of plumage variation for Montecincla supports four distinct groups (Table 2). All four species have distinguishing features in multiple color patches that separate these evolutionary units. Within-species variation is low and populations with similar features occur in geographically clustered sky islands (shown in Fig. 2). In Sholicola, S. major (A, B) showed striking differences from the other two species south of the Palghat Gap. The differences in plumage of between S. albiventris (C) and Sholicola sp. nov. (D; see below) are mainly in terms of the extent of white coloration on the belly. The newly described species has a considerably larger white belly patch (~24mm in length) than S. albiventris (~19mm) (Table 3).
Table 2.
Plumage differences between species of Montecincla
| Plumage feature | M. jerdoni | M. cachinnans | M. fairbanki | M. meridionale |
|---|---|---|---|---|
| Crown | Slaty brown | Slaty brown | Dark brown almost black | Grey brown |
| Chin | Black | Black | Grey | Pale grey |
| Ear coverts | Greyish white | Pale rufous | Pale grey | Brownish grey |
| Supercilium | White, long, reaching behind eye; black eye strip from lore | White, long reaching behind eye; black eye strip from lore | White, Long, extending behind eye; black stripe below lores through eyes | White, short, not extending behind eye |
| Nape | Slaty brown | Ashy brown | Brownish grey | Pale brown |
| Upper parts | Olive brown up to tail | Olive brown | Olive brown | Dull grey at nape brownish towards rump |
| Breast | Grey, with faint streaks | Bright rufous | Pale grey with faint streaks | Whitish grey with prominent dark streaks |
| Belly | Olive brown but centre of belly pale rufous | Ochraceous | Rufous to chestnut | White at centre with mild dark streaks, sides deep chestnut |
| Centre of belly | Pale rufous | Rufous | Rufous to chestnut | White centre |
| Flanks | Olive brown | Olive brown | Rufous to chestnut | Reddish chestnut |
Table 3.
Plumage differences between species of Sholicola
| Plumage feature | S. major | S. albiventris | S. ashambuensis |
|---|---|---|---|
| Crown | Slaty blue | Slaty blue | Slaty blue |
| Supraloral stripe | Faint blue | Whitish blue | Faint blue |
| Throat | Slaty blue | Slaty blue | Greyish blue |
| Breast | Slaty blue | Slaty blue | Greyish blue |
| Belly | Broad white patch | Narrow white patch from centre to vent | Narrow white patch extending anteriorly from vent to breast |
| Flanks | Pale rufous | Greyish blue | Greyish blue |
| Undertails coverts | Pale rufous | White | White |
| Upper parts | Slaty blue | Deep blue | Pale, slaty blue |
Morphometrics
The four species of Montecincla and the three species of Sholicola differed significantly from each other in morphospace occupied (Fig. 3). This differentiation within both Montecincla and Sholicola was largely driven by tarsus length, although the direction of increase was reversed – Montecincla in the southernmost region of Ashambu Hills (D) is larger than the other species further north, while in Sholicola the species in the southern hills were the smallest (Additional file 1: Tables S3, S4).
Fig. 3.
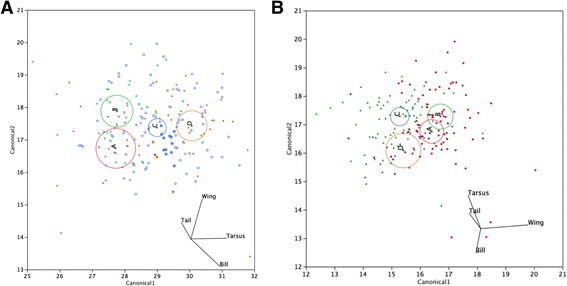
Discriminant Function Analysis showing morphometric differentiation between a) Montecincla and b) Sholicola populations/species. A-D labels refer to geographic regions (as in Fig. 2). Circles represent 95% confidence ellipses of the mean of each group, which are significantly different when not overlapping. The direction of the biplot rays show how variables contributed to the observed differences
Song
Spectrograms reveal the presence of unique song types in all four species of Montecincla and all three Sholicola species (Fig. 4). All four species of Montecincla show distinct features in song. Montecincla fairbanki sings at a higher bandwidth and song rate than the other populations, whereas M. meridionalis sings at a lower bandwidth than other species, while M. jerdoni has a higher song complexity with longer phrases than the other populations (Additional file 1: Table S5). Songs of all Sholicola species are also quantitatively distinct, with differences in song length and frequency across different species (more details in Purushotham and Robin [49]). The song of the newly described species of Sholicola is shorter in length and higher in frequency from S. albiventris (Additional file 1: Table S6).
Fig. 4.
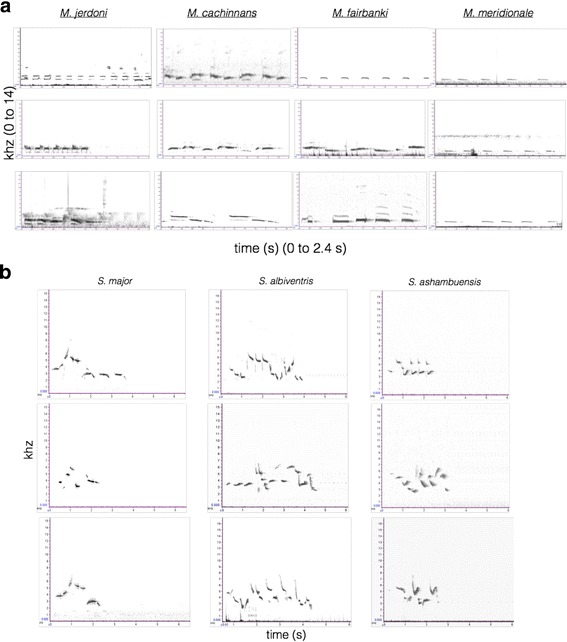
Spectrograms showing song variation across a) Montecincla and b) Sholicola species. Songs of three different individuals from each species are shown
Naming new taxa
As outlined above, the results of our analysis warrant new names for two genera and one species, in addition to elevating six subspecies to species level.
