Abstract
Colorectal tumors arise with genomic and epigenomic alterations through interactions between neoplastic cells, immune cells, and microbiota that vary along the proximal to distal axis of colorectum. Long interspersed nucleotide element-1 (LINE-1) hypomethylation in colorectal cancer has been associated with worse clinical outcome. Utilizing 1,317 colon and rectal carcinoma cases in two U.S.-nationwide prospective cohort studies, we examined patient survival according to LINE-1 methylation level stratified by tumor location. Cox proportional hazards model was used to assess a statistical interaction between LINE-1 methylation level and tumor location in colorectal cancer-specific mortality analysis, controlling for potential confounders including microsatellite instability, CpG island methylator phenotype, and KRAS, BRAF, and PIK3CA mutations. A statistically significant interaction was found between LINE-1 methylation level and tumor location in colorectal cancer-specific mortality analysis (Pinteraction = 0.011). The association of LINE-1 hypomethylation with higher colorectal cancer-specific mortality was stronger in proximal colon cancers (multivariable hazard ratio [HR], 1.66; 95% confidence interval [CI], 1.21 to 2.28) than in distal colon cancers (multivariable HR, 1.18; 95% CI, 0.81 to 1.72) or rectal cancers (multivariable HR, 0.87; 95% CI, 0.57 to 1.34). Our data suggest the interactive effect of LINE-1 methylation level and colorectal cancer location on clinical outcome.
Keywords: epigenetics, left-sided, molecular pathological epidemiology, prognosis, right-sided
INTRODUCTION
Evidence suggests that colorectal tumors arise with sets of genomic and epigenomic alterations through interactions between neoplastic cells, immune cells, and microbiota that vary along the proximal to distal axis of colorectum [1–6]. Consistent with a continuous change in the intestinal microbiota and luminal contents along the bowel subsites, host immunity against colorectal tumors and proportions of colorectal cancers with specific molecular features of colorectal cancer such as microsatellite instability (MSI), high-level CpG island methylator phenotype (CIMP), and BRAF and PIK3CA mutations change along the bowel subsites [7–12].
Methylation status of the long interspersed nucleotide element-1 (LINE-1), which constitutes approximately 18% of the entire human genome, serves as a surrogate for overall cellular DNA methylation status [13, 14]. The genome-wide DNA hypomethylation has been associated with an increased chromosomal instability that may cause low-level antitumor immunity in colorectal cancer [15–20]. In fact, LINE-1 hypomethylation in colorectal cancer has been associated with a lower density of T cells in tumor tissue and worse clinical outcome [21–23]. Studies have shown that the prognostic association of tumor LINE-1 methylation level in colorectal cancer differs by MSI status [24, 25]. We hypothesized that the prognostic association of LINE-1 hypomethylation in colorectal cancer might differ by tumor location.
To test this hypothesis, we utilized resources of 1,317 colorectal cancer cases in two U.S.-nationwide prospective cohort studies (the Nurses’ Health Study [NHS] and the Health Professionals Follow-up Study [HPFS]), and examined the interactive association of LINE-1 methylation level and tumor location in colorectal cancer mortality analysis, controlling for potential confounders including major molecular features of colorectal cancer. A better understanding of the prognostic association of tumor LINE-1 hypomethylation according to colorectal cancer location may offer new insights into the pathogenesis of colorectal cancer.
RESULTS
Tumor LINE-1 methylation level and colorectal cancer location
We measured tumor LINE-1 methylation level (ranging from 23.1 to 93.8% of 0 to 100% scale; mean 63.4%; standard deviation 9.8%) among 1,317 colon and rectal cancer cases within the NHS and the HPFS. Table 1 summarizes clinical, pathological, and tumor molecular features according to tumor LINE-1 methylation level in colorectal cancer. Of the 1,317 cases, 621 (47%) had proximal colon cancer, 409 (31%) had distal colon cancer, and 287 (22%) had rectal cancer. Low-level tumor LINE-1 methylation was associated with higher pN stage and metastatic disease, and inversely associated with poor tumor differentiation, MSI-high, MLH1 hypermethylation, CIMP-high, and BRAF mutation (P ≤ 0.003 with the adjusted α level of 0.003 for multiple hypothesis testing).
Table 1. Clinical, pathological, and tumor molecular features according to tumor LINE-1 methylation level in colorectal cancer.
| Characteristica | Total No. (n = 1,317) | Tumor LINE-1 methylation level | P valueb | ||
|---|---|---|---|---|---|
| High (≥65%) (n = 579) | Intermediate (55-64.9%) (n = 496) | Low (<55%) (n = 242) | |||
| Mean age ± SD (year) | 68.9 ± 8.8 | 69.9 ± 8.6 | 68.7 ± 8.7 | 67.1 ± 9.1 | 0.0001 |
| Sex | 0.09 | ||||
| Men | 593 (45%) | 255 (44%) | 214 (43%) | 124 (51%) | |
| Women | 724 (55%) | 324 (56%) | 282 (57%) | 118 (49%) | |
| Year of diagnosis | < 0.0001 | ||||
| Prior to 1995 | 468 (36%) | 162 (28%) | 202 (40%) | 104 (43%) | |
| 1996 to 2000 | 402 (30%) | 157 (27%) | 147 (30%) | 98 (40%) | |
| 2001 to 2008 | 447 (34%) | 260 (45%) | 147 (30%) | 40 (17%) | |
| Family history of colorectal cancer in a first-degree relative | 0.35 | ||||
| Absent | 1,048 (80%) | 471 (81%) | 391 (79%) | 186 (77%) | |
| Present | 264 (20%) | 107 (19%) | 102 (21%) | 55 (23%) | |
| Tumor location | 0.004 | ||||
| Proximal colon | 621 (47%) | 301 (52%) | 224 (45%) | 96 (40%) | |
| Distal colon | 409 (31%) | 151 (26%) | 166 (34%) | 92 (38%) | |
| Rectum | 287 (22%) | 127 (22%) | 106 (21%) | 54 (22%) | |
| Tumor differentiation | 0.002 | ||||
| Well to moderate | 1,178 (90%) | 501 (87%) | 461 (94%) | 216 (90%) | |
| Poor | 131 (10%) | 74 (13%) | 32 (6.5%) | 25 (10%) | |
| pT stage (depth of tumour invasion) | 0.45 | ||||
| pT1 (submucosa) | 143 (12%) | 62 (12%) | 63 (14%) | 18 (8.2%) | |
| pT2 (muscularis propria) | 249 (21%) | 115 (22%) | 90 (19%) | 44 (20%) | |
| pT3 (subserosa) | 757 (62%) | 327 (62%) | 288 (62%) | 142 (65%) | |
| pT4 (serosa or other organs) | 64 (5.3%) | 26 (4.9%) | 23 (5.0%) | 15 (6.9%) | |
| pN stage (number of positive lymph nodes) | 0.003 | ||||
| pN0 (0) | 740 (63%) | 348 (68%) | 280 (62%) | 112 (54%) | |
| pN1 (1-3) | 268 (23%) | 108 (21%) | 105 (23%) | 55 (26%) | |
| pN2 (≥4) | 163 (14%) | 56 (11%) | 65 (15%) | 42 (20%) | |
| TNM stagec | 0.0001 | ||||
| I | 308 (26%) | 141 (27%) | 124 (27%) | 43 (19%) | |
| II | 385 (32%) | 186 (36%) | 141 (31%) | 58 (26%) | |
| III | 342 (28%) | 131 (25%) | 137 (30%) | 74 (33%) | |
| IV | 168 (14%) | 60 (12%) | 58 (12%) | 50 (22%) | |
| MSI status | < 0.0001 | ||||
| MSI-low/MSS | 1,071 (84%) | 416 (75%) | 433 (89%) | 222 (93%) | |
| MSI-high | 209 (16%) | 141 (25%) | 51 (11%) | 17 (7.1%) | |
| MLH1 hypermethylation | < 0.0001 | ||||
| Absent | 1,073 (86%) | 440 (79%) | 413 (89%) | 220 (94%) | |
| Present | 179 (14%) | 116 (21%) | 50 (11%) | 13 (5.6%) | |
| CIMP status | < 0.0001 | ||||
| Low/negative | 1,032 (82%) | 412 (74%) | 403 (87%) | 217 (93%) | |
| High | 220 (18%) | 144 (26%) | 60 (13%) | 16 (6.9%) | |
| BRAF mutation | < 0.0001 | ||||
| Wild-type | 1,095 (85%) | 440 (79%) | 438 (89%) | 217 (91%) | |
| Mutant | 194 (15%) | 120 (21%) | 53 (11%) | 21 (8.8%) | |
| KRAS mutation | 0.88 | ||||
| Wild-type | 729 (58%) | 311 (59%) | 281 (58%) | 137 (58%) | |
| Mutant | 519 (42%) | 214 (41%) | 204 (42%) | 101 (42%) | |
| PIK3CA mutation | 0.44 | ||||
| Wild-type | 1,007 (84%) | 443 (84%) | 373 (83%) | 191 (86%) | |
| Mutant | 195 (16%) | 86 (16%) | 79 (17%) | 30 (14%) | |
CIMP, CpG island methylator phenotype; LINE-1, long interspersed nucleotide element-1; MSI, microsatellite instability; MSS, microsatellite stable; SD, standard deviation.
