Abstract
Reversible protein S-palmitoylation confers spatiotemporal control of protein function by modulating protein stability, trafficking and activity, as well as protein–protein and membrane–protein associations. Enabled by technological advances, global studies revealed S-palmitoylation to be an important and pervasive posttranslational modification in eukaryotes with the potential to coordinate diverse biological processes as cells transition from one state to another. Here, we review the strategies and tools to analyze in vivo protein palmitoylation and interrogate the functions of the enzymes that put on and take off palmitate from proteins. We also highlight palmitoyl proteins and palmitoylation-related enzymes that are associated with cellular differentiation and/or tissue development in yeasts, protozoa, mammals, plants and other model eukaryotes.
Keywords: cellular differentiation, fatty-acylation, lipidation, posttranslational modification, S-palmitoylation
Introduction
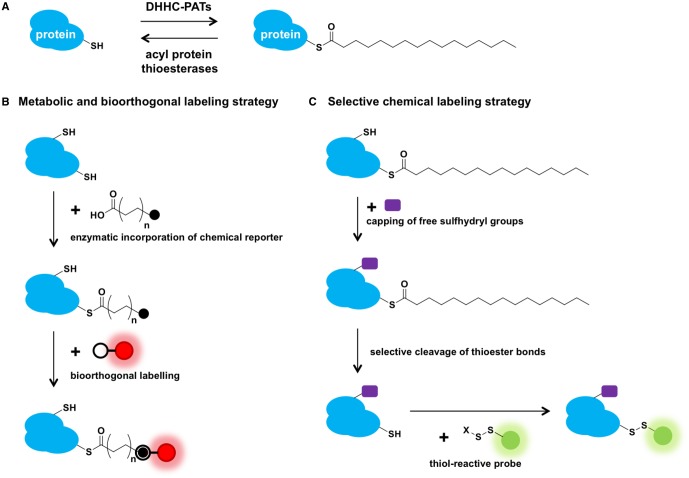
Protein S-acylation or S-palmitoylation involves the posttranslational addition of fatty acyl chains, typically a palmitate (C16:0), to cysteine residues of proteins via thioester linkages (Figure 1A). In contrast with other forms of protein lipidation, S-palmitoylation is uniquely reversible due to the high-energy thioester bond formed between the acyl group and the cysteine side chain, potentially allowing for rapid spatiotemporal control of protein function akin to protein phosphorylation. S-Palmitoylation predominantly serves to target proteins to specific membrane compartments and/or microdomains [1–4]. While typically not the primary membrane association signal, S-palmitoylation often acts in concert with other lipid modifications, such as N-myristoylation and prenylation, to determine the intracellular distribution of soluble proteins [2–4]. By influencing protein localization and trafficking, palmitoylation/depalmitoylation can have critical effects on protein function as epitomized by compartmentalized Ras signaling, where spatial segregation of distinct signal transduction modules diversifies the signaling outputs by a single protein [5,6]. Palmitoylation may also regulate protein activity in diverse ways, such as inducing protein conformational changes, modulating protein stability and protein–protein interactions and interacting with other posttranslational modifications [7–12]. These functional consequences of protein palmitoylation are not mutually exclusive, and a single palmitoylation event can simultaneously modulate multiple aspects of protein function [13].
Figure 1. Protein S-palmitoylation and analytical strategies.
(A) Dynamic S-palmitoylation is mediated by DHHC-containing PATs (DHHC-PATs) and acyl protein thioesterases that put on and take off palmitate from cysteine residues of proteins, respectively. (B) Metabolic and bioorthogonal labeling strategy using fatty acid chemical reporters as well as (C) the selective chemical labeling strategy for analysis of protein S-palmitoylation. In both strategies, the incorporation of detection or enrichment probes in the final step allows for direct detection or enrichment and mass spectrometry identification of tagged proteins.
S-Palmitoylation of intracellular proteins is mainly mediated by an evolutionarily conserved family of palmitoyl acyltransferases (PATs), which are identified by the characteristic catalytic Asp-His-His-Cys (DHHC) motif embedded within a cysteine-rich domain. First discovered in budding yeast Saccharomyces cerevisiae [14,15], PAT orthologs are found in all eukaryotes, ranging from 5 PATs in fission yeast Schizosaccharomyces pombe to ∼23 in mammals and 24 in Arabidopsis. While there is overlapping substrate specificity between multiple PATs [16,17], it is clear that there are additional specificity determinants in the variable domains of the PATs [18] as well as regulatory mechanisms that determine the context-dependent protein substrate pool and function of individual enzymes. Regulation of specific PATs at the transcriptional [19], translational [20] and posttranslational [21] levels has been reported with significant changes in the palmitoylation state of the corresponding protein substrates. Additionally, PAT activity can be determined by its subcellular trafficking [22], oligomerization state [23] or additional protein subunits [14,24,25]. Notably, these are isolated examples in specific host systems looking at a few protein substrates. In general, the in vivo substrate selectivity and regulation of PATs as well as the physiological significance of PAT-mediated protein palmitoylation remain to be fully characterized.
Until recently, depalmitoylation of cytosolic cysteine residues has been attributed to only two related acyl protein thioesterases APT1/LYPLA1 and APT2/LYPLA2 [26]. Since APT1 and APT2 are both palmitoylated, autoregulatory mechanisms are proposed where the depalmitoylating enzymes control their physical access to protein substrates [27,28]. Studies focusing on serine hydrolases led to the independent discovery of the ABHD17 proteins that depalmitoylate N-Ras and PSD-95 in mammalian cells [29,30]. ABHD12 and ABHD13 also exhibited depalmitoylating activities, albeit weaker than ABHD17, when tested against PSD-95 as the substrate [29]. Interestingly, an ABDH13 ortholog in Toxoplasma, TgPpt1, was demonstrated to depalmitoylate proteins and to have important functions in regulating host cell invasion by the parasite [31]. These findings suggest that the family of depalmitoylating enzymes may be larger and more diverse than previously appreciated. Like the PATs, the substrate selectivity and biological importance of depalmitoylating enzymes remain to be determined, but it is clear that cellular context matter and factors, such as cell type and physiology, can affect depalmitoylating activity toward specific protein substrates [29,30].
This review focuses on protein S-palmitoylation in the context of cellular differentiation, which is the process by which cells becomes more specialized to perform specific functions. We will introduce the main chemical strategies and various chemical tools to analyze in vivo protein palmitoylation, and to interrogate the activity of palmitoylating and depalmitoylating enzymes in different cellular states. We will then highlight palmitoyl proteins and palmitoylation-related enzymes that are involved with cellular and tissue development in yeasts, protozoa, mammals, plants and other model eukaryotes. Other forms of protein acylation such as N-myristoylation, N-palmitoylation and O-palmitoleoylation of secreted proteins (e.g. Wnt and Hedgehog) and the family of membrane-bound O-acyltransferases that mediate these modifications will not be included in this review [32,33].
Chemical strategies and tools to study protein S-palmitoylation
Advances in our understanding of protein S-palmitoylation over the last decade can be largely attributed to the development of chemical tools that enable rapid quantitative analyses of palmitoylated proteins. These chemical tools can be broadly categorized into two main strategies. In the first strategy (Figure 1B), cells are metabolically labeled with fatty acid chemical reporters, which are site-specifically installed onto cysteines of target proteins via native thioester linkages by the endogenous enzymatic machinery. Visualization or enrichment probes are subsequently introduced using bioorthogonal labeling reactions to allow detection or identification of modified proteins [34,35]. Because these bioorthogonal fatty acid chemical reporters can be competed away by endogenous substrates, they have been employed in pulse-chase experiments to monitor palmitate turnover kinetics on proteins [36,37]. Fatty acid chemical reporters with cross-linking functionalities have also been developed to interrogate protein–protein interactions that depend on protein palmitoylation [38]. The second strategy involves the selective chemical modification of thioester-linked cysteines in S-palmitoylated proteins (Figure 1C). After initial capping of free thiols on proteins, selective cleavage of thioester linkages liberates thiols that can be selectively labeled to facilitate detection and/or enrichment by biochemical methods or mass spectrometry [39,40]. Methods, such as acyl-biotin exchange (ABE) and acyl-resin-assisted capture, employ this strategy. Notably, the selective hydrolysis and removal of thioester-linked acyl groups facilitate the identification of palmitoylation sites since direct mass spectrometry detection of acylated peptides can be challenging [41]. In a recent variation of the latter strategy, acyl-PEG switch/exchange uses PEGylated thiol-reactive reagents to induce electrophoretic mobility shift of modified proteins to monitor the relative abundance of palmitoylated versus nonpalmitoylated forms of target proteins and the number of palmitoylation sites [29,42,43]. Both strategies have been successfully employed to detect and identify S-palmitoylated proteins in global and focused studies of various cell types.