Western Ghats laughingthrushes
The laughingthrushes of the Western Ghats have been treated as part of the genus Garrulax Lesson, 1831 by Ali and Ripley [6] and as Strophocincla Wolters, 1981 by Rasmussen and Anderton [50], which was followed by others [22, 26]. However, in most global avian checklists, they are currently placed in Trochalopteron [8, 21] following phylogenetic insights [27]. Genus-level treatments in laughingthrushes have clearly been very unstable given extensive morphological variation and labile trait evolution in these birds. Our phylogenetic results reveal that all previous genus designations are inappropriate for Western Ghats Laughingthrushes and here we erect a new genus to recognize their distinctiveness.
Furthermore, we propose elevation of the four subspecies to full species level. This fits assertions by others: Rasmussen and Anderton [50] tentatively delimited two species (one north and the other south of the Palghat Gap with two subspecies each) but suggested that the four allopatric populations may each warrant species-level status with more evidence. Additionally, Praveen and Nameer [26] conducted an analysis of plumage differences and suggested that this complex was best represented as four distinct species. In line with our results of multiple character systems, we hereby propose elevation of these subspecies as four full allopatric species (Additional file 1: Figure S4). This is essentially a revival of the taxonomic treatment at the time of the original descriptions of these taxa.
Montecincla genus novum
Order Passeriformes: Family Leiothrichidae
Suggested common name: Chilappan
Type species: C. [=Crateropus] cachinnaus [sic] Jerdon, 1839 = Strophocincla cachinnans = Montecincla cachinnans comb. nov.
Additional species included: Garrulax (?) Jerdoni [sic] Blyth, 1851 = Strophocincla cachinnans jerdoni = Montecincla jerdoni comb. nov.; Trochalopteron Fairbanki Blanford, 1869 = Strophocincla fairbanki = Montecincla fairbanki comb. nov.; and Trochalopterum meridionale Blanford, 1880 = Strophocincla fairbanki meridionalis = Montecincla meridionalis comb. nov.
ZooBank Registry: urn:lsid:zoobank.org:act:88B21D13-2639-4BFD-8729-74308F609E4E
Diagnosis: Montecincla gen. nov. can be differentiated from other genera of traditional laughingthrushes - Garrulax; Ianthocincla Gould, 1835; Trochalopteron; and from Turdoides babblers by the combination of a prominent white supercilium, olive brown upperparts, small size, brown wings, rufous flanks and dark bill. Montecincla lacks the complex upperwing colouration (consisting of strong barring or conspicuous wing panels) of Actinodura Gould, 1836, and of Liocichla Swinhoe, 1877. Montecincla also lacks the slender greyish body and long tail of sibias of the genus Heterophasia Blyth, 1842. Laniellus Swainson, 1832 (=Crocias Temminck, 1836) has a diagnostic whitish underside with bold, streaked flanks that are absent in Montecincla. Minla Hodgson, 1837 is a genus of small birds (14cm) with a colourful reddish tail or rump, while Montecincla is larger (20cm) with a dull olive-brown tail and rump. Leiothrix Swainson, 1832, is characterized by a colourful bill and a yellow throat, contrasting with the black bill and greyish-white or black throats of Montecincla.
Description: Montecincla chilappans are medium-sized (~20cm) songbirds with rounded tails. Their upperparts are olive grey and they have a prominent white supercilium. The breast is greyish-white in three species and rufous in one (Table 2).
Etymology: Montecincla, of feminine gender, is a combination of Latin “mons” (gen. “montis”), with the meaning “mountain”, and Greek “kinklos”, denoting an unidentified type of songbird (often assumed to be a thrush). We chose this appropriate moniker because the genus is confined to the higher mountains of the Western Ghats. Our suggested vernacular name “Chilappan” stems from the local name of this genus (in Malayalam language), denoting their joyful cackling calls.
Comments: We suggest Banasura Chilappan as a common name for Montecincla jerdoni in recognition of Mt. Banasura, one of its strongholds and its type locality. Similarly, we suggest the common names Nilgiri Chilappan for Montecincla cachinnans, Palani Chilappan for Montecincla fairbanki, and Ashambu Chilappan for Montecincla meridionalis.
Western Ghats blue robins or shortwings
The species were historically placed in the genus Phoenicura Swainson, 1831, later in Callene Blyth, 1847, then in Brachypteryx Horsfield, 1821 and more recently in Myiomela. None of these generic names are appropriate given our phylogenetic analysis. We thus erect a new genus to recognize the distinctness of Western Ghats Blue Robins (also known as Western Ghats Shortwings). In addition, we recognize three distinct species in this genus, including one new, previously unrecognized species.
Sholicola genus novum
Order Passeriformes: family Muscicapidae
Suggested common name: Sholakili
Type species: Phoenicura major Jerdon, 1841 = Myiomela major = Sholicola major comb. nov.
