Significance
How organisms maintain homeostasis when confronted with environmental stress is an important question in biology. The master nutrient response regulator target of rapamycin complex 1 (TORC1) regulates many progrowth cellular processes, including transcription of genes required for protein synthesis. RNA polymerase III (RNAPIII) plays a crucial role in regulation of protein synthesis by transcribing tRNA genes. RNAPIII activity is known to be dependent on TORC1, but the underlying molecular mechanisms remain to be fully elucidated. Here, we show that TORC1 promotes sumoylation of a specific set of proteins mainly involved in transcription. Subunits of RNAPIII, particularly Rpc82, are among the most prominent TORC1-dependent SUMO substrates. Mechanistically, we demonstrate that TORC1-dependent sumoylation of Rpc82 is required for efficient assembly and full activity of the RNAPIII holoenzyme.
Keywords: Sumo, transcription, RNA polymerase III, tRNA, TORC1
Abstract
Maintaining cellular homeostasis under changing nutrient conditions is essential for the growth and development of all organisms. The mechanisms that maintain homeostasis upon loss of nutrient supply are not well understood. By mapping the SUMO proteome in Saccharomyces cerevisiae, we discovered a specific set of differentially sumoylated proteins mainly involved in transcription. RNA polymerase III (RNAPIII) components, including Rpc53, Rpc82, and Ret1, are particularly prominent nutrient-dependent SUMO targets. Nitrogen starvation, as well as direct inhibition of the master nutrient response regulator target of rapamycin complex 1 (TORC1), results in rapid desumoylation of these proteins, which is reflected by loss of SUMO at tRNA genes. TORC1-dependent sumoylation of Rpc82 in particular is required for robust tRNA transcription. Mechanistically, sumoylation of Rpc82 is important for assembly of the RNAPIII holoenzyme and recruitment of Rpc82 to tRNA genes. In conclusion, our data show that TORC1-dependent sumoylation of Rpc82 bolsters the transcriptional capacity of RNAPIII under optimal growth conditions.
In yeast and in more complex eukaryotes, cell growth is restricted by the rate of mRNA translation and ribosome biogenesis, which depend on the transcription of ribosomal protein genes (RPGs), tRNAs and rRNAs. Synthesis of rRNA, tRNAs, and 5S rRNA represents 75% of total cellular transcription, whereas transcription of RPGs corresponds to 50% of RNA polymerase II (RNAPII) initiation events (1). These processes consume a significant portion of the cell’s resources, making nutrient availability a limiting factor to cell growth and proliferation (2). The conserved rapamycin-sensitive target of rapamycin complex 1 (TORC1) is a master regulator of the cellular nutrient response (2, 3). Under nitrogen-rich conditions, TORC1 promotes growth-related processes, like protein synthesis, ribosome biogenesis, and tRNA synthesis, while inhibiting catabolic processes, like autophagy (2). Conversely, inhibition of TORC1 activity by nitrogen depletion (or addition of the TORC1 inhibitor rapamycin) results in a metabolic switch from anabolism to catabolism, which involves many cellular processes, including down-regulation of transcription of RPGs, rRNA and tRNA genes (2, 3).
A key downstream target of TORC1 in regulation of tRNA transcription is the conserved RNAPIII inhibitor Maf1, which is phosphorylated and maintained in the cytoplasm under nitrogen-rich conditions (2, 3). Maf1 becomes hypophosphorylated under conditions that inhibit TORC1, allowing it to enter the nucleus where it associates with TFIIIB. The interaction between Maf1 and TFIIIB prevents the recruitment of RNAPIII and precludes transcription reinitiation at 5S rRNA and tRNA genes (4, 5). However, expression of an unphosphorylatable maf1 mutant does not completely repress tRNA expression in nutrient-replete cells (6), suggesting that dephosphorylation of Maf1 alone is not sufficient to fully inhibit RNAPIII. Indeed, inhibition of TORC1 also results in phosphorylation of the RNAPIII subunit Rpc53 by the kinases Kns1 and Mck1, which inhibits RNAPIII activity (7). Nonetheless, mutation of the Kns1/Mck1 phosphorylation sites on Rpc53 is not sufficient to completely prevent inactivation of RNAPIII by rapamycin (7), indicating that additional posttranslational modifications may exist that regulate RNAPIII under changing nutrient conditions.
The ubiquitin family member SUMO has important functions in maintaining cell homeostasis (8). For instance, we recently discovered that SUMO is important for transcription of nutrient-dependent genes, such as RPGs (9, 10). In the present study, we analyzed the plasticity of the SUMO proteome and the genome-wide localization of SUMO on chromatin in nutrient-rich and nutrient-poor conditions. We demonstrate that TORC1-dependent sumoylation of the RNAPIII subunit Rpc82 promotes assembly of the RNAPIII complex to stimulate tRNA transcription under optimal growth conditions.
Results
Starvation-Induced Remodeling of the SUMO Proteome.
To better understand the function of SUMO in regulation of nutrient responses, we incubated cells in nitrogen-rich and nitrogen-limiting media and purified SUMO under denaturing conditions. We found that nitrogen starvation resulted in an apparent increase in several SUMO conjugates (Fig. 1A), suggesting that nitrogen starvation alters the SUMO proteome. To gain further insight into starvation-induced SUMO proteome remodeling, we analyzed SUMO conjugates by mass spectrometry (MS). We found that 226 proteins were significantly enriched (group 1) and 76 proteins were significantly depleted (group 2) from SUMO pull-downs upon starvation (Dataset S1). Gene ontology (GO) analysis revealed that the proteins belonging to group 1 are enriched for transcriptional regulators (Fig. S1A and Dataset S1), such as the transcriptional activators Gcn4 and Rap1 and the transcriptional repressors Tup1 and Cyc8 (Fig. 1B), which we and others have previously shown to be SUMO targets (10–12). The vast majority of proteins in group 2 are involved in cellular processes that ultimately impinge upon mRNA translation (Fig. 1C and Dataset S1), including several subunits of RNAPIII, such as Rpc53, Rpc82, and Ret1 (Fig. 1B). These data show that nutritional status has a major impact on the SUMO proteome, particularly on sumoylation of proteins that regulate the cell’s translational capacity.
Fig. 1.
Inhibition of TORC1 alters the sumoylated protein landscape. (A) Nitrogen starvation alters the profile of protein sumoylation. Sumoylated proteins were purified under denaturing conditions from cells expressing HIS6-FLAG–tagged SUMO (6HF-SMT3, which encodes yeast SUMO) and analyzed by Western blotting using SUMO antibodies. (B) Differential sumoylation of proteins in response to nitrogen starvation. Sumoylation of Cyc8, Tup1, and Rap1 was increased, whereas sumoylation of RNAPIII components Rpc53, Rpc82, and Ret1 was decreased upon nitrogen starvation. (C) GO analysis showing that proteins that are desumoylated following nitrogen stress and are primarily involved in mRNA translation. The size of the nodes indicates the relative number of proteins associated with that specific GO term; the width of the edges indicates the relative number of proteins in common between the GO terms. (D) Differential sumoylation of proteins in response to inhibition of TORC1. Sumoylation of RNAPIII components Rpc53, Rpc82, and Ret1 was decreased upon rapamycin treatment. (E) Validation of the MS data. SUMO pull-downs were performed under denaturing conditions using cell lysates of cells grown under nitrogen-rich or nitrogen-starved conditions (Starv.), and the levels of copurifying Rpc82-HA, Rpc53-HA, and Ret1-HA were analyzed by Western blotting with anti-HA antibodies. Rpc82-HA signals (Lower graphs) were quantified using ImageJ. Error bars indicate the deviation from the average of two experiments. (F) Validation of MS data. Cells were treated with rapamycin (Rapa), and cell lysates were analyzed as in E.
