Abstract
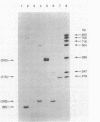
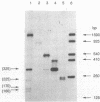
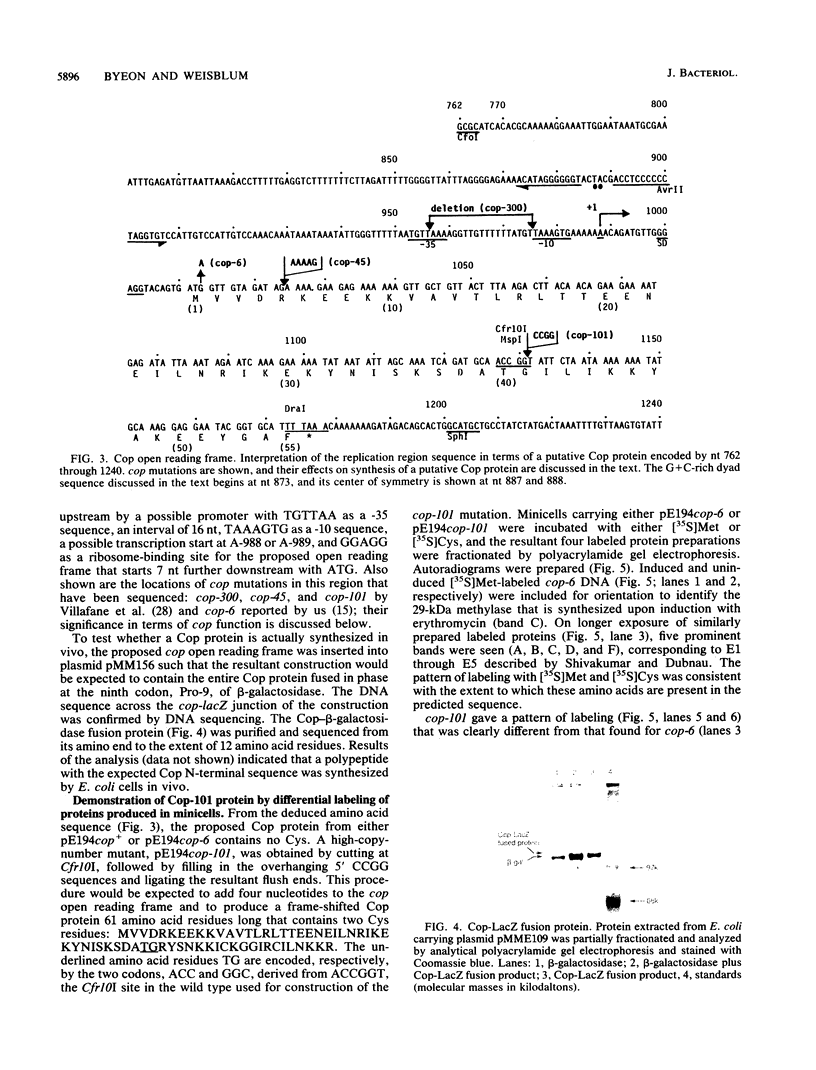
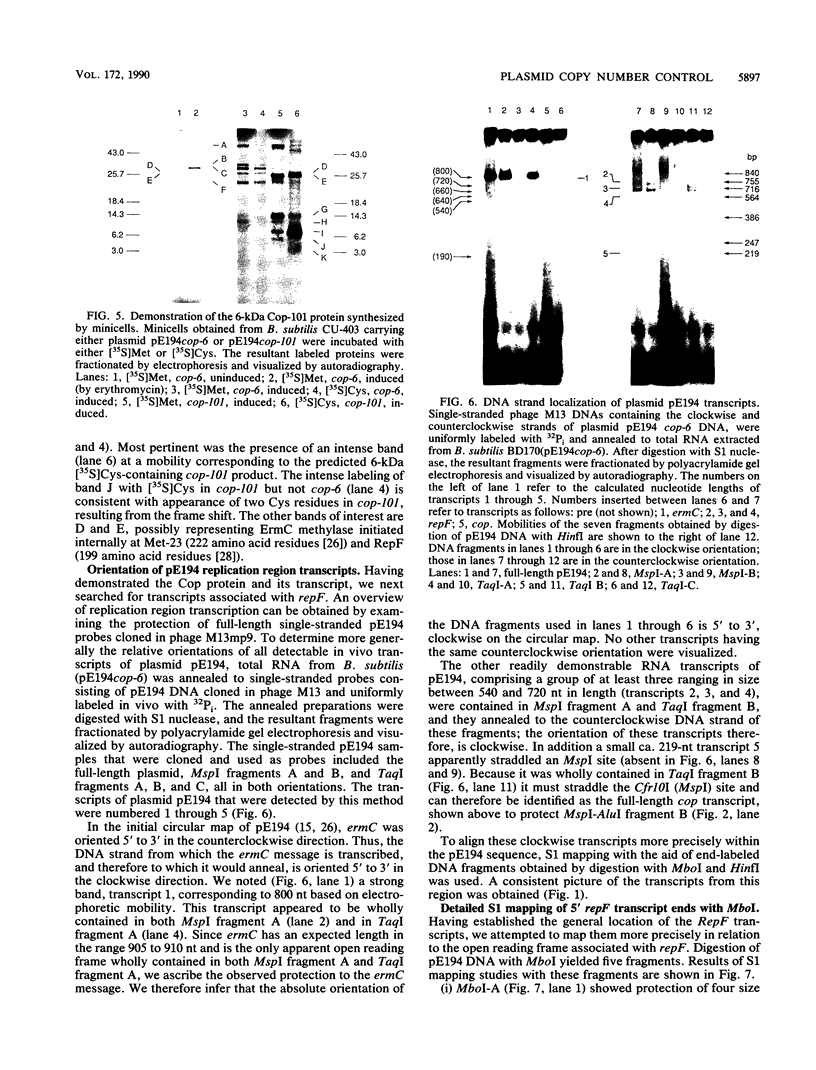
In vivo transcription of the replication region of plasmid pE194 yeidls two classes of mRNAs that encode Cop and RepF proteins, respectively. These transcripts are oriented 5' to 3' exclusively in the clockwise direction on the standard map. The cop region contains an open reading frame capable of encoding a 55-amino-acid protein that was demonstrated electrophoretically as a 6-kilodalton product synthesized in Bacillus subtilis minicells and chemically by N-terminal sequencing of a 116-kilodalton fusion protein with Escherichia coli beta-galactosidase. Four transcripts derived from the repF region were found, of which the longest, approximately 720 nucleotides, had the length, orientation, and transcription start site necessary to code for the full-length RepF protein (216 amino acid residues), deduced from the DNA sequence. The 5' ends of the shorter repF transcripts fall within the repF open reading frame. We propose that (i) cop specifies a protein rather than an RNA countertranscript, (ii) the Cop protein functions as a negative-acting element in pE194 replication by regulating synthesis of both RepF and of itself, and (iii) increased plasmid copy number can be explained in terms of cop region mutations that either reduce the intrinsic activity of Cop protein or the rate of its synthesis.
Full text
PDF






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Anagnostopoulos C., Spizizen J. REQUIREMENTS FOR TRANSFORMATION IN BACILLUS SUBTILIS. J Bacteriol. 1961 May;81(5):741–746. doi: 10.1128/jb.81.5.741-746.1961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Band L., Henner D. J. Bacillus subtilis requires a "stringent" Shine-Dalgarno region for gene expression. DNA. 1984;3(1):17–21. doi: 10.1089/dna.1.1984.3.17. [DOI] [PubMed] [Google Scholar]
- Bergemann A. D., Whitley J. C., Finch L. R. Homology of mycoplasma plasmid pADB201 and staphylococcal plasmid pE194. J Bacteriol. 1989 Jan;171(1):593–595. doi: 10.1128/jb.171.1.593-595.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berk A. J., Sharp P. A. Sizing and mapping of early adenovirus mRNAs by gel electrophoresis of S1 endonuclease-digested hybrids. Cell. 1977 Nov;12(3):721–732. doi: 10.1016/0092-8674(77)90272-0. [DOI] [PubMed] [Google Scholar]
- CRAVEN G. R., STEERS E., Jr, ANFINSEN C. B. PURIFICATION, COMPOSITION, AND MOLECULAR WEIGHT OF THE BETA-GALACTOSIDASE OF ESCHERICHIA COLI K12. J Biol Chem. 1965 Jun;240:2468–2477. [PubMed] [Google Scholar]