Additional species included: Callene albiventris Blanford, 1868 = Myiomela albiventris = Sholicola albiventris comb. nov.; Sholicola ashambuensis Robin, Vishnudas, Rheindt, Gupta, Hooper, Ramakrishnan & Reddy (see below).
ZooBank Registry: urn:lsid:zoobank.org:act:66837381-635E-476C-9454-092BB03B0A91
Diagnosis: Sholicola is sexually monomorphic, thus differing from the following closely related and sexually dimorphic genera: Cyornis Blyth, 1843; Eumyias Cabanis, 1850; Niltava Hodgson, 1837; and Cyanoptila Blyth, 1847. Being largely blue, it differs from Anthipes Blyth, 1847, which comprises brown birds with distinctive white throats. Members of Sholicola are resident birds with no known seasonal migration as in members of Niltava and Cyanoptila. The three species of Sholicola are phylogenetically distant from the genera they were previously placed under (Brachypteryx, Callene, Phoenicura, Myiomela). They also differ from these genera in having a sexually monomorphic plumage dominated by blue colouration.
Description: Sholicola is a genus of terrestrial blue flycatchers confined to high altitude (above 1200m) forests in the Shola-grassland complexes of the Western Ghats. Their general plumage is dominated by blue. They have a bluish-white band above the black lores, a slightly curved bill tip, well-developed rictal bristles, a short, nearly square tail, and long tarsi. The birds are mainly restricted to the understory, rarely venturing above three meters and feeding on insects singly or in pairs. There is no known sexual dimorphism in plumage though morphometric differences in wing length are recognized [10].
Etymology: Sholicola, of masculine gender, is a combination of Shola (the local name for montane forests in the Western Ghats) and the suffix –cola (from the Latin verb “colere”), meaning “dweller". We also suggest the common English name Sholakili, where -kili is the local name for “bird”.
Description of a new species in the genus Sholicola:
Sholicola ashambuensis, species nova
English name: Ashambu Sholakili
Holotype: Trivandrum Museum of Natural History (TMNH) No. 725, collected by H.S. Fergusson on 3 May 1903 in the Chemunji Hills, Travencure (Travancore).
Etymology: The adjective ashambuensis denotes the species’ geographical locality, the Ashambu Hills of southern India.
ZooBank Registry: urn:lsid:zoobank.org:act:CB9F32BC-750B-48AA-8F87-C9727144019D
Diagnosis: The following characters can be used to diagnose Sholicola ashambuensis.
Despite being smaller in general body size, S. ashambuensis has a considerably larger white belly patch (~24mm in length) than S. albiventris (~19mm). Only one S. ashambuensis specimen was available for measurements.
S. ashambuensis is a smaller bird (Additional file 1: Figure S5) with a shorter tarsus and longer bill (Additional file 1: Table S7; Additional file 1: Figure S6) than S. albiventris based on an examination of 76 S. albiventris and 21 S. ashambuensis.
S. ashambuensis has a distinct song (Additional file 1: Figure S7) with a higher mean maximum frequency, but narrower song bandwidth, shorter notes and shorter song bouts than S. albiventris, based on 119 S. ashambuensis songs and 203 S. albiventris songs.
When compared side-by-side, S. ashambuensis is paler blue than S. albiventris.
In addition to these phenotypic diagnostic characters, S. ashambuensis forms a reciprocally monophyletic clade based on DNA data from about 25 individuals. Further population genetic data with 15 microsatellites and 218 individuals (including 17 of S. ashambuensis) also support significant genetic differentiation [51].
Description of holotype: A small, overall dark-blue flycatcher with a large white belly patch reaching to the vent, black lores, and a bluish-white band above the lores. The holotype measures: bill – 16mm, wing– 81mm, tail – 61mm, tarsus – 21mm.
Distribution: Ashambu hills south of Shenkottah gap, southern India, mostly above 1200m elevation (Additional file 1: Figure S4).
Comments: The subtle plumage differentiation and limited fieldwork in the Ashambu Hills, including an absence of systematic capture-based studies, are perhaps the reason why S. ashambuensis has gone taxonomically unrecognized until now. The only museum specimen to our knowledge, the holotype (described above), was re-discovered (by CKV) after being locked up for about 120 years in a drawer of the Travancore Museum. Other specimens thought to be from the southern range are missing (and perhaps destroyed). There are possibly two more specimens in NHMUK (Tring) based on geography, but these were not examined by the authors.
Discussion
Using a modern systematic approach, we uncovered two deeply divergent lineages in the Indian avifauna and provided the first evidence of in-situ avian radiations within the Indian subcontinent. These new genera are not only phylogenetically distant from the clades with which they were previously classified, but also constitute lineages that diverged early from their closest relatives in larger Asian clades, many of which were only recently identified themselves. Our study underscores the importance of continued systematic studies in untangling taxonomic confusions to better understand local, regional, and global patterns of diversification.
The two WG lineages we examined, previously known as laughingthrushes and shortwings, do not belong to either of these two groups. Our results corroborate previous studies [27, 32, 33] that showed laughingthrushes to be polyphyletic. The WG Laughingthrushes do not group with any of the other traditional laughingthrush clades including species placed within Trochalopteron or Strophocincla. Rather, they are in a distinct clade of their own that is sister to several genera traditionally not considered laughingthrushes. We introduced a new genus name, Montecincla, to highlight the phylogenetic and biological distinction of the Western Ghats Laughingthrushes as well as to avoid confusion with alternative names. The taxonomic history of the WG Shortwings has gone through a similar level of confusion. Due to behavioral and plumage traits, they were traditionally thought to be shortwings (genus Brachypteryx). The first phylogenetic analysis of the WG species found that they were not closely related to Brachypteryx [9]. Our analysis conclusively shows that WG Shortwings are actually flycatchers and sister to a newly discovered clade of several genera of Asian blue flycatchers, the Niltavinae.
