Abstract
Longevity Assurance 5 (LASS5), a member of the LASS/Ceramide Synthases family, synthesizes C16-ceramide and is implicated in tumor biology. However, its precise role is not yet well understood. A yeast two-hybrid screen was performed using a human cDNA library to identify potential LASS5-interaction partners. One identified clone encodes succinate dehydrogenase subunit B (SDHB). Mammalian two-hybrid assays showed that LASS5 interacts with SDHB, and the result was also confirmed by GST pull-down and co-immunoprecipitation assays. The C-terminal fragment of SDHB was required for the interaction. LASS5 and SDHB were co-localized in COS-7 cells. LASS5 and SDHB expressions were found to be up-regulated in neuroglioma tissue. Transfection assays showed that LASS5 or SDHB expression repressed p53 or p21 reporter activity, respectively. Simultaneous LASS5 and SDHB expression resulted in stronger repression of p53 and p21 reporter activity, suggesting that LASS5 and SDHB interaction may synergistically affect transcriptional regulation of p53 and p21. Our data provide new molecular insights into potential roles of LASS5 and SDHB in tumor biology.
Keywords: LASS5, SDHB, neuroglioma, p53
INTRODUCTION
Longevity Assurance (LASS) genes, also known as Ceramide Synthases (CerSs), are members of a family of six related genes found in mice and humans. Each LASS protein regulates the synthesis of ceramides of particular acyl-chain lengths [1]. LASS5, which synthesizes C16-ceramide through a Lag1p motif [2], is implicated in diverse cellular processes. LASS5 and C16:0-ceramide are important mediators of cell stress responses. LASS5 mRNA levels are up-regulated in several models of stress, including responses to DNA damage, hypoxia/re-oxygenation injury, and treatment with the AMPK (AMP-activated protein kinase) inhibitor Compound C [3–5]. LASS5 is also implicated in tumor cell biology, mediating sensitivity to chemotherapeutic drugs [3] and acting as aneoantigen influencing cytotoxic T lymphocyte (CTL) immune surveillance [6]. Overexpression of LASS5 promotes apoptosis and autophagy [7, 8]. LASS5 expression changes in response to carcinogens [9] and is reportedly elevated in colorectal [10, 11], endometrial and colon cancer [12]. In Molt-4 LXSN leukemia cells, activation of LASS5 is responsible for de novo ceramide synthesis specifically drived by p53 [3]. p53 plays key roles to prevent tumor development by its cell-cycle and apoptosis functions. p53 can block cell cycle progression and/or induce apoptosis through specific transactivation of a number of target genes such as p21 [13]. p53 and p21 can induce apoptosis in response to genotoxic agents, and activate extrinsic and mitochondrial death pathway components [14].
Succinate dehydrogenase (SDH) is a highly conserved heterotetrameric protein. SDHA and SDHB catalytic subunits protrude into the mitochondrial matrix, anchored to the mitochondrial inner membrane by SDHC and SDHD subunits [15]. SDH is the first mitochondrial enzyme identified as a tumor suppressor in familial paragangliomas (PGL) [16]. PGL and Pheochromocytomas (PCCs) are neural-crest-derived tumors associated with germline and somatic mutations in SDH genes [15, 17]. Mutations in SDHB are reported in 4.6% of sporadic PPC/PGL patients [18]. SDHB mutations are also potentially associated with renal cell carcinoma [15, 19, 20]. Renal tumors in patients bearing SDHB mutations present various histological subtypes [15, 19–22]. Moreover, mutations in SDHB have been implicated in neuroblastomas [13,17], thyroid carcinoma [15,19], pituitary tumors [23], gastrointestinal stromal tumors (GISTs) [24], seminoma, prostatic adenocarcinoma, and lymphoepithelial carcinoma of the stomach [25]. Reduced SDHB expression is associated with growth and de-differentiation of colorectal cancer cells [26], and shorter survival time of patients with recurrent nasopharyngeal carcinoma [27]. In human ovarian carcinoma SDHB was shown to play a key role in cell proliferation, invasion, migration, and apoptosis [28]. However, no physical interaction between LASS and SDH families has been reported to date. In the present study we identified an association of human LASS5 and SDHB proteins. We showed that SDHB interacted with LASS5 in vitro and in cells. The C-terminal fragment of SDHB was required for the interaction. Both LASS5 and SDHB expressions were up-regulated in neuroglioma tissue compared to normal nervous ganglion tissue. LASS5 cooperated with SDHB to suppress the transcriptional activity of p53 and p21 reporter constructs. These observations yield new insights into the molecular mechanism underlying potential tumor biological roles of LASS5 and SDHB.
