Abstract
Background
There has been an emergence and expansion of tick-borne diseases in Europe, Asia and North America in recent years, including Lyme disease, tick-borne encephalitis and human anaplasmosis. The primary vectors implicated are hard ticks of the genus Ixodes. Although much is known about the host response to these bacterial and viral pathogens, there is limited knowledge of the cellular responses to infection within the tick vector. The bacterium Anaplasma phagocytophilum is able to bypass apoptotic processes in ticks, enabling infection to proceed. However, the tick cellular responses to infection with the flaviviruses tick-borne encephalitis virus (TBEV) and louping ill virus (LIV), which cause tick-borne encephalitis and louping ill respectively, are less clear.
Results
Infection and transcriptional analysis of the Ixodes ricinus tick cell line IRE/CTVM20 with the viruses LIV and TBEV, and the bacterium A. phagocytophilum, identified activation of common and distinct cellular pathways. In particular, commonly-upregulated genes included those that modulate apoptotic pathways, putative anti-pathogen genes, and genes that influence the tick innate immune response, including selective activation of toll genes.
Conclusion
These data provide an insight into potential key genes involved in the tick cellular response to viral or bacterial infection, which may promote cell survival and host resistance.
Electronic supplementary material
The online version of this article (doi:10.1186/s13071-017-2011-1) contains supplementary material, which is available to authorized users.
Keywords: Tick cell, Flavivirus, Anaplasma phagocytophilum, Ixodes ricinus, Apoptosis, Toll, Transcriptomics, Immunology
Background
The past few decades have seen the emergence and expansion of tick-borne diseases in Europe, Asia and North America [1]. These include Lyme disease, tick-borne encephalitis, human granulocytic anaplasmosis (HGA) and louping ill [2–5]. Hard ticks of the genus Ixodes are primarily implicated, including Ixodes ricinus, Ixodes persulcatus and Ixodes scapularis.
The flavivirus tick-borne encephalitis virus (TBEV) is responsible for an increasing number of cases of human tick-borne encephalitis (TBE) throughout Europe and Asia. Infection with the virus can cause severe encephalitic disease which can be fatal in some cases [6]. Louping ill virus (LIV), which is genetically and antigenically closely-related to TBEV, also causes severe encephalitis and is responsible for disease in sheep in the United Kingdom (UK) and discrete parts of mainland Europe [5]. Both viruses are transmitted by I. ricinus, although in Asia, TBEV is transmitted predominantly by I. persulcatus [7]. Anaplasma phagocytophilum is considered an emerging zoonotic bacterium, transmitted by I. ricinus ticks in Europe, and I. scapularis in the United States [8]. Anaplasma phagocytophilum infects vertebrate host granulocytes, leading to human, canine or equine granulocytic anaplasmosis and to tick-borne fever in ruminants [9–11].
The biological effect on I. ricinus ticks of infection with these pathogens has yet to be fully characterised, and genes associated with apoptosis and innate immune function are of particular interest, as these pathways are crucially involved in the cellular response to infection. The induction of apoptosis serves a range of functions in the vertebrate host, including control at the cellular level following infection [12]. Previous studies have shown that A. phagocytophilum is able to inhibit this process in I. scapularis ticks and human cells, through inhibition of different apoptotic pathways, leading to increased bacterial dissemination [13]. Subsequent studies have shown that the transcriptional response to A. phagocytophilum infection in an I. ricinus cell line was similar to that detected in I. scapularis midguts [14, 15], where the response did not associate the intrinsic apoptotic pathway with the inhibition of cellular apoptosis, but did suggest a role for the janus-associated kinase-signal transducer and activator of transcription (Jak-STAT) pathway via upregulation of Jak [15]. Along with the Jak-STAT pathway, the Toll pathway is known to constitute part of the innate immune response in arthropods [16]. A number of recent studies have investigated the response of tick cells to virus infection and provided preliminary data on the pathways activated by flaviviruses [17–19].
In this study, the transcriptional response of an I. ricinus cell line to LIV and TBEV infection was investigated, and compared to that observed following A. phagocytophilum infection. All infection experiments were undertaken simultaneously, and the dataset derived from A. phagocytophilum infection has previously been utilised to investigate apoptosis in a comparison with infection in I. scapularis cells [15]. The utilisation of a systems biology approach using high-throughput omics technology has enabled the generation of large datasets yielding evidence of differential gene expression associated with both apoptotic and innate immune pathways. Furthermore, evidence for increased expression of anti-pathogen genes is demonstrated. The application of Next Generation Sequencing (NGS) and subsequent transcriptomic analysis has provided an insight into the tick cell response to virus or bacterial infection, and enhanced our understanding of the tick-pathogen interface.
Methods
Virus and bacterial isolates
The virus isolates used were LIV strain LI3/1 (APHA reference: Arb 126), which was originally isolated from a sheep in Oban, Scotland, in 1962, and the TBEV strain Neudorfl H2J (APHA reference: Arb 131), originally isolated from an I. ricinus tick in Austria in the early 1950s. Both isolates were mouse brain homogenates, kindly provided by Professor John Stephenson (Public Health England, formerly Centre for Applied Microbiology and Research, Porton Down, UK). The TBEV isolate was originally isolated by Dr Christian Kunz, University of Vienna, Austria, and had subsequently been passaged four times in an outbred strain of mice. However, it remains genetically identical to the standard prototype Neudoerfl strain. The LIV isolate was originally isolated by Dr Hugh Reid, Moredun Institute, Scotland, and had been passaged four times in sheep and six times in an outbred strain of mice. The bacterial isolate was A. phagocytophilum NY-18, which was originally isolated from a human in 1996 [20, 21]. The isolate was subsequently passaged in I. scapularis tick cells prior to infection of I. ricinus cells.
Ixodes ricinus cell line
The I. ricinus embryo-derived tick cell line IRE/CTVM20 [22] (provided by the Tick Cell Biobank, The Pirbright Institute, UK) was maintained in a 1:1 mixture of supplemented L-15 (Leibovitz) medium and L-15B medium [23], as previously described [24]. Briefly, the supplemented L-15 medium contained 20% foetal bovine serum (FBS), 10% tryptose phosphate broth (TPB), 2 mM L-glutamine, 100 μg/ml streptomycin and 100 U/ml penicillin. The L-15B medium included 10% TPB, 5% FBS, 0.1% bovine lipoprotein concentrate, 2 mM L-glutamine, 100 μg/ml streptomycin, 100 U/ml penicillin and a range of vitamin and mineral supplements, as previously described [23].
Infection of tick cells with bacterial and viral isolates
IRE/CTVM20 cells were seeded in 12-well tissue culture plates (Greiner), at a cell density of 6 × 105 cells/well. Quadruplicate wells were infected with either louping ill virus (LIV), tick-borne encephalitis virus (TBEV) or A. phagocytophilum at a multiplicity of infection (MOI) of 1, alongside uninfected control wells. Plates were incubated at 28 °C without additional CO2; this was the optimal temperature for growth of this cell line, as utilised in previous studies [24–26]. Samples were taken at 0, 72, 120 and 168 hours post-infection (hpi), and cells and medium within each well were resuspended and transferred to sterile cryotubes at each time-point. Following centrifugation at 2000 rpm (366× g) for 15 min, the supernatants were transferred to fresh tubes for virus titration. The cell pellets were mixed with 0.5 ml of TRIzol® reagent (Invitrogen, Life Technologies, Paisley, UK) for RNA extraction.
Extraction of RNA from infected tick cells
Total RNA was extracted from the cell pellets combined with TRIzol® reagent (Invitrogen), according to the manufacturer’s instructions. The RNA was further purified using an RNeasy Mini kit (QIAgen, Manchester, UK), according to the manufacturer’s instructions, and quantified spectrophotometrically. Each RNA sample was assessed by RT-PCR for evidence of virus replication prior to further treatment of the remaining RNA in preparation for sequencing.