We present the first evidence of in-situ bird radiations in the Western Ghats mountains. Our examination of various character data – genetic, song, plumage and morphometric – all point to considerable differentiation among populations that we propose to comprise four species in the genus Montecincla and three species in the genus Sholicola. The complete distributions of both species complexes lie along a 400 km latitudinal gradient comprising the highest elevations, the sky islands of the Western Ghats. Further intensive examinations in this region may reveal patterns of similar endemic radiations in other taxa.
The diversification of Montecincla and Sholicola from their respective sister taxa supports the classic model of species colonizing the WG from the Himalayas. The estimated divergence dates for corresponding species on either side of common barriers are similar across these two groups, which is compelling evidence for a vicariance model. We interpret our estimated dates using the best calibration currently available and in light of the corresponding known climatic events that may have led to these biogeographic patterns. Given that there is much disagreement about the age of bird diversification (see [52–54]), we acknowledge that different divergence dating methods and calibrations may provide alternative estimates of these dates. However, the relative timings of these divergences will remain the same. Our analyses provides a compelling demonstration that these two lineages have nearly identical divergence times, regardless of the exact ages, implying a common mechanism.
The coincidence in timing of colonization of the WG by Montecincla (11.57 Ma) and Sholicola (11.83 Ma) supports a scenario of initial range expansion during a wet, cool period in the mid-to-late Miocene [15, 55, 56] that allowed these cool-adapted lineages to disperse across the moist forests that covered the Indian subcontinent followed by subsequent vicariance as the subcontinent became drier and more seasonal in the late Miocene [56–58], which likely led to local extinction in the central region and current isolation in the WG. Intriguingly, another species of laughingthrush found in the Western Ghats, Ianthocincla [Dryonastes] delesserti, represents a more recent colonization into this region (see Fig. 1a).
Diversification within the WG in the Pliocene and Pleistocene was likely driven by climatic events that expanded and contracted the cool montane forest in southwestern India. Within the WG, both lineages show similar divergence dates (4.7 and 4.33 Ma) across the Palghat Gap. This break in the mountain chain is a major habitat barrier for many taxa [59–61]. It is the widest and deepest valley in the range and is thought to be the result of an ancient geological fissure (~500 Myr old) [9]. The tight correspondence of divergence dates in Montecincla and Sholicola across this now dry gap indicates that suitable wet, cool habitat, which was needed for these taxa to disperse, likely only lasted for a brief period in the Pliocene. Divergences across narrower gaps are not consistent across these two lineages, or several other bird species [60]. Climatic fluctuations in the Pleistocene were more numerous and variable in strength and duration [58, 62], perhaps leading to more individual responses by species across the narrow valleys within the WG. A more thorough survey of endemic taxa and their diversification history is crucial for a better understanding of the evolutionary history and assembly of this unique ecosystem.
Conclusions
The discovery of two independent endemic bird radiations in the Shola Sky Islands highlights the evolutionary role that such habitats can play in the diversification of lineages and the need for additional systematic studies that can potentially find other taxa with similar patterns. An important result from our findings is the dramatic increase in the biodiversity inventory of the WG, from the previously recognized three species to seven endemic species, each with much narrower ranges. Tropical sky island species are thought to be the most susceptible to anthropogenic climate change [4], and with 50% of Shola habitat in the WG already lost [5], these findings provide a much needed impetus for conservation [63]. With the Western Ghats continuing to lose between 0.57%–0.91% [64, 65] of their forest habitats each year, endemic species face extraordinary conservation challenges. Studies such as this not only clarify the taxonomic and phylogenetic information needed to quantify biodiversity but also urge the need to assess the possible responses of these species to anthropogenic climate change.
Acknowledgements
We thank Forest Departments of Kerala and Tamil Nadu; for exceptional support, we specifically thank - Chief Wildlife Wardens of Kerala - R. Rajaraja Varma, V. Gopinathan, G. Harikumar, PCCF Kerala T.M. Manoharan, Bransdon Corrie, former PCCF Tamil Nadu R. Sunder Raju, DCF K.I. Pradeep Kumar, ACF Santhosh Kumar, Senior Wildlife Assistant Reney R. Pillai; Abhilash Babu, Sahas Barve, Ravi Kiran, Chetana Purushotham, Anusha Shankar, Sriranjini Swaminathan for field assistance; Chetana Purushotham and Suma M. Shirley for song analyses; Joli Borah, Himanshu Chattani, Aleeson E., Shilpa M., Jyothi Nair, Abhinav Sur, Nelum Wickramasinghe for lab assistance; Tanya Balcar, Sumin George, Reney R. Pillai, Suhel Quader, Prathim Roy, Robert Stewart, for various support; H.S. Sudhira and Gubbi Labs for help with maps; Robin Abraham, Edward Dickinson, K.P. Dinesh, Praveen J., Rajah Jayapal, Muhammed Jafer Palot, Aasheesh Pittie, Pamela Rasmussen, C. Sasikumar, Anindya Sinha, Richard Schodde, for discussion on nomenclature and taxonomy; Ishan Aggarwal, John Bates, Balaji Chattopadhyay, Nishma Dahal, K.P. Dinesh, Varad Giri, Peter Makovicky, Nandini Rajamani, Vivek Ramachandran, Krishnapriya Tamma and S.P. Vijaykumar for comments on the manuscript; Prasenjeet Yadav for photographs of the museum birds; Maya Ramaswamy for bird illustrations; S. Abu at Trivandrum Museum of Natural History; M. Adams at NHMUK; Vithoba Hegde, Rahul Khot, Deepak Apte at BNHS. This manuscript also greatly benefited from submission to Axios Review including comments from Dieter Thomas Tietze, George Sangster, Jon Fjeldså, Jason Weir, and an anonymous reviewer. We used color-blind safe colors from www.colorbrewer2.org where possible.
