Abstract
Triple-helix formation can in principle serve as a general method for sequence-specific recognition and physical separation of duplex DNA molecules. Realization of this goal depends on how much the triplex is destabilized by mismatches and other defects (specificity) and on finding conditions in which perfect complexes are stable and defect complexes are not (stringency). We have addressed the question of specificity by determining the difference in free energy between perfect and defect complexes by using UV melting curves and equilibrium competition experiments. We find that third strands that bind with either single-base bulges or single mismatches are destabilized relative to the perfect triplex by 2.5-2.9 and 3.2-4.0 kcal/mol (1 cal = 4.184 J), respectively, essentially equivalent to the corresponding values determined for duplex DNA and RNA. Also, we present a method, referred to as stringency clamping, which maintains specific binding under conditions far from normal stringency. To do this, we provide for the formation of a competing structure involving the third strand with stability between that of the perfect and imperfect complexes; the competitive interaction effectively prevents triplex formation at imperfect sites even far below their melting temperature. We illustrate the phenomenon with three different stringency clamps, two of which compete for the all-pyrimidine third strand through Watson-Crick pairing and one that competes through all-pyrimidine pairing at acidic pH. We demonstrate physical separation of two duplex DNA molecules differing by a single base pair in their target sequence for triple-helix formation.
Full text
PDF

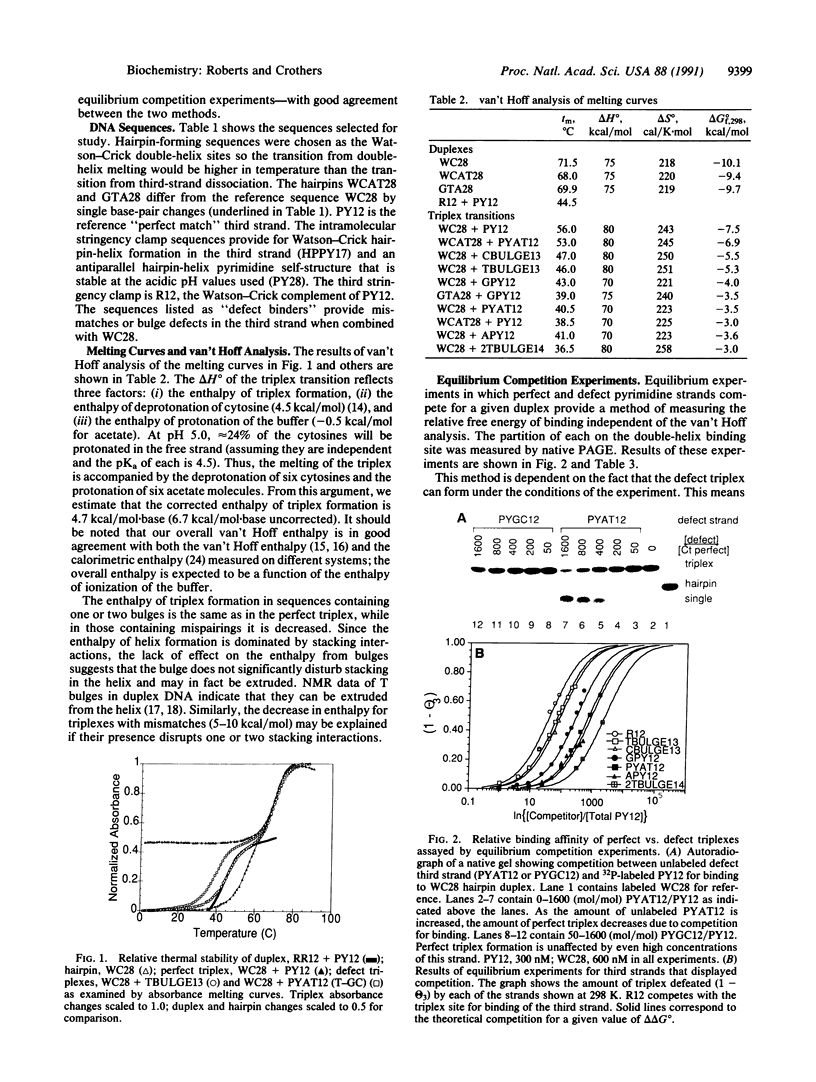


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arnott S., Bond P. J., Selsing E., Smith P. J. Models of triple-stranded polynucleotides with optimised stereochemistry. Nucleic Acids Res. 1976 Oct;3(10):2459–2470. doi: 10.1093/nar/3.10.2459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belotserkovskii B. P., Veselkov A. G., Filippov S. A., Dobrynin V. N., Mirkin S. M., Frank-Kamenetskii M. D. Formation of intramolecular triplex in homopurine-homopyrimidine mirror repeats with point substitutions. Nucleic Acids Res. 1990 Nov 25;18(22):6621–6624. doi: 10.1093/nar/18.22.6621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cantor C. R., Warshaw M. M., Shapiro H. Oligonucleotide interactions. 3. Circular dichroism studies of the conformation of deoxyoligonucleotides. Biopolymers. 1970;9(9):1059–1077. doi: 10.1002/bip.1970.360090909. [DOI] [PubMed] [Google Scholar]
