Summary
Background
The ability to generate defined null mutations in mice revolutionized the analysis of gene function in mammals. However, gene-deficient mice generated by using 129-derived embryonic stem cells may carry large segments of 129 DNA, even when extensively backcrossed to reference strains, such as C57BL/6J, and this may confound interpretation of experiments performed in these mice. Tissue plasminogen activator (tPA), encoded by the PLAT gene, is a fibrinolytic serine protease that is widely expressed in the brain. A number of neurological abnormalities have been reported in tPA-deficient mice.
Objectives
To study genetic contamination of tPA-deficient mice.
Materials and methods
Whole genome expression array analysis, RNAseq expression profiling, low- and high-density SNP analysis, bioinformatics, and genome editing was used to analyze gene expression in tPA-deficient mouse brains.
Results and conclusions
Genes differentially expressed in the brain of Plat−/− mice from two independent colonies highly backcrossed onto the C57BL/6J strain clustered near Plat on chromosome 8. SNP analysis attributed this anomaly to about 20 Mbp of DNA flanking Plat being of 129 origin in both strains. Bioinformatic analysis of these 129-derived chromosomal segments identified a significant number of mutations in genes co-segregating with the targeted Plat allele, including several potential null mutations. Using zinc finger nuclease technology, we generated novel “passenger mutation”-free isogenic C57BL/6J-Plat−/− and FVB/NJ-Plat−/− mouse strains by introducing an 11 bp deletion in the exon encoding the signal peptide. These novel mouse strains will be a useful community resource for further exploration of tPA function in physiological and pathological processes.
Keywords: brain, congenic mice, gene targeting, mutation, tissue plasminogen activator
Introduction
The development of methods to selectively disrupt genes in mice by homologous recombination in embryonic stem (ES) cells provided a potent new tool for analysis of gene function in mammals that has been of immeasurable value to a wide range of research fields [1]. Because germ line-competent ES cells were most successfully derived from various sub-strains of the 129 mouse strain, the large majority of gene disruptions initially were made in this strain. Due to the poor breeding characteristics, neurological and neuroanatomical abnormalities, and high tumor incidence of 129 mice, gene targeted mice made using 129 ES cells typically were backcrossed to inbred reference strains, such as C57BL/6J. However, due to the low probability of meiotic crossover occurring close to the targeted gene, a large region of DNA is likely to remain of 129 origin, even in “congenic” mice backcrossed for 10 generations (p = 0.91 for more than 1 centiMorgan of 129-derived DNA flanking each side of the targeted gene) [2]. Importantly in this respect, a recent direct comparison of the 129 and C57BL/6J genome sequences uncovered that the two strains differ by no less than 1395 deletions, insertions or nucleotide substitutions in protein coding regions, which resulted in 188 lost or gained stop codons, 875 frame shift variants, and 332 splice donor or acceptor variants [2]. This finding led the authors of the above study to conclude that essentially all constitutively gene-targeted mouse strains generated using 129-derived ES cells carry “passenger mutations” that confound data interpretation [2]. Indeed, several examples of erroneous attribution of observed phenotypes in gene-targeted mice generated using 129-derived ES cells have emerged recently [2-4]. As regards extracellular proteases, the reported resistance of matrix metalloproteinase (MMP)-13-deficient mice and, most likely, MMP-7-deficient and MMP-8-deficient mice, to LPS-induced lethality was demonstrated not to be a consequence of the loss of the MMP. Rather, it was caused by a null mutation in the neighboring Casp11 gene, which is present in 129 mice, but not in C57BL/6J mice [2]. A second issue with many ES cell-generated gene disrupted mice relates to the use of homologous recombination to generate the null mutation. This strategy often involves the deletion of large segments of DNA within the targeted gene that could result in the deletion of cis-acting regulatory elements and microRNAs, while the insertion of a powerful neomycin selection cassette could similarly modulate the expression of genes flanking the targeted locus.
Tissue plasminogen activator (tPA) is a fibrinolytic serine protease [5] that is widely expressed in the brain [6-10] where it is involved in a diverse array of physiological and pathophysiological processes unrelated to fibrinolysis that were identified in large part through the analysis of tPA-deficient mice [11-25].
During a transcriptomic profiling of genes differentially expressed in the brains of tPA-deficient mice extensively backcrossed to the C57BL/6J strain, we noted a highly significant overrepresentation of differentially expressed genes clustering around the Plat gene, encoding tPA, on chromosome 8. A similar anomalous clustering of differentially expressed genes was observed when we analyzed a second colony of tPA-deficient mice independently highly backcrossed to the C57BL/6J strain. Single nucleotide polymorphism (SNP) analysis provided evidence that at least 22 Mbp of DNA flanking Plat remained of 129 origin in each C57BL/6J-Plat−/− strain. Furthermore, bioinformatic analysis identified several mutations, including potential null mutations, in protein coding genes located on this chromosomal segment.
Using zinc finger nuclease (Zfn) technology, we engineered a null mutation in the Plat gene of C57BL/6J and FVB/N mice. We demonstrate that isogenic C57BL/6J-Plat−/− mice, free of 129 DNA, do not show anomalous clustering of differentially expressed genes in proximity to the targeted Plat gene. These novel isogenic Plat−/− mouse strains constitute a valuable community resource for further exploration of tPA functions in the brain and other tissues.
Materials and Methods
Mice
Mice were housed in standard barrier facilities under approved protocols.
Expression array analysis
12 to 14 week-old male “Melbourne” C57BL/6J-Plat−/− and age-matched C57BL/6J mice (Jackson Laboratory) were transcardially transfused with ice-cold PBS (8 per genotype). The hippocampus was dissected out and placed in “RNA later”. RNA was extracted using the “RNeasy Lipid Tissue” kit (Qiagen) according to manufacturer’s instructions. RNA was then pooled from pairs of mice to give 4 samples from each genotype for subsequent analysis. The extracted RNA was quality controlled, hybridized to Illumina microarray chips and analyzed by the SRC MicroRNA facility (University of Queensland, Australia) using MouseWG-6 v2.0 Expression BeadChip (Catalog ID: BD-201-0202). Genes were considered differentially expressed if the adjusted p value was less than 0.05. The original high-throughput microarray data is available through the following links: https://cloudstor.aarnet.edu.au/plus/index.php/s/PYxAHTIkijH4zYy and https://cloudstor.aarnet.edu.au/plus/index.php/s/jPAXctPNeiIXKvQ.
RNAseq
Six-week-old “Michigan” C57BL/6J-Plat−/− and “Bethesda” C57BL/6J-Plat−/− littermates (four mice per genotype), and age-matched wildtype C57BL/6J control mice (four mice per genotype) were euthanized by CO2 inhalation. The brains were dissected out, the olfactory bulb and cerebellum removed, and the brains snap frozen in liquid nitrogen. cDNA library preparation and Illumina high throughput sequencing was performed by The University of Michigan's DNA Sequencing Core. Bioinformatic analysis of these data was conducted by the University of Michigan Bioinformatics Core. All data from these studies has been deposited in the Gene Expression Omnibus (GEO) database with accession number, GSE76093. Genes were considered differentially expressed if presenting with an adjusted p value (q-value) of 0.05001 or less and fold-difference in expression of more than 0.3. Genes located on the Y chromosome were excluded from the analysis because the analyzed mice included both males and females.