Percentage (%) indicates the proportion of cases with a specific clinical, pathological, or tumor molecular feature in colorectal cancer cases with each tumor LINE-1 methylation level. There were cases that had missing values for any of the characteristics except for age, sex, year of diagnosis, and tumor location.
To assess associations between categorical data, the chi-square test was performed. To compare mean age, an analysis of variance was performed. We adjusted two-sided α level to 0.003 (= 0.05/15) by simple Bonferroni correction for multiple hypothesis testing.
TNM stage was based on the classification of the American Joint Committee on Cancer staging system.
Clinical, pathological, and tumor molecular features according to tumor LINE-1 methylation level in proximal colon, distal colon, and rectal cancers are summarized in Supplementary Table S1.
Tumor LINE-1 hypomethylation and patient survival according to colorectal cancer location
We examined the relationship between tumor LINE-1 methylation level and patient survival according to colorectal cancer location (Table 2). In the 1,317 colorectal cancer cases, there were 717 deaths, including 382 colorectal cancer-specific deaths, during a median patient follow-up of 12.0 years (interquartile range: 8.0 to 16.6) among censored cases.
Table 2. Tumor LINE-1 hypomethylation and patient survival according to colorectal cancer location.
| No. of cases | Colorectal cancer-specific mortality | Overall mortality | |||||
|---|---|---|---|---|---|---|---|
| No. of events | Univariable HR (95% CI) | Multivariable HR (95% CI)a | No. of events | Univariable HR (95% CI) | Multivariable HR (95% CI)a | ||
| Proximal colon cancer | |||||||
| LINE-1 hypomethylation (20% decrease as a unit) | 621 | 176 | 2.37 (1.74-3.23) | 1.66 (1.21- 2.28) | 344 | 1.53 (1.21-1.93) | 1.40 (1.10-1.78) |
| Pb | < 0.0001 | 0.002 | 0.0004 | 0.007 | |||
| Distal colon cancer | |||||||
| LINE-1 hypomethylation (20% decrease as a unit) | 409 | 111 | 1.30 (0.91-1.86) | 1.18 (0.81- 1.72) | 211 | 1.04 (0.79-1.36) | 1.14 (0.86-1.50) |
| Pb | 0.16 | 0.40 | 0.78 | 0.36 | |||
| Rectal cancer | |||||||
| LINE-1 hypomethylation (20% decrease as a unit) | 287 | 95 | 0.92 (0.60-1.41) | 0.87 (0.57- 1.34) | 162 | 0.79 (0.57-1.09) | 0.72 (0.52-1.00) |
| Pb | 0.69 | 0.53 | 0.15 | 0.052 | |||
| Pinteractionc | 0.0002 | 0.011 | 0.0007 | 0.002 | |||
CI, confidence interval; HR, hazard ratio; LINE-1, long interspersed nucleotide element-1.
The multivariable Cox regression model initially included sex, age, year of diagnosis, family history of colorectal cancer in parent or sibling, disease stage, tumor differentiation, microsatellite instability, CpG island methylator phenotype, and KRAS, BRAF, and PIK3CA mutations. A backward elimination with a threshold of P = 0.05 was used to select variables in the final models.
P value was calculated by the Wald test (two-sided).
For our primary hypothesis testing, we found a statistically significant interaction between LINE-1 methylation level and tumor location in colorectal cancer-specific mortality analysis (Pinteraction = 0.011; Table 2). Hazard ratio (HR) of colorectal cancer-specific mortality for 20% decrease in tumor LINE-1 methylation level was higher in proximal colon cancers (multivariable HR, 1.66; 95% confidence interval [CI], 1.21 to 2.28) than in distal colon cancers (multivariable HR, 1.18; 95% CI, 0.81 to 1.72) or rectal cancers (multivariable HR, 0.87; 95% CI, 0.57 to 1.34) (Table 2). In the Kaplan-Meier analysis and the log-rank test (Figure 1), tumor LINE-1 hypomethylation was associated with higher colorectal cancer-specific mortality in proximal colon cancer (P < 0.0001 for trend), but not in distal colon cancer (P = 0.12 for trend) or rectal cancer (P = 0.60 for trend).
Figure 1. Kaplan-Meier curves for colorectal cancer-specific survival according to tumor LINE-1 methylation level in proximal colon cancer (A), distal colon cancer (B), and rectal cancer (C).
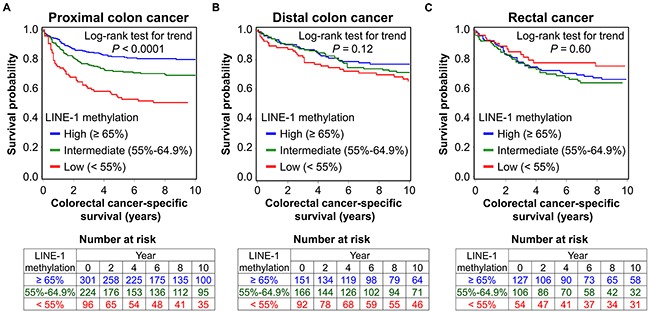
P value was calculated by the log-rank test for trend (two-sided). The tables (bottom) show the number of patients who remained alive and at risk of death at each time point after the diagnosis of colorectal cancer.