- Fink T. R., Crothers D. M. Free energy of imperfect nucleic acid helices. I. The bulge defect. J Mol Biol. 1972 Apr 28;66(1):1–12. doi: 10.1016/s0022-2836(72)80002-0. [DOI] [PubMed] [Google Scholar]
- François J. C., Saison-Behmoaras T., Barbier C., Chassignol M., Thuong N. T., Hélène C. Sequence-specific recognition and cleavage of duplex DNA via triple-helix formation by oligonucleotides covalently linked to a phenanthroline-copper chelate. Proc Natl Acad Sci U S A. 1989 Dec;86(24):9702–9706. doi: 10.1073/pnas.86.24.9702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gralla J., Crothers D. M. Free energy of imperfect nucleic acid helices. 3. Small internal loops resulting from mismatches. J Mol Biol. 1973 Aug 5;78(2):301–319. doi: 10.1016/0022-2836(73)90118-6. [DOI] [PubMed] [Google Scholar]
- Griffin L. C., Dervan P. B. Recognition of thymine adenine.base pairs by guanine in a pyrimidine triple helix motif. Science. 1989 Sep 1;245(4921):967–971. doi: 10.1126/science.2549639. [DOI] [PubMed] [Google Scholar]
- Hanvey J. C., Shimizu M., Wells R. D. Intramolecular DNA triplexes in supercoiled plasmids. Proc Natl Acad Sci U S A. 1988 Sep;85(17):6292–6296. doi: 10.1073/pnas.85.17.6292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Htun H., Dahlberg J. E. Single strands, triple strands, and kinks in H-DNA. Science. 1988 Sep 30;241(4874):1791–1796. doi: 10.1126/science.3175620. [DOI] [PubMed] [Google Scholar]
- Kalnik M. W., Norman D. G., Li B. F., Swann P. F., Patel D. J. Conformational transitions in thymidine bulge-containing deoxytridecanucleotide duplexes. Role of flanking sequence and temperature in modulating the equilibrium between looped out and stacked thymidine bulge states. J Biol Chem. 1990 Jan 15;265(2):636–647. [PubMed] [Google Scholar]
- Le Doan T., Perrouault L., Praseuth D., Habhoub N., Decout J. L., Thuong N. T., Lhomme J., Hélène C. Sequence-specific recognition, photocrosslinking and cleavage of the DNA double helix by an oligo-[alpha]-thymidylate covalently linked to an azidoproflavine derivative. Nucleic Acids Res. 1987 Oct 12;15(19):7749–7760. doi: 10.1093/nar/15.19.7749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Longfellow C. E., Kierzek R., Turner D. H. Thermodynamic and spectroscopic study of bulge loops in oligoribonucleotides. Biochemistry. 1990 Jan 9;29(1):278–285. doi: 10.1021/bi00453a038. [DOI] [PubMed] [Google Scholar]