Funding
The project was supported by a National Geographic Society Research and Exploration Grant to VVR; Indian DAE, NCBS, Ramanujan Fellowship to UR; U.S. National Science Foundation (DEB-0962078; DEB-1457624) to SR. FER was supported by Singaporean Ministry of Education Tier I grants (WBS R-154-000-570-133; WBS R-154-000-658-112).
Availability of data and materials
The datasets generated during and/or analysed for the current study are available in the GENBANK repository, Genbank accessions F234391-F234402. Phylogenetic alignments deposited into TreeBASE: XXX (http://purl.org/phylo/treebase/phylows/study/TB2:S20458).
Authors’ contributions
VVR, CKV, SR conceived of the study; VVR, PG carried out the molecular lab work; VVR, CKV, PG, DH, FER, SR analyzed the data; VVR, CKV, SR, UR coordinated the study; VVR, CKV, FER, SR drafted the manuscript; CKV, VVR carried out the field work. All authors gave final approval for publication.
Competing interests
The authors declare that they have no competing interests.
Consent for publication
Not applicable.
Ethics approval
Permission to handle birds and collect blood samples was granted by the Forest Departments of Kerala (WL10-1647/2011) and Tamil Nadu (WL5/8844/2011); NCBS institutional animal ethics clearances were obtained prior to the study.
Additional file
Sample collection sites across the sky islands of the Western Ghats. Figure S2. Ancestral area analysis using Lagrange for Montecincla. Area codes for terminal taxa are A-Africa, B-peninsular India, C-Himalayas, D-Southeast Asia, E-China, F-Sundaland, G-Philippines, and H-Assam. The most likely ancestral range reconstruction for relevant nodes (in red) are shown. Figure S3. Ancestral area analysis using Lagrange for Sholicola. Area codes for terminal taxa are I-Peninsular India, H-Himalayas, A-Southeast Asia, D-Eurasia, E-Australasia F- Africa + Southwest Asia, C-China, S-Sundaland, P-Philippines, E-Eurasia, and N-Australasia/New World. The most likely ancestral range reconstruction for relevant nodes (in red) are shown with their relative probabilities. Figure S4. Map showing the species distributions of Montecincla chilappans and Sholicola sholakillis. Figure S5. Photographs of Sholicola ashambuensis sp. nov. a) holotype in the Trivandrum Museum of Natural History; b) in comparison with other members of Sholicola: S. ashambuensis (right), with larger white belly contrasted with S. albiventris (middle), S. major (left). Figure S6. a) Discriminant Function Analysis of morphometric differences between Sholicola ashambuensis and S. albiventris based on 97 individuals shows 79.4% accurate classification and significant differences in the means (non overlapping 95% confidence interval circles); b) Box plots showing morphometric differences in bill and tarsus lengths in Sholicola ashambuensis and S. albiventris. Figure S7. Discriminant Function Analysis of Sholicola ashambuensis and S. albiventris songs based on 217 song recordings shows 80% accurate classification and significant differences in the means (non overlapping 95% confidence interval circles). Table S1: Samples used to reconstruct Montecincla phylogeny. Table S2: Samples used to reconstruct Sholicola phylogeny. Table S3: Morphometric variation in Montecincla species. Table S4: Morphometric variation in Sholicola species. Table S5: Song variation in Montecincla species in the Western Ghats Sky Islands. Table S6: Song variation in Sholicola species in the Western Ghats Sky Islands. Table S7: Comparison of morphometric measurements of Sholicola ashambuensis and S. albiventris. (PDF 3728 kb)
Contributor Information
V.V. Robin, Email: robinvijayan@gmail.com
C. K. Vishnudas, Email: ck.vishnudas@gmail.com
Pooja Gupta, Email: pooja.1923@gmail.com.
Frank E. Rheindt, Email: dbsrfe@nus.edu.sg
Daniel M. Hooper, Email: dhooper1@uchicago.edu
Uma Ramakrishnan, Email: uramakri@ncbs.res.in.
Sushma Reddy, Email: sreddy6@luc.edu.