In the secondary analysis, a similar interactive association between tumor LINE-1 methylation level and colorectal cancer location was observed in overall mortality analysis (Pinteraction = 0.002; Table 2). Tumor LINE-1 hypomethylation was associated with higher overall mortality in proximal colon cancer (for 20% decrease in tumor LINE-1 methylation level: multivariable HR, 1.40; 95% CI, 1.10 to 1.78), whereas tumor LINE-1 methylation level were not significantly associated with overall mortality in distal colon cancer (for 20% decrease in tumor LINE-1 methylation level: multivariable HR, 1.14; 95% CI, 0.86 to 1.50) or rectal cancer (for 20% decrease in tumor LINE-1 methylation level: multivariable HR, 0.72; 95% CI, 0.52 to 1.00).
Tumor LINE-1 hypomethylation and patient survival in strata of colorectal tumor location and MSI status
Given our previous study showing the interactive association between tumor LINE-1 methylation level and MSI status in relation to colorectal cancer mortality [25], we conducted an exploratory analysis to examine the relationship between tumor LINE-1 hypomethylation and patient survival in strata of colorectal cancer location and MSI status (Table 3). In proximal colon cancers, tumor LINE-1 hypomethylation appeared to be associated with higher colorectal cancer-specific mortality not only in MSI-high cancers (for 20% decrease in tumor LINE-1 methylation level: multivariable HR, 6.14; 95% CI, 2.27 to 16.6) but also in MSI-low/MSS cancers (for 20% decrease in tumor LINE-1 methylation level: multivariable HR, 1.44; 95% CI, 1.04 to 2.01).
Table 3. Tumor LINE-1 hypomethylation and patient survival according to colorectal cancer location and MSI status.
| No. of cases | Colorectal cancer-specific mortality | Overall mortality | |||||
|---|---|---|---|---|---|---|---|
| No. of events | Univariable HR (95% CI) | Multivariable HR (95% CI)a | No. of events | Univariable HR (95% CI) | Multivariable HR (95% CI)a | ||
| MSI-low/MSS proximal colon cancer | |||||||
| LINE-1 hypomethylation (20% decrease as a unit) | 440 | 153 | 1.77 (1.26-2.50) | 1.44 (1.04-2.01) | 259 | 1.41 (1.07-1.84) | 1.27 (0.97-1.66) |
| Pb | 0.001 | 0.030 | 0.014 | 0.08 | |||
| MSI-high proximal colon cancer | |||||||
| LINE-1 hypomethylation (20% decrease as a unit) | 181 | 23 | 4.51 (1.78-11.4) | 6.14 (2.27-16.6) | 85 | 1.62 (0.96-2.74) | 2.30 (1.30-4.06) |
| Pb | 0.002 | 0.0003 | 0.07 | 0.004 | |||
| MSI-low/MSS distal colon cancer | |||||||
| LINE-1 hypomethylation (20% decrease as a unit) | 388 | 108 | 1.21 (0.84-1.76) | 1.13 (0.76-1.66) | 203 | 0.97 (0.73-1.28) | 1.06 (0.80-1.41) |
| Pb | 0.31 | 0.55 | 0.82 | 0.69 | |||
| MSI-high distal colon cancer | |||||||
| LINE-1 hypomethylation (20% decrease as a unit) | 21 | 3 | 3.05 (0.74-12.6) | 2.76 (0.75-10.1) | 8 | 2.38 (0.89-6.36) | 2.00 (0.87-4.60) |
| Pb | 0.12 | 0.13 | 0.08 | 0.10 | |||
| Rectal cancer | |||||||
| LINE-1 hypomethylation (20% decrease as a unit) | 287 | 95 | 0.92 (0.60-1.40) | 0.87 (0.57-1.34) | 162 | 0.79 (0.57-1.09) | 0.72 (0.52-1.00) |
| Pb | 0.69 | 0.52 | 0.15 | 0.052 | |||
CI, confidence interval; HR, hazard ratio; LINE-1, long interspersed nucleotide element-1; MSI, microsatellite instability; MSS, microsatellite stable.
The multivariable Cox regression model initially included sex, age, year of diagnosis, family history of colorectal cancer in parent or sibling, disease stage, tumor differentiation, CpG island methylator phenotype, and KRAS, BRAF, and PIK3CA mutations. A backward elimination with a threshold of P = 0.05 was used to select variables in the final models.
P value was calculated by the Wald test (two-sided).
DISCUSSION
We conducted this study to test the hypothesis that the prognostic association of tumor LINE-1 hypomethylation might differ by colorectal cancer location. Utilizing the database of 1,317 colorectal cancer cases in the two U.S.-nationwide prospective cohort studies, we found a statistically significant interactive association of tumor LINE-1 methylation level and tumor location in colorectal cancer mortality analysis. The adverse prognostic association of tumor LINE-1 hypomethylation was stronger in proximal colon cancers than in distal colorectal cancers.
Colorectal cancers are a heterogeneous group of diseases that result from the accumulation of differing sets of genomic and epigenomic alterations, and tumor-host interactions and, hence, research on tumor biomarkers is important for clinical medicine and public health [26–32]. Few studies have considered colorectal cancer location in evaluating clinical outcome by tumor LINE-1 methylation level. In 76 patients with stage III proximal colon cancer, a lower tumor LINE-1 methylation level is associated with poor disease-free survival [23]. In 94 patients with stage I and II rectal cancer, a lower tumor LINE-1 methylation level is associated with poor recurrence-free and overall survival [33]. However, these previous studies were limited by small sample sizes, and did not examine the interactive association of tumor LINE-1 methylation level and tumor location with clinical outcome. By utilizing a large collection (n = 1,317) of colorectal cancer cases, our population-based data support the interactive association of tumor LINE-1 methylation level and tumor location in colorectal cancer mortality analysis.
Exact mechanisms by which colorectal tumors that exhibit LINE-1 hypomethylation have been associated with aggressive tumor behavior remain uncertain. It has been known that colorectal cancers develop through the accumulation of genetic and epigenetic alterations, influenced by microbial and other environmental exposures and host responses to the exposures [34–39]. Our previous study has shown that the frequencies of key molecular features such as MSI, CIMP-high, BRAF and PIK3CA mutations change gradually along the length of the colorectum [7], suggesting effects of gut contents and microbiota on colorectal tumorigenesis [40]. It is conceivable that tumor-host interactions in the tumor microenvironment including immune response and inflammation might influence the progression of colorectal tumors that exhibit LINE-1 hypomethylation. An increasing body of evidence suggests that microbiota can influence host immunity, and that microbiota, luminal contents, and colonic mucosal immunity may change from the proximal to distal colorectal segments [5, 6, 12, 41–44], future investigations are necessary to examine effects of microbiota on colorectal tumor progression.
Studies have shown that tumor LINE-1 hypomethylation in colorectal cancer is inversely associated with MSI-high, and that the prognostic association of tumor LINE-1 hypomethylation in colorectal cancer is stronger in MSI-high colorectal cancers than in MSS colorectal cancers [24, 25, 45–47], suggesting a complex biological interaction between MSI and LINE-1 hypomethylation in colorectal tumor progression. MSI-high colorectal cancers have been characterized by numerous somatic mutations [48], which might interact with genomic DNA methylation. Future studies are needed to clarify the underlying mechanisms of the association between tumor LINE-1 methylation and MSI status in colorectal cancer progression.
