Abstract
Mitochondria are double-membrane organelles with varying shapes influenced by metabolic conditions, developmental stage, and environmental stimuli1–4. Their dynamic morphology is realized through regulated and balanced fusion and fission processes5, 6. Fusion is crucial for the health and physiological functions of mitochondria, including complementation of damaged mitochondrial DNAs and maintenance of membrane potential6–8. Mitofusins (Mfns) are dynamin-related GTPases essential for mitochondrial fusion9, 10. They are embedded in the mitochondrial outer membrane and thought to fuse adjacent mitochondria via concerted oligomerization and GTP hydrolysis11–13. However, the molecular mechanisms behind this process remains elusive. Here we present crystal structures of engineered human Mfn1 containing the GTPase domain and a helical domain in different stages of GTP hydrolysis. The helical domain is composed of elements from widely dispersed sequence regions of Mfn1 and resembles the Neck of the bacterial dynamin-like protein. The structures reveal unique features of its catalytic machinery and explain how GTP binding induces conformational changes to promote G domain dimerization in the transition state. Disruption of G domain dimerization abolishes the fusogenic activity of Mfn1. Moreover, a conserved aspartate trigger was found in Mfn1 to affect mitochondrial elongation, likely through a GTP-loading-dependent domain rearrangement. Based on these results, we propose a mechanistic model for Mfn1-mediated mitochondrial tethering. Our study provides important insights in the molecular basis of mitochondrial fusion and mitofusin-related human neuromuscular disorders14.
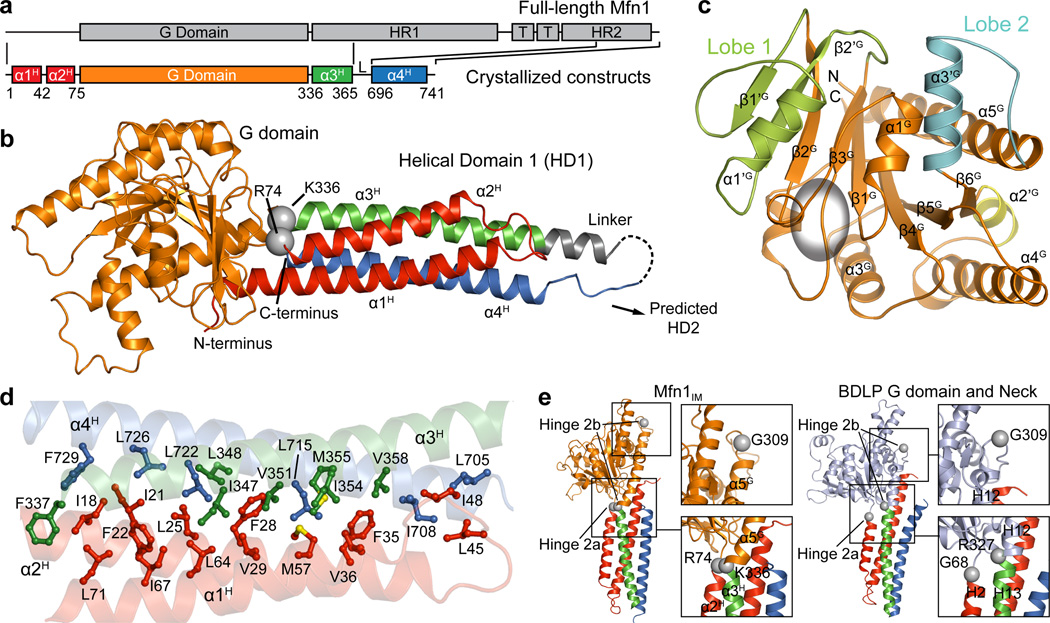
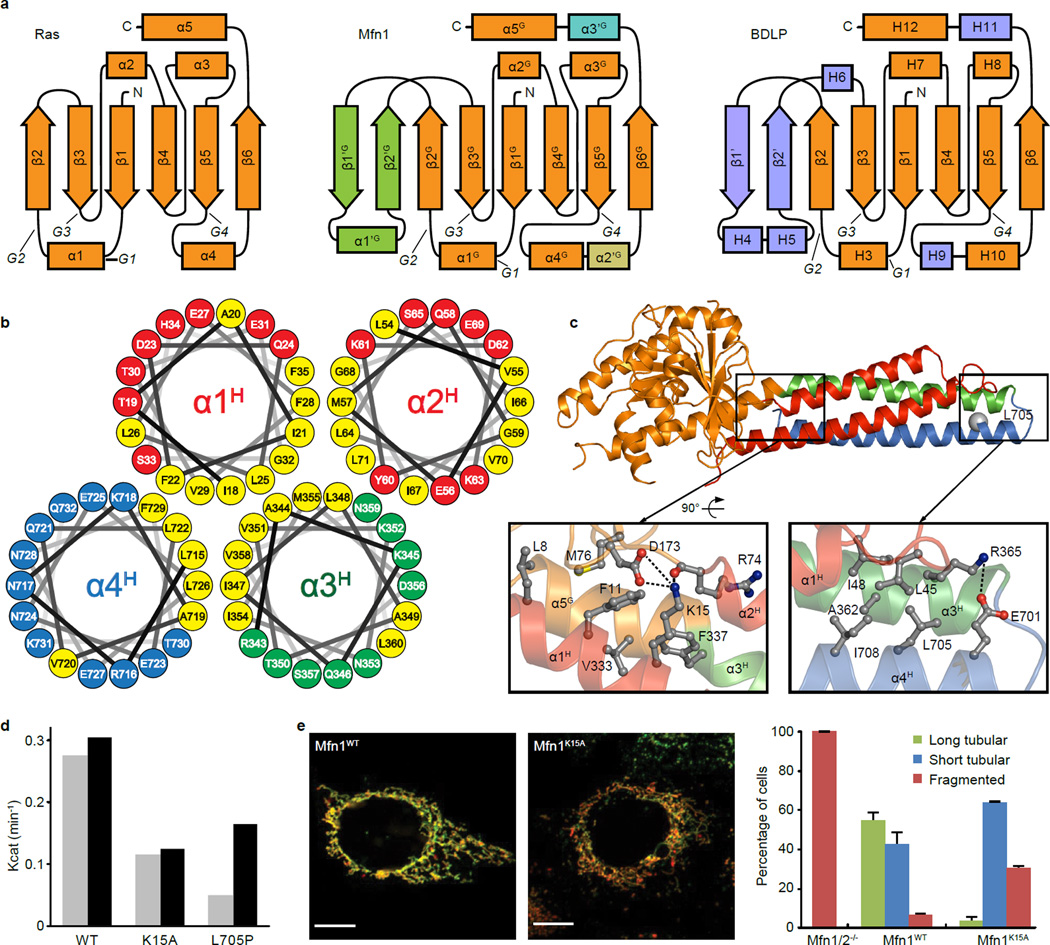
We constructed an internally modified human Mfn1 (Mfn1IM) composed of the GTPase (G) domain (residues 75–336) and a four-helix-bundle that we term helical domain 1 (HD1, Fig. 1a, 1b, Extended Data Fig. 1a–g and Extended Data Table 1). The G domain contains a central eight-strand β-sheet surrounded by eight α-helices. Compared to the canonical GTPase Ras, the G domain of Mfn1 has two extra lobes that shield the nucleotide binding pocket, and a specific short α-helix (α2’G) sitting between α4G and β6G (Fig. 1c). Lobe 1, containing two β-strands (β1’G and β2’G) and an α-helix (α1’G), is located between β2G and β3G, whereas Lobe 2, consisting of an α-helix (α3’G) and loop, is located between β6G and α5G (Extended Data Fig. 2a). The four α-helices of the HD1, derived from widely dispersed sequence regions, form a vast and conserved hydrophobic network (Fig. 1a, d and Extended Data Fig. 2b). HD1 is connected to the G domain via R74 at the C-terminal end of α2H and K336 between α5G and α3H (Fig. 1b). The N terminus of α1H substantially contacts the G domain (Extended Data Fig. 2c–e). On the other side of HD1, part of the artificial linker folds into an α-helix extending α3H (Fig. 1b). This is in agreement with the secondary structure prediction for the replaced residues (Extended Data Fig. 3), suggesting that α3H may be longer in full-length Mfn1.
Figure 1. Overall structure of Mfn1IM.
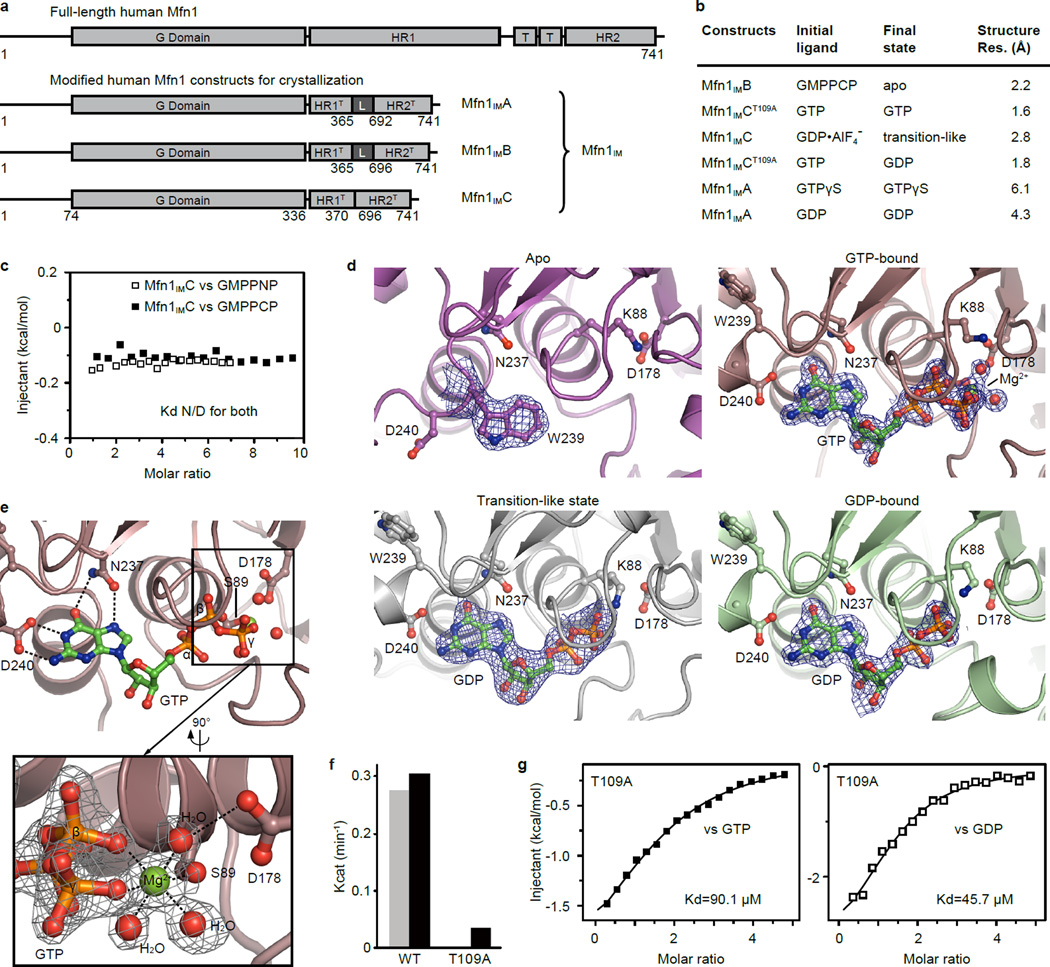
a, Schematic representation showing the organization of Mfn1IM based on full-length Mfn1 with the conventional terminology: G domain, GTPase domain; HR1, heptad repeat region 1; T, transmembrane region; HR2, heptad repeat region 2. Elements for Mfn1IM are assigned according to the structure. L stands for the artificial linker. Borders of each element are indicated by residue numbers.
b, Structure of Mfn1IM. α-helices of HD1 are differentially coloured to specify their distribution on the primary structure as in a. The artificial linker is in grey. Disordered loop is shown as dashed lines. The Cα atoms of R74 and K336 linking G domain and HD1 are shown as grey spheres.
c, The G domain of Mfn1IM. Lobe 1, Lobe 2 and α2’G are colour-specified. The core region corresponding to Ras is coloured orange and the GTPase active site is indicated by an ellipsoid.
d, Hydrophobic network within HD1. Side chains of the residues involved in the network are shown in the same colour as the helices they belong to.
e, Comparison between the putative Hinge 2 of Mfn1IM and BDLP.
