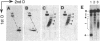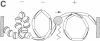Abstract
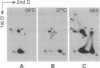
During transcription, positive DNA supercoils generated ahead of RNA polymerase could theoretically uncoil the negative DNA supercoils associated with nucleosomes and thereby decondense the chromatin fiber in preparation for RNA polymerase passage. Here we examine the effect of positive DNA supercoiling on the structure of yeast 2-microns minichromosomes. We utilized a conditional topoisomerase mutant expressing Escherichia coli topoisomerase I to convert the DNA supercoiling state from negative to positive in vivo. Minichromosomes containing positively supercoiled DNA exhibited a striking increase in DNase I sensitivity. They also displayed additional micrococcal nuclease cleavage sites but yielded nearly typical nucleosomal ladders after extensive digestion. Upon in vitro relaxation with eukaryotic topoisomerase I, the minichromosomes remained DNase I sensitive but were converted to negative DNA supercoiling with a slightly increased linking number compared to typical minichromosomes, thus indicating the presence of bound histones. Therefore, positive DNA supercoiling provides a mechanism for generating, but is not required for maintaining, a conformation in chromatin characteristic of highly transcribed genes.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baer B. W., Rhodes D. Eukaryotic RNA polymerase II binds to nucleosome cores from transcribed genes. Nature. 1983 Feb 10;301(5900):482–488. doi: 10.1038/301482a0. [DOI] [PubMed] [Google Scholar]
- Brill S. J., Sternglanz R. Transcription-dependent DNA supercoiling in yeast DNA topoisomerase mutants. Cell. 1988 Jul 29;54(3):403–411. doi: 10.1016/0092-8674(88)90203-6. [DOI] [PubMed] [Google Scholar]
- Broach J. R. The yeast plasmid 2 mu circle. Cell. 1982 Feb;28(2):203–204. doi: 10.1016/0092-8674(82)90337-3. [DOI] [PubMed] [Google Scholar]
- Clark D. J., Felsenfeld G. Formation of nucleosomes on positively supercoiled DNA. EMBO J. 1991 Feb;10(2):387–395. doi: 10.1002/j.1460-2075.1991.tb07960.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fagrelius T. J., Livingston D. M. Location of DNAase I sensitive cleavage sites in the yeast 2 micron plasmid DNA chromosome. J Mol Biol. 1984 Feb 15;173(1):1–13. doi: 10.1016/0022-2836(84)90400-5. [DOI] [PubMed] [Google Scholar]
- Giaever G. N., Wang J. C. Supercoiling of intracellular DNA can occur in eukaryotic cells. Cell. 1988 Dec 2;55(5):849–856. doi: 10.1016/0092-8674(88)90140-7. [DOI] [PubMed] [Google Scholar]
- Harland R. M., Weintraub H., McKnight S. L. Transcription of DNA injected into Xenopus oocytes is influenced by template topology. Nature. 1983 Mar 3;302(5903):38–43. doi: 10.1038/302038a0. [DOI] [PubMed] [Google Scholar]
- Izban M. G., Luse D. S. Transcription on nucleosomal templates by RNA polymerase II in vitro: inhibition of elongation with enhancement of sequence-specific pausing. Genes Dev. 1991 Apr;5(4):683–696. doi: 10.1101/gad.5.4.683. [DOI] [PubMed] [Google Scholar]
- Jantzen K., Fritton H. P., Igo-Kemenes T. The DNase I sensitive domain of the chicken lysozyme gene spans 24 kb. Nucleic Acids Res. 1986 Aug 11;14(15):6085–6099. doi: 10.1093/nar/14.15.6085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee M. S., Garrard W. T. Transcription-induced nucleosome 'splitting': an underlying structure for DNase I sensitive chromatin. EMBO J. 1991 Mar;10(3):607–615. doi: 10.1002/j.1460-2075.1991.tb07988.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levy A., Noll M. Chromatin fine structure of active and repressed genes. Nature. 1981 Jan 15;289(5794):198–203. doi: 10.1038/289198a0. [DOI] [PubMed] [Google Scholar]
- Liu L. F., Wang J. C. Supercoiling of the DNA template during transcription. Proc Natl Acad Sci U S A. 1987 Oct;84(20):7024–7027. doi: 10.1073/pnas.84.20.7024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Losa R., Brown D. D. A bacteriophage RNA polymerase transcribes in vitro through a nucleosome core without displacing it. Cell. 1987 Aug 28;50(5):801–808. doi: 10.1016/0092-8674(87)90338-2. [DOI] [PubMed] [Google Scholar]
