Abstract
To identify functionally important amino acids in the two zinc fingers of transcription factor ADR1 [alcohol dehydrogenase (ADH) II synthesis regulator], oligonucleotide-directed mutagenesis was used to substitute alanine for the original amino acid at each position in both fingers. The effects of these mutations on DNA binding and thermal stability of ADR1 in vitro and on activation of ADH2 expression in vivo were measured. The DNA binding activity was remarkably heatstable. Amino acids that are candidates for DNA contact sites were identified in the finger-tip and alpha-helical region of each finger, three in the first finger and two in the second. Unexpectedly, an acidic residue in the first finger was essential for transactivation, but its replacement by alanine had no effect on DNA binding. Substitution at several highly conserved positions did not affect ADR1 functions. The ADR1 zinc fingers appear to make relatively few energetically significant contacts to DNA, perhaps as few as three in the first finger and one in the second.
Full text
PDF

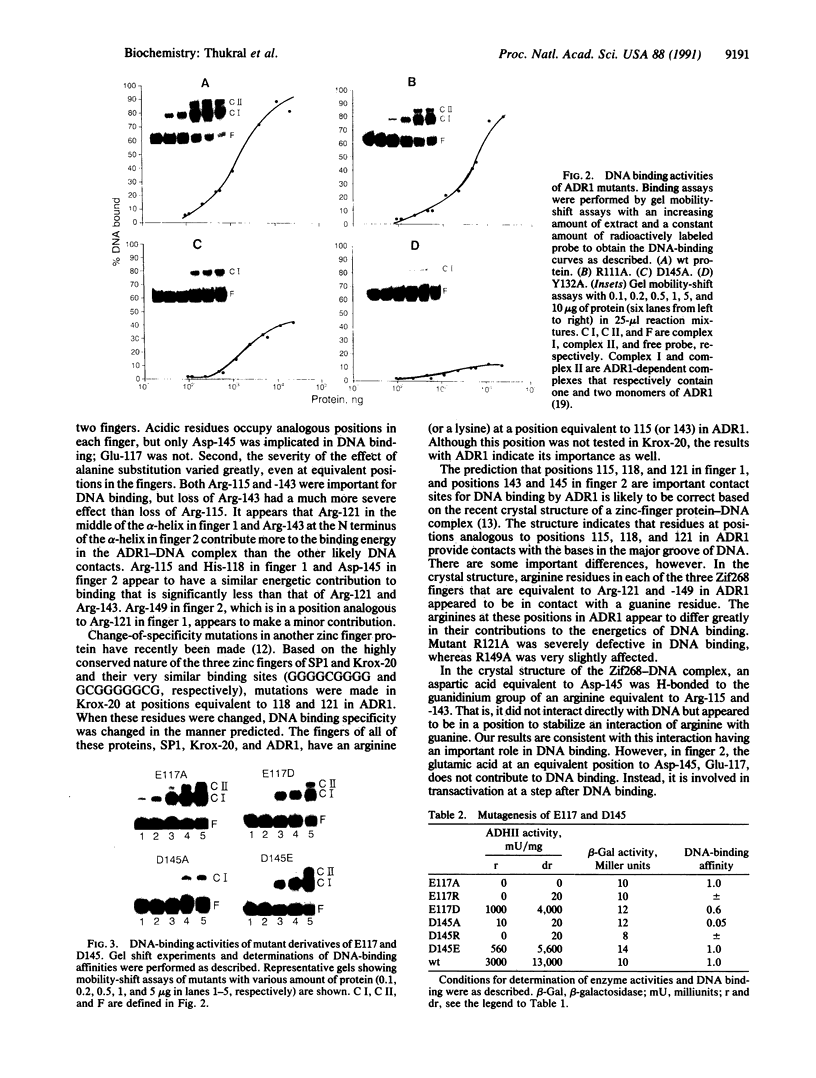

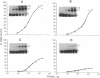
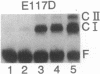
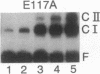
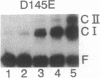
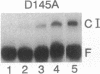
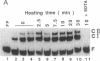
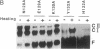
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Berg J. M. Zinc finger domains: hypotheses and current knowledge. Annu Rev Biophys Biophys Chem. 1990;19:405–421. doi: 10.1146/annurev.bb.19.060190.002201. [DOI] [PubMed] [Google Scholar]
- Blumberg H., Eisen A., Sledziewski A., Bader D., Young E. T. Two zinc fingers of a yeast regulatory protein shown by genetic evidence to be essential for its function. 1987 Jul 30-Aug 5Nature. 328(6129):443–445. doi: 10.1038/328443a0. [DOI] [PubMed] [Google Scholar]
- Blumberg H., Hartshorne T. A., Young E. T. Regulation of expression and activity of the yeast transcription factor ADR1. Mol Cell Biol. 1988 May;8(5):1868–1876. doi: 10.1128/mcb.8.5.1868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown R. S., Argos P. Fingers and helices. Nature. 1986 Nov 20;324(6094):215–215. doi: 10.1038/324215a0. [DOI] [PubMed] [Google Scholar]
- Brown R. S., Sander C., Argos P. The primary structure of transcription factor TFIIIA has 12 consecutive repeats. FEBS Lett. 1985 Jul 8;186(2):271–274. doi: 10.1016/0014-5793(85)80723-7. [DOI] [PubMed] [Google Scholar]
- Cherry J. R., Johnson T. R., Dollard C., Shuster J. R., Denis C. L. Cyclic AMP-dependent protein kinase phosphorylates and inactivates the yeast transcriptional activator ADR1. Cell. 1989 Feb 10;56(3):409–419. doi: 10.1016/0092-8674(89)90244-4. [DOI] [PubMed] [Google Scholar]
- Cunningham B. C., Wells J. A. High-resolution epitope mapping of hGH-receptor interactions by alanine-scanning mutagenesis. Science. 1989 Jun 2;244(4908):1081–1085. doi: 10.1126/science.2471267. [DOI] [PubMed] [Google Scholar]
- Eisen A., Taylor W. E., Blumberg H., Young E. T. The yeast regulatory protein ADR1 binds in a zinc-dependent manner to the upstream activating sequence of ADH2. Mol Cell Biol. 1988 Oct;8(10):4552–4556. doi: 10.1128/mcb.8.10.4552. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gibson T. J., Postma J. P., Brown R. S., Argos P. A model for the tertiary structure of the 28 residue DNA-binding motif ('zinc finger') common to many eukaryotic transcriptional regulatory proteins. Protein Eng. 1988 Sep;2(3):209–218. doi: 10.1093/protein/2.3.209. [DOI] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim K. S., Guarente L. Mutations that alter transcriptional activation but not DNA binding in the zinc finger of yeast activator HAPI. Nature. 1989 Nov 9;342(6246):200–203. doi: 10.1038/342200a0. [DOI] [PubMed] [Google Scholar]