MATERIALS AND METHODS
Cloning and Plasmid Construction
LASS5 clones
The full-length LASS5 open reading frame was amplified from a human Hela cell cDNA library using primers PORF1 and PORF2 (Table 1). The resulting amplicon was cloned into pMD18T-vector (Sagon) to generate pMD18T-LASS5. In order to obtain a fusion protein of LASS5 with green fluorescent protein (GFP), pEGFP-C2-LASS5, the full length LASS5 coding sequence was amplified from pMD18T-LASS5 using primers P1 and P2 (Table 1) incorporating EcoRI and SalI sites. The amplified and cleaved DNA fragment was cloned in pEGFP-C2. The full-length LASS5 coding sequence was also subcloned into expression vector pCMV-BD, containing the coding region of the GAL4 DNA-binding domain (BD), to make pCMV-BD-LASS5 expresses a fusion protein of GAL4 DNA-BD and LASS5. Myc epitope-tagged full-length (amino acids 1–392) and truncated (amino acids 69–170) LASS5 expression vectors were generated by PCR amplification from pMD18T-LASS5 using primers P1, P3, P4, P5 (Table 1) incorporating EcoRI and XhoI sites, followed by in-frame assembly in pcDNA3.1-His to create pcDNA3.1-His-LASS5. pGBKT7-HH (encoding amino acids 72–146, including the HOX domain) was constructed by ligating a PCR fragment amplified from pMD18T-LASS5 with primers P6 and P7 (Table 1), incorporating EcoRI and SalI sites, into pGBKT7. The same fragment, HH, was subcloned into the prokaryotic expression vector pGEX-4T-1 to produce a GST (glutathione S-transferase) –fusion protein. GFP-HH and Gal4 DNA-BD-HH fusion protein constructs, pEGFP-C2-HH and pCMV-BD-HH, respectively, were also generated. All constructs were confirmed by DNA sequencing (ABI PRISM3770 DNA Sequencer).
Table 1.
The specific oligonucleotide sequences of primers.
| Primers | Sequences |
|---|---|
|
| |
| pORF1 | 5′- TTCGGGGTGGGCGTAAGAT-3′ |
| pOFR2 | 5′- ATGGGAAGGGCCAAGAGGA-3′ |
| P1 | 5′- GCGTGAATTCCGGGGTGGGCGTAAGATG-3′ |
| P2 | 5′- GCAAGTCGACTACTCTTCAGCCCAGTAG-3′ |
| P3 | 5′- ACCCTCGAGCTCTTCAGCCCAGTAGCTG -3′ |
| P4 | 5′- GAGGAATTCATGGCCAAACCCTGTGCA C -3′ |
| P5 | 5′- ACTGCTCGAGGTCCCAGAACCAAGGTGA-3′ |
| P6 | 5′- GCCGAATTCTGTGCACTCTGTATTGGC-3′ |
| P7 | 5′- ATGGTCGACTCCACATGCTTTCACAG-3′ |
| PS1 | 5′- AAGAATTCGGGCTGACGTCAGGAGCCAA -3′ |
| PS2 | 5′- GGCGTCGACTAAACTGAAGCTTTCTTCTCC-3′ |
| GS1 | 5′-TGAAGGTCGGAGTCAACGGAT TTGGT-3′ |
| GS2 | 5′-CATGTGGGCCATGAGGTCCACCAC-3′ |
SDHB clones
One of 16 positive clones selected by yeast two-hybrid screening, encoding the C-terminal fragment of SDHB, was designated pACT2-SDHB-C. The full-length SDHB ORF was PCR-amplified from a human Hela cell cDNA library using primers PS1 and PS2 (Table 1) incorporating EcoRI and SalI sites. The amplicon was cloned in vector pMD18T and then subcloned in pCMV-AD to yield pCMV-AD-SDHB, encoding a GAL4 DNA-activating domain (AD) and SDHB fusion protein. To create Flag-tagged SDHB, full length and C-terminal SDHB coding sequences were cloned in-frame into pCMV-Tag2 with EcoRI/SalI sites. In addition, C-terminal fragment cut directly from pACT2 was inserted into pCMV-Tag2 vector with EcoRI/XhoI sites. Both full-length and truncated fragments of SDHB were also cloned into pDS-Red-Monomer-C1 (Monomer-SDHB). All constructs were confirmed by DNA sequencing (ABI PRISM3770 DNA Sequencer).
Yeast Two-Hybrid Screening
Yeast two-hybrid (Y2H) screening was performed according to the method described previously [29]. The LASS5 HOX domain construct pGBKT7-HH was used as bait. The yeast strain AH109 was transformed with pGBKT7-HH and a human heart cDNA library cloned into the pACT2 vector was screened. Clones of interest were sequenced. BLAST searches were conducted using the NCBI online service.
Cell Culture, Transfection
COS-7 cells were maintained in DMEM (Dulbecco’s modified Eagle’s medium; Gibco-BRL) supplemented with 10% fetal bovine serum (FBS) at 37°C and 5% CO2. Cells were transfected using LipofectAMINE (Invitrogen) according to the previous protocol [30].
Co-Immunoprecipitation Assays
COS-7 cells were co-transfected with pcDNA3.1-His-LASS5 and pCMV-tag2-SDHB, or the corresponding truncated expression plasmids. After 48 h, cells were harvested in lysis buffer according to the method described previously [29]. Western blot analysis was carried out with anti-Myc or anti-Flag monoclonal antibodies.
Expression of LASS5 (HOX) in E. coli and Preparation of Polyclonal Antibodies
E. coli BL-21 was transformed with pGEX-4T-1 (as a control) or pGEX-4T-1-HH plasmids. The bacteria were cultured in LB medium with ampicillin to an OD 600 of 0.4 to 0.6 at 37°C. Then GST or GST-HH fusion protein expression was induced with IPTG, cultures were incubated overnight at 27°C, and proteins were purified according to the manufacturer’s protocol (Amersham Pharmacia). Rabbits were first immunized with proteins emulsified in Incomplete Freund’s Adjuvant (IFA), then subsequently reimmunized twice with half the original dose of proteins in Complete Freund’s Adjuvant (CFA). Antibody titers were measured 4 to 5 days after each immunization. Ultimately, anaesthetized animals were euthanized and bled out for antiserum.