The overall topology of Mfn1IM is typical of the dynamin superfamily15–20 (Extended Data Fig. 4). Apart from the G domain, the Mfn1IM HD1 is particularly consistent with the Neck of the bacterial dynamin-like protein from Nostoc punctiforme (BDLP), which was suggested to mediate membrane fusion in bacteria21 (Extended Data Fig. 2a, 4, 5a). Given the compact organization of HD1 and the predicted secondary structure (Extended Data Fig. 3), the missing portion of Mfn1 (excluding TM) from Mfn1IM is likely to fold into a helix-rich domain resembling the Trunk and Paddle of BDLP21. We term this putative region helical domain 2 (HD2).
When bound to tubulated liposomes in the presence of GMPPNP, BDLP bears G domain-Neck and Neck-Trunk rearrangements via so-called Hinge 2 and Hinge 122. Intriguingly, K336 of Mfn1IM exactly overlaps with BDLP R327 at Hinge 2b, whilst Mfn1IM G309 and R74 also have counterparts in BDLP (G309 in Hinge 2b and G68 in Hinge 2a) at equivalent positions (Fig. 1e). Mutation of these Hinge 2-related residues diminished GTPase activity and mitochondrial elongation, although the G domain and HD1 exhibited only limited relative movement in different nucleotide-loading states (Extended Data Fig. 5b–e). Like in dynamins, the potential Hinge 1 between HD1 and predicted HD2 (YSVEER368–373 and EEEIAR692–697) lacks overall conservation among mitofusins (Extended Data Fig. 3). A proline insertion in EEEIAR692–697 abolished mitochondrial elongation activity (Extended Data Fig. 5e, f). Altogether, full-length Mfn1 possibly undergoes conformational changes similar to BDLP when mediating mitochondrial outer membrane (OMM) fusion via aforementioned hinges.
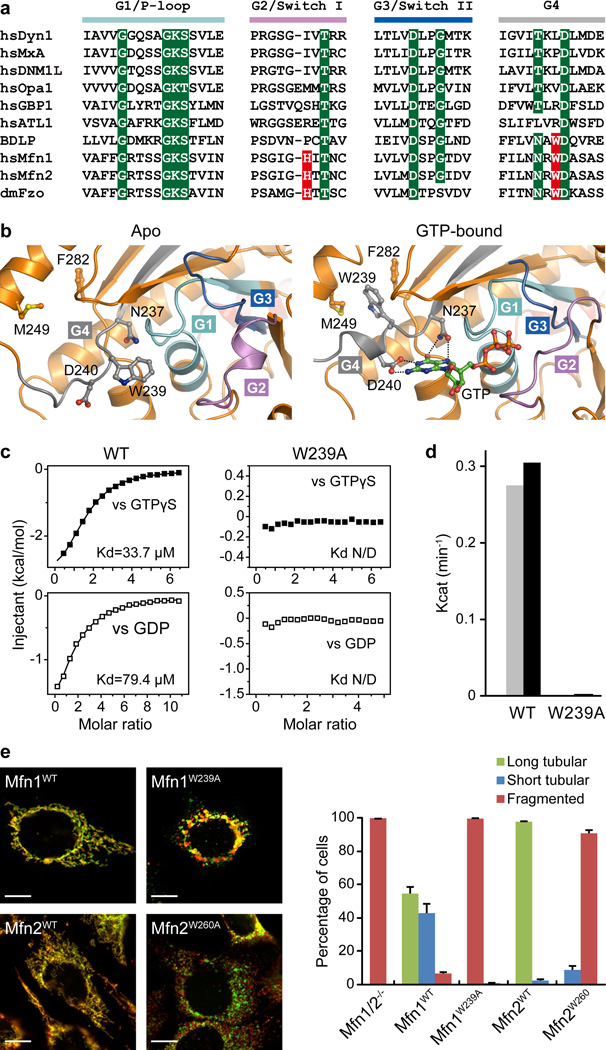
G1-G4 elements of GTPases are essential for binding and hydrolysis of GTP (Fig. 2a). Strikingly, in the nucleotide-free (apo) Mfn1IM structure, the nucleotide-binding pocket is occupied by the bulky side chain of W239 from G4, a residue conserved only in mitofusins and BDLP (Fig. 2b). Loading of GTP drives W239 away, causing it to wedge into a wide hydrophobic groove formed by M249 from α4G and F282 from β6G. This rearrangement allows the suitable positioning of N237 and D240 to dock the guanine base (Fig. 2b). Mutation of W239 to alanine abolished nucleotide-binding and GTPase activity of Mfn1IM (Fig. 2c, 2d). Both Mfn1W239A and the corresponding Mfn2 mutant Mfn2W260A were nonfunctional for mitochondrial elongation (Fig. 2e), manifesting the importance of this tryptophan switch. When accommodating a nucleotide, Mfn1IM utilizes α2’G to loosely buttress the guanine from a vertical orientation, and a large area of the nucleotide is thus exposed. This feature is shared by BDLP but not with Dynamin-121, 23, where the nucleotide is tightly wrapped (Extended Data Fig. 5g).
Figure 2. A tryptophan switch mediates nucleotide binding.
a, Sequence alignment of the G1-G4 elements for different dynamin superfamily members. Conventional motifs/residues are highlighted in green. Key residues found in this paper responsible for nucleotide binding and catalysis of Mfn1 are highlighted in red.
b, Structural details of the nucleotide-binding pocket in nucleotide-free (or apo, left) and GTP-bound (right) states. G1-G4 are colour-specified as in a.
c, Binding affinities to GTPγS and GDP for Mfn1IMWT and Mfn1IMW239A were measured by ITC. N/D: not deducible.
d, GTP turnover rates of Mfn1IMWT and Mfn1IMW239A. Results from two separated experiments are individually presented in grey and black for each protein.
e, Mitochondrial elongation assay for wild-type (WT) and the tryptophan switch mutants of Mfn1/2. In the representative images, red and green fluorescence stains indicate mitochondria and Myc-tagged protein, respectively, for the Mfn1 experiment, and vice versa for the Mfn2 experiment. The data are quantified on the right. For each construct, 100 cells were scored in biological triplicate. Error bars indicate standard errors. Scale bar is 10 µm.
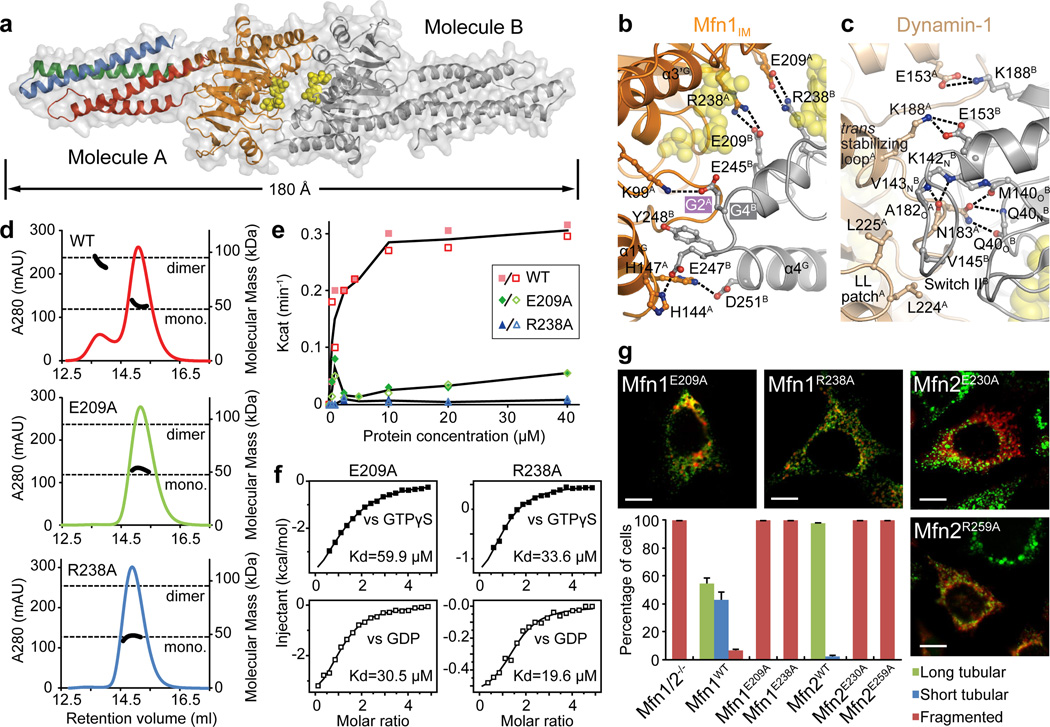
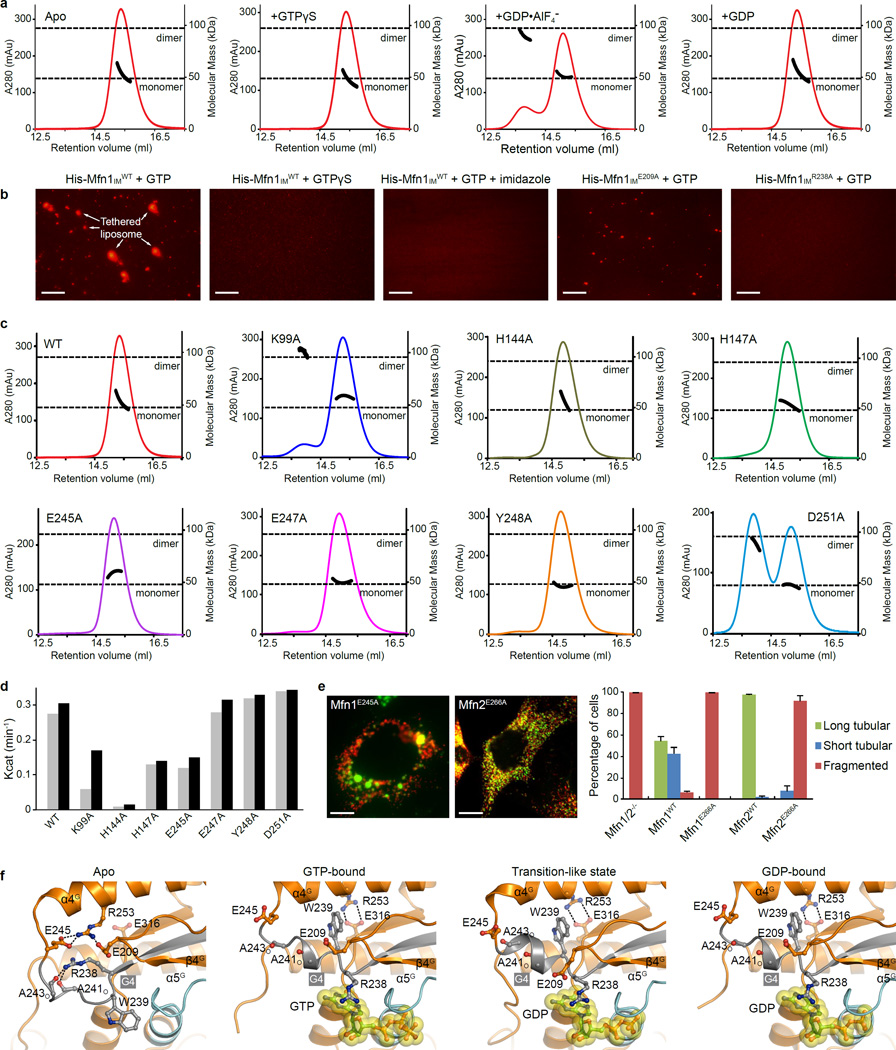
Whereas the apo and GTP structures are monomeric, Mfn1IM forms a homodimer in the presence of the transition state mimic GDP•AlF4− (Extended Data Fig. 6a). Dimerization is mediated by association of the G domains across the nucleotide binding pocket, and the HD1s protrude in opposite directions from the dimer axis (Fig. 3a). Major interactions of this 995 Å2 ‘G interface’ include a pair of symmetrical, parallel aligned salt bridges between R238 in the G4 element and E209 in the loop between β3G and α3G (Fig. 3b). Flanking this central salt bridge pair, close in trans contacts are also observed between K99-E245, H144-E247, and H147-D251. In addition, the side chain of Y248 inserts into the groove between the Switch I and α1’G of the other molecule (Fig. 3b). G domain dimerization has been found in several dynamin superfamily members in the transition state of GTP hydrolysis23–25. Compared to the ∼2,500 Å2 G interface of Dynamin-123 involving extensive hydrogen bonds and hydrophobic associations, the substantially smaller G interface of Mfn1IM is dictated by charged interactions, and no in trans stabilization of the nucleotides is observed (Fig. 3c).