- Klevit R. E., Herriott J. R., Horvath S. J. Solution structure of a zinc finger domain of yeast ADR1. Proteins. 1990;7(3):215–226. doi: 10.1002/prot.340070303. [DOI] [PubMed] [Google Scholar]
- Kunkel T. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Proc Natl Acad Sci U S A. 1985 Jan;82(2):488–492. doi: 10.1073/pnas.82.2.488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee M. S., Gippert G. P., Soman K. V., Case D. A., Wright P. E. Three-dimensional solution structure of a single zinc finger DNA-binding domain. Science. 1989 Aug 11;245(4918):635–637. doi: 10.1126/science.2503871. [DOI] [PubMed] [Google Scholar]
- Miller J., McLachlan A. D., Klug A. Repetitive zinc-binding domains in the protein transcription factor IIIA from Xenopus oocytes. EMBO J. 1985 Jun;4(6):1609–1614. doi: 10.1002/j.1460-2075.1985.tb03825.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nardelli J., Gibson T. J., Vesque C., Charnay P. Base sequence discrimination by zinc-finger DNA-binding domains. Nature. 1991 Jan 10;349(6305):175–178. doi: 10.1038/349175a0. [DOI] [PubMed] [Google Scholar]
- Omichinski J. G., Clore G. M., Appella E., Sakaguchi K., Gronenborn A. M. High-resolution three-dimensional structure of a single zinc finger from a human enhancer binding protein in solution. Biochemistry. 1990 Oct 9;29(40):9324–9334. doi: 10.1021/bi00492a004. [DOI] [PubMed] [Google Scholar]
- Pavletich N. P., Pabo C. O. Zinc finger-DNA recognition: crystal structure of a Zif268-DNA complex at 2.1 A. Science. 1991 May 10;252(5007):809–817. doi: 10.1126/science.2028256. [DOI] [PubMed] [Google Scholar]
- Párraga G., Horvath S. J., Eisen A., Taylor W. E., Hood L., Young E. T., Klevit R. E. Zinc-dependent structure of a single-finger domain of yeast ADR1. Science. 1988 Sep 16;241(4872):1489–1492. doi: 10.1126/science.3047872. [DOI] [PubMed] [Google Scholar]
- Párraga G., Horvath S., Hood L., Young E. T., Klevit R. E. Spectroscopic studies of wild-type and mutant "zinc finger" peptides: determinants of domain folding and structure. Proc Natl Acad Sci U S A. 1990 Jan;87(1):137–141. doi: 10.1073/pnas.87.1.137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schena M., Freedman L. P., Yamamoto K. R. Mutations in the glucocorticoid receptor zinc finger region that distinguish interdigitated DNA binding and transcriptional enhancement activities. Genes Dev. 1989 Oct;3(10):1590–1601. doi: 10.1101/gad.3.10.1590. [DOI] [PubMed] [Google Scholar]
- Shuster J., Yu J., Cox D., Chan R. V., Smith M., Young E. ADR1-mediated regulation of ADH2 requires an inverted repeat sequence. Mol Cell Biol. 1986 Jun;6(6):1894–1902. doi: 10.1128/mcb.6.6.1894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor W. E., Young E. T. cAMP-dependent phosphorylation and inactivation of yeast transcription factor ADR1 does not affect DNA binding. Proc Natl Acad Sci U S A. 1990 Jun;87(11):4098–4102. doi: 10.1073/pnas.87.11.4098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thukral S. K., Eisen A., Young E. T. Two monomers of yeast transcription factor ADR1 bind a palindromic sequence symmetrically to activate ADH2 expression. Mol Cell Biol. 1991 Mar;11(3):1566–1577. doi: 10.1128/mcb.11.3.1566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu R. X., Horvath S. J., Klevit R. E. ADR1a, a zinc finger peptide, exists in two folded conformations. Biochemistry. 1991 Apr 9;30(14):3365–3371. doi: 10.1021/bi00228a003. [DOI] [PubMed] [Google Scholar]
- Yu J., Donoviel M. S., Young E. T. Adjacent upstream activation sequence elements synergistically regulate transcription of ADH2 in Saccharomyces cerevisiae. Mol Cell Biol. 1989 Jan;9(1):34–42. doi: 10.1128/mcb.9.1.34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zoller M. J., Smith M. Oligonucleotide-directed mutagenesis of DNA fragments cloned into M13 vectors. Methods Enzymol. 1983;100:468–500. doi: 10.1016/0076-6879(83)00074-9. [DOI] [PubMed] [Google Scholar]