- Casadaban M. J., Chou J., Cohen S. N. In vitro gene fusions that join an enzymatically active beta-galactosidase segment to amino-terminal fragments of exogenous proteins: Escherichia coli plasmid vectors for the detection and cloning of translational initiation signals. J Bacteriol. 1980 Aug;143(2):971–980. doi: 10.1128/jb.143.2.971-980.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dempsey L. A., Dubnau D. A. Localization of the replication origin of plasmid pE194. J Bacteriol. 1989 May;171(5):2866–2869. doi: 10.1128/jb.171.5.2866-2869.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dreyfus M., Kotlarz D., Busby S. Point mutations that affect translation initiation in the Escherichia coli gal E gene. J Mol Biol. 1985 Apr 5;182(3):411–417. doi: 10.1016/0022-2836(85)90200-1. [DOI] [PubMed] [Google Scholar]
- Dubnau D., Davidoff-Abelson R., Scher B., Cirigliano C. Fate of transforming deoxyribonucleic acid after uptake by competent Bacillus subtilis: phenotypic characterization of radiation-sensitive recombination-deficient mutants. J Bacteriol. 1973 Apr;114(1):273–286. doi: 10.1128/jb.114.1.273-286.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gennaro M. L., Kornblum J., Novick R. P. A site-specific recombination function in Staphylococcus aureus plasmids. J Bacteriol. 1987 Jun;169(6):2601–2610. doi: 10.1128/jb.169.6.2601-2610.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gough J. A., Murray N. E. Sequence diversity among related genes for recognition of specific targets in DNA molecules. J Mol Biol. 1983 May 5;166(1):1–19. doi: 10.1016/s0022-2836(83)80047-3. [DOI] [PubMed] [Google Scholar]
- Gruss A., Ehrlich S. D. The family of highly interrelated single-stranded deoxyribonucleic acid plasmids. Microbiol Rev. 1989 Jun;53(2):231–241. doi: 10.1128/mr.53.2.231-241.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gryczan T. J., Hahn J., Contente S., Dubnau D. Replication and incompatibility properties of plasmid pE194 in Bacillus subtilis. J Bacteriol. 1982 Nov;152(2):722–735. doi: 10.1128/jb.152.2.722-735.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horinouchi S., Weisblum B. Nucleotide sequence and functional map of pE194, a plasmid that specifies inducible resistance to macrolide, lincosamide, and streptogramin type B antibodies. J Bacteriol. 1982 May;150(2):804–814. doi: 10.1128/jb.150.2.804-814.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lacks S. A., Lopez P., Greenberg B., Espinosa M. Identification and analysis of genes for tetracycline resistance and replication functions in the broad-host-range plasmid pLS1. J Mol Biol. 1986 Dec 20;192(4):753–765. doi: 10.1016/0022-2836(86)90026-4. [DOI] [PubMed] [Google Scholar]
- Mayford M., Weisblum B. Messenger RNA from Staphylococcus aureus that specifies macrolide-lincosamide-streptogramin resistance. Demonstration of its conformations and of the leader peptide it encodes. J Mol Biol. 1985 Oct 20;185(4):769–780. doi: 10.1016/0022-2836(85)90061-0. [DOI] [PubMed] [Google Scholar]