SNP analysis
Low-density SNP analysis of the entire mouse genome was performed by the Jackson Laboratory using the “JAX 150 SNP Panel” polymorphic between C57BL/6J and 129 (Supporting table S1). The panel consist of 154 SNPs that cover the 19 autosomes and the X chromosome with a density of ~15-20 Mbp/SNP. High density SNP analysis of mouse chromosome 8 was performed by the Jackson Laboratory using a chromosome 8-specific panel of 34 SNPs, polymorphic between C57BL/6J and 129, covering the length of the chromosome with a density of ~3.5 Mbp/SNP (Supporting table S2).
Generation of Plat−/− mice using Zfns
A pair of Zinc finger-FokI fragment fusion proteins (Zfns) binding, respectively, 5’-GACTGGCTTTCCCAT-3’ (nucleotides 9399-9413 of the Plat gene, NC_000074.6) and 5’-GACCAGGTGGGT’3’ (nucleotides 9419-9430 of the Plat gene) were generated by Sigma. mRNA encoding the Zfn pair was generated by in vitro transcription and microinjected into the male pronucleus of C57BL/6J and FVB/NJ zygotes, which were implanted into pseudopregnant mice. The ensuing founders were screened by PCR using the primer pair 5’-AAGAGTCATTGCTGGATGGG-3’ (forward) and 5’-GATCACTCCTGGGAACGTGT-3’ (reverse), which amplifies the 325 bp fragment constituting nt. 9221-9545 of the Plat gene. Founders positive for mutations in the signal peptide-encoding exon 2 of Plat were further characterized by DNA sequencing and screened for germ line transmission by breeding to C57BL/6J or FVB/NJ.
tPA Western blot analysis
Non-reduced and non-boiled protein lysates from brains were separated by SDS-PAGE and transferred to PVDF membranes. The membranes were blocked with 5% non-fat dry milk for 1 h. The blot was incubated with 2 μg/ml rabbit anti-human tPA (Molecular Innovation, ASHTPA-GF) – for 1 h at room temperature. Bound antibody was visualized by incubation for 1 h with donkey anti-rabbit-HRP (Jackson Immuno Research, 711-036-152) 1:5000.
Plasminogen-casein zymography
Plasminogen-casein zymography was performed as described [26]. Lysis zones were visualized by staining of gels with Coomassie brilliant blue.
tPA activity assay
Active tPA was determined using a commercial sandwich tPA capture ELISA kit (Molecular Innovations, NTBIOCPAI) as recommended by the manufacturer.
Statistical analysis
Distribution of differentially expressed genes over all chromosomes: We used a chi-square statistic with a reference distribution calculated using simulation (since many of the expected hit counts are very small). Specifically, the expected number of “hits” per chromosome, assuming uniform distribution, was calculated and compared to the observed number of hits per chromosome. This comparison utilized a chi-square statistic (but not a chi-square reference distribution). To obtain a reference distribution we generated Poisson counts from 100,000 randomizations with mean values equal to the expected number of hits per chromosome under uniformity, accounting for the differing lengths of the chromosomes.
Positions of differentially expressed genes on chromosome 8: To assess the probability of the observed hits on chromosome 8 being random, we divided the chromosome into 1 Mbp intervals and counted the number of hits per interval. We then used a chi-square statistic to compare this distribution of hit positions to what would be expected under a uniform distribution in which the hits were equally likely to occur at any point along chromosome 8. The reference distribution was calculated using simulation, using a uniform distribution to define random positions for hits along the length of chromosome 8.
Bioinformatic analysis
Bioinformatic analysis of “passenger mutations” co-segregating with the targeted Plat allele were identified using the Sanger “Mouse Genomes Project- - Query SNPs, indels or SVs” website (http://www.sanger.ac.uk/sanger/Mouse_SnpViewer/rel-1505) [27, 28].
Results
Transcriptomic profiling of brains of two independently generated strains of congenic C57BL/6J-Plat−/− mice reveals anomalous clustering of differentially expressed genes around Plat on chromosome 8
C57BL/6J-Plattm1Mlg mice [29] were housed in the Australian Centre for Blood Diseases, Monash University (hereafter termed Melbourne C57BL/6J-Plat−/− mice). In 2009, when the expression analysis was carried out, these mice were estimated to be backcrossed to C57BL/6J mice for 13 generations. RNA was isolated from the brains of Melbourne C57BL/6J-Plat−/− mice and from co-habitating age- and sex-matched C57BL/6J mice and hybridized to whole genome expression arrays. Quadruplicate analysis was performed, with each sample representing pools of RNA from two mice. Overall, only 13 genes reporting in the array were identified as differentially expressed in the brain of C57BL/6J-Plat−/− mice, when compared to the C57BL/6J control mice (Table 1). Surprisingly, 11 of these 13 genes (85 %) were located on chromosome 8, which also harbors the Plat gene (P<0.000001, see materials and methods for details on statistics). Furthermore, all of these 11 genes clustered around the Plat gene, on a chromosomal segment spanning approximately 19 Mbp upstream to 8 Mbp downstream of Plat (Fig. 1A). This clustering of differentially expressed genes within this 27 Mbp segment is also highly anomalous (P<0.000001), considering the total length of mouse chromosome 8 is approximately 129 Mbp and the total length of the mouse genome is more than 2600 Mbp.
Table 1.
Chromosomal location of genes differentially expressed in Melbourne C57BL/6J-Plat−/− mice.
| Chromosome | Length (Mbp) |
# Differentially expressed genes |
|---|---|---|
| 1 | 195 | 2 |
| 2 | 182 | 0 |
| 3 | 160 | 0 |
| 4 | 157 | 0 |
| 5 | 152 | 0 |
| 6 | 150 | 0 |
| 7 | 145 | 0 |
| 8 | 129 | 11 |
| 9 | 125 | 0 |
| 10 | 131 | 0 |
| 11 | 122 | 0 |
| 12 | 120 | 0 |
| 13 | 120 | 0 |
| 14 | 125 | 0 |
| 15 | 104 | 0 |
| 16 | 91 | 0 |
| 17 | 95 | 0 |
| 18 | 91 | 0 |
| 19 | 61 | 0 |
| X | 161 | 0 |
| Y | 95 | 0 |
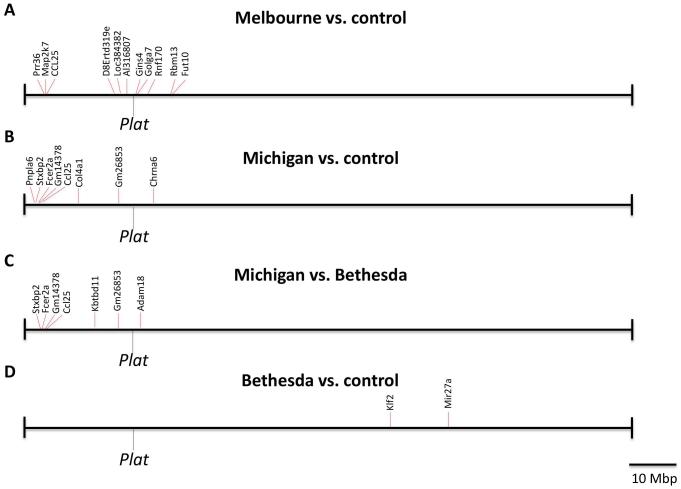
Fig. 1.
