Abstracts
Background
The production of extended-spectrum β-lactamases (ESBLs) confer resistance to the commonly used beta-lactam antimicrobials and ESBL–producing bacteria render treatment difficulty in human and veterinary medicine. ESBL–producing bacteria have emerged in livestock in recent years, which may raise concerns regarding possible transfer of such bacteria through the food chain. The swine industry is important in Taiwan, but investigations regarding the status of ESBL in swine are limited.
Results
We collected 275 fecal swab samples from piglets with diarrhea in 16 swine farms located in central and southern Taiwan from January to December 2015 and screened them for ESBL–producing Escherichia coli. ESBL producers were confirmed phenotypically by combination disc test and genotypically by polymerase chain reaction and DNA sequencing. The occurrence rate of ESBL–producing E. coli was 19.7% (54 of 275), and all were obtained in swine farms located in southern Taiwan. bla CTX-M-1-group and bla CTX-M-9-group were the two bla CTX-M groups found. bla CTX-M-55 (34 of 54; 63.0%) and bla CTX-M-15 (16 of 54; 29.6%), which belong to the bla CTX-M-1-group, were the two major bla gene types, whereas bla CTX-M-65 was the only type found in the bla CTX-M-9 group. Twenty-seven strains contained bla TEM-1, and the other 27 strains contained bla TEM-116. One strain found in Pingtung harbored three bla genes: bla TEM-116, bla CTX-M-55, and bla CTX-M-65. ESBL–producing E. coli exhibited a multidrug-resistant phenotype, and multilocus sequence typing revealed that the ST10 clonal complexes, including ST10, 167, 44, and 617 accounted for 35% (19 of 54) of these strains.
Conclusions
ESBL-producing E. coli from piglets with diarrhea were isolated from swine farms located in southern Taiwan. The most commonly detected bla were bla CTX-M-15 and bla CTX-M-55. The ST10 clonal complexes comprised most of our ESBL-producing E. coli strains. Fecal shedding from swine may contaminate the environment, resulting in public health concerns; thus, continued surveillance of ESBL is essential in swine and in other food animals.
Keywords: Extended spectrum β-lactamase, Escherichia coli, Multilocus sequence typing
Background
Diarrhea is a common clinical syndrome in the swine industry and may be classified into three entities. Sucking piglets usually exhibit neonatal diarrhea a few days after birth, and young piglet diarrhea occurs from the first week after birth to weaning [1]. Older piglets, commonly 2 weeks after weaning, may contract post-weaning diarrhea. Neonatal and post-weaning diarrhea are caused by pathogenic Escherichia coli, and causative agents of young piglet diarrhea may include transmissible gastroenteritis virus, rotavirus, coccidia, and E. coli [1]. Occurrence of diarrheal disease can be reduced by the vaccination of sows to let piglets obtain maternal antibodies. However, measures such as the use of antibiotic supplements in feed are also frequently practiced along with vaccination to reduce the incidence of diarrhea. If prudent usage of antibiotics is not taken into consideration, the massive, indiscriminate, and long-term use of antibiotics in veterinary practice may contribute to the selection and spread of drug-resistant bacteria [2, 3].
Production of extended-spectrum β-lactamases (ESBLs) confers resistance to the frequently used beta-lactam antimicrobial agents, including the third-generation cephalosporins such as ceftriaxone, ceftazidime, and ceftiofur. However, ESBLs are inhibited by the β-lactamases inhibitors clavulanic acid, sulbactam, and tazobactam [4]. TEM, SHV, and CTX-M-types are the three major families of ESBL [4]. All CTX-M-types enzymes are ESBLs, whereas the TEM- and SHV- types of ESBL arise by point mutation at specific residues from the natural TEM-1/TEM-2 and SHV-1 β-lactamase [5]. The production of ESBLs is mainly plasmid mediated, and such plasmids often carry genes that encode resistance to other classes of antimicrobials, such as fluoroquinolones and aminoglycosides [6]. ESBLs are widely distributed in Enterobacteriaceae, particularly in E. coli, and the rapid emergence and spread of ESBL-producing E. coli have been reported in food animals globally [7]. Such findings raise concerns about the possible transfer of ESBL producers through the food chain, thus presenting a hazard to public health [8].
The status of ESBL-producing E. coli in food animals in Taiwan has only been reported in cows [9]. Although a foot and mouth disease outbreak in 1997 had a great impact on the swine industry [10], swine are still among the most important agricultural products in Taiwan. The objective of this study is to analyze the fecal carriage of ESBL-producing E. coli isolated from piglets with diarrhea in 16 pig farms located in central and southern Taiwan. It is important to screen for the ESBL producers from food animals such as swine from a public health perspective.
Methods
Sample collection
A total of 275 fecal swab samples were collected from the piglets with diarrhea before weaning from 16 swine farms in Taiwan (one in Taichung, one in Nantou, one in Chunghua, two in Yunlin, four in Chiayi, one in Tainan, and six in Pingtung) from January to December 2015. These 16 farms belong to the same swine industry corporation. Isolating E. coli from feces in piglets with diarrhea and preparing “tailored vaccine” has been routinely practiced in these farms. These E. coli were cultured, inactivated and used as a vaccine component to feed pregnant sows. Neonatal piglets will presumably obtain maternal antibodies when sucking colostrum. Occurrence of neonatal diarrhea due to E. coli infection may be reduced as long as piglets have enough maternal antibody. We shared these fecal swab samples and inoculated on CHROMagar ESBL (CHROMagar, Paris, France) to screen for ESBL-producing E. coli. Any pink colony that appeared on the agar after incubation at 37 °C for 16–18 h was initially designated as ESBL-producing E. coli since they were resistant to cefotaxime and/or ceftazidime, and its identity as E. coli was confirmed with the RapIDTM ONE System (RapIDTM, Lenexa, KS, USA). Confirmed E. coli strains were stored at −80 °C for further study.