Confirmation of virus growth in tick cells
To assess the growth of LIV and TBEV in tick cells, both the cells and the cell supernatants were assessed at each time-point. To assess growth in cells, extracted RNA was reverse-transcribed using Moloney Murine Leukemia Virus Reverse Transcriptase (M-MLV RT; Promega, Southampton, UK) and random hexamers (Roche, Burgess Hill, UK), according to the manufacturers’ instructions. Positive controls were included, using RNA previously extracted from the virus isolates used as inoculum. Molecular standards with known copy numbers were prepared from the positive control cDNA samples, and were used to quantify the viral copy numbers in the test samples by qPCR using SYBR Green Jumpstart Taq Readymix (Sigma-Aldrich, Gillingham, UK) and TBEV/LIV-specific primers. Similarly, host standards were prepared from RNA extracted from uninfected IRE/CTVM20 cells. Acari primers (Additional file 1: Table S1) were used to quantify 16S copies in the test samples, to allow normalisation of results. To quantify virus titre of the cell supernatants at each time-point, the supernatants were assessed by plaque assay on Vero C1008 cells, using standard techniques [27].
Confirmation of bacterial growth in tick cells
Growth of A. phagocytophilum in tick cells was confirmed by qPCR and microscopy of Giemsa-stained slides. DNA was extracted from tick cells using Tri Reagent (Sigma-Aldrich) following the manufacturer’s instructions. Anaplasma phagocytophilum DNA levels were characterised by major surface protein 4 gene (msp4) real-time PCR with normalisation against the level of tick 16S ribosomal RNA (rRNA) as described previously [13]. Giemsa-stained cytocentrifuge smears were prepared from resuspended cell suspensions, and were inspected microscopically to monitor infection.
Next generation sequencing (NGS)
rRNA was depleted from extracted total RNA, using Terminator exonuclease (Epicentre [Illumina], Madison, USA) according to the manufacturer’s instructions. Prior to sequencing, the RNA was quantified (Nanodrop, Nanodrop Products, Thermo Scientific), to confirm that it was at a suitable concentration for Illumina sequencing (> 10 ng/μl).
Sequencing was undertaken of duplicate RNA samples from 0 hpi uninfected and 168 hpi infected cells only. RNA (200 ng) was reverse-transcribed to generate double-stranded cDNA, using the cDNA Synthesis System (Roche) and random hexamers. Illumina sequencing libraries were prepared using the Nextera XT system (Illumina, Madison, USA), and these were sequenced using an Illumina GA IIx instrument.
Bioinformatics and determination of differential gene expression
Sequence analysis was undertaken using multiplexed paired-end samples. Pre-analysis sequence quality checking was performed using the ‘FastQC’ programme (Barham Institute). The Ixodes scapularis genome was used as a reference for mapping (Broad Institute assembly lscaW1), and the programme ‘BowTie 2’ was used as an assembler, to align sequenced reads with the reference sequence. ‘TopHat’ was used to analyse the mapping results and identify splice junctions between exons. The ‘Cufflinks’ programme was used to provide an estimation of gene and isoform abundance and differential expression, allowing for splice variants and gaps due to the genome reference. Within Cufflinks, ‘Cuffmerge’ was used to merge Cufflinks assemblies, to provide normalisation of biological replicates. ‘Cuffquant’ was used to provide abundance estimation across normalised samples. The ‘Cuffdiff’ algorithm was used to account for biological variability between samples and identify differentially expressed genes; this included non-statistical analysis (log fold-change) and statistical analysis (test for variance), in order to identify statistically significant fold-changes in gene expression (P < 0.05, q < 0.05).
Genome coverage and depth statistics were generated using the ‘Qualimap’ programme [28, 29].
RNAseq data derived from LIV and TBEV-infected tick cells have been deposited in NCBI’s Gene Expression Omnibus, and are available through GEO Series accession number (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE85300). RNAseq data derived from A. phagocytophilum-infected tick cells has been previously analysed and published [15], and is available through GEO Series accession number (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE76906).
Molecular detection of transcripts
Selected differentially-expressed genes were detected by qPCR using specifically-designed primer pairs, and the primer sequences are detailed in Additional file 1: Table S1. The genes selected were Toll genes (ISCW022740, ISCW007727, ISCW007724 and ISCW017724) and Myeloid differentiation factor 88 (MyD88) gene (ISCW008802), along with Acari 16S rRNA as a housekeeping gene. Briefly, RNA was reverse transcribed with M-MLV RT (Promega) and random hexamers (Roche), in the presence of 14 units of RNAsin and 10 mM dithiothreitol (DTT), with incubation for 60 min at 42 °C.
Amplification using transcript-specific primer sets was performed using SYBR® Green JumpStartTM Taq ReadymixTM (Sigma-Aldrich) and an Mx3005p (Stratagene [Agilent], Santa Clara, USA). Amplification was achieved using specific primers for each mRNA transcript, which was quantified by comparison with a standard curve. The amount of 16S rRNA transcript was used to normalise each transcript within the sample. Transcript fold changes were calculated relative to the uninfected control cells at 0 hpi as previously described [30], with three test groups, A. phagocytophilum, LIV and TBEV, sampled at 72 hpi and 120 hpi. Results are reflected as transcript mean fold-change, with standard error of the mean (SEM). Upregulated transcripts have fold-change > 1, and downregulated transcripts have fold-change < 1. Statistical analysis was performed using a Student’s t-test with unequal variance. Statistically significant differences between the results from infected samples in comparison to the uninfected controls were denoted as follows: *P < 0.05; **P < 0.01.
Results
Pathogen growth in tick cells
Infection of IRE/CTVM20 cells with LIV, TBEV or A. phagocytophilum did not appear to inhibit cell growth or alter the cell morphology over the course of the experiment, as previously demonstrated [18, 25]. These observations were based on comparison between infected and uninfected cells using cell counts and examination of cell morphology under the microscope. Growth of LIV and TBEV was confirmed in both cell supernatants and cells (Fig. 1a) and of A. phagocytophilum in cells (Fig. 1b).
Fig. 1.
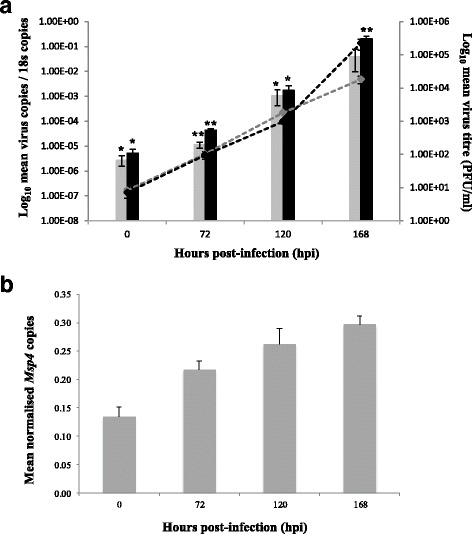
a Replication of LIV (grey) and TBEV (black) in IRE/CTVM20 cells, as demonstrated by log10 mean virus copies/16S copies detected in cellular-derived RNA (dashed lines), and virus titre (PFU/ml) quantified in cell supernatant (solid bars). b Replication of Anaplasma phagocytophilum in IRE/CTVM20 cells, as demonstrated by mean Msp4 copies normalised against mean 16S copies in cellular-derived RNA. Error bars represent the standard deviation. *P < 0.05; **P < 0.01
Infection with LIV, TBEV and A. phagocytophilum induces differential gene expression in a tick cell line
Initial gene expression analysis using RNA derived from A. phagocytophilum-infected tick cells has recently been published [15], but the extensive dataset generated has been further utilised here for comparison with flavivirus-induced gene expression. Infection of IRE/CTVM20 cells with LIV or TBEV induced differential gene expression (P < 0.05 and q < 0.05) as shown previously for A. phagocytophilum infection in the same cell line [15] and TBEV in another I. ricinus cell line IRE/CTVM19 [18]. These analyses have identified which of the annotated genes were differentially expressed in response to infection with LIV (samples 17 and 18) and TBEV (samples 19 and 20) when compared to uninfected control cells (samples 1–4). Details of the sequence depth and quality of alignment for these samples are provided in Additional file 2: Table S2, and suggest that the sequencing coverage was generally uniform across all of the samples included in the study, with between 38.34 and 50.65% of aligned pairs being within exon annotated sections of the reference genome. However, as we were not attempting to identify novel transcripts there was no requirement for ultra-deep sequencing. Additionally, the statistics for A. phagocytophilum (samples 15 and 16), of which some transcriptomic data has been previously published [15], have also been included in this table for comparison.