There are biological and clinical differences between colon and rectal cancers including treatment approaches and metastatic pattern [49]. Multiple studies have demonstrated that proportions of colorectal cancers with specific molecular features such as MSI, high-level CIMP, and BRAF and PIK3CA mutations gradually increase along the bowel subsites from rectum to ascending colon [7–9]. Hence, we included both colon and rectal carcinoma cases in the current study. This continuum model is a more advanced model than the simple colon-vs-rectum dichotomy model, because a continuous difference (along the colorectum) will surely lead to a difference in the dichotomy model. We have examined proximal colon vs. distal colon vs. rectum, considering statistical power. Future studies with larger sample size are needed to investigate the prognostic significance of tumor molecular features in the detailed colorectal cancer subsites.
Previous studies have shown that tumor LINE-1 hypomethylation in colorectal cancer is associated with worse clinical outcome, and that LINE-1 hypomethylated colorectal cancers are associated with young age of onset and a family history of colorectal cancer [21, 47, 50]. These findings suggest that LINE-1 methylation level may serve as a potential tumor biomarker for prognosis as well as for familial cancer risk assessment. In addition, our current study found that the adverse prognostic association of tumor LINE-1 hypomethylation was stronger in proximal colon cancers than in distal colorectal cancers. Our current data may help us further stratify patients with colorectal cancer into more individualized prognostic groups based on tumor location, and can guide future mechanistic studies.
We acknowledge limitations of our current study. First, data on cancer recurrence were limited in the two cohorts and were not examined. However, colorectal cancer-specific mortality is a reasonable cancer-specific outcome in the current study, which utilized the population-based data of long-term patient follow-up, since median survival for recurrent (local or metastatic) colorectal cancer was approximately 10 to 20 months during much of the time period of this study [51]. Second, the information on cancer treatment including chemotherapy use and regimens was also limited. However, distributions of chemotherapy use and its regimens would unlikely substantially differ according to tumor LINE-1 methylation level, because these data were not generally utilized for treatment decisions. The comprehensive genome-scale DNA methylation analysis in colorectal cancer by Hinoue et al. [52] has suggested distinct DNA methylation subgroups of colorectal cancer. Although we assessed well-established LINE-1 methylation as a surrogate of global DNA methylation status, this method might not capture methylation status in other CpG sites. Despite this limitation, LINE-1 hypomethylation was significantly associated with mortality in proximal colon cancer patients. This finding suggests the importance and clinical usefulness of LINE-1 methylation status, while more comprehensive methylation profile is warranted to gain new insights into roles of DNA methylation in carcinogenesis.
Strengths of this study include the use of the molecular pathological epidemiology [53–56] database of 1,317 colon and rectal carcinoma cases within the two U.S.-nationwide prospective cohort studies. Importantly, our colorectal cancer sample represented a group of patients in a large number of hospitals in diverse settings across the U.S., which increases the generalizability of our findings. Additionally, our database integrates clinical, pathologic, and key tumor molecular features of colorectal cancer. Finally, the sample size and the comprehensiveness of this population-based colorectal cancer database enabled us to achieve a sufficient statistical power to assess the interactive association of tumor LINE-1 methylation level and colorectal cancer location with colorectal cancer mortality, controlling for potential confounders.
In conclusion, we have shown that the adverse prognostic association of tumor LINE-1 hypomethylation is stronger in proximal colon cancers than in distal colorectal cancers. Our data suggest the interactive effect of tumor LINE-1 methylation level and tumor location on clinical outcome.
MATERIALS AND METHODS
Study population
We utilized the database of colon and rectal carcinoma cases in two U.S.-nationwide prospective cohort studies, the NHS (121,701 women enrolled in 1976) and the HPFS (51,529 men enrolled in 1986) [57, 58]. Every 2 years, we sent follow-up questionnaires to collect information on lifestyle factors, and asked whether they had received diagnoses of major disease including cancers. The National Death Index was used to identify unreported fatal colorectal cancer cases. Study physicians reviewed medical records for colorectal cancer cases, and assigned the cause of death for all deceased cases. We collected formalin-fixed paraffin-embedded tissue blocks from hospitals across the United States where participants with colorectal cancer had undergone tumor resection. A single pathologist (S.O.), who was unaware of other data, reviewed hematoxylin and eosin-stained tissue sections of all colorectal carcinoma cases, and recorded pathological features. Tumor differentiation was classified as well to moderate vs. poor (>50% vs. ≤50% glandular area). We analyzed available data on tumor LINE-1 methylation level, tumor location, and patient survival among 1,317 colorectal cancer cases diagnosed up to 2008. Patients were followed until death or January 1, 2012, whichever came first. The procedures and protocols of this study were approved by the institutional review boards for the Harvard T.H. Chan School of Public Health and the Brigham and Women's Hospital (Boston, MA, USA).
Assessment of tumor location
Study physicians, unaware of other data, reviewed medical and pathological reports, and recorded tumor location (cecum, ascending colon, hepatic flexure, transverse colon, splenic flexure, descending colon, sigmoid colon, rectosigmoid, and rectum). Proximal colon consists of cecum, ascending colon, hepatic flexure, and transverse colon, whereas distal colon consists of splenic flexure, descending colon, and sigmoid colon.
Analysis of LINE-1 methylation level
DNA was extracted from archival colorectal cancer tissue blocks. We performed bisulfite treatment of DNA, polymerase chain reaction (PCR), and a pyrosequencing assay to measure tumor LINE-1 methylation levels after assay validation [59]. We primarily used tumor LINE-1 methylation level as a continuous variable (20% decrease as a unit) in survival analyses. When we displayed tumor LINE-1 methylation level in relation to clinical, pathological, and tumor molecular features (Table 1), we categorized tumor LINE-1 methylation levels into low (<55% methylation) vs. intermediate (55-64.9% methylation) vs. high (≥65% methylation) to keep consistency with our previous studies [25, 50].
Analyses of MSI, CIMP, and KRAS, BRAF, and PIK3CA mutations
MSI status was analyzed with use of 10 microsatellite markers (D2S123, D5S346, D17S250, BAT25, BAT26, BAT40, D18S55, D18S56, D18S67, and D18S487) as previously described [60]. We defined MSI-high as the presence of instability in ≥30% of the markers, and MSI-low/microsatellite stable (MSS) as instability in <30% of the markers. Methylation analysis of eight promoter CpG islands specific for CIMP (CACNA1G, CDKN2A, CRABP1, IGF2, MLH1, NEUROG1, RUNX3, and SOCS1) was performed as previously described [61, 62]. PCR and pyrosequencing assay targeted for KRAS (codons 12, 13, 61, and 146) [63, 64], BRAF (codon 600) [60], and PIK3CA (exons 9 and 20) [65, 66] were performed as previously described.