ESBL testing
E. coli isolates were tested phenotypically for ESBL production by combination disc tests with cefotaxime and ceftazidime (30 μg), with and without clavulanic acid (10 μg), as stated by the guidelines of the Clinical and Laboratory Standards Institute [11]. The tested E. coli strains were plated on Muller-Hinton agar at a concentration of 0.5 McFarland standards and grown at 35 °C for 16–18 h. A difference of 5 mm or more in the inhibition zones for at least one cefotaxime or ceftazidime/clavulanic acid combination versus the corresponding cefotaxime or ceftazidime alone was used to define an ESBL producer. Klebsiella pneumoniae ATCC 700603 and E. coli ATCC 25922 were used as the positive and negative controls, respectively [11].
Detection of bla genes
The E. coli strains that were phenotypically confirmed to be ESBL producers were examined with polymerase chain reaction (PCR) to detect their bla genes. The tested strains were cultured for 16–18 h at 37 °C on tryptic soy agar plates (Difco/Becton Dickinson, Franklin Lakes, NJ, USA), and a loopful of bacterial cells was resuspended in 200 μL ddH2O and boiled for 10 min [12]. After centrifugation at 12,000 g for 10 min, the supernatant was saved as the source of template DNA for PCR. The primer sequences used to amplify bla TEM, bla SHV, bla CTX-M-1-group, bla CTX-M-2-group, bla CTX-M-8-group, bla CTX-M-9-group, and bla CTX-M-25-group, the annealing temperature, and the expected PCR product sizes are specified in Table 1. The PCR cycling program was set as follows using a LifeEco thermocycler (Bioer Technology, Hangzhou, China): initial denaturation at 95 °C for 5 min, followed by 35 cycles at 95 °C for 30 s, then the annealing temperature specified in Table 1 for 40 s, and 72 °C extension for 1 min. The reaction was then maintained at 72 °C for 10 min. Ten microliters of each PCR sample was loaded onto a 1.2% agarose gel and electrophoresed at 100 volts for 40 min. The gels were then stained with a fluorescent nucleic acid dye (Biotium, Hayward, CA, USA) and examined under a blue light LED illuminator (Smobio, Hsinchu City, Taiwan). The PCR products were then sliced from the agarose gel and subjected to further purification and sequenced by ABI 3130 x1 Genetic Analyzer (Applied Biosystems, Foster, CA, USA) in Center for Genomic Medicine, National Cheng Kung University, Tainan, Taiwan. The results were analyzed with MEGA 6.0 and examined with the NCBI BLAST program (http://www.ncbi.nlm.nih.gov/blast/) and β-lactamase database (http://www.ncbi.nlm.nih.gov/pathogens/submit-beta-lactamase).
Table 1.
Sequences of primers used for ESBL gene detection
| PCR target | primer | Sequences (5’–3’) | Annealing Tm (°C) | Predicted PCR size (bp) | Reference |
|---|---|---|---|---|---|
| bla TEM | TEM-F | TCGGGGAAATGTGCGCG | 55 | 972 | [37] |
| TEM-R | TGCTTAATCAGTGAGGCACC | ||||
| bla SHV | SHV-F | GCCTTTATCGGCCCTCACTCAA | 54 | 819 | [38] |
| SHV-R | TCCCGCAGATAAATCACCACAATG | ||||
| bla CTX-M-1-group | CTX-M-1-F | CCCATGGTTAAAAAATCACTGC | 54 | 942 | [39] |
| CTX-M-1-R | CAGCGCTTTTGCCGTCTAAG | ||||
| bla CTX-M-2-group | CTX-M-2-F | CGACGCTACCCCTGCTATT | 52 | 552 | [40] |
| CTX-M-2-R | CCAGCGTCAGATTTTTCAGG | ||||
| bla CTX-M-8-group | CTX-M-8-F | CAAAGAGAGTGCAACGGATG | 52 | 205 | [40] |
| CTX-M-8-R | ATTGGAAAGCGTTCATCACC | ||||
| bla CTX-M-9-group | CTX-M-9-F | ATGGTGACAAAGAGAGTGCAAC | 55 | 876 | [26] |
| CTX-M-9-R | TTACAGCCCTTCGGCGATGATT | ||||
| bla CTX-M-25-group | CTX-M-25-F | GCACGATGACATTCGGG | 52 | 327 | [40] |
| CTX-M-25-R | AACCCACGATGTGGGTAGC |
Antimicrobial susceptibility testing
The ESBL-producing E. coli strains were tested for susceptibility to antimicrobial agents using the disc agar diffusion method [11]. The antimicrobial agents tested included amikacin 30 μg, ampicillin 10 μg, amoxyclav 30 μg, ceftiofur 30 μg, cephalothin 30 μg, ciprofloxacin 10 μg, doxycycline 30 μg, enrofloxacin 5 μg, florfenicol 30 μg, gentamicin 30 μg, nalidixic acid 30 μg, streptomycin 10 μg, co-trimoxazole 25 μg, tetracycline 10 μg; all of the discs were purchased from Oxoid (Oxoid, Hampshire, UK).
E. coli genotyping
Our E. coli strains were analyzed genotypically by multilocus sequence typing. DNA fragments derived from adk, fumC, gyrB, icd, mdh, purA, and recA were amplified by PCR, sequenced, and then uploaded to the MLST website (http://enterobase.warwick.ac.uk/) for comparison [13]. Phylogenetic analysis was performed using BioNumerics Software version 7.0 (Applied Maths, Sint-Martens-Latem, Belgium).