The ability to visualize and profile dynamically S-palmitoylated proteins on a global scale has greatly expanded the scope of the modification by revealing new palmitoylated proteins and regulatory roles of protein palmitoylation in eukaryotic physiology and disease. Since the first global profiling of palmitoylated proteins in yeast by Roth et al. [16], dozens of palmitoyl proteomes consisting of ∼10% of the proteomes in yeasts, protozoans [44], plants [45,46] and mammalian systems have been reported [47–49]. Despite the lack of a ‘consensus sequence’ surrounding palmitoylation sites, the ever increasing number of palmitoyl proteomes led to the development and refinement of in silico predictive programs [50–53], which are robust in predicting modification sites for proteins with canonical palmitoylation motifs (e.g. dual acylation, cysteines near to prenylation motifs or transmembrane domains) and for those with validated palmitoylated homologs or orthologs. Our predictive ability for proteins with atypical modification sites will likely continue to improve as the list and diversity of experimentally validated proteins and palmitoylation sites expand.
Improving the quality, analysis and curation of palmitoyl proteomes will be important as the field moves toward comparative proteomics to identify palmitoylation events that are critical for various biological phenomena and disease states. Notably, the overlap between palmitoyl proteomes obtained using the chemical reporter strategy and those obtained using the selective chemical modification strategy is limited, reflecting the different sources of false positives from each technique (Table 1) [54]. With metabolic labeling, metabolism of the fatty acid chemical reporters can lead to enrichment of proteins with other forms of protein lipidation, although alk-16 (also known as 16-ODYA) has been shown to be preferentially incorporated into palmitoylated proteins [34,48]. The biotin switch strategy will enrich for proteins with any thioester-linked modifications, which also include SUMOylation and ubiquitination. Marrying these independent complementary approaches minimizes these pitfalls for more accurate global analyses of protein S-palmitoylation in cells [44]. Since bona fide palmitoylated proteins are more likely to be identified using multiple methods, Blanc et al. [54] combined the data from existing databases, global and focused palmitoylation studies to generate a high confidence SwissPalm database for improved prediction of palmitoylated proteins and modification sites. Comparison of SwissPalm with other databases uncovered potentially extensive cross-talk between palmitoylation and various posttranslational modifications [54].
Table 1. Chemical tools to study protein S-palmitoylation.
| Fatty acid probes | Principle of method | Detection | Identification of modified proteins | Variations/other applications |
|---|---|---|---|---|
| Metabolic labeling | ||||
| Radiolabelled (e.g. 3H, 13C, 125I) fatty acids | Autoradiographic detection | Sensitive and quantitative | X Indirect identification (e.g. immunoprecipitation and overexpression) | Used in pulse-chase experiments to determine turnover kinetics |
| X Cumbersome and requires long exposure periods or additional safety protocols | ||||
| X False positives as a result of fatty acid metabolism, including crosstalk between different forms of protein fatty acylation | X Requires prior knowledge of candidate proteins | |||
| Bioorthogonal fatty acid chemical reporters (e.g. azide-, alkyne-functionalized) | Introduction of custom detection and/or enrichment probes post-metabolic labeling via bioorthogonal reactions | Rapid, sensitive and quantitative non-radioactive detection | Selective enrichment of palmitoylproteomes | Used in pulse-chase experiments to determine turnover kinetics |
| Multiplex detection is possible with orthogonal reporters and detection probes | ||||
| X False positives as a result of fatty acid metabolism, including crosstalk between different forms of protein fatty acylation | X Challenging site ID due to difficult MS detection of acylated peptides | Fatty acid chemical reporters with crosslinking functionalities enable the interrogation of palmitoylation-specific protein-protein interactions | ||
| X False positives as a result of fatty acid metabolism, including crosstalk between different forms of protein fatty acylation | ||||
| Method | Principle of method | Detection | Identification of modified proteins | Variations/other applications |
| Selective chemical labeling of palmitoylated cysteines | ||||
| Acyl-biotin exchange (ABE) or Acyl-resin-assisted capture (acyl-Rac) | After initial capping of free thiols, selective cleavage of thioester bonds liberates free thiols for reaction with thiol-reactive reagents that enable detection and enrichment | Rapid, sensitive and quantitative non-radioactive detection | Selective enrichment of palmitoylproteomes | The population of modified proteins can be determined using thiol-reactive reagent that alters protein electophoretic mobility (acyl-PEG switch/exchange) |
| Identification of modification sites | ||||
| Acyl-PEG-exchange (APE) or Acyl-PEG-switch | X False positives with other thioester-linked PTMs and also with incomplete thiol capping | X False positives with other thioester-linked PTMs and also with incomplete thiol capping | ||
X symbol indicates the limitations of each strategy. PTMs, posttranslational modifications.
In addition to biochemical analyses, the ability to rapidly and selectively perturb the modification in cells will be valuable toward understanding the biological significance of palmitoylation in different physiological contexts. Chemical inhibitors provide a rapid and convenient method to perturb protein palmitoylation, especially in organisms and cells that are challenging to manipulate genetically. Readers are referred to a review that includes a dedicated section on the different classes of small molecule inhibitors targeting palmitoylating and depalmitoylating enzymes [55]. ‘Clickable’ forms of some inhibitors have also been used to discover and validate enzymes that modulate protein palmitoylation in cells [30,31,36]. It is important to note that although 2-bromopalmitate has been and remains one of the most widely used inhibitors of protein palmitoylation, it also inhibits fatty lipid metabolism and should not be used in isolation to prove or interrogate the function of protein S-palmitoylation. Undoubtedly, the development of potent and highly selective chemical inhibitors for PATs and depalmitoylating thioesterases will be pivotal toward dissecting their functional contributions and uncovering insights into the regulation of dynamic protein palmitoylation.
S-Palmitoylation in yeast cellular differentiation
The fission yeast S. pombe is an emerging model organism for palmitoylation studies due to its genetic tractability and relatively simple palmitoylation machinery compared with other eukaryotes. Studies have associated palmitoylation of several important signal transduction proteins with S. pombe sexual differentiation, which involves mating between haploid cells of opposite mating types, entry and progression through meiosis to yield haploid spores. Using the fission yeast system, where a single Ras ortholog is involved in two distinct signaling pathways with quantifiable phenotypes, the Chang laboratory showed that Ras1 signal transduction is spatially compartmentalized with cellular morphogenesis regulated by nonpalmitoylated Ras1 at endomembranes and mating requiring palmitoylated Ras1 at the plasma membrane [5]. Similar to the S. cerevisiase PAT ERF2 that preferentially modifies heterolipidated GTPases [16], the S. pombe Erf2 ortholog is the primary PAT that palmitoylates Ras1 and two other small GTPases — Rho2 and Rho3 [19,56]. High expression levels of SpErf2 and its noncatalytic protein cofactor SpErf4 during meiosis are required for Rho3 palmitoylation, and dysregulation of Rho3 palmitoylation triggers aberrant meiotic divisions in sensitized cells [19]. Rho2 palmitoylation is needed for morphogenesis and cell wall integrity of vegetative cells via the Pmk1 pathway that is antagonistically regulated by Rho3 [56]. Future work will reveal the regulatory roles and mechanisms of SpErf2-mediated palmitoylation in coordinating the signaling outputs of these small GTPases as cells exit vegetative growth and undergo sexual differentiation.