Fig. S1.
Nitrogen starvation and rapamycin treatment remodel the Sumo proteome. (A) GO analysis of the proteins that are more highly sumoylated in response to nitrogen stress reveals overrepresentation of transcriptional regulators. The size of the nodes indicates the relative number of proteins associated with that specific GO term; the width of the edges indicates the relative number of proteins in common between the GO terms. (B) Rapamycin treatment induces increased sumoylation of the bulk of cellular proteins. Cells expressing untagged SUMO (SMT3) or His6-FLAG–tagged SUMO (HF-SMT3) were grown to log phase and treated with either DMSO or 100 nM rapamycin for 30 min, and global protein sumoylation was analyzed as described in Fig. 1A.
Nitrogen starvation is well known to inhibit TORC1 (13). To directly test whether rewiring of the SUMO proteome depends on TORC1 activity, we treated cells with the TORC1 inhibitor rapamycin. Interestingly, this resulted in a similar change in global sumoylation as observed with nitrogen starvation (Fig. S1B). To identify the proteins that are differentially sumoylated upon inhibition of TORC1, we performed a quantitative MS experiment using stable isotope labeling with amino acids in cell culture (SILAC). Consistent with the nitrogen starvation data, Rpc82, Rpc53, and Ret1 were all depleted from the SUMO fraction upon rapamycin treatment (Fig. 1D and Dataset S2). Validation of the MS data by Western blotting confirmed that sumoylation of Rpc82 and Ret1, and to a lesser degree also Rpc53, was substantially decreased upon nitrogen starvation (Fig. 1E). Similar results were obtained with the TORC1 inhibitor rapamycin (Fig. 1F).
Taken together, these data show that TORC1 has an important function in shaping the SUMO proteome and that several proteins involved in transcription are sumoylated in a TORC1-dependent manner, including subunits of RNAPIII.
TORC1 Promotes Recruitment of SUMO to tRNA Genes.
To begin unraveling the physiological relevance of TORC1-dependent sumoylation in regulation of transcription, we first studied the global chromatin localization of SUMO by reanalyzing our previously published ChIP-seq dataset (10). Consistent with our finding that RNAPIII is a TORC1-dependent SUMO target, we found that SUMO is present at nearly all RNAPIII-transcribed genes, including the 275 genes encoding tRNAs (tDNA) as well as noncoding RNAs such as SCR1 and SNR52 (Fig. 2A, Upper, gray) (9). Meta-analysis of this dataset showed that SUMO overlaps the transcription unit of tRNA genes, forming a peak right at the transcription start site (TSS; Fig. 2B), resembling previously published meta-analyses of the chromatin localization of RNAPIII and its nascentome (14, 15). These high-throughput data were validated by ChIP followed by quantitative PCR (ChIP-qPCR) using primers specific for a panel of RNAPIII-transcribed genes (Fig. S2A).
Fig. 2.
SUMO is enriched at tRNA genes and regulates tRNA transcription. (A) Snapshots of SUMO ChIP-seq experiments (Upper, gray) and sRNA-seq experiments (Lower, yellow) in WT and ubc9-1 cells grown at permissive temperature and treated with either DMSO or rapamycin. “Class II genes” indicates genes that are transcribed by RNAPII, whereas “class III genes” indicates genes transcribed by RNAPIII. (B) Meta-analysis of SUMO levels at tRNA genes. Rapa, rapamycin; TSS, TSS. (C and D) Global expression levels of tRNAs (C) and other short RNAs (D) in WT and ubc9-1 cells grown at permissive temperature and treated with either DMSO or 100 nM of rapamycin (Rapa) for 30 min.
Fig. S2.
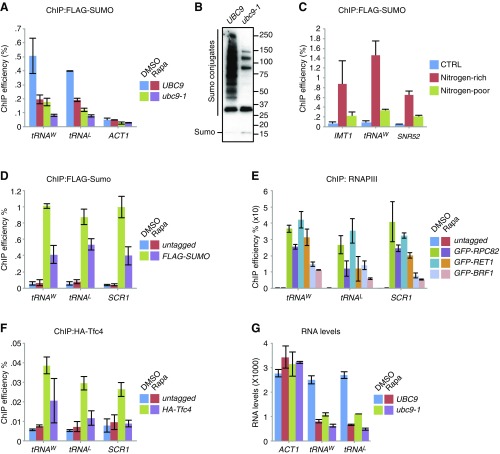
The Ubc9–SUMO pathway is required for efficient tRNA transcription. (A) Enrichment of SUMO at tRNA genes is dependent on TORC1 activity. WT cells and ubc9-1 mutants (both expressing 6HF-SMT3) were grown at permissive temperature to log phase and treated with either DMSO or 100 nM rapamycin (Rapa) for 30 min, and the relative level of SUMO at the indicated genes was determined by ChIP-qPCR. (B) Protein sumoylation is strongly reduced in ubc9-1 mutants. WT cells and ubc9-1 mutants (both expressing 6HF-SMT3) were grown at permissive temperature to log phase, after which global protein sumoylation was analyzed as described in Fig. 1A. (C) Enrichment of SUMO at tRNA genes is dependent on nitrogen availability. WT cells expressing either untagged SMT3 (CTRL) or 6HF-SMT3 were grown to log phase and either incubated in media supplemented with nitrogen and amino acids (Nitrogen-rich) or in media lacking these nitrogen sources (Nitrogen-poor), after which the relative level of SUMO at the indicated genes was determined by ChIP-qPCR. (D–F) The enrichment of the RNAPIII holoenzyme at class III genes is dependent on TORC1 activity. Indicated strains were grown to log phase, and the relative levels of 6HF-SUMO (D); GFP-Rpc82, GFP-Ret1, and GFP-Brf1 (E); and HA-Tfc4 (F) at tRNAL, tRNAW, and SCR1 were determined by ChIP-qPCR using antibodies against FLAG, HA, and GFP, respectively. Error bars, SEM of three independent experiments. (G) tRNA transcription is significantly reduced in ubc9-1 mutants. WT cells or ubc9-1 mutants were treated with DMSO of 100 nM of rapamycin for 30 min, after which RNA was purified and analyzed by qPCR using primer pairs against the indicated genes. Error bars, SEM of three independent experiments.
To further validate our data, we reanalyzed our previous SUMO ChIP-seq experiments that were performed with a temperature-sensitive ubc9-1 mutant (10). The ubc9-1 allele encodes a destabilized form of Ubc9 (16), resulting in a strong reduction in sumoylated protein levels even at the permissive temperature of 30 °C (Fig. S2B). Importantly, the levels of SUMO at RNAPIII-transcribed genes were significantly reduced in ubc9-1 cells compared with wild-type (WT) cells (Fig. 2 A and B and Fig. S2A). This shows that the SUMO ChIP-seq data are specific and not caused by a hyper-ChIPable chromatin artifact (17).
Our MS data showed that several subunits of RNAPIII are desumoylated upon nitrogen starvation or TORC1 inhibition, strongly suggesting that RNAPIII is a critical TORC1-dependent SUMO target at tRNA genes. Consistent with this hypothesis, SUMO ChIP-seq experiments using WT and ubc9-1 cells treated with rapamycin revealed that the level of SUMO at tRNA genes was reduced by ∼30% in ubc9-1 mutants compared with WT cells (Fig. 2 A and B). Treatment with rapamycin further reduced SUMO levels at tRNA genes, particularly in the ubc9-1 mutant. These data were confirmed by ChIP-qPCR (Fig. S2A). Nitrogen starvation also resulted in a reduction in SUMO levels at RNAPIII-transcribed genes (Fig. S2C).