Results
Fifty-four samples exhibited pink colonies on CHROMagar ESBL, initially indicating an identity of ESBL–producing E. coli. All of these strains were then confirmed biochemically with RapIDTM ONE System as E. coli, and they were not hemolytic when grown on blood agar. These 54 strains exhibited the ESBL phenotype when assayed by combination disc tests. From our results, we did not detect any ESBL–producing E. coli in diseased piglets from any of the five swine farms in Taichung, Nantou, Chunghua, and Yunlin, which were located in central Taiwan. ESBL–producing E. coli were all obtained in swine farms in southern Taiwan, including Chiayi, Tainan, and Pingtung, with the exception of one farm in Chiayi (farm ID CY-3) and one in Pingtung (farm ID PT-3). Geographic distribution of the swine farms and occurrence of ESBL were indicated in Fig. 1. Overall, the occurrence rate of ESBL-producing E. coli was 19.7% (54 of 275). Table 2 lists the occurrence of ESBL-producing E. coli in 16 swine farms.
Fig. 1.
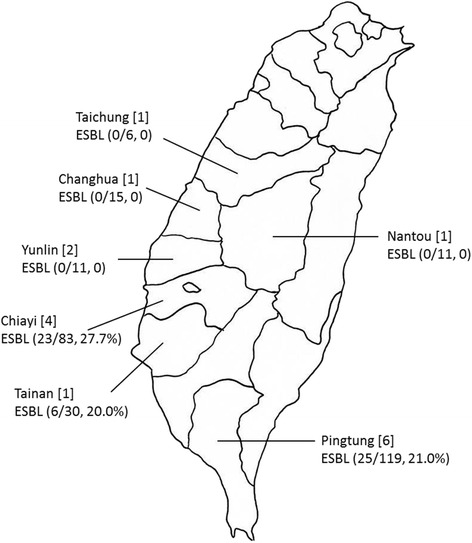
Geographic distribution of the swine farms in Taiwan included in this study. The number of square brackets indicates the number of swine farms included from each region. The number and occurrence rate of ESBL-producing E. coli are denoted in parentheses
Table 2.
Occurrence of ESBL–producing E. coli in 16 farms
| Farm location | Farm ID | No. of fecal samples | No. of ESBL-producing E. coli | Occurrence (%) |
|---|---|---|---|---|
| Taichung | TC-1 | 6 | 0 | 0 |
| Changhua | CH-1 | 15 | 0 | 0 |
| Nantou | NT-1 | 11 | 0 | 0 |
| Yunlin | YL-1 | 5 | 0 | 0 |
| YL-2 | 6 | 0 | 0 | |
| Chiayi | CY-1 | 40 | 19 | 47.5 |
| CY-2 | 8 | 2 | 25.0 | |
| CY-3 | 29 | 0 | 0 | |
| CY-4 | 6 | 2 | 33.3 | |
| Tainan | TN-1 | 30 | 6 | 20.0 |
| Pingtung | PT-1 | 10 | 2 | 20.0 |
| PT-2 | 36 | 16 | 44.4 | |
| PT-3 | 37 | 0 | 0 | |
| PT-4 | 17 | 2 | 11.8 | |
| PT-5 | 11 | 3 | 27.3 | |
| PT-6 | 8 | 2 | 25.0 |
Table 3 lists the bla genes and sequence type of ESBL-producing E. coli. bla CTX-M-1-group and bla CTX-M-9-group were the two bla CTX-M groups found in ESBL-producing E. coli. The bla CTX-M-1-group contained bla CTX-M-55 (34 of 54, 63.0%) and bla CTX-M-15 (16 of 54, 29.6%), whereas bla CTX-M-65 was the only type found from the bla CTX-M-9 group. All 54 strains contained bla TEM; 27 strains had bla TEM-1 and the other 27 strains contained bla TEM-116. One strain found in Pingtung harbored bla TEM-116, bla CTX-M-55, and bla CTX-M-65. The bla CTX-M-2-group, bla CTX-M-8-group, bla CTX-M-25-group, and bla SHV types of ESBL were not detected in this study.
Table 3.
The bla genes and sequence type of ESBL-producing E. coli
| Farm location | bla gene | Sequence Type |
|---|---|---|
| Chiayi | bla TEM-1 + bla CTX-M-15 (n = 9) | 4981 (n = 5), 1638 (n = 1), 3268 (n = 1), NTa (n = 2) |
| bla TEM-116 + bla CTX-M-15 (n = 3) | 1638 (n = 1), NT (n = 2) | |
| bla TEM-1 + bla CTX-M-55 (n = 5) | 167 (n = 4), NT (n = 1) | |
| bla TEM-6 + bla CTX-M-55 (n = 6) | 4981 (n = 1), 349 (n = 2), 44 (n = 2), NT (n = 1) | |
| Tainan | bla TEM-1 + bla CTX-M-55 (n = 2) | NT (n = 2) |
| bla TEM-116 + bla CTX-M-55 (n = 3) | 457 (n = 2), 617 (n = 1) | |
| bla TEM-116 (n = 1) | NT (n = 1) | |
| Pingtung | bla TEM-1 + bla CTX-M-15 (n = 2) | 617 (n = 2) |
| bla TEM-1 + bla CTX-M-55 (n = 9) | 10 (n = 3), 648 (n = 1), 69 (n = 3), 617 (n = 1), NT (n = 1) | |
| bla TEM-116 + bla CTX-M-15 (n = 2) | 10 (n = 1), 349 (n = 1) | |
| bla TEM-116 + bla CTX-M-55 (n = 8) | NT (n = 1), 10 (n = 1), 457 (n = 2), 617 (n = 1), 167 (n = 1), 69 (n = 1), 38 (n = 1) | |
| bla TEM-116 + bla CTX-M-55 + bla CTX-M-65 (n = 1) | NT (n = 1) | |
| bla TEM-116 (n = 3) | 44 (n = 1), 167 (n = 1), NT (n = 1) |
a: NT new type, there was no comparison standard in the databank
The results of the antibiotic susceptibility testing of the ESBL-producing E. coli isolated from Chiayi, Tainan, and Pingtung are shown in Table 4. The susceptibility testing showed that all 54 ESBL positive isolates were resistant to five antibiotics: ampicillin, cephalothin, ceftiofur, tetracycline, and enrofloxacin. Amikacin and gentamicin were active against 31 strains (57.4%) and 17 strains (31.5%) of ESBL producers, respectively. Overall, ESBL-producing E. coli exhibited a multi-drug-resistant phenotype.