The clustering of genes differentially expressed in brains of congenic tPA-deficient mice in the vicinity of Plat is unrelated to the loss of tPA. (A) RNA was extracted from congenic “Melbourne” C57BL/6J-Plat−/− mice and age and gender-matched C57BL/6J wildtype controls, and whole-genome transcriptome analysis was performed by using expression arrays. The location of Plat and of differentially expressed genes on chromosome 8 are shown. 11 of 13 differentially expressed genes (excluding Plat) in tPA-deficient brains (85 %) are located within a 27 Mbp interval, flanking Plat. (B) RNA was extracted from congenic “Michigan” C57BL/6J-Plat−/− mice and wildtype littermates and whole-genome transcriptome analysis was performed by RNAseq. The location of Plat and of differentially expressed genes on chromosome 8 are shown. Eight of 32 differentially expressed genes (excluding Plat) in tPA-deficient brains (25%) are located within a 30 Mbp interval flanking Plat. (C). RNA was extracted from “Michigan” C57BL/6J-Plat−/− mice and “Bethesda” C57BL/6J-Plat−/− littermates and whole-genome transcriptome analysis was performed by RNAseq. The location of Plat and of differentially expressed genes on chromosome 8 are shown. 7 of 19 differentially expressed genes (excluding Plat) (37%) are located within a 26 Mbp interval flanking Plat. (D) No anomalous clustering of differentially expressed genes in brains of isogenic “Bethesda” C57BL/6J-Plat−/− mice around Plat on chromosome 8. RNA was extracted from “Bethesda” C57BL/6J-Plat−/− mice and wildtype littermates and whole-genome transcriptome analysis was performed by RNAseq. The location of Plat and of differentially expressed genes on chromosome 8 are shown.
We next set out to examine if the abnormal distribution of differentially expressed genes in Melbourne C57BL/6J-Plat−/− mice was also observed in other congenic colonies of tPA-deficient mice. Specifically, we analyzed C57BL/6J-Plattm1Mlg mice housed at the University of Michigan (hereafter termed Michigan C57BL/6J-Plat−/− mice), estimated to be backcrossed for >10 generations to C57BL/6J in 2015 when the analysis was carried out. Brains of four Michigan C57BL/6J-Plat−/− mice and four wildtype C57BL/6J mice from Jackson Laboratories were subjected to transcriptomic profiling by RNAseq. We again observed an abnormally high frequency of differentially expressed genes (8 of 32, 25%) located on chromosome 8 (P<0.003) (Table 2). Furthermore, all eight genes were located approximately 18 Mbp upstream to 5 Mbp downstream of Plat (P<0.000001) (Fig. 1B).
Table 2.
Chromosomal location of genes differentially expressed in Michigan C57BL/6J-Plat−/− mice.
| Chromosome | Length (Mbp) |
# Differentially expressed genes |
|---|---|---|
| 1 | 195 | 0 |
| 2 | 182 | 1 |
| 3 | 160 | 0 |
| 4 | 157 | 1 |
| 5 | 152 | 2 |
| 6 | 150 | 0 |
| 7 | 145 | 2 |
| 8 | 129 | 8 |
| 9 | 125 | 3 |
| 10 | 131 | 3 |
| 11 | 122 | 3 |
| 12 | 120 | 0 |
| 13 | 120 | 2 |
| 14 | 125 | 1 |
| 15 | 104 | 0 |
| 16 | 91 | 0 |
| 17 | 95 | 3 |
| 18 | 91 | 0 |
| 19 | 61 | 3 |
| X | 161 | 0 |
Evidence for 129 origin of differentially expressed genes on chromosome 8 of tPA-deficient mice
To explore the basis of the anomalous clustering of differentially expressed genes around the Plat gene in the two independently backcrossed congenic strains of C57BL/6J-Plat−/− mice, we next contracted Jackson Laboratories to perform SNP analysis of Michigan C57BL/6J-Plat−/− mice and Melbourne C57BL/6J-Plat−/− mice. We first employed a JAX 150 SNP Panel polymorphic between C57BL/6J and 129, which covers the 19 autosomes and the X chromosome with a density of ~15-20 Mbp. Strikingly, of 154 SNPs analyzed in Michigan C57BL/6J-Plat−/− mice, only one was found to be 129-specific (rs3701395, Supporting table S1) and this SNP was located 19.7 Mbp upstream of Plat, just upstream of a cluster of five differentially expressed genes. This SNP was also 129-specific in Melbourne C57BL/6J-Plat−/− mice, as was the adjacent SNP (rs3701395 and rs3684251, Supporting table S1). Melbourne C57BL/6J-Plat−/− mice were also heterozygote for one chromosome 1 SNP (rs3697376, Supporting table S1). Remarkably, this SNP was located very close to both the chromosome 1-located genes differentially expressed in Melbourne C57BL/6J-Plat−/− mice (chromosome location-bp, rs3697376: 65042402. C430010P07Rik: 66719248-66817562. Mtap2: 66175273-66442583). Based on these findings, we next performed a high-density SNP analysis of chromosome 8, employing 34 SNPs, polymorphic between C57BL/6J and 129 (Fig. 2A and B and Supporting table S2). In Michigan C57BL/6J-Plat−/− mice, a continuous cluster of 7 SNPs analyzed, located 19-0.3 Mbp upstream of Plat were identified as being 129-specific (Fig. 2A). The identical cluster of 129-derived SNPs were found in Melbourne C57BL/6J-Plat−/− mice, which also displayed two additional 129 SNPs located just downstream of Plat (Fig. 2B). Notably, the chromosomal segments identified in the SNP analysis as being 129-derived in the two mouse strains harbored most of the genes differentially expressed on chromosome 8 of the Michigan C57BL/6J-Plat−/− mice and all of the identified genes differentially expressed in the Melbourne C57BL/6J-Plat−/− mice (Fig. 2A and B, and Supporting table S1).
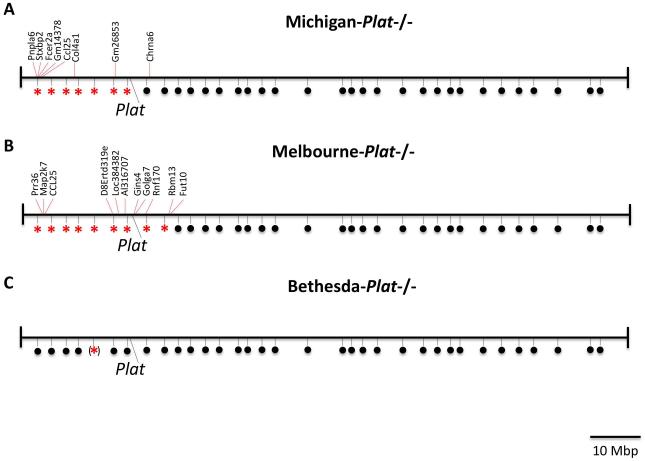
Fig. 2.