References
- 1.Myers N, Mittermeier RA, Mittermeier CG, da Fonseca G, Kent J. Biodiversity hotspots for conservation priorities. Nature. 2000;403:853–858. doi: 10.1038/35002501. [DOI] [PubMed] [Google Scholar]
- 2.Fjeldså J, Bowie RCK, Rahbek C. The role of mountain ranges in the diversification of birds. Annu Rev Ecol Evol Syst. 2012;43:249–265. doi: 10.1146/annurev-ecolsys-102710-145113. [DOI] [Google Scholar]
- 3.Reddy S. What's missing from avian global diversification analyses? Mol Phylogenet Evol. 2014;77:159–165. doi: 10.1016/j.ympev.2014.04.023. [DOI] [PubMed] [Google Scholar]
- 4.Freeman BG, Class Freeman AM. Rapid upslope shifts in New Guinean birds illustrate strong distributional responses of tropical montane species to global warming. Proc Natl Acad Sci U S A. 2014;111:4490–4494. doi: 10.1073/pnas.1318190111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Sukumar R, Suresh HS, Ramesh R. Climate change and its impact on tropical montane ecosystems in southern India. J Biogeogr. 1995;22:533–536. doi: 10.2307/2845951. [DOI] [Google Scholar]
- 6.Ali S, Ripley SD. Handbook of the Birds of India and Pakistan, Together with those of Nepal, Sikkim, Bhutan and Ceylon. Bombay: Oxford University Press; 1978. [Google Scholar]
- 7.Rasmussen P, Anderton JC. Birds of South Asia: The Ripley Guide. 2nd Edition. 2 vols. Barcelona: Lynx Edicions; 2012. [Google Scholar]
- 8.Clements JF, Schulenberg TS, Iliff MJ, Robertson D, Fredericks TA, Sullivan BL, et al. The eBird/Clements checklist of birds of the world: v2015. 2015. Downloaded from http://www.birds.cornell.edu/clementschecklist/. Accessed 10 Aug 2015.
- 9.Robin VV, Sinha A, Ramakrishnan U. Ancient geographical gaps and paleo-climate shape the phylogeography of an endemic bird in the sky islands of southern India. PLoS ONE. 2010;5:e13321. doi: 10.1371/journal.pone.0013321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Widdowson M, Cox K. Uplift and erosional history of the Deccan Traps, India: Evidence from laterites and drainage patterns of the Western Ghats and Konkan Coast. Earth Planet Sci Lett. 1996;137:57–69. doi: 10.1016/0012-821X(95)00211-T. [DOI] [Google Scholar]
- 11.Gunnell Y, Gallagher K, Carter A, Widdowson M, Hurford AJ. Denudation history of the continental margin of western peninsular India since the early Mesozoic – reconciling apatite fission-track data with geomorphology. Earth Planet Sci Lett. 2003;215:187–201. doi: 10.1016/S0012-821X(03)00380-7. [DOI] [Google Scholar]
- 12.Hora SL. Satpura hypothesis of the distribution of the Malayan fauna and flora to peninsular India. Proc Nat Instit Sci India. 1949;15:309–314. [Google Scholar]
- 13.Islam MA. Satpura Hypothesis and the distribution of laughing thrushes Garrulax Lesson of India. J Bomb Nat Hist Soc. 1990;86:318–322. [Google Scholar]
- 14.Ali S. The Satpura trend as an ornithogeographical highway. Proc Nat Instit Sci India. 1949;15:379–386. [Google Scholar]
- 15.Karanth K. Evolution of disjunct distributions among wet-zone species of the Indian subcontinent: Testing various hypotheses using a phylogenetic approach. Curr Sci. 2003;85:1276–1283. [Google Scholar]
- 16.Ripley SD, Beehler BM. Patterns of speciation in Indian birds. J Biogeogr. 1990;17:639–648. doi: 10.2307/2845145. [DOI] [Google Scholar]
- 17.Patnaik R. Fossil murine rodents as ancient monsoon indicators of the Indian subcontinent. Quatern Int. 2011;229:94–104. doi: 10.1016/j.quaint.2010.04.005. [DOI] [Google Scholar]
- 18.Robin VV, Nandini R. Shola habitats on sky islands: status of research on montane forests and grasslands in southern India. Curr Sci. 2012;103:1427–1437. [Google Scholar]
- 19.Daniels RJR, Joshi NV, Gadgil M. On the relationship between bird and woody plant species diversity in the Uttara Kannada district of south India. Proc Natl Acad Sci U S A. 1992;89:5311–5315. doi: 10.1073/pnas.89.12.5311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ripley SD. Avian relicts and double invasions in peninsular India and Ceylon. Evolution. 1949;3:150–159. doi: 10.2307/2405549. [DOI] [Google Scholar]
- 21.Dickinson EC, Christidis L. The Howard and Moore Complete Checklist of the Birds of the World Fourth Edition, Volume 2: Passerines. Dickinson EC, Christidis L, editors. Eastbourne, UK: Aves Press; 2014.
- 22.Collar NJ, Robson C. Family Timaliidae (Babblers) In: del Hoyo J, Elliott A, Christie DA, editors. Handbook of the Birds of the World, Vol. 12. Barcelona: Lynx Edicions; 2007. pp. 70–291. [Google Scholar]
- 23.Collar N, de Juana E, Sharpe CJ. White-bellied Blue Robin (Myiomela albiventris). In: del Hoyo J, Elliott A, Sargatal J, Christie DA, de Juana E, editors. Handbook of the Birds of the World Alive. Barcelona: Lynx Edicions. 2016. (retrieved from http://www.hbw.com/node/58510 on 7 March 2016).