Table 4.
Antimicrobial susceptibility test of ESBL–producing E. coli
| Chiayi, n = 23 (%) | Tainan, n = 6 (%) | Pingtung, n = 25 (%) | Total, N = 54 (%) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Antibiotic discs used | Sa | Ib | Rc | S | I | R | S | I | R | S | I | R |
| Ampicillin | 0 | 0 | 23 (100.0) | 0 | 0 | 6 (100.0) | 0 | 0 | 25 (100.0) | 0 | 0 | 54 (100.0) |
| Amoxicillin/clavulanic acid | 3 (13.0) | 8 (34.8) | 12 (52.2) | 2 (33.3) | 1 (16.7) | 3 (50.0) | 6 (24.0) | 11 (44.0) | 8 (32.0) | 11 (20.4) | 20 (37.0) | 23 (42.6) |
| Cephalothin | 0 | 0 | 23 (100.0) | 0 | 0 | 6 (100.0) | 0 | 0 | 25 (100.0) | 0 | 0 | 54 (100.0) |
| Ceftiofur | 0 | 0 | 23 (100.0) | 0 | 0 | 6 (100.0) | 0 | 0 | 25 (100.0) | 0 | 0 | 54 (100.0) |
| Amikacin | 15 (65.2) | 8 (34.8) | 0 | 5 (83.3) | 1 (16.7) | 0 | 11 (44.0) | 3 (12.0) | 11 (44.0) | 31 (57.4) | 12 (22.2) | 11 (20.4) |
| Gentamicin | 12 (52.2) | 3 (13.0) | 8 (34.8) | 1 (16.7) | 0 | 5 (83.3) | 4 (16.0) | 0 | 21 (84.0) | 17 (31.5) | 3 (5.6) | 34 (62.9) |
| Streptomycin | 0 | 1 (4.4) | 22 (95.6) | 0 | 0 | 6 (100.0) | 0 | 1 (4.0) | 24 (96.0) | 0 | 2 (3.7) | 52 (96.3) |
| Doxycycline | 3 (13.0) | 10 (43.5) | 10 (43.5) | 2 (33.3) | 2 (33.3) | 2 (33.3) | 2 (8.0) | 9 (36.0) | 14 (56.0) | 7 (13.0) | 21 (38.9) | 26 (48.1) |
| Tetracycline | 0 | 0 | 23 (100.0) | 0 | 0 | 6 (100.0) | 0 | 0 | 25 (100.0) | 0 | 0 | 54 (100.0) |
| Nalidixic acid | 1 (4.4) | 0 | 22 (95.6) | 0 | 0 | 6 (100.0) | 2 (8.0) | 0 | 23 (92.0) | 3 (5.6) | 0 | 51 (94.4) |
| Ciprofloxacin | 1 (4.4) | 0 | 22 (95.6) | 0 | 0 | 6 (100.0) | 3 (12.0) | 0 | 22 (88.0) | 4 (7.4) | 0 | 50 (92.6) |
| Enrofloxacin | 0 | 0 | 23 (100.0) | 0 | 0 | 6 (100.0) | 0 | 0 | 25 (100.0) | 0 | 0 | 54 (100.0) |
| Florfenicol | 0 | 1 (4.4) | 22 (95.6) | 0 | 0 | 6 (100.0) | 0 | 4 (16.0) | 21 (84.0) | 0 | 5 (9.3) | 49 (90.7) |
| Co-trimoxazole | 6 (26.1) | 0 | 17 (73.9) | 0 | 0 | 6 (100.0) | 2 (8.0) | 0 | 23 (92.0) | 8 (14.8) | 0 | 46 (85.2) |
a: susceptible; b: intermediate resistant; c: resistant
The most frequently seen sequence type of ESBL-producing E. coli was ST167 (ST10 clonal complex; 7 of 54; 13.0%), followed by ST4981 (6 of 54; 11.1%) and ST10 (ST10 clonal complex; 5 of 54; 9.3%). There were four strains of ST617 (ST10 clonal complex, 4/54, 7.4%), ST457, and ST69 (ST69 clonal complex) and three strains of ST44 (ST10 clonal complex; 3 of 54; 5.6%) and ST349 (ST349 clonal complex). ST1638 had two strains (2 of 54; 3.7%). ST38 (ST38 clonal complex), ST3268, and ST648 (ST648 clonal complex) had only one strain each. Nonetheless, we still had 13 strains whose sequence types were not matched to any type in the current databank. Figure 2 indicates the minimal spanning tree of the ESBL-producing E. coli STs based on the degree of allele sharing.
Fig. 2.