Following LIV and TBEV infection, RNAseq analysis identified 20,861 genes in IRE/CTVM20 cells, which was identical to results reported previously for A. phagocytophilum [15] (Table 1). When comparing infected cells at 168 hpi with uninfected cells at 0 hpi, analysis of differential expression identified 613 and 409 differentially expressed genes for LIV and TBEV infection, respectively, in comparison to 197 genes following A. phagocytophilum infection, as previously reported [15]. When compared to A. phagocytophilum infection, this suggests that flavivirus infection in tick cells leads to the differential expression of a greater number of genes, with 0.95, 2.96 and 1.98% of genes differentially expressed for A. phagocytophilum, LIV and TBEV, respectively. Although this suggests differences between the transcriptional responses to each pathogen, the limited number of samples in the study did not enable any correlation between levels of infection and levels of RNA expression to be inferred. However, it is possible that the low infection rate of A. phagocytophilum in IRE/CTVM20 cells observed (Fig. 1) may provide an alternative explanation for this difference. Following infection with all three pathogens, the majority of differentially expressed genes in tick cells were upregulated, rather than downregulated, with 78.7, 65.6 and 61.6% of genes upregulated following A. phagocytophilum, LIV and TBEV infection, respectively. The numbers of genes shared between pathogens which were significantly upregulated or downregulated are shown in Fig. 2a and b, respectively. Although the vast majority of differentially-expressed genes were involved in metabolic processes, many are important in a number of key pathways, including apoptosis, innate immune response and anti-pathogen responses. Selected key genes involved in these pathways are detailed in Fig. 3 and, again, a key observation is the greater number of genes differentially expressed following flavivirus infection, when compared to A. phagocytophilum infection, suggesting that virus infection may have a more profound effect on tick cells than A. phagocytophilum infection.
Table 1.
RNAseq statistics for I. ricinus IRE/CTVM20 cells, uninfected at 0 hpi, or infected with LIV or TBEV at 168 hpi (upper panel), and differential gene expression statistics at 168 hpi following infection with LIV or TBEV at 0 hpi, compared to uninfected cells at 0 hpi (lower panel)
| Parameter | Uninfected 0 hpi (replicate 1/replicate 2) |
LIV 168 hpi (replicate 1/replicate 2) |
TBEV 168 hpi (replicate 1/replicate 2) |
|---|---|---|---|
| Total reads | 11,976,631/10,334,781 | 16,075,150/13,693,034 | 13,339,640/18,910,604 |
| Aligned reads | 11,976,631/10,334,781 | 16,075,150/13,693,034 | 13,339,640/18,910,604 |
| PF_reads | 11,976,631/10,334,781 | 16,075,150/13,693,034 | 13,339,640/18,910,604 |
| PF_aligned reads | 11,976,631/10,334,781 | 16,075,150/13,693,034 | 13,339,640/18,910,604 |
| PF_HQ_aligned reads | 3,878,432/3,760,670 | 4,956,000/3,983,347 | 4,775,958/6,552,021 |
| Mean read length | 119/119 | 119/119 | 119/119 |
| PF_HQ_error rate | 0.76256/0.76193 | 0.763221/0.764392 | 0.766099/0.765034 |
| PF_indel rate | 0.000667/0.000668 | 0.0007/0.000667 | 0.000629/0.000636 |
| Total genes annotated | na | 20,861 | 20,861 |
| Total DE genes | na | 613 (2.96%) | 409 (1.98%) |
| Upregulated DE genes | na | 402 (65.6%) | 252 (61.6%) |
| Downregulated DE genes | na | 211 (34.4%) | 157 (38.4%) |
Abbreviations: na not available, HPI hours post-infection, PF pass-Illumina filter, HQ high quality, PF_HQ_ERROR_RATE percentage of I. ricinus sequence bases that mismatch the I. scapularis reference genome sequence in PF HQ aligned reads, PF_INDEL_RATE number of insertion and deletion events per 100 PF aligned bases. This uses the number of events as the numerator, not the number of inserted or deleted bases. Picard metrics definitions (https://broadinstitute.github.io/picard/picard-metric-definitions.html) were used, DE differentially expressed
Fig. 2.
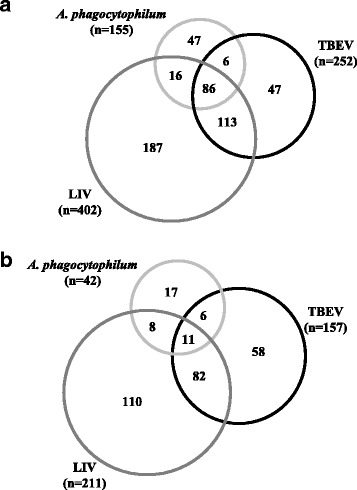
Distribution of differentially upregulated (a) and downregulated (b) genes, following infection of IRE/CTVM20 cells with A. phagocytophilum, LIV or TBEV
Fig. 3.
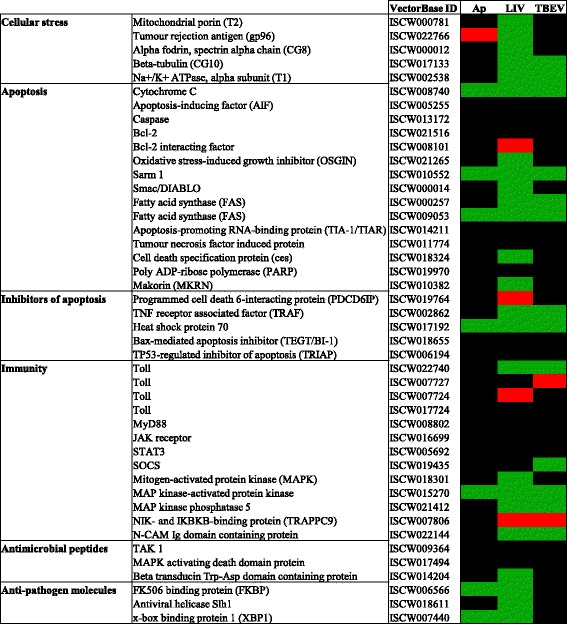
Summary of differential gene expression for selected genes, highlighted green (upregulated), red (downregulated) or black (no significant change), following infection of IRE/CTVM20 cells with A. phagocytophilum (Ap), LIV or TBEV
Upregulation of key regulators of apoptosis following infection with A. phagocytophilum, LIV or TBEV
By 168 hpi for all three pathogens, genes associated with cellular stress and apoptosis were shown to be differentially expressed (P < 0.05 and q < 0.05) when compared to uninfected cells at 0 hpi (Fig. 3). Significant upregulation of the gene encoding cytochrome c following infection with all three pathogens was observed. Crucially however, there was no detection of Caspase gene upregulation (Fig. 3) following infection with any of the pathogens. This suggests that not all components of the apoptotic pathway were transcriptionally induced, although this observation may be due to lack of sequence coverage for some genes. Significant upregulation of the gene encoding heat shock protein 70 (Hsp70) was induced by all three pathogens, and may suggest potential involvement in inhibition of apoptosis (Fig. 3) [31].