Paradoxically, our MS data also showed that Brf1 and Tfc4 (subunits of TFIIIB and TFIIIC, respectively) are hypersumoylated upon rapamycin treatment (Dataset S1). This finding appeared to be contradictory to the weaker binding of SUMO to the tDNA detected in rapamycin-treated cells. However, it has been recently shown that hypersumoylation of the transcriptional regulator Gcn4 prevents its binding to the chromatin (18), and hypersumoylation of Brf1 and Tfc4 may similarly reduce their binding to chromatin. To test this, we analyzed the association of SUMO, Rpc82, Ret1, Brf1, and Tfc4 to class III genes by ChIP-qPCR after rapamycin treatment. We found not only that the level of SUMO was lower at tRNAW, tRNAL, and SCR1 but that the recruitment of Rpc82, Ret1, Brf1, and Tfc4 was also lower after inhibition of TORC1 (Fig. S2 D–F). These data suggest that SUMO regulates the recruitment of the RNAPIII transcriptional complex to regulate tRNA transcription.
To study the effect of TORC1 and the Ubc9SUMO pathway on transcriptional output of RNAPIII, we performed transcriptome analysis using short-RNA sequencing (sRNA-seq; <200 nt) (Fig. 2A, Lower, yellow and Dataset S3). We extracted tRNA genes from the sRNA expression dataset and found that global tRNA transcription was much lower in ubc9-1 cells than in WT cells (Fig. 2C), whereas the expression of the remaining sRNAs was not significantly affected (Fig. 2D). These data show that sumoylation is required for efficient tRNA transcription under optimal growth conditions. Interestingly, the strong decrease in tRNA transcription observed in the ubc9-1 mutant was not further reduced by treatment with rapamycin (Fig. 2C), which supports our finding that Ubc9–SUMO and TORC1 function in the same pathway. RT-qPCR using primers specific for the immature forms of tRNAL and tRNAW confirmed these data, whereas the expression of ACT1, which is regulated by neither SUMO nor TORC1 (10), was unchanged (Fig. S2G).
Together, these data show that both TORC1 and Ubc9–SUMO promote tRNA transcription and suggest that they ultimately act on the same downstream target.
Sumoylation of Rpc82 Promotes Transcription of tRNA Genes.
To gain insight into the regulation of RNAPIII by SUMO, we first mapped the lysines in Rpc82, Ret1, and Rpc53 to which SUMO is conjugated (sumoylation of the RNAPIII basal transcription factors Tfc4 and Brf1 will be the topic of future studies). Using GPS-SUMO (19), we identified multiple potential SUMO acceptor sites (Table S1). We mutated these lysines into nonsumoylatable arginine residues and determined the sumoylation status of these mutants in vivo. Interestingly, lysine-to-arginine substitution of K403, K406, K591, and K596 in Rpc82 (Rpc82-4KR; Fig. 3A) completely abolished Rpc82 sumoylation (Fig. 3B), with a migration profile that was identical to WT Rpc82 expressed in a ubc9-1 mutant (Fig. 3C). Importantly, transcription of tRNA genes was reduced by ∼50% in rpc82-4KR cells compared with RPC82 cells (Fig. 3D), showing that sumoylation of Rpc82 is important for RNAPIII activity. Treatment with rapamycin did not further reduce tRNA transcription in the rpc82-4KR mutant (Fig. 3D), indicating that Ubc9–SUMO acts on Rpc82 either downstream of or in parallel to TORC1 to control tRNA expression. This interpretation is supported by our finding that deletion of MAF1 did not restore tRNA transcription in the rpc82-4KR mutant (Fig. 3E), strongly suggesting that SUMO exerts its effect on tRNA transcription downstream of Maf1.
Table S1.
SUMO sites predicted by GPS-SUMO
| Protein | Site | Sequence | Score | Type of SUMO site |
| Rpc82 | 403 | LPSKKLK | 2.021 | Nonconsensus |
| Rpc82 | 406 | KKLKTED | 3.588 | Ψ-K-X-E |
| Rpc82 | 591 | LLFKKEK | 1.005 | Ψ-K-X-E |
| Rpc82 | 596 | EKLKQEN | 3.886 | Ψ-K-X-E |
| Ret1 | 17 | KHVKDEA | 2.848 | Ψ-K-X-E |
| Ret1 | 448 | KRFKMER | 1.896 | Ψ-K-X-E |
| Ret1 | 927 | IIVKQED | 3.483 | Ψ-K-X-E |
| Rpc53 | 24 | PSLKDSS | 3.191 | Nonconsensus |
| Rpc53 | 51 | SKVKLEE | 3.417 | Ψ-K-X-E |
| Rpc53 | 115 | GFIKSEG | 1.403 | Ψ-K-X-E |
| Rpc53 | 216 | ETVKREI | 1.919 | Ψ-K-X-E |
| Rpc53 | 236 | PSVKTEP | 5.365 | Ψ-K-X-E |
| Rpc53 | 322 | PAVKEEK | 3.924 | Ψ-K-X-E |
| Rpc53 | 325 | KEEKEDM | 3.926 | Nonconsensus |
Fig. 3.
Sumoylation of Rpc82 promotes tRNA transcription. (A) Location of the putative SUMO sites in Rpc82. CC, coiled-coil domain; eWH, extended winged helix domain. Protein domain organization was derived from the structure of human RPC3 (31). (B) Rpc82-4KR is not sumoylated in vivo. Cells expressing RPC82-HA or rpc82-4KR-HA as well as HIS6-FLAG–tagged SUMO were treated with DMSO or 100 nM rapamycin (Rapa) for 30 min, after which SUMO was purified and eluates were analyzed by Western blotting with HA antibodies. Dashed line indicates where the image was cropped to remove irrelevant data. (C) Rpc82 sumoylation is abolished in the ubc9-1 mutant. WT cells or ubc9-1 mutants (both expressing normal RPC82-HA and 6HF-SMT3) were grown to log phase at permissive temperature and treated with either DMSO or 100 nM rapamycin (Rapa) for 30 min. SUMO was then purified and eluates were analyzed as in B. (D) tRNA transcription is significantly reduced in rpc82-4KR mutants. WT cells or rpc82-4KR mutants were grown and treated as in B, after which RNA was purified and analyzed by qPCR using primer pairs against the indicated genes. Error bars, SEM of three independent experiments. (E) Deletion of MAF1 does not restore tRNA transcription in rpc82-4KR mutants. The indicated strains were grown to log phase and treated with either DMSO or 100 nM rapamycin (Rapa), after which RNA was purified and analyzed by qPCR as in D. Error bars, SEM of three independent experiments.
In contrast to our findings with Rpc82, mutating the predicted SUMO sites in Rpc53 and Ret1 did not prevent, or only partially reduced, sumoylation of these proteins in vivo (Fig. S3 A and B). Accordingly, tRNA transcription in these mutants was either unaffected (rpc53-K24,51,115,216,236,322,325; “rpc53-7KR” in Fig. S3C) or apparently increased (ret1-K17,448,927R; “ret1-3KR” in Fig. S3D) compared with WT cells. The critical SUMO sites and the physiological significance of sumoylation of Rpc53 and Ret1 remain to be characterized, which is the aim of future studies.
Fig. S3.