Statistical analysis
All statistical analyses were conducted using SAS (version 9.3, SAS Institute, Cary, NC) and all P values were two-sided. Our primary hypothesis testing was a statistical interaction of tumor LINE-1 methylation level and colorectal cancer location in colorectal cancer-specific mortality analysis. Overall mortality was a secondary outcome. The statistical interaction was assessed by the Wald test on the cross-product term of tumor LINE-1 methylation level as a continuous variable and colorectal cancer location as an ordinal variable (proximal colon [1], distal colon [2], and rectum [3]) in a Cox proportional hazards regression model. A two-sided α level was set at 0.05 for our primary hypothesis testing. For a secondary or exploratory analysis, we adjusted the two-sided α level by simple Bonferroni correction for multiple hypothesis testing, in addition to the use of the two-sided α level of 0.05 for our primary hypothesis testing.
For analyses of colorectal cancer-specific mortality, deaths as a result of other causes were censored. The Kaplan-Meier method was used to describe the distribution of colorectal cancer-specific survival, and the log-rank test for trend was performed to assess a linear trend in survival probability across the ordinal categories (high [1] vs. intermediate [2] vs. low [3]) of tumor LINE-1 methylation level. To control for confounders, we used multivariable Cox proportional hazards regression models. In addition to the tumor LINE-1 hypomethylation variable (continuous; 20% decrease as a unit), the multivariable model initially included sex, age at diagnosis (continuous), year of diagnosis (continuous), family history of colorectal cancer in a first-degree relative (present vs. absent), disease stage (I/II vs. III/IV), tumor differentiation (well to moderate vs. poor), MSI (high vs. MSI-low/MSS), CIMP (high vs. low/negative), KRAS (mutant vs. wild-type), BRAF (mutant vs. wild-type), and PIK3CA (mutant vs. wild-type). A single analysis model could assess the prognostic association of tumor LINE-1 hypomethylation in each stratum of colorectal cancer location, using a reparameterization of the interaction term (of tumor LINE-1 hypomethylation and colorectal cancer location) as previously described [25, 57]. A backward elimination was carried out with P = 0.05 as a threshold, to select variables for the final model. For cases with missing information in any of the categorical covariates (family history of colorectal cancer in a first-degree relative [0.4%], disease stage [8.7%], tumor differentiation [0.6%], MSI [2.8%], CIMP [4.9%], KRAS [5.2%], BRAF [2.1%], and PIK3CA [8.7%]), we included these cases in the majority category of a given covariate to minimize the number of variables in the multivariable Cox models. We confirmed that excluding the cases with missing information in any of the covariates did not substantially alter results (data not shown). The proportionality of hazards assumption was assessed by a time-varying covariate, using an interaction term of colorectal cancer-specific survival and tumor LINE-1 methylation level (P = 0.97).
All univariable analyses for associations of tumor LINE-1 methylation level with clinical, pathological, and tumor molecular features in colorectal cancer, and these associations according to proximal colon, distal colon, and rectal cancer were considered as secondary exploratory analyses. Given the 13 covariates in overall colorectal cancer and the 12 covariates in each of the three cancer locations, we adjusted the two-sided α level to 0.003 (= 0.05/15) by simple Bonferroni correction for multiple hypothesis testing. To assess associations between categorical variables, the chi-square test was performed. To compare mean age, an analysis of variance assuming equal variances was performed.
Abbreviations
CI, confidence interval; CIMP, CpG island methylator phenotype; HPFS, Health Professionals Follow-up Study; HR, hazard ratio; LINE-1, long interspersed nucleotide element-1; MSI, microsatellite instability; MSS, microsatellite stable; NHS, Nurses’ Health Study; PCR, polymerase chain reaction; SD, standard deviation.
Use of standardized official symbols
We use HUGO (Human Genome Organisation)-approved official symbols for genes and gene products, including BRAF, CACNA1G, CDKN2A, CRABP1, IGF2, KRAS, MLH1, NEUROG1, PIK3CA, RUNX3, and SOCS1; all of which are described at www.genenames.org. Gene names are italicized, and gene product names are non-italicized.
SUPPLEMENTARY TABLE
Acknowledgments
We would like to thank the participants and staff of the Nurses’ Health Study and the Health Professionals Follow-up Study for their valuable contributions as well as the following state cancer registries for their help: AL, AZ, AR, CA, CO, CT, DE, FL, GA, ID, IL, IN, IA, KY, LA, ME, MD, MA, MI, NE, NH, NJ, NY, NC, ND, OH, OK, OR, PA, RI, SC, TN, TX, VA, WA, WY. The authors assume full responsibility for analyses and interpretation of these data.
Abbreviations
- CI
confidence interval
- CIMP
CpG island methylator phenotype
- HPFS
Health Professionals Follow-up Study
- HR
hazard ratio
- LINE-1
long interspersed nucleotide element-1
- MSI
microsatellite instability
- MSS
microsatellite stable
- NHS
Nurses’ Health Study
- PCR
polymerase chain reaction
- SD
standard deviation.
Footnotes
CONFLICTS OF INTEREST
Andrew T. Chan previously served as a consultant for Bayer Healthcare, Millennium Pharmaceuticals, Pozen Inc, and Pfizer Inc. This study was not funded by Bayer Healthcare, Millennium Pharmaceuticals, Pozen Inc, or Pfizer Inc. No other conflict of interest exists. The other authors declare that they have no conflicts of interest.
GRANT SUPPORT
This work was supported by U.S. National Institutes of Health (NIH) grants [P01 CA87969 to M.J. Stampfer; UM1 CA186107 to M.J. Stampfer; P01 CA55075 to W.C. Willett; UM1 CA167552 to W.C. Willett; P50 CA127003 to C.S.F.; R01 CA137178 to A.T.C.; R01 CA151993 to S.O.; R35 CA197735 to S.O.; K07 CA190673 to R.N.; R03 CA176717 to X.Z.; and K07 CA188126 to X.Z.]; and by grants from the Project P Fund, The Friends of the Dana-Farber Cancer Institute, Bennett Family Fund, and the Entertainment Industry Foundation through National Colorectal Cancer Research Alliance. K.M. is supported by a grant from Program for Advancing Strategic International Networks to Accelerate the Circulation of Talented Researchers from Japan Society for the Promotion of Science. The content is solely the responsibility of the authors and does not necessarily represent the official views of NIH. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Use of standardized official symbols
We use HUGO (Human Genome Organisation)-approved official symbols for genes and gene products, including BRAF, CACNA1G, CDKN2A, CRABP1, IGF2, KRAS, MLH1, NEUROG1, PIK3CA, RUNX3, and SOCS1; all of which are described at www.genenames.org. Gene names are italicized, and gene product names are non-italicized.
