Abstract
Drug abuse is a serious problem worldwide. Recently, hallucinogens have been reported as a potential preventative and auxiliary therapy for substance abuse. However, the use of hallucinogens as a drug abuse treatment has potential risks, as the fundamental mechanisms of hallucinogens are not clear. So far, no scientific database is available for the mechanism research of hallucinogens. We constructed a hallucinogen-specific chemogenomics database by collecting chemicals, protein targets and pathways closely related to hallucinogens. This information, together with our established computational chemogenomics tools, such as TargetHunter and HTDocking, provided a one-step solution for the mechanism study of hallucinogens. We chose salvinorin A, a potent hallucinogen extracted from the plant Salvia divinorum, as an example to demonstrate the usability of our platform. With the help of HTDocking program, we predicted four novel targets for salvinorin A, including muscarinic acetylcholine receptor 2, cannabinoid receptor 1, cannabinoid receptor 2 and dopamine receptor 2. We looked into the interactions between salvinorin A and the predicted targets. The binding modes, pose and docking scores indicate that salvinorin A may interact with some of these predicted targets. Overall, our database enriched the information of systems pharmacological analysis, target identification and drug discovery for hallucinogens.
Keywords: hallucinogen, salvinorin A, drug abuse, chemogenomics database, target identification, systems pharmacology
Graphical abstract

INTRODUCTION
Drug abuse is considered as a serious problem that threatens health, family, and social life significantly (1), killing about 200,000 people worldwide each year according to a recent report from United Nations (UN). Besides the statistics of deaths, the summary of disability-adjusted life-years (DALYs) among the population can be considered as a measurement of the gap between the current health status and an ideal health situation. The percentage of DALYs with substance abuse is 9.0, which means almost one tenth of a person‘s healthy lifetime was lost because of drug abuse (2). At this point, the analysis of existing treatments and the exploration of new therapy for drug abuse are necessary and urgent.
Hallucinogens, considered to be safe psychoactive substances and less possible to form addiction, are able to modify human mood, perception, consciousness, cognition and behavior (3). Recently, hallucinogens have been reported as potential preventative and auxiliary therapies for substance abuse (4).
Some hallucinogens have already been used to treat substance abuse. For example, a study showed that lysergic acid diethylamide (LSD) was used to combat alcoholism in the 1950s and was recently confirmed to significantly reduce alcohol consumption up to six months (5). Recently, Dr. Matt Johnson‘s group was engaged in a program to investigate the effect of psilocybin in long-term smokers and, so far, has gotten promising results (6). Ketamine was used to mitigate alcoholism and heroin addiction in 1985 and showed positive results (4). An observational study documenting Ayahuasca, in which the major ingredient is dimethyltryptamine (DMT), was used in Canada to treat addictions. Participants showed decreased use of alcohol, tobacco, and cocaine as well as significant improvements in hopefulness, empowerment, and mindfulness (7).
Hallucinogens have other therapeutic potentials in the treatment of mood, obsessive-compulsive, gastrointestinal disorders and pain (Table 1). A preclinical study showed pretreatment of bufotenin of mice with great protection from burn, tourniquet and endotoxin shock (8, 9). Several case studies reported that hallucinogens including LSD, psilocybin and mescaline improved symptoms of obsessive-compulsive disorder (10, 11). LSD and psilocybin can also dramatically reduce anxiety and stress among patients with terminal cancer according to a Phase III clinical study conducted by Heffter‘s research institute (3, 6, 12). Moreover, various studies pointed out great anti-depressive effect of ketamine (13). Ketamine can also act as an analgesic by blocking N-methyl-D-aspartate receptors (14). New applications of ketamine emerge nowadays, such as the less established treatment for post-traumatic epilepsy, and anesthesia in the emergency department and operating theater setting (14, 15). In summary, hallucinogens are wealthy to be studied for their promising therapeutic potentials.
Table 1.
Therapeutic value of hallucinogens
| Hallucinogen | Therapeutic value |
|---|---|
| Psilocybin | Terminal illness, Obsessive-compulsive disorder, Alcoholism, Smoking cessation |
| LSD | Obsessive-compulsive disorder, Alcoholism, Substance abuse, Terminal illness, Tools for cognitive neuroscience |
| Bufotenin | Protection against burn, tourniquet and endotoxin shock |
| Ketamine | Anti-depression, Pain, Mood disorder, Post-traumatic epilepsy, Anesthesia |
| Salvinorin A | Pain, Mood and personality disorders, Substance abuse, Gastrointestinal disturbances |
It was our objective to study mechanism of hallucinogens and ensure their safety in future clinical trials. However, as members of a broad variety containing nearly fifty natural or synthetic compounds, each hallucinogen shares little overlap of mechanisms and most of them are unclear. To solve this problem, we applied computational technologies for target identification, which has advantage with less cost and faster speed.
Salvinorin A (Figure 1), a potent hallucinogen, is the major ingredient of a Mexican plant, Salvia divinorum. It has potential therapeutic value for treatment of pain, mood and personality disorders, substance abuse and gastrointestinal disturbances (16). Recently, the Freeman‘s group has tested salvinorin A as a punisher of drug self-administration in monkeys. Compared to the lever of cocaine administration alone, the average percent of monkeys‘ choice for the associated salvinorin A lever decreased dose-dependently, which indicated the monkeys prefer to choose the lever with cocaine alone. The phenomenon became more obvious when cocaine was given with a higher dose of salvinorin A as punisher. They arrived at the conclusion that salvinorin A can punish drug choice, which suggested salvinorin A as a potential treatment in the development of abuse-deterrent formulations (17).
Figure 1. Representative ligands of opioid receptors.
Salvinorin A is a non-nitrogenous opioid receptor agonist that shares little structural similarity with other ligands of opioid receptors.
The well-known target of salvinorin A is kappa opioid receptor (OPRK) (18). However, salvinorin A has a unique structure and shares little structural similarity with other opioid ligands (Figure 1). As the first reported non-nitrogenous opioid receptor agonist, salvinorin A is also the first known psychoactive diterpenoid. The structural information indicated that nonalkaloids, such as salvinorin A, may be potential scaffolds for developing new drugs targeting on G-protein coupled receptors (GPCRs) (32).
However, it was reported that some experiencers went through a psychic depersonalization process, which is a unique sensation of being disconnected from one‘s body (19, 20). A double-blind, placebo-controlled study of vaporized/inhaled salvinorin A in eight hallucinogen-experienced adults showed that about half of the participants exhibited a positive affect (audible laughter) during peak drug effects and one described his feeling as —hysterically, almost uncontrollably laughing. (21)
Based on the complex effects of salvinorin A listed above, we believe the concept of —one drug, multiple targets interactions can provide reasonable explanations. We hypothesized that in addition to the OPRK, other potential targets can also interact with salvinorin A, and contribute to the complex effects of salvinorin A.
In this paper, we built a hallucinogen-specific chemogenomics knowledgebase and applied computational technologies for mechanisms study. Salvinorin A was chosen as a case study for demonstration.
MATERIALS AND METHODS
Database infrastructure and web interface
Hallucinogen specific knowledgebase is designed with a MySQL database (www.mysql.com) and an apache web server (www.apache.org) with OpenBabel (22) at the back end as the search engine for chemical structures. The web interface is written in PHP language (www.php.net).
Chemo-informatics Tools
To facilitate the in silico mechanism study and to further explore the therapeutic potential of hallucinogens for treatment of drug abuse, the hallucinogen specific knowledgebase also serves as a cloud computing server by integrating diverse chemo-informatics tools based on state-of-the-art machine learning algorithms. In this paper, we mainly applied two of our developed systems pharmacological analysis tools, HTDocking and TargetHunter, which are core functions in our hallucinogen-specific knowledgebase.