Mutating potential sumoylation sites in Rpc53 and Ret1 does not alter RNAPIII activity. (A) Mutations of putative SUMO sites do not abolish Rpc53 sumoylation. WT and ubc9-1 cells expressing RPC53-HA or rpc53-7KR-HA as well as HIS6-FLAG-SMT3 were treated with DMSO or 100 nM of rapamycin for 30 min; SUMO was purified and eluates were analyzed by Western blotting with HA antibodies. (B) Mutations of putative SUMO sites do not fully abolish Ret1 sumoylation. WT and ubc9-1 cells expressing RET1-HA or ret1-3KR-HA as well as HIS6-FLAG-SMT3 were treated with DMSO or 100 nM of rapamycin for 30 min, after which cell lysates were analyzed as in A. (C) tRNA synthesis is not affected in cells expressing rpc53-7KR. WT cells and rpc53-7KR mutants were grown and treated as in A, after which RNA was purified and analyzed by qPCR using primer pairs against the indicated genes. Error bars, SEM of three independent experiments. (D) Stimulation of tRNA synthesis in cells expressing ret1-3KR. WT cells and ret1-3KR mutants were grown and treated as in A, after which RNA was purified and analyzed by qPCR using primer pairs against the indicated genes. Error bars, SEM of three independent experiments.
Together, these data show that sumoylation of Rpc82 is important for tRNA transcription. We decided to focus on Rpc82 sumoylation for the remainder of this study, because the rpc82-4KR phenotype most closely resembled the severe tRNA transcription defect observed in the ubc9-1 mutant.
Sumoylation of Rpc82 Stabilizes the RNAPIII Transcriptional Complex.
One emerging function of SUMO is to promote the cohesion of entire protein complexes (9, 20). We therefore hypothesized that sumoylation of Rpc82 promotes stability of the RNAPIII complex. To test this hypothesis, we first purified WT Rpc82 and Rpc82-4KR from cells grown under nutrient-rich conditions and analyzed the associated proteins by quantitative MS. Interestingly, this experiment revealed that the interactions between Rpc82-4KR and other components of the RNAPIII enzyme, as well as the basal transcription factor TFIIIB, were strongly reduced (Fig. 4A and Dataset S4), suggesting that sumoylation of Rpc82 is important for integrity of the RNAPIII complex. These MS data were confirmed by coimmunoprecipitation experiments, which showed a substantial reduction in the interaction of Rpc82-4KR with Brf1-GFP and with Ret1-GFP compared with WT Rpc82 (Fig. 4 B and C). Furthermore, ChIP-qPCR experiments demonstrated that Rpc82-4KR was specifically lost from the chromatin at tRNAL and tRNAW genes, whereas the presence of other components of the RNAPIII complex, such as Ret1, was unaffected in the rpc82-4KR mutant (Fig. 4 D and E); this specific reduction in chromatin-bound Rpc82-4KR was not due to a nuclear entry defect or a defect in Rpc82-4KR protein stability, because Rpc82-4KR entered the nucleus normally (Fig. S4A) and the stability of Rpc82-4KR was comparable to that of WT Rpc82 (Fig. S4 B and C). These data show that the weaker interaction of Rpc82-4KR with the RNAPIII holoenzyme is not due to a trafficking defect or a defect with Rpc82-4KR protein stability.
Fig. 4.
Sumoylation of Rpc82 promotes its interaction with RNAPIII and TFIIIB. (A) Reduced binding of Rpc82-4KR to RNAPIII and TFIIIB. Cells were grown to log phase, and equal amounts of Rpc82-HA and Rpc82-4KR-HA were purified with HA antibodies in the presence of NEM to prevent desumoylation. Copurifying proteins were analyzed by quantitative MS. (B) Sumoylation of Rpc82 is important for the interaction with Brf1. Rpc82-HA and Rpc82-4KR-HA were purified with HA antibodies under native conditions in the presence of NEM, and the amount of copurifying Brf1-GFP was analyzed by Western blotting with GFP antibodies. GFP over HA ratios were determined by quantifying Western blot signals with ImageJ. Error bars indicate the deviation from the mean of two independent experiments. (C) Sumoylation of Rpc82 promotes the interaction with Ret1. Rpc82-HA and Rpc82-4KR-HA were purified from cells as described in B, and the amount of copurifying Ret1-GFP was analyzed by Western blotting with GFP antibodies. GFP over HA ratios were determined as in B. Error bars indicate the deviation from the mean of two independent experiments. (D and E) Sumoylation of Rpc82 promotes its recruitment to tRNAL. RPC82-HA RET1-GFP cells and rpc82-4KR-HA RET1-GFP mutants were grown to log phase, and the relative level of Rpc82 or Rpc82-4KR and Ret1 at tRNAL (D) or tRNAW (E) was determined by ChIP-qPCR using antibodies against HA and GFP, respectively. Error bars, SEM of three independent experiments. (F) Rpc82 SUMO sites in the context of the structure of RNAPIII. K401 and K406 lie in a low-density flexible loop of Rpc82 WH3 domain. K591 lies in the coiled-coil domain of Rpc82 and faces outwards in a cavity formed by Rpc160, Rpc82, Rpc34, Rpc31, and Rpc25. K596 lies under a low-density (flexible) domain of Rpc31. (G) The putative positioning of SUMO density at K591 (purple) suggests that SUMO may stabilize the active conformation of RNAPIII. SUMO may either coordinate or stabilize the multidomain cavity formed by Rpc160, Rpc82, Rpc34, Rpc31, and Rpc25, potentially favoring the closed-clamp (active) conformation of RNAPIII. (H) rpc82-4KR cells are sensitive to heat stress. WT and rpc82-4KR cells were spotted on plates supplemented and were incubated at 30 °C or 37 °C until colonies appeared.
Fig. S4.
Rpc82 K596 is occluded by a flexible part of Rpc31. (A) Normal trafficking of Rpc82-KR. rpc82Δ cells transformed with GFP-RPC82 or GFP-rpc82-4KR were grown in dropout medium to log phase and further incubated 30 min with DAPI (100 ng/mL). Cells were gently harvested by centrifugation and washed once with PBS before being prepared for microscopy. (B) Rpc82-HA and Rpc82-4KR-HA were purified with HA antibodies and resolved by SDS/PAGE. The presence of ubiquitin and Rpc82 was detected by anti-ubiquitin (WB:Ub) and anti-HA (WB:HA) Western blots. (C) Normal stability of Rpc82-4KR. WT and rpc82-4KR cells were grown in YPD to log phase and treated with 0.2 mg/mL of cycloheximide (CHX). At indicated times, the presence of Rpc82-HA and Rpc82-4KR-HA was detected by anti-HA Western blot. As a loading control, Cdc11 protein levels were assessed by anti-Cdc11 Western blot. HA signals were determined by quantifying Western blot signals with ImageJ and were normalized to time point “0.” Error bars represent the mean of two independent experiments. (D) Cartoon showing that K596 is occluded by a low-density (flexible) loop of Rpc31.
Together, these data indicate that Rpc82 sumoylation is important for efficient incorporation of Rpc82 into the RNAPIII holoenzyme.
Discussion
RNAPIII activity is tightly regulated by the cell. Compared with RNAPII, which is regulated by a plethora of modifications in its C-terminal domain (21), little is known about regulation of RNAPIII activity. Several types of stress signals result in inhibition of RNAPIII activity, including nutrient stress and DNA damage, and most of these signals appear to converge on Maf1 (3). However, it has recently become clear that RNAPIII activity is also directly regulated by the cell; for instance, TORC1 regulates Rpc53 activity by preventing the direct, inhibitory phosphorylation of Rpc53 by Kns1 and Mck1 (7). Our finding that the Ubc9–SUMO pathway directly controls RNAPIII activity, which occurs in parallel to or downstream of Maf1 (Fig. 3E), reveals a layer of complexity of RNAPIII regulation. Our findings are supported by previous high-throughput proteomic studies that showed that Rpc82 and Rpc53, as well as subunits of TFIIIB, are SUMO targets (22, 23), although the function of SUMO in regulation of these proteins was not characterized in these studies. Here, we have demonstrated that TORC1-dependent sumoylation of Rpc82 is required for efficient tRNA transcription under optimal growth conditions.