Cryptococcus neoformans is a facultative intracellular fungal pathogen that is able to survive and proliferate in the harsh environment of macrophage phagolysosomes. Recent investigations uncover a major role for PAT-mediated protein palmitoylation in the virulent potential of C. neoformans. The PFA4 gene encoding a DHHC-PAT was identified from screening genes that influenced host–pathogen interactions, and loss of PFA4 function has dramatic effects on morphology, stress tolerance and virulence potential of C. neoformans [57]. Comparative palmitoyl proteome profiling identified CnPFA4 protein substrates that are involved in cell wall synthesis, membrane transport, signal transduction and membrane trafficking, which is consistent with the pleiotropic defects observed for pfa4Δ cells [57]. CnPFA4 was also identified as the major PAT responsible for Ras1 palmitoylation, which is required for Ras1 localization at the plasma membrane and pathogenesis in a cryptococcosis murine model [58,59]. Ras1 palmitoylation is not required for C. neoformans sexual differentiation [58], suggesting palmitoylation-dependent compartmentalization of Ras1 signaling.
Protein palmitoylation can affect the function of nuclear proteins. In S. cerevisiae, PFA4-mediated palmitoylation of the telomere-binding protein RIF1 altered heterochromatin dynamics and transcriptional silencing [60]. It is unclear if RIF1 palmitoylation is regulated, but with 25% of the putative palmitoylated proteins in mouse and human cells being nuclear proteins [54], this finding raises the interesting possibility that palmitoylation of nuclear proteins may be important in direct modulation of global gene expression during cellular transitions.
S-Palmitoylation in the virulence and transmission of protozoan parasites
Apicomplexan parasites are a large group of obligate intracellular protozoan parasites. Most members of this group have complex asexual and sexual reproduction cycles within multiple hosts, with survival requiring rapid adjustment to distinct environments and precise spatiotemporal coordination of key cellular processes for host cell invasion, replication and egress. Recent work demonstrated the pervasive roles of protein S-palmitoylation in the developmental life cycles of Toxoplasma gondii and Plasmodium falciparum and, by extension, their pathogenesis. Treatment with 2-bromopalmitate yielded pleiotropic developmental defects and reduced the invasive capacity of T. gondii and P. falciparum [44,61]. Interestingly, chemical inhibition of depalmitoylation activity by palmitoyl protein thioesterase 1 TgPPT1 significantly enhanced host cell invasion by T. gondii [31]. Palmitoyl proteome analyses in T. gondii and P. falciparum reveal palmitoyl proteins that are needed for the function of specialized invasion organelles, providing further insights into the apparently essential roles of dynamic protein palmitoylation on host cell invasion by these parasites [44,62]. Many components of the invasion motor glideosome complex, including the well-known GAP45, are palmitoylated, disruption of which is associated with motility and invasion defects [44,62,63]. Palmitoylation of TgAMA1 and Pf/TgARO is required for proper apical localization of specialized invasion-associated secretory organelles called rhoptries [62,64,65]. Consistent with the importance of protein palmitoylation in rhoptry function, disruption of rhoptry-localized TgDHHC7 responsible for TgARO palmitoylation blocks host invasion [64,66]. Global studies of PATs in Toxoplasma and Plasmodium support functional specialization by the enzymes across different life cycle stages of the parasites, with PATs' function determined by differential subcellular localization, expression patterns and posttranslational modifications [66–68]. These studies also implicate protein palmitoylation in other developmental stages of apicomplexan parasites. For example, PfDHHC2 is essential for the progression through both asexual and sexual stages in the mammalian and mosquito hosts, respectively [69]. In Plasmodium berghei, PbDHHC10 expression is translationally repressed in gametocyetes and briefly translated during ookinete formation to mediate palmitoylation events that are required for crystalloid formation and parasite transmission from the mammalian host to the mosquito vector [70]. PbHHC3 and PbDHHC9 appear to have functionally overlapping roles in mediating parasite sexual differentiation in the insect host [71,72].
Protein palmitoylation also plays a key role in the life cycle of a different protozoan parasite Giardia lamblia, which is one of the major global causes of diarrheal disease. Encystation of the parasite to form cysts is critical for the survival of the parasite outside the host and its transmission. In addition to a changing palmitoyl proteome during encystation, Merino et al. [73] showed that genetic and chemical perturbation of protein palmitoylation by the different PATs in G. lamblia can negatively affect parasite differentiation into cysts. Identifying the palmitoyl proteins and the respective PATs participating in this key cellular transition will open up opportunities for therapeutic interventions.
S-Palmitoylation in mammalian tissue development
Meta-analysis of mammalian palmitoyl proteomes reveals dominant roles for protein palmitoylation in neural development and function, with a striking proportion of synaptic genes encoding for palmitoyl proteins [74], including neurotransmitter receptors, transporters, adhesion molecules, scaffolding proteins and vesicular trafficking proteins [75]. Reduced ZDHHC2 expression was observed in degenerating dopaminergic neurons and in patients with incipient Parkinson's disease [76]. Additionally, mutations in PATs, including ZDHHC17/ZDHHC13 (HIP14/HIP14L) and ZDHHC8, are associated with neurological disorders such as Huntington's disease and schizophrenia, respectively [18,75]. The spine density deficits in a schizophrenia mouse model are palmitoylation-dependent and can be rescued in vivo by overexpressing ZDHHC8 or one of its substrates — the constitutively active brain-specific splice isoform of Cdc42-palm [47,77–79]. By regulating its subcellular localization and RhoGDI (Rho guanine nucleotide dissociation inhibitor) binding, palmitoylation of Cdc42-palm is required for normal dendritic spine development during synaptogenesis [47,80]. Palmitoylation of other proteins, including paralemmin [81] and actin regulator LIMK1 [82], have also been shown to affect dendritic spine maturation that is critical for neuronal plasticity. The modulation of global protein palmitoylation in response to synaptic activity and cellular signals further supports the regulatory roles of protein palmitoylation in neuronal plasticity and development [47]. Palmitate turnover of the most abundant neural scaffolding protein PSD-95 is accelerated upon glutamate receptor activation, and this down-regulates receptor signaling activity [83]. On the other hand, suppression of neural activity triggers the ZDHHC2 translocation to postsynaptic membrane, leading to increased PSD-95 palmitoylation and synaptic accumulation [22]. Synaptic activity increases ZDHHC5-mediated palmitoylation of intracellular cadherin-binding protein δ-catenin and stabilizes synaptic cadherin adhesion complexes that are critical for synaptic plasticity [84]. In cultured neuronal stem cells, induction of neural differentiation led to the rapid degradation of ZDHHC5 and reduced flotillin palmitoylation [21], implicating protein palmitoylation in neural stem cell differentiation. Palmitoylation has been shown to stabilize EID1, an inhibitor of the CREB-binding protein/p300 epigenetic regulator for neural stem cell differentiation [85].