• HTDocking
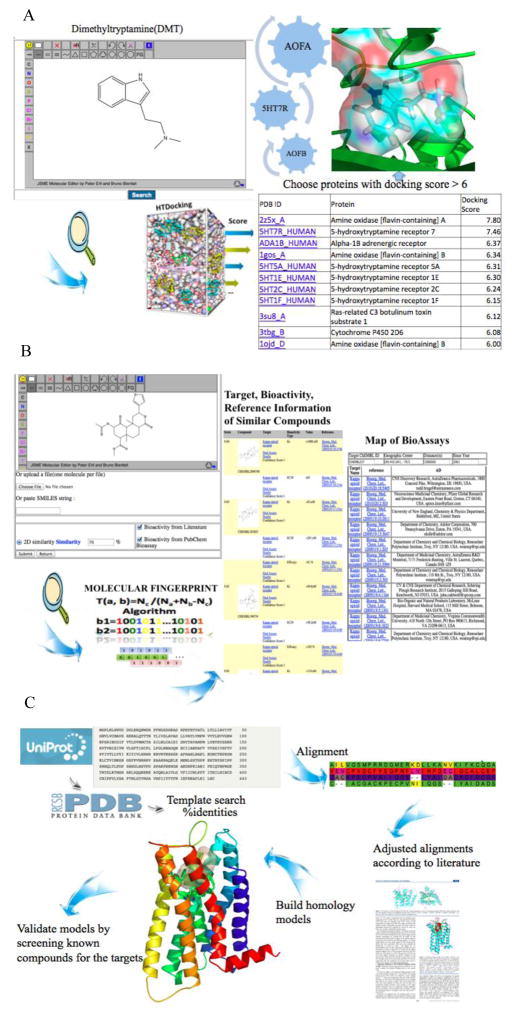
HTDocking is a web-interface computing tool that can run the molecular docking procedures automatically to predict possible interactions between compounds and targets (Figure 2A). In the current version of hallucinogen specific knowledgebase, crystal structures of hallucinogenic related protein targets have been collected from Protein Data Bank (PDB) (www.rcsb.org/pdb) (23). All of the original ligands are removed from the crystal structures. Water molecules and ions are also removed from the crystal structures if necessary. Pockets are defined according to the co-crystal structure ligands, or are automatically predicted by the MOLCAD module implemented in SYBYL-X 1.3 (24–26). In addition, homology models were established by Modeller for the targets that are lacking crystal structures (27). Docking scores calculated by AutoDock Vina (http://vina.scripps.edu/) are used to assess and rank potential targets of a small molecule submitted to the server. The score represents as - log10 (predicted Ki), where predicted Ki=exp (−ΔG×1000/(8.314×298.15) and ΔG (kJ/mole) is the best binding affinity value of a compound with a protein calculated by AutoDock Vina (28). Docking scores larger than 6.0 mean that the predicted Ki value of a compound with its target is within the micromole range or even better. We used SYBYL-X 1.3 to further analyze the detailed interactions (29).
Figure 2. Schematic diagram of HTDocking, TargetHunter, and 3D homology modeling.
(A) When a structure of small molecule is submitted to the HTDocking website, and the HTDocking program will automatically calculate the binding affinity values of this molecule against a list of protein targets. The predicted targets will be ranked by docking scores and can be retrieved through the website. (B) A structure of small molecule is submitted to the TargetHunter website together with a structure similarity threshold, and the TargetHunter program will automatically search similar compounds above that similarity based on the molecular fingerprint. The targets of this compound are predicted as the annotated targets of its similar compounds. The results will be ranked by similarity scores. TargetHunter will provide information including structures of similar compounds as well as their targets, bioassays and references. (C) The sequences of to be modeled proteins were retrieved from UniProt. Then a template search for each sequence was performed from the Protein Data Bank (PDB) according to the sequence identities. Sequence alignments of the protein to be modeled and the templates were done by using the Modeller 9.12 software. Alignments were adjusted according to the literature. Finally, the homology models were built.
• TargetHunter
TargetHunter is a ligand-based targets searching tool. It was designed based on the principle that structurally similar compounds have great possibility sharing similar targets and biological profiles (30–32). When a compound structure is submitted to the server, TargetHunter can provide not only the predicted targets, but also the structures of similar compounds. TargetHunter can also provide the information of potential collaborators whose labs have the established bioassays for experimental validation (Figure 2B). Detailed descriptions can be found in our published paper (33).
Data collection
The hallucinogenic related data was collected from literatures and a variety of public databases including the ChEMBL database (34), DrugBank (35), ClinicalTrials.gov (36), BindingDB (37) and SuperTarget (38). The 3D crystal structures of protein targets were collected from Protein Data Bank (23, 39), while the related signaling pathways were gathered from the KEGG pathway database (40). After the data collection, all these chemical structures, bioactivity values, pathways, bioassays, and references were imported into our hallucinogen-specific knowledgebase.
Homology Modeling
Homology model with a sequence identity of 35% or greater is a powerful tool and strategy for drug discoveries and functional studies. Homology modeling was used to build structure models for those targets with no crystal structures reported in PDB. Firstly, the whole sequences of targets were obtained from the UniProtKB / Swiss-Prot (www.uniprot.org/uniprot) (41). The templates were searched from Protein Data Bank according to sequence identities. When the sequences of the modeled protein and its template(s) have higher sequence identities, the 3D homology models will be more reliable. Secondly, the sequences and templates were aligned using the Modeller 9.12 software (42). In the meantime, we adjusted the alignments according to the literature. Finally, the homology models were built (Figure 2C).
For illustration, we used the construction of DRD2 homology model as an example. As there is no available crystal structure for the DRD2 receptor, we thus built the homology model of DRD2 for the following studies. The full sequence of the human DRD2 receptor (P14416, 443 residues) was obtained from UniProtKB. The crystal structure of human DRD3 (PDB entry: 3PBL, resolution: 2.89 Å) (43) was used as the template to construct the human DRD2 as described in previous papers (44, 45). In briefly, sequence similarity between DRD3 and DRD2 is 90% and up to 79% sequence identity is found in the trans-membrane (TM) regions where the ligand binding site is located. After sequence alignment, we manually checked to make sure that all TMs alignments for those conserved motifs in GPCRs were reasonable. Examples are —D/ERY in TM3, —CWxPx (or —D/E 6.30) in TM6, and —NPxxY in TM7. We also checked and manually adjusted the alignments of sequences for TMs to make sure that there were no —gaps in the helical sequences. DRD2 homology model has been validated by proSA-web Zscores, PROCHECK Ramachandran plots, molecular dockings, and molecular dynamics simulations, etc. The whole DRD2 model and its important residues involved in the binding pocket agreed with the experimental data and docking studies. More detailed procedures can be found in our published protocol (26).
RESULTS
Database Analysis
Forty-six hallucinogenic compounds together with 144 protein targets and 161 related pathways were collected from literature and databases including PubMed (www.ncbi.nlm.nih.gov/pubmed/), PubChem (pubchem.ncbi.nlm.nih.gov), DrugBank (www.drugbank.ca), ChEMBL (www.ebi.ac.uk/chembl), KEGG (www.genome.jp/kegg), and PDB (www.pdb.org). The database also archived 145 chemical compounds associated with these Hallucinogens targets reported in 6,721 assays and 23,598 references.
The largest component of targets were enzymes, occupying over more than half of hallucinogen related targets, including amine oxidases (AOF), Ras-related C3 botulinum toxin substrates (46), surase-isomaltase (SUIS), CYP enzymes and so on. Membrane receptors were counted as the second biggest component and consisted of approximately 10% of hallucinogen related targets, including serotonin receptors (5HTRs), muscarinic acetylcholine receptors (ACMs), opioid receptors (OPRs), dopamine receptors (DRDs), cannabinoid receptors (CNRs) and so on. Hallucinogen related targets also included transporters, ion channels, epigenetic regulators, transcription factors and unclassified proteins.