We found that sumoylation of Rpc82 is important for its incorporation into the RNAPIII holoenzyme. Interestingly, K403 and K406 lie in an 80 residue-long flexible loop of Rpc82 (Fig. 4F), which could serve as a platform that stabilizes interactions either within RNAPIII or with the basal transcription factors of RNAPIII. Furthermore, K591 lies in the coiled-coil domain of Rpc82 and faces outward toward a cavity that is formed by multiple RNAPIII components, including C160, C25, C31, and C34 (Fig. 4G). It is possible that sumoylation of K591 in this multidomain cavity stabilizes the interaction of Rpc82 with the RNAPIII complex to promote the closed clamp conformation of RNAPIII, which may be important for promoter opening and subsequent elongation (24). K596 is situated in the coiled-coil region of Rpc82, which in structural analyses is surrounded by a low-resolution density of Rpc31, which occludes the precise position of SUMO, making it difficult to predict how sumoylation of this residue might affect the interaction of Rpc82 with RNAPIII (Fig. S4D). Finally, whereas K403, K406, and K591 are highly conserved in fungi, K596 is not, which suggests that sumoylation of K596 may not be important for regulation of RNAPIII. Our model in which sumoylation of Rpc82 promotes the cohesion of the RNAPIII holoenzyme is supported by the fact that rpc82-4KR cells are sensitive to elevated temperatures, which are known to destabilize protein complexes (Fig. 4H).
Although we have shown here that sumoylation of Rpc82 promotes RNAPIII activity, we cannot exclude the possibility that sumoylation of other RNAPIII components may have differential effects on RNAPIII activity. Indeed, we found that mutation of the putative SUMO-targeted lysines in Ret1 appeared to promote rather than reduce the transcriptional activity of RNAPIII. Therefore, it is possible that different RNAPIII components may be modified by SUMO in response to specific intracellular and environmental signals and that the balance of sumoylation of different RNAPIII subunits ultimately determines its transcriptional output. Nonetheless, under optimal growth conditions SUMO has a net positive effect on RNAPIII, as evidenced by our finding that RNAPIII activity is strongly reduced in sumoylation-defective ubc9-1 mutants.
Taken together, we have discovered a mechanism for regulation of RNAPIII during changing nutrient conditions.
Methods
Yeast Strains and Plasmids.
S. cerevisiae strains were grown in appropriate media, depending on the experiment/genotype. Strains were derived directly from either the S288c strains RDKY3615 (25) or BY4741 using standard gene-replacement methods or intercrossing (Table S2 for strains and plasmids).
Table S2.
Strains and plasmids used in this study
| Strain no. | Relevant genotype | Source |
| BY4741 | MATa, his3Δ1, leu2Δ0, met15Δ, ura3Δ0 | (37) |
| RDKY3615 | MATa, ura3-52, leu2Δ1, trp1Δ63, his3-200, lys2-Bgl, hom3-10, ade2Δ1, ade8, YEL069C::URA3 | (25) |
| JEY9454 | RDKY3615 ubc9::NAT [pSE362-UBC9, HIS3] HIS6-FLAG-SMT3:KAN | (10) |
| JEY9458 | RDKY3615 ubc9::NAT [pSE362-ubc9-1, HIS3] HIS6-FLAG-SMT3:KAN | (10) |
| HZY1017 | BY4741 HIS6-FLAG-SMT3:KAN | (38) |
| JEY10612 | BY4741 ubc9::NAT [pSE362-UBC9, HIS3] , RPC82-HIS6-MYC2:HIS3 | This study |
| JEY10612 | BY4741 ubc9::NAT [pSE362-ubc9-1, HIS3] , RPC82-HIS6-MYC2:HIS3 | This study |
| JEY12038 | BY4741 HIS6-FLAG-SMT3:KAN, rpc53::NAT [pAG415-GPD_pr-HA-RPC53, LEU2] | This study |
| JEY12039 | BY4741 HIS6-FLAG-SMT3:KAN, rpc53::NAT [pAG415-GPD_pr-HA-rpc53-7KR, LEU2] | This study |
| JEY12045 | BY4741 HIS6-FLAG-SMT3:KAN, ret1::NAT [pAG415-GPD_pr-HA-RET1, LEU2] | This study |
| JEY12047 | BY4741 HIS6-FLAG-SMT3:KAN, ret1::NAT [pAG415-GPD_pr-HA-ret1-3KR, LEU2] | This study |
| JEY12061 | BY4741 HIS6-FLAG-SMT3:KAN, rpc82::NAT [pAG415-GPD_pr-HA-RPC82, LEU2] | This study |
| JEY12061 | BY4741 HIS6-FLAG-SMT3:KAN, rpc82::NAT [pAG415-GPD_pr-HA-rpc82-4KR, LEU2] | This study |
| JEY12068 | BY4741 HIS6-FLAG-SMT3:KAN, rpc82::NAT [pAG415-GPD_pr-HA-RPC82, LEU2], maf1::hph | This study |
| JEY12075 | BY4741 HIS6-FLAG-SMT3:KAN, rpc82::NAT [pAG415-GPD_pr-HA-rpc82-4KR, LEU2], maf1::hph | This study |
| JEY12079 | BY4741 rpc82::NAT [pAG415-GPD_pr-HA-RPC82, LEU2] RET1-GFP:HIS3 | This study |
| JEY12081 | BY4741 rpc82::NAT [pAG415-GPD_pr-HA-rpc82-4KR, LEU2] RET1-GFP:HIS3 | This study |
| JEY12092 | BY4741 rpc82::NAT [pAG415-GPD_pr-HA-RPC82, LEU2] BRF1-GFP:HIS3 | This study |
| JEY12093 | BY4741 rpc82::NAT [pAG415-GPD_pr-HA-rpc82-4KR, LEU2] BRF1-GFP:HIS3 | This study |
| JEY12219 | BY4741 rpc82::NAT [pAG415-GPD_pr-GFP-RPC82, LEU2] | This study |
| JEY12221 | BY4741 rpc82::NAT [pAG415-GPD_pr-HA-rpc82-4KR, LEU2] | This study |
ChIP, ChIP-Seq, and Small RNA-Seq.
ChIP and short RNA-seq were performed as described previously (10, 26) with minor modifications (SI Methods). The original SUMO ChIP-seq dataset has been reported previously (10). For primer sequences, Table S3.
Table S3.
Primers for ChIP and for expression analysis
| Target | Sequence |
| ChIP | |
| IMT1 | GAAGCGCGCAGGGCTCATAA |
| TAGCGCCGCTCGGTTTCGAT | |
| tDNA(W) | GGAATTTCCAAGATTTAATTGGAGTCG |
| TGGCTCAATGGTAGAGCTTTCG | |
| tDNA(L) | GTCTAAGGCGCCTGATTCAAGA |
| GTTCACTGCGGTCAAGAT | |
| SNR52 | GCACCAACTTTCACTTCTACAGCG |
| TTTGTAGTGCCCTCTTGGGCTA | |
| ACT1 | CTCAAAGCAAGCGTGGTATTT |
| TCTTTTCCATATCATCCCAGTTG | |
| SCR1 | GAATTCTGGCCGAGGAACAAATCC |
| CAGCTCTGCCCAGGACAAATTTAC | |
| Expression analysis | |
| tDNA(L) | GTCTAAGGCGCCTGATTCAAGA |
| GTTCACTGCGGTCAAGAT | |
| tDNA(W) | GGAATTTCCAAGATTTAATTGGAGTCG |
| TGGCTCAATGGTAGAGCTTTCG | |
| ACT1 | GTACTAACATCGATTGCTTCA |
| AAACCGGCTTTACACATAC |
Analysis of Sequencing Data.