Congenic tPA-deficient mice contain large contiguous segments of 129-derived DNA. High density SNP analysis of congenic “Michigan” C57BL/6J-Plat−/− mice (A), congenic “Melbourne” C57BL/6J-Plat−/− mice (B) and isogenic “Bethesda” C57BL/6J-Plat−/− mice (C). Position of SNPs specific for strain 129 mice versus C57BL/6J mice are indicated with red asterisks, and the position of SNPs specific for strain C57BL/6J mice versus strain 129 mice are indicated with black circles.
Bioinformatic analysis identifies multiple passenger mutations in congenic C57BL/6J-Plat−/− mice
Having located the specific regions of 129-derived DNA in Michigan C57BL/6J-Plat−/− and Melbourne C57BL/6J-Plat−/− mice allowed for the direct bioinformatic identification of “passenger mutations” co-segregating with the targeted Plat allele by using the Sanger “Mouse Genomes Project- - Query SNPs, indels or SVs” website (http://www.sanger.ac.uk/sanger/Mouse_SnpViewer/rel-1505) [27, 28]. Interestingly, the 129-derived regions of both Michigan C57BL/6J-Plat−/− mice and Melbourne C57BL/6J-Plat−/− mice harbored five protein-coding genes (Cd209a, Cd209b, Ccl25, Defb34, Defb46) predicted to be null in 129 mice and wildtype in C57BL/6J mice or vice versa, as evidenced by the presence of in frame stop codons, frame shift-inducing insertions and deletions (Table 5). An additional 8 genes (Arhgef18, Cd209e, BC068157, Gm560, Tubgcp3, Mcf2l, Myom2, and Mchp1) harbored one or more mutations causing non-conservative amino acid substitutions. Owing to additional 129-derived regions upstream of Plat, Melbourne C57BL/6J-Plat−/− mice also displayed mutations in Adam3.
Table 5.
Passenger mutations associated with 129 DNA regions in “Michigan” and “Melbourne” Plat−/− strains
| Gene | Position on Chr.8 (Mbp) |
Distance from Plat (Mbp) |
Coding region mutations in 129 vs C57BL/6J | Mutated in Michigan |
Mutated in Melbourne |
||
|---|---|---|---|---|---|---|---|
| Stop gain/loss | Frameshift | Missense | |||||
| Arhgef18 | 3.40 | 19.30 | T315M; S975P | + | + | ||
| Cd209a | 3.74 | 18.96 | after C64 | R13G; T36P | + | + | |
| Cd209e | 3.85 | 18.85 | W208Stop | + | + | ||
| Cd209b | 3.92 | 18.78 | R247Stop | C42S; I259V A175V; T369A; V389M; E428G; V439I; R524Q; P558L; T559A; S620P; P690S; |
+ | + | |
| BC068157 | 4.21 | 18.49 | F716L; P779T; S950A S82P; W86R; H116Y; E120K; P154S; |
+ | + | ||
| Ccl25 | 4.34 | 18.36 | after P98 | M169I | + | + | |
| Tubgcp3 | 12.64 | 10.06 | G190S | + | + | ||
| Mcf2l | 12.95 | 9.75 | Stop1102L | + | + | ||
| Myom2 | 15.10 | 7.60 | R123Q; M131L; S234P; L977I; S1311N A300V; A303V; S469L; C580F; T615A; |
+ | + | ||
| Mcph | 18.60 | 4.10 | T615K | + | + | ||
| Defb34 | 19.12 | 3.58 | after D25 | + | + | ||
| Defb46 | 19.24 | 3.46 | C32 to Ter | C21F | + | + | |
| ENSMUSG00000097230 | 19.79 | 2.91 | + | + | |||
| AY761185 | 20.94 | 1.76 | + | + | |||
| Plat | 22.70 | 0 | N/A | N/A | |||
| Adam3 | 24.70 | 2.00 | Stop823S | G741D | + | ||
C57BL/6J-Plat−/− mice, generated by zinc finger nuclease targeting of C57BL/6J mice, do not show anomalous clustering of differentially expressed genes around Plat
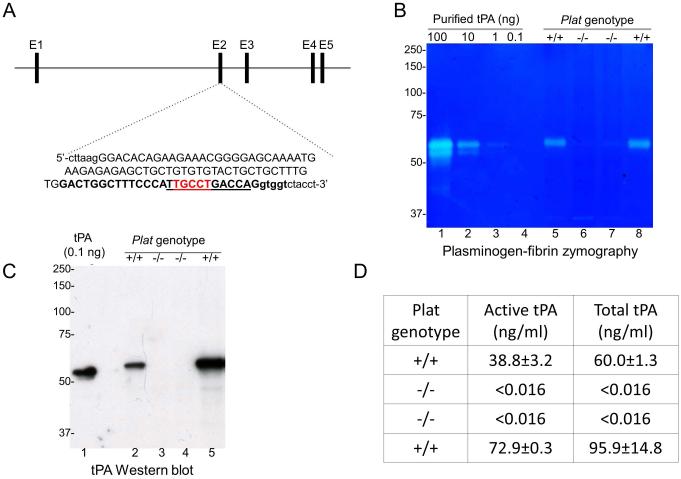
We next used custom-designed zinc finger nucleases (Zfns) to generate tPA-deficient mice de novo. A Zfn was engineered to introduce mutations into the signal peptide-encoding exon 2 of Plat (Fig. 3). mRNA encoding the Zfn was microinjected into fertilized C57BL/6J and FVB/NJ embryos that were implanted into pseudo-pregnant females. Analysis of born offspring identified a number of mice with mutations in exon 2 of Plat. This included C57BL/6J and FVB/NJ offspring with an identical 11 bp deletion in signal peptide-coding exon 2, which were selected for further analysis. This deletion removes nucleotides 52-62 of the Plat open reading frame and introduces a frameshift mutation after Pro17 of the tPA signal peptide and a premature stop signal (Fig. 3A). This gave rise to a null allele, as confirmed by plasminogen-casein zymography (Fig. 3B), Western blot (Fig. 3C), and tPA activity assays (Fig. 3D) of brain tissue from mice bred to homozygosity for the targeted allele and their wildtype littermates. These novel mouse strains hereafter are termed, respectively, Bethesda C57BL/6J-Plat−/− and Bethesda FVB/NJ-Plat−/−. As expected, low density SNP analysis of Bethesda C57BL/6J-Plat−/− mice showed all SNPs to be C57BL/6J-derived (Supporting table S1). High density SNP analysis of chromosome 8 of Bethesda C57BL/6J-Plat−/− mice (Supporting table S2 and Fig. 2) showed all SNPs, but one, to be C57BL/6J-specific. The single 129-specific SNP (rs3709624, Supporting table S2, indicated by red asterisk in Fig. 2) also genotyped as 129-specific in two C57BL/6J control mice purchased from Jackson Laboratories that were included in the analysis (“Control”, columns L and M in Supporting table S2), and we currently have no explanation for this finding.