GST pull-Down Assays
Flag-SDHB was transiently expressed in COS-7 cells for 48 h following transfection of pCMV-tag2-SDHB. The experiment was performed according to the previous method [29]. The proteins bound to the beads were separated by SDS-PAGE, transferred to PVDF membrane, and membrane bound proteins were detected using an anti-Flag antibody (Santa Cruz Biotechnology).
Transient Transfection Reporter Gene Assays and Luciferase Assay
pCMV-Tag2B-SDHB (or truncated SDHB vector) was co-transfected into HEK293 cells with p53-luc alone or p53-luc plus pCMV-Tag2B-LASS5 according to the previous protocol [30]. Experiments were performed in triplicate and each assay was repeated at least three times. For the mammalian two-hybrid assay, COS-7 cells were co-transfected with pCMV-BD-LASS5 (or truncated LASS5 vector) and pCMV-AD-SDHB (or truncated SDHB vector) together with a pFR-Luc reporter construct (Stratagene).
Fluorescence Co-Localization
pLASS5-EGFP and pMonomer-SDHB were transfected into COS-7 cells. Forty-eight hours after transfection cells were fixed with 4% paraformaldehyde for 15 min and washed with PBS three times. Nuclei were visualized with 4′, 6′-diamidino-2-phenylindole hydrochloride (DAPI). Fluorescence microscopy and image capture was performed using a charge-coupled device/camera mounted on a Nikon inverted fluorescence microscope and Ultraview imaging software.
RT-PCR and Western Blotting
Total RNA and protein were extracted using trizol reagent from one sample of normal nervous ganglion tissue and four samples of neuroglioma tissue (use of human tissue samples was approved by the ethics committee of Hunan Normal University). Approximately 20 ng of total RNA was used, and template for cDNA synthesis using the Multiscribe reverse transcription kit according to the manufacturer’s protocol. Gene-specific primer pairs P6/P7 for human LASS5 (HOX) and PS1/PS2 for SDHB were used as primers GS1/GS2 for glyceraldehyde 3-phosphate dehydrogenase (GAPDH) as an internal control. The sequences of all the PCR primer pairs are shown in Tabel 1. PCR products were electrophoresed and quantitated. Protein samples were subjected SDS-PAGE, followed by Western blotting with anti-LASS5 (HOX) antibody. Bands were visualized using peroxidase-conjugated secondary antibody and chemiluminescence (Pierce). Protein loading was verified by staining with Coomassie Blue.
RESULTS
Molecular Cloning and Domain Structure of Human LASS5
In order to study the role(s) of human LASS5 in normal cell function and disease, we cloned a full-length human LASS5 cDNA. We designed primers based on the sequence of a CERS5 (LASS5) transcript (NCBI Reference Sequence NM_147190.3) that includes a homeobox (HOX) domain. A Hela cell cDNA library was used as a template to obtain a 1272-bp PCR product representing the full length coding sequence of human LASS5. LASS5 encodes a 392 amino acid protein with a (calculated) relative molecular mass of 45.76 KDa. In addition to a HOX domain, LASS5 possesses a TRAM/LAG1/CLN8 (TLC) motif that is involved in acyl-CoA-dependent ceramide synthesis (Fig. 1A).
Fig. 1. LASS5 and SDHB mammalian cell two-hybrid assay, LASS5 and SDHB mRNA expression.
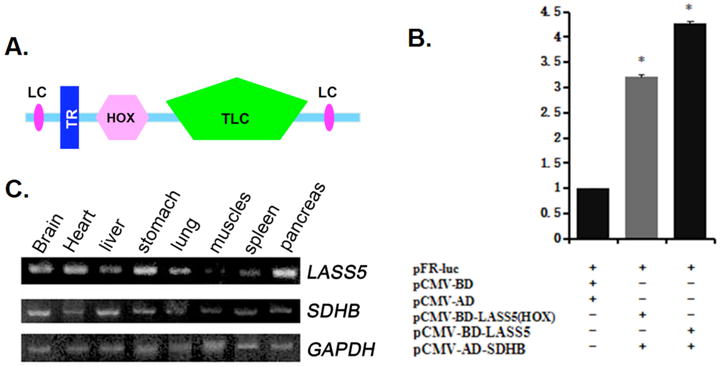
A. LASS5 protein domains inferred from Smart program analysis. LC, low complexity region. TR, transmembrane region. HOX, hox domain. TLC, TRAM/LAG1/CLN8 domain. B. Interaction between LASS5 and SDHB in mammalian cell two-hybrid assay. pFR-Luc reporter plasmid co-transfected with pCMV-BD and pCMV-AD, or pCMV-BD–LASS5 (HOX) and pCMV-AD-SDHB-C, or pCMV-BD–LASS5 and pCMV-AD-SDHB into COS-7 cells. 48 h later luciferase activity was determined. Data shown are means of three repeats of a transfection experiment following normalization transfection efficiency (β-galactosidease activity). Each transfection experiment was repeated at least three times. C. RT-PCR analysis of LASS5 and SDHB mRNA expression in adult human tissues. GAPDH expression was used as internal control.