Host cell survival supported by expression of innate immune activation and anti-pathogen genes
Viral infection is known to activate the Toll and Jak-STAT pathways [32]. There was evidence for activation of an immune response in I. ricinus tick cells via the Jak-STAT pathway following infection with all three pathogens, with a number of innate immunity genes differentially expressed at 168 hpi (Fig. 3, Table 2). Only A. phagocytophilum infection led to an increase in expression of the Jak receptor gene as previously described [15], although this was not significantly differentially expressed (P < 0.05, but q > 0.05) (Table 2), and there was also no observed differential expression of the gene encoding STAT3. However, the Jak-STAT modulatory suppressor of cytokine signalling (Socs) gene was upregulated significantly following TBEV infection, with some evidence for upregulation following LIV infection (P = 0.009, q = 0.055). There was also evidence for activation of the mitogen-activated protein kinase (MAPK) pathway; although only LIV infection caused upregulation of the gene encoding MAPK, all three pathogens induced significant upregulation of genes for MAPK-activated protein kinase and MAPK phosphatase 5 (Fig. 3, Table 2).
Table 2.
RNAseq results for I. ricinus IRE/CTVM20 cells infected with A. phagocytophilum (Ap), LIV or TBEV at 168 hpi for selected genes associated with innate immunity and anti-pathogen response. Statistical significance denoted by P- or q-values highlighted in bold (< 0.05); genes are considered differentially expressed when both P and q < 0.05
| Gene | VectorBase ID | Pathogen | Log2 fold-change (168 hpi) |
Statistical analysis (P-value/q-value) |
|---|---|---|---|---|
| Toll | ISCW022740 | Ap | 0.34 | 0.261/0.587 |
| LIV | 2.08 | 0.00005/0.001 | ||
| TBEV | 1.64 | 0.00005/0.001 | ||
| Toll | ISCW007727 | Ap | -1.96 | 0.085/0.358 |
| LIV | -3.85 | 0.0078/0.051 | ||
| TBEV | -2.67 | 0.0017/0.022 | ||
| Toll | ISCW007724 | Ap | -0.12 | 0.729/0.888 |
| LIV | -1.26 | 0.0008/0.009 | ||
| TBEV | -0.77 | 0.018/0.116 | ||
| Toll | ISCW017724 | Ap | 0.23 | 0.607/0.827 |
| LIV | 0.21 | 0.690/0.815 | ||
| TBEV | -0.08 | 0.886/0.938 | ||
| MyD88 | ISCW008802 | Ap | 0.44 | 0.323/0.632 |
| LIV | -0.36 | No test | ||
| TBEV | -0.02 | No test | ||
| JAK receptor | ISCW016699 | Ap | 0.85 | 0.035/0.232 |
| LIV | 0.14 | No test | ||
| TBEV | 0.30 | No test | ||
| STAT 3 | ISCW005692 | Ap | 0.23 | 0.451/0.731 |
| LIV | 0.43 | 0.141/0.328 | ||
| TBEV | 0.09 | 0.770/0.874 | ||
| SOCS | ISCW019435 | Ap | 0.38 | 0.356/0.654 |
| LIV | 1.10 | 0.009/0.055 | ||
| TBEV | 1.21 | 0.005/0.046 | ||
| Beta transducing Trp-Asp domain-containing protein | ISCW014204 | Ap | -0.26 | 0.507/0.768 |
| LIV | 1.21 | 0.003/0.024 | ||
| TBEV | 0.39 | No test | ||
| MAPK | ISCW018301 | Ap | 0.21 | 0.557/0.795 |
| LIV | 1.39 | 0.0006/0.007 | ||
| TBEV | 0.28 | 0.486/0.680 | ||
| MAPK-activated protein kinase | ISCW015270 | Ap | 1.93 | 0.00005/0.003 |
| LIV | 2.16 | 0.00005/0.001 | ||
| TBEV | 2.10 | 0.00005/0.001 | ||
| MAPK phosphatase 5 | ISCW021412 | Ap | 1.22 | 0.004/0.077 |
| LIV | 2.29 | 0.00005/0.001 | ||
| TBEV | 1.35 | 0.002/0.026 | ||
| FKBP | ISCW006566 | Ap | 1.10 | 0.0002/0.007 |
| LIV | 1.20 | 0.0001/0.002 | ||
| TBEV | 0.62 | 0.032/0.163 | ||
| Slh1 | ISCW018611 | Ap | -0.13 | 0.669/0.862 |
| LIV | 0.60 | 0.002/0.018 | ||
| TBEV | 0.24 | 0.270/0.503 | ||
| XBP1 | ISCW007440 | Ap | 1.39 | 0.00005/0.003 |
| LIV | 0.67 | 0.0007/0.008 | ||
| TBEV | 0.53 | 0.014/0.097 |
Only flavivirus infection led to significant upregulation of gene expression for toll gene ISCW022740 at 168 hpi (Fig. 3, Table 2). Toll genes ISCW007727 and ISCW7724 were significantly downregulated at 168 hpi following TBEV and LIV infection respectively (Fig. 3, Table 2). There was no change in gene expression for the downstream MyD88 gene at 168 hpi, although LIV infection appeared to upregulate the gene encoding an antimicrobial peptide (ISCW014204, Beta transducin Trp-Asp domain containing protein) (Fig. 3, Table 2). To further analyse toll gene activation as an initiator of innate immune response, selected genes involved in the Toll pathway were analysed, where RNA from the remaining intermediate time-points at 72 and 120 hpi were assessed using specific primers to determine transcript fold-change (normalised against 16S rRNA). Quantitative RT-PCR for toll gene ISCW022740 demonstrated that this transcript was significantly upregulated at 72 hpi following LIV infection (2.47-fold; SEM = 0.55; P = 0.047), with evidence for upregulation at 120 hpi following LIV infection (3.74-fold; SEM = 1.16; P = 0.118) and TBEV infection (2.38-fold; SEM = 0.59; P = 0.063), when compared to uninfected control cells at 0 hpi (Fig. 4a). In contrast, the remaining toll genes were all significantly downregulated at 72 and 120 hpi following flavivirus infection, in accordance with the RNAseq results at 168 hpi. There was no significant increase in expression of any of the toll genes following A. phagocytophilum infection, although toll gene ISCW007727 was significantly downregulated at 72 and 120 hpi (P = 0.022 and 0.025, respectively). Despite the upregulation of toll gene expression induced by flavivirus infection, the gene encoding down-stream MyD88, which is activated via toll engagement, was significantly downregulated at 72 and 120 hpi following infection with all three pathogens (Fig. 4b, Table 2). Therefore, although there was evidence for antimicrobial peptide gene activation, the selective or limited expression of genes within the Toll pathway suggests that Jak-STAT signalling and activation of the MAP kinase cascade dominate the immune response in this cell line.
Fig. 4.
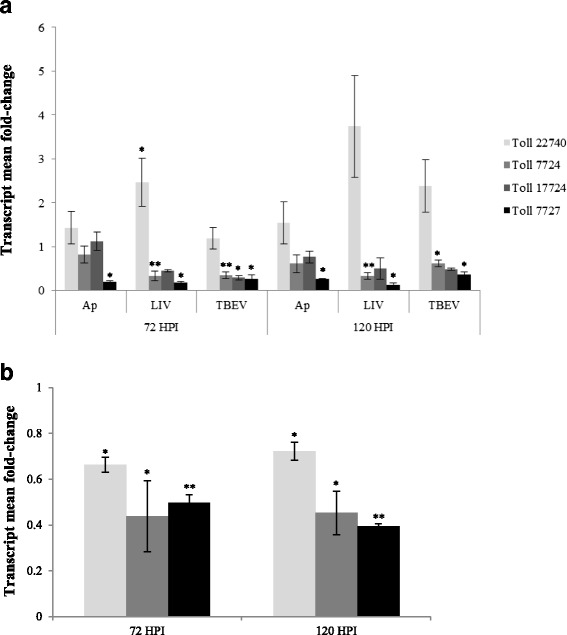
Pathogen-specific effect on biological processes in IRE/CTVM20 cells following infection with tick-borne pathogens at 72 and 120 hpi, as demonstrated by differential expression of (a) toll genes ISCW022740, ISCW007724, ISCW017724 and ISCW007727, and (b) MyD88 gene ISCW008802, where A. phagocytophilum, LIV and TBEV are represented by light grey bars, dark grey bars and black bars respectively. Statistical significance denoted by *P < 0.05; **P < 0.01
In addition to immune genes, a number of genes associated with anti-pathogen responses were differentially expressed (Fig. 3, Table 2). Anaplasma phagocytophilum and LIV induced differential expression of genes encoding FK506 binding protein (FKBP) and X-box binding protein 1 (XBP1) (Fig. 3, Table 2); both of these genes are important for immune gene expression, and were significantly upregulated. Additionally, the gene for antiviral helicase Slh1 was differentially expressed following LIV infection only, with significant upregulation suggestive of a direct antiviral response.