REFERENCES
- 1.Kuipers EJ, Grady WM, Lieberman D, Seufferlein T, Sung JJ, Boelens PG, van de Velde CJH, Watanabe T. Colorectal cancer. Nature Reviews Disease Primers. 2015;1:15065. doi: 10.1038/nrdp.2015.65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Sears CL, Garrett WS. Microbes, microbiota, and colon cancer. Cell Host Microbe. 2014;15:317–328. doi: 10.1016/j.chom.2014.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Keku TO, Dulal S, Deveaux A, Jovov B, Han X. The gastrointestinal microbiota and colorectal cancer. Am J Physiol Gastrointest Liver Physiol. 2015;308:G351–363. doi: 10.1152/ajpgi.00360.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Tahara T, Yamamoto E, Suzuki H, Maruyama R, Chung W, Garriga J, Jelinek J, Yamano HO, Sugai T, An B, Shureiqi I, Toyota M, Kondo Y, et al. Fusobacterium in colonic flora and molecular features of colorectal carcinoma. Cancer Res. 2014;74:1311–1318. doi: 10.1158/0008-5472.CAN-13-1865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Dejea CM, Wick EC, Hechenbleikner EM, White JR, Mark Welch JL, Rossetti BJ, Peterson SN, Snesrud EC, Borisy GG, Lazarev M, Stein E, Vadivelu J, Roslani AC, et al. Microbiota organization is a distinct feature of proximal colorectal cancers. Proc Natl Acad Sci U S A. 2014;111:18321–18326. doi: 10.1073/pnas.1406199111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Mima K, Sukawa Y, Nishihara R, Qian ZR, Yamauchi M, Inamura K, Kim SA, Masuda A, Nowak JA, Nosho K, Kostic AD, Giannakis M, Watanabe H, et al. Fusobacterium nucleatum and T Cells in Colorectal Carcinoma. JAMA Oncol. 2015;1:653–661. doi: 10.1001/jamaoncol.2015.1377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Yamauchi M, Morikawa T, Kuchiba A, Imamura Y, Qian ZR, Nishihara R, Liao X, Waldron L, Hoshida Y, Huttenhower C, Chan AT, Giovannucci E, Fuchs C, et al. Assessment of colorectal cancer molecular features along bowel subsites challenges the conception of distinct dichotomy of proximal versus distal colorectum. Gut. 2012;61:847–854. doi: 10.1136/gutjnl-2011-300865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Phipps AI, Limburg PJ, Baron JA, Burnett-Hartman AN, Weisenberger DJ, Laird PW, Sinicrope FA, Rosty C, Buchanan DD, Potter JD, Newcomb PA. Association between molecular subtypes of colorectal cancer and patient survival. Gastroenterology. 2015;148:77–87. doi: 10.1053/j.gastro.2014.09.038. e72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Rosty C, Young JP, Walsh MD, Clendenning M, Sanderson K, Walters RJ, Parry S, Jenkins MA, Win AK, Southey MC, Hopper JL, Giles GG, Williamson EJ, et al. PIK3CA activating mutation in colorectal carcinoma: associations with molecular features and survival. PLoS One. 2013;8:e65479. doi: 10.1371/journal.pone.0065479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ogino S, Nosho K, Irahara N, Meyerhardt JA, Baba Y, Shima K, Glickman JN, Ferrone CR, Mino-Kenudson M, Tanaka N, Dranoff G, Giovannucci EL, Fuchs CS. Lymphocytic reaction to colorectal cancer is associated with longer survival, independent of lymph node count, microsatellite instability, and CpG island methylator phenotype. Clin Cancer Res. 2009;15:6412–6420. doi: 10.1158/1078-0432.CCR-09-1438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ghazi S, Lindforss U, Lindberg G, Berg E, Lindblom A, Papadogiannakis N. Analysis of colorectal cancer morphology in relation to sex, age, location, and family history. J Gastroenterol. 2012;47:619–634. doi: 10.1007/s00535-011-0520-9. [DOI] [PubMed] [Google Scholar]
- 12.Kirby JA, Bone M, Robertson H, Hudson M, Jones DE. The number of intraepithelial T cells decreases from ascending colon to rectum. J Clin Pathol. 2003;56:158. doi: 10.1136/jcp.56.2.158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Yang AS, Estecio MR, Doshi K, Kondo Y, Tajara EH, Issa JP. A simple method for estimating global DNA methylation using bisulfite PCR of repetitive DNA elements. Nucleic Acids Res. 2004;32:e38. doi: 10.1093/nar/gnh032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Weisenberger DJ, Campan M, Long TI, Kim M, Woods C, Fiala E, Ehrlich M, Laird PW. Analysis of repetitive element DNA methylation by MethyLight. Nucleic Acids Res. 2005;33:6823–6836. doi: 10.1093/nar/gki987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gaudet F, Hodgson JG, Eden A, Jackson-Grusby L, Dausman J, Gray JW, Leonhardt H, Jaenisch R. Induction of tumors in mice by genomic hypomethylation. Science. 2003;300:489–492. doi: 10.1126/science.1083558. [DOI] [PubMed] [Google Scholar]
- 16.Holm TM, Jackson-Grusby L, Brambrink T, Yamada Y, Rideout WM, 3rd, Jaenisch R. Global loss of imprinting leads to widespread tumorigenesis in adult mice. Cancer Cell. 2005;8:275–285. doi: 10.1016/j.ccr.2005.09.007. [DOI] [PubMed] [Google Scholar]
- 17.Suzuki K, Suzuki I, Leodolter A, Alonso S, Horiuchi S, Yamashita K, Perucho M. Global DNA demethylation in gastrointestinal cancer is age dependent and precedes genomic damage. Cancer Cell. 2006;9:199–207. doi: 10.1016/j.ccr.2006.02.016. [DOI] [PubMed] [Google Scholar]
- 18.Rodriguez J, Frigola J, Vendrell E, Risques RA, Fraga MF, Morales C, Moreno V, Esteller M, Capella G, Ribas M, Peinado MA. Chromosomal instability correlates with genome-wide DNA demethylation in human primary colorectal cancers. Cancer Res. 2006;66:8462–9468. doi: 10.1158/0008-5472.CAN-06-0293. [DOI] [PubMed] [Google Scholar]
- 19.Rajagopalan H, Nowak MA, Vogelstein B, Lengauer C. The significance of unstable chromosomes in colorectal cancer. Nat Rev Cancer. 2003;3:695–701. doi: 10.1038/nrc1165. [DOI] [PubMed] [Google Scholar]
- 20.Mlecnik B, Bindea G, Angell HK, Sasso MS, Obenauf AC, Fredriksen T, Lafontaine L, Bilocq AM, Kirilovsky A, Tosolini M, Waldner M, Berger A, Fridman WH, et al. Functional network pipeline reveals genetic determinants associated with in situ lymphocyte proliferation and survival of cancer patients. Sci Transl Med. 2014;6:228ra237. doi: 10.1126/scitranslmed.3007240. [DOI] [PubMed] [Google Scholar]