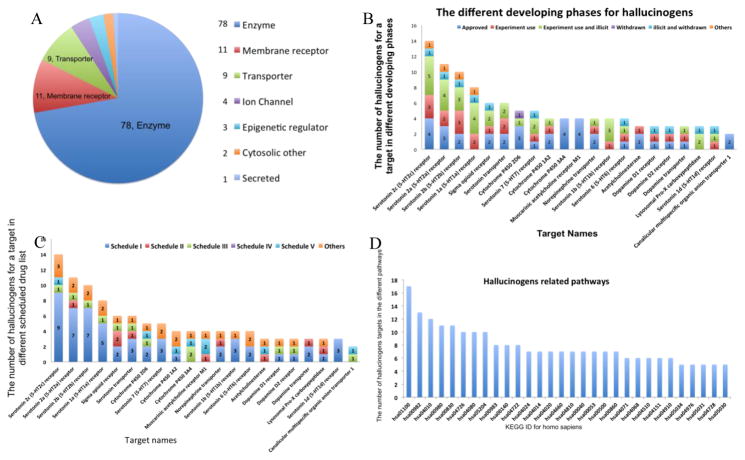
To further evaluate the current hallucinogens, the statistics on these hallucinogens targets were plotted according to their interacting targets in different scheduled drugs lists and development stages. As Figure 3A shown, the hallucinogens targets we collected were ranked according to the number of scheduled drugs in the United States Controlled Substances Act (CSA). More than 80% of hallucinogens were on the controlled drug list. Most of the targets only had one or two drugs that are not controlled substances. CSA is the statute prescribing federal U.S. drug policy under which the manufacture, importation, possession, uses and distribution of certain substances is regulated. The legislation created five schedules (classifications), with varying qualifications for a substance to be included in each. Two federal agencies, the Drug Enforcement Administration (47) and the Food and Drug Administration (FDA), determine which substances are added to or removed from the various schedules, although the statute passed by Congress created the initial listing (48). 5HTRs are ranked on the top with several number of hallucinogens acting on 5HT2C, 5HT2A, 5HT1B, and 5HT1A, including MMDA, 5-Methoxy-DMT, LSD, Mescaline, Psilocin, MDA, DOM and so on. This is reasonable because the two major classes of psychedelic hallucinogens, the indoleamines (e.g., LSD) and the phenethylamines (e.g., mescaline), have a common site of action as partial agonists at 5HT2A and other 5HT2 receptors in the central nervous system (49). Among these hallucinogens acting on 5HTRs, more than half are in the list of Schedule I drugs with high potential of drug abuse and no acceptable medical use (Figure 3C. 9 hallucinogens for 5HT2C, 7 for 5HT2A, 7 for 5HT2B, and 5 for 5HT1A). For example, LSD (lysergic acid diethylamide), a semi-synthetic psychedelic drug famous for its involvement in the counterculture of the 1960s, is on the list of Schedule I drugs (50). The others targeting 5HTRs are included in the list of Schedule II, III, and V, which have less potential of drug abuse, less potential leading to physical or psychological dependence and are currently acceptable for medical use with various restrictions. For the other receptors, such as OPRs, ACM1, acetyl cholinesterase (51) and dopamine receptor 2 (DRD2), obverse conditions were observed: less drugs are found in the schedule I, more hallucinogens are in the list of schedule II, III, V, or even approved drugs without scheduled control. For instance, only two hallucinogens acting on sigma opioid receptors are in the list of schedule I, including Benzylpiperazine and Tenocyclidine. Others are involved in the Schedule II (Cocaine and Phencyclidine), schedule III (Dextromethorphan), indicating for less potential to induce drug dependence and drug abuse.
Figure 3. Summary of targets and pathways for hallucinogens.
(A) 144 hallucinogens related targets were summarized according to 46 hallucinogenic compounds. (B) Hallucinogens in different list of schedule drugs according to the CSA (Controlled Substances Act) and their corresponding targets. These hallucinogens were classified by different schedule lists with distinct colors. The blue and orange bars indicated the Schedule I and unscheduled hallucinogens, respectively. The red, green, and purple and cyan bars denoted hallucinogens in Schedule II, III, IV, and V, respectively (C) Hallucinogens in different uses and their targets. These hallucinogens were classified by different uses with distinct colors. The blue and orange bars indicated the approved and unclassified hallucinogens, respectively. The red, green, and purple and cyan bars denoted hallucinogens in experiment use, experiment use and being illicit, being withdrawn, being illicit and withdrawn, respectively. (D) Hallucinogens related pathways. Hallucinogens and their targets were plotted according to the pathways the targets involved.
The similar results can be observed from Figure 3B, the statistic of the different developing phases for hallucinogens. Although 5HTRs are the targets of many hallucinogens, from Figure 3B we can see only a small portion of hallucinogens acting on 5HTRs are approved by FDA and allowed for medical use in the U.S. (4 for 5HT2C, 3 for 5HT2A, and 2 for 5HT2B). The other hallucinogens for 5HT receptors are only experimentally used in animal studies or being withdrawn from the market and then becoming illicit. On the other hand, four hallucinogens acting on ACM1 are all approved by the FDA to be used in market; two-thirds of hallucinogens targeting AChE are allowed for medical use in the America. These results may suggest for relatively more accepted safety targeting ACM1 and AChE than 5HTRs for hallucinogens.
The hallucinogens related pathways were mapped out according to the number of hallucinogen targets involved. The top 20 pathways associated with a large number of hallucinogen related proteins were listed and ranked in Figure 3D. It is no surprise that serotonergic synapse pathway (KEGGID: hsa04726) and dopaminergic synapse (KEGGID: hsa04728) are among the top of the pathway list, indicating that many hallucinogens share a common mechanism of action, such as interacting with 5HTRs and DRDs. Our data also shows that pathways related to drug addiction, such as amphetamine addiction (KEGGID: hsa05031), cocaine addiction (KEGGID: hsa05030), and alcoholism (KEGGID: hsa05034), are also enriched on the top list, which could imply that hallucinogens share some common pathways with other drugs of addictions. Furthermore, we also noticed that the neuron activity-related pathways, such as neuroactive ligand-receptor interaction (KEGGID: hsa04080) and neurotrophin signaling pathway (KEGGID: hsa04722), are among the top list, which indicates the hallucinogenic activity may also modulate the neurological pathways of normal neuron activity.
Database Validation
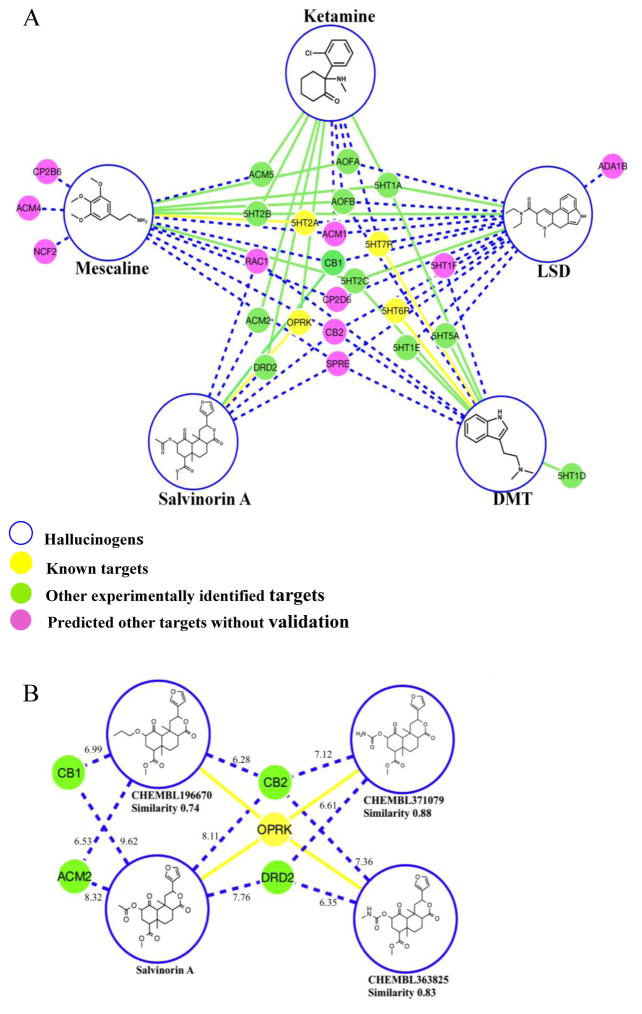
As described in the method section, we submitted many hallucinogenic compounds into HTDocking program of Hallucinogen Knowledgebase (www.cbligand.org/hallucinogen/) to validate our database. As an example, five representative hallucinogenic compounds that share little structural similarity were selected to perform the docking results. Based on the docking scores, we drew a schema to show the interactions between compounds and related targets. As shown in Figure 4A, five compounds were listed, including ketamine, LSD, mescaline, DMT and salvinorin A. All of the targets listed in the schema have docking scores above 6.0 with corresponding ligands, indicated that the Ki values should be within micromole range. For example, the docking score of salvinorin A with OPRK was 7.40. This was congruent with the fact that the OPRK is a known target of salvinorin A (Ki value of 16nM at OPRK) (18, 52). Similarly, as Figure 4A illustrates, 5HT2A, 5HT6R and 5HT7R were known targets of DMT. 5HT2A was also the target of mescaline and LSD. 5HT1A was the target of Ketamine, Mescaline, DMT and LSD. Like evidence listed above, our docking results matched the information we collected either from literature or databases, which meant this method is reliable.