Discussion
The I. scapularis genome is the only tick genome fully sequenced and annotated [33, 34], and constitutes a valuable tool for the investigation of interactions between tick-borne pathogens and Ixodes spp. ticks [35]. Transcriptional analysis of the I. scapularis genome has suggested the involvement of a number of host pathways involved in anti-pathogen responses, including Toll, Jak-STAT and the immune deficiency (Imd) pathway [36–38]. A systems biology approach has previously been utilised to study A. phagocytophilum infection in I. scapularis ticks, where infection was shown to inhibit apoptosis and upregulate the Jak-STAT pathway, promoting survival and enabling infection to become established [14, 38]. Although the response in tick cells to A. phagocytophilum has been well characterised, the response to tick-borne flaviviruses is not yet fully understood. The majority of genes differentially expressed within I. ricinus cells infected with LIV and TBEV were associated with metabolic processes (data not shown), an observation shared with previous proteomics studies analysing the infection of I. scapularis cells with the related tick-borne flavivirus Langat virus (LGTV) [19]. Similarly, TBEV infection in I. ricinus and I. scapularis cells has been shown to induce changes in expression of genes associated with a number of processes, including metabolism and immunity [18]. In Ixodes spp. tick cells infected with A. phagocytophilum, the upregulation of genes associated with apoptosis and cellular stress, including cytochrome c, has been previously reported [15], and flavivirus infection appears to activate some of the same genes that could be associated with apoptosis. However, the lack of Caspase gene activation suggests selective gene activation, which correlates with microscopic observations of tick cells during the 7-day course of infection with LIV, TBEV (this study), or A. phagocytophilum [15], where cell morphology remained unaltered. As vectors of these pathogens, I. ricinus ticks therefore have mechanisms for restricting the cellular damage caused by infection, including apoptosis. Hsp70 has previously been shown to inhibit apoptosis through a number of pathways [39–41], suggesting that upregulation of the hsp70 gene may be involved in the inhibition of apoptosis as a mechanism for host survival. Indeed, knockdown of hsp70 family genes has been shown to increase LGTV replication in another I. ricinus cell line, suggesting a potential antiviral role for these proteins [18]. All three pathogens induced a significant increase in expression of the hsp70 gene by 168 hpi, and the upregulation of Hsp70 at the protein level has previously been observed in A. phagocytophilum-infected tick cells [42]. However, although these data may suggest inhibition of apoptosis, it is possible that the differential expression of these genes could be associated with other cellular responses, such as Hsp70 involvement in protein folding and cellular stress [43].
Innate immune activation is essential for survival following infection, and the infection of tick cells with A. phagocytophilum has previously been shown to upregulate the Jak-STAT pathway, with a role in inhibition of apoptosis [14, 15], or the restriction of infection through regulation of antimicrobial peptide expression [37]. Although we could find no evidence of increased expression of Jak or STAT genes following flavivirus infection (Fig. 3), possibly due to limitations in sequence coverage, the differential expression of other genes within the pathway, including Socs genes, suggests that selective gene regulation following infection occurs. SOCS inhibit the Jak-STAT pathway, leading to reduced transcription of Jak-STAT genes, and the upregulation of Socs gene transcripts has previously been demonstrated in mouse brain following flavivirus infection [44]. Additionally, all three pathogens also induced significant increases in expression of genes within the MAPK cascade. Differential expression of genes involved in both the Jak-STAT and MAPK signalling pathways has also been observed in an Aedes albopictus mosquito cell line, following infection with another arthropod-borne virus, bluetongue virus (BTV) [45].
Innate immunity in ticks and tick cells has still not been fully elucidated, but evidence suggests that the Toll signalling pathway is also present in ticks [36]. The lack of detection of toll gene activation following A. phagocytophilum infection is unsurprising, since gram-negative bacteria are more likely to initiate the alternative Imd pathway [46]. However, the upregulation of toll gene ISCW022740 following flavivirus infection suggests that the toll pathway is activated during flavivirus infection in tick cells and ticks. This constitutes a host survival mechanism, since the toll pathway has been shown to be important for restricting infection with dengue virus (DENV) in Aedes aegypti mosquito cells [47]. However, the downregulation of other toll genes assessed suggests some degree of toll receptor specificity following flavivirus infection, and the downregulation of MyD88 gene expression following infection with all three pathogens suggests limitations to, or potential inhibition of, toll effector functions. Although not beneficial for host survival, this may enhance infection and benefit tick transmission of pathogens, since MyD88 has been shown to restrict the flaviviruses West Nile virus (WNV) and Japanese encephalitis virus (JEV), in mammalian cells [48, 49]. Furthermore, an absence of MyD88 in mice has been shown to enhance the infectivity of a strain of the spirochaete Borrelia burgdorferi when transmitted to I. scapularis ticks during feeding [50], suggesting that a lack of expression of this gene enabled an enhancement of infection within tick cells.
Immune activation leads to down-stream upregulation of a number of genes which may be considered to have anti-pathogen functionality. These include the xbp1 gene, which was upregulated following infection with all three pathogens (although only differentially expressed following A. phagocytophilum or LIV infection), consistent with the activation of an innate immune response. The transcription factor XBP1 has multiple functions, including a role in the production of pro-inflammatory cytokines in macrophages, and has been shown to be induced via Toll-like receptor (TLR) activation [51]. It has also been shown to contribute to host protection against immune activation in bacterially-infected Caenorhabditis elegans nematodes [52]. A similar trend was demonstrated for fkbp gene expression, which was also upregulated following infection with all three pathogens. FKBP51, a protein from the FKBP family of proteins, has been shown to interact with TNF receptor associated factor (TRAF) proteins in mammalian cells to facilitate the expression of type I interferon following viral infection [53]. Infection with Sindbis virus (SINV), an alphavirus, has also been shown to alter gene expression for FKBP1 in the midgut of Ae. aegypti mosquitoes [54], whilst FKBP1 paralogs have been shown to suppress infection with DENV and WNV in human HeLa cells [55]. FKBPs can therefore be considered to constitute host resistance factors with antiviral potential [56]. Additionally, the antiviral helicase Slh1 gene was significantly upregulated following LIV infection. Slh1 has been associated with antiviral defence in yeast (Saccharomyces cerevisiae), where it has been shown to inhibit translation of viral mRNAs [57]. This provides some evidence that a mechanism may exist within tick cells for limiting flavivirus replication independently of apoptosis, and in addition to immune gene activation.
Host cells respond by increasing apoptosis and activating immune genes for survival. Indeed, A. phagocytophilum infection of ISE6 tick cells has been shown to induce other pathways that increase host cell survival, whilst enabling pathogen transmission through limitations to the tick cell response via protein mis-folding, suggestive of co-evolution between pathogen and host [42, 58]. There is evidence that selective gene expression in response to bacteria or viruses occurs within tick cells, implying that different cellular recognition proteins are allied to alternative pathways for transcript activation. Viral-induced dsRNA is also known to be recognised in mosquito cells by Dicer-2, which is similar in functionality to mammalian retinoic acid-inducible gene-I (RIG-I), and also activates the Jak-STAT pathway [59]. There may also be activation of alternative immune pathways including RNAi responses in tick cells [17]. However, the response observed in tick cells may vary compared to the response in ticks, although previous studies in tick cell lines have demonstrated that the transcriptional response to A. phagocytophilum infection in I. scapularis ISE6 cells was similar to that observed in tick haemocytes, whilst the response in I. ricinus IRE/CTVM20 cells was more similar to that observed in the tick midgut [15]. Additionally, although these experiments were undertaken at a single temperature of 28 °C, previous studies have shown that variation in temperature can affect tick physiology and tick-pathogen interactions [60]. Therefore, variation in temperature may influence gene expression in tick cell lines, and may lead to alternative gene expression profiles.