Fig. 3.
Generation of isogenic C57BL/6J-Plat−/− and FVB/NJ-Plat−/− mice. (A) De novo generation of Plat null allele. Schematic structure of the proximal part of the mouse Plat gene (top), and the sequence of the signal peptide-encoding exon 2 (upper case letters) and flanking intron sequences (lower case letters) (bottom). The Zfn binding sites are in bold letters, with the Zfn cleavage site in red. The 11 bp deleted in the C57BL/6J and FVB/NJ strains carried forward for analysis are underlined. This deletion introduces a frameshift resulting in the production of a mRNA encoding amino acid 1-17 of tPA fused to a 19 amino acid nonsense peptide. B. Plasminogen-casein zymography. Lanes 1-4; purified human tPA. Lanes 5-8; protein extracts from brains of a litter of FVB/NJ mice containing 2 wildtype (lanes 5 and 8) and 2 Plat−/− littermates (lanes 6 and 7). C. tPA Western blot. Lanes 1; purified human tPA. Lanes 2-5; protein extracts from brains of a litter of mice containing 2 wildtype (lanes 2 and 5) and 2 Plat−/− littermates (lanes 3 and 4). D. Active tPA (middle column) and total tPA (right column) protein extracts from brains of a litter of mice containing 2 wildtype (top and bottom row) and 2 Plat−/− littermates (middle rows).
We next set out to provide direct experimental evidence that the differential expression of chromosome 8 genes in brains of Melbourne and Michigan C57BL/6J-Plat−/− mice were not a consequence of loss of tPA, but rather due to retention of a 129-derived chromosomal segment. For this purpose, we interbred the new Bethesda C57BL/6J-Plat−/− mice with Michigan C57BL/6J-Plat−/− mice to generate littermate pairs of Bethesda C57BL/6J-Plat−/− and Michigan C57BL/6J-Plat−/− mice. These mice, both devoid of tPA, were then subjected to transcriptomic profiling by RNAseq using quadruplicate analysis. A conspicuous clustering of genes differentially expressed in Bethesda C57BL/6J-Plat−/− mice and Michigan C57BL/6J-Plat−/− littermates around chromosome 8 was again observed, with 7 of 19 differentially expressed genes (37%) located on chromosome 8 (P<0.05), clustering 19 Mbp upstream to 2.9 Mbp downstream from Plat (P<0.000001) (Fig. 1C and Table 3). In sharp contrast, when the identical RNAseq analysis was performed comparing brains from Bethesda C57BL/6J-Plat−/− mice with their wildtype littermates, no overrepresentation of differentially expressed genes on chromosome 8 was found, with only 2 of 37 differentially expressed genes (5%) residing on chromosome 8 (P=N.S.), located, respectively, 50 and 61 Mbp from Plat (Fig. 1D and Table 4) (P=N.S.).
Table 3.
Chromosomal location of genes differentially expressed in Michigan and Bethesda C57BL/6J-Plat−/− mice.
| Chromosome | Length (Mbp) |
# Differentially expressed genes |
|---|---|---|
| 1 | 195 | 2 |
| 2 | 182 | 2 |
| 3 | 160 | 0 |
| 4 | 157 | 0 |
| 5 | 152 | 0 |
| 6 | 150 | 1 |
| 7 | 145 | 2 |
| 8 | 129 | 7 |
| 9 | 125 | 0 |
| 10 | 131 | 1 |
| 11 | 122 | 1 |
| 12 | 120 | 1 |
| 13 | 120 | 1 |
| 14 | 125 | 0 |
| 15 | 104 | 0 |
| 16 | 91 | 0 |
| 17 | 95 | 1 |
| 18 | 91 | 0 |
| 19 | 61 | 0 |
| X | 161 | 0 |
| Y | 95 | N.D. |
Table 4.
Chromosomal location of genes differentially expressed in Bethesda C57BL/6J-Plat−/− mice.
| Chromosome | Length (Mbp) |
Differentially expressed genes |
|---|---|---|
| 1 | 195 | 1 |
| 2 | 182 | 2 |
| 3 | 160 | 1 |
| 4 | 157 | 2 |
| 5 | 152 | 2 |
| 6 | 150 | 0 |
| 7 | 145 | 4 |
| 8 | 129 | 2 |
| 9 | 125 | 2 |
| 10 | 131 | 4 |
| 11 | 122 | 3 |
| 12 | 120 | 1 |
| 13 | 120 | 3 |
| 14 | 125 | 1 |
| 15 | 104 | 4 |
| 16 | 91 | 0 |
| 17 | 95 | 2 |
| 18 | 91 | 0 |
| 19 | 61 | 2 |
| X | 161 | 1 |
| Y | 95 | N.D. |
Discussion
This current study provides evidence that tPA-deficient mice highly backcrossed to C57BL/6J display differences in gene expression in the brain that are not related to the absence of tPA, but rather to way the Plat gene was targeted. Two colonies of highly backcrossed of C57BL/6J-Plat−/− mice, made by targeting Plat 129-derived ES cells, showed the same anomalous clustering of differentially expressed genes in the vicinity of Plat, when independently analyzed by whole genome array analysis and RNAseq. In the two analysis, respectively, 86 and 25 % of all differentially expressed genes were located within a segment of chromosome 8 roughly 19 Mbp upstream to 8 Mbp downstream of Plat. Considering the length of the mouse genome of more than 2600 Mbp, less than 1.5 % of differentially expressed genes would be expected to be located on this short segment of chromosome 8, which constitutes just 1.03 % of the mouse genome. Furthermore, an isogenic C57BL/6J-Plat−/− mouse strain, generated in this study by Zfn-mediated genome editing in C57BL/6J mice, did not show this anomalous clustering of differentially expressed genes. High-density SNP analysis showed that both Melbourne and Michigan C57BL/6J-Plat−/− mice contained a surprisingly large contiguous 129-derived chromosomal segment, and we provide direct experimental evidence that the presence of this chromosomal segment is responsible for the anomalous clustering of differentially expressed genes in C57BL/6J-Plat−/− mice. Importantly, the 129-derived chromosomal segment that co-segregates with the targeted Plat allele displayed a number of differences in coding regions, when compared to the corresponding chromosomal segment of C57BL/6J mice, with at least four protein-coding genes potentially being null in the 129 mice and wildtype in C57BL/6J mice or vice versa, and eight additional genes displaying non-conservative amino acid substitutions. Notably, some of these genes have reported roles in human brain function: ARHGEF18 is involved in neurite retraction [30] and was linked in a genome-wide association study to sexual dysfunction in individuals treated for major depression [31]. MCF2L participates in the formation and stabilization of glutaminergic synapses of cortical neurons [32], and SNPs in MCF2L and TUBGCP3 were identified as predictive for successful smoking cessation [33]. Homozygosity for mutations in MCPH1 causes microcephaly [34], while homozygous loss of ADAM3A is associated with pediatric high-grade glioma and diffuse intrinsic pontine gliomas [35].