Figure 4. Systems pharmacological analysis of salvinorin A and other known hallucinogens.
(A) Abbreviations, 5HT=5-hydroxytryptamine receptor, ACM=Muscarinic acetylcholine receptor, ADA1A=Alpha-1A adrenergic receptor, AOFA=Amine oxidase [flavin-containing] A, AOFB=Amine oxidase [flavin-containing] B, CNR=Cannabinoid receptor, OPRK=Kappa opioid receptor, DRD2=Dopamine receptor 2, CP2B6=Cytochrome P450 2B6, CP2D6=Cytochrome P450 2D6, NCF2=Neutrophil cytosol factor 2, RAC1=Ras-related C3 botulinum toxin substrate 1, SPRE=Sepiapterin reductase, SUIS=Sucrase-isomaltase (B) Three similar compounds of salvinorin A were chosen according to the results from TargetHunter. OPRK is a known target for all of them. Other four targets including CB1, CB2, ACM2 and DRD2 were predicted according to TargetHunter and HTDocking results.
Systems Pharmacological Analysis of Salvinorin A
Systems pharmacology is the study of drugs, drug-target interactions and drug effects in a macroscopic view, combining large-scale biological experiments data with computational analysis (53). Systems pharmacology focuses on big data analysis, which tends to believe in —multiple drugs, multiple targets interactions , unlike the traditional concept —one drug, one target interaction (54). With obvious advantages of rapid identification of the networks and connections among different drugs and targets, it works as a very useful method for quick analysis of drug repurposing. It is also beneficial to analyze the complex drug‘s adverse effects (53–56). Using this method, we explored possible targets of salvinorin A by integrating predicting results from both HTDocking and TargetHunter. Based on the principle that structurally similar compounds had a greater possibility of sharing similar targets and biological profiles (51–53), we further mapped out the potential targets of salvinorin A as the annotated targets of its similar compounds. In Figure 4B, we picked three compounds that are structurally similar to salvinorin A with similarities above 0.70 from TargetHunter program and these three compounds were reported to target OPRK (CHEMBL196670, Ki = 28.7nM; CHEMBL371079, Ki = 3.2nM; CHEMBL363825, Ki = 83.2nM) (69). We then submitted these three compounds and salvinorin A to the HTDocking program. The combined TargetHunter and HTDocking predictions were plotted (Figure 4B). After the systems pharmacological study, we continued molecular mechanism study, looking in depth into the relationship of each possible target with salvinorin A.
According to the prescreening HTDocking study, besides OPRK, we found four promising targets of salvinorin A (Figure 4A). ACM2 got a docking score of 8.32. The molecular docking study was based on the co-crystal structure of ACM2 (PDB entry: 3UON, resolution 3.00 Å) published in 2012 (57, 58). The original ligand was QNB, an antagonist of ACM2 (Kd = 0.2nM). Figure 5A shows the interactions between QNB and helix III, IV, V, VI, and VII of ACM2. Asp103, Tyr104, Ser107, Trp155, Phe181, Trp400, Tyr403, Asn404 and Cys429 were important residues within the binding pocket. The amine moiety of QNB formed a salt bridge (4.0 Å) with the side chain of Asp103 in TM3, which was proven to be important by modeling study, mutagenesis and covalent-labeling experiments (57, 58). Asn404 formed hydrogen bonds with both hydroxyl (2.6 Å) and carbonyl (3.2 A) group of QNB. However, according to the residues‘ mutation result, Asn404 had no contribution to interactions between endogenous acetylcholine and the receptor (57, 58). Phe181 and the benzene ring formed a weak π-π interaction. Indeed, Phe181 was the unique residue for ACM2, in which this position for other ACMs was leucine (57). These might explain the reason why QNB was not a highly selective ligand of ACM2.
Figure 5. Interactions between salvinorin A and predicted targets.
(A) The 3D homology model of CB1. The pocket was emphasized in pink. The important residues within pocket were highlighted with yellow. (B) The interactions between salvinorin A and CB1. The purple dash lines represent the π-π interactions between Phe57 and the furan moiety of the salvinorin A. The docking score of salvinorin A on CB1 is 9.60. (C) The co-crystal structure of ACM2 (PDB entry: 3UON, resolution: 3.00 Å), binding affinity Ki = 0.2nM. (D) The docking results between salvinorin A and ACM2. Salvinorin A have a similar position and pose with QNB, the original ligand of ACM2 within the binding pocket of ACM2. The docking score of salvinorin A on ACM2 is 8.32.
Salvinorin A was located in the same pocket as QNB at ACM2 (Figure 5B). The poses and interactions of QNB with ACM2 were highly similar to that of salvinorin A with ACM2. Asn404 and Trp155 formed strong hydrogen bonds with two separate parts of ester bond of salvinorin A (both within 3 Å). Similarly, Ser107 and Cys429 formed strong hydrogen bonds with salvinorin A (both with 3 Å). Trp400 and the ternary heterocyclic ring of salvinorin A formed hydrophobic contacts. Similarly, Trp400 had hydrophobic contact with the nitrogen heterocyclic ring of QNB from the original co-crystal (57). According to the residues mutations study, mutation of Cys429 reduced agonist binding affinity with ACM2 over 10 fold, while the mutation of Tyr104, Trp155, or Trp400 reduced both agonist and antagonist binding affinity over 10 fold (57). Based on these similar interactions, we could predict that ACM2 to be a potential target of salvinorin A.
Besides ACM2, our HTDocking results showed that CB1 and CB2 had a high possibility to interact with salvinorin A. The docking score of salvinorin A against CB2 was 8.11. The binding pocket of CB2 mainly located in helix III, V, VI and VII (Purple region in Figure 6A). The important residues of CB2 were Phe87, Phe91, Phe94, Asp101, Phe106, Lys109, Ile110, Val113, Phe117, Glu181, Leu182, Phe183, Trp194, Phe197, Trp258, Val261, Met265, Lys278, Lys279, Phe281 and Ser285 (Highlighted with green in Figure 6A). This model was validated by our group using virtual screening, molecular dynamics simulation, and residues‘ mutations coupled with experiments radio-ligand binding results (46).
Figure 6. Interactions between salvinorin A and predicted targets.
(A) The 3D homology model of CB2. The pocket was emphasized in purple. The important residues within the pocket were highlighted with green. (B) The interactions between salvinorin A and CB2. The docking score of salvinorin A on CB2 is 8.11. (C) The 3D homology model of DRD2. The pocket was emphasized in pink. The important residues within the pocket were highlighted with green. (D) The interactions between salvinorin A and DRD2. The docking score of salvinorin A on DRD2 is 7.76.
As shown in Figure 6B, in extracellular loop 2 region, Leu182 formed a hydrogen bond (2.8 Å) with the ester bond moiety of salvinorin A. As literature reported, the extracellular loop 2 region of CB2 proved to be essential for ligand bindings (59, 60). Besides, the amine part of Lys109 in helix III formed a hydrogen bond (2.8 Å) with the oxygen of salvinorin A. Lys109 is a conserved residue in GPCRs. Phe281 had hydrophobic contact with the ternary heterocyclic ring of salvinorin A. In summary, salvinorin A preferred hydrophobic interactions within CB2 binding pocket with residues including Phe91, Phe94, Phe106, Ile110, Trp194, Phe197, Trp258 and Phe281.