The pivotal role for PATs and their substrates in the development of other tissues that are not part of the nervous system is indicated by the associations between dysregulated protein palmitoylation with cancers, which reflect defective control over cellular proliferation and differentiation [74]. The involvement of palmitoyl proteins, associated enzymes in cellular transformation and tumorigenesis are covered in a detailed review by Yeste-Velasco et al. [55]. Here, we focus on the role of protein palmitoylation in cellular and tissue development, particularly in modulating growth factor and hormone signaling, which mediate intercellular communication and regulate gene expression programs that drive cellular differentiation in various developmental stages. Palmitoylation of the epidermal growth factor receptor (EGFR) by ZDHHC20 limits EGFR signaling by inhibiting autophosphorylation and recruitment of the downstream adapter protein Grb2 and increasing receptor turnover [86]. For estrogen receptors (ERs), progesterone receptors and the androgen receptors, palmitoylation by ZDHHC7 and ZDHHC21 is crucial for their plasma membrane localization and function in mediating rapid tissue-specific responses to steroid hormones [87–89]. Upon estradiol binding, ERα is depalmitoylated and dissociates from caveolin-1, after which it is available to downstream signaling targets in the RK/MAPK and PI3K/AKT pathways [89]. Mice that are deficient in ZDHHC21 activity showed defects in maintaining skin homeostasis and hair follicle differentiation that results in hair loss [90], though it remains to be determined if these phenotypes can be attributed to defective steroid hormone signaling. Other protein substrates of ZDHHC7 or ZDHHC21 include Scribble [91] as well as the death receptor Fas and kinase Lck, highlighting the additional roles of these two PATs in modulating cellular polarity and proliferation [92,93]. ZDHHC13 knockout mice with pleiotropic developmental abnormalities of the hair, skin and bone hint at yet to be identified protein palmitoylation events that affect normal mammalian cellular and tissue development [94].
S-Palmitoylation in plants
Compared with yeast and mammalian systems, the study of protein S-palmitoylation in plants is in the early stages. There are no reports of plant palmitoyl protein thioesterases to date. Of the 24 Arabidopsis thaliana DHHC-PATs [95], only a few have been characterized in any detail in terms of their biological roles. AtPAT10 is involved in vacuolar and tonoplast function, while AtPAT24/TIP1 has been implicated in developmental processes including pollen tube and root hair growth, shoot branching and cell polarity [96,97]. Recently, AtPAT13 and AtPAT14 have been shown to be involved in leaf senescence [98,99]. Identifying their protein substrates will be integral toward understanding how PATs and protein palmitoylation coordinate cellular differentiation and development in plants, but may require the use of independent methods to confidently establish PAT–substrate relationships [45]. Among the ∼500 candidate S-palmitoylated proteins in plants identified using the biotin switch strategy, protein kinases are overrepresented [45,46], suggesting that S-palmitoylation may have a major role in modulating phosphorylation signaling cascades in plants. These include RLK (receptor-like kinase) superfamily members that contain conserved cysteines adjacent to predicted transmembrane domains or N-myristoylated sites [45]. S-Palmitoylation of the LIP1 and LIP2 receptor-like cytoplasmic kinases is needed for directing pollen tube growth [100]. Non-RLKs such as the calcium-dependent protein kinase OSCPK2 in rice require both N-myristoylation and S-palmitoylation for proper subcellular localization [101]. Calcineurin B-like proteins, CBL1 and CBL2, which recruit Ser/Thr protein kinases during Ca2+ signaling, require S-palmitoylation for targeting to the right membrane compartments in the cell [102,103]. Palmitoylation-deficient CBL2 fails to localize to the tonoplast and led to seed germination defects [102]. Besides individual proteins, palmitoylation can also direct the cellular distribution of large protein complexes. Kumar et al. [104] demonstrated that all catalytic subunits within the cellulose synthase complex are S-acylated and that the modification is required for correct localization of the complex to the plasma membrane and normal cellulose synthesis.
S-Palmitoylation in other model organisms
In zebrafish, strong phenotypes observed with various PAT deficiencies support the importance of protein palmitoylation in cellular differentiation and animal development. ZDHHC13 modulates bone morphogenetic protein signaling for lineage specification during embryogenesis [105]. A preliminary study using 2-bromopalmitate further implicates PATs and protein palmitoylation in mediating the transition from maternal to zygotic transcriptional programs after embryo fertilization [106]. Knockdown of DHHC15b and DHHC16 negatively affects forebrain development via dysregulation of neuronal differentiation and neural stem cell proliferation, respectively [107,108]. The forebrain developmental defect observed in animals deficient in ZDHHC15b activity is further associated with poor learning ability [108].
In Caenorhabditis elegans, lysosome-related fibrous body-membrane organelles (FB-MOs) are important for asymmetric cytoplasmic partitioning. Worms with mutations in spe-10, which encodes a DHHC-containing protein localized to FB-MOs, showed defective spermatogenesis and are sterile, suggesting a role for SPE-10-mediated protein palmitoylation in establishing cellular polarity during cellular differentiation [109]. A systematic study of the other 14 DHHC-containing proteins in C. elegans using single and double RNAi knockdowns, however, did not yield any obvious phenotypes [110]. A similar functional analysis remains to be performed for Drosophila, in which tissue- and sex-specific expression of specific DHHC-containing genes have been observed [111].
Perspectives
This is an exciting time to be studying protein S-palmitoylation. Posttranslational modifications increase the proteome complexity, and pervasive reversible modifications like protein palmitoylation have the potential to orchestrate diverse biological processes involved as cells transition from one state to another. The tools are in place to monitor in vivo palmitoylation stoichiometry and dynamics in various eukaryotic systems. Increasingly, selective small molecule inhibitors complement genetic approaches to profile and rapidly interrogate the functional contributions of palmitoylating and depalmitoylating enzymes. Ongoing efforts to minimize false positives, validate and curate palmitoyl proteomes will continue to enhance in silico predictive programs and establish the framework for comparative proteomics studies. We are now poised to identify and dissect critical palmitoylation events that regulate and/or are regulated during cellular differentiation and navigate the complex regulatory networks governing eukaryotic physiology and disease.
Abbreviations
- ABE
acyl-biotin exchange
- DHHC
Asp-His-His-Cys
- EGFR
epidermal growth factor receptor
- ER
estrogen receptor
- FB-MOs
fibrous body-membrane organelles
- PAT
palmitoyl acyltransferase
- RLK
receptor-like kinase.
Competing Interests
The Authors declare that there are no competing interests associated with the manuscript.