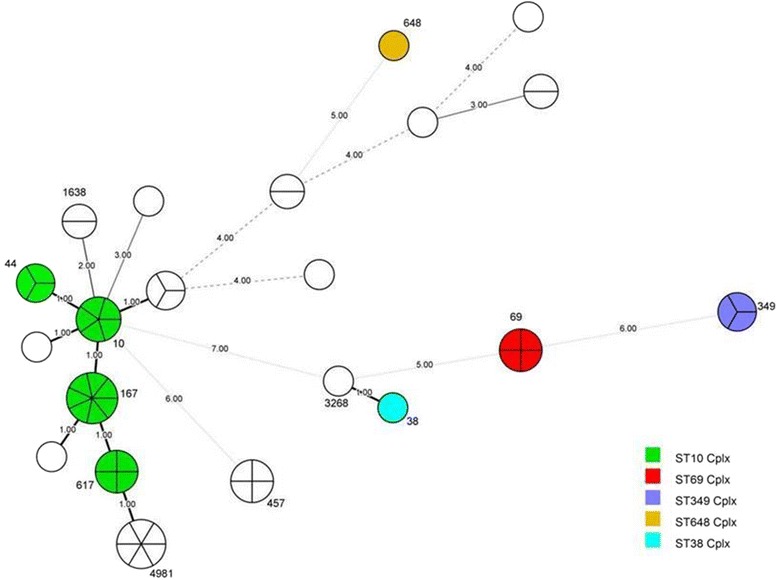
Minimal spanning tree (MSTree) of ESBL-producing E. coli. Each circle indicates one ST, subdivided into one sector for each isolate, and bordered by the ST number. White circles or sectors without an ST number denote a lack of comparison standard in the current databank. The numbers on the connecting line between STs within the MSTree indicate the number of different alleles. Solid lines represent an allele difference of 3 or less, whereas dotted lines and faint lines indicate an allele difference of 4 or more
Discussion
The ESBL-producing E. coli were all obtained from the swine farms located in southern Taiwan. There were five swine farms in central Taiwan (Taichung, Changhua, Yunlin, and Nantou) that participated in our study, and only 43 fecal samples (43 of 275; 15.6%) were collected from piglets with diarrhea and screened for ESBL. Although the scale of these farms was similar to that of those located in southern Taiwan, the hygienic procedures or disease control management of individual farms may contribute to such differences in diarrheal cases. The specificity of this chromogenic agar was 100% because all of the pink colonies, indicative of ESBL-producing E. coli, were phenotypically and genotypically positive for ESBL. A previous report also suggested the high sensitivity and specificity of CHROMagar ESBL in the detection of clinical ESBL-producing Enterobacteriaceae [14]. However, our strategy may also lose some ESBL producers that could grow on blood agar or MacConkey agar but not on CHROMagar ESBL.
The occurrence of ESBL-producing E. coli in food animals has been increasing around the world [2]. For example, more than 40% of the ESBL-producing E. coli were detected from piglets with post-weaning diarrhea in Heilongjiang Province, China [15]. The authors also compared their findings with those of a similar study in healthy pigs in China and concluded that ESBL-producing E. coli were more commonly found in sick animals [16]. Because we did not investigate the prevalence of ESBL in a healthy swine population, there was no basis of comparison for healthy and diseased swine in Taiwan. Although diseased pigs are not likely to enter slaughter or market, fecal shedding from such pigs can contaminate the piggery environment and provide a reservoir for the exchange of drug-resistance genes [17].
TEM-116–producing E. coli was first identified in Korean hospitals in a nationwide survey in 2002 [18]. Consequently, a high prevalence of TEM-116 was also reported in Spain [19]. In animals, TEM-116–producing E. coli has been detected in dogs [20, 21]. Our results, to the best of our knowledge, demonstrate for the first time the presence of bla TEM-116 genes in the ESBL–producing E. coli from porcine origin. Although most of our TEM-116–containing strains also had CTX-M, we did find four E. coli strains that harbored only TEM-116 that exhibited an ESBL phenotype. ESBL producers within the CTX-M group are becoming more common [22, 23]. In Europe, CTX-M-1 is broadly disseminated in animals, whereas CTX-M-14 is most prevalent in animals in Asian countries [8]. The most frequently found CTX-M types in our study were CTX-M-15 and CTX-M-55, whereas CTX-M-14 was reported in healthy and diseased swine in Korea and China [15, 16, 24]. CTX-M-55 was first isolated from patients in a hospital in Thailand; this novel CTX-M type was derived from CTX-M-15, with only a single substitution of valine instead of alanine at residue 77 [25]. The incidence of CTX-M-55 has been reported to exceed that of CTX-M-15 in outpatient infection cases in Chinese county hospitals [26]. Thus, the authors of that study hypothesized an animal-human transfer of CTX-M-55 because most of these outpatients in county hospitals live in rural areas and thus have more chances to come into contact with infected food animals and farm sewage [26]. In our study, CTX-M-55 was the predominant ESBL type isolated from swine with diarrhea in southern Taiwan. CTX-M-55–producing E. coli has also been detected in the milk of cows with clinical mastitis in the same region [9]. It is conceivable that ESBL-producing E. coli that possess CTX-M-55 have spread to the environment. One strain obtained in Pingtung possessed CTX-M-65 in addition to CTX-M-55 and TEM-116. Although the detection rate of CTX-M-65 was low compared to those of CTX-M-55 and CTX-M-15, the presence of CTX-M-65 has been reported in humans, animals, and vegetables [15, 24, 27]. These findings underscore the importance of screening and investigation of the genotypes of ESBL producers in food animals on a regular basis. Our investigation did not detect any CTX-M-8 or CTX-M-25, which were also not detected in previous studies [15, 28].
Antimicrobial susceptibility test revealed a high frequency of the resistance of ESBL-producing E. coli to most antimicrobial agents. Inappropriate use or overuse of antimicrobial agents, including third-generation cephalosporins, may be associated with the emergence of ESBL-producing E. coli in swine [29]. The selection of CTX-M-producing E. coli in swine by treatment with ceftiofur has been documented [30]. It is worthwhile to consider banning the use of third-generation of cephalosporins such as ceftiofur in food animals to decrease the occurrence of ESBL producers. For example, the occurrence of ESBL-producing E. coli was reduced when third-generation cephalosporins were banned in the Danish pig industry [31]. Forty-two percent of ESBL-producing E. coli were resistant to amoxicillin/clavulanic acid. Possible reasons that account for this phenotype may include hyper production of chromosomal class C β-lactamase, possession of plasmid-mediated TEM enzymes, production of oxacillinases, or production of inhibitor-resistant TEM by these isolates [32]. In addition, plasmid mediated AmpC may also cause resistance to amoxicillin/clavulanic acid [8].