Conclusions
The host response to flavivirus infection in tick cells is complex, and involves the interaction of a number of host mechanisms in order to promote cellular survival, whilst ensuring that virus replication can occur. Infection of I. ricinus cells with the tick-borne flaviviruses LIV and TBEV induced an increase in the expression of genes associated with apoptosis, along with the activation of immune genes, including toll, for survival. However, infection with these pathogens was also associated with an increase in gene expression for proteins associated with inhibition of apoptosis, which is promoted by pathogens to increase infection. Therefore, multiple pathways exist within tick cells to provide efficient mechanisms for pathogen control and host survival, and the identification of key genes may contribute to the identification of potential targets for future antiviral strategies. However, differential gene expression in virus-infected cells does not necessarily lead to significant changes in levels of the related proteins, therefore although not in scope for this study, future investigation will include proteomics analysis in order to enhance and validate these transcriptional studies.
Acknowledgements
Not applicable.
Funding
This work was jointly funded by the European Commission Seventh Framework Programme under ANTIGONE (project 278976), the Department for Environment, Food and Rural Affairs (Defra), Scottish Government and Welsh Government under project SE4112, EUH2020-funded Research Infrastructure Grant 'European Virus Archive Global (EVAg)' (H2020 grant agreement number 653316) and the INIA grant E-RTA2013-C04-04 (FEDER co-funded, Spain). Lesley Bell-Sakyi is supported by the United Kingdom Biotechnology and Biological Sciences Research Council’s Institute Strategic Programme Grant (BBS/E/I/0000174) to the Pirbright Institute.
Availability of data and materials
RNAseq data supporting these findings are deposited in NCBI’s Gene Expression Omnibus, and are available through GEO Series accession number (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE85300) (LIV and TBEV-infected tick cells). Previously published RNAseq data derived from A. phagocytophilum-infected tick cells [15], are available through GEO Series accession number (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE76906).
Authors’ contributions
ARF, NJ and JDF provided funding for this study; LBS provided the tick cell line; KLM, PA and RJE undertook the experimental work; CC, RJE and KLM performed the bioinformatics; KLM, CC, NJ and JDF analysed the results; KLM prepared the draft manuscript and all authors contributed to the final version. All authors read and approved the final manuscript.
Competing interests
The authors declare that they have no competing interests.
Consent for publication
Not applicable.
Ethics approval and consent to participate
Not applicable.
Abbreviations
- Ap
Anaplasma phagocytophilum
- BTV
Bluetongue virus
- cDNA
Complementary DNA
- DENV
Dengue virus
- DTT
Dithiothreitol
- FKBP
FK506 binding protein
- HGA
Human granulocytic anaplasmosis
- hpi
Hours post-infection
- HSP70
Heat shock protein 70
- Imd
Immune deficiency
- Jak-STAT
Janus kinase-signal transducer and activator of transcription
- JEV
Japanese encephalitis virus
- LGTV
Langat virus
- LIV
Louping ill virus
- MAPK
Mitogen-activated protein kinase
- M-MLV RT
Maloney murine leukemia virus reverse transcriptase
- MOI
Multiplicity of infection
- MSP4
Major surface protein 4
- MyD88
Myeloid differentiation factor 88
- NGS
next generation sequencing
- PFU
Plaque forming units
- RIG-I
retinoic acid-inducible gene-I
- rRNA
Ribosomal RNA
- SEM
Standard error of the mean
- SINV
Sindbis virus
- Slh1
Antiviral helicase
- SOCS
Suppressor of cytokine signalling
- TBE
Tick-borne encephalitis
- TBEV
Tick-borne encephalitis virus
- TLR
Toll-like receptor
- TPB
Tryptose phosphate broth
- TRAF
TNF receptor associated factor
- UK
United Kingdom
- WNV
West nile virus
- XBP1
X-box binding protein 1
Additional files
Primers used in this study for detection of viral RNA (LIV/TBEV) from infected I. ricinus IRE/CTVM20 cells, along with host gene transcripts in RNA extracted from I. ricinus IRE/CTVM20 cells infected with A. phagocytophilum, LIV or TBEV. (DOC 38 kb)
Build statistics: assessment of sequence depth and quality of alignment for samples taken from I. ricinus IRE/CTVM20 cells, uninfected at 0 hpi (samples 1–4), or infected with A. phagocytophilum (samples 15–16), LIV (samples 17–18) or TBEV (samples 19–20) at 168 hpi. (DOC 36 kb)
Contributor Information
Karen L. Mansfield, Email: Karen.Mansfield@apha.gsi.gov.uk
Charlotte Cook, Email: Charlotte.Cook@apha.gsi.gov.uk.
Richard J. Ellis, Email: Richard.Ellis@apha.gsi.gov.uk
Lesley Bell-Sakyi, Email: lesley.sakyi@pirbright.ac.uk.
Nicholas Johnson, Email: Nick.Johnson@apha.gsi.gov.uk.
Pilar Alberdi, Email: Maria.Alberdi@uclm.es.
José de la Fuente, Email: jose_delafuente@yahoo.com.
Anthony R. Fooks, Email: Tony.Fooks@apha.gsi.gov.uk
References
- 1.Ostfield RS, Brunner JL. Climate change and Ixodes tick-borne diseases of humans. Philos Trans R Soc Lond B Biol Sci. 2015;370:1665. doi: 10.1098/rstb.2014.0051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bacon RM, Kugeler KJ, Mead PS. Centers for Disease Control and Prevention (CDC). Surveillance for Lyme disease - United States, 1992–2006. MMWR Surveill Summ. 2008;57:1–9. [PubMed] [Google Scholar]
- 3.Leger E, Vourc'h G, Vial L, Chevillon C, McCoy KD. Changing distributions of ticks: causes and consequences. Exp Appl Acarol. 2013;59:219–44. doi: 10.1007/s10493-012-9615-0. [DOI] [PubMed] [Google Scholar]
- 4.Medlock JM, Hansford KM, Bormane A, Derdakova M, Estrada-Peña A, George JC, et al. Driving forces for change in geographical distribution of Ixodes ricinus ticks in Europe. Parasit Vectors. 2013;6:11. doi: 10.1186/1756-3305-6-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Jeffries CL, Mansfield KL, Phipps LP, Wakeley PR, Mearns R, Schock A, et al. Louping ill virus: an endemic tick-borne disease of Great Britain. J Gen Virol. 2014;95:1005–14. doi: 10.1099/vir.0.062356-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Mansfield KL, Johnson N, Phipps LP, Stephenson JR, Fooks AR, Solomon T. Tick-borne encephalitis virus - a review of an emerging zoonosis. J Gen Virol. 2009;90:1781–94. doi: 10.1099/vir.0.011437-0. [DOI] [PubMed] [Google Scholar]
- 7.Gritsun TS, Nuttall PA, Gould EA. Tick-borne flaviviruses. Adv Virus Res. 2003;61:317–71. doi: 10.1016/S0065-3527(03)61008-0. [DOI] [PubMed] [Google Scholar]
- 8.de la Fuente J, Estrada-Peña A, Venzal JM, Kocan KM, Sonenshine DE. Overview: ticks as vectors of pathogens that cause disease in humans and animals. Front Biosci. 2008;13:6938–46. doi: 10.2741/3200. [DOI] [PubMed] [Google Scholar]
- 9.Dumler JS, Barbet AF, Bekker CP, Dasch GA, Palmer GH, Ray SC, et al. Reorganization of the genera in the families Rickettsciaceae and Anaplasmataceae in the order Rickettsiales: unification of some species of Ehrlichia with Anaplasma, Cowdria with Ehrlichia and Ehrlichia with Neorickettsia, descriptions of six new species combinations and designation of Ehrlichia equi and ‘HGE agent’ as subjective synonyms of Ehrlichia phagocytophila. Int J Syst Evol Microbiol. 2001;51:2145–65. doi: 10.1099/00207713-51-6-2145. [DOI] [PubMed] [Google Scholar]