It should be noted that the 129-derived co-segregating chromosomal segments identified in the congenic C57BL/6J-Plat−/− mouse strains have the potential to confound data interpretation irrespectively of whether the wildtype mice used as controls are littermate-derived or age- and sex-matched C57BL/6J mice. Our study, thus, highlights the importance of reconstitution experiments in non-isogenic mice, where a wildtype phenotype can be restored via the introduction of exogenous tPA into tPA-deficient mice [11], or where tPA-overexpressing mice exhibit an opposite phenotype of that of tPA-deficient mice [36]. Similarly, with regards to plasmin-dependent functions of tPA, one can eliminate the confounding effects of co-inherited passenger mutations when tPA-deficient and plasminogen-deficient mice exhibit similar phenotypes [37]. It is important to stress, however, that the present study provides no evidence, directly or indirectly, that any of the published phenotypes in tPA-deficient mice is caused by 129-derived passenger mutations co-inherited with the targeted Plat allele. Nonetheless, it is also clear from the data presented in the current study that caution must be exerted when interpreting phenotypes observed in C57BL/6J-Plat−/− mice. Related to this, it is interesting to note that this region of chromosome 8, which is syntenic with human chromosome 8, has been suggested to be a potential hub for genes associated with neuropsychiatric disorders and with cancer [38]. Undoubtedly, the C57BL/6J-Plat−/− and FVB/NJ-Plat−/− mice generated here should provide two novel mouse strains of use for the research community for further exploration of tPA function that are free of co-inherited passenger mutations inevitable when using 129-derived ES cells. Nevertheless, the Zfn technology used to generate the two mouse strains is relatively new, and the degree to which “off-targeting” of the germ line occurs is not fully investigated [39, 40]. Potential off-target mutations in the two novel mouse strains, however, would only constitute a potential long-term problem if located in proximity to Plat.
It should also be stressed that in conditionally-targeted strains, the potential of passenger mutations to confound data interpretation can largely be eliminated by the use of appropriate littermate controls, including Cre-negative littermates that carry two conditionally targeted alleles and, therefore, are expected to carry the same set of passenger mutations near the gene of interest [2].
The plasminogen activation system was among the first proteolytic systems to be analyzed by gene targeting in mice. Consequently, targeting of genes encoding most components of the system was done using 129-derived ES cells. This includes, besides Plat, Plau[29], Plaur[41, 42], Plg[43, 44], Serpine1[45], SerpinF2[46], and Annat2[47]. In light of the speed, efficiency, and low cost of current genome editing technologies, the regeneration of isogenic strains carrying null mutations in these genes should be considered.
Supplementary Material
Essentials.
C57BL/6J-tPA-deficient mice are widely used to study tissue plasminogen activator (tPA) function.
Congenic C57BL/6J-tPA-deficient mice harbor large 129-derived chromosomal segments.
The 129-derived chromosomal segments contain gene mutations that may confound data interpretation.
“Passenger mutation”-free isogenic tPA-deficient mice were generated for exploration of tPA function.
Acknowledgements
We thank Kris Mann and Mark Warnock for excellent technical assistance, Dr. Richard McEachin the University of Michigan the Bioinformatics Core for Bioinformatics analysis of the RNAseq data, Dr. Kerby Shedden of the University of Michigan Center for Statistical Consultation and Research (CSCAR) for statistical analysis, and Dr. Mary Jo Danton for critically reviewing this manuscript. Supported by the NIDCR Intramural Research Program (T.H.B), by National Institutes of Health Grants HL055374 and NS079639 (D. A. L.). Bethesda C57BL/6J-Plat−/− mice and Bethesda FVB/NJ-Plat−/− mice can be obtained by contacting Thomas Bugge (Thomas.bugge@nih.gov).
Footnotes
Addendum
A. L. Samson, D. A. Lawrence, T. H. Bugge, R. L. Medcalf, and R. Szabo conceived and designed experiments: D. A. Lawrence and R. Szabo generated and analyzed Plat−/− mice. A. L. Samson, T. H. Bugge, R. L. Medcalf, and R. Szabo performed array analysis and analyzed data: A. L. Samson, D. A. Lawrence, and T. H. Bugge performed RNAseq and analyzed data: T. H. Bugge wrote paper.
References
- 1.Bronson SK, Smithies O. Altering mice by homologous recombination using embryonic stem cells. The Journal of biological chemistry. 1994;269:27155–8. [PubMed] [Google Scholar]
- 2.Vanden Berghe T, Hulpiau P, Martens L, Vandenbroucke RE, Van Wonterghem E, Perry SW, Bruggeman I, Divert T, Choi SM, Vuylsteke M, Shestopalov VI, Libert C, Vandenabeele P. Passenger Mutations Confound Interpretation of All Genetically Modified Congenic Mice. Immunity. 2015;43:200–9. doi: 10.1016/j.immuni.2015.06.011. 10.1016/j.immuni.2015.06.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kayagaki N, Warming S, Lamkanfi M, Vande Walle L, Louie S, Dong J, Newton K, Qu Y, Liu J, Heldens S, Zhang J, Lee WP, Roose-Girma M, Dixit VM. Non-canonical inflammasome activation targets caspase-11. Nature. 2011;479:117–21. doi: 10.1038/nature10558. 10.1038/nature10558. [DOI] [PubMed] [Google Scholar]
- 4.Kenneth NS, Younger JM, Hughes ED, Marcotte D, Barker PA, Saunders TL, Duckett CS. An inactivating caspase 11 passenger mutation originating from the 129 murine strain in mice targeted for c-IAP1. The Biochemical journal. 2012;443:355–9. doi: 10.1042/BJ20120249. 10.1042/BJ20120249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Collen D, Lijnen HR. The fibrinolytic system in man. Crit Rev Oncol Hematol. 1986;4:249–301. doi: 10.1016/s1040-8428(86)80014-2. [DOI] [PubMed] [Google Scholar]
- 6.Soreq H, Miskin R. Plasminogen activator in the rodent brain. Brain research. 1981;216:361–74. doi: 10.1016/0006-8993(81)90138-4. [DOI] [PubMed] [Google Scholar]
- 7.Krystosek A, Seeds NW. Plasminogen activator secretion by granule neurons in cultures of developing cerebellum. Proc Natl Acad Sci U S A. 1981;78:7810–4. doi: 10.1073/pnas.78.12.7810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Qian Z, Gilbert ME, Colicos MA, Kandel ER, Kuhl D. Tissue-plasminogen activator is induced as an immediate-early gene during seizure, kindling and long-term potentiation. Nature. 1993;361:453–7. doi: 10.1038/361453a0. 10.1038/361453a0. [DOI] [PubMed] [Google Scholar]