- 21.Ogino S, Nosho K, Kirkner GJ, Kawasaki T, Chan AT, Schernhammer ES, Giovannucci EL, Fuchs CS. A cohort study of tumoral LINE-1 hypomethylation and prognosis in colon cancer. J Natl Cancer Inst. 2008;100:1734–1738. doi: 10.1093/jnci/djn359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Nosho K, Baba Y, Tanaka N, Shima K, Hayashi M, Meyerhardt JA, Giovannucci E, Dranoff G, Fuchs CS, Ogino S. Tumour-infiltrating T-cell subsets, molecular changes in colorectal cancer, and prognosis: cohort study and literature review. J Pathol. 2010;222:350–366. doi: 10.1002/path.2774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ahn JB, Chung WB, Maeda O, Shin SJ, Kim HS, Chung HC, Kim NK, Issa JP. DNA methylation predicts recurrence from resected stage III proximal colon cancer. Cancer. 2011;117:1847–1854. doi: 10.1002/cncr.25737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Rhee YY, Kim MJ, Bae JM, Koh JM, Cho NY, Juhnn YS, Kim D, Kang GH. Clinical outcomes of patients with microsatellite-unstable colorectal carcinomas depend on L1 methylation level. Ann Surg Oncol. 2012;19:3441–3448. doi: 10.1245/s10434-012-2410-7. [DOI] [PubMed] [Google Scholar]
- 25.Inamura K, Yamauchi M, Nishihara R, Lochhead P, Qian ZR, Kuchiba A, Kim SA, Mima K, Sukawa Y, Jung S, Zhang X, Wu K, Cho E, et al. Tumor LINE-1 methylation level and microsatellite instability in relation to colorectal cancer prognosis. J Natl Cancer Inst. 2014:106. doi: 10.1093/jnci/dju195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Bartley AN, Parikh N, Hsu CH, Roe DJ, Buckmeier JA, Corley L, Phipps RA, Gallick G, Lance P, Thompson PA, Hamilton SR. Colorectal adenoma stem-like cell populations: associations with adenoma characteristics and metachronous colorectal neoplasia. Cancer Prev Res (Phila) 2013;6:1162–1170. doi: 10.1158/1940-6207.CAPR-13-0113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Sinicrope FA, Shi Q, Smyrk TC, Thibodeau SN, Dienstmann R, Guinney J, Bot BM, Tejpar S, Delorenzi M, Goldberg RM, Mahoney M, Sargent DJ, Alberts SR. Molecular markers identify subtypes of stage III colon cancer associated with patient outcomes. Gastroenterology. 2015;148:88–99. doi: 10.1053/j.gastro.2014.09.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Campbell PT, Newton CC, Newcomb PA, Phipps AI, Ahnen DJ, Baron JA, Buchanan DD, Casey G, Cleary SP, Cotterchio M, Farris AB, Figueiredo JC, Gallinger S, et al. Association between body mass index and mortality for colorectal cancer survivors: overall and by tumor molecular phenotype. Cancer Epidemiol Biomarkers Prev. 2015;24:1229–1238. doi: 10.1158/1055-9965.EPI-15-0094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Colussi D, Brandi G, Bazzoli F, Ricciardiello L. Molecular pathways involved in colorectal cancer: implications for disease behavior and prevention. Int J Mol Sci. 2013;14:16365–16385. doi: 10.3390/ijms140816365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kocarnik JM, Shiovitz S, Phipps AI. Molecular phenotypes of colorectal cancer and potential clinical applications. Gastroenterol Rep (Oxf) 2015;3:269–276. doi: 10.1093/gastro/gov046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Bardhan K, Liu K. Epigenetics and colorectal cancer pathogenesis. Cancers (Basel) 2013;5:676–713. doi: 10.3390/cancers5020676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Suzuki H, Yamamoto E, Maruyama R, Niinuma T, Kai M. Biological significance of the CpG island methylator phenotype. Biochem Biophys Res Commun. 2014;455:35–42. doi: 10.1016/j.bbrc.2014.07.007. [DOI] [PubMed] [Google Scholar]
- 33.Benard A, van de Velde CJ, Lessard L, Putter H, Takeshima L, Kuppen PJ, Hoon DS. Epigenetic status of LINE-1 predicts clinical outcome in early-stage rectal cancer. Br J Cancer. 2013;109:3073–3083. doi: 10.1038/bjc.2013.654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Ogino S, Galon J, Fuchs CS, Dranoff G. Cancer immunology--analysis of host and tumor factors for personalized medicine. Nat Rev Clin Oncol. 2011;8:711–719. doi: 10.1038/nrclinonc.2011.122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Bishehsari F, Mahdavinia M, Vacca M, Malekzadeh R, Mariani-Costantini R. Epidemiological transition of colorectal cancer in developing countries: environmental factors, molecular pathways, and opportunities for prevention. World J Gastroenterol. 2014;20:6055–6072. doi: 10.3748/wjg.v20.i20.6055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Chia WK, Ali R, Toh HC. Aspirin as adjuvant therapy for colorectal cancer--reinterpreting paradigms. Nat Rev Clin Oncol. 2012;9:561–570. doi: 10.1038/nrclinonc.2012.137. [DOI] [PubMed] [Google Scholar]
- 37.Ng JM, Yu J. Promoter hypermethylation of tumour suppressor genes as potential biomarkers in colorectal cancer. Int J Mol Sci. 2015;16:2472–2496. doi: 10.3390/ijms16022472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Martinez-Useros J, Garcia-Foncillas J. Obesity and colorectal cancer: molecular features of adipose tissue. J Transl Med. 2016;14:21. doi: 10.1186/s12967-016-0772-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Nosho K, Sukawa Y, Adachi Y, Ito M, Mitsuhashi K, Kurihara H, Kanno S, Yamamoto I, Ishigami K, Igarashi H, Maruyama R, Imai K, Yamamoto H, et al. Association of Fusobacterium nucleatum with immunity and molecular alterations in colorectal cancer. World J Gastroenterol. 2016;22:557–566. doi: 10.3748/wjg.v22.i2.557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yamauchi M, Lochhead P, Morikawa T, Huttenhower C, Chan AT, Giovannucci E, Fuchs C, Ogino S. Colorectal cancer: a tale of two sides or a continuum? Gut. 2012;61:794–797. doi: 10.1136/gutjnl-2012-302014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kohoutova D, Smajs D, Moravkova P, Cyrany J, Moravkova M, Forstlova M, Cihak M, Rejchrt S, Bures J. Escherichia coli strains of phylogenetic group B2 and D and bacteriocin production are associated with advanced colorectal neoplasia. BMC Infect Dis. 2014;14:733. doi: 10.1186/s12879-014-0733-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Li X, LeBlanc J, Truong A, Vuthoori R, Chen SS, Lustgarten JL, Roth B, Allard J, Ippoliti A, Presley LL, Borneman J, Bigbee WL, Gopalakrishnan V, et al. A metaproteomic approach to study human-microbial ecosystems at the mucosal luminal interface. PLoS One. 2011;6:e26542. doi: 10.1371/journal.pone.0026542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Hayashi H, Takahashi R, Nishi T, Sakamoto M, Benno Y. Molecular analysis of jejunal, ileal, caecal and recto-sigmoidal human colonic microbiota using 16S rRNA gene libraries and terminal restriction fragment length polymorphism. J Med Microbiol. 2005;54:1093–1101. doi: 10.1099/jmm.0.45935-0. [DOI] [PubMed] [Google Scholar]
- 44.Mowat AM, Agace WW. Regional specialization within the intestinal immune system. Nat Rev Immunol. 2014;14:667–685. doi: 10.1038/nri3738. [DOI] [PubMed] [Google Scholar]