CB1 shared 48% and 68% of the whole sequence identity and the trans-membrane domains sequence identity with CB2 (24). The same method was used to construct the CB1 homology model. We used the MOLCAD module implemented in SYBYL-X1.3 to explore its potential binding pocket. Based on the model reported by literatures, our predicted pocket of the receptor was proven reliable (24, 61). The binding site of CB1 mainly formed by residues from helix III, V, VI and VII, which was similar to CB2. The important residues included Ile150, Phe57, Phe60, His61, Lys75, Val79, Asp149, Leu216, Met220, Phe236, and Ser240 (Figure 5C) (61).
We observed a relatively high docking score of 9.62 when salvinorin A interacted with CB1. As shown in Figure 5D, amino part of His174 formed a hydrogen bond (4.7 Å) with the furan moiety of salvinorin A. The furan ring also had a π-π interaction (within 4 Å) with Phe170. Both the hydrogen bond and the π-π interaction contributed to the slightly extend of the furan moiety of salvinorin A to helix II. Ile267 located at the extracellular loop 2 region formed a hydrogen bond (3.3 Å) with salvinorin A. Both Val196 and Ser383 formed hydrogen bonds (within 4 Å) with salvinorin A. The insight interactions between salvinorin A and CB1, CB2 indicated that these two targets might also contribute to the pharmacological effects of salvinorin A.
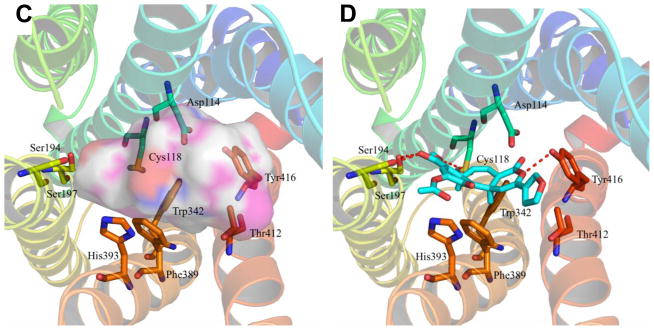
DRD2 plays a central role in pathophysiology of psychosis, movement disorders such as Parkinson‘s disease or multiple system atrophy, and addiction (62–64). It was also reported associative with pathological laughing (65). We got the docking score of 7.76 for salvinorin A within the DRD2 homology model. The homology model of DRD2 was established according to the template of DRD3 (66, 67). Figure 6D shows that Cys118, Ser194 and Ser197 formed strong hydrogen bonds with the ester moiety of salvinorin A (all hydrogen bonds are within 3 Å). Tyr416 formed a hydrogen bond with the oxygen (2.9 Å). Salvinorin A fits well within the binding pocket formed by helix III, V, VI and VII. The important residues in the binding pocket are Asp114, Cys118, Ser194, Ser197, Trp342, Phe389, His393, Thr412 and Tyr416 (Figure 6C) (66, 67). Based on these similarities, we suggested salvinorin A might bind to DRD2, which also contribute to the pharmacological effects of salvinorin A (detailed explanations were in discussion session).
DISCUSSION
Our hallucinogen-specific knowledgebase is the first comprehensive database for hallucinogenic drug research, which also includes several analysis tools (HTDocking, TargetHunter, BBB Predictor, properties explorer, the pains predictor, toxicity predictor). It could provide a quicker, less costly and relatively easier way for researchers to perform systems pharmacological analysis, off-target identifications and mechanism studies.
Benefited from our hallucinogen specific knowledgebase, we applied the integrated tools to explore possible targets of a special hallucinogen – salvinorin A. We found that in addition to OPRK, other targets such as CB1, CB2, ACM2 and DRD2 might also be involved in the pharmacological effects of salvinorin A. In fact, our predictions were supported by some literature. Seeman‘s group designed an in vitro experiment to test the binding affinity of salvinorin A against the D2 High receptor. The D2High receptor is an agonistic, high-affinity state of DRD2. They found salvinorin A had EC50 values of 89 nM at the D2High receptor and were blocked by 10μM S-sulpiride, an antagonist of DRD2. They claimed that the D2High receptors may be more pathophysiologically important than the overall expression of DRD2 (68). However, the overall effects of salvinorin A on DRD2 still need to be further tested.
As we know, the dopaminergic system and the cannabinoid system work as the essential parts for the mechanism of addictions (69–71). Based on our similarity search results and docking studies, we predicted that targeting CB1, CB2 or DRD2 as an explanation for the potential therapeutic profile of salvinorin A to treat drug abuse, which was supported by former studies. For instance, Capasso‘s group reported the inhibitory effect of salvinorin A on ileitis-induced hypermotility, which was conducted by the crosstalk between OPRK and CB1 under pathological conditions in the inflamed gut (72). However, we believed that the effects of salvinorin A on the normal healthy condition should also be considered. At this point, whether salvinorin A acts on CB1 or CB2 under health conditions should be tested in the future. Later, Fichna‘s group got interesting results of the effects of salvinorin A on colonic. They found differences between in vivo and in vitro phenomena, where the effects of salvinorin A on colonic motility in vitro were inhibited by antagonists of OPRK, CB1 and CB2, but the in vivo were largely mediated by OPRK (73). The explanations for these controversial results still need to be discussed.
ACM2 is expressed both in the central nervous system (CNS) and the periphery system, especially in the heart and smooth muscle tissues (74). We assumed that the effects of salvinorin A on colonic motility were partly because of its interactions with ACM2. Besides, Capasso‘s group also found that salvinorin A depressed enteric cholinergic transmission in the guinea-pig ileum. Whether this pharmacological effect was triggered through OPRK or ACM2 still needs further confirmation.
As for the uncontrollable laughing side effect of salvinorin A, we suggested that targeting on DRD2 or ACM2 might be one of the reasons. Literature reported that the dopaminergic system and the muscarinic transmission were closely related to pathological laughing and crying (PLC) (65). Above all related receptors, DRD2 was highly pointed out with great possibility to be blamed of. Clinical observations documented that levodopa, amantadine or ropinirole were successfully used to treat patients with PLC (75, 76). Ten out of twenty-five patients with PLC symptoms having less brain lesions were almost completely controlled by the administration of levodopa or amantadine (75). Those patients without improvement either have severe dementia or have injury to a large area of the brain and various neurons (75). Mariën‘s group also reported a patient with tumoural aetioloty who has PLC symptoms and was successfully treated with a specific DRD2 and DRD3 agonist, ropinirole (76). Schindehütte‘s group reported two drug-induced psychosis in Parkinson disease (PD) patients who developed pathological laughing after using ziprasidone, a DRD2 and 5HTRs antagonist, dose-dependently (77). Coupled with the fact of dopaminergic deficit caused by PD, the authors suggested that blockade of DRD2 reporters may induce pathological laughing (77). ACMs play an important role in PLD (65). Cohn‘s group investigated nine PD patients having PLC symptom, who showed improvement when given atropine or scopolamine (78). Zellers‘ group found a man who explored pyrethrin insecticide, a cholinesterase inhibitor, and appeared with the symptom of uncontrollable laughing (79).
Although the pathways of PLC are complicated and unclear, literature reported lesion of some certain area of the brain or binding to some receptors in the brain should be blamed (65, 75). By X-ray computed tomography scan, scientists revealed that the lesion of brain of PLC patients could be found in the basal ganglia, internal capsule, substantia nigra, cerebral peduncle, hypothalamus, bilateral thalamus, bilateral pyramidal tract, or extrapyramidal systems (75). A review article summarized some receptors that may work in PLC according to clinical observations or literature reports, including glutamatergic NMDA, GABA-A, ACM1, ACM2, and ACM3, DRD2, adrenergic alpha-1 and alpha-2, serotoninergic 5HT1A and 5HT1B/D, and sigma-1 (65). In our case, we suggested salvinorin A is not only an OPRK specific agonist, but that it may also act as a low-affinity modulator of DRD2 and ACM2, which led to the pathological laughing side effect.
