Abstract
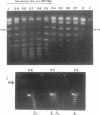
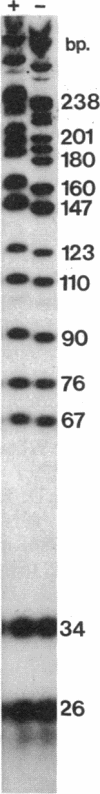
HMf, a histone from the hyperthermophilic archaeon Methanothermus fervidus binds double-stranded DNA molecules in vitro, forming compact structures that visibly resemble eukaryal nucleosomes. We show here that HMf binding increases the helical periodicity of DNA molecules to approximately 11 base pairs (bp) per turn and that DNA molecules in these nucleosome-like structures are constrained in positive toroidal supercoils. Based on the mass of HMf needed to cause a change in linking number (delta Lk), the maximum delta Lk introduced into circular DNA molecules of known sizes, and electron microscopy, we estimate that each HMf-DNA structure contains between 90 and 150 bp of DNA wrapped in 1.5 positive toroidal supercoils around a core of four HMf molecules. A model and pathway for the formation of these structures in vitro are presented and the possible role of positive toroidal wrapping of the M. fervidus genome in vivo is discussed.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Allmansberger R., Bokranz M., Kröckel L., Schallenberg J., Klein A. Conserved gene structures and expression signals in methanogenic archaebacteria. Can J Microbiol. 1989 Jan;35(1):52–57. doi: 10.1139/m89-008. [DOI] [PubMed] [Google Scholar]
- Bouthier de la Tour C., Portemer C., Nadal M., Stetter K. O., Forterre P., Duguet M. Reverse gyrase, a hallmark of the hyperthermophilic archaebacteria. J Bacteriol. 1990 Dec;172(12):6803–6808. doi: 10.1128/jb.172.12.6803-6808.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown J. W., Daniels C. J., Reeve J. N. Gene structure, organization, and expression in archaebacteria. Crit Rev Microbiol. 1989;16(4):287–338. doi: 10.3109/10408418909105479. [DOI] [PubMed] [Google Scholar]
- Broyles S. S., Pettijohn D. E. Interaction of the Escherichia coli HU protein with DNA. Evidence for formation of nucleosome-like structures with altered DNA helical pitch. J Mol Biol. 1986 Jan 5;187(1):47–60. doi: 10.1016/0022-2836(86)90405-5. [DOI] [PubMed] [Google Scholar]
- Drlica K., Rouviere-Yaniv J. Histonelike proteins of bacteria. Microbiol Rev. 1987 Sep;51(3):301–319. doi: 10.1128/mr.51.3.301-319.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dürrenberger M., Bjornsti M. A., Uetz T., Hobot J. A., Kellenberger E. Intracellular location of the histonelike protein HU in Escherichia coli. J Bacteriol. 1988 Oct;170(10):4757–4768. doi: 10.1128/jb.170.10.4757-4768.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Felsenfeld G. Chromatin. Nature. 1978 Jan 12;271(5641):115–122. doi: 10.1038/271115a0. [DOI] [PubMed] [Google Scholar]
- Hayes J. J., Tullius T. D., Wolffe A. P. The structure of DNA in a nucleosome. Proc Natl Acad Sci U S A. 1990 Oct;87(19):7405–7409. doi: 10.1073/pnas.87.19.7405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krogh S., Mortensen U. H., Westergaard O., Bonven B. J. Eukaryotic topoisomerase I-DNA interaction is stabilized by helix curvature. Nucleic Acids Res. 1991 Mar 25;19(6):1235–1241. doi: 10.1093/nar/19.6.1235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pettijohn D. E. Histone-like proteins and bacterial chromosome structure. J Biol Chem. 1988 Sep 15;263(26):12793–12796. [PubMed] [Google Scholar]
- Rhodes D., Klug A. Helical periodicity of DNA determined by enzyme digestion. Nature. 1980 Aug 7;286(5773):573–578. doi: 10.1038/286573a0. [DOI] [PubMed] [Google Scholar]
- Sandman K., Krzycki J. A., Dobrinski B., Lurz R., Reeve J. N. HMf, a DNA-binding protein isolated from the hyperthermophilic archaeon Methanothermus fervidus, is most closely related to histones. Proc Natl Acad Sci U S A. 1990 Aug;87(15):5788–5791. doi: 10.1073/pnas.87.15.5788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmid M. B. More than just "histone-like" proteins. Cell. 1990 Nov 2;63(3):451–453. doi: 10.1016/0092-8674(90)90438-k. [DOI] [PubMed] [Google Scholar]
- Searcy D. G., Delange R. J. Thermoplasma acidophilum histone-like protein. Partial amino acid sequence suggestive of homology to eukaryotic histones. Biochim Biophys Acta. 1980 Aug 26;609(1):197–200. doi: 10.1016/0005-2787(80)90212-9. [DOI] [PubMed] [Google Scholar]
- Travers A. A. Why bend DNA? Cell. 1990 Jan 26;60(2):177–180. doi: 10.1016/0092-8674(90)90729-x. [DOI] [PubMed] [Google Scholar]
- Tullius T. D., Dombroski B. A., Churchill M. E., Kam L. Hydroxyl radical footprinting: a high-resolution method for mapping protein-DNA contacts. Methods Enzymol. 1987;155:537–558. doi: 10.1016/0076-6879(87)55035-2. [DOI] [PubMed] [Google Scholar]
- Tullius T. D., Dombroski B. A. Iron(II) EDTA used to measure the helical twist along any DNA molecule. Science. 1985 Nov 8;230(4726):679–681. doi: 10.1126/science.2996145. [DOI] [PubMed] [Google Scholar]
- Vosberg H. P. DNA topoisomerases: enzymes that control DNA conformation. Curr Top Microbiol Immunol. 1985;114:19–102. doi: 10.1007/978-3-642-70227-3_2. [DOI] [PubMed] [Google Scholar]
- Wang J. C. DNA topoisomerases: why so many? J Biol Chem. 1991 Apr 15;266(11):6659–6662. [PubMed] [Google Scholar]
- Weil C. F., Cram D. S., Sherf B. A., Reeve J. N. Structure and comparative analysis of the genes encoding component C of methyl coenzyme M reductase in the extremely thermophilic archaebacterium Methanothermus fervidus. J Bacteriol. 1988 Oct;170(10):4718–4726. doi: 10.1128/jb.170.10.4718-4726.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams M. R., Gilson D., Marsh L., Morgan D. A., Nicholson R. I., Elston C. W., Griffiths K., Blamey R. W. The early results from a randomised study of radiotherapy versus Nolvadex (tamoxifen) as initial treatment for stage III breast cancer. Eur J Surg Oncol. 1988 Jun;14(3):235–240. [PubMed] [Google Scholar]
- Woese C. R., Kandler O., Wheelis M. L. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci U S A. 1990 Jun;87(12):4576–4579. doi: 10.1073/pnas.87.12.4576. [DOI] [PMC free article] [PubMed] [Google Scholar]