LASS5 Interacted with SDHB in Yeast Two-Hybrid and Mammalian Cell Two-Hybrid Assays
We sought to identify potential binding partners for LASS5 by performing a yeast two-hybrid screen. A GAL4 Activation Domain (AD)-fused human adult cardiac cDNA library was used for screening. We obtained a positive clone encoding a C-terminal fragment of SDHB. We also tested LASS5-SDHB interaction in a mammalian cell two-hybrid assay. Because of the intrinsic DNA binding activity of the LASS5 HOX domain, we constructed vectors expressing full-length and truncated LASS5 proteins fused with the DNA-binding domain (BD) of the yeast transcription factor GAL-4, under the control of a CMV promoter. Full-length and C-terminal SDHB protein expression vectors were constructed in pCMV-AD. We co-transfected COS-7 cells with pCMV-BD-LASS5/HOX and pCMV-AD-SDHB/SDHB-C and a Luciferase reporter construct, pFR-Luc. In these experiments LASS5 activated transcriptional activity of the reporter gene through interaction with SDHB (Fig. 1B).
LASS5 and SDHB were Normally Expressed in the Same Tissues, and Physically Interacted with Each other In Vitro and in Mammalian Cells
We examined LASS5 and SDHB mRNA expression in different human tissues, and observed overlapping expression patterns (Fig. 1C) though with variation in relative mRNA levels. This result indicated that the proteins could be co-localized in the same tissues.
We co-transfected COS-7 cells with Myc-tagged LASS5 and Flag-tagged SDHB vectors, including full length and truncated forms Cell extracts were immunoprecipitated with anti-Myc antisera, and then subjected to immunoblotting with anti-FLAG antisera (Fig. 2A). Flag–SDHB was detected only in samples derived from Myc–LASS5 transfected cells, suggesting a physical interaction between LASS5 and SDHB. Conversely, antibodies directed against the Flag-tag immunoprecipitated the Flag-SDHB protein together with the Myc-tagged LASS5 protein. The control antibody did not precipitate either protein. Experiments performed using a Myc-tagged HOX domain of LASS5 and Flag-tagged C-terminal fragment of SDHB (Fig. 2B) indicated that the HOX domain of LASS5 and the C-terminal region of SDHB are involved in the interaction of these two proteins.
Fig. 2. LASS5 and SDHB co-immunoprecipitation assays and GST pull-down assays.

A. COS-7 cells transfected with expression vectors encoding Myc-tagged LASS5 and Flag-tagged SDHB, alone or in combination as indicated. Upper two panels show immunoblot (IB) of cell lysates confirming expression of tagged proteins. Third panel: following immunoprecipitation (IP) using anti-Myc antibody, co-expressed Flag–SDHB was detected by anti-Flag IB. Fourth panel: following IP using anti-Flag antibody, co-expressed LASS5 was detected by anti-Myc IB. B. COS-7 cells transfected with expression vectors encoding LASS5 (HOX) and Flag–SDHB-C protein domains, alone or in combination as indicated. Upper two panels show immunoblot (IB) of cell lysates confirming expression of tagged proteins. Third panel: following immunoprecipitation (IP) using anti-Myc antibody, co-expressed Flag–SDHB-C was detected by anti-Flag IB. Fourth panel: following IP using anti-Flag antibody, co-expressed LASS5 (HOX) was detected by anti-Myc IB. C. 5 μg of GST or GST-LASS5 (HOX) fusion protein bound to glutathione sepharose beads (as indicated) was mixed with 100 μg of cell lysate containing Flag-SDHB. Bound protein was detected by SDS-PAGE and immunoblotting (IB) using anti-Flag antibody.
Results from two-hybrid and co-immunoprecipation studies suggested that LASS5 was capable of physically interacting with SDHB. To further confirma direct interaction between LASS5 and SDHB, we performed in vitro GST-pull down assays using purified full length or truncated versions of LASS5 and SDHB fusion proteins. As shown in Fig. 2C, SDHB was pulled-down by GST fused to LASS5 (HOX), indicating a direct SDHB-LASS5 physical interaction in vitro.
Subcellular Co-Localization of LASS5 and SDHB
We investigated whether LASS5 and SDHB exhibited subcellular co-localization by fluorescence microscopy following transfection of COS-7 cells with pEGFP-C2-LASS5 and pDsRed-monomer-C1-SDHB, or corresponding truncated versions. Both LASS5 and SDHB full length proteins localized in the cytoplasm (Fig. 3A). The proteins co-localized most strongly in a region of the cytoplasm adjacent to the nucleus. LASS5 (HOX) was cytoplasmic when transfected together with full length SDHB (Fig. 3B). Intriguingly, LASS5 (HOX) and SDHB-N truncated proteins were observed together in both cytoplasm and nucleus (Fig. 3C). Co-transfection of LASS5 (HOX) and SDHB-C fragments, by contrast, resulted in cytoplamic co-localization (Fig. 3D) as observed for the two full length proteins. These results suggested that the C-terminal domain of SDHB was involved in the cytoplasmic localization of this protein, and interaction with the LASS5 HOX domain, resulting in cytoplasmic co-localization of the full length LASS5 and SDHB proteins.
Fig. 3. LASS5 and SDHB colocalize in COS-7 cells.

A. COS-7 cells co-transfected with LASS5-EGFP and SDHB-DsRed-Monomer expression vectors. Nuclei stained with DAPI. The full length proteins colocalized in the cell cytoplasm. B. LASS5(HOX)-EGFP protein domain and SDHB-DsRed-Monomer colocalized in the cell cytoplasm. C. LASS5(HOX)-EGFP protein domain and SDHB(N)-DsRed-Monomer N-terminal protein colocalized in cytoplasm and nucleus. D. LASS5(HOX)-EGFP protein domain and SDHB(C)-DsRed-MonomerC-terminal protein colocalized in the cytoplasm. These observations suggest that LASS5 (HOX) cytoplasmic localization is dependent upon interaction with SDHB-C domain. Scale bar 5μm.