Figure 3. Dimerization of Mfn1IM via G domain.
a, Mfn1IM dimer in the transition-like state, with transparent surface representation. Molecule A is coloured as in Fig. 1b, molecule B in grey. GDP is shown as yellow spheres.
b, Details of the G interface of Mfn1IM. Only one side of the G interface is shown for other involved residues.
c, Details of the G interface of human Dynamin-1 in the transition state (Protein Data Bank code 2×2E). Region corresponding to b is shown.
d, Dimerization properties of Mfn1IMWT, Mfn1IME209A and Mfn1IMR238A in the presence of GDP•AlF4− were assayed in analytical gel filtration coupled to RALS. Calculated molecular masses at the absorption peak of 280 nm are plotted in black.
e, GTP turnover rates of Mfn1IMWT, Mfn1IME209A and Mfn1IMR238A were measured at 7 different protein concentrations. For each group, the averages of Kcat values from two separate experiments at each protein concentration are traced by line charts
f, Binding affinities to GTPγS and GDP for Mfn1IME209A and Mfn1IMR238A.
g, Mitochondrial elongation assay with quantification for Mfn1E209A, Mfn1R238A and related mutants Mfn2E230A, Mfn2R259A. For each construct, 100 cells were scored in biological triplicate; representative images are shown. Error bars indicate standard errors. Scale bar is 10 µm.
To verify the functional relevance of G domain-mediated dimerization, we performed mutagenesis studies on residues E209 and R238. Whereas GDP•AlF4− induced the formation of Mfn1IM dimers in analytical gel filtration coupled to right angle light scattering (RALS) assays, Mfn1IME209A and Mfn1IMR238A failed to dimerize (Fig. 3d). Moreover, the two mutants lacked stimulated protein-concentration-dependent GTPase activity (Fig. 3e), even though they bound guanine nucleotides with wild-type affinity (Fig. 3f). These results suggest that GTPase activation is mediated by dimerization via the G interface. In addition, Mfn1IME209A and Mfn1IMR238A showed suppressed liposome tethering activity in vitro (Extended Data Fig. 6b). Both Mfn1E209A and Mfn1R238A, as well as the corresponding Mfn2 mutants Mfn2E230A and Mfn2R259A, failed to elongate mitochondria (Fig. 3g). Thus, G domain association of mitofusins during the transition state of GTP hydrolysis is an indispensable step for OMM fusion. In addition, mutations of most other residues involved in the G interface also impinge dimerization, GTP hydrolysis and mitochondrial elongation to various extents (Extended Data Fig. 6c–e).
Nucleotide binding induces conformational changes in the residues defining the G dimer interface. In the nucleotide-free state, the conformation of Mfn1IM disfavours dimerization: R238 is blocked by the carbonyl oxygens of A241 and A243, whereas E209 and E245 are attracted by R253 (Extended Data Fig. 6f). GTP-loading-induced rearrangement of the G4 element translocates W239 to push R253 aside towards E316. As a result, E209, E245 and R238 are released and become solvent-facing, allowing for G domain dimerization.
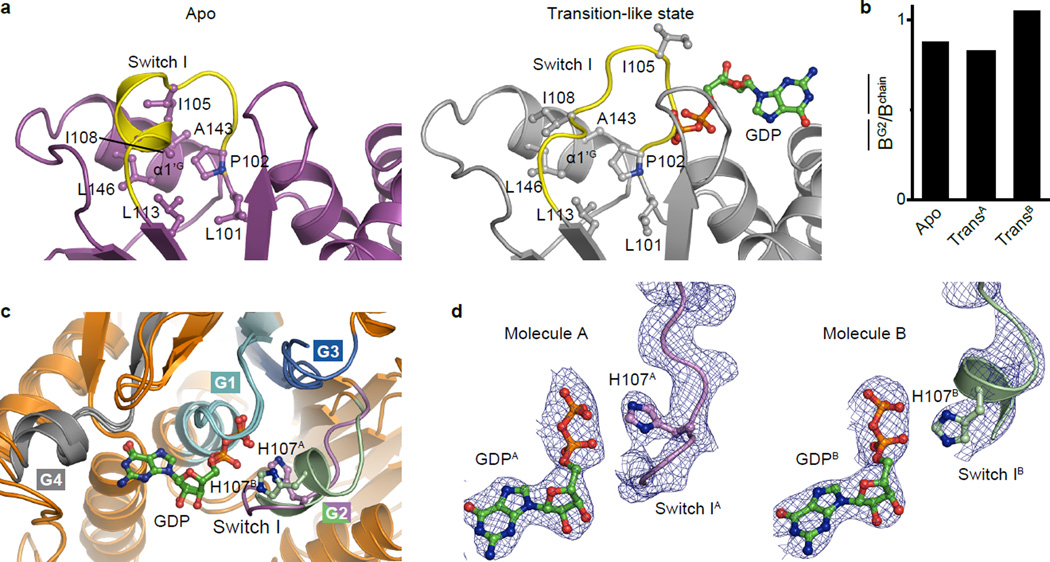
During GTP hydrolysis, the conformationally flexible switch regions must be stabilized to favour catalysis. In our Mfn1IM structure solved from co-crystallization with GDP•AlF4−, although the AlF4− moiety is absent, the Switch I holds a catalysis-compatible conformation analogous to those of other dynamin superfamily members in the transition state (Fig. 4a and Extended Data Fig. 7a–d). Interestingly, unlike many dynamin-related proteins, the Switch I of nucleotide-free Mfn1IM is fixed in a conformation distinct from that in the transition-like state by a hydrophobic network involving G2 element and α1’G, indicating that Switch I is rearranged through dimerization of the G domains (Fig. 2b, Fig. 3b and Extended Data Fig. 7a).
Figure 4. Catalytic machinery of Mfn1.
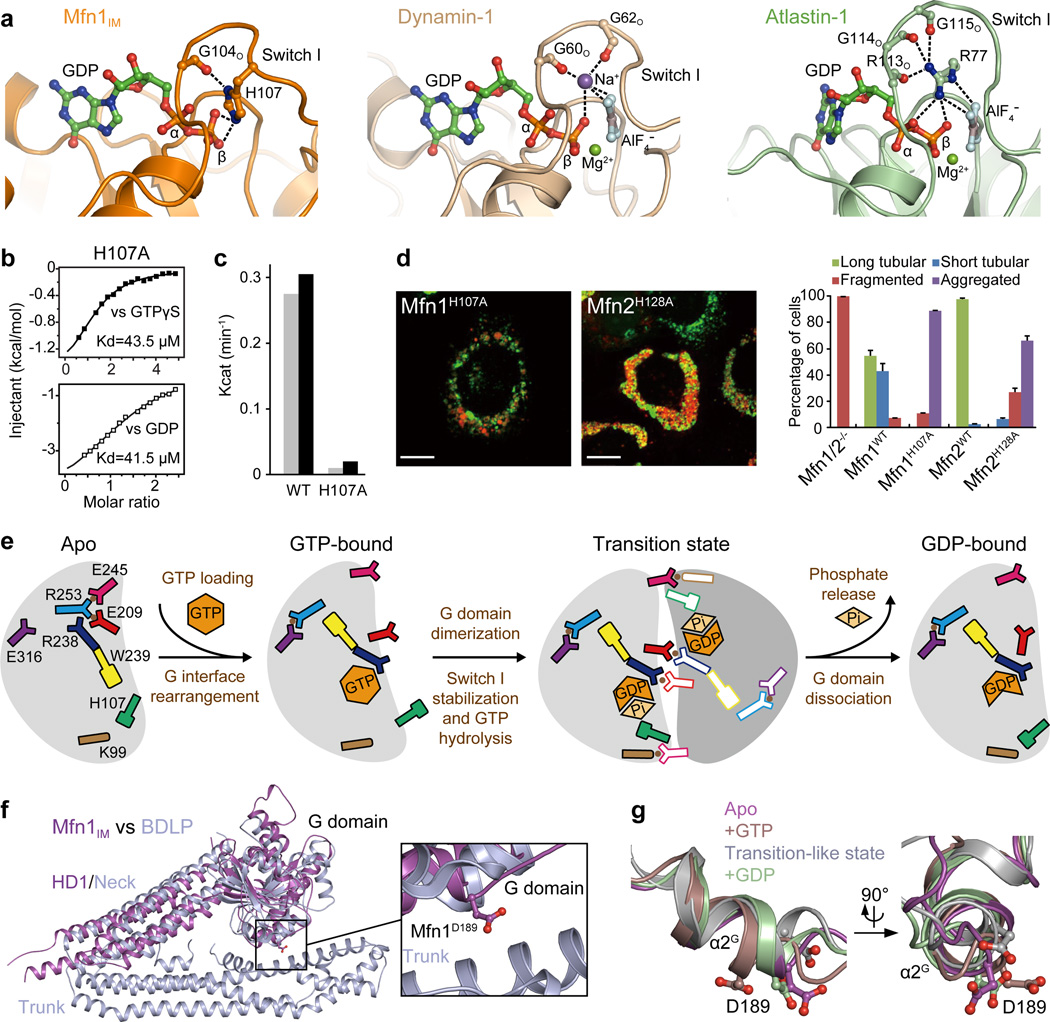
a, Comparison of the catalytic centres of Mfn1 (transition-like state), Dynamin-1 (2×2E) and Atlastin-1 (4IDO).
b, Binding affinities of Mfn1IMH107A to GTPγS and GDP.
c, GTP turnover rates of Mfn1IMWT and Mfn1IMH107A. Results from two separated experiments are presented for each protein.
d, Mitochondrial elongation assay with quantification for Mfn1H107A and related Mfn2H128A. For each construct, 100 cells were scored in biological triplicate; representative images are shown. Error bars indicate standard errors. Scale bar is 10 µm.
e, Schematic drawing summarizing the rearrangements in the G domain during GTP hydrolysis. G domains are coloured grey. Residues and nucleotides are colour/shape-specified. Salt bridges are specified by brown dots between involved residues. Pi stands for phosphate ion.
f, Superposition of nucleotide-free Mfn1IMB and BDLP (2J69) with the predicted-HD2-facing D189 of Mfn1 specified.
g, Structural comparison of Mfn1IM in different nucleotide-loading states at α2G reveals distinct orientation of D189 in the GTP-bound state from other states.