References
- 1.Levental I., Grzybek M. and Simons K. (2010) Greasing their way: lipid modifications determine protein association with membrane rafts. Biochemistry 49, 6305–6316 doi: 10.1021/bi100882y [DOI] [PubMed] [Google Scholar]
- 2.Rocks O., Peyker A., Kahms M., Verveer P.J., Koerner C., Lumbierres M. et al. (2005) An acylation cycle regulates localization and activity of palmitoylated Ras isoforms. Science 307, 1746–1752 doi: 10.1126/science.1105654 [DOI] [PubMed] [Google Scholar]
- 3.Kanaani J., Patterson G., Schaufele F., Lippincott-Schwartz J. and Baekkeskov S. (2008) A palmitoylation cycle dynamically regulates partitioning of the GABA-synthesizing enzyme GAD65 between ER-Golgi and post-Golgi membranes. J. Cell Sci. 121, 437–449 doi: 10.1242/jcs.011916 [DOI] [PubMed] [Google Scholar]
- 4.Schroeder H., Leventis R., Rex S., Schelhaas M., Nägele E., Waldmann H. et al. (1997) S-Acylation and plasma membrane targeting of the farnesylated carboxyl-terminal peptide of N-ras in mammalian fibroblasts. Biochemistry 36, 13102–13109 doi: 10.1021/bi9709497 [DOI] [PubMed] [Google Scholar]
- 5.Onken B., Wiener H., Philips M.R. and Chang E.C. (2006) Compartmentalized signaling of Ras in fission yeast. Proc. Natl Acad. Sci. U.S.A. 103, 9045–9050 doi: 10.1073/pnas.0603318103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Aran V. and Prior I.A. (2013) Compartmentalized Ras signaling differentially contributes to phenotypic outputs. Cell. Signal. 25, 1748–1753 doi: 10.1016/j.cellsig.2013.05.004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ho G.P.H., Selvakumar B., Mukai J., Hester L.D., Wang Y., Gogos J.A. et al. (2011) S-nitrosylation and S-palmitoylation reciprocally regulate synaptic targeting of PSD-95. Neuron 71, 131–141 doi: 10.1016/j.neuron.2011.05.033 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Tian L., Jeffries O., McClafferty H., Molyvdas A., Rowe I.C.M., Saleem F. et al. (2008) Palmitoylation gates phosphorylation-dependent regulation of BK potassium channels. Proc. Natl Acad. Sci. U.S.A. 105, 21006–21011 doi: 10.1073/pnas.0806700106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Valdez-Taubas J. and Pelham H. (2005) Swf1-dependent palmitoylation of the SNARE Tlg1 prevents its ubiquitination and degradation. EMBO J. 24, 2524–2532 doi: 10.1038/sj.emboj.7600724 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Abrami L., Leppla S.H. and van der Goot F.G. (2006) Receptor palmitoylation and ubiquitination regulate anthrax toxin endocytosis. J. Cell Biol. 172, 309–320 doi: 10.1083/jcb.200507067 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Yount J.S., Karssemeijer R.A. and Hang H.C. (2012) S-Palmitoylation and ubiquitination differentially regulate interferon-induced transmembrane protein 3 (IFITM3)-mediated resistance to influenza virus. J. Biol. Chem. 287, 19631–19641 doi: 10.1074/jbc.M112.362095 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ahearn I.M., Tsai F.D., Court H., Zhou M., Jennings B.C., Ahmed M. et al. (2011) FKBP12 binds to acylated H-ras and promotes depalmitoylation. Mol. Cell 41, 173–185 doi: 10.1016/j.molcel.2011.01.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Holland S.M., Collura K.M., Ketschek A., Noma K., Ferguson T.A., Jin Y. et al. (2016) Palmitoylation controls DLK localization, interactions and activity to ensure effective axonal injury signaling. Proc. Natl Acad. Sci. U.S.A. 113, 763–768 doi: 10.1073/pnas.1514123113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lobo S., Greentree W.K., Linder M.E. and Deschenes R.J. (2002) Identification of a Ras palmitoyltransferase in Saccharomyces cerevisiae. J. Biol. Chem. 277, 41268–41273 doi: 10.1074/jbc.M206573200 [DOI] [PubMed] [Google Scholar]
- 15.Roth A.F., Feng Y., Chen L. and Davis N.G. (2002) The yeast DHHC cysteine-rich domain protein Akr1p is a palmitoyl transferase. J. Cell Biol. 159, 23–28 doi: 10.1083/jcb.200206120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Roth A.F., Wan J., Bailey A.O., Sun B., Kuchar J.A., Green W.N. et al. (2006) Global analysis of protein palmitoylation in yeast. Cell 125, 1003–1013 doi: 10.1016/j.cell.2006.03.042 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Fukata M., Fukata Y., Adesnik H., Nicoll R.A. and Bredt D.S. (2004) Identification of PSD-95 palmitoylating enzymes. Neuron 44, 987–996 doi: 10.1016/j.neuron.2004.12.005 [DOI] [PubMed] [Google Scholar]
- 18.Lemonidis K., Sanchez-Perez M.C. and Chamberlain L.H. (2015) Identification of a novel sequence motif recognized by the ankyrin repeat domain of zDHHC17/13 S-acyltransferases. J. Biol. Chem. 290, 21939–21950 doi: 10.1074/jbc.M115.657668 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhang M.M., Wu P.-Y.J., Kelly F.D., Nurse P. and Hang H.C. (2013) Quantitative control of protein S-palmitoylation regulates meiotic entry in fission yeast. PLoS Biol. 11, e1001597 doi: 10.1371/journal.pbio.1001597 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Chai S., Cambronne X.A., Eichhorn S.W. and Goodman R.H. (2013) MicroRNA-134 activity in somatostatin interneurons regulates H-Ras localization by repressing the palmitoylation enzyme, DHHC9. Proc. Natl Acad. Sci. U.S.A. 110, 17898–17903 doi: 10.1073/pnas.1317528110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Li Y., Martin B.R., Cravatt B.F. and Hofmann S.L. (2012) DHHC5 protein palmitoylates flotillin-2 and is rapidly degraded on induction of neuronal differentiation in cultured cells. J. Biol. Chem. 287, 523–530 doi: 10.1074/jbc.M111.306183 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Noritake J., Fukata Y., Iwanaga T., Hosomi N., Tsutsumi R., Matsuda N. et al. (2009) Mobile DHHC palmitoylating enzyme mediates activity-sensitive synaptic targeting of PSD-95. J. Cell Biol. 186, 147–160 doi: 10.1083/jcb.200903101 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lai J. and Linder M.E. (2013) Oligomerization of DHHC protein S-acyltransferases. J. Biol. Chem. 288, 22862–22870 doi: 10.1074/jbc.M113.458794 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Swarthout J.T., Lobo S., Farh L., Croke M.R., Greentree W.K., Deschenes R.J. et al. (2005) DHHC9 and GCP16 constitute a human protein fatty acyltransferase with specificity for H- and N-Ras. J. Biol. Chem. 280, 31141–31148 doi: 10.1074/jbc.M504113200 [DOI] [PubMed] [Google Scholar]
- 25.Mitchell D.A., Hamel L.D., Ishizuka K., Mitchell G., Schaefer L.M. and Deschenes R.J. (2012) The Erf4 subunit of the yeast Ras palmitoyl acyltransferase is required for stability of the acyl-Erf2 intermediate and palmitoyl transfer to a Ras2 substrate. J. Biol. Chem. 287, 34337–34348 doi: 10.1074/jbc.M112.379297 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Duncan J.A. and Gilman A.G. (1998) A cytoplasmic acyl-protein thioesterase that removes palmitate from G protein alpha subunits and p21RAS. J. Biol. Chem. 273, 15830–15837 doi: 10.1074/jbc.273.25.15830 [DOI] [PubMed] [Google Scholar]