ChIP-seq data were analyzed as previously described (10). Short RNA-seq was performed in duplicates by BGI (www.genomics.cn/en/). Raw reads were filtered for low-quality reads, adapters, and contamination before reads were mapped to the R64-1–1 genome assembly using Tophat2 (v. 2.0.13) (27) using default options. The number of reads per gene in the Ensemble 75 were counted using featureCounts (v. 1.4.6-p3) (28) in paired-end mode and with –t «gene» and –M options. The counts were then normalized for sequencing depth by dividing each value by the total number of reads in million (generating counts per million, cpm). Raw data and expression values before and after normalization are available in Datasets S3 and S5. Expression values of RNAPIII-transcribed genes were extracted based on the gene list from the yeast database (www.yeastgenome.org/).
RNA Preparation and Reverse Transcription.
Total RNA purification and reverse transcription were performed as previously described (26).
SUMO Pull-Down Under Denaturing Conditions.
Pull-downs in denaturing conditions were performed as previously described (10, 11, 29) with minor modifications (SI Methods).
MS.
Label-free MS samples were processed at the Proteomics Core Facility at Oslo University Hospital–Rikshospitalet (www.med.uio.no/cir/english/services/proteomics/) (SI Methods). Briefly, cells were cultured until log phase in complete synthetic medium, washed, and incubated for 3 h in either complete synthetic medium or in synthetic drop-out medium lacking nitrogen and amino acids. Cells were harvested, washed with ice-cold water, and processed for purification of SUMO conjugates under denaturing conditions (SI Methods).
Modified SILAC experiments were performed and analyzed at the proteomic core facility at the EMBL in Heidelberg (https://www.embl.de/proteomics/proteomics_services/index.html). Briefly, protein extracts from DMSO- and rapamycin-treated cells were prepared in denaturing conditions (SI Methods). After digestion with LysC and trypsin, DMSO-treated samples were labeled light and rapamycin-treated samples were labeled heavy. Both samples were mixed and submitted to MALDI-TOF MS/MS (Dataset S2).
ChIP-MS experiments were performed as previously described (10) with minor modifications (SI Methods).
Cell Lysis, Immunoprecipitation, and Western Blotting.
IP and Western blotting was performed as previously described (26, 29, 30). All buffers contained 10 mM of fresh N-ethylmaleimide (Sigma) to prevent desumoylation by SUMO proteases.
For coimmunoprecipitations, log phase cells were collected and washed in lysis buffer [50 mM Hepes–KOH (pH 7.5), 150 mM NaCl, 1 mM EDTA, 0.05% Nonidet P-40, 0.5 mM DTT, 20% (vol/vol) glycerol, 10 mM NEM, protease inhibitors]. Pellets were resuspended in lysis buffer, and cells were lysed by vortexing with glass beads followed by centrifugation to remove cell debris. Equal amounts of proteins were used for immunoprecipitation using magnetic beads conjugated covalently to relevant antibodies. After extensive washing with lysis buffer, coimmunoprecipitated proteins were resolved by SDS/PAGE and analyzed by Western blotting with the indicated antibodies.
SI Methods
Sumo Pull-Down Under Denaturing Conditions for Western Blotting.
Cells (∼15 mL log phase cultures) were collected by centrifugation, washed with water, and flash-frozen in liquid nitrogen. The pellet was resuspended in lysis buffer (8 M urea, 50 mM Tris, pH 8.0, 0.05% SDS, 10 mM NEM, PIC), and cells were lysed by vortexing with glass beads. The cell lysate was then centrifuged to eliminate cell debris, and 5% of the supernatant was saved as input material. The rest of the supernatant was incubated with Ni-NTA agarose beads overnight at 4 °C. The resin was washed three times with wash buffer (8 M urea, 50 mM Tris, pH 8.0, 200 mM NaCl, 0.05% SDS, 5 mM imidazol). Sumoylated proteins were eluted from the resin by addition of loading buffer (8 M urea, 10 mM Mops, 10 mM EDTA, 1% SDS, 0.01% bromophenol blue, pH 6.8).
Sumo Pull-Down Under Denaturing Conditions for MS Analysis.
One liter of log-phase cultures of cells expressing the HIS6-FLAG–tagged protein of interested was grown until log phase, harvested by centrifugation, washed with cold water, and frozen in liquid nitrogen. The frozen pellet was resuspended in cold lysis buffer [1.85 M NaOH, 7.5% (vol/vol) β‐mercaptoethanol] to a final volume of 25 mL. After 20 min on ice, an equal volume of cold 55% trichloroacetic acid (TCA) was added and kept on ice for 20 min. Proteins were pelleted by centrifugation (20 min, 4 °C, 3,500 × g), and the pellet was washed with cold water. The pellet was resuspended in 25 mL of buffer A (6 M guanidine–HCl, 100 mM NaH2PO4, 10 mM Tris·HCl; adjusted to pH 8.0) supplemented with 0.05% Tween 20. The suspension was shaken for 1 h at room temperature. Insoluble material was removed by centrifugation (20 min, 4 °C, 15,000 × g), and the supernatant was transferred to a fresh tube and supplemented with imidazole to a final concentration of 10 mM. We added 150 µL of Ni‐NTA agarose beads (Qiagen) to the sample, and samples were incubated overnight at 4 °C under agitation. The beads were then washed with 35 bed volumes of buffer A supplemented with 0.05% Tween 20, 50 bed volumes of buffer C (8 M urea, 100 mM NaH2PO4, 10 mM Tris·HCl; adjusted to pH 6.3) supplemented with 0.05% Tween 20, and 50 bed volumes of buffer C. Proteins bound to the Ni‐NTA column were eluted in four subsequent steps with one bed volume of buffer C plus 250 mM imidazole, and the eluates were pooled. One volume of 55% TCA was added to the eluate and left on ice for 20 min before centrifugation (20 min, 4 °C, 3,500 × g). Proteins were resuspended in 25 µL of buffer HU [8 M urea, 5% (vol/vol) SDS, 200 mM Tris·HCl, pH 6.8, 1.5% (vol/vol) DTT] plus 10 µL of water and denatured for 10 min at 65 °C.
ChIP Assays.
Cells were fixed with 1% (vol/vol) formaldehyde for 30 min. Formaldehyde was quenched for 5 min by adding glycine to a final concentration of 125 mM. Cells were washed with cold Tris-buffered saline, resuspended in lysis buffer [50 mM Hepes–KOH (pH 7.5), 140 mM NaCl, 1 mM EDTA, 1% Triton X-100, 0.1% Na-deoxycholate, and protease inhibitor mixture], and sonicated twice for 15 min by performing alternating cycles of 30-s pulses followed by a 30-s cool-down period using a Diagenode Bioruptor Twin. After centrifugation, supernatants were immunoprecipitated at 4 °C. Beads were first washed with lysis buffer, followed by washing with lysis buffer containing 500 mM NaCl, then with wash buffer [10 mM Tris (pH 8.0), 250 mM LiCl, 0.5% Nonidet P-40, 0.5% Na-deoxycholate, and 1 mM EDTA], and finally with a buffer (pH 8.0) containing 10 mM Tris and 1 mM EDTA. Elutions were performed in 50 mM Tris (pH 8.0), 10 mM EDTA, and 1% (wt/vol) SDS at 65 °C, and the cross-link was reversed 5 h at 65 °C. After RNase and proteinase K treatment, DNA fragments were purified using QIAquick PCR purification kit (Qiagen) and analyzed by qPCR using an AB StepOnePlus machine (Applied Biosystems).