- 45.Estecio MR, Gharibyan V, Shen L, Ibrahim AE, Doshi K, He R, Jelinek J, Yang AS, Yan PS, Huang TH, Tajara EH, Issa JP. LINE-1 hypomethylation in cancer is highly variable and inversely correlated with microsatellite instability. PLoS One. 2007;2:e399. doi: 10.1371/journal.pone.0000399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Ogino S, Kawasaki T, Nosho K, Ohnishi M, Suemoto Y, Kirkner GJ, Fuchs CS. LINE-1 hypomethylation is inversely associated with microsatellite instability and CpG island methylator phenotype in colorectal cancer. Int J Cancer. 2008;122:2767–2773. doi: 10.1002/ijc.23470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Baba Y, Huttenhower C, Nosho K, Tanaka N, Shima K, Hazra A, Schernhammer ES, Hunter DJ, Giovannucci EL, Fuchs CS, Ogino S. Epigenomic diversity of colorectal cancer indicated by LINE-1 methylation in a database of 869 tumors. Mol Cancer. 2010;9:125. doi: 10.1186/1476-4598-9-125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330–337. doi: 10.1038/nature11252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Tamas K, Walenkamp AM, de Vries EG, van Vugt MA, Beets-Tan RG, van Etten B, de Groot DJ, Hospers GA. Rectal and colon cancer: Not just a different anatomic site. Cancer Treat Rev. 2015;41:671–679. doi: 10.1016/j.ctrv.2015.06.007. [DOI] [PubMed] [Google Scholar]
- 50.Ogino S, Nishihara R, Lochhead P, Imamura Y, Kuchiba A, Morikawa T, Yamauchi M, Liao X, Qian ZR, Sun R, Sato K, Kirkner GJ, Wang M, et al. Prospective study of family history and colorectal cancer risk by tumor LINE-1 methylation level. J Natl Cancer Inst. 2013;105:130–140. doi: 10.1093/jnci/djs482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Meyerhardt JA, Mayer RJ. Systemic therapy for colorectal cancer. N Engl J Med. 2005;352:476–487. doi: 10.1056/NEJMra040958. [DOI] [PubMed] [Google Scholar]
- 52.Hinoue T, Weisenberger DJ, Lange CP, Shen H, Byun HM, Van Den Berg D, Malik S, Pan F, Noushmehr H, van Dijk CM, Tollenaar RA, Laird PW. Genome-scale analysis of aberrant DNA methylation in colorectal cancer. Genome Res. 2012;22:271–282. doi: 10.1101/gr.117523.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Ogino S, Chan AT, Fuchs CS, Giovannucci E. Molecular pathological epidemiology of colorectal neoplasia: an emerging transdisciplinary and interdisciplinary field. Gut. 2011;60:397–411. doi: 10.1136/gut.2010.217182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Ogino S, Nishihara R, VanderWeele TJ, Wang M, Nishi A, Lochhead P, Qian ZR, Zhang X, Wu K, Nan H, Yoshida K, Milner DA, Jr, Chan AT, et al. Review Article: The Role of Molecular Pathological Epidemiology in the Study of Neoplastic and Non-neoplastic Diseases in the Era of Precision Medicine. Epidemiology. 2016;27:602–611. doi: 10.1097/EDE.0000000000000471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Nishihara R, VanderWeele TJ, Shibuya K, Mittleman MA, Wang M, Field AE, Giovannucci E, Lochhead P, Ogino S. Molecular pathological epidemiology gives clues to paradoxical findings. Eur J Epidemiol. 2015;30:1129–1135. doi: 10.1007/s10654-015-0088-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Ikram MA. Molecular pathological epidemiology: the role of epidemiology in the omics-era. Eur J Epidemiol. 2015;30:1077–1078. doi: 10.1007/s10654-015-0093-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Liao X, Lochhead P, Nishihara R, Morikawa T, Kuchiba A, Yamauchi M, Imamura Y, Qian ZR, Baba Y, Shima K, Sun R, Nosho K, Meyerhardt JA, et al. Aspirin use, tumor PIK3CA mutation, and colorectal-cancer survival. N Engl J Med. 2012;367:1596–1606. doi: 10.1056/NEJMoa1207756. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Nishihara R, Wu K, Lochhead P, Morikawa T, Liao X, Qian ZR, Inamura K, Kim SA, Kuchiba A, Yamauchi M, Imamura Y, Willett WC, Rosner BA, et al. Long-term colorectal-cancer incidence and mortality after lower endoscopy. N Engl J Med. 2013;369:1095–1105. doi: 10.1056/NEJMoa1301969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Irahara N, Nosho K, Baba Y, Shima K, Lindeman NI, Hazra A, Schernhammer ES, Hunter DJ, Fuchs CS, Ogino S. Precision of pyrosequencing assay to measure LINE-1 methylation in colon cancer, normal colonic mucosa, and peripheral blood cells. J Mol Diagn. 2010;12:177–183. doi: 10.2353/jmoldx.2010.090106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Ogino S, Nosho K, Kirkner GJ, Kawasaki T, Meyerhardt JA, Loda M, Giovannucci EL, Fuchs CS. CpG island methylator phenotype, microsatellite instability, BRAF mutation and clinical outcome in colon cancer. Gut. 2009;58:90–96. doi: 10.1136/gut.2008.155473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Ogino S, Kawasaki T, Brahmandam M, Cantor M, Kirkner GJ, Spiegelman D, Makrigiorgos GM, Weisenberger DJ, Laird PW, Loda M, Fuchs CS. Precision and performance characteristics of bisulfite conversion and real-time PCR (MethyLight) for quantitative DNA methylation analysis. J Mol Diagn. 2006;8:209–217. doi: 10.2353/jmoldx.2006.050135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Ogino S, Kawasaki T, Kirkner GJ, Kraft P, Loda M, Fuchs CS. Evaluation of markers for CpG island methylator phenotype (CIMP) in colorectal cancer by a large population-based sample. J Mol Diagn. 2007;9:305–314. doi: 10.2353/jmoldx.2007.060170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Ogino S, Kawasaki T, Brahmandam M, Yan L, Cantor M, Namgyal C, Mino-Kenudson M, Lauwers GY, Loda M, Fuchs CS. Sensitive sequencing method for KRAS mutation detection by Pyrosequencing. J Mol Diagn. 2005;7:413–421. doi: 10.1016/S1525-1578(10)60571-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Imamura Y, Lochhead P, Yamauchi M, Kuchiba A, Qian ZR, Liao X, Nishihara R, Jung S, Wu K, Nosho K, Wang YE, Peng S, Bass AJ, et al. Analyses of clinicopathological, molecular, and prognostic associations of KRAS codon 61 and codon 146 mutations in colorectal cancer: cohort study and literature review. Mol Cancer. 2014;13:135. doi: 10.1186/1476-4598-13-135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Nosho K, Kawasaki T, Ohnishi M, Suemoto Y, Kirkner GJ, Zepf D, Yan L, Longtine JA, Fuchs CS, Ogino S. PIK3CA mutation in colorectal cancer: relationship with genetic and epigenetic alterations. Neoplasia. 2008;10:534–541. doi: 10.1593/neo.08336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Liao X, Morikawa T, Lochhead P, Imamura Y, Kuchiba A, Yamauchi M, Nosho K, Qian ZR, Nishihara R, Meyerhardt JA, Fuchs CS, Ogino S. Prognostic role of PIK3CA mutation in colorectal cancer: cohort study and literature review. Clin Cancer Res. 2012;18:2257–2268. doi: 10.1158/1078-0432.CCR-11-2410. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