Efficient GTP hydrolysis requires the neutralization of negative charge developing between the β- and γ-phosphates in the transition state. A sodium ion coordinated by a QS motif in the P-loop takes over this role in Dynamin-123, whereas Atlastin-1 possesses a cis-arginine finger in the P-loop24 (Fig. 4a). However, these motifs are not present in Mfn1. Instead, a mitofusin-specific H107 in Switch I stretches to the β-phosphate of GDP with a similar position as the charge-compensating ion/residue of Dynamin-1 and Atlastin-1. Furthermore, the side chain of H107 is coordinated by the carbonyl oxygen of Gly104, which corresponds to G60 of Dynamin-1 and G114 of Atlastin-1 (Fig. 2a, Fig. 4a). Mutation of H107 to alanine did not perturb the binding of guanine nucleotides but eliminated the GTPase activity of Mfn1IM (Fig. 4b, c). Mfn1/2-null MEFs transfected with either Mfn1H107A or the corresponding mutant Mfn2H128A were defective in mitochondrial elongation (Fig. 4d). Thus, we propose this H107 finger as a charge-compensating factor during catalysis. The catalytic water responsible for the nucleophilic attack on the γ-phosphate was not observable, probably due to the absence of the AlF4− moiety.
Compared to other scission-related dynamin family members, Mfn1IM shows extraordinarily weak GTPase activity (Fig. 2d). The stable conformation of Switch I and the shielded GTP-binding pocket in the nucleotide-free state may account for this feature (Fig. 4e). Slow GTP turnover, exemplified by BDLP22 and EHD215, has been suggested to favour self-assembly over disassembly, and thus promote membrane fusion26.
We also purified a near-full-length construct termed Mfn1ΔTM that contains the G domain, HD1 and HD2 (Extended Data Fig. 8a, b). Mfn1ΔTM elutes as a stable dimer in RALS assays without nucleotide (Extended Data Fig. 8c). This dimer, possibly mediated by HD2, may relate to the 180 kDa stable complex of rat Mfn1 found in sedimentation studies27. Moreover, Mfn1ΔTM oligomerizes only when GDP•AlF4− is present (Extended Data Fig. 8d), consistent with the previous observation that the 180 kDa Mfn1 complex can form higher-order oligomers when incubated with GTP but not with GTPγS27. Mfn1ΔTM with either E209A or R238A failed to oligomerize (Extended Data Fig. 8e), indicating that oligomerization is dependent on the G interface.
In the apo and GMPPNP-bound states, the Neck and Trunk of BDLP take either ‘closed’ or ‘open’ conformation21, 22, and the contact between G domain and Trunk may stabilize the former. Given the structural congruence with BDLP, Mfn1 may also adopt this feature. A conserved D189 potentially participates in the plausible G domain-HD2 contact (Fig. 4f). This surface residue located on α2G is differently oriented between nucleotide-free and GTP-bound states, and in latter case its side chain swings away from the HD2 (Fig. 4g, Extended Data Fig. 8f, g). Mfn1D189A led to mitochondrial clumping (Extended Data Fig. 8h).
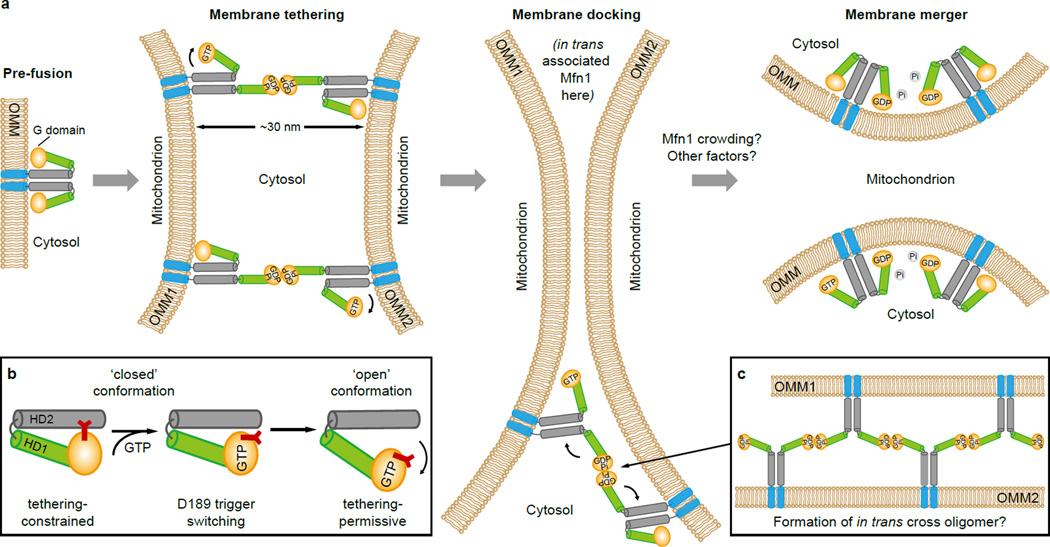
Our data highlight the role of G domain dimerization, regulated by guanine nucleotide, in membrane fusion mediated by mitofusin. GTP loading may induce a conformational change from the ‘closed’ tethering-constrained state to the ‘open’ tethering-permissive state, possibly involving the D189 trigger. In the ‘open’ state, stretched Mfn1 molecules allow efficient tethering of two OMMs with a distance of as far as ∼30 nm. During GTP hydrolysis, HD2 may fold back to the ‘closed’ state to bring opposing membranes in close proximity (Extended Data Fig. 9a–c). An analogous model based on the BDLP1 structure has been proposed28. This process may be reversible and controllable by local GTP concentration and Mfn1 density, so that excessive tethering can be avoided.
A key issue to be resolved is the reconciliation of this model with the previous Mfn1 tethering model, in which an antiparallel coiled coil at the C-terminus played a central role13, 29_ENREF_13. One possibility is that Mfn1 operates through sequential tethering events, an initial one utilizing nucleotide-regulated, G domain dimerization followed by closer apposition via the coiled coil. For this sequence to happen, a large energy barrier must be overcome to detach α4H from the large hydrophobic network in HD1. It should also be noted that the G domain association of Mfn1 may as well occur in cis, as both forms have been proposed to have functions in BDLP22 and Atlastin-130.
Methods
Protein expression and purification
cDNAs of all truncated human Mfn1 constructs, including those for crystallization and indicated mutants for biochemical assays were individually cloned into a modified pET28 vector. For constructs used in co-crystallization with GTP, an extra T109A mutation was introduced to Mfn1IMC to suppress the GTPase activity. Details of these constructs were summarized and illustrated in Extended Data Fig. 1a. For Mfn1ΔTM, residues 580–631 were replaced by an (SAA)5 linker. For Mfn1IM and corresponding mutants, recombinant proteins containing an N-terminal His6 tag followed by a cleavage site for PreScission protease (PSP) were expressed in E. coli Rosetta (DE3) cells. Transformed bacteria were cultured at 37°C before induced with 0.1 mM isopropyl-1-thio-β-D-galactopyranoside (IPTG) at an OD600 of 0.6, and grown overnight at 17∼18°C. Cells expressing Mfn1IM were lysed in 50 mM HEPES (pH 7.5), 400 mM NaCl, 5 mM MgCl2, 30 mM imidazole, 1 µM DNase I, 1 mM Phenylmethanesulfonyl fluoride (PMSF) and 2.5 mM β-mercaptoethanol (β-ME) using a cell disruptor (JNBIO) and subjected to centrifugation at 40,000g for 1 h. The supernatant was filtered and applied to a Ni-NTA (first Ni-NTA) column (GE healthcare) equilibrated with Binding Buffer 1 containing 20 mM HEPES (pH 7.5), 400 mM NaCl, 5 mM MgCl2, 30 mM imidazole and 2.5 mM β-ME. After washed with Binding Buffer 1, proteins were eluted with Elution buffer containing 20 mM HEPES (pH 7.5), 400 mM NaCl, 5 mM MgCl2, 300 mM imidazole and 2.5 mM β-ME. Eluted proteins were incubated with 2 µg GST-fused PSP to remove the His6-tag and dialysed overnight against Binding Buffer 2 containing 20 mM HEPES (pH 7.5), 400 mM NaCl, 5 mM MgCl2 and 2.5 mM β-ME. After dialysis, PSP was removed using a GST column. The protein was re-applied to a second Ni-NTA column equilibrated with Binding Buffer 2. Binding Buffer 1 was used to elute the proteins which were subsequently loaded onto a Superdex200 16/60 column (GE healthcare) equilibrated with Gel Filtration Buffer containing 20 mM HEPES (pH 7.5), 150 mM NaCl, 5 mM MgCl2 and 1 mM dithiothreitol (DTT). The proteins eluted in a discrete peak corresponding to a molecular mass of approximately 50 kDa. Cell lysis and protein purification were both performed at 4°C. The selenomethionine (SeMet) derivative of Mfn1IMB was expressed in E. coli Rosetta (DE3) in M9 minimal media. For 1 L bacteria culture, 100 mg lysine, 100 mg phenylalanine, 100 mg threonine, 50 mg isoleucine, 50 mg leucine, 50 mg valine and 60 mg SeMet were added when OD600 reached 0.5. The cells were then induced with 100 µM IPTG and cultured at 18°C for 24 h. Purification protocols for the SeMet derivative were the same as those of native protein. Mutants used in RALS, ITC and GTP hydrolysis assays were all based on Mfn1IMC, unless specified. Structure of Mfn1IMB in the nucleotide-free state (assigned PDB code: 5GO4) was used in Fig. 1 are representative.
Constructs based on Mfn1ΔTM possesses an extra Strep-tag was inserted between His6 tag and PSP cleavage site. Cells expressing Mfn1ΔTM (WT or mutants) were harvested and lysed in 50 mM HEPES (pH 7.5), 400 mM NaCl, 5 mM MgCl2, 30 mM imidazole, 1% Tween-20, 1 mM DNase I, 1 mM PMSF and 2.5 mM β-ME with a cell disruptor (JNBIO). After centrifugation at 40,000g for 1 h, the supernatant was filtered and applied to a Ni-NTA column equilibrated with Binding Buffer 3 containing 20 mM HEPES (pH 7.5), 400 mM NaCl, 5 mM MgCl2, 30 mM imidazole, 1% Tween-20, and 2.5 mM β-ME. Proteins were eluted with Elution Buffer after washed with Binding Buffer 1 and reapplied to a StrepTactin column (GE healthcare) equilibrated with Binding Buffer 2. Binding Buffer 2 containing extra 2.5 mM desthiobiotin was used to elute the proteins. After the tags and PSP were removed in the same way as for Mfn1IM, proteins were applied to gel filtration using a Superdex200 16/60 column equilibrated with Gel Filtration Buffer for GTPase assays or buffer containing 20 mM HEPES (pH 7.2), 30 mM NaCl, 5 mM MgCl2 and 1 mM DTT for RALS assays.