The ST10 clonal complexes (ST10, 167, 44, 617) comprised most of our ESBL-producing E. coli strains. There were 13 strains that did not match any ST in the current database; however, six of these had only a one to three-allele difference from ST10 clonal complexes. It is fair to say that ST10 was the dominant clonal complex in our study. A recent investigation indicated that ESBL-producing E. coli were commonly isolated from river waters in southern Taiwan and that ST10 and ST58 was the most frequently found clonal complexes [33]. The authors of that study also observed a substantial association of these ESBL-producing E. coli with the presence of chicken farms at that region. Geographically, food animal farms, including swine, chicken, and cattle, are primarily situated in southern Taiwan. It is conceivable that livestock may spread ESBL-producing E. coli from feces, thus contaminating the environment. We did not detect ESBL-producing E. coli ST131 (O25:H4) that possessed CTX-M-15, a leading cause of urinary tract infections and bacteremia in human medicine globally, in our study. However, swine and other food animals may play a role as vectors in the transmission of bacteria to humans [34], so continued surveillance of food animals for ESBL-producing E. coli is essential.
Our study has some limitations. Fecal samples from healthy piglets were not collected and there was no comparison for the occurrence of ESBL-producing E. coli between the healthy and diseased populations. The virulence factors like K88, K99 or 987 P fimbriae genes in our ESBL-producing E. coli isolates were not screened and they were not hemolytic when grown on blood agar. It is possible that the ESBL-producing E. coli in the present study was not the causative agent for the diarrhea of these piglets. We did not detect if these E. coli isolates produced AmpC-β-lactamases, which also hydrolyze third-generation cephalosporins. AmpC-producing E. coli were also found in increasing numbers in food-producing animals [8]. In addition, profiles of resistant plasmids were not characterized in our study. Plasmid analysis methods like PCR-based replicon typing could assign the incompatibility (Inc) groups [35], whereas replicon sequence typing could discriminate IncF plasmid variants [36]. Inclusion of plasmid characterization could have provided insights into the epidemiology of the ESBL plasmid in our study.
Conclusions
ESBL-producing E. coli from piglets with diarrhea were isolated from swine farms located in Chiayi, Tainan, and Pingtung. bla CTX-M-15 and bla CTX-M-55 were the most commonly detected bla genes. The ST10 clonal complexes comprised most of our ESBL-producing E. coli strains. Fecal shedding from swine may contaminate the environment, from a public health perspective, continued surveillance of ESBL is essential in swine and in other food animals.
Acknowledgements
The authors would like to thank Hsiuo-Tung Yeh for Fig. 1 graphic drawing and Dr. Lee-Jene Teng, Department of Clinical Laboratory Sciences and Medical Biotechnology, National Taiwan University, for providing Klebsiella pneumoniae ATCC 700603.
Funding
This research was supported by National Taiwan University, No. G049919.
Availability of data and materials
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
Authors’ contributions
W-CL performed all the experiments and was a major contributor in writing up this manuscript. K-SY coordinated this study and also helped draft this manuscript. Both authors read and approved the final manuscript.
Competing interests
The authors declare that they have no competing interests.
Consent for publication
Not applicable.
Ethics approval and consent to participate
Ethics approval was not applicable. Animal handling, including fecal sample collection, was performed or supervised by the approved veterinarians throughout routine veterinary health management. Consent was obtained for the samples to be collected at each farm.
Abbreviations
- ATCC
American type culture collection
- BLAST
Basic local alignment search tool
- CTX-M
Cefotaximase-Munich
- ESBL
Extended-spectrum β-lactamase
- h
hour
- MEGA 6.0
Molecular evolutionary genetics analysis software version 6.0
- min
minute
- MLST
Multilocus sequence typing
- NCBI
National center for biotechnology information
- PCR
Polymerase chain reaction
- s
second
- SHV
Sulfhydryl variable
- ST
Sequence type
- TEM
Temoneira
Contributor Information
Wan-Chen Lee, Email: r02629007@ntu.edu.tw.
Kuang-Sheng Yeh, Phone: 886-2-33661289, Email: ksyeh@ntu.edu.tw.