- 27.Kong E., Peng S., Chandra G., Sarkar C., Zhang Z., Bagh M.B. et al. (2013) Dynamic palmitoylation links cytosol-membrane shuttling of acyl-protein thioesterase-1 and acyl-protein thioesterase-2 with that of proto-oncogene H-ras product and growth-associated protein-43. J. Biol. Chem. 288, 9112–9125 doi: 10.1074/jbc.M112.421073 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Vartak N., Papke B., Grecco H.E., Rossmannek L., Waldmann H., Hedberg C. et al. (2014) The autodepalmitoylating activity of APT maintains the spatial organization of palmitoylated membrane proteins. Biophys. J. 106, 93–105 doi: 10.1016/j.bpj.2013.11.024 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Yokoi N., Fukata Y., Sekiya A., Murakami T., Kobayashi K. and Fukata M. (2016) Identification of PSD-95 depalmitoylating enzymes. J. Neurosci. 36, 6431–6444 doi: 10.1523/JNEUROSCI.0419-16.2016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Lin D.T. and Conibear E. (2015) ABHD17 proteins are novel protein depalmitoylases that regulate N-Ras palmitate turnover and subcellular localization. eLife 4, e11306 PMID: [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Child M.A., Hall C.I., Beck J.R., Ofori L.O., Albrow V.E., Garland M. et al. (2013) Small-molecule inhibition of a depalmitoylase enhances Toxoplasma host-cell invasion. Nat. Chem. Biol. 9, 651–656 doi: 10.1038/nchembio.1315 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Nile A.H. and Hannoush R.N. (2016) Fatty acylation of Wnt proteins. Nat. Chem. Biol. 12, 60–69 doi: 10.1038/nchembio.2005 [DOI] [PubMed] [Google Scholar]
- 33.Resh M.D. (2016) Fatty acylation of proteins: the long and the short of it. Prog. Lipid Res. 63, 120–131 doi: 10.1016/j.plipres.2016.05.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Charron G., Zhang M.M., Yount J.S., Wilson J., Raghavan A.S., Shamir E. et al. (2009) Robust fluorescent detection of protein fatty-acylation with chemical reporters. J. Am. Chem. Soc. 131, 4967–4975 doi: 10.1021/ja810122f [DOI] [PubMed] [Google Scholar]
- 35.Thinon E. and Hang H.C. (2015) Chemical reporters for exploring protein acylation. Biochem. Soc. Trans. 43, 253–261 doi: 10.1042/BST20150004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Martin B.R., Wang C., Adibekian A., Tully S.E. and Cravatt B.F. (2011) Global profiling of dynamic protein palmitoylation. Nat. Methods 9, 84–89 doi: 10.1038/nmeth.1769 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Zhang M.M., Tsou L.K., Charron G., Raghavan A.S. and Hang H.C. (2010) Tandem fluorescence imaging of dynamic S-acylation and protein turnover. Proc. Natl Acad. Sci. U.S.A. 107, 8627–8632 doi: 10.1073/pnas.0912306107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Peng T. and Hang H.C. (2015) Bifunctional fatty acid chemical reporter for analyzing S-palmitoylated membrane protein–protein interactions in mammalian cells. J. Am. Chem. Soc. 137, 556–559 doi: 10.1021/ja502109n [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Drisdel R.C. and Green W.N. (2004) Labeling and quantifying sites of protein palmitoylation. Biotechniques 36, 276–285 PMID: [DOI] [PubMed] [Google Scholar]
- 40.Forrester M.T., Hess D.T., Thompson J.W., Hultman R., Moseley M.A., Stamler J.S. et al. (2011) Site-specific analysis of protein S-acylation by resin-assisted capture. J. Lipid Res. 52, 393–398 doi: 10.1194/jlr.D011106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ji Y., Leymarie N., Haeussler D.J., Bachschmid M.M., Costello C.E. and Lin C. (2013) Direct detection of S-palmitoylation by mass spectrometry. Anal. Chem. 85, 11952–11959 doi: 10.1021/ac402850s [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Percher A., Ramakrishnan S., Thinon E., Yuan X., Yount J.S. and Hang H.C. (2016) Mass-tag labeling reveals site-specific and endogenous levels of protein S-fatty acylation. Proc. Natl Acad. Sci. U.S.A. 113, 4302–4307 doi: 10.1073/pnas.1602244113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Howie J., Reilly L., Fraser N.J., Vlachaki Walker J.M., Wypijewski K.J., Ashford M.L.J. et al. (2014) Substrate recognition by the cell surface palmitoyl transferase DHHC5. Proc. Natl Acad. Sci. U.S.A. 111, 17534–17539 doi: 10.1073/pnas.1413627111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Jones M.L., Collins M.O., Goulding D., Choudhary J.S. and Rayner J.C. (2012) Analysis of protein palmitoylation reveals a pervasive role in Plasmodium development and pathogenesis. Cell Host Microbe 12, 246–258 doi: 10.1016/j.chom.2012.06.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Hemsley P.A., Weimar T., Lilley K.S., Dupree P. and Grierson C.S. (2013) A proteomic approach identifies many novel palmitoylated proteins in Arabidopsis. New. Phytol. 197, 805–814 doi: 10.1111/nph.12077 [DOI] [PubMed] [Google Scholar]
- 46.Srivastava V., Weber J.R., Malm E., Fouke B.W. and Bulone V. (2016) Proteomic analysis of a poplar cell suspension culture suggests a major role of protein S-acylation in diverse cellular processes. Front. Plant Sci. 7, 477 doi: 10.3389/fpls.2016.00477 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Kang R., Wan J., Arstikaitis P., Takahashi H., Huang K., Bailey A.O. et al. (2008) Neural palmitoyl-proteomics reveals dynamic synaptic palmitoylation. Nature 456, 904–909 doi: 10.1038/nature07605 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Martin B.R. and Cravatt B.F. (2009) Large-scale profiling of protein palmitoylation in mammalian cells. Nat. Methods 6, 135–138 doi: 10.1038/nmeth.1293 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Yount J.S., Moltedo B., Yang Y.-Y., Charron G., Moran T.M., López C.B. et al. (2010) Palmitoylome profiling reveals S-palmitoylation-dependent antiviral activity of IFITM3. Nat. Chem. Biol. 6, 610–614 doi: 10.1038/nchembio.405 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Ren J., Wen L., Gao X., Jin C., Xue Y. and Yao X. (2008) CSS-Palm 2.0: an updated software for palmitoylation sites prediction. Protein Eng. Des. Sel. 21, 639–644 doi: 10.1093/protein/gzn039 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kumari B., Kumar R. and Kumar M. (2014) Palmpred: an SVM based palmitoylation prediction method using sequence profile information. PLoS ONE 9, e89246 doi: 10.1371/journal.pone.0089246 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Li Y.-X., Shao Y.-H. and Deng N.-Y. (2011) Improved prediction of palmitoylation sites using PWMs and SVM. Protein Pept. Lett. 18, 186–193 doi: 10.2174/092986611794475084 [DOI] [PubMed] [Google Scholar]
- 53.Wang X.-B., Wu L.-Y., Wang Y.-C. and Deng N.-Y. (2009) Prediction of palmitoylation sites using the composition of k-spaced amino acid pairs. Protein Eng. Des. Sel. 22, 707–712 doi: 10.1093/protein/gzp055 [DOI] [PubMed] [Google Scholar]
- 54.Blanc M., David F., Abrami L., Migliozzi D., Armand F., Burgi J. et al. (2015) SwissPalm: protein palmitoylation database. F1000Research 4, 261 doi: 10.12688/f1000research.6464.1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Yeste-Velasco M., Linder M.E. and Lu Y.J. (2015) Protein S-palmitoylation and cancer. Biochim. Biophys. Acta 1856, 107–120 doi: 10.1016/j.bbcan.2015.06.004 [DOI] [PubMed] [Google Scholar]