CONCLUSIONS
In our study, we established a comprehensive hallucinogen-specific knowledgebase for scientific purpose. We used our knowledgebase to study the mechanisms of hallucinogens. The prediction of Salvinorin A‘s mechanism was used as a case study in this paper. The results showed that besides OPRK, four new receptors were predicted for the complex effects of salvinorin A, including ACM2, CB1, CB2 and DRD2 We further conducted literature search to find evidences to support our prediction (Table 2). These predictions could be used to guide further experimental validation.
Table 2.
Predicted targets for salvinorin A and literature reports.
| Targets | Docking Score | Related reference |
|---|---|---|
| Muscarinic acetylcholine receptor 2 | 8.32 | |
| Cannabinoid receptor 1 | 9.62 | Capasso R. et al. (2008); Fichna J. et al. (2009) |
| Cannabinoid receptor 2 | 8.11 | Capasso R. et al. (2008); Fichna J. et al. (2009) |
| Dopamine receptor 2 | 7.76 | Lauterbach EC. et al. (2013); Seeman P. et al.(2009) |
Highlights.
A comprehensive hallucinogenic-specific knowledgebase was established for research purpose.
Salvinorin A was predicted to act on not only kappa opioid receptor, but also muscarinic acetylcholine receptor, dopamine receptor, and cannabinoid receptors.
The pathological laughing side effect of salvinorin A, which haven‘t been reported from other kappa opioid receptor agonists, should be blamed to its binding to dopamine receptor 2 or muscarinic acetylcholine receptor 2.
Acknowledgments
This project is funded by NIDA (P30 DA035778) for Xiang-Qun Xie‘s lab. We give thanks to Dr. Rao Rapaka from NIDA for his enlightenment of this project.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Center NDI. The Economic Impact of Illicit Drug Use on American Society. Washington DC: United States Department of Justice; 2011. [Google Scholar]
- 2.Tan G, Lou Z, Jing J, Li W, Zhu Z, Zhao L, Zhang G, Chai Y. Screening and analysis of aconitum alkaloids and their metabolites in rat urine after oral administration of aconite roots extract using LC-TOFMS-based metabolomics. Biomedical chromatography : BMC. 2011;25(12):1343. doi: 10.1002/bmc.1607. [DOI] [PubMed] [Google Scholar]
- 3.Nichols DE. Hallucinogens. Pharmacology & therapeutics. 2004;101(2):131–81. doi: 10.1016/j.pharmthera.2003.11.002. [DOI] [PubMed] [Google Scholar]
- 4.Vargas-Perez H, Doblin R. The potential of psychedelics as a preventative and auxiliary therapy for drug abuse. Current drug abuse reviews. 2013;6(1):1–231. doi: 10.2174/18744737112059990023. [DOI] [PubMed] [Google Scholar]
- 5.Dyck E. Hitting Highs at Rock Bottom‘: LSD Treatment for Alcoholism, 1950–1970. Social History of Medicine. 2006;19(2):313–29. [Google Scholar]
- 6.Nichols DE. The Heffter Research Institute: Past and Hopeful Future. Journal of psychoactive drugs. 2014;46(1):20–6. doi: 10.1080/02791072.2014.873688. [DOI] [PubMed] [Google Scholar]
- 7.Thomas G, Lucas P, Capler NR, Tupper KW, Martin G. Ayahuasca-assisted therapy for addiction: results from a preliminary observational study in Canada. Current drug abuse reviews. 2013;6(1):30–42. doi: 10.2174/15733998113099990003. [DOI] [PubMed] [Google Scholar]
- 8.Wallach J. Endogenous hallucinogens as ligands of the trace amine receptors: a possible role in sensory perception. Medical hypotheses. 2009;72(1):91–4. doi: 10.1016/j.mehy.2008.07.052. [DOI] [PubMed] [Google Scholar]
- 9.Markley K, Smallman E, Thornton S. Protection against burn, tourniquet and endotoxin shock by histamine, 5 - hydroxytryptamine and 5 - hydroxytryptamine derivatives. British journal of pharmacology. 1971;42(1):13–24. doi: 10.1111/j.1476-5381.1971.tb07082.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Delgado PL, Moreno FA. Hallucinogens, serotonin and obsessive-compulsive disorder. Journal of psychoactive drugs. 1998;30(4):359–66. doi: 10.1080/02791072.1998.10399711. [DOI] [PubMed] [Google Scholar]
- 11.Brandrup E, Vanggaard T. LSD treatment in a severe case of compulsive neurosis. Acta Psychiatrica Scandinavica. 1977;55(2):127–41. doi: 10.1111/j.1600-0447.1977.tb00149.x. [DOI] [PubMed] [Google Scholar]
- 12.Ahn KH, Bertalovitz AC, Mierke DF, Kendall DA. Dual role of the second extracellular loop of the cannabinoid receptor 1: ligand binding and receptor localization. Molecular pharmacology. 2009;76(4):833–42. doi: 10.1124/mol.109.057356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Monteggia LM, Zarate C. Antidepressant actions of ketamine: from molecular mechanisms to clinical practice. Current opinion in neurobiology. 2015;30:139–43. doi: 10.1016/j.conb.2014.12.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Sawynok J. Topical and peripheral ketamine as an analgesic. Anesthesia & Analgesia. 2014;119(1):170–8. doi: 10.1213/ANE.0000000000000246. [DOI] [PubMed] [Google Scholar]
- 15.Szaflarski JP, Nazzal Y, Dreer LE. Post-traumatic epilepsy: current and emerging treatment options. Neuropsychiatric disease and treatment. 2014;10:1469. doi: 10.2147/NDT.S50421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Cunningham CW, Rothman RB, Prisinzano TE. Neuropharmacology of the naturally occurring κ-opioid hallucinogen salvinorin A. Pharmacological reviews. 2011;63(2):316–47. doi: 10.1124/pr.110.003244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Freeman KB, Naylor JE, Prisinzano TE, Woolverton WL. Assessment of the kappa opioid agonist, salvinorin A, as a punisher of drug self-administration in monkeys. Psychopharmacology. 2014:1–8. doi: 10.1007/s00213-014-3436-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Roth BL, Baner K, Westkaemper R, Siebert D, Rice KC, Steinberg S, Ernsberger P, Rothman RB. Salvinorin A: a potent naturally occurring nonnitrogenous κ opioid selective agonist. Proceedings of the National Academy of Sciences. 2002;99(18):11934–9. doi: 10.1073/pnas.182234399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Casselman I, Nock CJ, Wohlmuth H, Weatherby RP, Heinrich M. From local to global—Fifty years of research on Salvia divinorum. Journal of ethnopharmacology. 2014;151(2):768–83. doi: 10.1016/j.jep.2013.11.032. [DOI] [PubMed] [Google Scholar]
- 20.Zawilska JB, Wojcieszak J. Salvia divinorum: from Mazatec medicinal and hallucinogenic plant to emerging recreational drug. Human Psychopharmacology: Clinical and Experimental. 2013;28(5):403–12. doi: 10.1002/hup.2304. [DOI] [PubMed] [Google Scholar]
- 21.MacLean KA, Johnson MW, Reissig CJ, Prisinzano TE, Griffiths RR. Dose-related effects of salvinorin A in humans: dissociative, hallucinogenic, and memory effects. Psychopharmacology. 2013;226(2):381–92. doi: 10.1007/s00213-012-2912-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.OLBoyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR. Open Babel: An open chemical toolbox. J Cheminf. 2011;3:33. doi: 10.1186/1758-2946-3-33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat T, Weissig H, Shindyalov IN, Bourne PE. The protein data bank. Nucleic acids research. 2000;28(1):235–42. doi: 10.1093/nar/28.1.235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Feng Z, Alqarni MH, Yang P, Tong Q, Chowdhury A, Wang L, Xie X-Q. Modeling, Molecular Dynamics Simulation, and Mutation Validation for Structure of Cannabinoid Receptor 2 Based on Known Crystal Structures of GPCRs. Journal of Chemical Information and Modeling. 2014;54(9):2483. doi: 10.1021/ci5002718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Chen J-Z, Wang J, Xie X-Q. GPCR structure-based virtual screening approach for CB2 antagonist search. Journal of chemical information and modeling. 2007;47(4):1626–37. doi: 10.1021/ci7000814. [DOI] [PubMed] [Google Scholar]