- 24.Rasmussen P. Biogeographic and conservation implications of revised species limits and distributions of South Asian birds. Zool Med Leiden. 2005;79-3:137–146. [Google Scholar]
- 25.Sibley CG, Monroe BJ. Distribution and Taxonomy of Birds of the World. New Haven: Yale University Press; 1991. [Google Scholar]
- 26.Praveen J, Nameer PO. Strophocincla Laughingthrushes of South India: a case for allopatric speciation and impact on their conservation. J Bomb Nat Hist Soc. 2012;109:46–52. [Google Scholar]
- 27.Moyle RG, Andersen MJ, Oliveros CH, Steinheimer FD, Reddy S. Phylogeny and biogeography of the core babblers (Aves: Timaliidae) Syst Biol. 2012;61:631–651. doi: 10.1093/sysbio/sys027. [DOI] [PubMed] [Google Scholar]
- 28.Sangster G, Alström P, Forsmark E, Olsson U. Multi-locus phylogenetic analysis of Old World chats and flycatchers reveals extensive paraphyly at family, subfamily and genus level (Aves: Muscicapidae) Mol Phylogenet Evol. 2010;57:380–392. doi: 10.1016/j.ympev.2010.07.008. [DOI] [PubMed] [Google Scholar]
- 29.Price TD, Hooper DM, Buchanan CD, Johansson US, Tietze DT, Alström P, et al. Niche filling slows the diversification of Himalayan songbirds. Nature. 2014;509:222–225. doi: 10.1038/nature13272. [DOI] [PubMed] [Google Scholar]
- 30.Kimball R, Braun EL, Barker FK, Bowie RCK, Braun MJ, Chojnowski JL, et al. A well-tested set of primers to amplify regions spread across the avian genome. Mol Phylogenet Evol. 2009;50:654–660. doi: 10.1016/j.ympev.2008.11.018. [DOI] [PubMed] [Google Scholar]
- 31.Reddy S, Driskell AC, Rabosky DL, Hackett SJ, Schulenberg TS. Diversification and the adaptive radiation of the vangas of Madagascar. Proc R Soc B. 2012;279:2062–2071. doi: 10.1098/rspb.2011.2380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Luo X, Qu YH, Han LX, Li SH, Lei FM. A phylogenetic analysis of laughingthrushes (Timaliidae: Garrulax) and allies based on mitochondrial and nuclear DNA sequences. Zool Scripta. 2009;38:9–22. doi: 10.1111/j.1463-6409.2008.00355.x. [DOI] [Google Scholar]
- 33.Gelang M, Cibois A, Pasquet E, Olsson U, Alström P, Ericson PGP. Phylogeny of babblers (Aves, Passeriformes): major lineages, family limits and classification. Zool Scripta. 2009;38:225–236. doi: 10.1111/j.1463-6409.2008.00374.x. [DOI] [Google Scholar]
- 34.Lanfear R, Calcott B, Ho SYW, Guindon S. PartitionFinder: Combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol. 2012;29:1695–1701. doi: 10.1093/molbev/mss020. [DOI] [PubMed] [Google Scholar]
- 35.Zuccon D, Ericson PGP. A multi-gene phylogeny disentangles the chat-flycatcher complex (Aves: Muscicapidae) Zool Scripta. 2010;39:213–224. doi: 10.1111/j.1463-6409.2010.00423.x. [DOI] [Google Scholar]
- 36.Stamatakis A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics. 2006;22:2688–2690. doi: 10.1093/bioinformatics/btl446. [DOI] [PubMed] [Google Scholar]
- 37.Swofford DL. PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods) Sunderland, Massachusetts: Version 4. Sinauer Associates; 2003. [Google Scholar]
- 38.Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Hohna S, et al. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012;61:539–542. doi: 10.1093/sysbio/sys029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Rambaut A, Suchard MA, Xie D, Drummond AJ. Tracer v1.6. 2014. Available from: http://tree.bio.ed.ac.uk/software/tracer/.
- 40.Wilgenbusch JC, Warren DL, Swofford DL. AWTY: a system for graphical exploration of MCMC convergence in Bayesian phylogenetic inference. 2004. Available from: http://ceb.csit.fsu.edu/awty. [DOI] [PubMed]
- 41.Drummond AJ, Suchard MA, Xie D, Rambaut A. Bayesian Phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 2012;29:1969–1973. doi: 10.1093/molbev/mss075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ree RH, Smith SA. Maximum likelihood inference of geographic range evolution by dispersal, local extinction, and cladogenesis. Syst Biol. 2008;57:4–14. doi: 10.1080/10635150701883881. [DOI] [PubMed] [Google Scholar]
- 43.Yang Z. The BPP, program for species tree estimation and species delimitation. Curr Zool. 2015;61:854–865. doi: 10.1093/czoolo/61.5.854. [DOI] [Google Scholar]
- 44.De Beer S, Lockwood G, Raijmakers JHFA, Raijmakers JMH, Scott WA, Oschadleus HD, et al. SAFRING bird ringing manual. Cape Town, South Africa: University of Cape Town; 2000. [Google Scholar]
- 45.Ralph CJ, Geupel GR, Pyle P, Martin TE, DeSante DF. Handbook of Field Methods for Monitoring Landbirds. Gen. Tech. Rep. PSW-GTR-144-www. Albany, CA: Pacific Southwest Research Station, Forest Service, U.S. Department of Agriculture; 1993. [Google Scholar]
- 46.Lougheed SC, Arnold TW, Bailey RC. Measurement error of external and skeletal variables in birds and its effect on principal components. Auk. 1991;108:432–436. [Google Scholar]
- 47.Robin VV, Katti M, Purushotham C, Sancheti A, Sinha A. Singing in the sky: song variation in an endemic bird on the sky islands of southern India. Anim Behav. 2011;82:513–520. doi: 10.1016/j.anbehav.2011.05.032. [DOI] [Google Scholar]
- 48.Bioacoustics Research Program. Raven Pro: Interactive Sound Analysis Software (Version 1.3) [Computer software]. Ithaca, NY: The Cornell Lab of Ornithology. 2008. Available from http://www.birds.cornell.edu/raven.