- Mertens G., Reeve J. N. Synthesis of cell envelope components by anucleate cells (minicells) of Bacillus subtilis. J Bacteriol. 1977 Mar;129(3):1198–1207. doi: 10.1128/jb.129.3.1198-1207.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Messing J. New M13 vectors for cloning. Methods Enzymol. 1983;101:20–78. doi: 10.1016/0076-6879(83)01005-8. [DOI] [PubMed] [Google Scholar]
- Minton N. P., Oultram J. D., Brehm J. K., Atkinson T. The replication proteins of plasmids pE194 and pLS1 have N-terminal homology. Nucleic Acids Res. 1988 Apr 11;16(7):3101–3101. doi: 10.1093/nar/16.7.3101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Novick R. P. Plasmid incompatibility. Microbiol Rev. 1987 Dec;51(4):381–395. doi: 10.1128/mr.51.4.381-395.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Novick R. P. Staphylococcal plasmids and their replication. Annu Rev Microbiol. 1989;43:537–565. doi: 10.1146/annurev.mi.43.100189.002541. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scott J. R. Regulation of plasmid replication. Microbiol Rev. 1984 Mar;48(1):1–23. doi: 10.1016/b978-0-12-048850-6.50006-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shivakumar A. G., Gryczan T. J., Kozlov Y. I., Dubnau D. Organization of the pE194 genome. Mol Gen Genet. 1980;179(2):241–252. doi: 10.1007/BF00425450. [DOI] [PubMed] [Google Scholar]
- Sozhamannan S., Dabert P., Moretto V., Ehrlich S. D., Gruss A. Plus-origin mapping of single-stranded DNA plasmid pE194 and nick site homologies with other plasmids. J Bacteriol. 1990 Aug;172(8):4543–4548. doi: 10.1128/jb.172.8.4543-4548.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villafane R., Bechhofer D. H., Narayanan C. S., Dubnau D. Replication control genes of plasmid pE194. J Bacteriol. 1987 Oct;169(10):4822–4829. doi: 10.1128/jb.169.10.4822-4829.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weisblum B., Graham M. Y., Gryczan T., Dubnau D. Plasmid copy number control: isolation and characterization of high-copy-number mutants of plasmid pE194. J Bacteriol. 1979 Jan;137(1):635–643. doi: 10.1128/jb.137.1.635-643.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young F. E., Smith C., Reilly B. E. Chromosomal location of genes regulating resistance to bacteriophage in Bacillus subtilis. J Bacteriol. 1969 Jun;98(3):1087–1097. doi: 10.1128/jb.98.3.1087-1097.1969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- del Solar G. H., de al Campa A. G., Pérez-Martín J., Choli T., Espinosa M. Purification and characterization of RepA, a protein involved in the copy number control of plasmid pLS1. Nucleic Acids Res. 1989 Apr 11;17(7):2405–2420. doi: 10.1093/nar/17.7.2405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- te Riele H., Michel B., Ehrlich S. D. Single-stranded plasmid DNA in Bacillus subtilis and Staphylococcus aureus. Proc Natl Acad Sci U S A. 1986 Apr;83(8):2541–2545. doi: 10.1073/pnas.83.8.2541. [DOI] [PMC free article] [PubMed] [Google Scholar]