References
- 1.Biehl LG, Hoefling DC. Diagnosis, treatment, and prevention of diarrhea in 7-to 14-day-old pigs. J Am Vet Med Assoc. 1986;188:1144–1146. [PubMed] [Google Scholar]
- 2.Seiffert SN, Hilty M, Perreten V, Endimiani A. Extended-spectrum cephalosporin-resistant gram-negative organisms in livestock; an emerging problem for human health. Drug Resist Updat. 2013;16:22–45. doi: 10.1016/j.drup.2012.12.001. [DOI] [PubMed] [Google Scholar]
- 3.Marshall BM, Levy SB. Food animals and antimicrobials: impacts on human health. Clin Microbiol Rev. 2011;24:718–733. doi: 10.1128/CMR.00002-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bush K, Jacoby GA. Updated functional classification of β-lactamases. Antimicrob Agents Chemother. 2010;54:969–976. doi: 10.1128/AAC.01009-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Gniadkowski M. Evolution of extended-spectrum β-lactamase by mutation. Clin Microbiol Infect. 2008;14(Suppl. 1):11–32. doi: 10.1111/j.1469-0691.2007.01854.x. [DOI] [PubMed] [Google Scholar]
- 6.Pitout JD, Laupland KB. Extended-spectrum β-lactamase-producing Enterobacteriaceae: an emerging public-health concern. Lancet Infect Dis. 2008;8:159–166. doi: 10.1016/S1473-3099(08)70041-0. [DOI] [PubMed] [Google Scholar]
- 7.Smet A, Martel A, Persoons D, Dewulf J, Heyndrickx M, Herman L, Haesebrouck F, Butaye P. Broad-spectrum β-lactamases among Enterobacteriaceae of animal origin: molecular aspects, mobility and impact on public health. FEMS Microbiol Rev. 2010;34:295–316. doi: 10.1111/j.1574-6976.2009.00198.x. [DOI] [PubMed] [Google Scholar]
- 8.Ewers C, Bethe A, Semmler T, Guenther S, Wieler LH. Extended-spectrum β-lactamase-producing and AmpC-producing Escherichia coli from livestock and companion animals, and their putative impact on public health: a global perspective. Clin Microbiol Infect. 2012;18:646–655. doi: 10.1111/j.1469-0691.2012.03850.x. [DOI] [PubMed] [Google Scholar]
- 9.Su Y, Yu CY, Tsai Y, Wang SH, Lee C, Chu C. Fluoroquinolone-resistant and extended-spectrum beta-lactamase-producing Escherichia coli from the milk of cows with clinical mastitis in Southern Taiwan. J Microbiol Immunol Infect. 2014. doi: 1016/j.jmii.2014.1010.1003. [DOI] [PubMed]
- 10.Shieh HK. The FMD, situiation in Taiwan. J Chin Soc Vet Sci. 1997;23(5):395–402. [Google Scholar]
- 11.CLSI . Performance standards for antimicrobial susceptibility testing: twenty-first informational supplement M100-21. Wayne, PA, USA: CLSI; 2011. [Google Scholar]
- 12.Shaheen BW, Oyarzabal OA, Boothe DM. The role of class 1 and 2 integrons in mediating antimicrobial resistance among canine and feline clinical E. coli isolates from the US. Vet Microbiol. 2010;144:363–370. doi: 10.1016/j.vetmic.2010.01.018. [DOI] [PubMed] [Google Scholar]
- 13.Wirth T, Falush D, Lan R, Colles F, Mensa P, Wieler LH, Karch H, Reeves PR, Maiden MCJ, Ochman H, et al. Sex and virulence in Escherichia coli: an evolutionary perspective. Mol Microbiol. 2006;60:1136–1151. doi: 10.1111/j.1365-2958.2006.05172.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Saito R, Koyano S, Nagai R, Okamura N, Moriya K, JKoike K. Evaluation of a chromogenic agar medium for the detection of extended-spectrum β-lactamase-producing Enterobacteriaceae. Lett Appl Microbiol. 2010;51:704–706. doi: 10.1111/j.1472-765X.2010.02945.x. [DOI] [PubMed] [Google Scholar]
- 15.Xu G, An W, Wang H, Zhang X. Prevalence and characteristics of extended-spectrum β-lactamase genes in Escherichia coli isolated from piglets with post-weaning diarrhea in Heilongjiang province, China. Front Microbiol. 2015;6:1103. doi: 10.3389/fmicb.2015.01103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zheng H, Zeng Z, Chen S, Liu Y, Yao Q, Deng Y, Chen X, Lv L, Zhuo C, Chen Z, et al. Prevalence and characterisation of CTX-M β-lactamases amongst Escherichia coli isolates from healthy food animals in China. Int J Antimicrob Agents. 2012;39:305–310. doi: 10.1016/j.ijantimicag.2011.12.001. [DOI] [PubMed] [Google Scholar]
- 17.Whitehead TR, Cotta MA. Stored swine manure and swine faeces as reservoirs of antibiotic resistance genes. Lett Appl Microbiol. 2013;56:264–267. doi: 10.1111/lam.12043. [DOI] [PubMed] [Google Scholar]
- 18.Jeong SH, Bae IK, Lee JH, Sohn SG, Kang GH, Jeon GJ, Kim YH, Jeong BC, Lee SH. Molecular characterization of extended-spectrum beta-lactamases produced by clinical isolates of Klebsiella pneumoniae and Escherichia coli from a korean nationwide survey. J Clin Microbiol. 2004;42:2902–2906. doi: 10.1128/JCM.42.7.2902-2906.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Romero EDV, Padilla TP, Hernández AH, Grande RP, Vázquez MF, García IG, García-Rodríguez JA, Bellido JLM. Prevalence of clinical isolates of Escherichia coli and Klebsiella spp. producing multiple extended-spectrum β-lactamases. Diagn Microbiol Infect Dis. 2007;59:433–437. doi: 10.1016/j.diagmicrobio.2007.06.007. [DOI] [PubMed] [Google Scholar]
- 20.Ewers C, Grobbel M, Stamm I, Kopp PA, Diehl I, Semmler T, Fruth A, Beutlich J, Guerra B, Wieler LH, et al. Emergence of human pandemic O25: H4-ST131 CTX-M-15 extended-spectrum- β-lactamase-producing Escherichia coli among companion animals. J Antimicrob Chemother. 2010;65:651–660. doi: 10.1093/jac/dkq004. [DOI] [PubMed] [Google Scholar]
- 21.Rzewuska M, Stefanska I, Kizerwetter-Swida M, Chrobak-Chmiel D, Szczygielska P, Lesniak M, Binek M. Characterization of extended-spectrum-β-lactamases produced by Escherichia coli strains isolated from dogs in Poland. Pol J Microbiol. 2015;64(3):285–288. [PubMed] [Google Scholar]