- 49.Purushotham CB, Robin VV. Sky island bird populations isolated by ancient genetic barriers are characterized by different song traits than those isolated by recent deforestation. Ecol Evol. 2016;20:7334–7343. doi: 10.1002/ece3.2475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Rasmussen PC, Anderton JC. Birds of South Asia - The Ripley Guide. Lynx Edicions: Barcelona; 2005. [Google Scholar]
- 51.Robin VV, Gupta P, Thatte P, Ramakrishnan U. Islands within islands: two montane palaeo-endemic birds impacted by recent anthropogenic fragmentation. Mol Ecol. 2015;24:3572–3584. doi: 10.1111/mec.13266. [DOI] [PubMed] [Google Scholar]
- 52.Jarvis ED, Mirarab S, Aberer AJ, Li B, Houde P, Li C. Whole-genome analyses resolve early branches in the tree of life of modern birds. Science. 2014;346:1320–1331. doi: 10.1126/science.1253451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Claramunt S, Cracraft J. A new time tree reveals Earth history's imprint on the evolution of modern birds. Sci Adv. 2015;1:e1501005–5. doi: 10.1126/sciadv.1501005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Prum RO, Berv JS, Dornburg A, Field DJ, Townsend JP, Lemmon EM, et al. A comprehensive phylogeny of birds (Aves) using targeted next-generation DNA sequencing. Nature. 2015;526:569–573. doi: 10.1038/nature15697. [DOI] [PubMed] [Google Scholar]
- 55.Potter PE, Szatmari P. Global Miocene tectonics and the modern world. Earth-Sci Rev. 2009;96:279–295. doi: 10.1016/j.earscirev.2009.07.003. [DOI] [Google Scholar]
- 56.Pound MJ, Haywood AM, Salzmann U, Riding JB. Global vegetation dynamics and latitudinal temperature gradients during the mid to late Miocene (15.97–5.33 Ma) Earth Sci Rev. 2012;112:1–22. doi: 10.1016/j.earscirev.2012.02.005. [DOI] [Google Scholar]
- 57.Sanyal P, Bhattacharya SK, Kumar R, Ghosh SK, Sangode SJ. Mio–Pliocene monsoonal record from Himalayan foreland basin (Indian Siwalik) and its relation to vegetational change. Palaeogeogr Palaeoclimatol Palaeoecol. 2004;205:23–41. doi: 10.1016/j.palaeo.2003.11.013. [DOI] [Google Scholar]
- 58.Zhisheng A, Guoxiong W, Jianping L, Youbin S, Yimin L, Weijian Z, et al. Global Monsoon Dynamics and Climate Change. Annu Rev Earth Planet Sci. 2015;43:29–77. doi: 10.1146/annurev-earth-060313-054623. [DOI] [Google Scholar]
- 59.Vidya TNC, Fernando P, Melnick DJ, Sukumar R. Population differentiation within and among Asian elephant (Elephas maximus) populations in southern India. Heredity. 2004;94:71–80. doi: 10.1038/sj.hdy.6800568. [DOI] [PubMed] [Google Scholar]
- 60.Robin VV, Vishnudas CK, Gupta P, Ramakrishnan U. Deep and wide valleys drive nested phylogeographic patterns across a montane bird community. Proc R Soc B. 2015;282:20150861–20150867. doi: 10.1098/rspb.2015.0861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Van Bocxlaer I, Biju SD, Willaert B, Giri VB, Shouche YS, Bossuyt F. Mountain-associated clade endemism in an ancient frog family (Nyctibatrachidae) on the Indian subcontinent. Mol Phylogenet Evol. 2012;62:839–847. doi: 10.1016/j.ympev.2011.11.027. [DOI] [PubMed] [Google Scholar]
- 62.Clift PD, Hodges KV, Heslop D, Hannigan R, Van Long H, Calves G. Correlation of Himalayan exhumation rates and Asian monsoon intensity. Nat Geosci. 2008;1:875–880. doi: 10.1038/ngeo351. [DOI] [Google Scholar]
- 63.Padma TV. India faces uphill battle on biodiversity. Nature. 2013;504(7479):200. doi: 10.1038/504200a. [DOI] [PubMed] [Google Scholar]
- 64.Menon S, Bawa KS. Applications of geographic information systems, remote-sensing, and a landscape ecology approach to biodiversity conservation in the Western Ghats. Curr Sci. 1997;73:134–145. [Google Scholar]
- 65.Prasad SN, Vijayan L, Balachandran S. Conservation planning for the Western Ghats of Kerala: I. A GIS approach for location of biodiversity hot spots. Curr Sci. 1998;75:211–219. [Google Scholar]
- 66.Molnar P. Mio-Pliocene growth of the Tibetan Plateau and evolution of East Asian climate. Palaeontol Electron. 2005;8:2A. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets generated during and/or analysed for the current study are available in the GENBANK repository, Genbank accessions F234391-F234402. Phylogenetic alignments deposited into TreeBASE: XXX (http://purl.org/phylo/treebase/phylows/study/TB2:S20458).