Protein crystallization
Purified Mfn1IM constructs were pre-incubated with corresponding nucleotides in 10-fold concentration relative to protein for 2 h before crystallized at 20°C via hanging drop vapour diffusion by mixing equal volumes of protein (15∼30 mg/ml) and reservoir solution. Crystals of native and SeMet Mfn1IMB (apo) grew from 0.2 M sodium citrate tribasic dehydrate and 20∼22% PEG 3350 overnight in the presence of GMPPCP. These crystals were then soaked in reservoir solution mixed with an equal volume of Gel Filtration Buffer supplemented with 18∼20% PEG 400 for cryo-protection. GTP bound Mfn1IMCT109A grew in 1 mM ZnCl2, 100 mM Tris-HCl (pH 8.0∼9.0), 15∼16% PEG 3350, 10∼15% glycerol overnight and cryo-protected by reservoir solution containing extra 15% ethylene glycol. The crystals of Mfn1IMC in transition state were obtained in 100 mM MES (pH 6.0), 240 mM NaCl, 36% pentaerythriol propoxylate (5/4 PO/OH) in the presence of 10-fold GDP, 10-fold AlCl3 and 100-fold NaF. GDP-bound Mfn1IMCT109A grew in 0.2 M ammonium citrate tribasic and 20% PEG 3350 for 5 days in the present of 10-fold GTP. Extra 15% ethylene glycol were used for cryo-protection. GTPγS-bound Mfn1IMA were crystallized in 0.1 M MIB (pH 6.0∼8.0, PACT suite, QIAGEN), 25% PEG 1500 and directly flash-frozen in liquid nitrogen. Complex of Mfn1IMA with GDP was crystallized in 50∼80 mM sodium citrate and 23∼25% PEG 3350 after a week. Extra 20% MPD was used for cryo-protection. All crystals were stored in liquid nitrogen before diffraction assays. We also obtained crystals of apo Mfn1IMA and apo Mfn1IMB in conditions without nucleotides, but these crystals did not diffract X-ray well.
Structure determination
All diffraction data sets were collected at beamline BL17U1 of the Shanghai Synchrotron Radiation Facility (SSRF) and processed with the XDS suite31. Initial phases were calculated and refined by the single anomalous dispersion method using SHELXC/D/E32 with the graphical interface HKL2MAP33 from a diffraction data set of SeMet-substituted Mfn1IMB crystal in the apo form. Other structures were solved by molecular replacement using MolRep34 and Phaser35 with the structure of SeMet Mfn1IMB as the search model. Models were built with COOT36 and refined with Refmac37 and Phenix38. Structural validation was carried out using MolProbity39. Structural illustrations were prepared using PyMOL40. The 6.1 Å Mfn1IMA•GTPγS structure and 4.3 Å Mfn1IMA•GDP structure were not further refined after molecular replacement. X-ray data collection and refinement statistics can be found in Extended Data Table 1. The area of the G interface was calculated using the PISA server41. The Ramachandran statistics determined by PROCHECK42 are as follows: 98.3% in favoured region, 1.7% allowed, no outlier for apo Mfn1IMB; 98.7% favoured, 1.0% allowed, 0.3% outlier for GTP-bound Mfn1IMCT109A; 95.8% favoured, 4.1% allowed, 0.1% outlier for transition-like state Mfn1IMC and 98.9% favoured, 1.7% allowed, 0 outlier for GDP-bound Mfn1IMCT109A.
Right angle light scattering assay
A coupled RALS-refractive index detector (Malvern) was connected in line to an analytical gel filtration column Superdex200 10/300 to determine absolute molecular masses of the applied protein samples. For each experiment, 100 µM purified Mfn1IMC (WT or mutants) or 10∼30 µM Mfn1ΔTM (WT or mutants) was incubated with or without 1 mM corresponding ligand for 6 h at 25°C prior to the measurement. The column was equilibrated with 20 mM HEPES (pH 7.2), 30 mM NaCl, 5 mM MgCl2 and 1 mM DTT. Data were analysed with the provided OMNISEC software. All experiments were repeated at least twice and the data showed satisfying consistency.
GTP hydrolysis assay
GTP hydrolysis assays for Mfn1IMC, Mfn1ΔTM and corresponding mutants were carried out at 37°C in 20 mM HEPES (pH 7.5), 150 mM NaCl, and 5 mM MgCl2 and 1 mM DTT as described earlier43. For measuring stimulated GTP turnover of Mfn1IMCWT, Mfn1IMCE209A, and Mfn Mfn1IMCR238A, protein at concentrations of 0.5 µM, 1 µM, 2.5 µM, 5 µM, 10 µM, 20 µM and 40 µM were individually mixed with 1∼2 mM GTP and hydrolysis rates were determined from a linear fit to the initial rate of the reaction (<40% GTP hydrolysed). For other experiments, 20 µM protein and 1 mM GTP were used.
Nucleotide binding study
The equilibrium dissociation constants for Mfn1IMC and indicated mutants to guanine nucleotides were determined by isothermal titration calorimetry (ITC) using a MicroCal ITC200 (Malvern) at 25°C. 2 mM nucleotide was titrated at 2 µl step against 60∼80 µM protein in the buffer containing 20 mM HEPES (pH 7.5), 150 mM NaCl, 5 mM MgCl2, 2.5 mM β-ME. Resulting heat changes upon each injection was integrated and the values were fitted to a standard single-site binding model using Origin7. All experiments were repeated at least twice and the data showed satisfying consistency.
Mitochondrial elongation assay
To examine the effect of point mutations, Mfn1-Myc and Mfn2-Myc variants were expressed in Mfn1/2-null MEFs from the pQCXIP retroviral vector. MEFs were generated in-house. The cell line is free of myoplasma and has been authenticated by genotyping with PCR to confirm deletion of the Mfn1 and Mfn2 genes. Point mutants were constructed by overlapping PCR with primers encoding the point mutation. All mutations were confirmed by DNA sequencing. Retroviral supernatants were produced from 293T cells transfected with the retroviral vector and the packaging plasmid pCLEco. Mfn1/2-null MEFs were maintained in Dulbecco’s Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum and penicillin/streptomycin at 37°C and 5%CO2. After retroviral transduction of Mfn1/2-null MEFs, puromycin selection was applied for 2 days. In the case of Mfn1 constructs, Mfn1/2-null cells stably expressing mito-DsRed were used, green fluorescence is from immunostaining against the Myc epitope; for Mfn2 constructs, Mfn1/2-null cells stably expressing mito-GFP were used, red fluorescence is from immunostaining against the Myc epitope. Cells were plated onto 8-well chambered slides for analysis. Immunostaining and Western blot analysis with the 9E10 antibody against Myc was performed to confirm proper expression of the mitofusin variant. Mitochondrial morphology was scored by analysis of mito-DsRed or mito-GFP as described previously12.
Liposome tethering assay
POPC (1-palmitoyl-2-oleyl-sn-glycero-3-phosphocholine), DOPS (1,2-dioleyl-sn-glycero-3-phospho-L-serine), DOGS-NTA-Ni2+ (1,2-dioleoyl-sn-glycero-3-[(N-(5-amino-1-carboxypentyl)iminodiacetic acid)succinyl] (nickel salt)) and Rho-DPPE (1,2-dipalmitoyl-sn-glycero-3-phosphoethanolamine-N-(lissamine rhodamine B sulfonyl)) (Avanti Polar Lipids) were mixed in a molar ratio of 78.5:15:5:1.5. Lipid film formed by evaporating chloroform under mild Nitrogen stream was dehydrated in a vacuum drier before re-suspended in Buffer L (20 mM HEPES, pH 7.5, 150 mM NaCl, 2 mM β-ME) to a final concentration of 10 mM. liposomes were prepared as previously described30. An internal His12 tag was inserted between α3H and α4H of Mfn1IMC and corresponding mutants. For each reaction, 0.5 µM protein was mixed with 1 mM prepared liposomes in Buffer L for 30 minuets at 4°C before 5 mM GTP or GTPγS were subsequently added. After incubation for another 40 minutes at 37°C, the proteoliposomes were imaged by a fluorescent microscope. Otherwise, the proteoliposomes were incubated with 300 mM imidazole (pH 7.5) for another 20 minutes to release the protein and then imaged. All experiments were repeated at least twice and the data showed satisfying consistency.
Data availability
The X-ray crystallographic coordinates and structure factor files for Mfn1IM structures have been deposited in the Protein Data Bank (http://www.rcsb.org) under following accession numbers shown in the brackets: apo MfnIMB (5GO4), GTP-bound Mfn1IMCT109A (5GOF), transition-like state Mfn1IMC (5GOM), and GDP-bound Mfn1IMCT109A (5GOE). All other data generated or analysed during this study are included in this published article, and are available from the corresponding author upon reasonable request.
Extended Data
Extended Data Figure 1. Mfn1 constructs and their biochemical properties.
a, Schematic representation showing the strategy of generating human Mfn1 constructs for crystallization. Indication of the labels and numbers are the same as in Fig. 1a. HR1T and HR2T stand for truncated HR1 and HR2. We removed the TM and flanking residues from human Mfn1 and inserted artificial linkers as illustrated. Three different constructs used for crystallization are named Mfn1IMA, Mfn1IMB and Mfn1IMC, respectively (collectively termed Mfn1IM).
b, Summary of the crystal structures. “Initial ligands” stands for the ligands added to the protein solution before crystallization, whereas “Final state” means the contents from the refined structure. Resolutions for the structure are specified.
c, Isothermal titration calorimetry (ITC) results showing that Mfn1IM constructs have no binding affinity to GMPPNP or GMPPCP. Kd N/D: Kd value is not deducible. Only the result of the Mfn1IMC construct is shown here as representative.
d, Electron density of the guanine nucleotides in corresponding structures. The election density maps are all shown at a contour level of 1.2σ. The residues involved in ligand coordination are shown as ball-and-stick models.
e, Details of the Mfn1IM active site in the GTP-bound state. Key hydrogen bonds for coordinating the GTP were indicated by dotted lines. In lower panel, details of the Mg2+ coordination is depicted. The electron density for Mg2+ ion, water and GTP was shown as grey mesh at a contour level of 1.2σ.
f, GTP turnover rates of Mfn1IMWT and Mfn1IMT109A. Mfn1IMT109A shows greatly impaired GTPase activity that facilitates the co-crystallization with GTP. Results from two separated experiments are presented for each protein.
g, ITC results showing that Mfn1IMT109A binds both GTP and GDP.
Extended Data Figure 2. Overall structure of Mfn1IM.
a, The topology diagrams of the G domains of Ras, Mfn1 and BDLP. Secondary structural elements were not drawn to scale and positions of G1-G4 motifs are indicated. Elements of Mfn1 are named and coloured as in Fig. 1c. For BDLP, elements extra than Ras in light blue. The helices of BDLP are named as in ref21.
b, Helical wheel diagrams of HD1. Hydrophobic residues are coloured yellow and other residues are in the same colour with the corresponding helices as in Fig. 1d. The plots are arranged according to the positions of the four helices of HD1 in the crystal structure, showing a massive hydrophobic core of HD1.
c, Intramolecular association of Mfn1IM. For the G domain-HD1 interaction, L8, M76, V333 and F337 embrace F11, whereas K15 forms a salt bridge with D173 and a hydrogen bond with the main chain oxygen of R74. Mfn1L705P mutant was previously found to be non-functional in mediating mitochondrial fusion13. L705 is surrounded by several hydrophobic residues including I45, I48, A362 and I708, as well as a salt bridge formed by R365 and E701. The proline mutation of L705 may disrupt α4H and the local hydrophobic interactions, thereby impeding the folding of the protein.
d, GTP turnover rates of Mfn1IMWT and Mfn1IMK15A and Mfn1IML705P. Results from two separated experiments are presented for each protein.
e, Mitochondria elongation assays of Mfn1WT and Mfn1K15A. The Myc-tagged Mfn1 constructs were assayed for mitochondrial elongation activity by expression in Mfn1/2-null MEFs, which have completely fragmented mitochondria. Overexpression of Mfn1WT in Mfn1/2-null MEFs induces the formation of mostly tubular mitochondria, indicating normal elongation activity, whereas Mfn1K15A induces substantially less mitochondrial tubulation. Green fluorescence is from immunostaining against the Myc epitope; red fluorescence is from mito-DsRed. The data are quantified on the right. For each construct, 100 cells were scored in biological triplicate; representative images are shown. Error bars indicate standard errors. Scale bar is 10 µm.
Extended Data Figure 3. Sequence alignment of mitofusins and BDLP.