Expression Levels of LASS5 and SDHB were Up-Regulated in Neuroglioma Tissue
We compared LASS5 and SDHB expression levels in normal nervous ganglion tissue and neuroglioma. RT-PCR was performed on total RNA isolated from four neuroglioma samples and one normal control specimen. At the same time, total protein was also extracted and subjected to Western Blot analysis. As shown in Fig. 4A, LASS5 and SDHB mRNA expression was elevated significantly above normal levels in all four neuroglioma samples. Western blot analysis showed LASS5 protein levels were significantly increased in all neuroglioma tissues relative to the control (Fig. 4B).
Fig. 4. Expression level of LASS5 was up-regulaedin glioma tissue samples, and LASS5 and SDHB inhibited p53 and p21 promoter activity.

A. RT-PCR analysis of LASS5 and SDHB mRNA expression levels in normal nervous ganglion (control, Cont) and four independent neuroglioma tissue samples. GAPDH expression assayed as internal control. Amplification products are 244 bp (LASS5) and 880-bp (SDHB). Bar graph shows quantitation of LASS5 and SDHB mRNA expression levels in tumor samples relative to control (CTRL). B. LASS5 protein expression in normal tissue and glioma samples analyzed by Western blot. Actin used as an internal control. Bar graph shows quantitation of LASS5 protein expression in tumor samples relative to control (CTRL). Unpaired student t test used to determine significance. *p<0.05. C. Cos-7 cells co-transfected with p53-Luc reporter and expression vectors pCMV-Tag2-LASS5, pCMV-Tag2-SDHB, pCMV-Tag2-LASS5(HOX), pCMV-Tag2-SDHB-C as indicated. D. Cos-7 cells co-transfected with p21-Luc reporter and expression vectors pCMV-Tag2-LASS5, pCMV-Tag2-SDHB, pCMV-Tag2-LASS5(HOX), pCMV-Tag2-SDHB-C as indicated. The data are the mean of three repeats of a transfection experiment. *p<0.05.
LASS5/SDHB Down-Regulated p53 and p21 Promoter Activities
The tumor suppressor protein p53 plays pivotal roles in promoting cell cycle arrest and apoptosis. p53 and p21, which mediates p53-dependent growth arrest, are both regulated post-translationally by stress response signaling pathways. They can induce apoptosis in response to genotoxic agents, and activate extrinsic and mitochondrial death pathway components [14]. p53 was shown, moreover, to specifically drive de novo ceramide synthesis by activation of LASS5 in Molt-4 LXSN leukemia cells [3]. To investigate the role of LASS5/SDHB in cellular signal transduction, we measured expression of p53 and p21 reporter constructs following transfection of HEK293 cells with LASS5 and SDHB expression vectors. As shown in Fig. 4, LASS5 (HOX) domain and full length SDHB protein both repress p53 (Fig. 4C) and p21 (Fig. 4D) promoter activities. Full length LASS5 protein was not active. The SDHB-C domain repressed p21 promoter activity, but not that of p53. Simultaneous expression of LASS5 (HOX) and full length SDHB had the largest repressive effects on both p53 and p21 promoters (Fig. 4C, D).
DISCUSSION
The LASS protein family is highly conserved from yeast to mammals, indicating that these proteins are important mediators of fundamental cellular processes. LASS5 contains a DNA-binding HOX domain involved in transcriptional regulation and a TLC-motif, characteristic of family members, that includes five membrane-spanning helices [31, 32]. It has been suggested that LASS proteins may be associated with oncogenic processes including cell proliferation, apoptosis, and possibly tumor development. LASS2 has been shown to suppress tumorigenesis in mice, with approximately half of LASS2 null mutants developing spontaneous pheochromocytoma (PCC) [33]. Although knowledge of LASS protein involvement in cancer biology is advancing, precise roles of distinct LASS family members, including LASS5, are not yet well understood. Here, we cloned a full length human LASS5 cDNA and showed that LASS5 interacted with SDHB through the latter protein’s N-terminal domain. Mammalian cell two-hybrid assays, GST-pull down experiments, and co-immunoprecipitation studies showed that the HOX domain of LASS5 functioned as a protein-protein interaction domain with SDHB. Furthermore, LASS5 and SDHB proteins displayed overlapping subcellular localization in COS-7 cells. LASS5 (HOX) truncated protein localization was shifted from cytoplasmic to nuclear in the presence of SDHB(C) truncated protein. Intriguingly, both LASS5 (HOX) and SDHB expression could inhibit p53 and p21 promoter activities in transfected cells.
We examined the expression level of LASS5 mRNA and protein in human patient neuroglioma samples and normal nervous ganglion tissue. RT-PCR analysis showed that LASS5 mRNA was significantly up-regulated in neuroglioma (Fig. 4A). Western blotting confirmed upregulation of LASS5 protein (Fig. 4B). These observations are compatible with published observations of colorectal cancer [10,11], endometrial and colon cancer [12], and in livers of LASS2-knockout mice [33] in which spontaneous PCC can arise [34]. Previous studies reported down-regulation of SDHB expression in nasopharyngeal carcinoma, colorectal cancer, and childhood non-Hodgkin lymphoma (NHL) [26, 27, 35]. In the experiments reported here we observed significant upregulation of SDHB mRNA in neuroglioma (Fig. 4A). LASS5 and SDHB were both considered to be tumor suppressors. We suggest that during neuroglioma development LASS5 and SDHB may be coordinately up-regulated, and interact directly with one another to prevent further cancer progression. This hypothesis remains to be explored.