Gene Synthesis.
RET1; ret1-K17,448,927R; RPC82; rpc82-K403,406,591,596R; RPC53; and rpc53-K24,51,115,216,236,322,325R alleles were synthesized and subcloned into pDONR221 by Genscript and confirmed by sequencing.
ChIP–MS.
RPC82 and rpc82-4KR cells were processed as for a regular ChIP assay until the IP step (ChIP Assays). For this step, 700 µg of cross-linked material was mixed with 20 µL of HA magnetic beads (Roche). After an overnight incubation at 4 °C, beads were washed three times with ChIP lysis buffer + 500 mM NaCl, once with ChIP lysis buffer, and once with TE.
MS.
Proteolytic digestion.
SUMO pull-downs were precipitated in TCA, centrifuged (16,000 × g, 15 min, 4 °C), and the pellets washed with –20 °C acetone. Precipitated proteins were then resolubilized in 50 mM NH4HCO3 containing 2% (vol/vol) Rapigest surfactant (Waters), reduced using 5 mM DTT (45 min, 56 °C), and digested with trypsin (Promega) at an enzyme:substrate ratio of 1:50–1:100 (overnight, 37 °C). Rpc82 immunoprecipitations were proteolytically digested on-bead. Following immunoprecipitation, washed agarose beads were incubated in 50 mM NH4HCO3 containing 0.05% ProteaseMax surfactant (Promega). Samples were then reduced by addition of DTT (5 μM, incubated at 90 °C for 30 min) and left to cool at room temperature for 30 min before proteins were digested using 3 μg sequencing-grade modified trypsin (Promega; 37 °C, overnight).
All digested samples were acidified to 0.5–1% trifluoroacetic acid (TFA) and centrifuged at 22,000 × g for 10 min to remove ProteaseMax or Rapigest. Samples were then desalted using c18 StageTips and eluted in 80 μL 80% (vol/vol) acetonitrile (ACN)/0.1% formic acid (FA) before drying down in a Speedivac to <10 μL for MS analysis.
MS analyses.
All MS analyses were performed in triplicate on an Easy nLC1000 LC system (Thermo Electron) connected to a quadrupole Orbitrap (QExactive) mass spectrometer (Thermo Electron) with a nanoelectrospray ion source (EasySpray; Thermo Electron). Peptides were separated using an EasySpray analytical column (C18, 2 µm beads, 100 Å, 75 μm inner diameter; Thermo). The flow rate used was 0.3 μL/min, and the solvent gradient used was 2–30% (vol/vol) ACN in 0.1% FA over 120 min, followed by a column wash stage using 90% (vol/vol) ACN in 0.1% FA for 20 min. Column temperature was maintained at 60 °C. All MS analyses were operated in the data-dependent mode to automatically select the most intense peptides for MS/MS acquisition using high-energy collision dissociation. Target ions already selected for MS/MS were dynamically excluded for 30 s.
MS raw files were submitted to MaxQuant software version 1.5.3.8 (32) for protein identification. Only tryptic peptides were considered. A maximum of two missed cleavages were permitted, with a first search peptide tolerance of 20 ppm and a main search peptide tolerance of 5 ppm. No fixed modifications were searched, whereas protein N-acetylation and methionine oxidation were set as variable modifications (additionally, for SUMO pull-down analyses, the EQIGG tag on lysine residues that results from tryptic cleavage of SUMO was included as a variable modification). The minimum number of unique peptides for protein identification was set to 1, and the FDR cutoff was 0.01 (1%) for both peptide and protein identifications. The Uniprot S. cerevisiae database was used. Generation of reversed sequences was selected to assign FDR rates.
Quantitative data were analyzed using the Perseus software version 1.5.3.0 (33). Identifications from contaminant and reversed sequences were removed from the protein lists. Protein intensities were transformed to log2 for further analysis, and zero intensity values were replaced by noise values of the normal distribution of each sample. Samples were then compared using a Student’s t test method (see Datasets S1 and S4 for t test parameters and for lists of proteins that were significantly enriched to different samples).
All data were submitted to the ProteomeXchange consortium via the PRIDE database (see main article for accession numbers). SUMO pull-down data consist of experiments S1 (control pull-down, nitrogen rich), S2 (SUMO pull-down, nitrogen rich), S3 (control pull-down, nitrogen starved), and S4 (SUMO pull-down, nitrogen starved). Rpc82 immunoprecipitation data consist of Experiments Ctrl (untagged Rpc82), HA (HA-tagged WT Rpc82), and KR (HA-tagged Rpc82-4KR).
GO Enrichment Analyses.
GO enrichment analyses were performed using the online bioinformatic tools available via the Database for Annotation, Visualization and Integrated Discovery (DAVID; version 6.7; https://david.ncifcrf.gov/home.jsp) (34), using the Biological Process (BP) GO category. Protein lists were searched using gene names as assigned by MaxQuant. The background dataset used for analyses was the S. cerevisiae genome. Enrichment data generated using DAVID were visualized using the Enrichment Map plugin for Cytoscape (version 3.3.0) (35, 36). Enrichment map parameters used were a P value cutoff of 0.001 and a Q value cutoff of 0.05.
Primer Sequences.
The primer sequences are indicated in Table S3.
Supplementary Material
Acknowledgments
We thank Dr. Christoph W. Müller and Niklas A. Hoffmann (EMBL) for scientific discussions and for providing RNAPIII models that were used in Discussion. We thank Prof. Tony Hunter and Dr. Zheng Wang (The Salk Institute for Biological Studies) for sharing unpublished data and reagents. This project is supported by Norwegian Research Council Grants 221694 (to J.M.E.) and 221920 (to P.C.) and Norwegian Cancer Society Project Number 3311782 (to J.M.E.).
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
Data deposition: All ChIP sequencing data have been submitted to the European Nucleotide Archive (ENA), www.ebi.ac.uk/ena (accession no. PRJEB7579); all RNA sequencing data have been submitted to the Gene Expression Omnibus database (GEO), www.ncbi.nlm.nih.gov/geo/ (accession no. GSE85622); and all mass spectrometry datasets have been submitted to the ProteomeXchange consortium via the PRIDE database, www.ebi.ac.uk/pride (accession no. PXD005142).
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1615093114/-/DCSupplemental.