Sequence alignment of mitofusins and BDLP. Amino acid sequences of human (hs) Mfn1 (UniProt accession Q8IWA4) and Mfn2 (O95140), mouse (mm) Mfn1 (Q811U4) and Mfn2 (Q80U63), fruit fly (Drosophila melanogaster, dm) Marf (Q7YU24), fruit fly Fzo (O18412) and the bacterial dynamin-like protein (BDLP) from Nostoc punctiforme (B2IZD3) are aligned using Clustal W44. Residues with a conservation of 100% are in red shades, greater than 80% in green shades and 50% in grey shades, respectively. α-helices are shown as cylinders and β-strands as arrows for both nucleotide-free human Mfn1 (above the sequences) and nucleotide-free BDLP (2J,under the sequences). In the case of human Mfn1, the secondary structure signs are coloured as in Fig. 1b and labelled as in Figs. 1b–d for Mfn1IM regions. Secondary structural elements of the missing HD2 and TM predicted from the PHYRE2 server45 (exclusively α-helices) are depicted as shaded cylinders with dashed outlines. For BDLP, the secondary structure signs are coloured grey and labelled according to the previous report21. The G1-G4 elements are specified in the sequences. Key residues on human Mfn1 are also indicated, including those involved in the hydrophobic core of HD1 (♦), Hinges (▼), guanine nucleotide binding and hydrolysis (●), G interface (▲), and the plausible HD1-HD2 conformational change (■).
Extended Data Figure 4. Structural comparison of Mfn1IM with other dynamin family members.
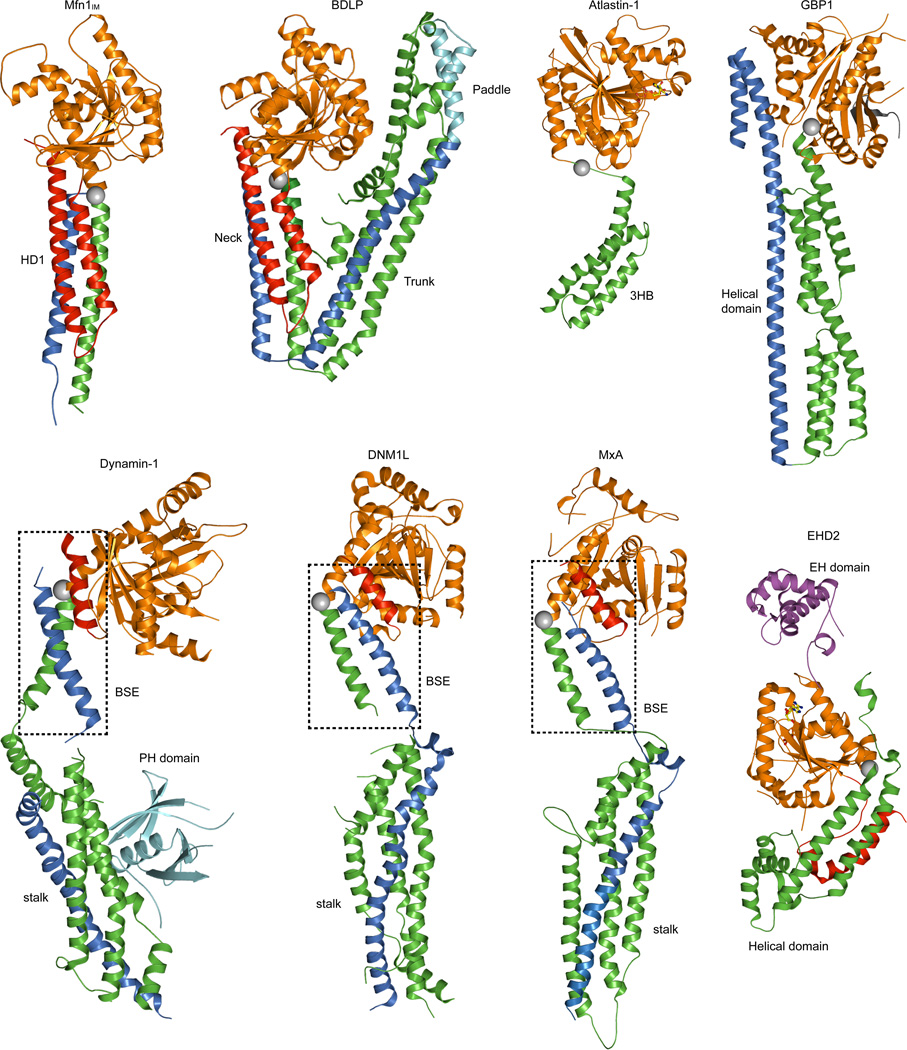
Structural comparison of nucleotide-free Mfn1IMB with nucleotide-free BDLP (PDB accession code 2J69)21, GDP-bound Atlastin-1 (3Q5D)19, nucleotide-free GBP1 (1DG3)46, nucleotide-free Dynamin-1 (3SNH)17, nucleotide-free DNM1L (4BEJ)47, nucleotide-free MxA (3SZR)16, and AMPPNP-bound EHD2 (2QPT)15. For these molecules, the region N-terminal to the G domain is in red, G domain itself in orange, the conventional middle domain in green, and the conventional GTPase Effector Domain (GED) in marine, the Paddle region of BDLP and the Pleckstrin homology (PH) domain of Dynamin-1 in cyan, and the Eps15 homology (EH) domain of EHD2 in magenta. The Hinges between the G domains and middle domains are depicted by grey spheres. Nucleotides are shown as ball-and-stick models.
Extended Data Figure 5. Structural comparison of Mfn1IM with BDLP and Dynamin-1.
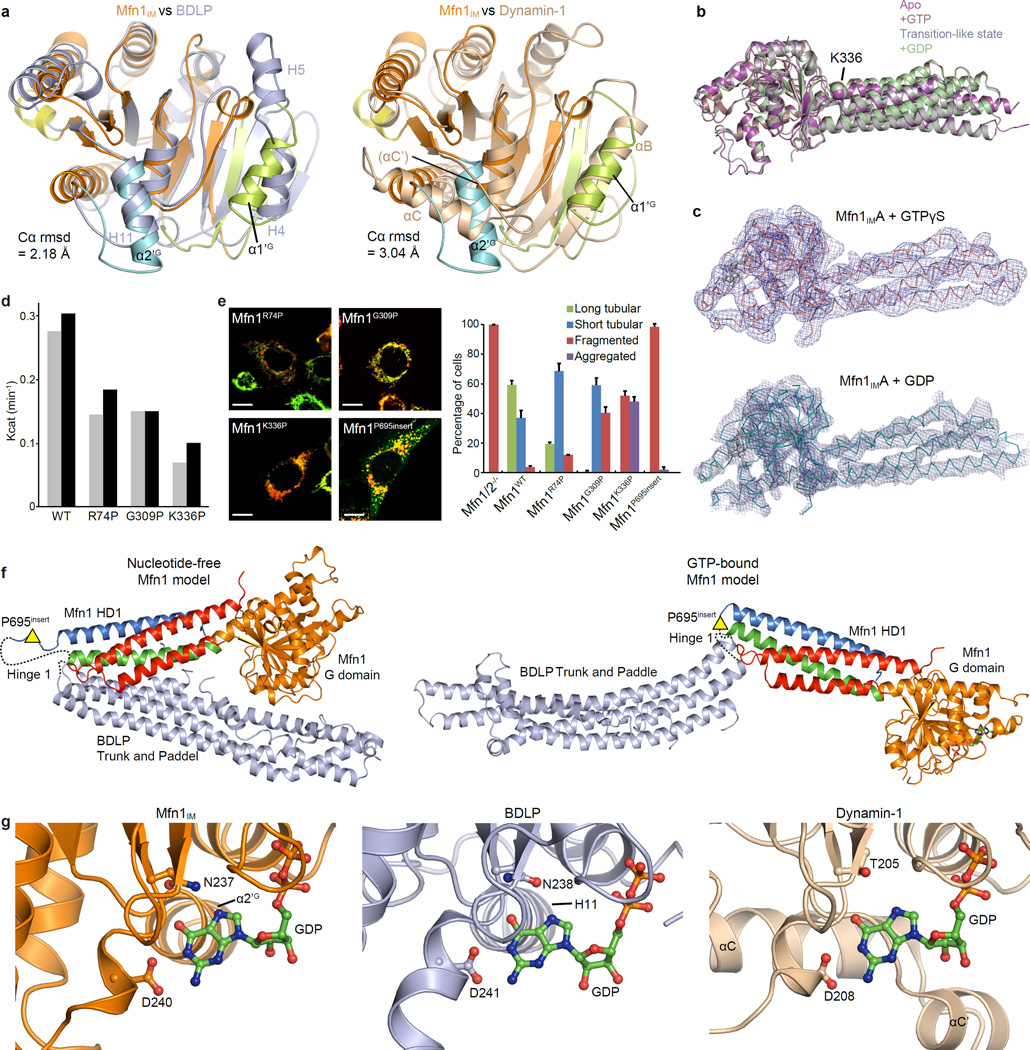
a, Structural comparison of the G domains between Mfn1IM and BDLP (left) or Dynamin-1 (right) in the nucleotide-free state. The Mfn1 G domain (coloured as in Fig. 1b) is separately superimposed with G domains of BDLP (2J69, light blue) and of rat Dynamin-1 (2AKA, wheat). The root mean standard deviation (rmsd) of aligned Cα atoms are indicated. α-helices on the two lobes are labelled for the three molecules. The G domain of Mfn1IM resembles the BDLP G domain, except that at lobe 1, BDLP has two separate α-helices (H4 and H5) rather than the single α1’G of Mfn1IM. Dynamin-1 is similar to Mfn1IM in lobe 1, but at lobe 2 the αC tilts 60° from its counterpart α2’G in Mfn1IM. In addition, the long loop N-terminal to αC (termed α’C with brackets) in Dynamin-1 folds into a short α-helix (αC’) when guanine nucleotide is loaded, which is not the case in Mfn1IM.
b, Structural comparison of Mfn1IM in different nucleotide-loading states. Structures of nucleotide-free Mfn1IMB, GTP-bound Mfn1IMCT109A, transition-like state Mfn1IMC and GDP-bound Mfn1IMCT109A are colour-specified and superimposed on their G domains.
c, Architectures of Mfn1IMA•GTPγS and Mfn1IMA•GDP. Shown here are the corresponding Cα traces and electron density maps with contour level at 1.2σ. The outlines of the molecules are clearly discernable from the resulting electron density maps, showing that the HD1 does not have large-scale movement relative to the G domain compared with structures shown in b. These two structures are presented to exclude the possible influence of the T109A mutation for the orientation of HD1 in the GTP- and GDP-bound structures with higher resolution.
d, GTP turnover rates of Mfn1IMWT and the Hinge 2 mutants. Results from two separated experiments are presented for each protein. Note that although all three Hinge 2 mutants had similar GTPase activities, Mfn1K336P showed much more significantly reduced activity in mediating mitochondrial elongation.
e. Mitochondrial elongation assay for Mfn1WT and the hinge mutants. For each construct, 100 cells were scored in biological triplicate; representative images are shown. Error bars indicate standard errors. Scale bar is 10 µm.
f, Full-length Mfn1 models showing the plausible Hinge 1 between HD1 and HD2. Models were based on nucleotide-free (2J69, upper) and GMPPNP-bound (2W6D, lower) BDLP. G domain and HD1 are coloured as in Fig. 1b, and HD2 in light blue. Hinge 1 was shown as dashed lines. The approximate position for the P695 insert was indicated by yellow triangles.
g, Extra support of the guanine base in Mfn1, BDLP (2J68) and Dynamin-1 (5D3Q). GDP-bound structures of Mfn1IMCT109A (coloured as in Fig. 1c), BDLP (light blue) and Dynamin-1 (wheat) are shown in ribbon-type representations. The nucleotides and the residues on G4 element involved in guanine base coordination are shown as ball-and-stick models. The α-helices in the G domains that support the guanine base are specified. Part of the G domains are removed for clarity. Note that for Mfn1IM and BDLP, the α2’G and H11 only loosely associate with one side of the guanine base, whereas αC of Dynamin-1 (corresponding to α2’G in Mfn1IM) tightly wraps the guanine base from a parallel orientation together with a specific element αC’.