Cells treated with actinomycin D or γ-irradiation rapidly upregulate p53 expression, which is followed by gradual accumulation of ceramide. Inhibiting p53 upregulation is sufficient to prevent ceramide accumulation, suggesting that p53 may regulate ceramide abundance [36]. Ceramides act as second messengers in apoptotic signaling pathways [37]. De novo ceramide synthesis was induced by ionizing radiation (IR) in HeLa cells, in which LASS2, LASS5 and LASS6 generated competing anti- and pro-apoptotic ceramides in mitochondrial membranes. Overexpression of LASS2 partially protected cells from IR-induced apoptosis, whereas overexpression of LASS5 increased apoptosis [6]. Both LASS4 and LASS5 were up-regulated in endometrial and colon cancer cells and down-regulated following induction of apoptosis [12]. Down-regulation of LASS5 was furthermore shown to attenuate Compound C-mediated ceramide production, Bax redistribution, and apoptosis in MCF7 breast carcinoma cells [5].
SDH genes, including SDHB, may be associated with a variety of tumors [15–17, 19, 23–25]. Two models have been proposed to explain how SDH loss-of-function mutations lead to tumor formation [38]. In the first model, SDH dysfunction leads to succinate accumulation that in turn induces a pseudo-hypoxia response mediated by the HIF transcription factor [39]. HIF induces the expression of genes involved in angiogenesis, proliferation, cell survival and glycolysis [40]. The second model proposes that SDH mutations contribute to phaechromocytoma when, during embryogenesis, neuronal precursor cells lacking SDH function fail to undergo apoptosis in response to growth factor with drawal [41]. This model was supported by a study of PDH3 knock-out mice which showed reduced apoptosis of sympathetic neurons [42].
p53 is a transcription factor highly inducible by many stress signals such as DNA damage, oncogene activation, and nutrient deprivation. Cell-cycle arrest and apoptosis are the most prominent outcomes of p53 activation. Many studies showed that p53 cell-cycle and apoptosis functions are important for preventing tumor development [13]. p53 can block cell cycle progression and/or induce apoptosis through specific transactivation of a number of target genes such as p21, the growth arrest and DNA damage-inducible 45 (GADD45), cyclin G, and Bcl-2-associated X protein (Bax) [13, 43, 44].
In the present study, we showed that LASS5 and SDHB protein domains could directly interact with one another and co-localized intracellularly. Expressing LASS5 (HOX) domain with the SDHB (C) domain shifted localization from the cytoplasm to the nucleus. We also demonstrated the ability of LASS5 (HOX) protein domain and full length SDHB protein to inhibit both p53 and p21 promoters. These observations provide additional insights into mechanisms by which SDHB and LASS5 may influence tumor formation through inhibition of the p53 pathway.
Acknowledgments
The authors thank all the members of the Center for Heart Development of Hunan Normal University for their assistance and encouragement. The authors thank Dr. Adam M. Richman (Children’s National Medicine Center, USA) for reviewing a draft of the manuscript. This study was supported in part by grants from the National Natural Science Foundation of China (Nos. 81470377, 81270156, 81470449, 31272396, 81270291, 81370451, 81400304, 31472060, 81370341, 31572349, 30971663, 30871417, 2015DK3001, 2015DK3002, 14a093), Postgraduate research and innovation project in Hunan Province (No. CX2012B213), and the Cooperative Innovation Center of Engineering and New Products for Developmental Biology of Hunan Province (No. 2013-448-6).
Biography

Y. Li
Footnotes
ETHICS STATEMENT
The study was approved by the ethics committees of Hunan Normal University and adhered to the tenets of the Declaration of Helsinki. Informed consent was obtained from the patients or their guardians.
CONFLICT OF INTEREST
The authors confirm that this article content has no conflicts of interest.