- Manzini G., Xodo L. E., Gasparotto D., Quadrifoglio F., van der Marel G. A., van Boom J. H. Triple helix formation by oligopurine-oligopyrimidine DNA fragments. Electrophoretic and thermodynamic behavior. J Mol Biol. 1990 Jun 20;213(4):833–843. doi: 10.1016/S0022-2836(05)80267-0. [DOI] [PubMed] [Google Scholar]
- Moser H. E., Dervan P. B. Sequence-specific cleavage of double helical DNA by triple helix formation. Science. 1987 Oct 30;238(4827):645–650. doi: 10.1126/science.3118463. [DOI] [PubMed] [Google Scholar]
- Plum G. E., Park Y. W., Singleton S. F., Dervan P. B., Breslauer K. J. Thermodynamic characterization of the stability and the melting behavior of a DNA triplex: a spectroscopic and calorimetric study. Proc Natl Acad Sci U S A. 1990 Dec;87(23):9436–9440. doi: 10.1073/pnas.87.23.9436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rajagopal P., Feigon J. NMR studies of triple-strand formation from the homopurine-homopyrimidine deoxyribonucleotides d(GA)4 and d(TC)4. Biochemistry. 1989 Sep 19;28(19):7859–7870. doi: 10.1021/bi00445a048. [DOI] [PubMed] [Google Scholar]
- Rajagopal P., Feigon J. Triple-strand formation in the homopurine:homopyrimidine DNA oligonucleotides d(G-A)4 and d(T-C)4. Nature. 1989 Jun 22;339(6226):637–640. doi: 10.1038/339637a0. [DOI] [PubMed] [Google Scholar]
- Tibanyenda N., De Bruin S. H., Haasnoot C. A., van der Marel G. A., van Boom J. H., Hilbers C. W. The effect of single base-pair mismatches on the duplex stability of d(T-A-T-T-A-A-T-A-T-C-A-A-G-T-T-G) . d(C-A-A-C-T-T-G-A-T-A-T-T-A-A-T-A). Eur J Biochem. 1984 Feb 15;139(1):19–27. doi: 10.1111/j.1432-1033.1984.tb07970.x. [DOI] [PubMed] [Google Scholar]
- Voloshin O. N., Mirkin S. M., Lyamichev V. I., Belotserkovskii B. P., Frank-Kamenetskii M. D. Chemical probing of homopurine-homopyrimidine mirror repeats in supercoiled DNA. Nature. 1988 Jun 2;333(6172):475–476. doi: 10.1038/333475a0. [DOI] [PubMed] [Google Scholar]
- Woodson S. A., Crothers D. M. Preferential location of bulged guanosine internal to a G.C tract by 1H NMR. Biochemistry. 1988 Jan 12;27(1):436–445. doi: 10.1021/bi00401a065. [DOI] [PubMed] [Google Scholar]
- Xodo L. E., Manzini G., Quadrifoglio F. Spectroscopic and calorimetric investigation on the DNA triplex formed by d(CTCTTCTTTCTTTTCTTTCTTCTC) and d(GAGAAGAAAGA) at acidic pH. Nucleic Acids Res. 1990 Jun 25;18(12):3557–3564. doi: 10.1093/nar/18.12.3557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de los Santos C., Rosen M., Patel D. NMR studies of DNA (R+)n.(Y-)n.(Y+)n triple helices in solution: imino and amino proton markers of T.A.T and C.G.C+ base-triple formation. Biochemistry. 1989 Sep 5;28(18):7282–7289. doi: 10.1021/bi00444a021. [DOI] [PubMed] [Google Scholar]