- 26.Xie XQ, Chen JZ, Billings EM. 3D structural model of the G - protein - coupled cannabinoid CB2 receptor. Proteins: Structure, Function, and Bioinformatics. 2003;53(2):307–19. doi: 10.1002/prot.10511. [DOI] [PubMed] [Google Scholar]
- 27.Webb B, Sali A. Comparative protein structure modeling using Modeller. Current protocols in bioinformatics. 2014:5.6. 1–5.6. 32. doi: 10.1002/0471250953.bi0506s47. [DOI] [PubMed] [Google Scholar]
- 28.Jain AN. Scoring noncovalent protein-ligand interactions: a continuous differentiable function tuned to compute binding affinities. J Comput Aided-Mol Des. 1996;10(5):427–40. doi: 10.1007/BF00124474. [DOI] [PubMed] [Google Scholar]
- 29.SYBYL-X 1.3. Tripos International; 1699 South Hanley Rd., St. Louis, Missouri, 63144, USA: 2010. [Google Scholar]
- 30.Schuffenhauer A, Floersheim P, Acklin P, Jacoby E. Similarity metrics for ligands reflecting the similarity of the target proteins. Journal of chemical information and computer sciences. 2003;43(2):391–405. doi: 10.1021/ci025569t. [DOI] [PubMed] [Google Scholar]
- 31.Mitchell JB. The relationship between the sequence identities of alpha helical proteins in the PDB and the molecular similarities of their ligands. Journal of chemical information and computer sciences. 2001;41(6):1617–22. doi: 10.1021/ci010364q. [DOI] [PubMed] [Google Scholar]
- 32.Boström J, Hogner A, Schmitt S. Do structurally similar ligands bind in a similar fashion? Journal of medicinal chemistry. 2006;49(23):6716–25. doi: 10.1021/jm060167o. [DOI] [PubMed] [Google Scholar]
- 33.Wang L, Ma C, Wipf P, Liu H, Su W, Xie X-Q. TargetHunter: an in silico target identification tool for predicting therapeutic potential of small organic molecules based on chemogenomic database. The AAPS journal. 2013;15(2):395–406. doi: 10.1208/s12248-012-9449-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gaulton A, Bellis LJ, Bento AP, Chambers J, Davies M, Hersey A, Light Y, McGlinchey S, Michalovich D, Al-Lazikani B. ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic acids research. 2012;40(D1):D1100–D7. doi: 10.1093/nar/gkr777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Knox C, Law V, Jewison T, Liu P, Ly S, Frolkis A, Pon A, Banco K, Mak C, Neveu V. DrugBank 3.0: a comprehensive resource for omics‘ research on drugs. Nucleic acids research. 2011;39(suppl 1):D1035–D41. doi: 10.1093/nar/gkq1126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zarin DA, Tse T, Williams RJ, Califf RM, Ide NC. The ClinicalTrials. gov results database—update and key issues. New England Journal of Medicine. 2011;364(9):852–60. doi: 10.1056/NEJMsa1012065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wassermann AM, Bajorath J. BindingDB and ChEMBL: online compound databases for drug discovery. Expert opinion on drug discovery. 2011;6(7):683–7. doi: 10.1517/17460441.2011.579100. [DOI] [PubMed] [Google Scholar]
- 38.Hecker N, Ahmed J, von Eichborn J, Dunkel M, Macha K, Eckert A, Gilson MK, Bourne PE, Preissner R. SuperTarget goes quantitative: update on drug–target interactions. Nucleic acids research. 2011:gkr912. doi: 10.1093/nar/gkr912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Bernstein FC, Koetzle TF, Williams GJ, Meyer EF, Brice MD, Rodgers JR, Kennard O, Shimanouchi T, Tasumi M. The protein data bank. European Journal of Biochemistry. 1977;80(2):319–24. doi: 10.1111/j.1432-1033.1977.tb11885.x. [DOI] [PubMed] [Google Scholar]
- 40.Kanehisa M, Goto S, Sato Y, Furumichi M, Tanabe M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic acids research. 2011:gkr988. doi: 10.1093/nar/gkr988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Consortium U. The universal protein resource (UniProt) Nucleic acids research. 2008;36(suppl 1):D190–D5. doi: 10.1093/nar/gkm895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Martí-Renom MA, Stuart AC, Fiser A, Sánchez R, Melo F, Šali A. Comparative protein structure modeling of genes and genomes. Annu Rev Bioph Biom. 2000;29(1):291–325. doi: 10.1146/annurev.biophys.29.1.291. [DOI] [PubMed] [Google Scholar]
- 43.Chien EY, Liu W, Zhao Q, Katritch V, Han GW, Hanson MA, Shi L, Newman AH, Javitch JA, Cherezov V. Structure of the human dopamine D3 receptor in complex with a D2/D3 selective antagonist. Science. 2010;330(6007):1091–5. doi: 10.1126/science.1197410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Feng Z, Hou T, Li Y. Structure-based drug design for dopamine D3 receptor. Combinatorial chemistry & high throughput screening. 2012;15(10):775–91. doi: 10.2174/138620712803901135. [DOI] [PubMed] [Google Scholar]
- 45.Feng Z, Hou T, Li Y. Selectivity and activation of dopamine D3R from molecular dynamics. Journal of molecular modeling. 2012;18(12):5051–63. doi: 10.1007/s00894-012-1509-x. [DOI] [PubMed] [Google Scholar]
- 46.Masaru I, Yoshihisa K. Muscarinic Acetylcholine Receptors. Current Pharmaceutical Design. 2006;12(28):3573–81. doi: 10.2174/138161206778522056. [DOI] [PubMed] [Google Scholar]
- 47.Wishart DS, Knox C, Guo AC, Cheng D, Shrivastava S, Tzur D, Gautam B, Hassanali M. DrugBank: a knowledgebase for drugs, drug actions and drug targets. Nucleic acids research. 2008;36(suppl_1):D901–D6. doi: 10.1093/nar/gkm958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Library of Congress. Congressional Research S. Controlled Substances Act: Regulatory Requirements. 2008. [Google Scholar]
- 49.Aghajanian GK, Marek GJ. Serotonin and hallucinogens. Neuropsychopharmacology. 1999;21(2):16S–23S. doi: 10.1016/S0893-133X(98)00135-3. [DOI] [PubMed] [Google Scholar]
- 50.Quigley A, Waun J. Lysergic Acid Diethylamide (LSD) 2011:2704–5. [Google Scholar]
- 51.Pacher P, Nagayama T, Mukhopadhyay P, Bátkai S, Kass DA. Measurement of cardiac function using pressure–volume conductance catheter technique in mice and rats. Nature protocols. 2008;3(9):1422–34. doi: 10.1038/nprot.2008.138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Sheffler DJ, Roth BL. Salvinorin A: the magic mint‘hallucinogen finds a molecular target in the kappa opioid receptor. Trends in Pharmacological Sciences. 2003;24(3):107–9. doi: 10.1016/S0165-6147(03)00027-0. [DOI] [PubMed] [Google Scholar]
- 53.Berger SI, Iyengar R. Role of systems pharmacology in understanding drug adverse events. Wiley Interdisciplinary Reviews: Systems Biology and Medicine. 2011;3(2):129–35. doi: 10.1002/wsbm.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Boran AD, Iyengar R. Systems approaches to polypharmacology and drug discovery. Current opinion in drug discovery & development. 2010;13(3):297. [PMC free article] [PubMed] [Google Scholar]