- 9.Sappino AP, Madani R, Huarte J, Belin D, Kiss JZ, Wohlwend A, Vassalli JD. Extracellular proteolysis in the adult murine brain. The Journal of clinical investigation. 1993;92:679–85. doi: 10.1172/JCI116637. 10.1172/JCI116637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Salles FJ, Strickland S. Localization and regulation of the tissue plasminogen activator-plasmin system in the hippocampus. The Journal of neuroscience : the official journal of the Society for Neuroscience. 2002;22:2125–34. doi: 10.1523/JNEUROSCI.22-06-02125.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Tsirka SE, Gualandris A, Amaral DG, Strickland S. Excitotoxin-induced neuronal degeneration and seizure are mediated by tissue plasminogen activator. Nature. 1995;377:340–4. doi: 10.1038/377340a0. 10.1038/377340a0. [DOI] [PubMed] [Google Scholar]
- 12.Siddiq MM, Tsirka SE. Modulation of zinc toxicity by tissue plasminogen activator. Molecular and cellular neurosciences. 2004;25:162–71. doi: 10.1016/j.mcn.2003.10.007. 10.1016/j.mcn.2003.10.007. [DOI] [PubMed] [Google Scholar]
- 13.Wang YF, Tsirka SE, Strickland S, Stieg PE, Soriano SG, Lipton SA. Tissue plasminogen activator (tPA) increases neuronal damage after focal cerebral ischemia in wild-type and tPA-deficient mice. Nature medicine. 1998;4:228–31. doi: 10.1038/nm0298-228. [DOI] [PubMed] [Google Scholar]
- 14.Frey U, Muller M, Kuhl D. A different form of long-lasting potentiation revealed in tissue plasminogen activator mutant mice. The Journal of neuroscience : the official journal of the Society for Neuroscience. 1996;16:2057–63. doi: 10.1523/JNEUROSCI.16-06-02057.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Huang YY, Bach ME, Lipp HP, Zhuo M, Wolfer DP, Hawkins RD, Schoonjans L, Kandel ER, Godfraind JM, Mulligan R, Collen D, Carmeliet P. Mice lacking the gene encoding tissue-type plasminogen activator show a selective interference with late-phase long-term potentiation in both Schaffer collateral and mossy fiber pathways. Proc Natl Acad Sci U S A. 1996;93:8699–704. doi: 10.1073/pnas.93.16.8699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Medina MG, Ledesma MD, Dominguez JE, Medina M, Zafra D, Alameda F, Dotti CG, Navarro P. Tissue plasminogen activator mediates amyloid-induced neurotoxicity via Erk1/2 activation. The EMBO journal. 2005;24:1706–16. doi: 10.1038/sj.emboj.7600650. 10.1038/sj.emboj.7600650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Matys T, Pawlak R, Matys E, Pavlides C, McEwen BS, Strickland S. Tissue plasminogen activator promotes the effects of corticotropin-releasing factor on the amygdala and anxiety-like behavior. Proc Natl Acad Sci U S A. 2004;101:16345–50. doi: 10.1073/pnas.0407355101. 10.1073/pnas.0407355101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Pawlak R, Magarinos AM, Melchor J, McEwen B, Strickland S. Tissue plasminogen activator in the amygdala is critical for stress-induced anxiety-like behavior. Nature neuroscience. 2003;6:168–74. doi: 10.1038/nn998. 10.1038/nn998. [DOI] [PubMed] [Google Scholar]
- 19.Nagai T, Yamada K, Yoshimura M, Ishikawa K, Miyamoto Y, Hashimoto K, Noda Y, Nitta A, Nabeshima T. The tissue plasminogen activator-plasmin system participates in the rewarding effect of morphine by regulating dopamine release. Proc Natl Acad Sci U S A. 2004;101:3650–5. doi: 10.1073/pnas.0306587101. 10.1073/pnas.0306587101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Maiya R, Zhou Y, Norris EH, Kreek MJ, Strickland S. Tissue plasminogen activator modulates the cellular and behavioral response to cocaine. Proc Natl Acad Sci U S A. 2009;106:1983–8. doi: 10.1073/pnas.0812491106. 10.1073/pnas.0812491106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Noel M, Norris EH, Strickland S. Tissue plasminogen activator is required for the development of fetal alcohol syndrome in mice. Proc Natl Acad Sci U S A. 2011;108:5069–74. doi: 10.1073/pnas.1017608108. 10.1073/pnas.1017608108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Yepes M, Sandkvist M, Moore EG, Bugge TH, Strickland DK, Lawrence DA. Tissue-type plasminogen activator induces opening of the blood-brain barrier via the LDL receptor-related protein. The Journal of clinical investigation. 2003;112:1533–40. doi: 10.1172/JCI19212. 10.1172/JCI19212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Seeds NW, Basham ME, Haffke SP. Neuronal migration is retarded in mice lacking the tissue plasminogen activator gene. Proc Natl Acad Sci U S A. 1999;96:14118–23. doi: 10.1073/pnas.96.24.14118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Nicole O, Docagne F, Ali C, Margaill I, Carmeliet P, MacKenzie ET, Vivien D, Buisson A. The proteolytic activity of tissue-plasminogen activator enhances NMDA receptor-mediated signaling. Nature medicine. 2001;7:59–64. doi: 10.1038/83358. [DOI] [PubMed] [Google Scholar]
- 25.Pang PT, Teng HK, Zaitsev E, Woo NT, Sakata K, Zhen S, Teng KK, Yung WH, Hempstead BL, Lu B. Cleavage of proBDNF by tPA/plasmin is essential for long-term hippocampal plasticity. Science. 2004;306:487–91. doi: 10.1126/science.1100135. [DOI] [PubMed] [Google Scholar]
- 26.Sawaya R, Highsmith R. Plasminogen activator activity and molecular weight patterns in human brain tumors. Journal of neurosurgery. 1988;68:73–9. doi: 10.3171/jns.1988.68.1.0073. [DOI] [PubMed] [Google Scholar]
- 27.Yalcin B, Wong K, Agam A, Goodson M, Keane TM, Gan X, Nellaker C, Goodstadt L, Nicod J, Bhomra A, Hernandez-Pliego P, Whitley H, Cleak J, Dutton R, Janowitz D, Mott R, Adams DJ, Flint J. Sequence-based characterization of structural variation in the mouse genome. Nature. 2011;477:326–9. doi: 10.1038/nature10432. 10.1038/nature10432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Keane TM, Goodstadt L, Danecek P, White MA, Wong K, Yalcin B, Heger A, Agam A, Slater G, Goodson M, Furlotte NA, Eskin E, Nellaker C, Whitley H, Cleak J, Janowitz D, Hernandez-Pliego P, Edwards A, Belgard TG, Oliver PL, McIntyre RE, Bhomra A, Nicod J, Gan X, Yuan W, van der Weyden L, Steward CA, Bala S, Stalker J, Mott R, Durbin R, Jackson IJ, Czechanski A, Guerra-Assuncao JA, Donahue LR, Reinholdt LG, Payseur BA, Ponting CP, Birney E, Flint J, Adams DJ. Mouse genomic variation and its effect on phenotypes and gene regulation. Nature. 2011;477:289–94. doi: 10.1038/nature10413. 10.1038/nature10413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Carmeliet P, Schoonjans L, Kieckens L, Ream B, Degen J, Bronson R, De Vos R, van den Oord JJ, Collen D, Mulligan RC. Physiological consequences of loss of plasminogen activator gene function in mice. Nature. 1994;368:419–24. doi: 10.1038/368419a0. [DOI] [PubMed] [Google Scholar]