References
- 1.Mullen TD, Hannun YA, Obeid LM. Ceramide synthases at the centre of sphingolipid metabolism and biology. Biochem J. 2012;441:789–802. doi: 10.1042/BJ20111626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Lahiri S, Futerman AH. LASS5 is a bona fide dihydroceramide synthase that selectively utilizes palmitoyl CoA as acyl donor. J Biol Chem. 2005;280:33735–8. doi: 10.1074/jbc.M506485200. [DOI] [PubMed] [Google Scholar]
- 3.Levy M, Futerman AH. Mammalian ceramide synthases. IUBMB Life. 2010;62:347–56. doi: 10.1002/iub.319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Jin J, Hou Q, Mullen TD, et al. Ceramide generated by sphingomyelin hydrolysis and the salvage pathway is involved in hypoxia/reoxygenation-induced Bax redistribution to mitochondria in NT-2 cells. J Biol Chem. 2008;283:26509–17. doi: 10.1074/jbc.M801597200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Jin J, Mullen TD, Hou Q, et al. AMPK inhibitor Compound C stimulates ceramide production and promotes Bax redistribution and apoptosis in MCF7 breast carcinoma cells. J Lipid Res. 2009;50:2389–97. doi: 10.1194/jlr.M900119-JLR200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Mesicek J, Lee H, Feldman T, et al. Ceramide synthases 2, 5, and 6 confer distinct roles in radiation-induced apoptosis in HeLa cells. Cell Signal. 2010;22:1300–7. doi: 10.1016/j.cellsig.2010.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Yamane M, Miyazawa K, Moriya S, et al. D, L-Threo-1-phenyl-2-decanoylamino-3-morpholino-1-porpanol (DL-PDMP) increases endopasmic reticulum stress, autophagy and apoptosis accompanying ceramide accumulation via ceramide synthase 5 protein expression in A549 cells. Biochimie. 2011;93:1446–59. doi: 10.1016/j.biochi.2011.04.016. [DOI] [PubMed] [Google Scholar]
- 8.van Hall T, Wolpert EZ, van Veelen P, et al. Selective cytotoxic T-lymphocyte targeting of tumor immune escape variants. Nat Med. 2006;12:417–24. doi: 10.1038/nm1381. [DOI] [PubMed] [Google Scholar]
- 9.Kim JY, Kwon J, Kim JE, et al. Identification of potential biomarkers of genotoxicity and carcinogenicity in L5178Y mouse lymphoma cells by cDNA microarray analysis. Environ Mol Mutagen. 2005;45:80–9. doi: 10.1002/em.20077. [DOI] [PubMed] [Google Scholar]
- 10.Kijanka G, Hector S, Kay EW, et al. Human IgG antibody profiles differentiate between symptomatic patients with and without colorectal cancer. Gut. 2010;59:69–78. doi: 10.1136/gut.2009.178574. [DOI] [PubMed] [Google Scholar]
- 11.Chen L, Chen H, Li Y, et al. Endocannabinoid and ceramide levels are altered in patients with colorectal cancer. Oncol Rep. 2015;34:447–54. doi: 10.3892/or.2015.3973. [DOI] [PubMed] [Google Scholar]
- 12.Mojakgomo R, Mbita Z, Dlamini Z. Linking the ceramide synthases (CerSs) 4 and 5 with apoptosis, endometrial and colon cancers. Exp Mol Pathol. 2015;98:585–92. doi: 10.1016/j.yexmp.2015.03.019. [DOI] [PubMed] [Google Scholar]
- 13.Chen J. The Cell-Cycle Arrest and Apoptotic Functions of p53 in Tumor Initiation and Progression. Cold Spring Harb Perspect Med. 2016:a026104. doi: 10.1101/cshperspect.a026104. 6pii. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Miled C, Pontoglio M, Grbay S, et al. A genomic map of p53 binding sites identifies novel p53 targets involved in an apoptotic network. Cancer Res. 2005;65:5096–104. doi: 10.1158/0008-5472.CAN-04-4232. [DOI] [PubMed] [Google Scholar]
- 15.Bardella C, Pollard PJ, Tomlinson I. SDH mutations in cancer. Biochim Biophys Acta. 2011;1807:1432–43. doi: 10.1016/j.bbabio.2011.07.003. [DOI] [PubMed] [Google Scholar]
- 16.Petropoulos AE, Luetje CM, Camarata PJ, et al. Genetic analysis in the diagnosis of familial paragangliomas. Laryngoscope. 2000;110:1225–9. doi: 10.1097/00005537-200007000-00030. [DOI] [PubMed] [Google Scholar]
- 17.Hoekstra AS, Bayley JP. The role of comlex II in disease. Biochim Biophys Acta. 2013;1827:543–51. doi: 10.1016/j.bbabio.2012.11.005. [DOI] [PubMed] [Google Scholar]
- 18.Brito JP, Asi N, Bancos I, et al. Testing for germline mutations in sporadic pheochromocytoma/paraganglioma: a systematic review. Clin Endocrinol (Oxf) 2015;82:338–45. doi: 10.1111/cen.12530. [DOI] [PubMed] [Google Scholar]
- 19.Ricketts CJ, Forman JR, Rattenberry E, et al. Tumor risks and genotype–phenotype–proteotype analysis in 358 patients with germline mutations in SDHB and SDHD. Hum Mutat. 2010;31:41–51. doi: 10.1002/humu.21136. [DOI] [PubMed] [Google Scholar]
- 20.Ricketts C, Woodward ER, Killick P, et al. Germline SDHB mutations and familial renal cell carcinoma. J Natl Cancer Inst. 2008;100:1260–2. doi: 10.1093/jnci/djn254. [DOI] [PubMed] [Google Scholar]
- 21.Schimke RN, Collins DL, Stolle CA. Paraganglioma, neuroblastoma, and a SDHB mutation: resolution of a 30-year-old mystery. Am J Med Genet A. 2010;152A:1531–5. doi: 10.1002/ajmg.a.33384. [DOI] [PubMed] [Google Scholar]
- 22.Henderson A, Douglas F, Perros P, et al. SDHB-associated renal oncocytoma suggests a broadening of the renal phenotype in hereditary paragangliomatosis. Fam Cancer. 2009;8:257–60. doi: 10.1007/s10689-009-9234-z. [DOI] [PubMed] [Google Scholar]