- 10.Stuen S. Anaplasma phagocytophilum - the most widespread tick-borne infection in animals in Europe. Vet Res Commun. 2010;31:79–84. doi: 10.1007/s11259-007-0071-y. [DOI] [PubMed] [Google Scholar]
- 11.Stuen S, Granquist EG, Silaghi C. Anaplasma phagocytophilum - a widespread multi-host pathogen with highly adaptive strategies. Front Cell Infect Microbiol. 2013;3:31. doi: 10.3389/fcimb.2013.00031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Everett H, McFadden G. Apoptosis: an innate immune response to virus infection. Trends Microbiol. 1999;7:160–5. doi: 10.1016/S0966-842X(99)01487-0. [DOI] [PubMed] [Google Scholar]
- 13.Ayllón N, Villar M, Busby AT, Kocan KM, Blouin EF, Bonzón-Kulichenko E, et al. Anaplasma phagocytophilum inhibits apoptosis and promotes cytoskeleton rearrangement for infection of tick cells. Infect Immun. 2013;81:2415–25. doi: 10.1128/IAI.00194-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ayllón N, Villar M, Galindo RC, Kocan KM, Šíma R, López JA, et al. Systems biology of tissue-specific response to Anaplasma phagocytophilum reveals differentiated apoptosis in the tick vector Ixodes scapularis. PLoS Genet. 2015;11:e1005120. doi: 10.1371/journal.pgen.1005120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Alberdi P, Mansfield KL, Manzano-Román R, Cook C, Ayllón N, Villar M, et al. Tissue-specific signatures in the transcriptional response to Anaplasma phagocytophilum infection of Ixodes scapularis and Ixodes ricinus tick cell lines. Front Cell Infect Microbiol. 2016;6:20. doi: 10.3389/fcimb.2016.00020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.De Gregorio E, Spellman PT, Tzou P, Rubin GM, Lemaitre B. The Toll and Imd pathways are the major regulators of the immune response in Drosophila. Embo J. 2002;21:2568–79. doi: 10.1093/emboj/21.11.2568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Schnettler E, Tykalová H, Watson M, Sharma M, Sterken MG, Obbard DJ, et al. Induction and suppression of tick cell antiviral RNAi responses by tick-borne flaviviruses. Nucleic Acids Res. 2014;42:9436–46. doi: 10.1093/nar/gku657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Weisheit S, Villar M, Tykalová H, Popara M, Loecherbach J, Watson M, et al. Ixodes scapularis and Ixodes ricinus tick cell lines respond to infection with tick-borne encephalitis virus: transcriptomic and proteomic analysis. Parasit Vectors. 2015;8:599. doi: 10.1186/s13071-015-1210-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Grabowski JM, Perera R, Roumani AM, Hedrick VE, Inerowicz HD, Hill CA, et al. Changes in the proteome of Langat-infected Ixodes scapularis ISE6 cells; metabolic pathways associated with flavivirus infection. PLoS Negl Trop Dis. 2016;10:e0004180. doi: 10.1371/journal.pntd.0004180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Aguero-Rosenfeld ME, Horowitz HW, Wormser GP, McKenna DF, Nowakowski J, Muñoz J, et al. Human granulocytic ehrlichiosis: A case series from a medical center in New York State. Ann Intern Med. 1996;125:904–8. doi: 10.7326/0003-4819-125-11-199612010-00006. [DOI] [PubMed] [Google Scholar]
- 21.Asanovich KM, Bakken JS, Madigan JE, Aguero-Rosenfeld M, Wormser GP, Dumler JS. Antigenic diversity of granulocytic Ehrlichia isolates from humans in Wisconsin and New York and a horse in California. J Infect Dis. 1997;176:1029–34. doi: 10.1086/516529. [DOI] [PubMed] [Google Scholar]
- 22.Bell-Sakyi L, Zweygarth E, Blouin EF, Gould EA, Jongejan F. Tick cell lines: tools for tick and tick-borne disease research. Trends Parasitol. 2007;23:450–7. doi: 10.1016/j.pt.2007.07.009. [DOI] [PubMed] [Google Scholar]
- 23.Munderloh UG, Kurtti TJ. Formulation of medium for tick cell culture. Exp Appl Acarol. 1989;7:219–29. doi: 10.1007/BF01194061. [DOI] [PubMed] [Google Scholar]
- 24.Dyachenko V, Geiger C, Pantchev N, Majzoub M, Bell-Sakyi L, Krupka I, et al. Isolation of canine Anaplasma phagocytophilum strains from clinical blood samples using the Ixodes ricinus cell line IRE/CTVM20. Vet Microbiol. 2013;162:980–6. doi: 10.1016/j.vetmic.2012.10.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ruzek D, Bell-Sakyi L, Kopecky K, Grubhoffer L. Growth of tick-borne encephalitis virus (European subtype) in cell lines from vector and non-vector ticks. Virus Res. 2008;137:142–6. doi: 10.1016/j.virusres.2008.05.013. [DOI] [PubMed] [Google Scholar]
- 26.Mazelier M, Rouxel RN, Zumstein M, Mancini R, Bell-Sakyi L, Lozach P-Y. Uukuniemi virus as a tick-borne virus model. J Virol. 2016;90:6784–98. doi: 10.1128/JVI.00095-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Mansfield KL, Horton DL, Johnson N, Li L, Barrett AD, Smith DJ, et al. Flavivirus-induced antibody cross-reactivity. J Gen Virol. 2011;92:2821–9. doi: 10.1099/vir.0.031641-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.García-Alcalde F, Okonechnikov K, Carbonell J, Cruz LM, Götz S, Tarazona S, et al. Qualimap: evaluating next-generation sequencing alignment data. Bioinformatics. 2012;28:2678–9. doi: 10.1093/bioinformatics/bts503. [DOI] [PubMed] [Google Scholar]
- 29.Okonechnikov K, Conesa A, García-Alcalde F. Qualimap 2: advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics. 2016;32:292–4. doi: 10.1093/bioinformatics/btv566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.McKimmie C, Johnson N, Fooks AR, Fazakerley JK. Viruses selectively upregulate Toll-like receptors in the central nervous system. Biochem Biophys Res Commun. 2005;336:925–33. doi: 10.1016/j.bbrc.2005.08.209. [DOI] [PubMed] [Google Scholar]
- 31.Beere HM, Wol BB, Cain K, Mosser DD, Mahboubi A, Kuwana T, et al. Heat-shock protein 70 inhibits apoptosis by preventing recruitment of procaspase-9 to the Apaf-1 apoptosome. Nat Cell Biol. 2000;2:469–75. doi: 10.1038/35019501. [DOI] [PubMed] [Google Scholar]
- 32.Smith AA, Pal U. Immunity-related genes in Ixodes scapularis - perspectives from genome information. Front Cell Infect Microbiol. 2014;4:116. doi: 10.3389/fcimb.2014.00116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Geraci NS, Spencer Johnston J, Paul Robinson J, Wikel SK, Hill CA. Variation in genome size of argasid and ixodid ticks. Insect Biochem Mol Biol. 2007;37:399–408. doi: 10.1016/j.ibmb.2006.12.007. [DOI] [PubMed] [Google Scholar]
- 34.Gulia-Nuss M, Nuss AB, Meyer JM, Sonenshine DE, Roe RM, Waterhouse RM, et al. Genomic insights into the parasitic vector of Lyme disease. Ixodes scapularis Nat Comm. 2016;7:10507. doi: 10.1038/ncomms10507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Genomic Resources Development Consortium. Contreras M, de la Fuente J, Estrada-Peña A, Grubhoffer L, Tobes R. Transcriptome sequence divergence between Lyme disease tick vectors, Ixodes scapularis and Ixodes ricinus. Mol Ecol Resour. 2014;14:1095. doi: 10.1111/1755-0998.12298. [DOI] [PubMed] [Google Scholar]
- 36.Kopáček P, Hajdušek O, Burešová V, Daffre S. Tick innate immunity. In: Söderhäll K, editor. Invertebrate Immunity. Austin: Landes Biosciences and Springer Science + Business Media; 2010. pp. 137–62. [Google Scholar]