Extended Data Figure 6. Dimerization of Mfn1IM G domains in the transition-like state.
a, Oligomerization states of Mfn1IM in different nucleotide-loading conditions by RALS. Mfn1IM is monomeric in nucleotide-free, GTPγS-bound and GDP-bound states, and forms dimers in the presence of GDP•AlF4−. Data are presented as in Fig. 3d.
b, liposome tethering assay for Mfn1IMWT and corresponding mutants. Representative images from 5 separate experiments are shown. Mfn1IMWT tethered liposomes carrying fluorescence in the presence of GTP hydrolysis-dependent manner as large aggregated liposomes were observed (first left). In GTPγS-present condition the liposome aggregation was largely attenuated, suggesting that tethering is dependent on GTP hydrolysis (second left). When proteins were washed off the liposome by imidazole, the liposomes became homogeneously scattered (middle), indicating that the liposomes were tethered but did not merger. Mfn1IME209A and Mfn1IMR238A displayed suppressed tethering activity (right two). Scale bar is 50 µM.
c, Dimerization test of the G interface mutants in the presence of GDP•AlF4−.
d, GTP turnover rates of the G interface mutants compared with Mfn1IMWT. Results from two separated experiments are presented for each protein.
e, Mitochondrial elongation assay for Mfn1E245A and related Mfn2E266A. For each construct, 100 cells were scored in biological triplicate; representative images are shown. Error bars indicate standard errors. Scale bar is 10 µm. Both mutants lost fusogenic activity.
f, Rearrangement of residues in the G interface upon nucleotide binding. Structures shown from left to right are: nucleotide-free Mfn1IMB; GTP-bound Mfn1IMCT109A; transition-like state Mfn1IMC; and GDP-bound Mfn1IMCT109A. Key residues involved in the structural rearrangement of the G interface are shown as ball-and-stick models. Yellow surface representation is used for GTP and GDP.
Extended Data Figure 7. Analysis of the Switch I conformations.
a, Configuration of Switch I of Mfn1IM in nucleotide-free and the transition-like states (molecule A of the dimer is used). Switch I is coloured yellow. Residues involved in the hydrophobic networks are shown as ball-and-stick models. Note the rearrangements of this region between the two states.
b, Stability of Switch I region of Mfn1IM at different states. The stability of Switch I is reflected by the mean B factor of the main chain atoms of Switch I compared to that of the whole peptide chain. TransA and TransB stand for molecule A and B of the Mfn1IMC dimer in the transition-like state. The Switch I regions in both nucleotide-free and transition-like (TransA) states have relatively stable conformations with regard to the whole molecule.
c, Superposition of the GTPase catalysis centres of two molecules of the Mfn1IMC dimer in the transition-like state. The G1-G4 elements are coloured and labelled as in Fig. 2b, except the G2 element of the molecule B in pale green. H107 and is shown as ball-and-stick models.
d, The electron density of the Switch I regions in the two molecules of the Mfn1IMC dimer. The density is shown as blue mesh at a contour level of 1.2σ for both molecule A (left) and molecule B (right). H107 is shown as ball-and-stick models.
Extended Data Figure 8. Characterization of Mfn1ΔTM and the D189 trigger.
a, Schematic representation showing the strategy of generating the Mfn1ΔTM construct. Colour as in Fig. 1a, and HD2 is in purple.
b, Comparison of GTPase activity between Mfn1IMC and Mfn1ΔTM. Results from two separated experiments are presented for each protein.
c, RALS analysis of Mfn1ΔTM showing that it is a stable dimer in nucleotide-free state.
d, Analytical gel filtration results of Mfn1ΔTM in the GTPγS, GDP•AlF4− and GDP-bound states.
e, Analytical gel filtration results of Mfn1ΔTME209A and Mfn1ΔTMR238A in nucleotide-free and GDP•AlF4−-bound states. Note that in the GDP•AlF4−-bound state, no peak at the exclusion volume is observed, indicating that both mutants do not oligomerize.
f, Structural comparison of Mfn1IM in different nucleotide-loading states at α2G. Note the distinct orientation of D189 in the GTP-bound state, and the uniformly oriented D193. D193 is a conserved residue that also faces the predicted HD2. Colour as in Fig. 4g.
g, Electron density of D189 and D193 on α2G in MfnIM structures contoured at 1.0σ. Note the difference in orientations of α2G in these structures as revealed by the density maps. Although the side chain of D189 is not fully traceable in some non-GTP-bound cases, their locations would differ from that in the GTP-bound form.
h, Mitochondrial elongation assay for the mutants in the plausible G domain-HD2 contact. For each construct, 100 cells were scored in biological triplicate; representative images are shown. Error bars indicate standard errors. Scale bar is 10 µm. Note the clumping mitochondria for Mfn1D189A and anticipated normal mitochondria for Mfn1D193A. R455, R460, Q473 and R594 are conserved residues in the predicted HD2 which were screened for contacting D189 based on sequence alignment of mitofusins and BDLP. Corresponding mutants increased mitochondrial fragmentation or aggregation. It seems that either they are not the right residues interacting with D189, or single point mutation was not sufficient to break the plausible interaction.
Extended Data Figure 9. Proposed model for Mfn1-mediated OMM fusion.
a, Model for nucleotide-regulated OMM fusion mediated by Mfn1. The G domain, HD1, predicted HD2 and TM are indicated in the upper left Mfn1 molecule, and coloured orange, green, grey and blue, respectively. During GTP hydrolysis, HD2s of tethered Mfn1 molecules may fold back via intrinsic mechanistic potential analogous to the BSE-stalk of MxA protein (Chen et al, our unpublished data) to bring opposing membrane in close proximity. Repeating tethering reactions by appropriate numbers of Mfn1 would promote docking of opposing OMMs, presumably as described in a recent in vitro electron cryo-tomography study where discrete electron densities representing yeast Fzo1 displayed a ring-like arrangement surrounding docked OMMs48. If this ‘docking ring’ exists in mammals, Mfn1 may contribute to its formation through hydrolysis-dependent in trans oligomerization (shown in c). Subsequent membrane merger may rely on local membrane curvature, as reported in many cellular events such as synaptic vesicle fusion and cell-to-cell fusion49, 50. As the space between docked OMMs (∼2 nm) is too small to accommodate Mfn1 molecules48, these molecules may gather at the rim of the docking site, resulting in a crowding effect that possibly generates bending on local OMMs to facilitate fusion51, 52.
b, Schematic drawing shows the GTP-loading-induced conformational rearrangement of Mfn1 HD1-HD2 region via the D189 trigger.
c, Possible organization of the plausible in trans cross oligomer of Mfn1 around the docking site. This process is dependent on GTP hydrolysis.
Extended Data Table 1.
Crystallographic data collection and refinement statistics
| Initial ligand State |
Mfn1IMB GMPPCP apo |
Mfn1IMB GMPPCP apo |
Mfn1IMCT109A GTP GTP-bound |
Mfn1IMC GDP•AIF4− Transition-like |
Mfn1IMCT109A GTP GDP-bound |
Mfn1IMA GTPyS GTPyS-bound |
Mfn1IMA GDP GDP-bound |
|---|---|---|---|---|---|---|---|
| Data collection | |||||||
| Data Set | Native | SeMet | Native | Native | Native | Native | Native |
| Space group | P212121 | P212121 | P212121 | C2 | P212121 | C2221 | P3121 |
| Cell dimensions | |||||||
| a, b, c(Å) | 51.8, 110.9, 112.4 | 51.6, 110.2, 111.3 | 70.6, 72.4, 95.3 | 104.1,46.0, 146.2 | 70.4, 72.9, 95.0 | 127.6, 143.1, 159.0 | 93.6,93.6, 114.1 |
| α, β, γ(°) | 90 | 90 | 90 | 92.2 | 90 | 90 | 120 |
| Wavelength (Å) | 0.91800 | 0.97915 | 0.97915 | 0.91800 | 0.97915 | 0.91800 | 0.91800 |
| Resolution (Å) | 47.0-2.2 (6.55-2.2) | 49.7-2.3 (6.84-2.3) | 44.7-1.6(4.79-1.6) | 48.7-2.8(8.30-2.8) | 44.7-1.8(5.37-1.8) | 47.7-6.1 (16.77-6.1) | 46.8-4.3(12.49-4.3) |
| Rsym* | 0.068 (0.438) | 0.060 (0.494) | 0.043 (0.583) | 0.066 (0.629) | 0.048 (0.517) | 0.0470 (0.522) | 0.064 (0.537) |
| I/σ(I) | 18.08(4.27) | 21.22 (4.14) | 25.54 (3.33) | 20.55 (3.02) | 25.08(4.12) | 19.28(3.32) | 17.81 (4.62) |
| Completeness (%) | 99.4 (98.1) | 99.5 (97.7) | 89.4 (99.7) | 99.1 (97.0) | 98.9 (98.8) | 98.3 (99.8) | 97.2 (95.0) |
| Redundancy | 7.9 (8.0) | 7.7 (7.5) | 7.1 (7.2) | 7.4 (7.4) | 7.9(8.1) | 5.4 (5.4) | 10.4(10.8) |
| Refinement | |||||||
| Resolution (Å) | 39.5-2.2 | 29.1-1.6 | 48.7-2.8 | 29.0-1.8 | |||
| No. reflections | 33374 | 57847 | 24264 | 45448 | |||
| Rwork/Rfree | 0.175/0.214 | 0.179/0.215 | 0.201/0.258 | 0.177/0.204 | |||
| No. atoms | |||||||
| Protein | 3291 | 3224 | 6484 | 3137 | |||
| Ligand/ion | 0 | 34 | 64 | 28 | |||
| Water | 184 | 427 | 4 | 273 | |||
| B-factors | |||||||
| Protein | 57.5 | 33.6 | 106.9 | 43.4 | |||
| Ligand/ion | 0 | 30.8 | 84.0 | 36.3 | |||
| Water | 55.3 | 43.2 | 68.9 | 48.3 | |||
| R.m.s. deviations | |||||||
| Bond lengths (Å) | 0.007 | 0.005 | 0.012 | 0.006 | |||
| Bond angles (°) | 0.753 | 0.812 | 1.471 | 0.763 |
Numbers in parentheses represent values from the highest resolution shell.
Acknowledgments
We thank the staff at beamline BL17U1 of SSRF for the help with the collection of diffraction data, Prof. Wen-Lin Huang and Prof. Hui-Yun Wang (Sun Yat-sen Univeristy, Guangzhou) for the technical assistance, Prof. Junjie Hu and Xiangyang Guo (Nankai Univeristy, Tianjin) for the advice on liposome tethering assay, and Prof. Oliver Daumke (Max-Delbrück-Centrum für Molekulare Medizin, Berlin) for the comments on the manuscript. This work was supported by grants of National Basic Research Program of China (2013CB910500), National Natural Science Foundation of China (31200553), Natural Science Foundation of Guangdong Province (2014TQ01R584 and 2014A030312015), New Century Excellent Talents in University (NCET-12-0567) and the Recruitment Program of Global Youth Experts to S.G., and the National Institutes of Health (GM110039 and GM119388) to D.C.C.