- 23.Kim E, Rath EM, Tsang VH, et al. Structural and functional consequences of succinate dehydrogenase subunit B mutations. Encocr Relat Cancer. 2015;22:387–97. doi: 10.1530/ERC-15-0099. [DOI] [PubMed] [Google Scholar]
- 24.Janeway KA, Kim SY, Lodish M, et al. Defects in succinate dehydrogenase in gastrointestinal stromal tumors lacking KIT and PDGFRA mutations. Proc Natl Acad Sci U S A. 2011;108:314–8. doi: 10.1073/pnas.1009199108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Miettinen M, Sarlomo-Rikala M, McCue P, et al. Mapping of succinate dehydrogenase losses in 2258 epithelial neoplasms. Appl Immunohistochem Mol Morphol. 2014;22:31–6. doi: 10.1097/PAI.0b013e31828bfdd3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Dai Z, Pan S, Chen C, et al. Down-regulation of succinate dehydrogenase subunit B and up-regulation of pyruvate dehydrogenase kinase 1 predicts poor prognosis in recurrent nasopharyngeal carcinoma. Tumour Biol. 2016;37:5145–52. doi: 10.1007/s13277-015-4107-6. [DOI] [PubMed] [Google Scholar]
- 27.Zhang D, Wang W, Xiang B, et al. Reduced succinate dehydrogenase B expression is associated with growth and de-differentiation of colorectal cancer cells. Tumour Biol. 2013;34:2337–47. doi: 10.1007/s13277-013-0781-4. [DOI] [PubMed] [Google Scholar]
- 28.Chen L, Liu T, Zhang S, et al. Succinate dehydrogenase subunit B inhibits the AMPK-HIF-1α pathway in human ovarian cancer in vitro. J Ovarian Res. 2014;7:115. doi: 10.1186/s13048-014-0115-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Xiang Z, Yuan W, Luo N, et al. A novel human zinc finger protein ZNF540 interacts with MVP and inhibits transcriptional activities of the ERK signal pathway. Biochem Biophys Res Commun. 2006;347:288–96. doi: 10.1016/j.bbrc.2006.06.076. [DOI] [PubMed] [Google Scholar]
- 30.Li Y, Wang Y, Zhang C, et al. ZNF322, a novel human C2H2 Kruppel-like zinc-finger protein, regulates transcriptional activation in MAPK signaling pathways. Biochem Biophys Res Commun. 2004;325:1383–92. doi: 10.1016/j.bbrc.2004.10.183. [DOI] [PubMed] [Google Scholar]
- 31.Guillas I, Jiang JC, Vionnet C, et al. Human homologues of LAG1 reconstitute Acyl-CoA-dependent ceramide synthesis in yeast. J Biol Chem. 2003;278:37083–91. doi: 10.1074/jbc.M307554200. [DOI] [PubMed] [Google Scholar]
- 32.Winter E, Ponting CP. TRAM, LAG1 and CLN8: members of a novel family of lipid-sensing domains? Trends Biochem Sci. 2002;27:381–3. doi: 10.1016/s0968-0004(02)02154-0. [DOI] [PubMed] [Google Scholar]
- 33.Pewzner-Jung Y, Park H, Laviad E, et al. A critical role for ceramide synthase 2 in liver homeostasis: I. Alterations in lipid metabolic pathways. J Biol Chem. 2010;285:10902–10. doi: 10.1074/jbc.M109.077594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Park WJ, Brenner O, Kogot-Levin A, et al. Development of pheochromocytoma in ceramide synthase 2 null mice. Endocr Relat Cancer. 2015;22:623–32. doi: 10.1530/ERC-15-0058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Kusao I, Troelstrup D, Shiramizu B. Possible Mitochondria-Associated Enzymatic Role in Non-Hodgkin Lymphoma Residual Disease. Cancer Growth Metastasis. 2008;1:3–8. doi: 10.4137/cgm.s2046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Dbaibo GS, Pushkareva MY, Rachid RA, et al. p53-dependent ceramide response to genotoxic stress. J Clin Invest. 1998;102:329–39. doi: 10.1172/JCI1180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Obeid LM, Linardic CM, Karolak LA, et al. Programmed cell death induced by ceramide. Science. 1993;259:1769–71. doi: 10.1126/science.8456305. [DOI] [PubMed] [Google Scholar]
- 38.Bardella C, Pollard PJ, Tomlinson I. SDH mutations in cancer. Biochim Biophys Acta. 2022;1807:1432–43. doi: 10.1016/j.bbabio.2011.07.003. [DOI] [PubMed] [Google Scholar]
- 39.Selak MA, Armour SM, MacKenzie ED, et al. Succinate links TCA cycle dysfunction to oncogenesis by inhibiting HIF-alpha prolyl hydroxylase. Cancer Cell. 2005;7:77–85. doi: 10.1016/j.ccr.2004.11.022. [DOI] [PubMed] [Google Scholar]
- 40.Kaelin WG, Jr, Ratcliffe PJ. Oxygen sensing by metazoans: the central role of the HIF hydroxylase pathway. Mol Cell. 2008;30:393–402. doi: 10.1016/j.molcel.2008.04.009. [DOI] [PubMed] [Google Scholar]
- 41.Lee S, Nakamura E, Yang H, et al. Neuronal apoptosis linked to EglN3 prolyl hydroxylase and familial pheochromocytoma genes: developmental culling and cancer. Cancer Cell. 2005;8:155–67. doi: 10.1016/j.ccr.2005.06.015. [DOI] [PubMed] [Google Scholar]
- 42.Bishop T, Gallagher D, Pascual A, et al. Abnormal sympathoadrenal development and systemic hypotension in PHD3−/− mice. Mol Cell Biol. 2008;28:3386–400. doi: 10.1128/MCB.02041-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Chumakov PM. Function of the p53 gene: choice between life and death. Biochemistry (Mosc) 2000;65:28–40. [PubMed] [Google Scholar]
- 44.Zhang C, Liu G, Huang M. Ribonucleotide reductase metallocofactor: assembly, maintenance and inhibition. Front Biol (Beijing) 2014;9:104–13. doi: 10.1007/s11515-014-1302-6. [DOI] [PMC free article] [PubMed] [Google Scholar]