- 30.Tsuji T, Ohta Y, Kanno Y, Hirose K, Ohashi K, Mizuno K. Involvement of p114-RhoGEF and Lfc in Wnt-3a- and dishevelled-induced RhoA activation and neurite retraction in N1E-115 mouse neuroblastoma cells. Molecular biology of the cell. 2010;21:3590–600. doi: 10.1091/mbc.E10-02-0095. 10.1091/mbc.E10-02-0095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kurose K, Hiratsuka K, Ishiwata K, Nishikawa J, Nonen S, Azuma J, Kato M, Wakeno M, Okugawa G, Kinoshita T, Kurosawa T, Hasegawa R, Saito Y. Genome-wide association study of SSRI/SNRI-induced sexual dysfunction in a Japanese cohort with major depression. Psychiatry research. 2012;198:424–9. doi: 10.1016/j.psychres.2012.01.023. 10.1016/j.psychres.2012.01.023. [DOI] [PubMed] [Google Scholar]
- 32.Hayashi T, Yoshida T, Ra M, Taguchi R, Mishina M. IL1RAPL1 associated with mental retardation and autism regulates the formation and stabilization of glutamatergic synapses of cortical neurons through RhoA signaling pathway. PloS one. 2013;8:e66254. doi: 10.1371/journal.pone.0066254. 10.1371/journal.pone.0066254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Rose JE, Behm FM, Drgon T, Johnson C, Uhl GR. Personalized smoking cessation: interactions between nicotine dose, dependence and quit-success genotype score. Molecular medicine. 2010;16:247–53. doi: 10.2119/molmed.2009.00159. 10.2119/molmed.2009.00159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Jackson AP, Eastwood H, Bell SM, Adu J, Toomes C, Carr IM, Roberts E, Hampshire DJ, Crow YJ, Mighell AJ, Karbani G, Jafri H, Rashid Y, Mueller RF, Markham AF, Woods CG. Identification of microcephalin, a protein implicated in determining the size of the human brain. American journal of human genetics. 2002;71:136–42. doi: 10.1086/341283. 10.1086/341283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Barrow J, Adamowicz-Brice M, Cartmill M, MacArthur D, Lowe J, Robson K, Brundler MA, Walker DA, Coyle B, Grundy R. Homozygous loss of ADAM3A revealed by genome-wide analysis of pediatric high-grade glioma and diffuse intrinsic pontine gliomas. Neuro-oncology. 2011;13:212–22. doi: 10.1093/neuonc/noq158. 10.1093/neuonc/noq158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Sashindranath M, Sales E, Daglas M, Freeman R, Samson AL, Cops EJ, Beckham S, Galle A, McLean C, Morganti-Kossmann C, Rosenfeld JV, Madani R, Vassalli JD, Su EJ, Lawrence DA, Medcalf RL. The tissue-type plasminogen activator-plasminogen activator inhibitor 1 complex promotes neurovascular injury in brain trauma: evidence from mice and humans. Brain : a journal of neurology. 2012;135:3251–64. doi: 10.1093/brain/aws178. 10.1093/brain/aws178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Tsirka SE, Bugge TH, Degen JL, Strickland S. Neuronal death in the central nervous system demonstrates a non-fibrin substrate for plasmin. Proc Natl Acad Sci U S A. 1997;94:9779–81. doi: 10.1073/pnas.94.18.9779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Tabares-Seisdedos R, Rubenstein JL. Chromosome 8p as a potential hub for developmental neuropsychiatric disorders: implications for schizophrenia, autism and cancer. Molecular psychiatry. 2009;14:563–89. doi: 10.1038/mp.2009.2. 10.1038/mp.2009.2. [DOI] [PubMed] [Google Scholar]
- 39.Gabriel R, Lombardo A, Arens A, Miller JC, Genovese P, Kaeppel C, Nowrouzi A, Bartholomae CC, Wang J, Friedman G, Holmes MC, Gregory PD, Glimm H, Schmidt M, Naldini L, von Kalle C. An unbiased genome-wide analysis of zinc-finger nuclease specificity. Nature biotechnology. 2011;29:816–23. doi: 10.1038/nbt.1948. 10.1038/nbt.1948. [DOI] [PubMed] [Google Scholar]
- 40.Pattanayak V, Ramirez CL, Joung JK, Liu DR. Revealing off-target cleavage specificities of zinc-finger nucleases by in vitro selection. Nature methods. 2011;8:765–70. doi: 10.1038/nmeth.1670. 10.1038/nmeth.1670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Bugge TH, Suh TT, Flick MJ, Daugherty CC, Romer J, Solberg H, Ellis V, Dano K, Degen JL. The receptor for urokinase-type plasminogen activator is not essential for mouse development or fertility. The Journal of biological chemistry. 1995;270:16886–94. doi: 10.1074/jbc.270.28.16886. [DOI] [PubMed] [Google Scholar]
- 42.Dewerchin M, Nuffelen AV, Wallays G, Bouche A, Moons L, Carmeliet P, Mulligan RC, Collen D. Generation and characterization of urokinase receptor-deficient mice. The Journal of clinical investigation. 1996;97:870–8. doi: 10.1172/JCI118489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Bugge TH, Flick MJ, Daugherty CC, Degen JL. Plasminogen deficiency causes severe thrombosis but is compatible with development and reproduction. Genes & development. 1995;9:794–807. doi: 10.1101/gad.9.7.794. [DOI] [PubMed] [Google Scholar]
- 44.Ploplis VA, Carmeliet P, Vazirzadeh S, Van Vlaenderen I, Moons L, Plow EF, Collen D. Effects of disruption of the plasminogen gene on thrombosis, growth, and health in mice. Circulation. 1995;92:2585–93. doi: 10.1161/01.cir.92.9.2585. [DOI] [PubMed] [Google Scholar]
- 45.Carmeliet P, Kieckens L, Schoonjans L, Ream B, van Nuffelen A, Prendergast G, Cole M, Bronson R, Collen D, Mulligan RC. Plasminogen activator inhibitor-1 gene-deficient mice. I. Generation by homologous recombination and characterization. The Journal of clinical investigation. 1993;92:2746–55. doi: 10.1172/JCI116892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Lijnen HR, Okada K, Matsuo O, Collen D, Dewerchin M. Alpha2-antiplasmin gene deficiency in mice is associated with enhanced fibrinolytic potential without overt bleeding. Blood. 1999;93:2274–81. [PubMed] [Google Scholar]
- 47.Ling Q, Jacovina AT, Deora A, Febbraio M, Simantov R, Silverstein RL, Hempstead B, Mark WH, Hajjar KA. Annexin II regulates fibrin homeostasis and neoangiogenesis in vivo. The Journal of clinical investigation. 2004;113:38–48. doi: 10.1172/JCI200419684. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.