- 37.Liu L, Dai J, Zhao YO, Narasimhan S, Yang Y, Zhang L, Fikrig E. Ixodes scapularis JAK-STAT pathway regulates tick antimicrobial peptides, thereby controlling the agent of human granulocytic anaplasmosis. J Infect Dis. 2012;206:1233–41. doi: 10.1093/infdis/jis484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Alberdi P, Ayllón N, Cabezas-Cruz A, Bell-Sakyi L, Zweygarth E, Stuen S, et al. Infection of Ixodes spp. tick cells with different Anaplasma phagocytophilum isolates induces the inhibition of apoptotic cell death. Ticks Tick-borne Dis. 2015;6:758–67. doi: 10.1016/j.ttbdis.2015.07.001. [DOI] [PubMed] [Google Scholar]
- 39.Mosser DD, Caron AW, Bourget L, Denis-Larose C, Massie B. Role of the human heat shock protein hsp70 in protection against stress-induced apoptosis. Mol Cell Biol. 1997;17:5317–27. doi: 10.1128/MCB.17.9.5317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Ravagnan L, Gurbuxani S, Susin SA, Maisse C, Daugas E, Zamzami N, et al. Heat-shock protein 70 antagonizes apoptosis-inducing factor. Nat Cell Biol. 2001;3:839–43. doi: 10.1038/ncb0901-839. [DOI] [PubMed] [Google Scholar]
- 41.Shukla AK, Pragya P, Chaouhan HS, Tiwari AK, Patel DK, Abdin MZ, et al. Heat shock protein-70 (Hsp-70) suppresses paraquat-induced neurodegeneration by inhibiting JNK and caspase-3 activation in Drosophila model of Parkinson’s disease. PLoS One. 2014;9:e98886. doi: 10.1371/journal.pone.0098886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Villar M, Ayllón N, Alberdi P, Moreno A, Moreno M, Tobes R, et al. Integrated metabolics, transcriptomics and proteomics identifies metabolic pathways affected by Anaplasma phagocytophilum infection in tick cells. Mol Cell Proteomics. 2015;14:12. doi: 10.1074/mcp.M115.051938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.King AM, MacRae TH. Insect heat shock proteins during stress and diapause. Annu Rev Entomol. 2015;60:59–75. doi: 10.1146/annurev-ento-011613-162107. [DOI] [PubMed] [Google Scholar]
- 44.Mansfield KL, Johnson N, Cosby SL, Solomon T, Fooks AR. Transcriptional upregulation of SOCS 1 and suppressors of cytokine signalling 3 mRNA in the absence of suppressors of cytokine signalling 2 mRNA after infection with West Nile virus or tick-borne encephalitis virus. Vector Borne Zoonotic Dis. 2010;10:649–53. doi: 10.1089/vbz.2009.0259. [DOI] [PubMed] [Google Scholar]
- 45.Xing S, Du J, Gao S, Tian Z, Zheng Y, Liu G, et al. Analysis of the miRNA expression profile in an Aedes albopictus cell line in response to bluetongue virus infection. Infect Genet Evol. 2016;39:74–84. doi: 10.1016/j.meegid.2016.01.012. [DOI] [PubMed] [Google Scholar]
- 46.Kaneko T, Goldman WE, Mellroth P, Steiner H, Fukase K, Kusumoto S, et al. Monomeric and polymeric gram-negative peptidoglycan but not purified LPS stimulate the Drosophila IMD pathway. Immunity. 2004;20:637–49. doi: 10.1016/S1074-7613(04)00104-9. [DOI] [PubMed] [Google Scholar]
- 47.Xi Z, Ramirez JL, Dimopoulos G. The Aedes aegypti toll pathway controls dengue virus infection. PLoS Path. 2008;4:e1000098. doi: 10.1371/journal.ppat.1000098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Aleyas AG, George JA, Han YW, Rahman MM, Kim SJ, Han SB, et al. Functional modulation of dendritic cells and macrophages by Japanese encephalitis virus through MyD88 adaptor molecule-dependent and independent pathways. J Immunol. 2009;183:2462–74. doi: 10.4049/jimmunol.0801952. [DOI] [PubMed] [Google Scholar]
- 49.Szretter KJ, Daffis S, Patel J, Suthar MS, Klein RS, Gale M, et al. The innate immune adaptor molecule MyD88 restricts West Nile virus replication and spread in neurons of the central nervous system. J Virol. 2010;84:12125–38. doi: 10.1128/JVI.01026-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Bockenstedt LK, Liu N, Schwartz I, Fish D. MyD88 deficiency enhances acquisition and transmission of Borrelia burgdorferi by Ixodes scapularis ticks. Infect Immun. 2006;74:2154–60. doi: 10.1128/IAI.74.4.2154-2160.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Martinon F, Chen X, Lee AH, Glimcher LH. Toll-like receptor activation of XBP1 regulates innate immune responses in macrophages. Nat Immunol. 2010;11:411–8. doi: 10.1038/ni.1857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Richardson CE, Kooistra T, Kim DH. An essential role for XBP-1 in host protection against immune activation in C. elegans. Nature. 2010;463:1092–5. doi: 10.1038/nature08762. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Akiyama T, Shiraishi T, Qin J, Konno H, Akiyama N, Shinzawa M, et al. Mitochondria-nucleus shuttling FK506-binding protein 51 interacts with TRAF proteins and facilitates the RIG-I-like receptor-mediated expression of Type I IFN. PLoS One. 2014;9:e95992. doi: 10.1371/journal.pone.0095992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Sanders HR, Foy BD, Evans AM, Ross LS, Beaty BJ, Olson KE, et al. Sindbis virus induces transport processes and alters expression of innate immunity pathway genes in the midgut of the disease vector. Aedes aegypti. Insect Biochem Mol Biol. 2005;35:1293–307. [DOI] [PubMed]
- 55.Krishnan MN, Ng A, Sukumaran B, Gilfoy FD, Uchil PD, Sultana H, et al. RNA interference screen for human genes associated with West Nile virus infection. Nature. 2008;455:242–5. doi: 10.1038/nature07207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Rider MA, Zou J, Vanlandingham D, Nuckols JT, Higgs S, Zhang Q, et al. Quantitative proteomic analysis of the Anopheles gambiae (Diptera: Culicidae) midgut infected with O’nyong-Nyong virus. J Med Entomol. 2013;50:1077–88. doi: 10.1603/ME12155. [DOI] [PubMed] [Google Scholar]
- 57.Searfoss AM, Wickner RB. 3’ poly(A) is dispensable for translation. PNAS. 2000;97:9133–7. doi: 10.1073/pnas.97.16.9133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.de la Fuente J, Estrada-Peña A, Cabezas-Cruz A, Kocan KM. Anaplasma phagocytophilum uses common strategies for infection of ticks and vertebrate hosts. Trends Microbiol. 2016;24:173–80. doi: 10.1016/j.tim.2015.12.001. [DOI] [PubMed] [Google Scholar]
- 59.Paradkar PN, Trinidad L, Voysey R, Duchemin JB, Walker PJ. Secreted Vago restricts West Nile virus infection in Culex mosquito cells by activating the Jak-STAT pathway. Proc Natl Acad Sci USA. 2012;109:18915–20. [DOI] [PMC free article] [PubMed]
- 60.Obonyo M, Munderloh UG, Fingerle BW, Kurtti TJ. Borrelia burgdorferi in tick cell culture modulates expression of outer surface proteins A and C in response to temperature. J Clin Microbiol. 1999;37:2137–41. doi: 10.1128/jcm.37.7.2137-2141.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
RNAseq data supporting these findings are deposited in NCBI’s Gene Expression Omnibus, and are available through GEO Series accession number (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE85300) (LIV and TBEV-infected tick cells). Previously published RNAseq data derived from A. phagocytophilum-infected tick cells [15], are available through GEO Series accession number (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE76906).