- 55.Xie L, Xie L, Kinnings SL, Bourne PE. Novel computational approaches to polypharmacology as a means to define responses to individual drugs. Annual review of pharmacology and toxicology. 2012;52:361–79. doi: 10.1146/annurev-pharmtox-010611-134630. [DOI] [PubMed] [Google Scholar]
- 56.Metz JT, Hajduk PJ. Rational approaches to targeted polypharmacology: creating and navigating protein–ligand interaction networks. Current opinion in chemical biology. 2010;14(4):498–504. doi: 10.1016/j.cbpa.2010.06.166. [DOI] [PubMed] [Google Scholar]
- 57.Haga K, Kruse AC, Asada H, Yurugi-Kobayashi T, Shiroishi M, Zhang C, Weis WI, Okada T, Kobilka BK, Haga T. Structure of the human M2 muscarinic acetylcholine receptor bound to an antagonist. Nature. 2012;482(7386):547–51. doi: 10.1038/nature10753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.McPherson DW, Lambert CR, Jahn K, Sood V, McRee RC, Zeeberg B, Reba RC, Knapp FF., Jr Resolution and in Vitro and Initial in Vivo Evaluation of Isomers of Iodine-125-Labeled 1-Azabicyclo [2.2. 2] oct-3-yl. alpha.-Hydroxy-. alpha.-(1-iodo-1-propen-3-yl)-. alpha.-phenylacetate: A High-Affinity Ligand for the Muscarinic Receptor. Journal of medicinal chemistry. 1995;38(20):3908–17. doi: 10.1021/jm00020a004. [DOI] [PubMed] [Google Scholar]
- 59.Shire D, Calandra B, Delpech M, Dumont X, Kaghad M, Le Fur G, Caput D, Ferrara P. Structural features of the central cannabinoid CB1 receptor involved in the binding of the specific CB1 antagonist SR 141716A. Journal of Biological Chemistry. 1996;271(12):6941–6. doi: 10.1074/jbc.271.12.6941. [DOI] [PubMed] [Google Scholar]
- 60.Shearman LP, Stribling DS, Camacho RE, Rosko KM, Wang J, Tong S, Feng Y, Marsh DJ, Yu H, Guan X. Characterization of a novel and selective cannabinoid CB1 receptor inverse agonist, Imidazole 24b, in rodents. European journal of pharmacology. 2008;579(1):215–24. doi: 10.1016/j.ejphar.2007.10.033. [DOI] [PubMed] [Google Scholar]
- 61.Kapur A, Hurst DP, Fleischer D, Whitnell R, Thakur GA, Makriyannis A, Reggio PH, Abood ME. Mutation studies of Ser7. 39 and Ser2. 60 in the human CB1 cannabinoid receptor: evidence for a serine-induced bend in CB1 transmembrane helix 7. Molecular pharmacology. 2007;71(6):1512–24. doi: 10.1124/mol.107.034645. [DOI] [PubMed] [Google Scholar]
- 62.Kapur S, Mamo D. Half a century of antipsychotics and still a central role for dopamine D 2 receptors. Progress in Neuro-Psychopharmacology and Biological Psychiatry. 2003;27(7):1081–90. doi: 10.1016/j.pnpbp.2003.09.004. [DOI] [PubMed] [Google Scholar]
- 63.Lavalaye J, Linszen DH, Booij J, Reneman L, Gersons BP, van Royen EA. Dopamine D 2 receptor occupancy by olanzapine or risperidone in young patients with schizophrenia. Psychiatry Research: Neuroimaging. 1999;92(1):33–44. doi: 10.1016/s0925-4927(99)00032-3. [DOI] [PubMed] [Google Scholar]
- 64.Volkow ND. Imaging the addicted brain: from molecules to behavior. Journal of Nuclear Medicine. 2004;45(11):13N–24N. [PubMed] [Google Scholar]
- 65.Lauterbach EC, Cummings JL, Kuppuswamy PS. Toward a more precise, clinically—informed pathophysiology of pathological laughing and crying. Neuroscience & Biobehavioral Reviews. 2013;37(8):1893–916. doi: 10.1016/j.neubiorev.2013.03.002. [DOI] [PubMed] [Google Scholar]
- 66.Lane JR, Chubukov P, Liu W, Canals M, Cherezov V, Abagyan R, Stevens RC, Katritch V. Structure-based ligand discovery targeting orthosteric and allosteric pockets of dopamine receptors. Molecular pharmacology. 2013;84(6):794–807. doi: 10.1124/mol.113.088054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Vass M, Agai-Csongor E, Horti F, Keser GrM. Multiple Fragment Docking and Linking in Primary and Secondary Pockets of Dopamine Receptors. ACS medicinal chemistry letters. 2014;5(9):1010–4. doi: 10.1021/ml500201u. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Seeman P, Guan HC, Hirbec H. Dopamine D2High receptors stimulated by phencyclidines, lysergic acid diethylamide, salvinorin A, and modafinil. Synapse. 2009;63(8):698–704. doi: 10.1002/syn.20647. [DOI] [PubMed] [Google Scholar]
- 69.Ledent C, Valverde O, Cossu G, Petitet F, Aubert J-F, Beslot F, Böhme GA, Imperato A, Pedrazzini T, Roques BP. Unresponsiveness to cannabinoids and reduced addictive effects of opiates in CB1 receptor knockout mice. Science. 1999;283(5400):401–4. doi: 10.1126/science.283.5400.401. [DOI] [PubMed] [Google Scholar]
- 70.Nestler EJ. Is there a common molecular pathway for addiction? Nature neuroscience. 2005;8(11):1445–9. doi: 10.1038/nn1578. [DOI] [PubMed] [Google Scholar]
- 71.Volkow ND, Wang G-J, Fowler JS, Tomasi D. Addiction circuitry in the human brain. Annual review of pharmacology and toxicology. 2012;52:321. doi: 10.1146/annurev-pharmtox-010611-134625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Capasso R, Borrelli F, Cascio M, Aviello G, Huben K, Zjawiony J, Marini P, Romano B, Di Marzo V, Capasso F. Inhibitory effect of salvinorin A, from Salvia divinorum, on ileitis-induced hypermotility: cross-talk between m-opioid and cannabinoid CB1 receptors. British journal of pharmacology. 2008;155(5):681–9. doi: 10.1038/bjp.2008.294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Fichna J, Schicho R, Andrews C, Bashashati M, Klompus M, McKay D, Sharkey K, Zjawiony J, Janecka A, Storr M. Salvinorin A inhibits colonic transit and neurogenic ion transport in mice by activating κ-opioid and cannabinoid receptors. Neurogastroenterology & Motility. 2009;21(12):1326–e128. doi: 10.1111/j.1365-2982.2009.01369.x. [DOI] [PubMed] [Google Scholar]
- 74.Wess J. Muscarinic Acetylcholine Receptor Knockout Mice: Novel Phenotypes and Clinical Implications*. Annu Rev Pharmacol Toxicol. 2004;44:423–50. doi: 10.1146/annurev.pharmtox.44.101802.121622. [DOI] [PubMed] [Google Scholar]
- 75.Udaka F, Yamao S, Nagata H, Nakamura S, Kameyama M. Pathologic laughing and crying treated with levodopa. Archives of neurology. 1984;41(10):1095–6. doi: 10.1001/archneur.1984.04050210093023. [DOI] [PubMed] [Google Scholar]
- 76.Mariën P, De Smet HJ, Wijgerde E, Verhoeven J, Crols R, De Deyn PP. Posterior fossa syndrome in adults: a new case and comprehensive survey of the literature. Cortex. 2013;49(1):284–300. doi: 10.1016/j.cortex.2011.06.018. [DOI] [PubMed] [Google Scholar]
- 77.Schindehütte J, Trenkwalder C. Treatment of drug-induced psychosis in Parkinson's disease with ziprasidone can induce severe dose-dependent off-periods and pathological laughing. Clinical neurology and neurosurgery. 2007;109(2):188–91. doi: 10.1016/j.clineuro.2006.07.003. [DOI] [PubMed] [Google Scholar]
- 78.COHN R. FORCED CRYING AND LAUGHING: Treatment with Atropine and Atropine-like Agents. AMA Archives of Neurology & Psychiatry. 1951;66(6):738–43. doi: 10.1001/archneurpsyc.1951.02320120071009. [DOI] [PubMed] [Google Scholar]
- 79.Zellers GL, Frank M, Dougherty J. Pathological laughter. Annals of emergency medicine. 1990;19(3):327–9. doi: 10.1016/s0196-0644(05)82056-6. [DOI] [PubMed] [Google Scholar]