- Lowary P. T., Widom J. Higher-order structure of Saccharomyces cerevisiae chromatin. Proc Natl Acad Sci U S A. 1989 Nov;86(21):8266–8270. doi: 10.1073/pnas.86.21.8266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luchnik A. N., Hisamutdinov T. A., Georgiev G. P. Inhibition of transcription in eukaryotic cells by X-irradiation: relation to the loss of topological constraint in closed DNA loops. Nucleic Acids Res. 1988 Jun 10;16(11):5175–5190. doi: 10.1093/nar/16.11.5175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morse R. H., Pederson D. S., Dean A., Simpson R. T. Yeast nucleosomes allow thermal untwisting of DNA. Nucleic Acids Res. 1987 Dec 23;15(24):10311–10330. doi: 10.1093/nar/15.24.10311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Norton V. G., Imai B. S., Yau P., Bradbury E. M. Histone acetylation reduces nucleosome core particle linking number change. Cell. 1989 May 5;57(3):449–457. doi: 10.1016/0092-8674(89)90920-3. [DOI] [PubMed] [Google Scholar]
- Pfaffle P., Gerlach V., Bunzel L., Jackson V. In vitro evidence that transcription-induced stress causes nucleosome dissolution and regeneration. J Biol Chem. 1990 Oct 5;265(28):16830–16840. [PubMed] [Google Scholar]
- Prior C. P., Cantor C. R., Johnson E. M., Littau V. C., Allfrey V. G. Reversible changes in nucleosome structure and histone H3 accessibility in transcriptionally active and inactive states of rDNA chromatin. Cell. 1983 Oct;34(3):1033–1042. doi: 10.1016/0092-8674(83)90561-5. [DOI] [PubMed] [Google Scholar]
- Reed K. C., Mann D. A. Rapid transfer of DNA from agarose gels to nylon membranes. Nucleic Acids Res. 1985 Oct 25;13(20):7207–7221. doi: 10.1093/nar/13.20.7207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saavedra R. A., Huberman J. A. Both DNA topoisomerases I and II relax 2 micron plasmid DNA in living yeast cells. Cell. 1986 Apr 11;45(1):65–70. doi: 10.1016/0092-8674(86)90538-6. [DOI] [PubMed] [Google Scholar]
- Seligy V. L., Thomas D. Y., Miki B. L. Saccharomyces cerevisiae plasmid, Scp or 2 mum: intracellular distribution, stability and nucleosomal-like packaging. Nucleic Acids Res. 1980 Aug 11;8(15):3371–3391. doi: 10.1093/nar/8.15.3371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sutton A., Broach J. R. Signals for transcription initiation and termination in the Saccharomyces cerevisiae plasmid 2 micron circle. Mol Cell Biol. 1985 Oct;5(10):2770–2780. doi: 10.1128/mcb.5.10.2770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villeponteau B., Lundell M., Martinson H. Torsional stress promotes the DNAase I sensitivity of active genes. Cell. 1984 Dec;39(3 Pt 2):469–478. doi: 10.1016/0092-8674(84)90454-9. [DOI] [PubMed] [Google Scholar]
- Volkert F. C., Broach J. R. Site-specific recombination promotes plasmid amplification in yeast. Cell. 1986 Aug 15;46(4):541–550. doi: 10.1016/0092-8674(86)90879-2. [DOI] [PubMed] [Google Scholar]
- Weintraub H. Assembly and propagation of repressed and depressed chromosomal states. Cell. 1985 Oct;42(3):705–711. doi: 10.1016/0092-8674(85)90267-3. [DOI] [PubMed] [Google Scholar]
- Weintraub H., Cheng P. F., Conrad K. Expression of transfected DNA depends on DNA topology. Cell. 1986 Jul 4;46(1):115–122. doi: 10.1016/0092-8674(86)90865-2. [DOI] [PubMed] [Google Scholar]
- Weintraub H., Groudine M. Chromosomal subunits in active genes have an altered conformation. Science. 1976 Sep 3;193(4256):848–856. doi: 10.1126/science.948749. [DOI] [PubMed] [Google Scholar]
- Williams S. P., Athey B. D., Muglia L. J., Schappe R. S., Gough A. H., Langmore J. P. Chromatin fibers are left-handed double helices with diameter and mass per unit length that depend on linker length. Biophys J. 1986 Jan;49(1):233–248. doi: 10.1016/S0006-3495(86)83637-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu M., Barnard M. B., Rose S. M., Cockerill P. N., Huang S. Y., Garrard W. T. Transcription termination and chromatin structure of the active immunoglobulin kappa gene locus. J Biol Chem. 1986 Mar 15;261(8):3838–3845. [PubMed] [Google Scholar]
- Yang L., Jessee C. B., Lau K., Zhang H., Liu L. F. Template supercoiling during ATP-dependent DNA helix tracking: studies with simian virus 40 large tumor antigen. Proc Natl Acad Sci U S A. 1989 Aug;86(16):6121–6125. doi: 10.1073/pnas.86.16.6121. [DOI] [PMC free article] [PubMed] [Google Scholar]