References
- 1.Warner JR. The economics of ribosome biosynthesis in yeast. Trends Biochem Sci. 1999;24(11):437–440. doi: 10.1016/s0968-0004(99)01460-7. [DOI] [PubMed] [Google Scholar]
- 2.Loewith R, Hall MN. Target of rapamycin (TOR) in nutrient signaling and growth control. Genetics. 2011;189(4):1177–1201. doi: 10.1534/genetics.111.133363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Moir RD, Willis IM. Regulation of pol III transcription by nutrient and stress signaling pathways. Biochim Biophys Acta. 2013;1829(3-4):361–375. doi: 10.1016/j.bbagrm.2012.11.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Desai N, et al. Two steps in Maf1-dependent repression of transcription by RNA polymerase III. J Biol Chem. 2005;280(8):6455–6462. doi: 10.1074/jbc.M412375200. [DOI] [PubMed] [Google Scholar]
- 5.Vannini A, et al. Molecular basis of RNA polymerase III transcription repression by Maf1. Cell. 2010;143(1):59–70. doi: 10.1016/j.cell.2010.09.002. [DOI] [PubMed] [Google Scholar]
- 6.Moir RD, et al. Protein kinase A regulates RNA polymerase III transcription through the nuclear localization of Maf1. Proc Natl Acad Sci USA. 2006;103(41):15044–15049. doi: 10.1073/pnas.0607129103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lee J, Moir RD, McIntosh KB, Willis IM. TOR signaling regulates ribosome and tRNA synthesis via LAMMER/Clk and GSK-3 family kinases. Mol Cell. 2012;45(6):836–843. doi: 10.1016/j.molcel.2012.01.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Enserink JM. Sumo and the cellular stress response. Cell Div. 2015;10:4. doi: 10.1186/s13008-015-0010-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Chymkowitch P, Nguéa P A, Enserink JM. SUMO-regulated transcription: challenging the dogma. BioEssays. 2015;37(10):1095–1105. doi: 10.1002/bies.201500065. [DOI] [PubMed] [Google Scholar]
- 10.Chymkowitch P, et al. Sumoylation of Rap1 mediates the recruitment of TFIID to promote transcription of ribosomal protein genes. Genome Res. 2015;25(6):897–906. doi: 10.1101/gr.185793.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Oeser ML, et al. Dynamic sumoylation of a conserved transcription corepressor prevents persistent inclusion formation during hyperosmotic stress. PLoS Genet. 2016;12(1):e1005809. doi: 10.1371/journal.pgen.1005809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Rosonina E, Duncan SM, Manley JL. Sumoylation of transcription factor Gcn4 facilitates its Srb10-mediated clearance from promoters in yeast. Genes Dev. 2012;26(4):350–355. doi: 10.1101/gad.184689.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kennedy BK, Lamming DW. The mechanistic target of rapamycin: The grand conducTOR of metabolism and aging. Cell Metab. 2016;23(6):990–1003. doi: 10.1016/j.cmet.2016.05.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kumar Y, Bhargava P. A unique nucleosome arrangement, maintained actively by chromatin remodelers facilitates transcription of yeast tRNA genes. BMC Genomics. 2013;14:402. doi: 10.1186/1471-2164-14-402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Jordán-Pla A, et al. Chromatin-dependent regulation of RNA polymerases II and III activity throughout the transcription cycle. Nucleic Acids Res. 2015;43(2):787–802. doi: 10.1093/nar/gku1349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Betting J, Seufert W. A yeast Ubc9 mutant protein with temperature-sensitive in vivo function is subject to conditional proteolysis by a ubiquitin- and proteasome-dependent pathway. J Biol Chem. 1996;271(42):25790–25796. doi: 10.1074/jbc.271.42.25790. [DOI] [PubMed] [Google Scholar]
- 17.Teytelman L, Thurtle DM, Rine J, van Oudenaarden A. Highly expressed loci are vulnerable to misleading ChIP localization of multiple unrelated proteins. Proc Natl Acad Sci USA. 2013;110(46):18602–18607. doi: 10.1073/pnas.1316064110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Akhter A, Rosonina E. Chromatin association of Gcn4 is limited by post-translational modifications triggered by its DNA-binding in Saccharomyces cerevisiae. Genetics. 2016;204(4):1433–1445. doi: 10.1534/genetics.116.194134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhao Q, et al. GPS-SUMO: A tool for the prediction of sumoylation sites and SUMO-interaction motifs. Nucleic Acids Res. 2014;42(Web Server issue):W325–W330. doi: 10.1093/nar/gku383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Psakhye I, Jentsch S. Protein group modification and synergy in the SUMO pathway as exemplified in DNA repair. Cell. 2012;151(4):807–820. doi: 10.1016/j.cell.2012.10.021. [DOI] [PubMed] [Google Scholar]
- 21.Jeronimo C, Collin P, Robert F. The RNA polymerase II CTD: The increasing complexity of a low-complexity protein domain. J Mol Biol. 2016;428(12):2607–2622. doi: 10.1016/j.jmb.2016.02.006. [DOI] [PubMed] [Google Scholar]
- 22.Makhnevych T, et al. Global map of SUMO function revealed by protein-protein interaction and genetic networks. Mol Cell. 2009;33(1):124–135. doi: 10.1016/j.molcel.2008.12.025. [DOI] [PubMed] [Google Scholar]
- 23.Panse VG, Hardeland U, Werner T, Kuster B, Hurt E. A proteome-wide approach identifies sumoylated substrate proteins in yeast. J Biol Chem. 2004;279(40):41346–41351. doi: 10.1074/jbc.M407950200. [DOI] [PubMed] [Google Scholar]
- 24.Hoffmann NA, et al. Molecular structures of unbound and transcribing RNA polymerase III. Nature. 2015;528(7581):231–236. doi: 10.1038/nature16143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Chen C, Kolodner RD. Gross chromosomal rearrangements in Saccharomyces cerevisiae replication and recombination defective mutants. Nat Genet. 1999;23(1):81–85. doi: 10.1038/12687. [DOI] [PubMed] [Google Scholar]
- 26.Chymkowitch P, et al. Cdc28 kinase activity regulates the basal transcription machinery at a subset of genes. Proc Natl Acad Sci USA. 2012;109(26):10450–10455. doi: 10.1073/pnas.1200067109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kim D, et al. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013;14(4):R36. doi: 10.1186/gb-2013-14-4-r36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Liao Y, Smyth GK, Shi W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30(7):923–930. doi: 10.1093/bioinformatics/btt656. [DOI] [PubMed] [Google Scholar]
- 29.Sacher M, Pfander B, Jentsch S. Identification of SUMO-protein conjugates. Methods Enzymol. 2005;399:392–404. doi: 10.1016/S0076-6879(05)99027-7. [DOI] [PubMed] [Google Scholar]
- 30.Kats ES, Enserink JM, Martinez S, Kolodner RD. The Saccharomyces cerevisiae Rad6 postreplication repair and Siz1/Srs2 homologous recombination-inhibiting pathways process DNA damage that arises in asf1 mutants. Mol Cell Biol. 2009;29(19):5226–5237. doi: 10.1128/MCB.00894-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lefèvre S, et al. Structure-function analysis of hRPC62 provides insights into RNA polymerase III transcription initiation. Nat Struct Mol Biol. 2011;18(3):352–358. doi: 10.1038/nsmb.1996. [DOI] [PubMed] [Google Scholar]
- 32.Cox J, Mann M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol. 2008;26(12):1367–1372. doi: 10.1038/nbt.1511. [DOI] [PubMed] [Google Scholar]
- 33.Tyanova S, et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat Methods. 2016;13(9):731–740. doi: 10.1038/nmeth.3901. [DOI] [PubMed] [Google Scholar]
- 34.Dennis G, Jr, et al. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 2003;4(5):3. [PubMed] [Google Scholar]
- 35.Merico D, Isserlin R, Stueker O, Emili A, Bader GD. Enrichment map: A network-based method for gene-set enrichment visualization and interpretation. PLoS One. 2010;5(11):e13984. doi: 10.1371/journal.pone.0013984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Shannon P, et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):2498–2504. doi: 10.1101/gr.1239303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Brachmann CB, et al. Designer deletion strains derived from Saccharomyces cerevisiae S288C: A useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast. 1998;14(2):115–132. doi: 10.1002/(SICI)1097-0061(19980130)14:2<115::AID-YEA204>3.0.CO;2-2. [DOI] [PubMed] [Google Scholar]
- 38.Zhou W, Ryan JJ, Zhou H. Global analyses of sumoylated proteins in Saccharomyces cerevisiae. Induction of protein sumoylation by cellular stresses. J Biol Chem. 2004;279(31):32262–32268. doi: 10.1074/jbc.M404173200. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.