- 22.Zhao WH, Hu ZQ. Epidemiology and genetics of CTX-M extended-spectrum β-lactamases in Gram-negative bacteria. Crit Rev Microbiol. 2013;39(1):79–101. doi: 10.3109/1040841X.2012.691460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Bonnet R. Growing group of extended-spectrum betya-lactamases: the CTX-M enzymes. Antimicrob Agents Chemother. 2004;48:11–14. doi: 10.1128/AAC.48.1.1-14.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Tamang MD, Nam HM, Kim SR, Chae MH, Jang GC, Jung SC, Lim SK. Prevalence and molecular characterization of CTX-M β-lactamase-producing Escherichia coli isolated from healthy swine and cattle. Foodborne Pathog Dis. 2013;10:13–20. doi: 10.1089/fpd.2012.1245. [DOI] [PubMed] [Google Scholar]
- 25.Kiratisin P, Apisarnthanarak A, Saifon P, Laesripa C, Kitphati R, Mundy LM. The emergence of a novel ceftazidime-resistant CTX-M extended-spectrum beta-lactamase, CTX-M-55, in both community-onset and hospital-acquired infections in Thailand. Diagn Microbiol Infect Dis. 2007;58:349–355. doi: 10.1016/j.diagmicrobio.2007.02.005. [DOI] [PubMed] [Google Scholar]
- 26.Zhang J, Zheng B, Zhao L, Wei Z, Ji J, Li L, Xiao Y. Nationwide high prevalence of CTX-M and an increase of CTX-M-55 in Escherichia coli isolated from patients with community-onset infections in Chinese county hospitals. BMC Infect Dis. 2014;14:659–669. doi: 10.1186/s12879-014-0659-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Riccobono E, Di Pilato V, Di Maggio T, Revollo C, Bartoloni A, Pallecchi L, Rossolini GM. Characterization of IncI1 sequence type 71 epidemic plasmid lineage responsible for the recent dissemination of CTX-M-65 extended-spectrum beta-lactamase in the Bolivian Chaco region. Antimicrob Agents Chemother. 2007;59:5340–5347. doi: 10.1128/AAC.00589-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Geser N, Stephan R, Hächler H. Occurrence and characteristics of extended-spectrum β-lactamase (ESBL) producing Enterobacteriaceae in food producing animals, minced meat and raw milk. BMC Vet Res. 2012;8:21. doi: 10.1186/1746-6148-8-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Barton MD. Impact of antibiotic use in the swine industry. Curr Opin Microbiol. 2014;19:9–15. doi: 10.1016/j.mib.2014.05.017. [DOI] [PubMed] [Google Scholar]
- 30.Cavaco LM, Abatih E, Aarestrup FM, Guardabassi L. Selection and persistence of CTX-M-producing Escherichia coli in the intestinal flora of pigs treated with amoxicillin, ceftiofur, or cefquinome. Antimicrob Agents Chemother. 2008;52:3612–3616. doi: 10.1128/AAC.00354-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Agersø Y, Aarestrup FM. Voluntary ban on cephalosporin use in Danish pig production has effectively reduced extended-spectrum cephalosporinase-producing Escherichia coli in slaughter pigs. J Antimicrob Chemother. 2013;68:569–572. doi: 10.1093/jac/dks427. [DOI] [PubMed] [Google Scholar]
- 32.Leflon-Guibout V, Speldooren V, Heym B, Nicolas-Chanoine M-H. Epidemiological survey of amoxicillin-clavulanate resistance and corresponding molecular mechanisms in Escherichia coli isolates in France: new genetic features of blaTEM genes. Antimicrob Agents Chemother. 2000;44:2709–2714. doi: 10.1128/AAC.44.10.2709-2714.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Chen PA, Hung CH, Huang PC, Chen JR, Huang IF, Chen WL, Chiou YH, Hung WY, Wang JL, Cheng MF. Characteristics of CTX-M extended-spectrum β-lactamase-producing Escherichia coli strains isolated from multiple rivers in Southern Taiwan. Appl Environ Microbiol. 2016;82(6):1889–1897. doi: 10.1128/AEM.03222-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Dahms C, Hübner N, Kossow A, Mellmann A, Dittmann K, Kramer A. Occurrence of ESBL-producing Escherichia coli in livestock and farm workers in Mecklenburg-Western Pomerania, Germany. PLoS One. 2015;10:e0143326. doi: 10.1371/journal.pone.0143326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Carattoli A, Bertini A, Villa L, Falbo V, Hopkins KL, Threlfall EJ. Identification of plasmids by PCR-based replicon typing. J Microbiol Methods. 2005;63:219–228. doi: 10.1016/j.mimet.2005.03.018. [DOI] [PubMed] [Google Scholar]
- 36.Villa L, García-Fernández A, Fortini D, Carattoli A. Replicon sequence typing of IncF plasmids carrying virulence and resistance determinants. J Antimicrob Chemother. 2010;65:2518–2529. doi: 10.1093/jac/dkq347. [DOI] [PubMed] [Google Scholar]
- 37.Stucliffe JG. Nucleotide-sequence of ampicillin resistance gene of Escherichia coli plasmid pBR322. Proc Natl Acad Sci U S A. 1978;75:3737–3741. doi: 10.1073/pnas.75.8.3737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Chia JH, Chu C, Su LH, Chiu CH, Kuo AJ, Sun CF, Wu TL. Development of a multiplex PCR and SHV melting-curve mutation detection system for detection of some SHV and CTX-M-lactamases of Escherichia coli, Klebsiella pneumoniae, and Enterobacter cloacae in Taiwan. J Clin Microbiol. 2005;43:4486–4491. doi: 10.1128/JCM.43.9.4486-4491.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yu Y, Ji S, Chen Y, Zhou W, Wei Z, Li L, Ma Y. Resistance of strains producing extended-spectrum β-lactamases and genotype distribution in China. J Infect. 2007;54:53–57. doi: 10.1016/j.jinf.2006.01.014. [DOI] [PubMed] [Google Scholar]
- 40.Woodford N, fagan EJ, Ellington MJ. Multiplex PCR for rapid detection of genes encoding CTX-M extended-spectrum β-lactamases. J Antimicrob Chemother. 2006;57:154–155. doi: 10.1093/jac/dki412. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.