- 56.Sánchez-Mir L., Franco A., Martín-García R., Madrid M., Vicente-Soler J., Soto T. et al. (2014) Rho2 palmitoylation is required for plasma membrane localization and proper signaling to the fission yeast cell integrity mitogen-activated protein kinase pathway. Mol. Cell Biol. 34, 2745–2759 doi: 10.1128/MCB.01515-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Santiago-Tirado F.H., Peng T., Yang M., Hang H.C. and Doering T.L. (2015) A single protein S-acyl transferase acts through diverse substrates to determine cryptococcal morphology, stress tolerance, and pathogenic outcome. PLoS Pathog. 11, e1004908 doi: 10.1371/journal.ppat.1004908 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Nichols C.B., Ferreyra J., Ballou E.R. and Alspaugh J.A. (2009) Subcellular localization directs signaling specificity of the Cryptococcus neoformans Ras1 protein. Eukaryot. Cell 8, 181–189 doi: 10.1128/EC.00351-08 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Nichols C.B., Ost K.S., Grogan D.P., Pianalto K., Hasan S. and Alspaugh J.A. (2015) Impact of protein palmitoylation on the virulence potential of Cryptococcus neoformans. Eukaryot. Cell 14, 626–635 doi: 10.1128/EC.00010-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Park S., Patterson E.E., Cobb J., Audhya A., Gartenberg M.R. and Fox C.A. (2011) Palmitoylation controls the dynamics of budding-yeast heterochromatin via the telomere-binding protein Rif1. Proc. Natl Acad. Sci. U.S.A. 108, 14572–14577 doi: 10.1073/pnas.1105262108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Alonso A.M., Coceres V.M., De Napoli M.G., Nieto Guil A.F., Angel S.O. and Corvi M.M. (2012) Protein palmitoylation inhibition by 2-bromopalmitate alters gliding, host cell invasion and parasite morphology in Toxoplasma gondii. Mol. Biochem. Parasitol. 184, 39–43 doi: 10.1016/j.molbiopara.2012.03.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Foe I.T., Child M.A., Majmudar J.D., Krishnamurthy S., van der Linden W.A., Ward G.E. et al. (2015) Global analysis of palmitoylated proteins in Toxoplasma gondii. Cell Host Microbe 18, 501–511 doi: 10.1016/j.chom.2015.09.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Rees-Channer R.R., Martin S.R., Green J.L., Bowyer P.W., Grainger M., Molloy J.E. et al. (2006) Dual acylation of the 45 kDa gliding-associated protein (GAP45) in Plasmodium falciparum merozoites. Mol. Biochem. Parasitol. 149, 113–116 doi: 10.1016/j.molbiopara.2006.04.008 [DOI] [PubMed] [Google Scholar]
- 64.Beck J.R., Fung C., Straub K.W., Coppens I., Vashisht A.A., Wohlschlegel J.A. et al. (2013) A Toxoplasma palmitoyl acyl transferase and the palmitoylated armadillo repeat protein TgARO govern apical rhoptry tethering and reveal a critical role for the rhoptries in host cell invasion but not egress. PLoS Pathog. 9, e1003162 doi: 10.1371/journal.ppat.1003162 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Cabrera A., Herrmann S., Warszta D., Santos J.M., John Peter A.T., Kono M. et al. (2012) Dissection of minimal sequence requirements for rhoptry membrane targeting in the malaria parasite. Traffic 13, 1335–1350 doi: 10.1111/j.1600-0854.2012.01394.x [DOI] [PubMed] [Google Scholar]
- 66.Frénal K., Tay C.L., Mueller C., Bushell E.S., Jia Y., Graindorge A. et al. (2013) Global analysis of apicomplexan protein S-acyl transferases reveals an enzyme essential for invasion. Traffic 14, 895–911 doi: 10.1111/tra.12081 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Hodson N., Invergo B., Rayner J.C. and Choudhary J.S. (2015) Palmitoylation and palmitoyl-transferases in Plasmodium parasites. Biochem. Soc. Trans. 43, 240–245 doi: 10.1042/BST20140289 [DOI] [PubMed] [Google Scholar]
- 68.Wetzel J., Herrmann S., Swapna L.S., Prusty D., John Peter A.T.J., Kono M. et al. (2015) The role of palmitoylation for protein recruitment to the inner membrane complex of the malaria parasite. J. Biol. Chem. 290, 1712–1728 doi: 10.1074/jbc.M114.598094 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Santos J.M., Kehrer J., Franke-Fayard B., Frischknecht F., Janse C.J. and Mair G.R. (2015) The Plasmodium palmitoyl-S-acyl-transferase DHHC2 is essential for ookinete morphogenesis and malaria transmission. Sci. Rep. 5, 16034 doi: 10.1038/srep16034 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Santos J.M., Duarte N., Kehrer J., Ramesar J., Avramut M.C., Koster A.J. et al. (2016) Maternally supplied S-acyl-transferase is required for crystalloid organelle formation and transmission of the malaria parasite. Proc. Natl Acad. Sci. U.S.A. 113, 7183–7188 doi: 10.1073/pnas.1522381113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Tay C.L., Jones M.L., Hodson N., Theron M., Choudhary J.S. and Rayner J.C. (2016) Study of Plasmodium falciparum DHHC palmitoyl transferases identifies a role for PfDHHC9 in gametocytogenesis. Cell Microbiol. 18, 1596–1610 doi: 10.1111/cmi.12599 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Hopp C.S., Balaban A.E., Bushell E.S.C., Billker O., Rayner J.C. and Sinnis P. (2016) Palmitoyl transferases have critical roles in the development of mosquito and liver stages of Plasmodium. Cell Microbiol. 18, 1625–1641 doi: 10.1111/cmi.12601 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Merino M.C., Zamponi N., Vranych C.V., Touz M.C. and Rópolo A.S. (2014) Identification of Giardia lamblia DHHC proteins and the role of protein S-palmitoylation in the encystation process. PLoS Negl. Trop. Dis. 8, e2997 doi: 10.1371/journal.pntd.0002997 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Sanders S.S., Martin D.D.O., Butland S.L., Lavallée-Adam M., Calzolari D., Kay C. et al. (2015) Curation of the mammalian palmitoylome indicates a pivotal role for palmitoylation in diseases and disorders of the nervous system and cancers. PLoS Comput. Biol. 11, e1004405 doi: 10.1371/journal.pcbi.1004405 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Fukata Y. and Fukata M. (2010) Protein palmitoylation in neuronal development and synaptic plasticity. Nat. Rev. Neurosci. 11, 161–175 doi: 10.1038/nrn2788 [DOI] [PubMed] [Google Scholar]
- 76.Brichta L., Shin W., Jackson-Lewis V., Blesa J., Yap E.-L., Walker Z. et al. (2015) Identification of neurodegenerative factors using translatome–regulatory network analysis. Nat. Neurosci. 18, 1325–1333 doi: 10.1038/nn.4070 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Moutin E., Nikonenko I., Stefanelli T., Wirth A., Ponimaskin E., De Roo M. et al. (2016) Palmitoylation of cdc42 promotes spine stabilization and rescues spine density deficit in a mouse model of 22q11.2 deletion syndrome. Cereb. Cortex doi: 10.1093/cercor/bhw183 [DOI] [PubMed] [Google Scholar]
- 78.Mukai J., Tamura M., Fénelon K., Rosen A.M., Spellman T.J., Kang R. et al. (2015) Molecular substrates of altered axonal growth and brain connectivity in a mouse model of schizophrenia. Neuron 86, 680–695 doi: 10.1016/j.neuron.2015.04.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Mukai J., Dhilla A., Drew L.J., Stark K.L., Cao L., MacDermott A.B. et al. (2008) Palmitoylation-dependent neurodevelopmental deficits in a mouse model of 22q11 microdeletion. Nat. Neurosci. 11, 1302–1310 doi: 10.1038/nn.2204 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Nishimura A. and Linder M.E. (2013) Identification of a novel prenyl and palmitoyl modification at the CaaX motif of Cdc42 that regulates RhoGDI binding. Mol. Cell Biol. 33, 1417–1429 doi: 10.1128/MCB.01398-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Arstikaitis P., Gauthier-Campbell C., Herrera R.C.G., Huang K., Levinson J.N., Murphy T.H. et al. (2008) Paralemmin-1, a modulator of filopodia induction is required for spine maturation. Mol. Biol. Cell 19, 2026–2038 doi: 10.1091/mbc.E07-08-0802 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.George J., Soares C., Montersino A., Beique J.-C. and Thomas G.M. (2015) Palmitoylation of LIM kinase-1 ensures spine-specific actin polymerization and morphological plasticity. eLife 4, e06327 doi: 10.7554/eLife.06327 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.El-Husseini A.E.-D., Schnell E., Dakoji S., Sweeney N., Zhou Q., Prange O. et al. (2002) Synaptic strength regulated by palmitate cycling on PSD-95. Cell 108, 849–863 doi: 10.1016/S0092-8674(02)00683-9 [DOI] [PubMed] [Google Scholar]