Footnotes
Author Contributions
S.G. and D.C.C. conceived the project. Y.-L.C. made the constructs, purified proteins, and performed crystallographic and biochemical experiments. S.M. carried out mitochondrial elongation assays. Y.C. did ITC measurements and helped with collection of X-ray diffraction data. J.-X.F., B.Y and Y.-J.L. did cloning and purification for some of the Mfn1IM mutants. D.-D.G. performed some of the SEC-RALS experiments, D.-D.G., J.-Y.Y. and S.L. helped with crystallization experiments. Y.-L.C. S.L. and S.G. solved the structures. Y.-L.C., D.C.C. and S.G. wrote the paper.
Author Information
The authors declare no competing financial interests. Correspondence and requests for materials should be addressed to S.G. (gaosong@sysucc.org.cn).
References
- 1.Shaw JM, Nunnari J. Mitochondrial dynamics and division in budding yeast. Trends in cell biology. 2002;12:178–184. doi: 10.1016/s0962-8924(01)02246-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Karbowski M, Youle RJ. Dynamics of mitochondrial morphology in healthy cells and during apoptosis. Cell death and differentiation. 2003;10:870–880. doi: 10.1038/sj.cdd.4401260. [DOI] [PubMed] [Google Scholar]
- 3.Youle RJ, van der Bliek AM. Mitochondrial fission, fusion, and stress. Science. 2012;337:1062–1065. doi: 10.1126/science.1219855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Mishra P, Chan DC. Metabolic regulation of mitochondrial dynamics. J Cell Biol. 2016;212:379–387. doi: 10.1083/jcb.201511036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Yaffe MP. The machinery of mitochondrial inheritance and behavior. Science. 1999;283:1493–1497. doi: 10.1126/science.283.5407.1493. [DOI] [PubMed] [Google Scholar]
- 6.Chan DC. Fusion and fission: interlinked processes critical for mitochondrial health. Annu Rev Genet. 2012;46:265–287. doi: 10.1146/annurev-genet-110410-132529. [DOI] [PubMed] [Google Scholar]
- 7.Nakada K, et al. Inter-mitochondrial complementation: Mitochondria-specific system preventing mice from expression of disease phenotypes by mutant mtDNA. Nature medicine. 2001;7:934–940. doi: 10.1038/90976. [DOI] [PubMed] [Google Scholar]
- 8.Friedman JR, Nunnari J. Mitochondrial form and function. Nature. 2014;505:335–343. doi: 10.1038/nature12985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Santel A, Fuller MT. Control of mitochondrial morphology by a human mitofusin. J Cell Sci. 2001;114:867–874. doi: 10.1242/jcs.114.5.867. [DOI] [PubMed] [Google Scholar]
- 10.Praefcke GJ, McMahon HT. The dynamin superfamily: universal membrane tubulation and fission molecules? Nature reviews. Molecular cell biology. 2004;5:133–147. doi: 10.1038/nrm1313. [DOI] [PubMed] [Google Scholar]
- 11.Rojo M, Legros F, Chateau D, Lombes A. Membrane topology and mitochondrial targeting of mitofusins, ubiquitous mammalian homologs of the transmembrane GTPase Fzo. J Cell Sci. 2002;115:1663–1674. doi: 10.1242/jcs.115.8.1663. [DOI] [PubMed] [Google Scholar]
- 12.Chen H, et al. Mitofusins Mfn1 and Mfn2 coordinately regulate mitochondrial fusion and are essential for embryonic development. J Cell Biol. 2003;160:189–200. doi: 10.1083/jcb.200211046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Koshiba T, et al. Structural basis of mitochondrial tethering by mitofusin complexes. Science. 2004;305:858–862. doi: 10.1126/science.1099793. [DOI] [PubMed] [Google Scholar]
- 14.Ranieri M, et al. Mitochondrial fusion proteins and human diseases. Neurol Res Int. 2013;2013:293893. doi: 10.1155/2013/293893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Daumke O, et al. Architectural and mechanistic insights into an EHD ATPase involved in membrane remodelling. Nature. 2007;449:923–927. doi: 10.1038/nature06173. [DOI] [PubMed] [Google Scholar]
- 16.Gao S, et al. Structure of myxovirus resistance protein a reveals intra- and intermolecular domain interactions required for the antiviral function. Immunity. 2011;35:514–525. doi: 10.1016/j.immuni.2011.07.012. [DOI] [PubMed] [Google Scholar]
- 17.Faelber K, et al. Crystal structure of nucleotide-free dynamin. Nature. 2011;477:556–560. doi: 10.1038/nature10369. [DOI] [PubMed] [Google Scholar]
- 18.Ford MG, Jenni S, Nunnari J. The crystal structure of dynamin. Nature. 2011;477:561–566. doi: 10.1038/nature10441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Byrnes LJ, Sondermann H. Structural basis for the nucleotide-dependent dimerization of the large G protein atlastin-1/SPG3A. Proc Natl Acad Sci U S A. 2011;108:2216–2221. doi: 10.1073/pnas.1012792108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bian X, et al. Structures of the atlastin GTPase provide insight into homotypic fusion of endoplasmic reticulum membranes. Proc Natl Acad Sci U S A. 2011;108:3976–3981. doi: 10.1073/pnas.1101643108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Low HH, Löwe J. A bacterial dynamin-like protein. Nature. 2006;444:766–769. doi: 10.1038/nature05312. [DOI] [PubMed] [Google Scholar]
- 22.Low HH, Sachse C, Amos LA, Löwe J. Structure of a bacterial dynamin-like protein lipid tube provides a mechanism for assembly and membrane curving. Cell. 2009;139:1342–1352. doi: 10.1016/j.cell.2009.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chappie JS, Acharya S, Leonard M, Schmid SL, Dyda F. G domain dimerization controls dynamin’s assembly-stimulated GTPase activity. Nature. 2010;465:435–440. doi: 10.1038/nature09032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Byrnes LJ, et al. Structural basis for conformational switching and GTP loading of the large G protein atlastin. EMBO J. 2013;32:369–384. doi: 10.1038/emboj.2012.353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rennie ML, McKelvie SA, Bulloch EM, Kingston RL. Transient dimerization of human MxA promotes GTP hydrolysis, resulting in a mechanical power stroke. Structure. 2014;22:1433–1445. doi: 10.1016/j.str.2014.08.015. [DOI] [PubMed] [Google Scholar]
- 26.Hoppins S, Nunnari J. The molecular mechanism of mitochondrial fusion. Biochimica et biophysica acta. 2009;1793:20–26. doi: 10.1016/j.bbamcr.2008.07.005. [DOI] [PubMed] [Google Scholar]
- 27.Ishihara N, Eura Y, Mihara K. Mitofusin 1 and 2 play distinct roles in mitochondrial fusion reactions via GTPase activity. J Cell Sci. 2004;117:6535–6546. doi: 10.1242/jcs.01565. [DOI] [PubMed] [Google Scholar]
- 28.Knott AB, Perkins G, Schwarzenbacher R, Bossy-Wetzel E. Mitochondrial fragmentation in neurodegeneration. Nature reviews. Neuroscience. 2008;9:505–518. doi: 10.1038/nrn2417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Franco A, et al. Correcting mitochondrial fusion by manipulating mitofusin conformations. Nature. 2016;540:74–79. doi: 10.1038/nature20156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Liu TY, et al. Cis and trans interactions between atlastin molecules during membrane fusion. Proc Natl Acad Sci U S A. 2015;112:E1851–E1860. doi: 10.1073/pnas.1504368112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kabsch W. Xds. Acta crystallographica. Section D, Biological crystallography. 2010;66:125–132. doi: 10.1107/S0907444909047337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sheldrick GM. A short history of SHELX. Acta crystallographica. Section A, Foundations of crystallography. 2008;64:112–122. doi: 10.1107/S0108767307043930. [DOI] [PubMed] [Google Scholar]
- 33.Pape T, Schneider TR. HKL2MAP: a graphical user interface for macromolecular phasing with SHELX programs. J Appl Cryst. 2004;37:843–844. [Google Scholar]
- 34.Vagin A, Teplyakov A. MOLREP: an automated program for molecular replacement. J Appl Cryst. 1997;30:1022–1025. [Google Scholar]
- 35.McCoy AJ, et al. Phaser crystallographic software. J Appl Cryst. 2007;40:658–674. doi: 10.1107/S0021889807021206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Emsley P, Cowtan K. Coot: model-building tools for molecular graphics. Acta crystallographica. Section D, Biological crystallography. 2004;60:2126–2132. doi: 10.1107/S0907444904019158. [DOI] [PubMed] [Google Scholar]
- 37.Murshudov GN, Vagin AA, Dodson EJ. Refinement of macromolecular structures by the maximum-likelihood method. Acta crystallographica. Section D, Biological crystallography. 1997;53:240–255. doi: 10.1107/S0907444996012255. [DOI] [PubMed] [Google Scholar]
- 38.Adams PD, et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta crystallographica. Section D, Biological crystallography. 2010;66:213–221. doi: 10.1107/S0907444909052925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Chen VB, et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta crystallographica. Section D, Biological crystallography. 2010;66:12–21. doi: 10.1107/S0907444909042073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.DeLano WL. The PyMOL Molecular Graphics System. 2002 < http://www.pymol.org.
- 41.Krissinel E, Henrick K. Inference of macromolecular assemblies from crystalline state. Journal of molecular biology. 2007;372:774–797. doi: 10.1016/j.jmb.2007.05.022. [DOI] [PubMed] [Google Scholar]
- 42.Laskowski RA, MacArthur MW, Moss DS, Thornton JM. PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Cryst. 1993;26:283–291. [Google Scholar]
- 43.Gao S, et al. Structural basis of oligomerization in the stalk region of dynamin-like MxA. Nature. 2010;465:502–506. doi: 10.1038/nature08972. [DOI] [PubMed] [Google Scholar]
- 44.Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic acids research. 1994;22:4673–4680. doi: 10.1093/nar/22.22.4673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJ. The Phyre2 web portal for protein modeling, prediction and analysis. Nature protocols. 2015;10:845–858. doi: 10.1038/nprot.2015.053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Prakash B, Praefcke GJ, Renault L, Wittinghofer A, Herrmann C. Structure of human guanylate-binding protein 1 representing a unique class of GTP-binding proteins. Nature. 2000;403:567–571. doi: 10.1038/35000617. [DOI] [PubMed] [Google Scholar]
- 47.Frohlich C, et al. Structural insights into oligomerization and mitochondrial remodelling of dynamin 1-like protein. EMBO J. 2013;32:1280–1292. doi: 10.1038/emboj.2013.74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Brandt T, Cavellini L, Kuhlbrandt W, Cohen MM. A mitofusin-dependent docking ring complex triggers mitochondrial fusion. eLife. 2016;5 doi: 10.7554/eLife.14618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.McMahon HT, Kozlov MM, Martens S. Membrane curvature in synaptic vesicle fusion and beyond. Cell. 2010;140:601–605. doi: 10.1016/j.cell.2010.02.017. [DOI] [PubMed] [Google Scholar]
- 50.Richard JP, et al. Intracellular curvature-generating proteins in cell-to-cell fusion. The Biochemical journal. 2011;440:185–193. doi: 10.1042/BJ20111243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Stachowiak JC, et al. Membrane bending by protein-protein crowding. Nature cell biology. 2012;14:944–949. doi: 10.1038/ncb2561. [DOI] [PubMed] [Google Scholar]
- 52.Kozlov MM, et al. Mechanisms shaping cell membranes. Current opinion in cell biology. 2014;29:53–60. doi: 10.1016/j.ceb.2014.03.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The X-ray crystallographic coordinates and structure factor files for Mfn1IM structures have been deposited in the Protein Data Bank (http://www.rcsb.org) under following accession numbers shown in the brackets: apo MfnIMB (5GO4), GTP-bound Mfn1IMCT109A (5GOF), transition-like state Mfn1IMC (5GOM), and GDP-bound Mfn1IMCT109A (5GOE). All other data generated or analysed during this study are included in this published article, and are available from the corresponding author upon reasonable request.