- 84.Brigidi G.S., Sun Y., Beccano-Kelly D., Pitman K., Mobasser M., Borgland S.L. et al. (2014) Palmitoylation of δ-catenin by DHHC5 mediates activity-induced synapse plasticity. Nat. Neurosci. 17, 522–532 doi: 10.1038/nn.3657 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Chen X., Du Z., Li X., Wang L., Wang F., Shi W. et al. (2016) Protein palmitoylation regulates neural stem cell differentiation by modulation of EID1 activity. Mol. Neurobiol. 53, 5722–5736 doi: 10.1007/s12035-015-9481-y [DOI] [PubMed] [Google Scholar]
- 86.Runkle K.B., Kharbanda A., Stypulkowski E., Cao X.-J., Wang W., Garcia B.A. et al. (2016) Inhibition of DHHC20-mediated EGFR palmitoylation creates a dependence on EGFR signaling. Mol. Cell 62, 385–396 doi: 10.1016/j.molcel.2016.04.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Pedram A., Razandi M., Sainson R.C.A., Kim J.K., Hughes C.C. and Levin E.R. (2007) A conserved mechanism for steroid receptor translocation to the plasma membrane. J. Biol. Chem. 282, 22278–22288 doi: 10.1074/jbc.M611877200 [DOI] [PubMed] [Google Scholar]
- 88.Pedram A., Razandi M., Deschenes R.J. and Levin E.R. (2012) DHHC-7 and -21 are palmitoylacyltransferases for sex steroid receptors. Mol. Biol. Cell 23, 188–199 doi: 10.1091/mbc.E11-07-0638 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Acconcia F., Ascenzi P., Bocedi A., Spisni E., Tomasi V., Trentalance A. et al. (2005) Palmitoylation-dependent estrogen receptor α membrane localization: regulation by 17β-estradiol. Mol. Biol. Cell 16, 231–237 doi: 10.1091/mbc.E04-07-0547 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Mill P., Lee A.W.S., Fukata Y., Tsutsumi R., Fukata M., Keighren M. et al. (2009) Palmitoylation regulates epidermal homeostasis and hair follicle differentiation. PLoS Genet. 5, e1000748 doi: 10.1371/journal.pgen.1000748 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Chen B., Zheng B., DeRan M., Jarugumilli G.K., Fu J., Brooks Y.S. et al. (2016) ZDHHC7-mediated S-palmitoylation of Scribble regulates cell polarity. Nat. Chem. Biol. 12, 686–693 doi: 10.1038/nchembio.2119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Rossin A., Durivault J., Chakhtoura-Feghali T., Lounnas N., Gagnoux-Palacios L. and Hueber A.-O. (2015) Fas palmitoylation by the palmitoyl acyltransferase DHHC7 regulates Fas stability. Cell Death Diff. 22, 643–653 doi: 10.1038/cdd.2014.153 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Akimzhanov A.M. and Boehning D. (2015) Rapid and transient palmitoylation of the tyrosine kinase Lck mediates Fas signaling. Proc. Natl Acad. Sci. U.S.A. 112, 11876–11880 doi: 10.1073/pnas.1509929112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Saleem A.N., Chen Y.-H., Baek H.J., Hsiao Y.-W., Huang H.-W., Kao H.-J. et al. (2010) Mice with alopecia, osteoporosis, and systemic amyloidosis due to mutation in Zdhhc13, a gene coding for palmitoyl acyltransferase. PLoS Genet. 6, e1000985 doi: 10.1371/journal.pgen.1000985 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Batistic O. (2012) Genomics and localization of the Arabidopsis DHHC-cysteine-rich domain S-acyltransferase protein family. Plant Physiol. 160, 1597–1612 doi: 10.1104/pp.112.203968 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Hurst C.H. and Hemsley P.A. (2015) Current perspective on protein S-acylation in plants: more than just a fatty anchor? J. Exp. Bot. 66, 1599–1606 doi: 10.1093/jxb/erv053 [DOI] [PubMed] [Google Scholar]
- 97.Hemsley P.A. (2015) The importance of lipid modified proteins in plants. New Phytol. 205, 476–489 doi: 10.1111/nph.13085 [DOI] [PubMed] [Google Scholar]
- 98.Li Y., Scott R., Doughty J., Grant M. and Qi B. (2016) Protein S-acyltransferase 14: a specific role for palmitoylation in leaf senescence in Arabidopsis. Plant Physiol. 170, 415–428 doi: 10.1104/pp.15.00448 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Lai J., Yu B., Cao Z., Chen Y., Wu Q., Huang J. et al. (2015) Two homologous protein S-acyltransferases, PAT13 and PAT14, cooperatively regulate leaf senescence in Arabidopsis. J. Exp. Bot. 66, 6345–6353 doi: 10.1093/jxb/erv347 [DOI] [PubMed] [Google Scholar]
- 100.Liu J., Zhong S., Guo X., Hao L., Wei X., Huang Q. et al. (2013) Membrane-bound RLCKs LIP1 and LIP2 are essential male factors controlling male-female attraction in Arabidopsis. Curr. Biol. 23, 993–998 doi: 10.1016/j.cub.2013.04.043 [DOI] [PubMed] [Google Scholar]
- 101.Martin M.L. and Busconi L. (2000) Membrane localization of a rice calcium-dependent protein kinase (CDPK) is mediated by myristoylation and palmitoylation. Plant J. 24, 429–435 doi: 10.1046/j.1365-313x.2000.00889.x [DOI] [PubMed] [Google Scholar]
- 102.Batistič O., Rehers M., Akerman A., Schlücking K., Steinhorst L., Yalovsky S. et al. (2012) S-acylation-dependent association of the calcium sensor CBL2 with the vacuolar membrane is essential for proper abscisic acid responses. Cell Res. 22, 1155–1168 doi: 10.1038/cr.2012.71 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Batistic O., Sorek N., Schultke S., Yalovsky S. and Kudla J. (2008) Dual fatty acyl modification determines the localization and plasma membrane targeting of CBL/CIPK Ca2+ signaling complexes in Arabidopsis. Plant Cell 20, 1346–1362 doi: 10.1105/tpc.108.058123 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Kumar M., Wightman R., Atanassov I., Gupta A., Hurst C.H., Hemsley P.A. et al. (2016) S-Acylation of the cellulose synthase complex is essential for its plasma membrane localization. Science 353, 166–169 doi: 10.1126/science.aaf4009 [DOI] [PubMed] [Google Scholar]
- 105.Chen X., Shi W., Wang F., Du Z., Yang Y., Gao M. et al. (2014) Zinc finger DHHC-type containing 13 regulates fate specification of ectoderm and mesoderm cell lineages by modulating Smad6 activity. Stem Cells Dev. 23, 1899–1909 doi: 10.1089/scd.2014.0068 [DOI] [PubMed] [Google Scholar]
- 106.Du Z., Chen X., Li X., He K., Ji S., Shi W. et al. (2016) Protein palmitoylation activate zygotic gene expression during the maternal-to-zygotic transition. Biochem. Biophys. Res. Commun. 475, 194–201 doi: 10.1016/j.bbrc.2016.05.074 [DOI] [PubMed] [Google Scholar]
- 107.Shi W., Chen X., Wang F., Gao M., Yang Y., Du Z. et al. (2016) ZDHHC16 modulates FGF/ERK dependent proliferation of neural stem/progenitor cells in the zebrafish telencephalon. Dev. Neurobiol. 76, 1014–1028 doi: 10.1002/dneu.22372 [DOI] [PubMed] [Google Scholar]
- 108.Wang F., Chen X., Shi W., Yao L., Gao M., Yang Y. et al. (2015) Zdhhc15b regulates differentiation of diencephalic dopaminergic neurons in zebrafish. J. Cell. Biochem. 116, 2980–2991 doi: 10.1002/jcb.25256 [DOI] [PubMed] [Google Scholar]
- 109.Gleason E.J., Lindsey W.C., Kroft T.L., Singson A.W. and L'Hernault S.W. (2006) spe-10 encodes a DHHC–CRD zinc-finger membrane protein required for endoplasmic reticulum/Golgi membrane morphogenesis during Caenorhabditis elegans spermatogenesis. Genetics 172, 145–158 doi: 10.1534/genetics.105.047340 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Edmonds M.J. and Morgan A. (2014) A systematic analysis of protein palmitoylation in Caenorhabditis elegans. BMC Genomics 15, 841 doi: 10.1186/1471-2164-15-841 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Bannan B.A., Van Etten J., Kohler J.A., Tsoi Y., Hansen N.M., Sigmon S. et al. (2008) The Drosophila protein palmitoylome: characterizing palmitoyl-thioesterases and DHHC palmitoyl-transferases. Fly 2, 198–214 doi: 10.4161/fly.6621 [DOI] [PMC free article] [PubMed] [Google Scholar]